Abstract
The human Glucose Transporter 1 (hGLUT1 or SLC2A1) is a facilitative membrane transporter found in the liver, intestines, kidney, and brain, where it transports sugars such as D-glucose and D-galactose. Genetic variations in hGLUT1 are associated with a broad range of diseases and metabolic disorders. For example, hGLUT1 is upregulated in various cancer types (e.g., breast carcinoma) to support the increased anaerobic glycolysis and the Warburg Effect. Thus, hGLUT1 is an emerging therapeutic target, which also transports commonly used cancer biomarkers (e.g., 18F-DG). In this study, we use computational prediction followed by experimental testing, to characterize hGLUT1. We construct homology models of hGLUT1 in a partially occluded outward open (‘occluded’) conformation based on the X-ray structure of the E. coli xylose transporter, XylE. Comparison of the binding site of the occluded models to experimentally determined hGLUT structures revealed a hydrophobic pocket adjacent to the sugar-binding site, which was tested experimentally via site-directed mutagenesis. Virtual screening of various libraries of purchasable compounds against the occluded models, followed by experimental testing with cellular assays revealed seven previously unknown hGLUT1 ligands with IC50 values ranging from 0.45 μM to 59 μM. These ligands represent three unique chemotypes that are chemically different from any other known hGLUT1 ligands. The newly characterized hydrophobic pocket can potentially be utilized by the new ligands for increased affinity. Furthermore, the previously unknown hGLUT1 ligands can serve as chemical tools to further characterize hGLUT1 function or lead molecules for future drug development.
Keywords: Glucose transporter, SLC2A1, sugar transport, Solute Carrier transporter, cancer metabolism
INTRODUCTION
D-Glucose is an essential molecule serving as energy source for various biological functions, such as transcription, cell proliferation, motility, and neuronal activity.1 Sugar transporters, such as members of the Solute Carrier (SLC) transporter families SLC2 and SLC5, regulate the cellular concentration of sugars using a variety of transport mechanisms.2,3 The human Glucose Transporter 1 (hGLUT1 or SLC2A1) is a facilitative glucose transporter that maintains the basal-level cellular glucose concentration in multiple tissues, including the liver, intestines, kidney, and brain.4,5
Genetic variations in hGLUT1 have been associated with different metabolic disorders as well as with cancer. For example, deletion of four highly conserved residues (Q282-S285), are linked to hemolytic anemia likely caused by cation leak.6 hGLUT1 is also upregulated in some cancer types7,8 (e.g., breast cancer) and it is thought to support the shift of metabolism from oxidative respiration to anaerobic glycolysis, a phenomenon known as the Warburg effect.9,10 hGLUT1 transports biomarkers of cancer (e.g. 18F-DG11) and serves as a drug target, where a potential drug can be an inhibitor that deprives cancer cells from D-glucose or a cytotoxic substrate that acts on different intracellular target.12–14
hGLUT1 interacts with a broad range of organic molecules; it selectively transports monosaccharides such as D-glucose and D-galactose, as well as dehydroascorbic acid,15,16 but does not transport fructose. Various natural products inhibit hGLUT1. They include caffeine,17 which was suggested to interact with a putative nucleotide-binding site on the cytosolic face of the transporter; polyphenols,18,19 such as phloretin, a hGLUT1 inhibitor found in apple tree; cytochalasin B and forskolin, an alkaloid mycotoxin and a phytogenic labdane diterpene, respectively, are promiscuous inhibitors that also strongly inhibit hGLUT1.20 Although the identification of these inhibitors contributed to understanding structure/function relationships in hGLUT1, many of these inhibitors do not exhibit drug-like properties and may be toxic.21 Therefore, small molecules targeting hGLUT1 are needed as chemical tools to further characterize this transporter, as well as lead molecules for future drug discovery.
It is thought that hGLUT1 transport solutes via a “rocker-switch”, alternating-access mechanism, in which the transporter alternately exposes its binding site to either the extracellular space or cytoplasm and simultaneously closing the opposing face of the transporter, while transitioning through an intermediate state in which the sugar-binding site is occluded.22–26 A comprehensive description of the different conformational states of hGLUT1 including its binding mode to small molecule ligands is needed to design chemical probes targeting this biomedically important protein.
Recently, an atomic structure of the hGLUT1 has been determined in an inward-open conformation bound to a nonyl-β-D-glucoside inhibitor.27 This structure confirmed that hGLUT1 is member of the Major Facilitator Superfamily (MFS), one of the most common structural families of membrane proteins in all kingdoms of life.28 The structure revealed a wide, solvent-exposed channel that connects the binding site to the cytoplasm (Fig. 1A). Additionally, atomic structures of various hGLUT1 homologs have been determined in multiple conformations, revealing different snapshots of the transport cycle. For example, the structures of hGLUT3, which shares 86% sequence similarity to hGLUT1, were determined in the outward-open and occluded conformations;29 the structures of the fructose transporter GLUT5 from rat and bovine (75% similarity to hGLUT1) were determined in the inward-open and outward-open conformations,30 respectively; the structures of xylose transporter XylE from Escherichia coli (sequence similarity of 63%), were solved in the inward- and outward-open conformations,22,31 and partially occluded outward-open (‘occluded’) conformation bound to the natural substrate D-glucose (Fig 1B).32 Notably, the binding site of the occluded XylE structure is solvent-excluded and well resolved; it is also compact, which limits the size of ligands and their binding modes. Overall, this unique conformation provides crucial information on the hydrogen-bond network involved in substrate recognition and specificity (Fig. 1B, bottom). Taken together, the occluded XylE structure is a suitable template to model hGLUT1 in occluded conformation, which is still unknown and can be used for virtual screening against its unique binding site.
Figure 1. Crystal structures of hGLUT1 and E. coli XylE.
Shown is side view of (A) hGLUT1 in an inward-open conformation (4PYP) and (B) XylE in a partially occluded outward open conformation (4GBZ). The occluded XylE structure has more compact ligand binding site and protein-ligand interactions than the inward-open hGLUT1 structure.
Here, we use an integrated computational and experimental approach to identify previously unknown hGLUT1 ligands. We first performed a comprehensive comparison of various GLUT structures, including structures of hGLUT1 and its mammalian and bacterial homologs. We then generated models of hGLUT1 in occluded conformation, which has not been characterized yet, based on the occluded XylE structure. The hGLUT1 model was tested using site-directed mutagenesis with a focus on binding site residues predicted to make key interactions with the substrate as well as residues in a newly described hydrophobic pocket. We then performed virtual screening of various compound libraries against the occluded hGLUT1 models,33 where top-scoring compounds were confirmed in cell-based assay. Finally, we discuss the potential utility of the newly discovered hGLUT1 ligands as useful chemical tools to or lead molecules for drug development.
MATERIALS AND METHODS
Homology Modeling
Structures of multiple hGLUT1 homologs have been determined in a variety of conformations. Our goal was to model an unknown conformation of hGLUT1 and generate models that likely to be sufficiently accurate for successful virtual screening. We used a variety of considerations to select the most relevant template (Supplementary Materials). We modeled hGLUT1 based on the structure of the E. coli xylose transporter XylE in the occluded conformation (PDB: 4GC032). Protein sequences of the human GLUT family (hGLUT 1–12) and XylE were obtained from UniProt34 and aligned with the multiple sequence alignment program MUSCLE.35 We ensured the functionally important residues and the transmembrane helices TM1, TM3, TM5, TM7, and TM10 are aligned to the corresponding regions of hGLUT1–4 (Fig. S1). Initial 100 models were generated using MODELLER36 v9.12 with two cycles of optimization and assessed with Z-DOPE, a normalized atomic distance-dependent statistical potential based on known protein structures.37 Twenty top-scoring models were visually inspected and three models were subjected to virtual screening. Volume of the sugar-binding site was calculated with POVME 2.0.38 MMGBSA binding free energy estimation, which omits the contribution of entropy, was performed in Schrödinger’s Maestro 10.2 program.39
Molecular Docking
Due to the small size of the sugar-binding site of the occluded hGLUT1 models, we chose to screen the models against the “Fragment Now” (downloaded on 2013-02-14; 454,934 molecules) and the NCI-2007 (downloaded on 2014-03-19; 93,378 molecules) libraries of the ZINC database,40 which included compounds with low molecular weight and are readily available. Rigid conformers of the ligands were generated using OpenEye OMEGA41 2.5.1.4. Each ligand conformer was docked to the hGLUT1 model using OpenEye FRED42 (default settings) and scored using the native ChemGauss4 scoring function.43 Tanimoto coefficients (Tc) of the selected top-ranking docked ligands were compared to known hGLUT1 ligands in the ChEMBL database,44 using the ECFP-4 fingerprints45 implemented in the RDKit package (www.rdkit.org). The ligands were analyzed with a pan assay interference (PAINS) filter to identify frequent false positive compounds in large screens, such as rhodanines and enones.46
Stably expressed hGLUT1 CHO cell line establishment
Stably transfected Flp-in Chinese hamster ovary (CHO) cell line overexpressing hGLUT1 (CHO-hGLUT1) was established by transfecting pcDNA5/FRT vector (Life Technologies, Carlsbad, CA) containing the full-length human SLC2A1 cDNA using Lipofectamine 2000 according to manufacturer’s instructions (Life Technologies). Transfected cells were maintained in HAM’s F12 containing 10% (v/v) FBS, 100 units/mL penicillin, 100 μg/mL streptomycin, and 500 μg/mL hygromycin B at 37 °C and 5% CO2. Site-directed mutagenesis in the SLC2A1 gene was performed as previously described.47 The primers for the mutagenesis were listed in the supplementary Table S3. mRNA expression level of hGLUT1 in CHO cells was measured by real-time PCR using SYBR Green dyes and compared to CHO cell transfected with empty vector (CHO-EV) (Fig. S2A).
Detection of hGLUT1 in CHO cells
To detect hGLUT1 in whole cell, CHO cells overexpressing hGLUT1 and mutants were lysed in CelLytic™ M Cell Lysis Reagent (Sigma-Aldrich, St. Louis) with protease inhibitor cocktail (Sigma-Aldrich). Cell extracts (10 μg) were separated by SDS-PAGE (10%) and transferred onto a PVDF membrane. For membrane protein isolation, the cell surface protein biotinylation and purification kit (Pierce, Rockford, IL; #89881) was used. Briefly, CHO cells were biotinylated using Sulfo-NHS-SS Biotin for 30 min at 4 °C. Cells were then lysed and the lysates were run on the Streptavidin column to collect the biotinylated fractions that correspond to cell surface proteins. Subsequently, the collected fractions were eluted using SDS and run on SDS-PAGE. Blots were blocked in 5% non-fat milk in TBS with 0.05% Tween 20 and detected with primary antibody: anti-GLUT1 rabbit monoclonal antibody (Abcam, Cambridge, MA; ab115730), anti-Na+/K+-ATPase rabbit monoclonal antibody (Abcam; ab76020), and anti-β-actin mouse monoclonal antibody (Proteintech, Chicago, IL; 66009-1-Ig). Anti-rabbit IgG (CST, #7074) and anti-mouse IgG HRP-linked antibodies (CST, #7076) were subsequently used to detect the primary antibodies. Bound secondary antibodies were visualized by Immobilon Western Chemi HRP Substrate (Millipore, Billerica) and analyzed using the Tanon 5200 Chemiluminescent Imaging System (Tanon, Shanghai, China).
[3H]-2-deoxy-D-glucose ([3H]-2-DG) Uptake Assay
The stably transfected CHO cells were seeded in poly 24-well plates and were grown to 80–90% confluence. Cells were rinsed with PBS buffer and incubated in 280 μL of PBS buffer containing 61.73 μM [3H]-2-DG (PerkinElmer, Boston, MA) in the presence and absence of 50 μM test compound for 5 mins. The known hGLUT1 inhibitor phloretin (Accela ChemBio, San Diego, CA) was included as a positive control. Unradiolabelled 2-DG (40 mM) (Sigma, St. Louis, MO) was included in order to calculate specific inhibition of predicted inhibitors. Compounds were obtained from the National Cancer Institute (NCI) or purchased from commercial vendors. The reaction was terminated by washing cells twice with ice-cold PBS, followed by addition of 200 μL lysis buffer (0.1% (w/v) SDS, 0.1 N NaOH). Intracellular radioactivity was determined by scintillation counting. CHO-hGLUT1 cells showed a 3.92-fold increase in probe uptake relative to the CHO-EV cells and demonstrated to be a suitable system for sugar uptake inhibition assay (Fig. S2B). Probe-independent specific inhibition of the tested compound, which excludes hGLUT1 inhibition due to [3H]-2-DG, was calculated as follow:
where R is the intracelullar radioactivity of the treatment group, R2DG is the intracellular radioactivity of [3H]-2-DG treated group, and D0 is the intracellular radioactivity of the control group of [3H]-2-DG. Dose-dependent inhibition was measured under the same conditions as the single-point measurements. Cells were incubated with compounds at 0.1, 0.3, 1, 3, 10, 30, 100, 300, 1000, and 3000 μM. Ligand efficiency (LE) of the compounds was calculated with the following equation:
where R is the ideal gas constant, T is temperature at 25 °C, and HA is the number of heavy atoms.
RESULTS AND DISCUSSION
Analysis of the mammalian GLUTs and XylE structures
In homology modeling, a target protein is modeled based on a structure of a related protein (i.e. template structure). Characterizing different conformational states of hGLUT1 is important for understanding the transport process, as well as for identifying small molecule ligands via virtual screening, where virtual screenings against different conformational states are expected to capture different portions of the chemical space.48,49 To select a template for homology modeling relevant for virtual screening, we first compared the structures of hGLUT1 to those of XylE (E. coli), GLUT3 (human), and GLUT5 (rat and bovine), taking into account the binding site similarity, the quality of the structure in the binding site region, and the uniqueness of the conformation (Methods, Supplementary Material).
hGLUT1 and XylE are homologs that are closely related in their structure and function. For example, the atomic structures of XylE and hGLUT1 in the inward-open conformation have RMSD of 1.5 Å,27,31 despite sharing sequence identity of 26% (63% similarity). Both transporters have a conserved binding site (i.e., 12 of the 16 binding site residues are identical; 2 are similar), and share highly similar substrate specificity.50 Interestingly, XylE structures were solved with the natural substrates bound in a unique binding site conformation not available for any of the other hGLUT1 homologs.32
Recently, a D-glucose-bound atomic structure of hGLUT3 was determined in an occluded conformation29 similar to the XylE32 occluded structures (RMSD of 1.6 Å); however, two detergent molecules of 1-glycerol monooleate were found in the sugar-binding site of hGLUT3 (Fig. S3B), thereby altering its conformation. Particularly, TM1, TM5, TM7, and TM10 are shifted, affecting the overall shape of the binding site. Therefore, despite the fact that hGLUT3 is closer in sequence to hGLUT1, XylE is an appropriate template for modeling hGLUT1, and targeting the binding site in this conformation with virtual screening is expected to identify ligands less likely to be captured by other conformations.
The structures of hGLUT1 and XylE in the inward-open and occluded conformations, respectively, have two key differences that are relevant for transporter-ligand recognition: first, the conformation of transmembrane helix 10 (TM10) that forms part of the sugar-binding site varies among these two conformations, leading to difference in the size of the sugar-binding site. In the inward-open structures TM10 does not block the cytosolic channel that links the sugar-binding site to the cytosol, while in the occluded conformation TM10 shifts by 4.5 Å, obstructing the sugar-binding site from the cytosol. This major conformational change places W392 (W388 in hGLUT1), a key residue that coordinates sugar binding via a hydrogen bond, at the opening of the cytosolic channel, and the indole sidechain of W392 completely blocks the entry to the channel (Fig. 1B, bottom). This movement of TM10 leads to a much smaller binding site in the occluded structures (< 70 Å3) (Fig. 1B, bottom) as compared to the channel-like, elongated pocket in the inward-open structures (> 540 Å3) (Table S1; Fig. 1A, bottom).
Second, the occluded XylE structures involve more hydrogen bonds between the transporter and the sugar than the inward-open hGLUT1 structure (Fig 1, bottom). D-glucose makes 14 hydrogen bonds with XylE (involving residues Q168, Q175, Q288, Q289, N294, G388, W392, and Q415), while the inhibitor nonyl-β-D-glycoside makes 6 hydrogen bonds with hGLUT1 (Q282, Q283, N288, N317, and E380). Importantly, although the allowed ligand size in the sugar-binding site is smaller, the occluded structure appears to allow higher per heavy-atom estimated binding free energy than the corresponding site in the inward-open hGLUT1 (Table S1), making them the ideal structures for identifying small and efficient ligands using structure-based drug discovery approach.
Homology modeling of hGLUT1
hGLUT1 was modeled using MODELLER, based on the XylE structure in the occluded conformation (Fig. S1; Methods). The hGLUT1 homology models obtained Z-DOPE score between −0.38 and −0.76, suggesting that 65%–75% of their Cα atoms are within 3.5 Å of their correct positions.51 Key binding site residues, including Q161, Q282, Q283, N288, N317, W388, and N415, adopt similar conformation to those of the corresponding residues in XylE (Q168, Q288, Q289, N294, N325, W392, and N419) that form critical polar interactions with D-glucose.32,50 Moreover, our hGLUT1 models are highly similar to the new hGLUT3 structure (RMSD of 1.6 Å) (Fig. S3). These data increase our confidence in the models and suggest that they are likely to be sufficiently accurate for a successful virtual screening campaign.
Notably, our models reveal a hydrophobic pocket, termed the H-pocket, adjacent to the polar residues within the sugar-binding site (Figs. 2; S1). This pocket constitutes part of the sugar-binding site and is lined by the conserved residues G27, T30, I164, V165, I168, and F291. Two of the H-pocket residues, T30 and I168, are adjacent to G27 and G167 of TM1 and TM5, respectively, and might also have a structural role and helix-helix packing. Moreover, the primary difference between the inward-open and occluded conformations involves shift of TM1, TM5, and TM7 near the sugar-binding site. These conformational changes lead to significant differences in the size and shape of the H-pocket between the two conformations (Figs. 2C,D). In particular, the H-pocket in the occluded structures is significantly larger (4GC0: 37.5 Å3; Fig. 2B) compared to the corresponding pocket in the inward-open crystal structure (4PYP: 10.0 Å3; Fig. 2C).
Figure 2. The hydrophobic H-pocket adjacent to the hGLUT1 sugar-binding site.
Homology model of hGLUT1 based on the occluded XylE structure is in white and inward-open hGLUT1 structure is shown in gold color. (A) The hydrophobic moiety of multiple docked ligands (e.g., PUG-1) is predicted to reside in the H-pocket lined by the hydrophobic residues G27, T30, I164, V165, I168, and F291 (yellow sticks). (B) The H-pocket is larger in the hGLUT1 model in the occluded conformation (green spheres; 37 Å3) than in (C) the inward-open hGLUT1 structure (magenta spheres; 10 Å3) due to the rearrangement of the transmembrane helices (TM1, TM5, and TM7). (D) The occluded H-pocket (green surface) is larger and shaped differently than the inward-open H-pocket (purple spheres).
Site-directed mutagenesis of key residues
Next, we used site-directed mutagenesis to validate the model binding site, focusing on the previously undescribed H-pocket residues T30, I168, and F291 (Table S2), as their sidechain lines the H-pocket. All hGLUT1 mutants were expressed at comparable quantity to the wild-type transporter in the transfected CHO cells (Fig. 3A). The level of protein expression in whole cell extract (Fig. 3A) is also correlated with the real-time PCR measurement (Fig. S2C), indicating that all mutants and wild-type hGLUT1 have similar mRNA and protein expression levels in the transfected CHO cells. Importantly, all mutants were properly expressed on the cell surface (Fig. 3B), as confirmed by cell surface protein biotinylation experiment.
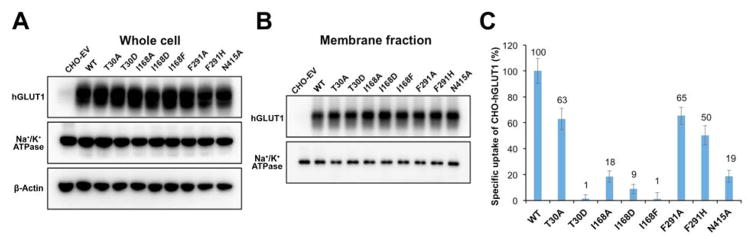
Figure 3.
Western blots of (A) hGLUT1 in whole cell lysate and (B) biotinylated hGLUT1 in membrane protein fractions prepared from CHO cells stably transfected with empty vector (EV), wild-type (WT), and mutated hGLUT1 plasmids. Na+/K+-ATPase and β-actin were used as plasma membrane and cytoplasmic protein markers, respectively. (C) Specific [3H]-2-deoxy-D-glucose uptake of WT and mutant hGLUT1 in stably transfected CHO cell lines. Experiments were performed in triplicate. Error bars represent SEM.
The activity of the hGLUT1 mutants was measured in the [3H]-2-DG uptake assay (Fig. 3C). As a positive control, we mutated N415, a conserved polar residue within the sugar-binding site that coordinates Q283 and N288 through an extensive hydrogen-bonding network to interact with the substrate. The N415A mutant has shown substantially reduced substrate uptake activity (18% remaining activity) compared to the wild-type transporter. We describe each of the mutants in turn.
T30 is located on TM1, 4.9 Å away from the substrate and its sidechain faces the sugar-binding site. In addition, the β-hydroxyl group of T30 interacts with the backbone carbonyl of F26 (i-4 position), which together with G27 may contribute to the bending of TM1.52 This bending is important for alternating between the inward-open and outward-open states.27,50 As expected, mutant T30A has a moderate effect on transport (63% activity compared to wild-type). The removal of the T30 hydroxyl hydrogen bond with F26 carbonyl may indirectly strengthen the TM1 integrity and increase the rigidity of the hinge. The T30D mutation abolishes substrate uptake (1% activity), suggesting that the mutation critically alters TM1 hinge flexibility and reduces substrate binding through steric hindrance and electrostatics effect, or both.
I168 is located in TM5, 4.1 Å away from the ligand. This residue is adjacent to the hinge residue G167, suggesting that I168 has a more direct influence on TM5 hinge bending and the global conformational changes of hGLUT1 that are associated with transport. Three I168 mutations with varying potential impact on structure and function were examined. I168A significantly affected transport (18% activity), nonetheless exhibiting the weakest effect among the three mutants. Due to alanine’s high α-helix propensity, I168A has a direct influence on G167 that likely contributes to increased rigidity of TM5, which is critical for transport. In addition, mutant I168D also showed significant effect on transport (9% activity), likely by altering the electrostatic properties of the binding site. Finally, I168F abolishes substrate uptake (1% activity), likely due to disruption of substrate binding via a significant steric hindrance.
F291 is located at the bend of TM7 and does not interact with the substrate directly. Interestingly, the hGLUTs include residues with long or bulky sidechains at this position: phenylalanine appears in four hGLUTs; leucine appears in three hGLUTs and in XylE; tyrosine and methionine both appear in two hGLUTs; and asparagine appears in one hGLUT. In the outward-open hGLUT3 structure29 TM7 shifts away and places F291 aside, exposing the sugar-binding site to the extracellular environment, while in occluded and inward-open structures TM7 shifts in and places F291 over the sugar-binding site, shielding it from the extracellular environment. F291A showed a moderate decrease in transport of radiolabeled substrate (65% activity). This modification of the gate residue from a bulky sidechain to a small hydrophobic sidechain likely affects the efficiency of the gate. Moreover, the mutant F291H exhibited even lower substrate uptake (50% activity). In this modification, even though we used a bulky residue to replace F291, the introduction of a positive charge alters the local environment and the gate function.
Virtual screening of compound libraries
As an initial control experiment, we docked known substrates (D-xylose and D-glucose) to the binding site of the occluded XylE structure and the hGLUT1 models. The docking recapitulated the crystallographic binding pose in the XylE structures, increasing the confidence in our model and approach (Fig. S4). To identify putative, chemically unique ligands for hGLUT1 in the occluded conformation, we performed virtual screening using the NCI-2007 and the Fragment-Like library from the ZINC40 database against our hGLUT1 models. The 548,312 screened compounds from each of the three hGLUT1 models were ranked based on their docking score (median value of −9.4 with standard deviation of 1.2). The 250 top-ranking compounds from each model (0.05% of the screening set) obtained scores that are three standard deviations lower than the median, and were therefore further analyzed. In particular, because erroneous docking may occur in structure-based virtual screenings, we carefully inspected the docking poses of these top-ranking compounds to remove molecules with questionable pose or strained conformations, and prioritized them for experimental testing.33,48 We focused on molecules that interact with the conserved polar residues of the sugar-binding site (i.e., Q161, Q282, Q283, N288, N317, and W388), which correspond to residues in XylE that make similar interactions (i.e., Q168, Q288, Q289, N294, N325, and W392), as well as those molecules that interact with the H-pocket. We purchased 19 molecules for experimental testing.
Experimental characterization of ligands with cis-inhibition
19 molecules were initially tested at 50 μM for their ability to inhibit [3H]-2-DG uptake. Eight exhibited significant level of specific inhibition close to the known hGLUT1 inhibitor phloretin at 50 μM (Table S4). These newly identified hGLUT1 ligands are predicted to form critical hydrogen bonds similar to those of D-glucose, the primary substrate of hGLUT1. These eight confirmed hits also passed the PAINS filter that identifies frequent false positives, such as promiscuous binders, in cell-based assays (e.g., rhodanines).46 We further measured the IC50 values of these inhibitors (Table 1; Fig. S5). Notably, the ligand efficiency (LE) of these compounds is higher (Table 1; average LE = 0.368 ± 0.126 kcal/mol/HA) than those of the known hGLUT1 inhibitors, such as cytochalasin B and forskolin (LE = 0.265 and 0.270 kcal/mol/HA, respectively).53 One compound, PUG-1, exhibits IC50 of 450 nM (LE = 0.481 kcal/mol/HA), where all other ligands inhibit hGLUT1 with IC50 values in the low μM range (12–60 μM). Furthermore, six of the confirmed inhibitors are chemically different from any known hGLUT1 inhibitors, exhibiting very low Tanimoto coefficient (Tc) ranging from 0.089 to 0.153 (Table 1). This highlights the utility of structure-based virtual screening in exploring novel chemical spaces that would unlikely to be explored otherwise.
Table 1.
IC50 of screening hits in [3H]-2-deoxy-D-glucose uptake assay.

|

|

|

|
|
|---|---|---|---|---|
|
|
||||
| Name | PUG-1 | PUG-2 | PUG-3 | PUG-4 |
|
|
||||
| IC50(μM) | 0.45 | 58.6 | 27.1 | 30.0 |
|
|
||||
| LEa | 0.481 | 0.339 | 0.346 | 0.363 |
|
|
||||
| Tcb | 0.152 | 0.145 | 0.099 | 0.153 |
|
| ||||

|

|

|

|
|
|
|
||||
| Name | PUG-5 (Phlorizin) | PUG-6 | PUG-7 | PUG-8 |
|
|
||||
| IC50(μM) | 23.0 | 37.9 | 11.8 | 22.5 |
|
|
||||
| LE | 0.204 | 0.215 | 0.420 | 0.576 |
|
|
||||
| Tc | 0.531 (Phloretin) | 0.357 (Phloretin) | 0.097 | 0.089 |
LE as ligand efficiency
Tc marks the Tanimoto coefficient (Tc) of the query molecule to the closest known ligand of hGLUT1 found in the ChEMBL database, calculated using the ECFP-4 fingerprints. Tc value of < 0.35 suggests that the ligand is chemically novel hGLUT1 ligand
We divided the newly identified ligands into the following four groups based on their chemical structures: Group 1 (PUG-1 and -2) includes xanthine-derivative compounds. PUG-1 is a hypoxanthine compound that inhibits hGLUT1 significantly (IC50 of 450 nM). PUG-2 (9-phenylxanthine) has modest inhibitory effect (IC50 of 58 μM). In our model, PUG-1 and PUG-2 are predicted to form multiple hydrogen bonds with the sugar-binding site polar residues while also occupying the H-pocket (Figs. 2A; 4A; S5A, B), which may provide additional affinity and selectivity. Notably, PUG-1 is predicted to interact with TM10 through hydrogen-bonding with the indole group of W388. This arrangement may provide stronger binding affinity between the ligand and TM10, potentially increases its activation energy of conformation transition and locks the bound transporter in an inactive, occluded state.
Figure 4. Newly discovered ligands of hGLUT1.
Predicted binding mode of (A) PUG-1 and (B) PUG-4 with the hGLUT1 models in the occluded conformation. The ligands (green sticks) occupy similar space to that of the sugar in the XylE template structure, and make critical hydrogen bonds with key residues (yellow dotted lines). An alternative rotamer of F291 was also used in the virtual screening (gold sticks). These ligands inhibit radiolabeled 2-DG uptake with significant IC50 values, as measured in a CHO-hGLUT1 cell line.
Group 2 consists of N-substituted adenine-derivative compounds (PUG-3 and -4). Both members of this group share a purine core similar to adenine, and a hydrophobic substitution at the 1-amino position. These molecules have similar inhibitory effect on hGLUT1, with IC50 values of 27 μM and 30 μM for PUG-3 and PUG-4, respectively. Similar to Group 1 ligands, Group 2 ligands are predicted to form hydrogen bonds with the polar residues of the sugar-binding site through their adenine core, as well as van Der Waals interactions with the H-pocket (Figs. 4B; S5C, D).
Group 3 includes glycosylated polyphenolic compounds (PUG-5 and -6). This family of compounds possesses multiple phenolic groups and is likely to form hydrogen bonding similar to the natural hGLUT1 substrates. PUG-5, phlorizin, is an inhibitor of SGLT1 (Ki = 1 μM54), a member of the Na+-dependent glucose transporter family SLC5 and an analog of the approved diabetes drug canagliflozin and dapagliflozin.55 Importantly, it is a glycosylated analog of phloretin, an inhibitor of hGLUT1 with IC50 of 50 μM56; this finding supports our strategy with homology modeling and virtual screening. PUG-6 is similar to PUG-5, which includes a double bond between the phenol moieties. The ligands are predicted to extend from the sugar-binding site to the nearby exofacial channel, which is gated by the side chain of F291. Hence, it is plausible that each of the two ligands select for a distinct F291 conformer for binding (Fig. S5E, F; gold stick of F291).
Group 4 is the pyridazinone-like compounds (PUG-7 and -8). These compounds are small fragments with very high ratio of carbon-to-heteroatom that bear chemical features different from those described above. PUG-7 and PUG-8 do not possess hydrophobic moiety that can reach into the H-pocket to harness the additional binding affinity (Fig. S5G, H). Although these compounds are the most chemically novel compounds among the newly discovered ligands (PUG-7 and -8 have Tc of 0.097 and 0.089, respectively), they possess unusual chemical features such as the sulfur linkage and the hydroquinone-like moiety that may have unpredictable reactivity.
CONCLUSIONS
hGLUT1 is a facilitative membrane transporter that controls cellular concentrations of various sugars, such as D-glucose and D-galactose, and plays a key role in altered metabolism in cancer. Novel small molecule ligands for hGLUT1 are needed for further characterization of the physiological role of this transporter, as well as for developing drugs against various disorders. Here, we characterize hGLUT1 using an integrated computational and experimental approach that has been successfully used to discover ligands for a variety of other human solute carrier transporters.57–61 We analyzed crystal structures of hGLUT1 and its homolog structures, and generated models of hGLUT1 based on the occluded XylE structures. Previously uncharacterized residues were tested experimentally via site-directed mutagenesis and cis-inhibition assays. We then performed virtual screening of small molecule libraries against the occluded hGLUT1 models, which was followed by experimental testing with cis-inhibition assay, to identify previously unknown potent ligands for this transporter.
Three key findings are reported in this study. First, several previously unknown small molecule hGLUT1 ligands were identified at a high hit rate, six of which are chemically different from any other known hGLUT1 ligands (Table 1). This finding provides novel chemical tools to further characterize the cellular functions of this important transporter, as well as potential hits for future lead and drug development. Second, structural analysis of atomic structures of hGLUT1 homologs and the newly constructed models, which was followed by experimental testing with site-directed mutagenesis, revealed key residues and regions that may facilitate ligand binding (Fig. 2). For example, the H-pocket is a hydrophobic pocket that differs in its size and shape between the occluded and inward-open conformations, which may influence the association and dissociation of bound ligands during the substrate transport. This result provides a framework for generating conformation specific ligands with increased efficiency and affinity. Third, phlorizin, a known inhibitor of members of the SLC5 family SGLT1 and SGLT2, which are evolutionarily unrelated to the GLUTs, was identified as a potent hGLUT1 inhibitor. Interestingly, a phlorizin analog, canagliflozin, is an approved anti-SGLT1/2 drug for the treatment of diabetes.55 Taken together, this result suggests that both transporters may share additional ligands whose positive and negative therapeutic effects are mediated via polypharmacology and future drugs can potentially be rationally designed to bind both structurally distinct transporters.
Supplementary Material
Figure S1. Alignment of hGLUT1–4 and XylE.
Figure S2. hGLUT1 mRNA expression level in stable CHO cell line and [3H]-2-deoxy-D-glucose uptake validation.
Figure S3. Structures of mammalian GLUTs.
Figure S4. Comparison of docked and observed poses of known ligands.
Figure S5. Predicted binding modes of newly discovered hGLUT1 ligands.
Table S1. Sugar-binding site of various hGLUT1 and XylE structures.
Table S2. Candidates of sugar-binding site residue for site-directed mutagenesis.
Table S3. The sequences of the primers for mutagenesis.
Table S4. Specific inhibition of initial screening hits in [3H]-2-deoxy-D-glucose uptake assay.
Acknowledgments
This work has been supported in part by the National Institutes of Health (R01 GM108911) to A. Schlessinger and P. Ung, Department of Defense (W81XWH-15-1-0539) to A. Schlessinger, and the National Natural Science Foundation of China (No. 81470839) to W. Song, L. Cheng, X. Zhao, H. Hu, and L. Chen. We thank the generous supply of the experimental chemicals by the National Cancer Institute at the National Institute of Health. We thank C. Colas (Mount Sinai) and G. Madej (UCLA) for helpful discussions. We appreciate OpenEye Scientific Software, Inc. for granting us access to its high-performance molecular modeling applications through its academic license program. This work was supported in part through the computational resources and staff expertise provided by the Department of Scientific Computing at the Icahn School of Medicine at Mount Sinai.
Footnotes
This material is available free of charge via the Internet at http://pubs.acs.org.
References
- 1.Maher F, Vannucci SJ, Simpson IA. Glucose transporter proteins in brain. FASEB J. 1994;8:1003–1011. doi: 10.1096/fasebj.8.13.7926364. [DOI] [PubMed] [Google Scholar]
- 2.Kasahara M, Hinkle PC. Reconstitution and purification of the D-glucose transporter from human erythrocytes. Journal of Biological Chemistry. 1977;252:7384–7390. [PubMed] [Google Scholar]
- 3.Thorens B, Mueckler M. Glucose transporters in the 21st Century. Am J Physiol Endocrinol Metab. 2010;298:E141–145. doi: 10.1152/ajpendo.00712.2009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Mueckler M, Thorens B. The SLC2 (GLUT) family of membrane transporters. Mol Aspects Med. 2013;34:121–138. doi: 10.1016/j.mam.2012.07.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Macintyre AN, Gerriets VA, Nichols AG, Michalek RD, Rudolph MC, Deoliveira D, Anderson SM, Abel ED, Chen BJ, Hale LP, Rathmell JC. The glucose transporter Glut1 is selectively essential for CD4 T cell activation and effector function. Cell Metab. 2014;20:61–72. doi: 10.1016/j.cmet.2014.05.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Weber YG, Storch A, Wuttke TV, Brockmann K, Kempfle J, Maljevic S, Margari L, Kamm C, Schneider SA, Huber SM, Pekrun A, Roebling R, Seebohm G, Koka S, Lang C, Kraft E, Blazevic D, Salvo-Vargas A, Fauler M, Mottaghy FM, Munchau A, Edwards MJ, Presicci A, Margari F, Gasser T, Lang F, Bhatia KP, Lehmann-Horn F, Lerche H. GLUT1 mutations are a cause of paroxysmal exertion-induced dyskinesias and induce hemolytic anemia by a cation leak. J Clin Invest. 2008;118:2157–2168. doi: 10.1172/JCI34438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Calvo MB, Figueroa A, Pulido EG, Campelo RG, Aparicio LA. Potential role of sugar transporters in cancer and their relationship with anticancer therapy. Int J Endocrinol 2010. 2010 doi: 10.1155/2010/205357. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Krzeslak A, Wojcik-Krowiranda K, Forma E, Jozwiak P, Romanowicz H, Bienkiewicz A, Brys M. Expression of GLUT1 and GLUT3 glucose transporters in endometrial and breast cancers. Pathol Oncol Res. 2012;18:721–728. doi: 10.1007/s12253-012-9500-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Warburg O, Wind F, Negelein E. The Metabolism of Tumors in the Body. J Gen Physiol. 1927;8:519–530. doi: 10.1085/jgp.8.6.519. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Vander Heiden MG, Cantley LC, Thompson CB. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science. 2009;324:1029–1033. doi: 10.1126/science.1160809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Larson SM, Schwartz LH. 18F-FDG PET as a candidate for “qualified biomarker”: functional assessment of treatment response in oncology. J Nucl Med. 2006;47:901–903. [PubMed] [Google Scholar]
- 12.Rastogi S, Banerjee S, Chellappan S, Simon GR. Glut-1 antibodies induce growth arrest and apoptosis in human cancer cell lines. Cancer Lett. 2007;257:244–251. doi: 10.1016/j.canlet.2007.07.021. [DOI] [PubMed] [Google Scholar]
- 13.Chan DA, Sutphin PD, Nguyen P, Turcotte S, Lai EW, Banh A, Reynolds GE, Chi JT, Wu J, Solow-Cordero DE, Bonnet M, Flanagan JU, Bouley DM, Graves EE, Denny WA, Hay MP, Giaccia AJ. Targeting GLUT1 and the Warburg effect in renal cell carcinoma by chemical synthetic lethality. Sci Transl Med. 2011;3:94ra70. doi: 10.1126/scitranslmed.3002394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Liu Y, Cao Y, Zhang W, Bergmeier S, Qian Y, Akbar H, Colvin R, Ding J, Tong L, Wu S, Hines J, Chen X. A small-molecule inhibitor of glucose transporter 1 downregulates glycolysis, induces cell-cycle arrest, and inhibits cancer cell growth in vitro and in vivo. Molecular Cancer Therapeutics. 2012;11:1672–1682. doi: 10.1158/1535-7163.MCT-12-0131. [DOI] [PubMed] [Google Scholar]
- 15.Rumsey SC, Kwon O, Xu GW, Burant CF, Simpson I, Levine M. Glucose transporter isoforms GLUT1 and GLUT3 transport dehydroascorbic acid. Journal of Biological Chemistry. 1997;272:18982–18989. doi: 10.1074/jbc.272.30.18982. [DOI] [PubMed] [Google Scholar]
- 16.Lefevre PG. Sugar transport in the red blood cell: structure-activity relationships in substrates and antagonists. Pharmacol Rev. 1961;13:39–70. [PubMed] [Google Scholar]
- 17.Sage JM, Cura AJ, Lloyd KP, Carruthers A. Caffeine inhibits glucose transport by binding at the GLUT1 nucleotide-binding site. Am J Physiol Cell Physiol. 2015;308:C827–834. doi: 10.1152/ajpcell.00001.2015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Naftalin RJ, Afzal I, Cunningham P, Halai M, Ross C, Salleh N, Milligan SR. Interactions of androgens, green tea catechins and the antiandrogen flutamide with the external glucose-binding site of the human erythrocyte glucose transporter GLUT1. Br J Pharmacol. 2003;140:487–499. doi: 10.1038/sj.bjp.0705460. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Frerichs H, Ball EG. Studies on the Metabolism of Adipose Tissue. Xvi. Inhibition by Phlorizin and Phloretin of the Insulin-Stimulated Uptake of Glucose. Biochemistry. 1964;3:981–985. doi: 10.1021/bi00895a024. [DOI] [PubMed] [Google Scholar]
- 20.Lu L, Lundqvist A, Zeng CM, Lagerquist C, Lundahl P. D-Glucose, forskolin and cytochalasin B affinities for the glucose transporter Glut1. Study of pH and reconstitution effects by biomembrane affinity chromatography. J Chromatogr A. 1997;776:81–86. doi: 10.1016/s0021-9673(97)00151-9. [DOI] [PubMed] [Google Scholar]
- 21.Carruthers A, DeZutter J, Ganguly A, Devaskar SU. Will the original glucose transporter isoform please stand up! Am J Physiol Endocrinol Metab. 2009;297:E836–848. doi: 10.1152/ajpendo.00496.2009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Wisedchaisri G, Park MS, Iadanza MG, Zheng H, Gonen T. Proton-coupled sugar transport in the prototypical major facilitator superfamily protein XylE. Nat Commun. 2014;5:4521. doi: 10.1038/ncomms5521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Jardetzky O. Simple allosteric model for membrane pumps. Nature. 1966;211:969–970. doi: 10.1038/211969a0. [DOI] [PubMed] [Google Scholar]
- 24.Radestock S, Forrest LR. The alternating-access mechanism of MFS transporters arises from inverted-topology repeats. J Mol Biol. 2011;407:698–715. doi: 10.1016/j.jmb.2011.02.008. [DOI] [PubMed] [Google Scholar]
- 25.Majumdar DS, Smirnova I, Kasho V, Nir E, Kong X, Weiss S, Kaback HR. Single-molecule FRET reveals sugar-induced conformational dynamics in LacY. Proc Natl Acad Sci U S A. 2007;104:12640–12645. doi: 10.1073/pnas.0700969104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Smirnova I, Kasho V, Kaback HR. Lactose permease and the alternating access mechanism. Biochemistry. 2011;50:9684–9693. doi: 10.1021/bi2014294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Deng D, Xu C, Sun P, Wu J, Yan C, Hu M, Yan N. Crystal structure of the human glucose transporter GLUT1. Nature. 2014;510:121–125. doi: 10.1038/nature13306. [DOI] [PubMed] [Google Scholar]
- 28.Pao SS, Paulsen IT, Saier MH., Jr Major facilitator superfamily. Microbiol Mol Biol Rev. 1998;62:1–34. doi: 10.1128/mmbr.62.1.1-34.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Deng D, Sun P, Yan C, Ke M, Jiang X, Xiong L, Ren W, Hirata K, Yamamoto M, Fan S, Yan N. Molecular basis of ligand recognition and transport by glucose transporters. Nature. 2015;526:391–396. doi: 10.1038/nature14655. [DOI] [PubMed] [Google Scholar]
- 30.Nomura N, Verdon G, Kang HJ, Shimamura T, Nomura Y, Sonoda Y, Hussien SA, Qureshi AA, Coincon M, Sato Y, Abe H, Nakada-Nakura Y, Hino T, Arakawa T, Kusano-Arai O, Iwanari H, Murata T, Kobayashi T, Hamakubo T, Kasahara M, Iwata S, Drew D. Structure and mechanism of the mammalian fructose transporter GLUT5. Nature. 2015;526:397–401. doi: 10.1038/nature14909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Quistgaard EM, Low C, Moberg P, Tresaugues L, Nordlund P. Structural basis for substrate transport in the GLUT-homology family of monosaccharide transporters. Nat Struct Mol Biol. 2013;20:766–768. doi: 10.1038/nsmb.2569. [DOI] [PubMed] [Google Scholar]
- 32.Sun L, Zeng X, Yan C, Sun X, Gong X, Rao Y, Yan N. Crystal structure of a bacterial homologue of glucose transporters GLUT1–4. Nature. 2012;490:361–366. doi: 10.1038/nature11524. [DOI] [PubMed] [Google Scholar]
- 33.Shoichet BK. Virtual screening of chemical libraries. Nature. 2004;432:862–865. doi: 10.1038/nature03197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.UniProt C. The universal protein resource (UniProt) Nucleic Acids Res. 2008;36:D190–195. doi: 10.1093/nar/gkm895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004;32:1792–1797. doi: 10.1093/nar/gkh340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Sali A, Blundell TL. Comparative protein modelling by satisfaction of spatial restraints. J Mol Biol. 1993;234:779–815. doi: 10.1006/jmbi.1993.1626. [DOI] [PubMed] [Google Scholar]
- 37.Shen MY, Sali A. Statistical potential for assessment and prediction of protein structures. Protein Sci. 2006;15:2507–2524. doi: 10.1110/ps.062416606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Durrant JD, Votapka L, Sorensen J, Amaro RE. POVME 2.0: An Enhanced Tool for Determining Pocket Shape and Volume Characteristics. J Chem Theory Comput. 2014;10:5047–5056. doi: 10.1021/ct500381c. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Schrödinger Release 2015-2: Maestro, 10.2. Schrödinger, LLC; New York, NY, USA: http://www.schrodinger.com. [Google Scholar]
- 40.Irwin JJ, Shoichet BK. ZINC--a free database of commercially available compounds for virtual screening. J Chem Inf Model. 2005;45:177–182. doi: 10.1021/ci049714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Hawkins PC, Skillman AG, Warren GL, Ellingson BA, Stahl MT. Conformer generation with OMEGA: algorithm and validation using high quality structures from the Protein Databank and Cambridge Structural Database. J Chem Inf Model. 2010;50:572–584. doi: 10.1021/ci100031x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.OEDocking, 3.0.1; OMEGA, 2.5.1.4. OpenEye Scientific Software; Santa Fe, NM, USA: http://www.eyesopen.com. [Google Scholar]
- 43.McGann M. FRED and HYBRID docking performance on standardized datasets. J Comput Aided Mol Des. 2012;26:897–906. doi: 10.1007/s10822-012-9584-8. [DOI] [PubMed] [Google Scholar]
- 44.Gaulton A, Bellis LJ, Bento AP, Chambers J, Davies M, Hersey A, Light Y, McGlinchey S, Michalovich D, Al-Lazikani B, Overington JP. ChEMBL: a large-scale bioactivity database for drug discovery. Nucleic Acids Res. 2012;40:D1100–1107. doi: 10.1093/nar/gkr777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Varani K, Portaluppi F, Merighi S, Ongini E, Belardinelli L, Borea PA. Caffeine alters A2A adenosine receptors and their function in human platelets. Circulation. 1999;99:2499–2502. doi: 10.1161/01.cir.99.19.2499. [DOI] [PubMed] [Google Scholar]
- 46.Baell JB, Holloway GA. New substructure filters for removal of pan assay interference compounds (PAINS) from screening libraries and for their exclusion in bioassays. Journal of Medicinal Chemistry. 2010;53:2719–2740. doi: 10.1021/jm901137j. [DOI] [PubMed] [Google Scholar]
- 47.Chen L, Durkin KA, Casida JE. Structural model for gamma-aminobutyric acid receptor noncompetitive antagonist binding: widely diverse structures fit the same site. Proc Natl Acad Sci U S A. 2006;103:5185–5190. doi: 10.1073/pnas.0600370103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Carlsson J, Coleman RG, Setola V, Irwin JJ, Fan H, Schlessinger A, Sali A, Roth BL, Shoichet BK. Ligand discovery from a dopamine D3 receptor homology model and crystal structure. Nat Chem Biol. 2011;7:769–778. doi: 10.1038/nchembio.662. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Schlessinger A, Wittwer MB, Dahlin A, Khuri N, Bonomi M, Fan H, Giacomini KM, Sali A. High selectivity of the gamma-aminobutyric acid transporter 2 (GAT-2, SLC6A13) revealed by structure-based approach. Journal of Biological Chemistry. 2012;287:37745–37756. doi: 10.1074/jbc.M112.388157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Madej MG, Sun L, Yan N, Kaback HR. Functional architecture of MFS D-glucose transporters. Proc Natl Acad Sci U S A. 2014;111:E719–727. doi: 10.1073/pnas.1400336111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Eramian D, Eswar N, Shen MY, Sali A. How well can the accuracy of comparative protein structure models be predicted? Protein Sci. 2008;17:1881–1893. doi: 10.1110/ps.036061.108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Ballesteros JA, Deupi X, Olivella M, Haaksma EE, Pardo L. Serine and threonine residues bend alpha-helices in the chi(1) = g(-) conformation. Biophys J. 2000;79:2754–2760. doi: 10.1016/S0006-3495(00)76514-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Hellwig B, Joost HG. Differentiation of erythrocyte-(GLUT1), liver-(GLUT2), and adipocyte-type (GLUT4) glucose transporters by binding of the inhibitory ligands cytochalasin B, forskolin, dipyridamole, and isobutylmethylxanthine. Mol Pharmacol. 1991;40:383–389. [PubMed] [Google Scholar]
- 54.Diedrich DF. Competitive inhibition of intestinal glucose transport by phlorizin analogs. Arch Biochem Biophys. 1966;117:248–256. doi: 10.1016/0003-9861(66)90409-7. [DOI] [PubMed] [Google Scholar]
- 55.Chao EC, Henry RR. SGLT2 inhibition--a novel strategy for diabetes treatment. Nat Rev Drug Discov. 2010;9:551–559. doi: 10.1038/nrd3180. [DOI] [PubMed] [Google Scholar]
- 56.Kasahara T, Kasahara M. Expression of the rat GLUT1 glucose transporter in the yeast Saccharomyces cerevisiae. Biochem J. 1996;315(Pt 1):177–182. doi: 10.1042/bj3150177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Schlessinger A, Matsson P, Shima JE, Pieper U, Yee SW, Kelly L, Apeltsin L, Stroud RM, Ferrin TE, Giacomini KM, Sali A. Comparison of human solute carriers. Protein Sci. 2010;19:412–428. doi: 10.1002/pro.320. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Schlessinger A, Geier E, Fan H, Irwin JJ, Shoichet BK, Giacomini KM, Sali A. Structure-based discovery of prescription drugs that interact with the norepinephrine transporter, NET. Proc Natl Acad Sci U S A. 2011;108:15810–15815. doi: 10.1073/pnas.1106030108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Colas C, Grewer C, Otte NJ, Gameiro A, Albers T, Singh K, Shere H, Bonomi M, Holst J, Schlessinger A. Ligand Discovery for the Alanine-Serine-Cysteine Transporter (ASCT2, SLC1A5) from Homology Modeling and Virtual Screening. PLoS Comput Biol. 2015;11:e1004477. doi: 10.1371/journal.pcbi.1004477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Colas C, Pajor AM, Schlessinger A. Structure-Based Identification of Inhibitors for the SLC13 Family of Na(+)/Dicarboxylate Cotransporters. Biochemistry. 2015;54:4900–4908. doi: 10.1021/acs.biochem.5b00388. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Geier EG, Schlessinger A, Fan H, Gable JE, Irwin JJ, Sali A, Giacomini KM. Structure-based ligand discovery for the Large-neutral Amino Acid Transporter 1, LAT-1. Proc Natl Acad Sci U S A. 2013;110:5480–5485. doi: 10.1073/pnas.1218165110. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Figure S1. Alignment of hGLUT1–4 and XylE.
Figure S2. hGLUT1 mRNA expression level in stable CHO cell line and [3H]-2-deoxy-D-glucose uptake validation.
Figure S3. Structures of mammalian GLUTs.
Figure S4. Comparison of docked and observed poses of known ligands.
Figure S5. Predicted binding modes of newly discovered hGLUT1 ligands.
Table S1. Sugar-binding site of various hGLUT1 and XylE structures.
Table S2. Candidates of sugar-binding site residue for site-directed mutagenesis.
Table S3. The sequences of the primers for mutagenesis.
Table S4. Specific inhibition of initial screening hits in [3H]-2-deoxy-D-glucose uptake assay.