Abstract
Background
The distinct types of hematological malignancies have different biological mechanisms and prognoses. For instance, myelodysplastic syndrome (MDS) is generally indolent and low risk; however, it may transform into acute myeloid leukemia (AML), which is much more aggressive.
Methods
We develop a novel network analysis approach that uses expression of eigengenes to delineate the biological differences between these two diseases.
Results
We find that specific genes in the extracellular matrix pathway are underexpressed in AML. We validate this finding in three ways: (a) We train our model on a microarray dataset of 364 cases and test it on an RNA Seq dataset of 74 cases. Our model showed 95% sensitivity and 86% specificity in the training dataset and showed 98% sensitivity and 91% specificity in the test dataset. This confirms that the identified biological signatures are independent from the expression profiling technology and independent from the training dataset.
(b) Immunocytochemistry confirms that MMP9, an exemplar protein in the extracellular matrix, is underexpressed in AML. (c) MMP9 is hypermethylated in the majority of AML cases (n=194, Welch’s t-test p-value <10−138), which complies with its low expression in AML.
Our novel network analysis approach is generalizable and useful in studying other complex diseases (e.g., breast cancer prognosis). We implement our methodology in the Pigengene software package, which is publicly available through Bioconductor.
Conclusions
Eigengenes define informative biological signatures that are robust with respect to expression profiling technology. These signatures provide valuable information about the underlying biology of diseases, and they are useful in predicting diagnosis and prognosis.
Electronic supplementary material
The online version of this article (doi:10.1186/s12920-017-0253-6) contains supplementary material, which is available to authorized users.
Keywords: Gene expression, Network analysis, Leukemia, Extracellular matrix, Homeobox, Hematological malignancy
Background
Acute myeloid leukemia (AML) is an aggressive type of blood cancer and accounts for 1.2% of cancer deaths in the United States [1]. It is the most common acute leukemia, which is characterized by the rapid growth of immature white blood cells. These cells interfere with the production of normal blood cells in the bone marrow. Without treatment, AML can lead to death within months after diagnosis [2]. Myelodysplastic syndrome (MDS) are a set of less aggressive diseases; however, about 30 to 40% of MDS cases can transform into AML [3]. Therefore, it is critical to delineate the exact mechanisms of this transformation [4].
Possible molecular mechanisms include genetic mutations [5, 6], chromosomal abnormalities [7], and epigenetic changes [8, 9]. For example, mutation and abnormal expression of mRNA splicing genes such as SRSF2 [10] and SF3B1 [11] are associated with the prognosis of MDS. Overexpression of Bcl-2 increases resistance of MDS cells to apoptosis [12], and it can play a role in the transformation into leukemia [13]. Similarly, the abnormal expression of some miRNAs such as miR-125 and miR-155 can lead to aberrant self-renewal of HSC [14], a characteristic of AML.
Although investigating the differences between AML and MDS at the molecular level has provided valuable insight, the research in this area has only scratched the surface of the problem. In particular, the current knowledge is far from adequate for the development of strategies for preventing or predicting the transformation of MDS into AML [9]. Researchers have proposed gene expression profiling as a systematic approach to explore the biology and clinical heterogeneity of MDS.
Most notably, Microarray Innovations in Leukemia (MILE), an international research consortium, assessed the clinical utility of gene expression profiling for the diagnosis and classification of leukemia subtypes [15, 16]. They investigated 3334 leukemia patients, including 202 AML with normal karyotype (AML-NK) and 164 MDS cases in their study, and they developed a classifier to distinguish MDS from AML. While their classifier could correctly predict 93% of AML cases from expression profiles, it failed to identify half of MDS cases [16]. This emphasized the heterogeneity of MDS and underlined the need for more sophisticated approaches for analyzing expression profiles. Specifically, the following challenges limited the performance of the classification:
The classifier was based only on the 100 most differentially expressed genes. However, the biological processes in a hematopoietic cell often depend on the coordination of many more genes. Because the status of the cell is determined by the level of expression of hundreds of transcripts, restricting the analysis to only 100 genes could decrease the statistical power to a great extent [17]. Also, a random gene might be considered differentially expressed due to biological or technical noise or due to the difference in the analyzed cell types. Such a gene would convolute a classification based on differentially expressed genes [18].
The produced data were inconsistent because of multiple platforms and approaches used across different institutions [9]. For instance, if a signature was defined using the level of expression in a microarray dataset, it would be very challenging to interpret and use that signature in an RNA-Seq dataset produced in a different laboratory [19].
We hypothesized that gene network analysis addresses both of the above challenges because it models the interactions between genes in a comprehensive structure [20, 21] (Additional file 1: Note S1). Recently, Liu reviewed the computational methods that employ a gene network approach to identify biomarkers from high-throughput data [22]. Gene networks provide a systematic way to organize complex data, and to identify biomarkers that are useful in improving diagnosis, prognosis and therapy of diseases.
To address the above-mentioned challenges in analysis of expression profiles, we developed Pigengene, a novel methodology that is inspired by—and builds upon—coexpression network analysis and Bayesian networks. Briefly, we identify gene modules using coexpression network analysis [23]. We summarize the biological information of each module in one eigengene using principal component analysis (PCA) [24]. Our approach is fundamentally different from applying PCA directly on the entire expression profile, which can lead to significant loss of information [25]. We innovatively use eigengenes as biological signatures (features) to identify the mechanisms underlying the disease. For instance, we use eigengenes to train a Bayesian network that models the probabilistic dependencies between all modules. Alternatively, we infer a decision tree to predict the disease type based on eigengenes. The main idea of our methodology is illustrated in Fig. 1.
Fig. 1.
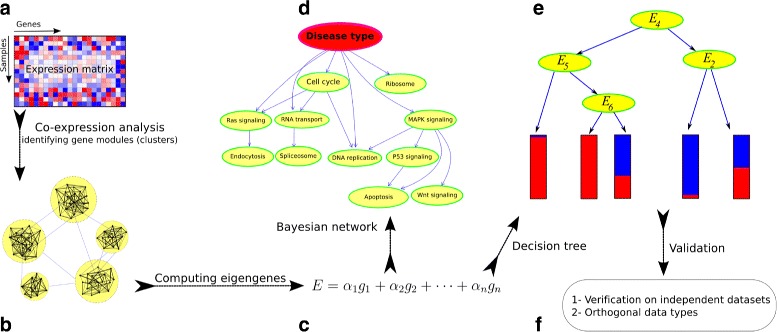
Schematic view of the Pigengene methodology. a The input is a gene expression profile (matrix) provided by RNA-Seq or microarray. b The coexpression network is built according to the correlation between gene pairs. c For each module, an eigengene is computed as a weighted average of the expression of all genes in that module. d Optionally, a Bayesian network is fitted to the eigengenes to delineate the relationships between modules. e A decision tree is fitted to the eigengenes and used for classification. f The results are validated on independent expression datasets and also evaluated using other data types. For instance, DNA methylation profiles can confirm gene-silencing events [43]
We used our methodology to classify patients in the MILE dataset. The accuracy of our model reached 95% for AML and 86% for MDS thus significantly outperforming the previously reported accuracy of 93 and 50%, respectively [16] (Table 1). To show the generalizability of the proposed approach, we report the results of applying it to several cohorts of breast cancer.
Table 1.
The confusion matrices show the accuracy of our decision tree on the training (MILE) and test (BCCA) datasets
| Dataset | MILE (train) | BCCA (test) | ||
|---|---|---|---|---|
| Disease | AML-NK | MDS | AML-NK | MDS |
| Full tree (155 genes) | 191 (95%) | 141 (86%) | 51 (98%) | 20 (91%) |
| Reduced tree (14 genes) | 181 (90%) | 137 (84%) | 51 (98%) | 20 (91%) |
| Mills et al. [16] | 188 (93%) | 82 (50%) | ||
| Reference diagnosis | 202 | 164 | 52 | 22 |
The percentages of correctly identified cases with respect to the reference diagnosis are shown in parentheses. Compared to Mills et al., our decision tree is 36% more sensitive to MDS. The sensitivity to AML is comparable in both approaches
Results
We identified 33 gene modules as clusters of genes that are coexpressed in the 202 AML cases from the MILE dataset [23] (Additional file 1: Note S2). The sizes of the modules vary in the range of 21 to 888, with a mean and median of 153 and 75, respectively (Additional file 1: Figure S1).
Analysis of gene modules
Overrepresentation analysis reveals that some of the modules are associated with canonical pathways and biological processes. For instance, module 6 is enriched with genes that are related to the cell cycle. That is, out of 421 genes in the Reactome cell cycle pathway [26], 81 (19%) are grouped in module 6, which consists of 255 genes (p-value of the hypergeometric test <10−37). Similarly, module 12 is associated with extracellular matrix, module 14 with cytotoxic pathway (CD8+ T cells), module 15 with DNA replication, and module 21 with translation (Additional file 1: Figures S2 and S3 and Additional file 2: Table S2).
Module 33 is the smallest module containing 21 genes. We named it HIST1 because almost all of its genes (20, 95%) encode proteins from the linker histone, or H1, family (Additional file 3: Table S1). Half of the 39 genes in module 28 are from the homeobox family. Considering that this module contains 10 HOXA and 9 HOXB genes, we named it HOXA&B module. It is highly enriched with the homeobox genes that have been reported to be associated with the development and prognosis of AML [27, 28] (Additional file 1: Figure S4, Additional file 3: Table S1 and Additional file 4: Table S3).
Eigengenes are associated with the disease
We summarized the biological information of each module in one eigengene (Additional file 5: Table S4). An eigengene of a module is a weighted average of expression of all genes in that module. The weights were adjusted such that the loss in the biological information is minimized (Methods) [24, 29]. In the MILE dataset, all module eigengenes present significantly different expression in AML vs. MDS. The adjusted Welch’s t-test p-values are in the range of 10−61 to 10−6, with a median of 10−24 (Additional file 1: Figure S5) [30].
We hypothesized that the eigengenes are important biological signatures that can predict the disease type solely based on gene expression. To validate this hypothesis, we developed an innovative approach to infer the values of these eigengenes using the RNA-Seq data from an independent dataset produced at the British Columbia Cancer Agency (BCCA) (Methods). We used this approach to investigate the patterns common in the MILE and BCCA datasets. Interestingly, eight eigengenes achieve significant p-values (<0.01, Bonferroni adjusted) on both BCCA and MILE datasets, indicating that these biological signatures are independent from the profiling platform (Fig. 2 and Table 2). The eight differentially expressed modules include module 28 (HOXA&B), module 21 (translation), module 12 (extracellular matrix), and module 14 (CD8+ T cells).
Fig. 2.
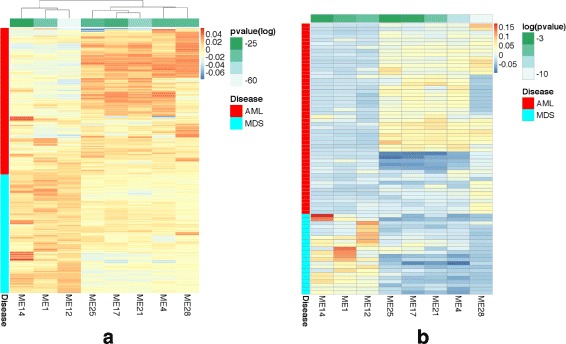
The eight eigengenes that are differentially expressed in both datasets. We computed an eigengene for each module as a weighted average of the expression of all genes in that module. The intensity of the colors in each heatmap corresponds to the normalized average expression. a Each column corresponds to an eigengene. Each row shows the expression of a case from the MILE microarray dataset. AML cases were first clustered and then plotted together with MDS cases for comparison. Three (five) eigengenes have clearly higher expression in MDS (AML) cases. Table 2 reports the adjusted p-values, shown in the green strips at the top. b The corresponding expression values are similar in the BCCA RNA-Seq dataset, which indicates that these biological signatures are robust and independent from the profiling platform
Table 2.
These eigengenes were differentially expressed in AML vs. MDS cases
| Module | 1 | 4 | 12 | 14 | 17 | 21 | 25 | 28 |
|---|---|---|---|---|---|---|---|---|
| P-value (MILE) | 10−37 | 10−32 | 10−61 | 10−23 | 10−28 | 10−43 | 10−32 | 10−33 |
| P-value (BCCA) | 10−3 | 10−5 | 10−3 | 10−2 | 10−3 | 10−7 | 10−3 | 10−11 |
They had adjusted p-values (Welch’s t-test) less than 0.01 in both the MILE and BCCA datasets
We fitted a Bayesian network to the eigengenes to determine the relationships of the modules with each other and with the type of hematological malignancy (Additional file 1: Note S3) [31]. Descendants of the “Disease” node, the variable that models the type of malignancy, are enriched with genes known to be associated with AML (Fig. 3). The relatively high dependency between these eigengenes and the disease type suggests that they have useful biological information that can explain the differences between the two diseases.
Fig. 3.
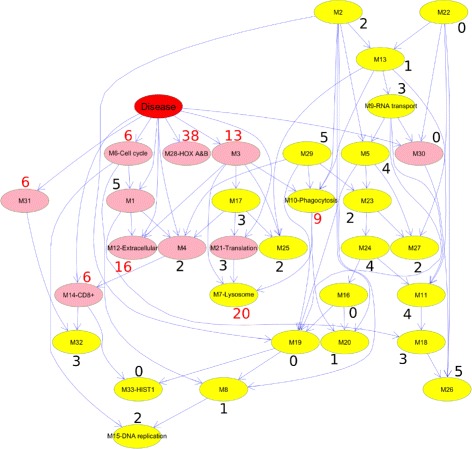
The Bayesian network fitted to the eigengenes. Each node represents an eigengene of a module. The arcs model the probabilistic dependencies between the modules [86]. The “Disease” node is set to 1 for AML and 0 for MDS, and its children are highlighted in pink. Some modules are labeled based on their association with a biological process or a pathway (Additional file 2: Table S2). We used Miller et al., survey to identify the 427 genes reported to be associated with AML in at least three studies [81] (Additional file 3: Table S1). For each module, the percentage of AML-related genes is noted. The percentages that exceed 5% are shown in red. As expected, most of the children of Disease are enriched in genes known to be associated with AML. Specifically, the average of percentages over the children of the Disease node is 10%, which is twice the average of all modules (5%). Also, hypergeometric tests showed that modules 3, 7, 12, and 28 are statistically enriched with AML-related genes (Bonferroni adjusted p-values are 10−7, 10−13, 10−3, and 10−8, respectively). All four of these modules are descendants of the Disease node
AML and MDS are different in their expression of extracellular matrix, HOXA, and HOXB genes
We fitted a decision tree to the eight children of the Disease node in our Bayesian network (R package C50 version 0.1.0-24) [32]. We used only MILE data to infer the topology of the tree and the corresponding parameters. The algorithm automatically selected the extracellular matrix and HOXA&B eigengenes (modules 12 and 28, respectively). The inferred decision tree had high predictive accuracy (Fig. 4). Specifically, 191 AML-NK cases (95%) and 141 MDS cases (86%) were correctly identified (Additional file 6: Table S5).
Fig. 4.
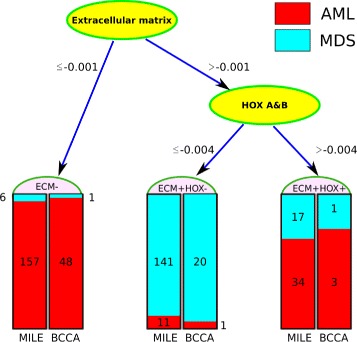
A simple decision tree for distinguishing AML from MDS cases. If the normalized extracellular matrix eigengene of a case is less than -0.001, it is classified as AML. Otherwise, the HOXA&B eigengene determines whether the case is AML (>−0.004) or MDS (≤−0.004). The number of cases classified in each leaf is noted for both the MILE (left) and the BCCA (right) datasets. Only the middle leaf corresponds to MDS. At the fixed thresholds shown above, this tree correctly classified 328 cases (90%) in the MILE dataset (the training set) and 71 cases (96%) in the BCCA dataset (the test set)
The majority of AML cases (157, 78%) were identified because of their low expression of extracellular matrix genes (i.e., their normalized eigengene value was less than −0.001). For the rest of the cases, which expressed the extracellular matrix eigengene, the tree considered the expression of the HOXA&B eigengene. If it was over −0.004, the case was classified as AML. The tree shows that for a case to be MDS, it must have relatively high expression of the extracellular matrix (Fig. 5 and Additional file 1: Figure S6) and low expression of HOXA&B (Fig. 6 and Additional file 1: Figure S7).
Fig. 5.
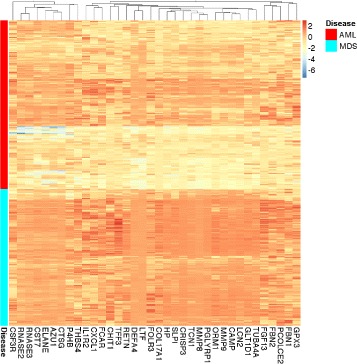
Comparing the expression of extracellular region genes. Each column shows the expression of a gene from the extracellular matrix module that is associated with the “extracellular region” in the cellular component category of Gene Ontology (GO). For clarity, each column is scaled by subtracting its mean and dividing by its standard deviation. Each row corresponds to a sample from the MILE dataset. These 36 genes are generally underexpressed in AML compared to MDS. The expression of all 133 genes in the extracellular matrix module have a similar pattern (Additional file 1: Figure S6)
Fig. 6.
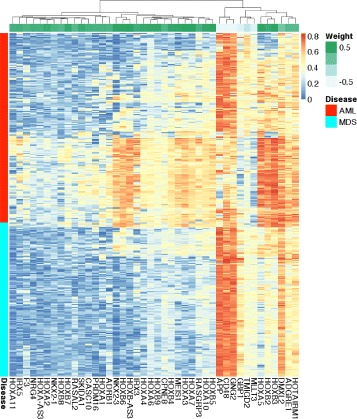
Comparing the expression of genes in the HOXA&B module. Expression of every member of HOXA&B module is shown in one column. Each row corresponds to a sample from the MILE dataset. The majority of HOXA and HOXB genes in this module are not expressed in MDS. Their expression in AML are variable indicating the heterogeneity of the disease. They anticorrelate with GNG2, CD48, and APP, which have the least negative weight (-0.7) in the corresponding eigengene (the green strip at the top). These patterns are similar in the BCCA dataset (Additional file 1: Figure S7)
Misclassification of MDS was associated with risk factor
The International Prognostic Scoring System (IPSS) score [3] is the standard tool for MDS risk stratification [33]. It ranges from 0 to 3.5, and a higher value indicates a poorer prognosis. There are 30 MDS cases (18%) in the MILE dataset with poor prognosis (IPSS ≥1.5). This set has a significant overlap with the 23 cases “misclassified” by our decision tree (Additional file 6: Table S5). Specifically, 15 MDS cases with poor prognosis show AML signatures and are classified as AML by the tree (hypergeometric test p-value <10−7). This suggests that underexpression of the extracellular genes and overexpression of the HOXA genes in an MDS case can be considered as a risk factor. Because transition into AML is more likely for such an MDS case, a monitoring assay can be developed based on these signatures.
Validating AML signatures in an independent dataset
We validated the performance of the tree on classifying 74 cases in the BCCA dataset. To this end, we inferred the values of extracellular matrix and HOXA&B eigengenes in the BCCA dataset (Methods). With the same above-mentioned thresholds that performed well for the MILE dataset, the tree correctly identified 51 (98%) of the AML-NK and 20 (91%) of the MDS cases. The high accuracy of our decision tree was helpful in correcting a clerical error in annotating the dataset. In particular, two BCCA cases (B118 and B129), originally labeled with MDS, have signatures very similar to AML (Additional file 5: Table S4). Interestingly, a second review revealed that their correct diagnosis is in fact tAML (therapy–related AML) and AML–M1, respectively.
Although the decision tree was trained using only AML-NK subtype in the MILE dataset, its performance in differentiating some other subtypes of AML from MDS in the BCCA dataset is remarkable. In particular, all of the four AML-t(8;21) cases (100%), all of the four AML cases with complex karyotype cases (100%), all of the four AML cases with 11q23 abnormality (100%), and 9 out of 11 AML-inv(16) cases (82%) are all correctly classified as AML. However, cases from other subtypes, such as AML-t(15;17), AML-M6, and tAML, do not always show strong extracellular or HOXA&B signatures of AML-NK and are frequently misclassified as MDS (Additional file 6: Table S5). This is expected, because these three subtypes of AML are distinct and too different from AML-NK. In particular, leukemic cells in AML-t(15;17) and AML-M6 are relatively more differentiated [34], and may produce some extracellular matrix proteins.
A minimal gene set for clinical testing
Considering the good performance of the decision tree, it is useful to develop a clinical test based on gene expression. The extracellular matrix and HOXA&B modules contain 113 and 42 genes, respectively. To infer the corresponding eigengenes, the expression of 155 genes are needed in total. If the number of genes is reduced without significant loss of accuracy, the test will be easier to use in clinical settings. Because the genes are correlated with each other in each module, shrinking the tree is expected to have little—or no—effect on the accuracy of classification.
Using a greedy approach, we excluded the majority of the 155 genes, and obtained a decision tree that need the expression values of only 14 genes (9%) (). The performance of the reduced tree is comparable to the original tree (Table 1). On the training set, the accuracy dropps by only 5% for AML and by 2% for MDS. On the test set, however, the reduced tree is as accurate as the full tree (Additional file 6: Table S5).
The list of 14 genes used in the reduces tree included PGLYRP1, MMP9, CEACAM6, ARG1, MMP8, ANXA3, RGL4, SLPI, HP, CEACAM1, MGAM, SYNE1 from the extracellular matrix module, and HOXB-AS3 and HOXA3 from HOXA&B module (Fig. 7).
Fig. 7.
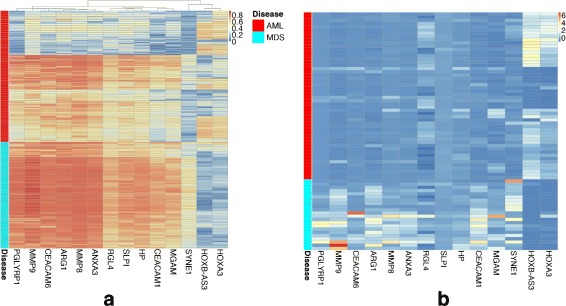
Comparing the expression of the 14 genes used in the reduced tree. Expression of the 14 genes that contribute to the reduced (compact) decision tree in the a MILE and b BCCA datasets are shown. The 12 extracellular matrix and the two HOXA genes are generally silenced and overexpressed in AML, respectively
The significance of the extracellular matrix pathway in AML
The relationship between HOX genes and AML and their role in leukemogenesis are extensively studied [27, 27, 28]. Researchers have also explained the significance of the extracellular matrix pathway in the prognosis of cancers in general [35]. However, its role in the development of AML and other leukemias is more complicated. In addition to regulating cell growth [36], proliferation [37], differentiation [38], and apoptosis [39], it also mediates the migration of hematopoietic stem cells through the vessels [40]. Module 12 is enriched with extracellular matrix genes (Additional file 1: Figure S8). We investigated these genes, which defined a significant signature in our decision tree (Fig. 4).
Gene Ontology Cellular Component (GO-CC) analysis showed that 36 of 113 genes in module 12 code for proteins in the extracellular region (Additional file 1: Figure S8 and Additional file 7: Table S7). Moreover, 77 of the genes in this module are associated with at least one of the following categories: extracellular vesicular exosome (44 genes), extracellular region (36), extracellular space (30 genes), and plasma membrane (31 genes). We noted that 18 genes (16%) are located on chromosome 19. Almost all of these 113 genes are underexpressed in AML (Fig. 5 and Additional file 1: Figure S6). The enriched biological processes include: immune system process (adjusted p-value <10−9), killing by host of symbiont cells (<10−3), killing of cells in other organism involved in symbiotic interaction (<10−2), defense response to fungus (<10−3), antibacterial humoral response (<10−2), extracellular matrix disassembly (<10−2), and response to lipopolysaccharide (<10−2) (Additional file 8: Table S9) [41].
One particularly interesting gene from this module was MMP9, which had a relatively high contribution to the eigengene. Its weight is 0.92, the highest in the extracellular matrix pathway (Reactome [42]), and the eighth in the module (Additional file 7: Table S7). MMP9 is a member of the matrix metalloproteinase (MMP) family, which has 23 members.
They remodel and degrade the extracellular matrix by cleaving its components [42]. In addition to MMP9, this module includes two other members of MMP family, namely MMP8 (weight = 0.91) and MMP25 (weight = 0.87). All of these three genes are underexpressed in AML (Additional file 1: Figure S9a). One way to confirm that these genes are silenced in AML would be to check epigenetic factors such as DNA methylation, which generally anticorrelates with gene expression [43]. We compared 194 AML cases of Acute Myeloid Leukemia (LAML) dataset from The Cancer Genome Atlas (TCGA) with 368 control cases, and we confirmed that these three genes were heavily methylated in AML (Additional file 1: Figure S9b and Additional file 9).
Validating gene expression changes at the protein level
Given the strong discriminating capability of extracellular matrix gene expression in differentiating AML from MDS (Fig. 4), we attempted to determine whether a simple immunohistochemical stain would provide such a differentiation. We selected MMP9 to test this, as it provided the highest-weighted contribution to the eigengene (0.92) within the extracellular matrix set of genes. We obtained 10 previously diagnosed AML cases and 10 previously diagnosed MDS cases, and performed immunostaining on the diagnostic bone marrow biopsies. As seen in Fig. 8, MMP9 staining is drastically lower in the AML samples compared to the MDS cases.
Fig. 8.
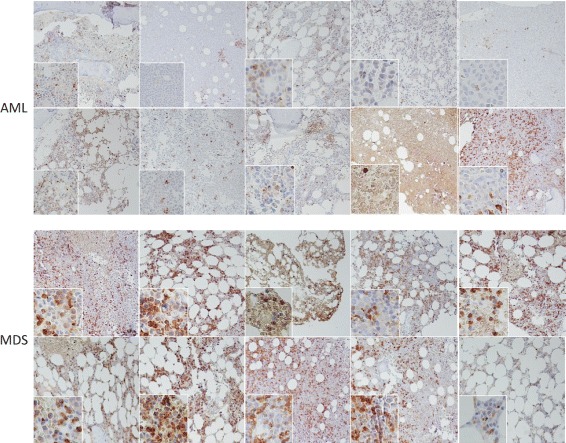
Expression of MMP9 in AML and MDS. The bone marrow of 10 patients with AML (upper panels) and 10 patients with MDS lower panels) was immunostained in parallel with an MMP9 antibody. MMP9 expression is reduced to absent in AML blasts. Where staining is seen in AML, it is only present in mature myeloid cells but not leukemic cells
Validating the identified coexpression pattern in other AML-related datasets
The 113 genes in the extracellular matrix module are correlated and underexpressed in AML. To validate that the observed coexpression pattern is specifically associated with AML, we investigated the expression of these 113 genes in a large collection of human datasets. Specifically, we used Search-Based Exploration of Expression Compendium (SEEK) [44] to objectively compare the coexpression of these genes across a collection of 5210 datasets. SEEK automatically scored and ranked the datasets based on the significance of coexpression of our 113 genes. SEEK also computed empirical p-values to assess the statistical significance of scores. Specifically, random scores for each dataset was computed based on 5000 queries of 113 random genes, and a p-value was reported as the fraction of random scores that were higher than the reported score. The collection contains 61 AML-related datasets (1.2%), which mostly score high in the ranked list (Additional file 10: Table S8). In particular, all of the five top datasets are related to AML (GEO accession numbers: GSE15434 [45], GSE16015 [46], GSE12417 [47], GSE21261 [48], and GSE30599 [49]; with 251, 107, 405, 96, and 29 samples, respectively). The coexpression scores are 0.31, 0.29, 0.28, 0.28, and 0.27, respectively; and the adjusted empirical p-values are smaller than 10−37 for each of these five datasets. A hypergeometric test confirmed that the coexpression of the queried genes is significantly associated with AML (p-value <10−9). Thereby, our unbiased and objective SEEK analysis indicates that these genes define an expression signature that is specific to AML.
Generalizability to studying other cancers
The described pipeline can also be applied to analyze other types of cancers and answer different biological questions. To demonstrate this, we applied our approach to a prognostic question in breast cancer research. Is it possible to identify low-risk breast cancer cases based solely on gene expression and thereby avoid overtreating a subset of patients who likely would not benefit from the additional toxic therapy [50]? In this type of prognostic setting, the emphasis lies on achieving very high specificity for predicted low-risk cases. For instance, the TRANSBIG Consortium [51] considers a test to be clinically practicable and reliable for ER+ breast cancer only if at least 88% of cases classified as low-risk have more than a 10-year overall survival. However, the only clinical test with such high precision is Oncotype DX, which is applicable to only one clinical subtype of breast cancer, stage I ER+ tumors [50]. Unfortunately, this method cannot be generalized to other breast cancer subtypes [52].
We analyzed 1374 ER+ cases from three datasets to train and validate our model (Methods and Additional file 1: Note S4). The low-risk specificity of our model is above 89% for all three datasets (Table 3 and Additional file 1: Figure S11). Two modules with 319 and 193 genes, respectively, were automatically selected (Additional file 11: Table S6). The larger module is associated with the mitotic cell cycle (p-value <10−71) and chromosome segregation (p-value <10−28). This module has 16 genes in common with the genes in the PAM50 assay, which is widely used in clinical settings to identify breast cancer subtypes [53]. These common genes include UBE2T, BIRC5, CCNB1, CEP55, MELK, UBE2C, CENPF, PTTG1, EXO1, ANLN, CCNE1 CDC20, MKI67, KIF2C, MAPT, and FGFR4. This is a significant overlap (p-value of the hypergeometric test <10−10).
Table 3.
Accuracy of predicting breast cancer risk
| Dataset | METABRIC discovery | METABRIC validation | MILLER (test) | |||
|---|---|---|---|---|---|---|
| Low risk | High risk | Low risk | High risk | Low risk | High risk | |
| Predicted low risk | 157 (94%) | 11 (7%) | 107 (89%) | 13 (11%) | 68 (93%) | 5 (7%) |
| Predicted medium risk | 278 (68%) | 134 (33%) | 236 (70%) | 99 (30%) | 55 (68%) | 26 (32%) |
| Predicted high risk | 21 (35%) | 39 (65%) | 33 (42%) | 46 (58%) | 29 (62%) | 18 (38%) |
The confusion matrices show the performance of our decision tree on three datasets. The percentage of predicted cases with respect to the total number of predictions in each group is shown in parentheses. From a clinical standpoint, it is important to achieve a high precision (positive predictive value) for low risk cases (shown in bold) to confidently recommend a less agressive treatment regimen for a subset of patients. The probability of surviving more than 10 years is above 89% for the predicted low risk cases in all the three datasets (Additional file 1: Figure S11)
The smaller module is associated with translational control (Additional file 1: Figure S12). The expression of the majority of the genes (122, 63%) is correlated with poor prognosis. Notable genes include AKT1, GSK3B, MTOR, RAF1, and SRC from the epidermal growth factor receptor (ErbB) signaling pathway [54]. In contrast, the high expression of 71 genes (37%) in this module—including 16 ribosome-related genes such as RPL22, RPL26, RPS27, RPS27A, RPL13A, RPL21 and RPLP0—correlate with good prognosis. This may be predicted, as the loss of function or abnormal expression of proteins involved in ribosomal biogenesis is associated with activation of the tumor suppressor p53 pathway [55, 56]. A possible mechanism of p53 activation could be through binding free (non-ribosome-bound) ribosomal proteins with MDM2, which modulates the inhibitory activity of MDM2 on p53 [55].
None of the 193 genes from the smaller module is in common with PAM50. This suggests that the corresponding eigengene can be considered as a novel biological signature to assess breast cancer prognosis, and it can be a basis for improving clinical tests. Overall, our model is biologically plausible because regulated cell cycle and controlled translation are generally associated with better prognostic outcome [57].
Discussion
Biological processes in a cell often require coordination between multiple genes and proteins, not just one gene or a single protein. Accordingly, we used network analysis to delineate the differences in gene expression profiles of AML and MDS in a systematic and robust way (Additional file 1: Note S1). We compared the expression at the module level to minimize the effect of artifacts such as a random change in expression of an isolated gene and other biological or technical noise (Additional file 1: Figure S10).
The results of our study underline the association of the extracellular matrix pathway with AML, and also confirm that the overexpression of homeobox genes is a biological characteristic of AML. These two signatures are biologically related [58]. Homeobox genes encode transcription factors that regulate the development of body structures during the embryonic period [59]. They also have key roles in adult tissue remodeling and pathogenesis [60]. In particular, specific homeobox genes can regulate the extracellular matrix through the expression of matrix-degrading proteinases [61]. For instance, the expression of the HOXA3 and HOXB3 are upregulated during wound healing to remodel the extracellular matrix and to increase endothelial cell migration [62]. Overexpression of HOXA7, which is associated with poor prognosis of AML [63], can modify the interactions between hematopoietic progenitor cells and the extracellular matrix in the bone marrow. This alteration can be responsible for blocking the differentiation process in AML cells [64].
The two signatures are highly associated with AML to such a degree that they can be used to design an accurate clinical test for differential diagnosis between AML and MDS. Furthermore, the following confirmatory evidence supports our findings on the significance of the extracellular matrix pathway in AML:
Our decision tree can accurately predict the diagnosis in a validating dataset (BCCA) without the need to change the parameters that were fitted to the training dataset (MILE). Our results confirm that the model was not overfitted to the training dataset.
SEEK analysis confirms that the genes in the extracellular matrix module are coexpressed in several other AML-related datasets.
The three MMP genes in the extracellular matrix, MMP9, MMP8, and MMP25, are methylated in AML.
Immunocytochemistry showed that MMP9 is underexpressed in AML at the protein level.
MMP9 is an important gene in our analysis, and it has a distinct expression profile between the two diseases. MMP9 acts as a cell surface transducer by cleaving the extracellular matrix and other proteins, including chemokines, cytokines, and growth factor receptors. In this way, it can regulate key signaling pathways in cell growth, migration, invasion, inflammation, and angiogenesis [65]. While MMP9 was previously reported to have a critical role in AML invasion and metastasis [66–69], the relationship between its expression and the prognosis of hematological malignancies is complicated. For instance, Aref et al. report that 43 pretreatment AML cases had significantly lower expression of MMP9 as compared to 10 controls. However, after chemotherapy, MMP9 was expressed significantly higher in relapsed cases as compared to complete remission cases [70].
In this context, the high expression of MMP9 in MDS, which we showed is more than AML, is interesting. Correspondingly, Travaglino et al. measured MMP2 and MMP9 in myeloid cells of 143 MDS cases using immunocytochemistry. They found that high MMP levels are associated with longer overall survival [71]. One possible interpretation is that by deregulating the extracellular matrix, MMP9 may interrupt the survival signalling in MDS and lead to apoptosis. In contrast, lowering MMP9 expression may prolong the life of the MDS cells and facilitate the transition into AML. MMP9 processing of the matrix may also have an impact on blast cell invasion, dissemination, and homing [70]. However, functional studies will be needed to determine the mechanism and impact of MMP9 on myeloid cancers. A competing theory would be that the observed differences in the extracellular matrix activity might be due to differences in the underlying cell-types.
Our approach has novel methodological contributions to gene expression analysis. While other scholars have used weights (loadings) of eigengenes to study genes in a module [24], we are the first to use values of eigengenes directly as biological signatures. We developed an approach to infer and compare eigengenes across datasets. Our approach is fundamentally different from applying PCA directly on the entire expression profile, which is not a promising approach because the first few PCs may not have enough information on the modules’ structure [25].
An analysis based on a limited number of genes with the best p-value can be convoluted by random, dramatic expression changes due to biological or technical noise [17]. In contrast, because an eigengene is a weighted average expression of several genes, our systematic and holistic approach is much more robust than the alternative approaches that select one or a few genes from each module [72, 73]. We show that our methodology is generalizable and useful in studying other malignancies by applying it to several breast cancer datasets.
Conclusions
Eigengenes are robust informative biological signatures. They are useful in predicting the diagnosis and prognosis, and also, in delineating the molecular characteristics of diseases. For instance, we used large-scale network analysis to show that underexpression of particular genes in the extracellular matrix pathway is a specific characteristic of AML.
Methods
The AML gene expression datasets
We downloaded the expression profiles of 202 AML-NK and 164 MDS cases from Gene Expression Omnibus (GEO) (series number GSE15061) [16], Additional file 12. The dataset is part of the expression MIcroarray analysis for diagnosis of LEukaemia (MILE) series. For simplicity, we refer to this expression profile as the MILE dataset, which was used to train our model. To validate our model and test the accuracy of classification, we used RNA-Seq data from 133 AML and 22 MDS cases analyzed at British Columbia Cancer Agency. For simplicity, we refer to this expression profile as the BCCA dataset, which is independent from the MILE dataset. From the 133 AML cases, 52 were AML-NK and thus were comparable with the 202 cases from the MILE dataset (Additional file 6: Table S5). We used Sailfish (version 0.6.3) [74] to compute reads per kilobase per million mapped reads (RPKM) values [75] for each gene, and considered the natural logarithm of RPKM to measure gene expression.
Breast cancer datasets
We used 640 ER+ cases from the Molecular Taxonomy of Breast Cancer International Consortium (METABRIC) [76] discovery dataset for training. We evaluated the resulting model on 533 different cases from the METABRIC validation dataset. We also validated the prognostic value of the inferred biological signatures using 201 cases from a second independent dataset produced by Miller et al. [77] (GEO accession number GSE3494). The details of our analysis on these three datasets is presented in Additional file 1: Note S4.
Detailed description of the Pigengene methodology
Preprocessing The input to the Pigengene methodology includes two gene expression profiles corresponding to two biological conditions (e.g., AML and MDS in this paper). Optionally, the user can provide a validating dataset (e.g., BCCA dataset). The train and validation datasets do not need to be assayed using the same platform. Thas is, one dataset can be microarray and the other one can be RNA-Seq. Figure 1 shows the main steps of the Pigengene methodology. More specificity, the first step of the analysis is to exclude the genes that have too little variation or negligible expression across the conditions. This can be done using a differential expressed analysis, which computes a p-value for each gene with the null hypothesis that it is similarly expressed in the two conditions. Consistent with the common approach in the gene network analysis [78, 79], we kept only the top one-third genes with the best p-values in our analysis.
Constructing the coexpression network: We used the WGCNA package to construct a coexpression gene network, in which each node (vertex) is a gene and the edge (connection) between two genes is weighted based on the correlation between their expression values (Additional file 1: Note S2). WGCNA uses a hierarchical clustering approach to identify gene modules from the coexpression network.
Computing eigengenes: We used principal component analysis (PCA) to compute an eigengene for each module. First, we balanced the number of AML and MDS cases using oversampling, so that both disease types had comparable representatives in the analysis. Specifically, we repeated the data of each AML and MDS case 9 and 11 times, and obtained 1818 and 1804 samples from each type, respectively. Then, we applied the moduleEigengenes() function from the WGCNA package on the oversampled data. We ran it with the default parameters, and computed an eigengene for each of the modules identified earlier. This function computed the first principal component of each module, which maximized the explained variance ensuring the loss in the biological information was minimized. [24, 29] (Additional file 5: Table S4).
Inferring the decision tree: We use eigengenes as features to infer a decision tree (R package C50 version 0.1.0-24) [32]. While the C50 package uses a heuristic approach to select the best set of features, its default arguments does not result in optimal performance when too many features are provided. The solutions include: 1) using a Bayesian network to determine the relationships of the modules with each other and with the type of hematological malignancy (Additional file 1: Note S3) [31], 2) using a feature scoring algorithm such as FeaLect [80], and 3) adjusting the C50 parameters, for example, enforcing the number of samples in each node to be at least 10%. The first and the third solutions are implemented in the Pigengene package through the bnNum argument of the one.step.pigengene() function and the minPerLeaf argument of the make.decision.tree() function, respectively. These two approaches resulted in the same decision tree presented in this paper (Fig. 4).
Inferring the values of eigengenes in an independent dataset: When a validation dataset is available (i.e., the BCCA dataset in our study), the values of the eigengenes need to be inferred in the validation dataset. We computed eigengenes using the MILE dataset, which is a microarray dataset. It was challenging to compute the values of the same eigengenes for BCCA cases because the BCCA dataset was produced using a different platform (i.e., RNA-Seq) [19]. The simple approach of applying PCA on the BCCA data would fail; It would result in different weights (loadings), and the eigengenes would not be comparable between the two datasets. Instead, we inferred the values of the eigengenes for BCCA cases using the same weights obtained from the MILE dataset. Specifically, for each module, we identified the genes that are common in both datasets. Then, we scaled the expression of those genes by subtracting their mean and dividing by their standard deviation. We used the scaled expression values to compute the eigengene (the weighted average of expression) for each BCCA case. The project.eigen() function from our Pigengene package facilitates this approach.
Reducing the number of genes needed for the decision tree: Our decision tree used the eigengenes of HOXA&B and extracellular matrix modules, which were weighted averages of the expression of 42 and 155 genes, respectively. To reduce the number of genes, we repeated the following greedy procedure [72]: We excluded the gene with the lowest absolute weight, inferred the eigengenes using the remaining genes, and used the updated eigengenes as input to the decision tree. In each iteration, we used the same tree structure and thresholds, and we measured the accuracy of classification. We repeated this procedure until the tree needed only 14 genes, because excluding any more genes would result in a significant decline in the accuracy of the classification. The sufficiency of these 14 related genes indicates that they contain the core biological information needed for classification. The compact.tree() function from our Pigengene package facilitates this approach.
Code availability
“Pigengene”, a documented R package that implements our approach, is publicly available through Bioconductor: http://bioconductor.org/packages/Pigengene. The results presented in this paper can be reproduced using version 0.99.19. To apply our methodology in other studies, we strongly recommend using the most recent version. We encourage users to use the Bioconductor mailing list to send bug reports and seek technical help.
Acknowledgements
We would like to thank the GSC sequencing and library generation teams for sequencing and Vennie Chou for immunohistochemistry. We thank Qian Zhu for her assistance in running SEEK. We used DNA methylation data generated by the TCGA Research Network: http://cancergenome.nih.gov/. We acknowledge the Texas Advanced Computing Center (TACC) at The University of Texas at Austin for providing high-performance computing (HPC) resources: http://www.tacc.utexas.edu.
Funding
Work in the lab of AK was funded by grants from the Terry Fox Research Institute (122869), CIHR (MOP-133455, MOP-97744), and Genome BC (121AML). AK is supported by the John Auston BC Cancer Foundation Clinical Investigator award. Work in the lab of HZ was supported by an internal grant from Texas State University.
Availability of data and materials
All data and code required to reproduce the presented results are available, either through publicly available repositories or as supplementary materials. Specifically, the MILE leukemia and Miller breast cancer datasets are available from GEO with accession numbers GSE3494 and GSE15061, respectively, and the METABRIC dataset is available from the European Genome-phenome Archive with study accession number EGAS00000000083. DNA methylation data are available from TCGA (LAML dataset). Additionally, eigengene values for the MILE and BCCA datasets are available as supplementary materials. The Pigengene software package is available through Bioconductor.
Authors’ contributions
AK and HZ conceived the experiments, AF, HZ, RA, and RD conducted the experiments, LC, GD and MH acquired data, and AK analyzed the results. All authors reviewed the manuscript. All authors read and approved the final manuscript.
Competing interests
The authors declare that they have no competing interests.
Consent for publication
Not applicable because no sequencing data or identifying information is being published.
Ethics approval and consent to participate
This study was approved by the University of British Columbia (UBC) BC Cancer Agency Research Ethics Board (UBC-BCCA REB) under protocol H13-02687 “Genomic analysis of molecular changes in myeloid malignancy”. The informed consent of participants was provided prior to specimen acquisition under the guidelines of the Leukemia/Bone Marrow Transplant Program at Vancouver General Hospital, as approved by the UBC-BCCA REB (protocol H04-61292). For historically-banked anonymized specimens (Legacy cell bank specimens), a waiver of consent was provided by the UBC-BCCA REB (protocol H09-01779). This protocol states: Genomic data obtained from these samples may be posted on access restricted sites as required for publications. This is covered by transfer contracts governed by our Technology Development Office. Transfer of material outside of the institution would also be covered by MTAs.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Abbreviations
- AML
Acute myeloid leukemia
- AML–NK AML
With normal karyotype
- BCCA
British Columbia cancer agency
- GEO
Gene expression omnibus
- GO–CC
Gene ontology cellular component
- HSC
Hematopoietic stem cells
- IPSS
Prognostic scoring system
- MDS
Myelodysplastic syndrome
- METABRIC
Molecular taxonomy of breast cancer international consortium
- MILE
Microarray innovations in leukemia
- PCA
Principal component analysis
- RPKM
Reads per kilobase per million mapped reads
- SEEK
Search-based exploration of expression compendium
- TCGA
The cancer genome atlas
Additional files
Supplementary Notes and Figures. (PDF 1656 kb)
Table S2. Overrepresented pathways. The canonical pathways that are overrepresented in the modules are available as part of the online supplementary materials. Each sheet in the excel file corresponds to a module. The statistics for each pathway (gene set) is reported on one row, in particular, the p-value of a hypergeometric test with the null hypothesis that the genes from the pathway were observed in the module by chance. The other columns include the name of the pathway (Set Name), the number of genes observed in the module (In Module), the number of the genes from the pathway that are present in our pool of 9,166 genes (Set Size), percentage of those genes that are in the module (% In Module), the name of the collection in MSIGDB [83] (Source), and a link to more information on the pathway (Description). There was no statistically significant pathway with p-value less than 10−4 for the modules that are not included. (XLS 131 kb)
Table S1. Module assignments. The module assignments for all of the 9,166 genes are available as part of the online supplementary materials. The columns of the table include:
• Symbol: The gene name.
• ENTREZ ID: The gene ID in the Global Query Cross-Database Search System.
• Probe representative: The representative probe mapped to this gene.
• Adjusted p-value: For the null hypothesis that expression is similar in AML and MDS (Supplementary Note 2).
• Module ID: The assigned module.
• IMC: Intramodular connectivity.
• EMC: Extramodular connectivity.
• Rank: The rank of the gene in the module based on its intramodular connectivity.
• # of references The number of studies that reported the gene being relevant to AML according to Miller et al,. survey [81].
Intramodular and extramodular connectivities were computed by intramodularConnectivity() function from WGNCA package. They measure the overall correlation of the gene with other genes within and outside the module, respectively. The genes with the most connectivity in a module are considered as the “hubs” of that module [82]. See WGNCA manual for more information. (XLS 1618 kb)
Table S3. Enrichment in AML-associated genes. The enrichment of modules in genes associated with AML is reported as part of the online supplementary materials. Miller et al,. systematically surveyed 25 published reports of gene expression profiling in AML [81]. We used this survey to score the modules based on their know association with AML. For instance, the # of Reported Genes-2 and Score-2 columns respectively report the number and the percentage of genes in each module that were reported to be related to AML in at least 2 studies according to Miller et al,. survey. The Module ID, Module Size, and the most overrepresented pathway in each module are also reported. These data are plotted in Additional file 1: Figure S4. (XLS 39.5 kb)
Table S4. Eigengenes. The eigengenes computed for all gene modules are available as part of the online supplementary materials. Each row reports the values for a sample and columns correspond to modules. The first sheet was computed using moduleEigengenes function from WGCNA package, which applied PCA on the gene expression in the MILE dataset. The second sheet shows the inferred expression of these 33 eigengenes in the BCCA dataset (Methods). Module zero contains the set of “outlier” genes that did not correlated with each other or with the rest of the genome. WGCNA could not confidently assign them to any module and we did not use them in our analysis. (XLS 334 kb)
Table S5. Classification results. The results of classification of the BCCA and the MILE cases using our decision tree are available as part of the online supplementary materials. The first sheet reports classification results of 176 BCCA patients. For each case sample ID, age, gender and the corresponding cohort are reported. The other columns of the table include:
• Sequenced Tissue: Bone marrow (BM) or peripheral blood (PB).
• RIN: The RNA Integrity Number, a value in the range of 1–10 measuring the quality of the RNA samples [84]. A higher value corresponds to less degradation of RNA.
• Subtype: The AML subtypes were diagnosed based on chromosomal abnormalities and other factors. AML-NK stands for AML with normal karyotype. tAML and tMDS are therapy related.
• Type: All AML subtypes were labeled as AML. MDS and tMDS were labeled as MDS, and 21 gray-highlighted cases that had clinical and pathological characteristics of both diseases were labeled as AML-MDS.
• Prediction (Full Tree) The label predicted by the decision tree using the inferred extracellular matrix and HOXA&B eigengenes. relevant misclassified.
• Prediction (Shrunk Tree) The label predicted by the shrunk decision tree using expression of only 14 genes from the extracellular matrix and HOXA&B modules.
• Signature The leaf of the full decision tree determined based on the expression of the extracellular matrix (ECM) and HOXA&B eigengenes.
Similarly, the second sheet reports classification results of 366 patients from the MILE dataset. Additional information such as IPSS, blast, and karyotype categories are included from MILE study [16]. The misclassified cases by either the full or the shrunk tree are highlighted in yellow. (XLS 155 kb)
Table S7. The extracellular matrix module. For each gene in the extracellular matrix module, the cellular component ontology (GO-CC) is available as part of the online supplementary materials. The genes are ordered according to their weight (also known as “membership” [85]) in the module eigengene, e.g., the first gene has the highest contribution to the eigengene. See the description of Additional file 2: Table S1 for the definition of other columns. (XLS 37 kb)
Table S9. Biological processes. We used PANTHER (Version 10) [41] to identify the GO biological processes that are enriched in the 113 genes in our extracellular module. The columns of the resulting table include the name of GO biological process, the number of genes in the corresponding category (Homo sapiens), the number of overlapping genes, the expected overlap, the fold enrichment, and the Bonferroni-adjusted p-value. (XLS 20 kb)
Supplementary Data 1. DNA-methylation. The DNA methylation of MMP9, MMP8, and MMP25 genes in TCGA dataset are available as part of the online supplementary materials. The two sheets include the patient barcodes for 194 AML and 368 control samples. The β values were reported at cg04656101 (equivalent to chr20:44,645,014 in hg19), cg01092036 (chr11:102,595), and cg02680314 (chr16:3,097,388), respectively (Additional file 1: Figure S9b). (XLS 102 kb)
Table S8. Validating coexpression patterns in 5,210 datasets. The descriptions and names of 5,210 datasets used to perform coexpression analysis are available as part of the online supplementary materials. SEEK sorted the datasets based on their coexpression score computed using the 113 genes from our extracellular matrix module. The AML-related datasets, highlighted in yellow, are frequent at the top of the list. The details of coexpression score and p-value computation are defined in the corresponding publication [44]. (XLS 1044 kb)
Table S6. The cell cycle and translational control modules. Two modules with 319 and 193 genes were automatically selected by breast cancer survival analysis. In each sheet, gene symbols, Entrez IDs, and weights in the corresponding modules are reported. Genes are sorted based on their weights. (XLS 55 kb)
Supplementary Code 1. Geo2R. The Geo2R script, used to compute p-values for probes, is available as part of the online supplementary materials. The process starts by downloading MILE data from GEO. (TXT 5.91 kb)
Contributor Information
Amir Foroushani, Email: a_f240@txstate.edu.
Rupesh Agrahari, Email: r_a157@txstate.edu.
Roderick Docking, Email: rdocking@bcgsc.ca.
Linda Chang, Email: lchang@bccrc.ca.
Gerben Duns, Email: gduns@bcgsc.ca.
Monika Hudoba, Email: monika.hudoba@vch.ca.
Aly Karsan, Email: akarsan@bcgsc.ca.
Habil Zare, Email: zare@txstate.edu.
References
- 1.Jemal A, Thomas A, Murray T, Thun M. Cancer statistics, 2002. CA: Cancer J Clin. 2002;52(1):23–47. doi: 10.3322/canjclin.52.1.23. [DOI] [PubMed] [Google Scholar]
- 2.Longo DL, Döhner H, Weisdorf DJ, Bloomfield CD. Acute myeloid leukemia. N Engl J Med. 2015;373(12):1136–52. doi: 10.1056/NEJMra1406184. [DOI] [PubMed] [Google Scholar]
- 3.Greenberg PL, Tuechler H, Schanz J, Sanz G, Garcia-Manero G, Solé F, Bennett JM, Bowen D, Fenaux P, Dreyfus F, et al. Revised international prognostic scoring system for myelodysplastic syndromes. Blood. 2012;120(12):2454–65. doi: 10.1182/blood-2012-03-420489. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.List A, Bennett J, Sekeres M, Skikne B, Fu T, Shammo J, Nimer S, Knight R, Giagounidis A. Extended survival and reduced risk of aml progression in erythroid-responsive lenalidomide-treated patients with lower-risk del (5q) mds. Leukemia. 2014;28(5):1033–40. doi: 10.1038/leu.2013.305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Harada Y, Harada H. Molecular mechanisms that produce secondary mds/aml by runx1/aml1 point mutations. J Cell Biochem. 2011;112(2):425–32. doi: 10.1002/jcb.22974. [DOI] [PubMed] [Google Scholar]
- 6.Shukron O, Vainstein V, Kündgen A, Germing U, Agur Z. Analyzing transformation of myelodysplastic syndrome to secondary acute myeloid leukemia using a large patient database. Am J Hematol. 2012;87(9):853–60. doi: 10.1002/ajh.23257. [DOI] [PubMed] [Google Scholar]
- 7.Meggendorfer M, De Albuquerque A, Nadarajah N, Alpermann T, Kern W, Steuer K, Perglerová K, Haferlach C, Schnittger S, Haferlach T. Karyotype evolution and acquisition of flt3 or ras pathway alterations drive progression of myelodysplastic syndrome to acute myeloid leukemia. Haematologica. 2015;100(12):487. doi: 10.3324/haematol.2015.127985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Yamazaki J, Estecio MR, Lu Y, Long H, Malouf GG, Graber D, Huo Y, Ramagli L, Liang S, Kornblau SM, et al. The epigenome of aml stem and progenitor cells. Epigenetics. 2013;8(1):92–104. doi: 10.4161/epi.23243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Raza A, Galili N. The genetic basis of phenotypic heterogeneity in myelodysplastic syndromes. Nat Rev Cancer. 2012;12(12):849–59. doi: 10.1038/nrc3321. [DOI] [PubMed] [Google Scholar]
- 10.Wang C, Sashida G, Saraya A, Ishiga R, Koide S, Oshima M, Isono K, Koseki H, Iwama A. Depletion of sf3b1 impairs proliferative capacity of hematopoietic stem cells but is not sufficient to induce myelodysplasia. Blood. 2014;123(21):3336–43. doi: 10.1182/blood-2013-12-544544. [DOI] [PubMed] [Google Scholar]
- 11.Wu SJ, Kuo YY, Hou HA, Li LY, Tseng MH, Huang CF, Lee FY, Liu MC, Liu CW, Lin CT, et al. The clinical implication of srsf2 mutation in patients with myelodysplastic syndrome and its stability during disease evolution. Blood. 2012;120(15):3106–11. doi: 10.1182/blood-2012-02-412296. [DOI] [PubMed] [Google Scholar]
- 12.Parker JE, Mufti GJ, Rasool F, Mijovic A, Devereux S, Pagliuca A. The role of apoptosis, proliferation, and the bcl-2–related proteins in the myelodysplastic syndromes and acute myeloid leukemia secondary to mds. Blood. 2000;96(12):3932–8. [PubMed] [Google Scholar]
- 13.Shimazaki K, Ohshima K, Suzumiya J, Kawasaki C, Kikuchi M. Evaluation of apoptosis as a prognostic factor in myelodysplastic syndromes. Br J Haematol. 2000;110(3):584–90. doi: 10.1046/j.1365-2141.2000.02228.x. [DOI] [PubMed] [Google Scholar]
- 14.Rhyasen G, Starczynowski D. Deregulation of micrornas in myelodysplastic syndrome. Leukemia. 2012;26(1):13–22. doi: 10.1038/leu.2011.221. [DOI] [PubMed] [Google Scholar]
- 15.Haferlach T, Kohlmann A, Wieczorek L, Basso G, Te Kronnie G, Béné M-C, De Vos J, Hernández JM, Hofmann WK, Mills KI, et al. Clinical utility of microarray-based gene expression profiling in the diagnosis and subclassification of leukemia: report from the international microarray innovations in leukemia study group. J Clin Oncol. 2010;28(15):2529–37. doi: 10.1200/JCO.2009.23.4732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Mills KI, Kohlmann A, Williams PM, Wieczorek L, Liu W-M, Li R, Wei W, Bowen DT, Loeffler H, Hernandez JM, et al. Microarray-based classifiers and prognosis models identify subgroups with distinct clinical outcomes and high risk of aml transformation of myelodysplastic syndrome. Blood. 2009;114(5):1063–72. doi: 10.1182/blood-2008-10-187203. [DOI] [PubMed] [Google Scholar]
- 17.Halsey LG, Curran-Everett D, Vowler SL, Drummond GB. The fickle p value generates irreproducible results. Nat Methods. 2015;12(3):179–85. doi: 10.1038/nmeth.3288. [DOI] [PubMed] [Google Scholar]
- 18.Choi Y, Kendziorski C. Statistical methods for gene set coexpression analysis. Bioinformatics. 2009;25(21):2780–6. doi: 10.1093/bioinformatics/btp502. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Zhao S, Fung-Leung WP, Bittner A, Ngo K, Liu X. Comparison of rna-seq and microarray in transcriptome profiling of activated t cells. PloS One. 2014;9(1):e78644. doi: 10.1371/journal.pone.0078644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Sinoquet C, Mourad R. Probabilistic Graphical Models for Genetics, Genomics and Postgenomics. Oxford, UK: Oxford University Press; 2014. [Google Scholar]
- 21.Bing H, Xue-wen C. bneat: a bayesian network method for detecting epistatic interactions in genome-wide association studies. BMC Genomics. 2011;12(Suppl 2):9. doi: 10.1186/1471-2164-12-S2-S9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Liu ZP. Identifying network-based biomarkers of complex diseases from high-throughput data. Biomarkers Med. 2016;10(6):633–50. doi: 10.2217/bmm-2015-0035. [DOI] [PubMed] [Google Scholar]
- 23.Langfelder P, Horvath S. Wgcna: an r package for weighted correlation network analysis. BMC Bioinforma. 2008;9(1):559. doi: 10.1186/1471-2105-9-559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Oldham MC, Horvath S, Geschwind DH. Conservation and evolution of gene coexpression networks in human and chimpanzee brains. Proc Nat Acad Sci. 2006;103(47):17973–8. doi: 10.1073/pnas.0605938103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Yeung KY, Ruzzo WL. Principal component analysis for clustering gene expression data. Bioinformatics. 2001;17(9):763–74. doi: 10.1093/bioinformatics/17.9.763. [DOI] [PubMed] [Google Scholar]
- 26.Milacic M, Haw R, Rothfels K, Wu G, Croft D, Hermjakob H, D’Eustachio P, Stein L. Annotating cancer variants and anti-cancer therapeutics in reactome. Cancers. 2012;4(4):1180–211. doi: 10.3390/cancers4041180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Alharbi RA, Pettengell R, Pandha HS, Morgan R. The role of hox genes in normal hematopoiesis and acute leukemia. Leukemia. 2013;27(5):1000–8. doi: 10.1038/leu.2012.356. [DOI] [PubMed] [Google Scholar]
- 28.Garzon R, Volinia S, Papaioannou D, Nicolet D, Kohlschmidt J, Yan PS, Mrózek K, Bucci D, Carroll AJ, Baer MR, et al. Expression and prognostic impact of lncrnas in acute myeloid leukemia. Proc Nat Acad Sci. 2014;111(52):18679–84. doi: 10.1073/pnas.1422050112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Jolliffe I. Principal Component Analysis. Hoboken, NJ: Wiley Online Library; 2002. [Google Scholar]
- 30.Welch BL. The generalization of student’s problem when several different population variances are involved. Biometrika. 1947;34(1/2):28–35. doi: 10.2307/2332510. [DOI] [PubMed] [Google Scholar]
- 31.Scutari M. Learning bayesian networks with the bnlearn r package. J Stat Softw. 2010;35(1):1–22. [Google Scholar]
- 32.Quinlan JR. C4.5: Programming for Machine Learning. Amsterdam, Netherlands: Elsevier; 1993. [Google Scholar]
- 33.Bejar R. Prognostic models in myelodysplastic syndromes. ASH Educ Program Book. 2013;2013(1):504–10. doi: 10.1182/asheducation-2013.1.504. [DOI] [PubMed] [Google Scholar]
- 34.Bruserud Ø, Gjertsen BT, Huang T-S. Induction of differentiation and apoptosis—a possible strategy in the treatment of adult acute myelogenous leukemia. The Oncologist. 2000;5(6):454–62. doi: 10.1634/theoncologist.5-6-454. [DOI] [PubMed] [Google Scholar]
- 35.Lu P, Weaver VM, Werb Z. The extracellular matrix: a dynamic niche in cancer progression. J Cell Biol. 2012;196(4):395–406. doi: 10.1083/jcb.201102147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Kim SH, Turnbull J, Guimond S. Extracellular matrix and cell signalling: the dynamic cooperation of integrin, proteoglycan and growth factor receptor. J Endocrinol. 2011;209(2):139–51. doi: 10.1530/JOE-10-0377. [DOI] [PubMed] [Google Scholar]
- 37.Armstrong SJ, Wiberg M, Terenghi G, Kingham PJ. Ecm molecules mediate both schwann cell proliferation and activation to enhance neurite outgrowth. Tissue Eng. 2007;13(12):2863–70. doi: 10.1089/ten.2007.0055. [DOI] [PubMed] [Google Scholar]
- 38.Ingber DE, Folkman J. Mechanochemical switching between growth and differentiation during fibroblast growth factor-stimulated angiogenesis in vitro: role of extracellular matrix. J Cell Biol. 1989;109(1):317–30. doi: 10.1083/jcb.109.1.317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Ilić D, Almeida EA, Schlaepfer DD, Dazin P, Aizawa S, Damsky CH. Extracellular matrix survival signals transduced by focal adhesion kinase suppress p53-mediated apoptosis. J Cell Biol. 1998;143(2):547–60. doi: 10.1083/jcb.143.2.547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Mahlknecht U, Schönbein C. Histone deacetylase inhibitor treatment downregulates vla-4 adhesion in hematopoietic stem cells and acute myeloid leukemia blast cells. Haematologica. 2008;93(3):443–6. doi: 10.3324/haematol.11796. [DOI] [PubMed] [Google Scholar]
- 41.Mi H, Poudel S, Muruganujan A, Casagrande JT, Thomas PD. Panther version 10: expanded protein families and functions, and analysis tools. Nucleic Acids Res. 2016;44(D1):336–42. doi: 10.1093/nar/gkv1194. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Lu P, Takai K, Weaver VM, Werb Z. Extracellular matrix degradation and remodeling in development and disease. Cold Spring Harb Perspect Biol. 2011;3(12):005058. doi: 10.1101/cshperspect.a005058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Suzuki MM, Bird A. Dna methylation landscapes: provocative insights from epigenomics. Nat Rev Genet. 2008;9(6):465–76. doi: 10.1038/nrg2341. [DOI] [PubMed] [Google Scholar]
- 44.Zhu Q, Wong AK, Krishnan A, Aure MR, Tadych A, et al. Targeted exploration and analysis of large cross-platform human transcriptomic compendia. Nat Methods. 2015;12(3):211–4. doi: 10.1038/nmeth.3249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Kohlmann A, Bullinger L, Thiede C, Schaich M, Schnittger S, Döhner K, Dugas M, Klein H, Döhner H, Ehninger G, et al. Gene expression profiling in aml with normal karyotype can predict mutations for molecular markers and allows novel insights into perturbed biological pathways. Leukemia. 2010;24(6):1216–20. doi: 10.1038/leu.2010.73. [DOI] [PubMed] [Google Scholar]
- 46.Haferlach C, Mecucci C, Schnittger S, Kohlmann A, Mancini M, Cuneo A, Testoni N, Rege-Cambrin G, Santucci A, Vignetti M, et al. Aml with mutated npm1 carrying a normal or aberrant karyotype show overlapping biologic, pathologic, immunophenotypic, and prognostic features. Blood. 2009;114(14):3024–32. doi: 10.1182/blood-2009-01-197871. [DOI] [PubMed] [Google Scholar]
- 47.Metzeler KH, Hummel M, Bloomfield CD, Spiekermann K, Braess J, Sauerland MC, Heinecke A, Radmacher M, Marcucci G, Whitman SP, et al. An 86-probe-set gene-expression signature predicts survival in cytogenetically normal acute myeloid leukemia. Blood. 2008;112(10):4193–201. doi: 10.1182/blood-2008-02-134411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Miesner M, Haferlach C, Bacher U, Weiss T, Macijewski K, Kohlmann A, Klein HU, Dugas M, Kern W, Schnittger S, et al. Multilineage dysplasia (mld) in acute myeloid leukemia (aml) correlates with mds-related cytogenetic abnormalities and a prior history of mds or mds/mpn but has no independent prognostic relevance. Blood. 2010;116(15):2742–51. doi: 10.1182/blood-2010-04-279794. [DOI] [PubMed] [Google Scholar]
- 49.Grossmann V, Bacher U, Kohlmann A, Artusi V, Klein HU, Dugas M, Schnittger S, Alpermann T, Kern W, Haferlach T, et al. Ezh2 mutations and their association with picalm-mllt10 positive acute leukaemia. Br J Haematol. 2012;157(3):387–90. doi: 10.1111/j.1365-2141.2011.08986.x. [DOI] [PubMed] [Google Scholar]
- 50.Marchionni L. Impact of Gene Expression Profiling Tests on Breast Cancer Outcomes. Collingdale, PA: DIANE Publishing; 2009. [PMC free article] [PubMed] [Google Scholar]
- 51.Tuma RS. A big trial for a new technology: Transbig project takes microarrays into clinical trials. J Nat Cancer Inst. 2004;96(9):648–9. doi: 10.1093/jnci/96.9.648. [DOI] [PubMed] [Google Scholar]
- 52.Li J, Lenferink AE, Deng Y, Collins C, Cui Q, Purisima EO, O’Connor-McCourt MD, Wang E. Identification of high-quality cancer prognostic markers and metastasis network modules. Nat Commun. 2010;1:34. doi: 10.1038/ncomms1033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Zhao X, Rodland EA, Tibshirani R, Plevritis S. Molecular subtyping for clinically defined breast cancer subgroups. Breast Cancer Res. 2015;17(1):29. doi: 10.1186/s13058-015-0520-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Citri A, Yarden Y. Egf–erbb signalling: towards the systems level. Nat Rev Mol Cell Biol. 2006;7(7):505–16. doi: 10.1038/nrm1962. [DOI] [PubMed] [Google Scholar]
- 55.Raiser DM, Narla A, Ebert BL. The emerging importance of ribosomal dysfunction in the pathogenesis of hematologic disorders. Leuk lymphoma. 2014;55(3):491–500. doi: 10.3109/10428194.2013.812786. [DOI] [PubMed] [Google Scholar]
- 56.Zhou X, Liao WJ, Liao JM, Liao P, Lu H. Ribosomal proteins: functions beyond the ribosome. J Mol Cell Biol. 2015;7(2):92–104. doi: 10.1093/jmcb/mjv014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Nakayama KI, Nakayama K. Ubiquitin ligases: cell-cycle control and cancer. Nat Rev Cancer. 2006;6(5):369–81. doi: 10.1038/nrc1881. [DOI] [PubMed] [Google Scholar]
- 58.Boudreau N, Bissell MJ. Extracellular matrix signaling: integration of form and function in normal and malignant cells. Curr Opin Cell Biol. 1998;10(5):640–6. doi: 10.1016/S0955-0674(98)80040-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Kessel M, Gruss P, et al. Murine developmental control genes. Science. 1990;249(4967):374–9. doi: 10.1126/science.1974085. [DOI] [PubMed] [Google Scholar]
- 60.Abate-Shen C. Deregulated homeobox gene expression in cancer: cause or consequence? Nat Rev Cancer. 2002;2(10):777–85. doi: 10.1038/nrc907. [DOI] [PubMed] [Google Scholar]
- 61.Rhoads K, Arderiu G, Charboneau A, Hansen SL, Hoffman W, Boudreau N. A role for hox a5 in regulating angiogenesis and vascular patterning. Lymphatic Res Biol. 2005;3(4):240–52. doi: 10.1089/lrb.2005.3.240. [DOI] [PubMed] [Google Scholar]
- 62.Mace KA, Hansen SL, Myers C, Young DM, Boudreau N. Hoxa3 induces cell migration in endothelial and epithelial cells promoting angiogenesis and wound repair. J Cell Sci. 2005;118(12):2567–77. doi: 10.1242/jcs.02399. [DOI] [PubMed] [Google Scholar]
- 63.Afonja O, Smith Jr JE, Cheng DM, Goldenberg AS, Amorosi E, Shimamoto T, Nakamura S, Ohyashiki K, Ohyashiki J, Toyama K, et al. Meis1 and hoxa7 genes in human acute myeloid leukemia. Leuk Res. 2000;24(10):849–55. doi: 10.1016/S0145-2126(00)00059-X. [DOI] [PubMed] [Google Scholar]
- 64.Leroy P, Berto F, Bourget I, Rossi B. Down-regulation of hox a7 is required for cell adhesion and migration on fibronectin during early hl-60 monocytic differentiation. J Leukoc Biol. 2004;75(4):680–8. doi: 10.1189/jlb.0503246. [DOI] [PubMed] [Google Scholar]
- 65.Bauvois B. New facets of matrix metalloproteinases mmp-2 and mmp-9 as cell surface transducers: outside-in signaling and relationship to tumor progression. Biochim Biophys Acta (BBA)-Rev Cancer. 2012;1825(1):29–36. doi: 10.1016/j.bbcan.2011.10.001. [DOI] [PubMed] [Google Scholar]
- 66.Hatfield JK, Reikvam H, Bruserud O. The crosstalk between the matrix metalloprotease system and the chemokine network in acute myeloid leukemia. Curr Med Chem. 2010;17(36):4448–61. doi: 10.2174/092986710794183033. [DOI] [PubMed] [Google Scholar]
- 67.Paupert J, Mansat-De Mas V, Demur C, Salles B, Muller C. Cell-surface mmp-9 regulates the invasive capacity of leukemia blast cells with monocytic features. Cell Cycle. 2008;7(8):1047–53. doi: 10.4161/cc.7.8.5645. [DOI] [PubMed] [Google Scholar]
- 68.Feng S, Cen J, Huang Y, Shen H, Yao L, Wang Y, Chen Z. Matrix metalloproteinase-2 and-9 secreted by leukemic cells increase the permeability of blood-brain barrier by disrupting tight junction proteins. PLoS One. 2011;6(8):20599. doi: 10.1371/journal.pone.0020599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Bernal T, Moncada-Pazos Á, Soria-Valles C, Gutiérrez-Fernández A. Effects of azacitidine on matrix metalloproteinase-9 in acute myeloid leukemia and myelodysplasia. Exp Hematol. 2013;41(2):172–9. doi: 10.1016/j.exphem.2012.10.005. [DOI] [PubMed] [Google Scholar]
- 70.Aref S, El-Sherbiny M, Mabed M, Menessy A, El-Refaei M. Urokinase plasminogen activator receptor and soluble matrix metalloproteinase-9 in acute myeloid leukemia patients: a possible relation to disease invasion. Hematology. 2003;8(6):385–91. doi: 10.1080/10245330310001621323. [DOI] [PubMed] [Google Scholar]
- 71.Travaglino E, Benatti C, Malcovati L, Porta MGD, Gallì A, Bonetti E, Rosti V, Cazzola M, Invernizzi R. Biological and clinical relevance of matrix metalloproteinases 2 and 9 in acute myeloid leukaemias and myelodysplastic syndromes. Eur J Haematol. 2008;80(3):216–26. doi: 10.1111/j.1600-0609.2007.01012.x. [DOI] [PubMed] [Google Scholar]
- 72.Hastie T, Tibshirani R, Eisen MB, Alizadeh A, Levy R, Staudt L, Chan WC, Botstein D, Brown P, et al. Gene shaving as a method for identifying distinct sets of genes with similar expression patterns. Genome Biol. 2000;1(2):1–0003. doi: 10.1186/gb-2000-1-2-research0003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Ma S, Song X, Huang J. Supervised group lasso with applications to microarray data analysis. BMC Bioinforma. 2007;8(1):60. doi: 10.1186/1471-2105-8-60. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Patro R, Mount SM, Kingsford C. Sailfish enables alignment-free isoform quantification from rna-seq reads using lightweight algorithms. Nat Biotechnol. 2014;32(5):462–4. doi: 10.1038/nbt.2862. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B. Mapping and quantifying mammalian transcriptomes by RNA-seq. Nat Methods. 2008;5(7):621–8. doi: 10.1038/nmeth.1226. [DOI] [PubMed] [Google Scholar]
- 76.Curtis C, Shah SP, CHin SF, Turashvili G, Rueda OM, et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature. 2012;486(7403):346–52. doi: 10.1038/nature10983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Miller LD, Smeds J, George J, Vega VB, Vergara L, Ploner A, Pawitan Y, Hall P, Klaar S, Liu ET, et al. An expression signature for p53 status in human breast cancer predicts mutation status, transcriptional effects, and patient survival. Proc Nat Acad Sci USA. 2005;102(38):13550–5. doi: 10.1073/pnas.0506230102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Zhang B, Gaiteri C, Bodea LG, Wang Z, McElwee J, Podtelezhnikov AA, Zhang C, Xie T, Tran L, Dobrin R, et al. Integrated systems approach identifies genetic nodes and networks in late-onset alzheimer’s disease. Cell. 2013;153(3):707–20. doi: 10.1016/j.cell.2013.03.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Tu Z, Zhang B, Zhu J. Network integration of genetically regulated gene expression to study complex diseases. Integrating Omics Data. 2015;88:88–109. doi: 10.1017/CBO9781107706484.005. [DOI] [Google Scholar]
- 80.Zare H, Haffari G, Gupta A, Brinkman RR. Scoring relevancy of features based on combinatorial analysis of lasso with application to lymphoma diagnosis. BMC Genomics. 2013;14(Suppl 1):14. doi: 10.1186/1471-2164-14-S1-S14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Miller BG, Stamatoyannopoulos JA. Integrative meta-analysis of differential gene expression in acute myeloid leukemia. PLoS One. 2010;5(3):e9466. doi: 10.1371/journal.pone.0009466. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Dong J, Horvath S. Understanding network concepts in modules. BMC Syst Biol. 2007;1(1):24. doi: 10.1186/1752-0509-1-24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Liberzon A, Subramanian A, Pinchback R, Thorvaldsdóttir H, Tamayo P, Mesirov JP. Molecular signatures database (msigdb) 3.0. Bioinformatics. 2011;27(12):1739–40. doi: 10.1093/bioinformatics/btr260. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Schroeder A, Mueller O, Stocker S, Salowsky R, Leiber M, Gassmann M, Lightfoot S, Menzel W, Granzow M, Ragg T. The rin: an rna integrity number for assigning integrity values to rna measurements. BMC Mol Biol. 2006;7(1):3. doi: 10.1186/1471-2199-7-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Ranola JM, Langfelder P, Lange K, Horvath S. Cluster and propensity based approximation of a network. BMC Syst Biol. 2013;7(1):21. doi: 10.1186/1752-0509-7-21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Jensen FV. An Introduction to Bayesian Networks. London: UCL press; 1996. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
All data and code required to reproduce the presented results are available, either through publicly available repositories or as supplementary materials. Specifically, the MILE leukemia and Miller breast cancer datasets are available from GEO with accession numbers GSE3494 and GSE15061, respectively, and the METABRIC dataset is available from the European Genome-phenome Archive with study accession number EGAS00000000083. DNA methylation data are available from TCGA (LAML dataset). Additionally, eigengene values for the MILE and BCCA datasets are available as supplementary materials. The Pigengene software package is available through Bioconductor.


