Abstract
Fibroblast growth factor receptors (FGFR) are transmembrane kinase proteins with growing importance in cancer biology given the frequency of molecular alterations and vast interface with multiple other signaling pathways. Furthermore, numerous FGFR inhibitors in clinical development demonstrate the expanding therapeutic relevance of this pathway. Indeed, results from early phase clinical trials already indicate that a subset of patients with advanced tumors derive benefit from FGFR targeted therapies. FGFR gene aberrations and FGFR gene rearrangements are relatively rare in solid malignancies. The recently described FGFR3-TACC3 fusion protein has a constitutively active tyrosine kinase domain and promotes aneuploidy. We summarize the prevalence data on FGFR3-TACC3 fusions among different histological tumor types and the preliminary evidence that this rearrangement represents a targetable molecular aberration in some patients with solid tumors.
Keywords: FGFR3-TACC3 fusion, non-small cell lung cancer, phosphatidylinositol 3-Kinase (PI3K), aneuploidy, glioblastoma multiforme
INTRODUCTION
The growing knowledge base of tumor genomics has led to never seen advances in the field of medical oncology. [1] The evolving molecularly targeted treatments of late-stage melanoma, gastro-intestinal stromal tumors, and non-small-cell lung cancer (NSCLC) exemplify these advances. [2–4]
The fibroblast growth factor receptor (FGFR) family consists of four subtypes of transmembrane tyrosine kinase receptors that play an important role in cell growth, differentiation and angiogenesis via binding of up to 22 known different FGF family ligands. [5] Upon FGFR activation through dimerization of receptor monomers and transphosphorylation of kinase domain loop tyrosine residues cytoplasmatic downstream molecules contribute to carcinogenic events mediated by PI3K/AKT, STAT and RAS/MAPK pathways (Figure 1A). [5, 6]
Figure 1.
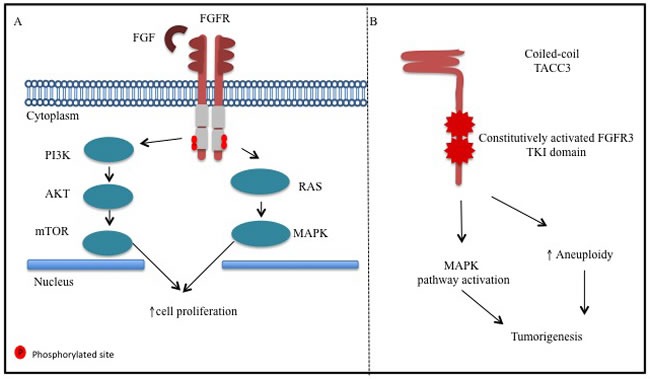
A. Fibroblast growth factor receptor (FGFR) has intra-cellular tyrosine kinase activity triggered by fibroblast growth factor (FGF) ligand. Its activation leads to FGFR transphosphorylation and activation of protein of ras oncogene (RAS)/mitogen-activated protein kinase (MAPK) and phosphatidylinositol 3-Kinase (PI3K) signaling pathways. B. Fibroblast growth factor receptor3-transforming acidic coiled-coil containing protein 3(FGFR3-TACC3) fusion protein harbors constitutively activated tyrosine kinase domain, which activates mitogen-activated protein kinase (MAPK) pathway. Also, FGFR3-TACC3 localizes to mitotic spindle poles, induces mitotic, chromosomal segregation defects and triggers aneuploidy.
Anomalous signaling through FGFR can occur through overexpression of receptors, activating mutations, gene amplification, or by FGFR-containing translocations in wide array of solid tumors as discussed below. Most importantly, these aberrations can represent important therapeutic targets in solid tumors and have supported the clinical development of several FGFR inhibitors and anti-FGFR drug conjugate (ADC) antibodies (Tables 3, 4, 5, 6, 7, 8, 9). Herein we review the current literature and describe the cancer pathobiology, the prevalence, the pre-clinical, and the clinical data supporting drug development targeting the recently described FGFR3-TACC3 fusion.
Table 3. Ongoing selected clinical trials targeting the FGFR pathway in solid tumor and hematologic malignancies.
| Drug | Mechanism of action | Phase | Study population | Clinicaltrials.gov Identification |
|---|---|---|---|---|
| ARQ 087 | FGFR1-3 TKI | 1/2 | Solid tumors with FGFR genetic alterations, including intrahepatic cholangiocarcinoma with FGFR2 gene fusion | NCT01752920 |
| AZD4547 | FGFR1-3 TKI | 2 | FGFR1 or FGFR2 amplified breast, squamous lung and stomach cancer | NCT01795768 |
| AZD4547 | FGFR1-3 TKI | 1 | In the dose expansion phase participant must have solid tumors with FGFR1 and/or FGFR2 gene amplified sqNSCLC, FGFR1 gene low & high amplified or gastric adenocarcinoma, including the lower esophagus/gastro-esophageal junction, FGFR2 gene low & high amplified | NCT00979134 |
| AZD4547 | FGFR1-3 TKI | 2 | Advanced Gastric Adenocarcinoma (Including adenocarcinoma of the lower third of the esophagus or the gastro-esophageal junction) with FGFR2 polysomy or gene amplification. | NCT01457846 |
| AZD4547 | FGFR1-3 TKI | 2a | ER+ breast cancer patients With FGFR1 polysomy or gene amplification who have progressed following treatment with prior endocrine therapy | NCT01202591 |
| AZD4547 | FGFR1-3 TKI | 2a | Refractory metastatic ER+ breast cancer | NCT01791985 |
| AZD4547 | FGFR1-3 TKI | 1 | Japanese patients with advanced solid tumors | NCT01213160 |
| BAY1163877 | FGFR1-3 TKI | 1 | In the dose expansion cohort patient must have histological or cytological sqNSCLC, lung adenocarcinoma, head and neck cancer or bladder cancer | NCT01976741 |
| BAY1187982 | Anti-FGFR2 antibody drug conjugate | 1 | Advanced solid tumors known to express FGFR2 | NCT02368951 |
| BAY1179470 | Anti FGFR2 antibody | 1 | Refractory solid tumors with at least moderate FGFR2 expression in the tumor tissue from archival samples is confirmed | NCT01881217 |
| BGJ398 | FGFR1-3 TKI | 2a | FGFR1-3 translocated, mutated, or amplified squamous cell carcinoma of the head and neck | NCT02706691 |
| BGJ398 | FGFR1-3 TKI | 2 | Solid tumor (except with a primary diagnosis of UC, cholangiocarcinoma, endometrial cancer, and GBM) or hematologic malignancies with FGFR genetic alteration | NCT02160041 |
| BGJ398 | FGFR1-3 TKI | 1 | Advanced solid tumors with FGFR1 or FGFR2 amplification or FGFR3 mutation, for which no further effective standard anticancer treatment exists or UC with FGFR3 mutations or gene fusions progressing after platinum-based chemotherapy or intolerant to platinum therapy or for whom platinum is contraindicated | NCT01004224 |
| BGJ398 combined with chemotherapy | FGFR1-3 TKI | 1b/2 | Advanced and metastatic pancreatic cancer | NCT02575508 |
Abbreviations: Fibroblast growth factor (FGF), fibroblast growth factor receptor (FGFR), not reported (NR), estrogen receptor positive (ER+), tyrosine kinase inhibitor (TKI), bacillus calmette-Guerin (BCG), small cell lung cancer (SCLC), Non-small cell lung cancer (NSCLC), squamous non-small cell lung cancer (sqNSCLC), pilot study (PS), glioblastoma multiforme (GBM), hepatocellular carcinoma (HCC), Malignant pleural mesothelioma (MPM), immunohistochemistry (IHC), urothelial carcinoma (UC).www.clinicaltrials.gov accessed on April 22nd 2016.
Table 4. Ongoing selected clinical trials targeting the FGFR pathway in solid tumor and hematologic malignancies continued.
| Drug | Mechanism of action | Phase | Study population | Clinicaltrials.gov Identification |
|---|---|---|---|---|
| BGJ398 | FGFR1-3 TKI | 2 | Advanced or metastatic cholangiocarcinoma with FGFR2 Gene Fusions or Other FGFR genetic alterations who failed or are intolerant to platinum-based Chemotherapy | NCT02150967 |
| BGJ398/BYL719 | FGFR1-3 TKI | 1b | Refractory solid tumor with PIK3CA mutations in all patients in dose escalation and expansion with or without documented genetic alterations in FGFR depending upon dose expansion cohort | NCT01928459 |
| BGJ398 | FGFR1-3 TKI | 2 | Histologically confirmed GBM and/or other glioma subtypes with FGFR1-TACC1, FGFR3-TACC3 fusion and/or activating mutation in FGFR1, 2 or 3 | NCT01975701 |
| BGJ 398 | FGFR1-3 TKI | 1 | Advanced solid tumor Having alterations of the FGF-R pathway | NCT01697605 |
| BLU-554 | FGFR4 TKI | 1 | Refractory HCC or refractory advanced solid tumor other than HCC that has evidence of aberrant FGF19/FGFR4 pathway activity | NCT02508467 |
| BT-701 | Anti FGFR-3 antibody | 2 | Refractory UC of the bladder cancer or transitional cell carcinoma arising in another location of the urinary tract, including urethra, ureter, and renal pelvis with positive FGFR3 expression on IHC | NCT02401542 |
| Debio 1347-101 | FGFR1-3 inhibitor | 1 | Advanced solid malignancies, whose tumors have an alteration of the FGFR 1, 2 or 3 genes, for whom standard treatment does not exist or is not indicated | NCT01948297 |
| Dovitinib | Multikinase inhibitor including FGFR1-3 | 2 | Refractory advanced/metastatic scirrhous gastric carcinoma | NCT01576380 |
| Dovitinib | Multikinase inhibitor including FGFR1-3 | 2 | FGFR1 amplified and non-amplified metastatic HER2 negative breast cancer | NCT00958971 |
| Dovitinib | Multikinase inhibitor including FGFR1-3 | 2 | BCG refractory UC patients with tumor fibroblast growth factor receptor 3(FGFR3) mutations or over-expression | NCT01732107 |
Abbreviations: Fibroblast growth factor (FGF), fibroblast growth factor receptor (FGFR), not reported (NR), estrogen receptor positive (ER+), tyrosine kinase inhibitor (TKI), bacillus calmette-Guerin (BCG), small cell lung cancer (SCLC), Non-small cell lung cancer (NSCLC), squamous non-small cell lung cancer (sqNSCLC), pilot study (PS), glioblastoma multiforme (GBM), hepatocellular carcinoma (HCC), Malignant pleural mesothelioma (MPM), immunohistochemistry (IHC), urothelial carcinoma (UC). www.clinicaltrials.govaccessed on April 22nd 2016.
Table 5. Ongoing selected clinical trials targeting the FGFR pathway in solid tumor and hematologic malignancies continued.
| Drug | Mechanism of action | Phase | Study population | Clinicaltrials.gov Identification |
|---|---|---|---|---|
| Dovitinib | Multikinase inhibitor including FGFR1-3 | 2 | Either FGFR2 mutated or wild-type advanced and/or metastatic endometrial cancer | NCT01379534 |
| Dovitinib | Multikinase inhibitor including FGFR1-3 | 2 | Refractory solid tumors with mutations or translocations of FGFR, PDGFR, VEGF, cKIT, FLT3, CSFR1, Trk and RET | NCT01831726 |
| Dovitinib | Multikinase inhibitor including FGFR1-3 | 2 | Advanced urothelial cancer with mutated or wild FGFR3 mutated | NCT00790426 |
| Divotinib | Multikinase inhibitor including FGFR1-3 | 2 | Metastatic or unresectable gastric cancer harboring FGFR2 amplification after failure of first or second sine chemotherapy | NCT01719549 |
| Dovitinib | Multikinase inhibitor including FGFR1-3 | NR | Refractory sqNSCLC with FGFR amplification (FISH > 5 copies of genes) | NCT01861197 |
| Dovitinib | Multikinase inhibitor including FGFR1-3 | 2 | Refractory progressive NSCLC and colorectal cancer status post antiangiogenic treatment | NCT01676714 |
| Dovitinib | Multikinase inhibitor including FGFR1-3 | PS | Refractory renal cell carcinoma | NCT01791387 |
Abbreviations: Fibroblast growth factor (FGF), fibroblast growth factor receptor (FGFR), not reported (NR), estrogen receptor positive (ER+), tyrosine kinase inhibitor (TKI), bacillus calmette-Guerin (BCG), small cell lung cancer (SCLC), Non-small cell lung cancer (NSCLC), squamous non-small cell lung cancer (sqNSCLC), pilot study (PS), glioblastoma multiforme (GBM), hepatocellular carcinoma (HCC), Malignant pleural mesothelioma (MPM), immunohistochemistry (IHC), urothelial carcinoma (UC).www.clinicaltrials.gov accessed on April 22nd 2016.
Table 6. Ongoing selected clinical trials targeting the FGFR pathway in solid tumor and hematologic malignancies continued.
| Drug | Mechanism of action | Phase | Study population | Clinicaltrials.gov Identification |
|---|---|---|---|---|
| Dovitinib | Multikinase inhibitor including FGFR1-3 | 2 | Refractory gastrointestinal stromal tumors | NCT01440959 |
| E7090 | FGF/FGFR pathway inhibitor | 1 | Refractory solid tumors dose expansion will enroll patients with tumor expressing genetic abnormality in FGF/FGFR pathway. | NCT02275910 |
| FGF401 | FGFR4-TKI | 1/2 | Hepatocellular carcinoma or solid malignancies characterized by positive FGFR4 and Klotho Berta (KLB) expression | NCT02325739 |
| FPA144 | FGFR2b antibody | 1 | Refractory solid tumors | NCT02318329 |
| GSK3052230 | FGF ligand trap (extra-cellular domain of FGFR1 fused with the Fc region of IgG1) | 1b | Refractory progressive sqNSCLC with FGFR1 gene amplification or MPM with measurable disease | NCT01868022 |
| INCB054828 | FGFR 1-3 TKI | 1 | Refractory solid tumors; on dose expansion subjects with sqNSCLC, gastric cancer, UC, endometrial cancer, multiple myeloma, or MPNs that have a tumor or malignancy that has been evaluated and confirmed to harbor genetic alterations in FGF or FGFR genes | NCT02393248 |
| JNJ-42756493 | Pan FGFR TKI | 2 | Metastatic or surgically unresectable UC that harbor specific FGFR genomic alterations | NCT02365597 |
| JNJ-42756493 | Pan FGFR TKI | 1 | Refractory HCC and for expansion phase participants must have FGF19 amplification in addition | NCT02421185 |
| JNJ-42756493 | Pan FGFR TKI inhibitor | 2a | Asian patients with advanced Non-small-cell lung cancer, urothelial cancer, gastric cancer, esophageal cancer or cholangiocarcinoma with FGFR gene mutation or translocation. | NCT02699606 |
| JNJ-42756493 | Pan FGFR TKI | 1 | Refractory solid tumors and lymphomas | NCT01962532 |
Abbreviations: Fibroblast growth factor (FGF), fibroblast growth factor receptor (FGFR), not reported (NR), estrogen receptor positive (ER+), tyrosine kinase inhibitor (TKI), bacillus calmette-Guerin (BCG), small cell lung cancer (SCLC), Non-small cell lung cancer (NSCLC), squamous non-small cell lung cancer (sqNSCLC), pilot study (PS), glioblastoma multiforme (GBM), hepatocellular carcinoma (HCC), Malignant pleural mesothelioma (MPM), immunohistochemistry (IHC), urothelial carcinoma (UC).www.clinicaltrials.gov accessed on April 22nd 2016.
Table 7. Ongoing selected clinical trials targeting the FGFR pathway in solid tumor and hematologic malignancies continued.
| Drug | Mechanism of action | Phase | Study population | Clinicaltrials.gov Identification |
|---|---|---|---|---|
| JNJ-42756493 | Pan FGFR TKI | 1 | Solid malignancy or lymphoma that is metastatic or unresectable, and for which standard curative treatment is no longer effective | NCT01703481 |
| Lucitanib | VEGFR-FGFR Tyrosine Kinase Inhibitor | 1/2a | Advanced solid tumors, relapsed or refractory to standard therapy. For the dose expansion, patients should have tumors bearing FGFR1 or 11q 12-14 amplification, assessed by FISH or CGH array, or “sensitive” to antiangiogenic treatment | NCT01283945 |
| Lucitanib | Multikinase inhibitor including FGFR | 2 | FGFR1-amplified or non-amplified ER+ metastatic breast cancer | NCT02053636 |
| Lucitanib | Multikinase inhibitor including FGFR | 2 | Metastatic breast cancer | NCT02202746 |
| Lucitanib | Multikinase inhibitor including FGFR | 2 | SCLC or NSCLC with tumor tissue based genetic alterations: FGFR1, FGFR2, FGFR3, VEGFA, or PDGFRα amplification; any FGFR1, FGFR2, or FGFR3 gene fusion; FGFR1, FGFR2, or FGFR3 activating mutation | NCT02109016 |
| LY3076226, | FGFR3 Antibody-Drug Conjugate | 1 | Advanced refractory solid tumors with FGFR3 alterations. | NCT02529553 |
| Nintedanib | Multikinase inhibitor including FGFR | PS | Advanced refractory NSCLC with mutations, rearrangement and fusion involving RET oncogene, or abnormalities (non-synonymous SNV or amplification) in the nintedanib target genes VEGFR1-3, TP53, PDGFR-A, PDGFR-B, and FGFR1-3. | NCT02299141 |
| Nintedanib | Multikinase inhibitor including FGFR | 2 | Refractory salivary gland tumors | NCT02558387 |
| Nintedanib | Multikinase inhibitor including FGFR | 2 | Refractory small Cell Lung Cancer Patients Who Have Previously Benefited From First-line Platinum-based Chemotherapy | NCT02152059 |
Abbreviations: Fibroblast growth factor (FGF), fibroblast growth factor receptor (FGFR), not reported (NR), estrogen receptor positive (ER+), tyrosine kinase inhibitor (TKI), bacillus calmette-Guerin (BCG), small cell lung cancer (SCLC), Non-small cell lung cancer (NSCLC), squamous non-small cell lung cancer (sqNSCLC), pilot study (PS), glioblastoma multiforme (GBM), hepatocellular carcinoma (HCC), Malignant pleural mesothelioma (MPM), immunohistochemistry (IHC), urothelial carcinoma (UC).www.clinicaltrials.gov accessed on April 22nd 2016.
Table 8. Ongoing selected clinical trials targeting the FGFR pathway in solid tumor and hematologic malignancies continued.
| Drug | Mechanism of action | Phase | Study population | Clinicaltrials.gov Identification |
|---|---|---|---|---|
| Nintedanib | Multikinase inhibitor including FGFR | PS | Refractory sqNSCLC | NCT01948141 |
| Nintedanib | Multikinase inhibitor including FGFR | 2 | Advanced FGFR3 mutated, FGFR3 overexpressed, or FGFR3 wild type UC of urinary bladder, urethra, ureter, and renal pelvis and who have failed platinum-based chemotherapy | NCT02278978 |
| Orantinib | Multikinase inhibitor including FGFR | 1/2 | Refractory HCC | NCT00784290 |
| Pazopanib | Multikinase inhibitor including FGFR | PS | Refractory solid tumors FGFR2 amplified and sensitive to pazopanib by Avatar scan | NCT02691767 |
| Pazopanib | Multikinase inhibitor including FGFR | PS | Refractory solid tumors harboring FGFR2 amplification or FGFR2 mutation | NCT02450136 |
| PRN1371 | FGFR1-4 TKI | 1 | Adults with advanced solid tumors | NCT02608125 |
| Ponatinib | Multikinase inhibitor including pan-FGFR | 2 | Advanced biliary cancer with FGFR2 fusion | NCT02265341 |
| Ponatinib | Multikinase inhibitor including pan-FGFR | 2 | Refractory solid tumor or chronic hematologic solid malignancy with activating genomic alterations in FGFR (mutations, fusions or amplifications [> 6 copies]) or activating genomic alterations in KIT, platelet-derived growth factor receptor alpha [PDGFRα], ret proto-oncogene [RET], ABL proto-oncogene 1, non-receptor tyrosine kinase [ABL1] and fms-related tyrosine kinase 3 [FLT3] | NCT02272998 |
Abbreviations: Fibroblast growth factor (FGF), fibroblast growth factor receptor (FGFR), not reported (NR), estrogen receptor positive (ER+), tyrosine kinase inhibitor (TKI), bacillus calmette-Guerin (BCG), small cell lung cancer (SCLC), Non-small cell lung cancer (NSCLC), squamous non-small cell lung cancer (sqNSCLC), pilot study (PS), glioblastoma multiforme (GBM), hepatocellular carcinoma (HCC), Malignant pleural mesothelioma (MPM), immunohistochemistry (IHC), urothelial carcinoma (UC).www.clinicaltrials.gov accessed on April 22nd 2016.
Table 9. Ongoing selected clinical trials targeting the FGFR pathway in solid tumor and hematologic malignancies continued.
| Drug | Mechanism of action | Phase | Study population | Clinicaltrials.gov Identification |
|---|---|---|---|---|
| Regorafenib | Multikinase inhibitor including FGFR | 2 | Refractory epithelial ovarian carcinoma (serous, clear cell, endometrioid, mucinous, mixed, carcinosarcoma and others), fallopian tube and primary peritoneal carcinoma) | NCT02736305 |
| Sunitinib | Multikinase inhibitor including FGFR | PS | Refractory solid tumors harboring RET fusion positive or FGFR2 amplification | NCT02450123 |
| Sunitinib | Multikinase inhibitor including FGFR | PS | RET fusion positive or FGFR2 fusion/other FGFR mutation Refractory solid tumor and/or specific sensitivity to Sunitinib by Avatar scan | NCT02691793 |
| TAS120 | FGFR TKI | 1 | Advanced metastatic solid tumors with or without abnormalities of FGF/FGFR who have failed all standard therapies or for whom standard therapy does not exist or multiple myeloma with amplification, mutation or translocation or other associated abnormalities of FGF/FGFR who have failed all standard therapies or for whom standard therapy does not exist | NCT02052778 |
| U3-1784 | FGFR4 monoclonal antibody | 1 | Refractory solid tumors | NCT02690350 |
| XL228 | Multi-targeted kinase inhibitor IGF-1R, Src, FGFR, and BCR-Abl | 1 | Refractory solid tumors, lymphoma, or multiple myeloma | NCT00526838 |
Abbreviations: Fibroblast growth factor (FGF), fibroblast growth factor receptor (FGFR), not reported (NR), estrogen receptor positive (ER+), tyrosine kinase inhibitor (TKI), bacillus calmette-Guerin (BCG), small cell lung cancer (SCLC), Non-small cell lung cancer (NSCLC), squamous non-small cell lung cancer (sqNSCLC), pilot study (PS), glioblastoma multiforme (GBM), hepatocellular carcinoma (HCC), Malignant pleural mesothelioma (MPM), immunohistochemistry (IHC), urothelial carcinoma (UC).www.clinicaltrials.gov accessed on April 22nd 2016.
FGFR ABERRATIONS
FGFR alterations relatively rare and were present in ~7% of a cohort of 4853 tumor samples including 47 different histological types. [7] When limiting to histologies with at least 75 samples analyzed, the frequencies of FGFR aberrations were higher among urothelial (31.7%), breast (17.4%), endometrial (11.3%), and endometrial/ovarian carcinomas (8.1%). [7] The majority of FGFR genomic alterations were FGFR amplifications and mutations (92%).
Evidence from early phase clinical trials support that FGFR aberrations can represent targetable events in solid tumors. In a phase I trial, twenty one patients with refractory squamous non-small cell lung cancer (sqNSCLC) harboring FGFR1-amplification were treated with the small molecule FGFR inhibitor BGJ398 (100 or 125 mg once daily in 28-day cycles). [8] Of 17 evaluable patients, 4 had radiologic tumor reduction and 3 had stable disease indicating that a subset of patients with FGFR pathway aberrations indeed benefits for FGFR targeted therapy. In another phase I trial BGJ398 also showed evidence of clinical activity in patients with solid tumors harboring FGFR aberrations, including 4 of 5 patients with urothelial cell carcinomas (4 of which originated in the bladder) with FGFR3-activating mutations. Additionally, two patients with FGFR1-amplified sqNSCLC achieved confirmed partial response. Tumor reductions were also observed in cholangiocarcinoma with an FGFR2 gene fusion, and FGFR1-amplified breast cancer. [9] In summary, given the rarity of FGFR aberrations the preliminary evidence of antitumor activity of FGFR targeted therapies comes from subpopulations of small early phase clinical trials. A small number of partial responses to FGFR targeted therapies have been documented among patients with sqNSCLC harboring FGFR1 amplification (BGJ398), cholangiocarcinoma harboring FGFR2 translocations (BGJ398), glioblastoma positive for FGFR3 translocation (JNJ-42756493), bladder cancer harboring FGFR3 mutations and translocations are sensitive to targeted therapies (JNJ-42756493 and BGJ398). [8–11]
FGFR3-TACC3 TRANSLOCATION
Fusions have been described in the FGFR1-3 genes with multiple partners (i.e., TACC1, TACC2, TACC3, BAIAP2L1, BICC1, NPM1, PPAPDC1A, AFF3, SLC45A3 and AHCYL1) in a wide spectrum of tumors (i.e., cholangiocarcinoma, breast, and prostate cancer, sqNSCLC, gastric adenocarcinoma, colorectal adenocarcinoma, carcinoma of unknown primary and glioblastoma). [7, 12–15]
Other rare FGFR fusions described included FGFR2-TACC3 (1), FGFR2-NPM1(3), FGFR2-TACC2(2), FGFR2-BICC1(2), FGFR2-C10orf68(1), FGFR3-JAKMIP1(1), FGFR2-KIAA1598(1), FGFR2-NCALD(1), FGFR2-NOL4(1), FGFR1-NTM(1), FGFR2-PPAPDCA(1), FGFR3-TNIP2(1), and FGFR3-WHSC1(1). Of note no FGFR4 gene fusion was observed in this large series. [7] Preliminary pre-clinical data support that these fusions represent important therapeutic targets. For instance stable cell lines harboring FGFR3-BAIAP2L1, FGFR3-TACC3, and FGFR2-CCDC6 fusions showed expression of active FGFR fusion kinases and activation of downstream mitogen-activated protein kinase ERK1/2 and the transcription factor STAT1. [12] The presence of FGFR fusions (FGFR3-BAIAP2L1) not only enhanced tumor cell proliferation, but also led to significant sensitivity to small kinase inhibitors (PD173074) in pre-clinical cellular and xenograft bladder cancer models, in contrast with FGFR3 mutant cell lines, which were not sensitive to kinase inhibition. [12] FGFR1-TACC1 fusion targeted therapy also showed anti-tumor effects in pre-clinical GBM model. [13] Cholangiocarcinoma mice model harboring FGFR2-AHCYL1, FGFR2-CCDC6, and FGFR2-BICC1 was sensitive to treatment with FGFR kinase inhibitors BGJ398 and PD173074. [16, 17]
TACC3 gene has been identified as an important partner of these FGFR fusions and associated with the pathogenesis of several solid tumors. [18–20] TACC3 (transforming acidic coiled-coil containing protein 3) belongs to the TACC gene family, which also includes TACC1 and TACC2. TACC3 protein has a coiled-coil domain at the C terminus, known as the TACC domain, which promotes stability and organization of mitotic spindle. [21]
FGFR3-TACC3 fusions were first described in glioblastoma multiforme (GBM) and bladder urothelial tumors. [13, 15] Subsequent manuscripts underscored the low frequency of FGFR3-TACC3 gene fusions across different tumor types.
Among the 4853 tumors samples analyzed by Helsten et al. only 28 exhibited FGFR gene fusions, and 14 samples had FGFR3-TACC3 fusions. Accompanying genomic aberrations observed primarily affected TP53 tumor suppressor gene, AKT/mTOR/PTEN pathway and cell cycle control genes (i.e., CNNE1, CDK2 and 4) (Table 1). [7]
Table 1. Co-existing genomic aberrations among 14 FGFR3-TACC3 fusion cases.[7].
| Histology | Cell cycle | PI3Kinase | TP53 | DNA repair | Transcription/Histone methylation | Growth factor receptor | RAS/RAF/MAPK | Other |
|---|---|---|---|---|---|---|---|---|
| CUP | CDK4-amplification | MDM2-amp ARID1A-A343_A348>A | ||||||
| Cervical cancer | PIK3CA-E545K | |||||||
| Cervical cancer | TP53-R248Q | ATM-V2115fs*5 | ||||||
| Endometrial adenocarcinoma | PIK3CA-E365K | TP53-W91* | FLT3-E978* | KMT2A-complex rearrange | ||||
| Gallbladder carcinoma | CCNE1-amplification | TP53-C141* | MYC-amplification | MCL1-amplification | ||||
| Glioma | TP53-E258K |
NF1 Y2285fs*5, Q270* MCL1-amplification NFKBIA-amplification PTPN11-E76K |
||||||
| Glioma | CDK4-amplification | PTEN-splice site 209+1delGT | MDM2-amplification | |||||
| NSCLC Not specified |
CDKN2A-loss | MDM2-amplification | ||||||
| Pancreatic exocrine carcinoma | TP53-I195F | ATM-R805* | MYC-amplification | SMAD4-loss | ||||
| Renal cell carcinoma | CDKN2A/B-loss | TP53-R196* | MYC-amplification |
TOP1-amplification SRC-amplification AURKA-amplification VHL-P25L |
||||
| Urothelial carcinoma | CCND1-amplification | PIK3CA-H1047R | TP53-R280T | |||||
| Urothelial carcinoma | TP53-K132N |
JUN-amplification IRS2-amplification MCL1-amplification |
||||||
| Urothelial carcinoma |
CCND1-amplification CDKN2A/B-loss |
PIK3R2-E543* | ERBB2 amplification | MAP3K1-truncation, exon 15 |
MDM2-amplification NF1-I1351M |
|||
| Urothelial carcinoma | CDKN2A/B-loss |
IRS2-amplification MDM2-amplification |
Abbreviations: Non-small cell lung cancer (NSCLC), carcinoma of unknown primary (CUP), mouse double minute 2 homolog (MDM2), cyclin E1 gene (CCNE1), cyclin D1 gene(CCND1), cyclin dependent kinase(CDK), cyclin dependent kinase inhibitor (CDKN), phosphatidylinositol 3-Kinase (PI3K), Erb-B2 receptor tyrosine kinase 2 (ERBB2), tumor protein P53(TP53), phosphatase and tensin homolog(PTEN), V-Myc Avian Myelocytomatosis viral oncogene homolog (MYC), ataxia telangiectasia mutated(ATM), mitogen-activated protein kinase 1 (MAP3K1), AT rich interactive domain 1A (ARID1A), lysine (K)-specific methyltransferase 2A(KMT2A), myeloid cell leukemia 1(MCL1), neurofibromin 1(NF1), Nuclear Factor Of Kappa Light Polypeptide Gene Enhancer In B-Cells Inhibitor, alpha (NFKBIA), protein tyrosine phosphatase, non-Receptor Type 11(PTPN11), SMAD family member 4(SMAD4), topoisomerase (DNA) I(TOP1), SRC proto-oncogene(SRC), aurora kinase A(AURKA), von hippel-lindau tumor suppressor (VHL), Jun proto-oncogene(JUN), insulin receptor substrate 2(IRS2).
FGFR3-TACC3 fusions are formed by rare intrachromosomal rearrangements located within 150 kb of the FGFR3 gene on chromosome 4p16 (Figure 2). [13] It has been proposed that the fusion, which occurs via a tandem duplication event, leads to loss of an miR-99a binding site within the 3′-untranslated region (3′- UTR) of FGFR3, releasing FGFR3 signaling from miR-99a-dependent inhibition and enhancing tumor progression relative to wild type FGFR3. [22] Given the close proximity of the FGFR3 and TACC3 genes, identification of the fusion by FISH is technically challenging. [23] Therefore the most common methods have involved transcriptomic analysis by RNA-seq and RT-PCR in addition to whole-genome sequencing. [7, 13, 23] Di Stefano et al. have also utilized immunostaining of the N terminus of FGFR3 and found uniform overexpression of FGFR3 in a subset of glioblastoma multiforme (GBM) cases with FGRF3-TACC fusions that were identified by RT-PCR. [23] Those findings demonstrate that detection of FGFR3 amplification by IHC or IF may serve as a method to screen for FGRF3-TACC fusions which is clinically relevant since RNA sequencing is not common practice. [24] The fusion protein exhibits constitutive kinase activity, induces mitotic, chromosomal segregation defects and triggers aneuploidy (Figure 1B). [13] The presence of the TACC coiled-coil domain increases the activity of FGFR3 through more promiscuous constitutive phosphorylation of tyrosine kinase residues within the FGFR3 protein, and preferential MAPK kinase activation (Figure 1A). [25] Moreover, preclinical and clinical results have shown that FGFR inhibitors can block the tyrosine kinase activity of this fusion protein.
Figure 2. FGFR3-TACC3 gene fusion.
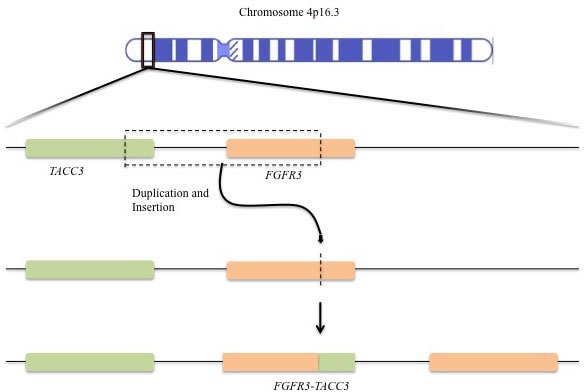
Tandem duplication and insertion leads the fusion of the tyrosine kinase domain of FGFR3 to the TACC domain of TACC3.
Treatment of nasopharyngeal carcinoma cells carrying the FGFR3-TACC3 fusion with FGFR inhibitor PD173074 inhibited cell proliferation. [26] RT4 urothelial carcinoma line harboring FGFR3-TACC3 fusion also exhibited sensitivity to this same FGFR inhibitor in a xenograft model. [12] Similar results were seen in GBM in vivo and in vitro models carrying the FGFR3-TACC3 fusions, which was not only associated oncogenic transformation but also exhibited significant sensitivity to FGFR inhibitor JNJ-42756493. [13, 22, 23]
The clinical relevance of FGFR3-TACC3 has been underscored by preliminary results from clinical studies and case reports of tumor responses to the treatment with FGFR inhibitors. For instance, the phase I trial with FGFR inhibitor JNJ-42756493 including 65 patients with advanced solid tumors included 4 patients with FGFR3-TACC3 translocation. [10] Three partial responses (two confirmed and one unconfirmed) were seen among 3 patients with urothelial, two of which cancer harbored FGFR3-TACC3 fusion. One of these patients stayed on treatment for about 10 months. Another confirmed partial response was observed in a patient with glioblastoma with FGFR3-TACC3. Tumor shrinkage was also seen in a patient with adrenal carcinoma with FGFR3-TACC3/FGFR2- CCDC6 (FGFR3-TACC3 being the predominant translocation), who received treatment for 10 months before disease progression. In agreement with preclinical results, these results suggest that FGFR3-TACC3 fusion is indeed an actionable therapeutic target.
PREVALENCE OF FGFR3-TACC3 FUSION IN SOLID TUMORS
Gliomas
The first report of this fusion emerged from a small subset of patients with GBM with 2 out of 97 samples harboring FGFR3-TACC3. [13] A subsequent study with 795 cases of gliomas (584 GBMs and 211 grade II-III gliomas) described 17 cases (2.9%) of FGFR3-TACC3 genomic fusions among the GBM group, and 3 cases among lower grade gliomas. [23] Amplicons ranged from 928 bp (for FGFR3exon18-TACC3exon13) to 1,706 bp (for FGFR3ex18-TACC3ex4). All tested FGFR3-TACC3 positive tumors showed strong expression of FGFR3 by tumor cells on immunohistochemistry (IHC). Epidermal growth factor receptor gene (EGFR) amplification showed significant negative correlation with FGFR3-TACC3 status; CDK4 and MDM2 amplification had significant positive correlation with FGFR3-TACC3 (CDK4 amplification was seen in 7/16 FGFR3-TACC3 positive cases). No statistically significant correlation was seen between FGFR3-TACC3 fusions and other genetic and epigenetic alterations (CDKN2A deletion, TERT promoter mutations, gain of chromosome 7p, loss of chromosome 10q, and methylation of the MGMT promoter). Parker et al. also reported higher frequency of FGFR3-TACC3 fusions in GBMs when compared to gliomas. [22] Four out of 48 (8.3%) GBMs tested were positive for this fusion, whereas none of the 43 low-grade glioma samples showed this translocation. In addition, 2 out 157 GBM (1.2%) and 2 out of 461 low grade gliomas (0.4%) samples from The Cancer Genome Atlas (TCGA) dataset tested positive for FGFR3-TACC3 fusions which reinforced the notion that this fusion is more common in high grade gliomas. [14] In another series 3 out of 59 patients with primary GBM harbored this fusion gene. [27] Wu et al. reported 2 cases of GBM harboring FGFR3-TACC3 fusion from the TGCA database. [12] In summary, it is estimated that 1.2-8.3% of GBMs will carry this translocation.
Urothelial cancer
The TCGA project reported a comprehensive genomic analysis of 131 high-grade muscle invasive urothelial bladder carcinomas including gene fusions. Three cases of tumors with FGFR3-TACC3 fusion were reported (~2.3%). The breakpoints were in intron 10 of TACC3 and intron 16 (2 cases) or exon 17 (1 case) of FGFR3. [28] Two cases among 99 samples analyzed showed two distinct FGFR3-TACC3 genomic fusions; first, intron 17 of FGFR3 with intron 10 of TACC3 resulting in exon 17 of FGFR3 being spliced 5′ to exon 11 of TACC3 in the fuse mRNA; second, intron 17 of FGFR3 with exon 4 of TACC3 at the gene level and in the fused mRNA, exon 17 and fragment of intron 17 in FGFR3, and fragment of exon 4 in TACC3 were merged into a novel exon. [29] Analysis of 250 samples of bladder urothelial carcinoma by RNA-seq detected FGFR3-TACC3 in 5 specimens (2%). [14] Helsten et al. described 126 cases of urothelial carcinomas and 4 cases (3%) of FGFR3-TACC3 translocations were observed. [7] Williams et al. reported results of analysis of 2 tumor samples positive for FGFR3 exon 18 and TACC3 exon 13 fusions among 32 selected bladder carcinoma samples. [15] Taken together, a total of 14 cases of urothelial carcinoma carrying FGFR3-TACC3 translocation have been described with an estimated incidence of 2.6% among the 539 cases described above.
Non-small cell lung cancer
RNA sequencing of 492 sqNSCLC from the TCGA showed only 3 cases of FGFR3-TACC3 fusions; none of 513 cases of adenocarcinomas harbored this translocation. [14] Another study describing genomic analysis of 675 cases of NSCLC showed only one case of FGFR3-TACC3 fusion (breakpoint not specified), this was associated with existing CDKN2A-loss and MDM2 amplification. [7] Wu et al. reported through analysis of the TGCA dataset 4 cases of sqNSCLC harboring FGFR3-TACC3 fusion. [12] Capelletti et al. identified 3 out 576 patients with adenocarcinoma of the lung harboring FGFR3-TACC3. [30] Fusion variants identified included exon 17 FGFR3 to exon exon 8 of TACC3; exon 17 of FGFR3 and exon 11 of TACC3; and exon 17 of FGFR3 to exon 4 TACC3 resulting in an overall prevalence of 0.5%. An additional analysis of 1,328 NSCLC samples revealed 15 FGFR3-TACC3 fusion variants identified through RT-PCR [31]. Histological distribution was as follows: 6/1016 (0.6%) adenocarcinomas and 9/312 (2.9%) squamous cell carcinoma of the lung suggesting the FGFR3-TACC3 fusions are more frequent in sqNSCLC. FGFR3-TACC3 fusion correlated with positive smoking history and male gender in univariate analysis. Tumor size > 3cm correlated with FGFR3 fusions independently in multivariate modeling. FGFR3-TACC3 fusions were distributed as follows: E18:E11 2 cases, E17:E11 9 cases, E17:E8 1 case, E17:E5 1 case, and E17:E10 2 cases.
Another cohort of patients from Korea was analyzed and only 2 cases of FGFR3-TACC3 fusion were detected (E17:E8 and E18:E9). The same author reviewed the samples of the TCGA database and found 4 out of 178 samples positive for this fusion. [32] Two more cases of FGFR3 E18 fused with TACC3 E10 were reported in sqNSCLC. [33] The TCGA dataset reported genomic analysis through RT-PCR of 230 previously untreated adenocarcinomas of the lung and 178 previously untreated sqNSCLC; FGFR fusions were not reported in either series. [34, 35] In summary, FGFR3-TACC3 fusions are rare in NSCLC but seem to be more common in sqNSCLC histological type with an estimated prevalence consistently lower than 2.9%.
Cervical cancer
Three cases of TACC3-FGFR3 translocations break points in patients with advanced cervical carcinoma have been described with early evidence of clinical benefit from FGFR targeted treatment. [36] One of the cases was a metastatic recurrent to the lung adenosquamous carcinoma of the cervix. The metastatic tumor showed FGFR3-TACC3 fusion (break point intron 17 and TACC3 intron 10). Associated genomic aberrations were: AKT1 missense mutation, mTOR point mutation, ATRX truncating nonsense mutation. The patient was treated with FGFR targeted therapy achieving stable disease for 4 cycles. In the second case, the patient had well differentiated squamous cell carcinoma of the cervix stage. Tumor tissue from the original biopsy showed FGFR3-TACC3 fusion (break points FGFR3 intron 18 and TACC3 intron 7) as well as BRAF 3′ tandem duplication, activating PIK3CA missense mutation, CDNK2A loss, and activating missense mutations in KRAS and HRAS. The third case was a squamous cell carcinoma of the cervix in which the following genomic aberrations were identified on hysterectomy specimen: FGFR3-TACC3 fusion (breakpoints at FGFR3 intron 17 and TACC intron 10) and partial loss of STK11, and RB1 loss.
Xiang et al. performed transcriptomic (RNA) followed by cDNA analysis of 285 cases of early stage carcinoma of the cervix; 11 cases of FGFR3-TACC3 translocations were described (3.9%) with 4 variants [FGFR3 (1_758) fused with TACC3 (549_838), FGFR3(1_758) fused with TACC3(648_838), FGFR3 (1_758) fused with TACC3 (648_838), and the most frequent variant FGFR3 (1_768) fused with TACC3(538_838)] [37]. There were four cases of squamous cell carcinoma, 3 cases of adenocarcinoma 3 cases of adenosquamous, and 1 small cell carcinoma. Other sporadic cases of FGFR3-TACC3 translocation in cervical carcinoma have been reported as part of genomic aberrations analysis of solid tumors: FGFR3-TACC3 fusion (intron 17-intron 7) one case of cervical carcinoma. [7]
Head and neck cancer
Among 411 head and neck SCC tumor samples analyzed using RNA sequencing data through the TGCA only 2 harbored FGFR3-TACC3 fusion. [14] Wu et al. reported two additional cases of head and neck cancer harboring this fusion. [12] An additional analysis of 279 head and neck SCC tumor samples from the TGCA dataset (172 oral cavity, 33 oropharynx, 72 laryngeal tumors) and cases of FGFR3-TACC3 were appreciated in two of the HPV+ samples. [38]
Gastrointestinal malignancies
FGFR3-TACC3 fusions are rare event as illustrated by analysis RNA sequencing of 856 tumor samples [hepatocellular carcinoma (194), colon (286), rectum (91), and gastric adenocarcinomas (285)] ; in which no tumor showed this translocation. [14]
Other malignancies
Two studies analyzed a total of 1594 cases of breast cancer no FGFR3-TACC3 fusion was reported. [7, 14] Single cases of carcinoma of unknown primary, endometrial carcinoma, renal cell carcinoma, gallbladder, papillary kidney tumor, and prostate adenocarcinoma harboring the FGFR3-TACC3 fusion have been described. [7, 14]
DISCUSSION
In the recent years the field of medical oncology has witnessed unprecedented advances in the understanding of cancer biology. FGF/FGFR pathway is a thriving area of targeted drug development in a wide array of tumors (Tables 3, 4, 5, 6, 7, 8, 9). FGFR3-TACC3 was first described in 2012 in two patients with GBM. [13] Since then FGFR3-TACC3 fusion has been reported in numerous solid tumors including urothelial carcinoma, NSCLC, thyroid, and cervical carcinoma (Table 2). [12] In addition dataset analysis through bioinformatics, namely the TCGA dataset, yield other rarer instances in which this aberration can be found such as prostate cancer, head and neck cancer, and kidney papillary cancer. [14]
Table 2. Cross-sectional studies and case series reporting positive FGFR3-TACC3 fusions (excluding TCGA dataset samples).
| Authors | Tumor type | Number of cases analyzed | Number of cases harboring FGFR3-TACC3 fusion | FGFR3 breakpoint | TACC3 breakpoint | Comments |
|---|---|---|---|---|---|---|
| Helsten et al.[7] | Urothelial carcinoma | 126 | 4 | NR | NR | Organ site was not specified |
| Williams et al.[15] | Bladder carcinoma | 32 | 2 | Exon 18 | Exon 13 | |
| Singh et al.[13] | GBM | 97 | 2 | Exon 17 | Intron 7 | |
| Di Stefano et al.[23] | Gliomas | 795 | 20 | Exon 17,18 | Exon 4, 5, 6, 8, 10, 11 | 17 patients had GBM and 3 patients grade III or II gliomas |
| Parker et al. [22] | GBM | 48 | 4 | -- | -- | |
| Bao Z. et al [27] | GBM | 59 | 3 | Exon 17 | Exons 8,10,11 | |
| Helsten et al. [7] | NSCLC (subtype not specified) | 675 | 1 | NR | NR | NSCLC subtype was not specified |
| Capelletti et al.[30] | Adenocarcinoma of the lung | 576 | 3 | Exon 17 | Exon 4, 8, 11 | |
| Wang et al. [31] | NSCLC (6 adenocarcinomas 9 sqNSCLC) | 1328 | 15 | Exon 17,18 | Exon 5,8,10,11 | 6 cases of adenocarcinoma and 9 cases of SCC; FGFR3-TACC3 correlated independently with tumor size > 3 cm |
| Kim et al.[32] | sqNSCLC | 104 | 2 | Exon 17, 18 | Exon 8,9 | Author also found 4 more cases at the TGCA dataset |
| Majewski et al.[33] | sqNSCLC | 95 | 2 | Exon 18 | Exon 10 | Fusion was identified in 2 SCC cases |
| Carneiro et al.[36] | Cervical cancer: SSC and adenosquamous cell carcinoma, | -- | 3 | Intron 17,18 | Intron 7,10 | |
| Xiang et al. [37] | Cervical cancer: SCC, adenocarcinoma, adenosquamous, small cell carcinoma | 285 | 11 | -- | -- | All early stage tumors |
| Helsten et al. [7] | Cervical adenocarcinoma and cervical carcinoma not specified | 48 | 2 | Intron 17 | Intron 7 | Fusion reported for carcinoma NOS |
| Helsten et al. [7] | Carcinoma of unknown primary | 267 | 1 | NR | NR | |
| Helsten et al. [7] | Endometrial Carcinoma | 80 | 1 | NR | NR | |
| Helsten et al.[7] | Gallbladder carcinoma | 47 | 1 | NR | NR | |
| Helsten et al. [7] | Glioma | 144 | 1 | NR | NR | |
| Helsten et al. [7] | Pancreatic exocrine tumor | 172 | 1 | Exon 18 | Intron 10 | |
| Helsten et al. [7] | Renal cell carcinoma | 87 | 1 | NR | NR | |
| Parish et al.[39] | Solid tumors | 391 | 1 | NR | NR |
Non-small cell lung cancer (NSCLC), squamous non-small cell lung cancer (sqNSCLC), glioblastoma multiforme (GBM), fibroblast growth factor receptor 3-transforming acidic coiled-coil containing protein 3(FGFR3-TACC3), Not reported (NR), squamous cell carcinoma (SCC), The Cancer Genome Atlas (TCGA).
The clinical relevance of FGFR3-TACC3 has been highlighted by 3 out of 4 partial responses among patients with tumors harboring FGFR3-TACC3 fusions treated with FGFR inhibitor JNJ-42756493. [10] In agreement with preclinical results, these results suggest that FGFR3-TACC3 fusion is indeed an actionable therapeutic target. Furthermore taking into account the obvious limitation of the sparse clinical data thus far tumors harboring FGFR3-TACC3 fusions seem to be more sensitive FGFR targeted therapies when compared to other FGFR aberrations. This is particularly important, as there are limited treatment options for patients with aggressive tumors in which FGFR3-TACC3 fusions have been described such as GBM and bladder cancer. Future studies should take into account that FGFR3-TACC3 fusion has shown positive correlations with PI3K/AKT/mTOR pathway, cell cycle control (CDK4, CDK2, and CCND1), and MDM2 aberrations (Table 1). [23, 36, 39] The corollary to these concomitant findings is that one could hypothesize that the combined approach targeting FGFR3-TACC3 and its downstream oncogenic proteins may further enhance the efficacy of FGFR aberrant targeted therapy. Notwithstanding its rarity, FGFR3-TACC3 fusions are present in wide array of solid tumor types and its analysis should be an integral part of screening procedures in FGFR targeted trials in solid tumors.
Elucidation of the potential interaction between FGFR and its fusions with the immune system is also warranted. Pre-clinical data suggest that FGFR3 mutations are exclusive to non-inflamed bladder cancers (~ 30% of tumors), a subgroup characterized by absence of CD8 tumor infiltrating lymphocytes (CD8 TILs), worsen prognosis, and lower chance of responding to PD-L1 blockade. [40–42] The exclusion of CD8 TILs from the tumor microenvironment limits the benefit from immunotherapies in melanoma and might carry similar relevance to bladder cancer. [42] Nevertheless, these preliminary results highlight a possible link between FGFR pathway aberrations and immune modulation of tumor microenvironment that could be explored therapeutically.
Footnotes
CONFLICTS OF INTEREST
There is no conflict of interest.
REFERENCES
- 1.Carneiro BA, Costa R, Taxter T, Chandra S, Chae YK, Cristofanilli M, Giles FJ. Is Personalized Medicine Here? Oncology (Williston Park) 2016;30:293–303. 307. [PubMed] [Google Scholar]
- 2.Chapman PB, Hauschild A, Robert C, Haanen JB, Ascierto P, Larkin J, Dummer R, Garbe C, Testori A, Maio M, Hogg D, Lorigan P, Lebbe C, et al. Improved survival with vemurafenib in melanoma with BRAF V600E mutation. The New England journal of medicine. 2011;364:2507–2516. doi: 10.1056/NEJMoa1103782. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Maemondo M, Inoue A, Kobayashi K, Sugawara S, Oizumi S, Isobe H, Gemma A, Harada M, Yoshizawa H, Kinoshita I, Fujita Y, Okinaga S, Hirano H, Yoshimori K, Harada T, Ogura T, et al. Gefitinib or chemotherapy for non-small-cell lung cancer with mutated EGFR. The New England journal of medicine. 2010;362:2380–2388. doi: 10.1056/NEJMoa0909530. [DOI] [PubMed] [Google Scholar]
- 4.Demetri GD, von Mehren M, Blanke CD, Van den Abbeele AD, Eisenberg B, Roberts PJ, Heinrich MC, Tuveson DA, Singer S, Janicek M, Fletcher JA, Silverman SG, Silberman SL, Capdeville R, Kiese B, Peng B, et al. Efficacy and safety of imatinib mesylate in advanced gastrointestinal stromal tumors. The New England journal of medicine. 2002;347:472–480. doi: 10.1056/NEJMoa020461. [DOI] [PubMed] [Google Scholar]
- 5.Touat M, Ileana E, Postel-Vinay S, Andre F, Soria JC. Targeting FGFR Signaling in Cancer. Clin Cancer Res. 2015;21:2684–2694. doi: 10.1158/1078-0432.CCR-14-2329. [DOI] [PubMed] [Google Scholar]
- 6.Gallo LH, Nelson KN, Meyer AN, Donoghue DJ. Functions of Fibroblast Growth Factor Receptors in cancer defined by novel translocations and mutations. Cytokine Growth Factor Rev. 2015;26:425–449. doi: 10.1016/j.cytogfr.2015.03.003. [DOI] [PubMed] [Google Scholar]
- 7.Helsten T, Elkin S, Arthur E, Tomson BN, Carter J, Kurzrock R. The FGFR Landscape in Cancer: Analysis of 4,853 Tumors by Next-Generation Sequencing. Clin Cancer Res. 2016;22:259–67. doi: 10.1158/1078-0432.CCR-14-3212. [DOI] [PubMed] [Google Scholar]
- 8.Nogova L, Cassier PA, Hidalgo M, Delord JP, Schuler MH, Lim WT, Camidge DR, Buettner R, Heukamp LC, Gardizi M, Scheffler S, Kambartel K, Ringeisen FP, Sen S, et al. Targeting FGFR1-amplified lung squamous cell carcinoma with the selective pan-FGFR inhibitor BGJ398. J Clin Oncol. 2014;32:5. [Google Scholar]
- 9.Lecia V, Sequist PC, Varga V, Tabernero J, Schellens JH, Delord JP, LoRusso P, Camidge DR, Medina MH, Schuler M, Campone M, Tian GG, Wong S, Corral J, et al. Phase I study of BGJ398, a selective pan-FGFR inhibitor in genetically preselected advanced solid tumors. [abstract]. Proceedings of the 105th Annual Meeting of the American Association for Cancer Research; 2014 Apr 5-9; San Diego, CA Philadelphia (PA): AACR. Cancer Res 2014;74:Abstract nr CT326. [DOI] [Google Scholar]
- 10.Tabernero J, Bahleda R, Dienstmann R, Infante JR, Mita A, Italiano A, Calvo E, Moreno V, Adamo B, Gazzah A, Zhong B, Platero SJ, Smit JW, et al. Phase I Dose-Escalation Study of JNJ-42756493, an Oral Pan-Fibroblast Growth Factor Receptor Inhibitor, in Patients With Advanced Solid Tumors. J Clin Oncol. 2015;33:3401–3408. doi: 10.1200/JCO.2014.60.7341. [DOI] [PubMed] [Google Scholar]
- 11.Milind M, Javle RTS, Zhu A, Sadeghi S, Choo SP, Borad MJ, Lowery MA, El-Khoueiry A, Macarulla T, Philip PA, Oh DY, Van Cutsem E, Yeh KH, Isaacs R, McGarry C, Sen S, Bekaii-Saab TS. A phase 2 study of BGJ398 in patients (pts) with advanced or metastatic FGFR-altered cholangiocarcinoma (CCA) who failed or are intolerant to platinum-based chemotherapy. J Clin Oncol. 2016;34 [Google Scholar]
- 12.Wu YM, Su F, Kalyana-Sundaram S, Khazanov N, Ateeq B, Cao X, Lonigro RJ, Vats P, Wang R, Lin SF, Cheng AJ, Kunju LP, Siddiqui J, Tomlins SA, Wyngaard P, Sadis S, et al. Identification of targetable FGFR gene fusions in diverse cancers. Cancer discovery. 2013;3:636–647. doi: 10.1158/2159-8290.CD-13-0050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Singh D, Chan JM, Zoppoli P, Niola F, Sullivan R, Castano A, Liu EM, Reichel J, Porrati P, Pellegatta S, Qiu K, Gao Z, Ceccarelli M, Riccardi R, Brat DJ, Guha A, et al. Transforming fusions of FGFR and TACC genes in human glioblastoma. Science (New York, NY) 2012;337:1231–1235. doi: 10.1126/science.1220834. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Stransky N, Cerami E, Schalm S, Kim JL, Lengauer C. The landscape of kinase fusions in cancer. Nature communications. 2014;5:4846. doi: 10.1038/ncomms5846. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Williams SV, Hurst CD, Knowles MA. Oncogenic FGFR3 gene fusions in bladder cancer. Human molecular genetics. 2013;22:795–803. doi: 10.1093/hmg/dds486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Arai Y, Totoki Y, Hosoda F, Shirota T, Hama N, Nakamura H, Ojima H, Furuta K, Shimada K, Okusaka T, Kosuge T, Shibata T. Fibroblast growth factor receptor 2 tyrosine kinase fusions define a unique molecular subtype of cholangiocarcinoma. Hepatology. 2014;59:1427–1434. doi: 10.1002/hep.26890. [DOI] [PubMed] [Google Scholar]
- 17.Wang Y, Ding X, Wang S, Moser CD, Shaleh HM, Mohamed EA, Chaiteerakij R, Allotey LK, Chen G, Miyabe K, McNulty MS, Ndzengue A, Knudson RA, Greipp PT, Clark KJ, Torbenson MS, et al. Antitumor effect of FGFR inhibitors on a novel cholangiocarcinoma patient derived xenograft mouse model endogenously expressing an FGFR2-CCDC6 fusion protein. Cancer letters. 2016 doi: 10.1016/j.canlet.2016.05.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Huang ZL, Lin ZR, Xiao YR, Cao X, Zhu LC, Zeng MS, Zhong Q, Wen ZS. High expression of TACC3 in esophageal squamous cell carcinoma correlates with poor prognosis. Oncotarget. 2015;6:6850–6861. doi: 10.18632/oncotarget.3190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Ha GH, Kim JL, Breuer EK. TACC3 is essential for EGF-mediated EMT in cervical cancer. PloS one. 2013;8:e70353. doi: 10.1371/journal.pone.0070353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Lauffart B, Vaughan MM, Eddy R, Chervinsky D, DiCioccio RA, Black JD, Still IH. Aberrations of TACC1 and TACC3 are associated with ovarian cancer. BMC Womens Health. 2005;5:8. doi: 10.1186/1472-6874-5-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Hood FE, Royle SJ. Pulling it together: The mitotic function of TACC3. Bioarchitecture. 2011;1:105–109. doi: 10.4161/bioa.1.3.16518. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Parker BC, Annala MJ, Cogdell DE, Granberg KJ, Sun Y, Ji P, Li X, Gumin J, Zheng H, Hu L, Yli-Harja O, Haapasalo H, Visakorpi T, Liu X, Liu CG, Sawaya R, et al. The tumorigenic FGFR3-TACC3 gene fusion escapes miR-99a regulation in glioblastoma. The Journal of clinical investigation. 2013;123:855–865. doi: 10.1172/JCI67144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Di Stefano AL, Fucci A, Frattini V, Labussiere M, Mokhtari K, Zoppoli P, Marie Y, Bruno A, Boisselier B, Giry M, Savatovsky J, Touat M, Belaid H, Kamoun A, Idbaih A, Houillier C, et al. Detection, Characterization, and Inhibition of FGFR-TACC Fusions in IDH Wild-type Glioma. Clin Cancer Res. 2015;21:3307–3317. doi: 10.1158/1078-0432.CCR-14-2199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Byron SA, Van Keuren-Jensen KR, Engelthaler DM, Carpten JD, Craig DW. Translating RNA sequencing into clinical diagnostics: opportunities and challenges. Nat Rev Genet. 2016;17:257–271. doi: 10.1038/nrg.2016.10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Nelson KN, Meyer AN, Siari A, Campos AR, Motamedchaboki K, Donoghue DJ. Oncogenic Gene Fusion FGFR3-TACC3 is Regulated by Tyrosine Phosphorylation. Mol Cancer Res. 2016;14:458–69. doi: 10.1158/1541-7786.MCR-15-0497. [DOI] [PubMed] [Google Scholar]
- 26.Yuan L, Liu ZH, Lin ZR, Xu LH, Zhong Q, Zeng MS. Recurrent FGFR3-TACC3 fusion gene in nasopharyngeal carcinoma. Cancer biology & therapy. 2014;15:1613–1621. doi: 10.4161/15384047.2014.961874. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Bao ZS, Chen HM, Yang MY, Zhang CB, Yu K, Ye WL, Hu BQ, Yan W, Zhang W, Akers J, Ramakrishnan V, Li J, Carter B, Liu YW, Hu HM, Wang Z, et al. RNA-seq of 272 gliomas revealed a novel, recurrent PTPRZ1-MET fusion transcript in secondary glioblastomas. Genome research. 2014;24:1765–1773. doi: 10.1101/gr.165126.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Comprehensive molecular characterization of urothelial bladder carcinoma. Nature. 2014;507:315–322. doi: 10.1038/nature12965. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Guo G, Sun X, Chen C, Wu S, Huang P, Li Z, Dean M, Huang Y, Jia W, Zhou Q, Tang A, Yang Z, Li X, Song P, Zhao X, Ye R, et al. Whole-genome and whole-exome sequencing of bladder cancer identifies frequent alterations in genes involved in sister chromatid cohesion and segregation. Nature genetics. 2013;45:1459–1463. doi: 10.1038/ng.2798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Capelletti M, Dodge ME, Ercan D, Hammerman PS, Park SI, Kim J, Sasaki H, Jablons DM, Lipson D, Young L, Stephens PJ, Miller VA, Lindeman NI, Munir KJ, Richards WG, Janne PA. Identification of recurrent FGFR3-TACC3 fusion oncogenes from lung adenocarcinoma. Clin Cancer Res. 2014;20:6551–6558. doi: 10.1158/1078-0432.CCR-14-1337. [DOI] [PubMed] [Google Scholar]
- 31.Wang R, Wang L, Li Y, Hu H, Shen L, Shen X, Pan Y, Ye T, Zhang Y, Luo X, Zhang Y, Pan B, Li B, Li H, Zhang J, Pao W, et al. FGFR1/3 tyrosine kinase fusions define a unique molecular subtype of non-small cell lung cancer. Clin Cancer Res. 2014;20:4107–4114. doi: 10.1158/1078-0432.CCR-14-0284. [DOI] [PubMed] [Google Scholar]
- 32.Kim Y, Hammerman PS, Kim J, Yoon JA, Lee Y, Sun JM, Wilkerson MD, Pedamallu CS, Cibulskis K, Yoo YK, Lawrence MS, Stojanov P, Carter SL, McKenna A, Stewart C, Sivachenko AY, et al. Integrative and comparative genomic analysis of lung squamous cell carcinomas in East Asian patients. J Clin Oncol. 2014;32:121–128. doi: 10.1200/JCO.2013.50.8556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Majewski IJ, Mittempergher L, Davidson NM, Bosma A, Willems SM, Horlings HM, de Rink I, Greger L, Hooijer GK, Peters D, Nederlof PM, Hofland I, de Jong J, Wesseling J, Kluin RJ, Brugman W, et al. Identification of recurrent FGFR3 fusion genes in lung cancer through kinome-centred RNA sequencing. The Journal of pathology. 2013;230:270–276. doi: 10.1002/path.4209. [DOI] [PubMed] [Google Scholar]
- 34.Comprehensive molecular profiling of lung adenocarcinoma. Nature. 2014;511:543–550. doi: 10.1038/nature13385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Comprehensive genomic characterization of squamous cell lung cancers. Nature. 2012;489:519–525. doi: 10.1038/nature11404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Carneiro BA, Elvin JA, Kamath SD, Ali SM, Paintal AS, Restrepo A, Berry E, Giles FJ, Johnson ML. FGFR3-TACC3: A novel gene fusion in cervical cancer. Gynecologic oncology reports. 2015;13:53–56. doi: 10.1016/j.gore.2015.06.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Xiang L, Li J, Jiang W, Shen X, Yang W, Wu X, Yang H. Comprehensive analysis of targetable oncogenic mutations in chinese cervical cancers. Oncotarget. 2015;6:4968–4975. doi: 10.18632/oncotarget.3212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature. 2015;517:576–582. doi: 10.1038/nature14129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Parish A, Schwaederle M, Daniels G, Piccioni D, Fanta P, Schwab R, Shimabukuro K, Parker BA, Helsten T, Kurzrock R. Fibroblast growth factor family aberrations in cancers: clinical and molecular characteristics. Cell cycle (Georgetown, Tex) 2015;14:2121–2128. doi: 10.1080/15384101.2015.1041691. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Sharma P, Shen Y, Wen S, Yamada S, Jungbluth AA, Gnjatic S, Bajorin DF, Reuter VE, Herr H, Old LJ, Sato E. CD8 tumor-infiltrating lymphocytes are predictive of survival in muscle-invasive urothelial carcinoma. Proceedings of the National Academy of Sciences of the United States of America. 2007;104:3967–3972. doi: 10.1073/pnas.0611618104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Sweis RF SS, Gajewski T. Molecular drivers of the non-T cell-inflamed tumor microenvironment in urothelial bladder cancer. J Clin Oncol. 2015;33 doi: 10.1158/2326-6066.CIR-15-0274. abstract 4511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Spranger S, Bao R, Gajewski TF. Melanoma-intrinsic beta-catenin signalling prevents anti-tumour immunity. Nature. 2015;523:231–235. doi: 10.1038/nature14404. [DOI] [PubMed] [Google Scholar]


