ABSTRACT
This study characterized cefoxitin-resistant and -susceptible Salmonella enterica serovar Heidelberg strains from humans, abattoir poultry, and retail poultry to assess the molecular relationships of isolates from these sources in Québec in 2012. Isolates were collected as part of the Canadian Integrated Program for Antimicrobial Resistance Surveillance (CIPARS). All isolates were subjected to antimicrobial susceptibility testing, PCR for CMY-2, pulsed-field gel electrophoresis (PFGE), and whole-genome sequencing (WGS). A total of 113 S. Heidelberg isolates from humans (n = 51), abattoir poultry (n = 18), and retail poultry (n = 44) were studied. All cefoxitin-resistant isolates (n = 65) were also resistant to amoxicillin-clavulanic acid, ampicillin, ceftiofur, and ceftriaxone, and all contained the CMY-2 gene. PFGE analysis showed that 111/113 (98.2%) isolates clustered together with ≥90% similarity. Core genome analysis using WGS identified 13 small clusters of isolates with 0 to 4 single nucleotide variations (SNVs), consisting of cefoxitin-resistant and -susceptible human, abattoir poultry, and retail poultry isolates. CMY-2 plasmids from cefoxitin-resistant isolates all belonged to incompatibility group I1. Analysis of IncI1 plasmid sequences revealed high identity (95 to 99%) to a previously described plasmid (pCVM29188_101) found in Salmonella Kentucky. When compared to pCVM29188_101, all sequenced cefoxitin-resistant isolates were found to carry 1 of 10 possible variant plasmids. Transmission of S. Heidelberg may be occurring between human, abattoir poultry, and retail poultry sources, and transmission of a common CMY-2 plasmid may be occurring among S. Heidelberg strains with variable genetic backgrounds.
KEYWORDS: Québec, Salmonella Heidelberg, beta-lactamase, cefoxitin, plasmid analysis, whole-genome sequencing
INTRODUCTION
Nontyphoidal Salmonella enterica subsp. enterica is a major cause of human gastroenteritis worldwide (1). In Canada, Salmonella enterica serovar Heidelberg is the third most frequently isolated serovar in humans, behind S. Enteritidis and S. Typhimurium, which rank as numbers 1 and 2, respectively (2). S. Heidelberg is often more invasive than other nontyphoidal serovars and is capable of causing extraintestinal infections and severe septicemia (3). In 2012, 9.24% of S. Heidelberg isolates in Canada were recovered from extraintestinal samples in comparison to 3.97% of S. Enteriditis and 2.21% of S. Typhimurium extraintestinal isolates from the same year (4). In such invasive infections, antimicrobial treatment is important to ensure positive patient outcomes.
Infection with S. Heidelberg is linked primarily to the consumption of poultry and is rarely transmitted from person to person (3, 5). Within the Canadian agriculture sector, this serovar is repeatedly isolated from farm, abattoir, and retail poultry samples but less frequently from bovine and porcine samples (6).
Extended-spectrum cephalosporins are used in veterinary medicine for the treatment and prevention of disease in livestock (7). In Canada, ceftiofur, a third-generation cephalosporin, has been employed in an extralabel manner in broiler chickens to prevent omphalitis caused by Escherichia coli. The Canadian Integrated Program for Antimicrobial Resistance Surveillance (CIPARS) (8) observed a strong correlation (r = 0.91, P < 0.0001) between the prevalence of ceftiofur-resistant S. Heidelberg isolates found in retail chicken samples and those found in humans (9). After a voluntary withdrawal of the antimicrobial by provincial hatcheries in 2005, a decrease in ceftiofur-resistant S. Heidelberg isolates from chicken meat and humans was observed, which was subsequently followed by an increase in resistant isolates in 2007 that paralleled a partial reinstitution of ceftiofur use (9). Classified by Health Canada as antimicrobials of “high importance” (category 2) (cefoxitin) and “very high importance” (category 1) (ceftriaxone) in human medicine, cefoxitin and ceftriaxone are broad-spectrum second-generation and third-generation cephalosporins, respectively, that are used to treat a wide variety of infections (9, 10).
Resistance of microorganisms to beta-lactam antimicrobials, including cephalosporins, is commonly due to the production of beta-lactamase enzymes that hydrolyze the antimicrobial (11). AmpC beta-lactamases, which belong to Ambler class C enzymes, encode resistance to penicillins, cephalosporins (including cephamycins), and monobactams (12). Although AmpC genes are found intrinsically in some microorganisms, in Salmonella they are due to acquired mechanisms (13). The AmpC beta-lactamase blaCMY-2, which is commonly found on plasmids, encodes resistance to cefoxitin and is a significant source of beta-lactam resistance in Salmonella found worldwide (14, 15). The blaCMY-2-containing plasmids can also harbor resistance genes to other classes of antimicrobials, thus potentially conferring multidrug resistance (13).
In this study, we examined the molecular characteristics of S. Heidelberg to assess the relationships of isolates from human, abattoir poultry, and retail poultry sources from Québec by using high-quality core genome single-nucleotide variant (hqSNV) analysis and blaCMY-2 plasmid analysis.
RESULTS
Initial characterization of isolates.
There were 128 S. Heidelberg isolates tested from human sources, with 31 (24.2%) resistant to cefoxitin, identified in Québec in 2012. A total of 113 isolates (Table 1) were examined in this study, of which 31 of 51 (60.8%) human isolates were cefoxitin resistant. Thirty-six (70.6%) human isolates were identified from stool, 11 (21.6%) from blood, and 1 (2.0%) from urine, and 3 (5.9%) were of unknown origin. All 18 of the selected abattoir poultry isolates were recovered from chicken cecal content, of which 10 (55.6%) were resistant to cefoxitin. Among the 44 retail poultry isolates examined, 24 (54.5%) were cefoxitin resistant. Thirty-one (70.5%) retail isolates were from a sample of fresh meat (24 from chicken, 7 from turkey), while the remaining 13 (29.5%) were from prepackaged, frozen chicken products (e.g., nuggets).
TABLE 1.
Characteristics of Salmonella Heidelberg identified in this study
| Strain ID | Provincea | Yr | Source | Resistance phenotypesb | Resistance genotypesc | STd | Plasmid(s)e | Size of blaCMY-2 plasmid contig(s) (bp)f | ST of blaCMY-2 plasmidg | Group of blaCMY-2 plasmidh | No. on minimum spanning tree |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 12-0315 | QC | 2012 | Human stool | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColRNAI | 100,217; 1,331 | 12 | C | 2 |
| 12-0467 | QC | 2012 | Human stool | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1 | 45,026; 27,243; 20,599; 4,881; 1,656 | 12 | A | 4 |
| 12-0469 | QC | 2012 | Human stool | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC | 65,621; 24,827; 7,381; 1,656 | 12 | A | 5 |
| 12-1195 | QC | 2012 | Human stool | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC | 48,418; 28,410; 23,779; 1,869; 1,759 | 12 | 8 | |
| 12-2458 | QC | 2012 | Human stool | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC | 100,376; 1,108 | 12 | C | 12 |
| 12-2460 | QC | 2012 | Human stool | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColRNAI | 99,125 | 12 | C | 13 |
| 12-2552 | QC | 2012 | Human stool | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC, Col8282 | 103,169; 1,108 | Untypeable | C | 14 |
| 12-2554 | QC | 2012 | Human stool | AMC, AMP, FOX, TIO, CRO | blaCMY-2, blaTEM-1B | 15 | IncI1, IncX1, Col156 | 71,837; 19,355 | 2 | 15 | |
| 12-2694 | QC | 2012 | Human stool | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC, ColRNAI | 65,427; 27,778; 3,694 | 12 | A | 16 |
| 12-3755 | QC | 2012 | Human stool | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC | 97,373 | 12 | A | 1 |
| 12-4179 | QC | 2012 | Human stool | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC | 99,601 | 12 | C | 24 |
| 12-4374 | QC | 2012 | Human stool | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC, IncFII | 96,728 | 12 | A | 1 |
| 12-4585 | QC | 2012 | Human stool | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC, IncFII | 96,616 | 12 | A | 1 |
| 12-5152 | QC | 2012 | Human stool | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC | 67,970; 27,244; 1,656 | 12 | A | 26 |
| 12-5444 | QC | 2012 | Human stool | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC | 95,989; 1,108 | 12 | A | 29 |
| 12-6245 | QC | 2012 | Human stool | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | ColRNAI | Plasmid integrated into chromosome | Plasmid integrated into chromosome | NAh | 33 |
| 12-6507 | QC | 2012 | Human stool | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, ColpVC, ColRNAI | 96,095; 1,116 | 12 | A | 35 |
| 12-7080 | QC | 2012 | Human stool | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC | 96,725 | 12 | A | 1 |
| 12-7329 | QC | 2012 | Human stool | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC, ColRNAI | 99,244 | 12 | 38 | |
| 12-1016 | QC | 2012 | Human stool | AMC, AMP, FOX, TIO, CRO, (CHL) | blaCMY-2 | 15 | IncI1, IncX1, ColpVC, ColRNAI | 96,047; 1,108 | 12 | A | 6 |
| 12-5643 | QC | 2012 | Human stool | AMC, AMP, FOX, TIO, CRO, NAL | blaCMY-2 | 15 | IncI1, IncX1 | 94,220; 4,699; 1,110 | 12 | A | 32 |
| 12-1667 | QC | 2012 | Human stool | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 10 |
| 12-2695 | QC | 2012 | Human stool | Susceptible | None | 15 | IncX1, Col8282 | NA | NA | NA | 17 |
| 12-3136 | QC | 2012 | Human stool | Susceptible | None | 15 | IncX1 | NA | NA | NA | 17 |
| 12-3227 | QC | 2012 | Human stool | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 18 |
| 12-3327 | QC | 2012 | Human stool | Susceptible | None | 15 | IncX1 | NA | NA | NA | 17 |
| 12-3383 | QC | 2012 | Human stool | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 20 |
| 12-3458 | QC | 2012 | Human stool | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 20 |
| 12-3461 | QC | 2012 | Human stool | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 20 |
| 12-4367 | QC | 2012 | Human stool | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 25 |
| 12-5334 | QC | 2012 | Human stool | Susceptible | None | 15 | IncFII | NA | NA | NA | 27 |
| 12-5632 | QC | 2012 | Human stool | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 31 |
| 12-5634 | QC | 2012 | Human stool | Susceptible | None | 15 | IncX1, ColRNAI | NA | NA | NA | 13 |
| 12-6510 | QC | 2012 | Human stool | Susceptible | None | 15 | IncX1, ColpVC, ColRNAI | NA | NA | NA | 36 |
| 12-7145 | QC | 2012 | Human stool | Susceptible | None | 15 | IncX1, ColRNAI | NA | NA | NA | 13 |
| 12-7730 | QC | 2012 | Human stool | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 39 |
| 12-1063 | QC | 2012 | Human blood | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1 | 81,373; 13,154; 4,699; 1,108 | 12 | A | 7 |
| 12-1847 | QC | 2012 | Human blood | AMC, AMP, FOX, TIO, CRO | blaCMY-2, blaTEM-1B | 15 | IncI1, IncX1 | 103,791 | 12 | B | 11 |
| 12-1959 | QC | 2012 | Human blood | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC, ColRNAI | 96,095; 1,108 | 12 | A | 6 |
| 12-3918 | QC | 2012 | Human blood | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1 | 69,872; 27,889; 3,871 | 12 | A | 23 |
| 12-5335 | QC | 2012 | Human blood | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColRNAI | 99,577 | 12 | C | 28 |
| 12-7092 | QC | 2012 | Human blood | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1 | 99,675 | 12 | C | 34 |
| 12-0466 | QC | 2012 | Human blood | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 3 |
| 12-3330 | QC | 2012 | Human blood | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 19 |
| 12-5542 | QC | 2012 | Human blood | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 30 |
| 12-6342 | QC | 2012 | Human blood | Susceptible | None | 15 | IncX1, ColRNAI | NA | NA | NA | 34 |
| 12-7327 | QC | 2012 | Human blood | Susceptible | None | 15 | ColRNAI | NA | NA | NA | 37 |
| 13-0067 | QC | 2012 | Human urine | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, ColpVC, ColRNAI | 99,067; 1,108 | 12 | C | 40 |
| 12-1666 | QC | 2012 | Human (unknown) | AMC, AMP, FOX, TIO, CRO | blaCMY-2, blaTEM-1B | 15 | IncI1, IncX1 | 101,107 | 12 | A | 9 |
| 12-3757 | QC | 2012 | Human (unknown) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC | 96,280 | 12 | A | 21 |
| 12-3792 | QC | 2012 | Human (unknown) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1 | 100,625; 1,219 | 12 | A | 22 |
| N13-02936 | ON | 2011 | Animal cecal content (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColRNAI | 99,572 | 12 | C | 88 |
| N13-02941 | ON | 2011 | Animal cecal content (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1 | 67,413; 27,194; 1,658; 1,215 | 12 | A | 89 |
| N13-02943 | ON | 2011 | Animal cecal content (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC | 98,544; 1,214 | 12 | C | 90 |
| N13-02944 | NB | 2012 | Animal cecal content (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC | 96,280 | 12 | A | 91 |
| N13-02945 | ON | 2012 | Animal cecal content (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1 | 96,628 | 12 | A | 92 |
| N13-02946 | NB | 2012 | Animal cecal content (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1 | 96,123; 1,108 | 12 | A | 93 |
| N13-02947 | ON | 2012 | Animal cecal content (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColRNAI | 96,740 | 12 | A | 94 |
| N13-02948 | ON | 2012 | Animal cecal content (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColRNAI | 95,999; 1,108 | 12 | A | 94 |
| N13-02949 | ON | 2012 | Animal cecal content (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC | 98,943; 1,108 | 12 | C | 95 |
| N13-02934 | NB | 2011 | Animal cecal content (chicken) | AMC, AMP, FOX, TIO, CRO, CHL, SUL | aadA1, aadA2, blaCMY-2, cmlA1, sul3 | 15 | IncI1, IncX1, ColpVC | 112,670 | 26 | 87 | |
| N13-01291 | QC | 2012 | Animal cecal content (chicken) | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 42 |
| N13-01311 | QC | 2012 | Animal cecal content (chicken) | Susceptible | None | 15 | IncX1 | NA | NA | NA | 55 |
| N13-01312 | QC | 2012 | Animal cecal content (chicken) | Susceptible | None | 15 | IncX1, IncX4 | NA | NA | NA | 56 |
| N13-01323 | QC | 2012 | Animal cecal content (chicken) | Susceptible | None | 15 | IncX1, ColpVC, ColRNAI | NA | NA | NA | 64 |
| N13-01324 | QC | 2012 | Animal cecal content (chicken) | Susceptible | None | 15 | IncX1, ColpVC, ColRNAI | NA | NA | NA | 65 |
| N13-01325 | QC | 2012 | Animal cecal content (chicken) | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 66 |
| N13-01330 | QC | 2012 | Animal cecal content (chicken) | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 70 |
| N13-01348 | QC | 2012 | Animal cecal content (chicken) | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 79 |
| N13-01290 | QC | 2012 | Food aliquot (turkey) | AMC, AMP, FOX, TIO, CRO, STR, SUL, TET | aadA1, blaCMY-2, blaTEM-1B, strA, strB, sul1, tetB | 15 | IncI1, ColpVC, ColRNAI, Col156, IncHI2, IncHI2A | 100,326 | 12 | C | 41 |
| N13-01313 | QC | 2012 | Food aliquot (turkey) | AMC, AMP, FOX, TIO, CRO, STR, SUL, TET | aadA1, blaCMY-2, blaTEM-1B, strA, strB, sul1, tetB | 15 | IncI1, ColpVC, ColRNAI, Col156, IncHI2, IncHI2A | 82,151; 12,485; 5,833 | 25 | 57 | |
| N13-01327 | QC | 2012 | Food aliquot (turkey) | AMC, AMP, FOX, TIO, CRO, TET | blaCMY-2, blaTEM-1B, tetB | 15 | IncI1, IncX1 | 103,902 | Untypeable | 68 | |
| N13-01354 | QC | 2012 | Food aliquot (turkey) | AMC, AMP, FOX, TIO, CRO, STR | blaCMY-2, blaTEM-1B, strA | 15 | IncI1, IncX1 | 103,768 | Untypeable | C | 84 |
| N13-01326 | QC | 2012 | Food aliquot (turkey) | AMP | blaTEM-1B | 15 | IncI1, IncX1, ColRNAI | NA | NA | NA | 67 |
| N13-01331 | QC | 2012 | Food aliquot (turkey) | TET | tetB | 15 | IncI1 | NA | NA | NA | 71 |
| N13-01366 | QC | 2012 | Food aliquot (turkey) | TET | tetB | 15 | IncI1 | NA | NA | NA | 86 |
| N13-01292 | QC | 2012 | Food aliquot (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1 | 100,044; 1,214 | 12 | A | 43 |
| N13-01295 | QC | 2012 | Food aliquot (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1 | 100,327; 1,210 | 12 | C | 28 |
| N13-01297 | QC | 2012 | Food aliquot (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1 | 42,796; 27,247; 20,599; 5,280; 3,886 | 12 | A | 47 |
| N13-01314 | QC | 2012 | Food aliquot (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC | 96,628; 1,219 | 12 | A | 58 |
| N13-01315 | QC | 2012 | Food aliquot (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC | 97,364 | 12 | A | 59 |
| N13-01316 | QC | 2012 | Food aliquot (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC, ColRNAI | 96,083; 1,108 | 12 | A | 60 |
| N13-01320 | QC | 2012 | Food aliquot (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | ColRNAI, Col156 | Plasmid integrated into chromosome | Plasmid integrated into chromosome | NA | 62 |
| N13-01321 | QC | 2012 | Food aliquot (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1 | 132,420 | 66 | 63 | |
| N13-01336 | QC | 2012 | Food aliquot (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColRNAI | 99,636; 1,214 | 12 | C | 74 |
| N13-01342 | QC | 2012 | Food aliquot (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC, Col156 | 98,725 | 12 | C | 77 |
| N13-01352 | QC | 2012 | Food aliquot (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColRNAI | 100,330 | 12 | C | 82 |
| N13-01333 | QC | 2012 | Food aliquot (chicken) | AMP | blaTEM-1B | 15 | IncI1, IncX1, ColpVC, ColRNAI, Col8282 | NA | NA | NA | 73 |
| N13-01293 | QC | 2012 | Food aliquot (chicken) | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 44 |
| N13-01294 | QC | 2012 | Food aliquot (chicken) | Susceptible | None | 15 | IncX1, ColpVC, ColRNAI | NA | NA | NA | 45 |
| N13-01296 | QC | 2012 | Food aliquot (chicken) | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 46 |
| N13-01298 | QC | 2012 | Food aliquot (chicken) | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 48 |
| N13-01317 | QC | 2012 | Food aliquot (chicken) | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 61 |
| N13-01329 | QC | 2012 | Food aliquot (chicken) | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 69 |
| N13-01332 | QC | 2012 | Food aliquot (chicken) | Susceptible | None | 15 | None | NA | NA | NA | 72 |
| N13-01337 | QC | 2012 | Food aliquot (chicken) | Susceptible | None | 15 | ColpVC | NA | NA | NA | 75 |
| N13-01338 | QC | 2012 | Food aliquot (chicken) | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 76 |
| N13-01346 | QC | 2012 | Food aliquot (chicken) | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 78 |
| N13-01351 | QC | 2012 | Food aliquot (chicken) | Susceptible | None | 15 | None | NA | NA | NA | 81 |
| N13-01353 | QC | 2012 | Food aliquot (chicken) | Susceptible | None | 15 | IncX1, ColpVC, ColRNAI | NA | NA | NA | 83 |
| N13-01303 | QC | 2012 | Food prepackaged (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC | 99,670 | 12 | C | 50 |
| N13-01304 | QC | 2012 | Food prepackaged (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1 | 73,672; 20,599; 5,280; 1,108 | 12 | A | 51 |
| N13-01307 | QC | 2012 | Food prepackaged (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC, ColRNAI | 96,083; 1,108 | 12 | A | 16 |
| N13-01308 | QC | 2012 | Food prepackaged (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC, ColRNAI | 96,616 | 12 | A | 16 |
| N13-01309 | QC | 2012 | Food prepackaged (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColRNAI | 99,236 | 12 | C | 54 |
| N13-01318 | QC | 2012 | Food prepackaged (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2, blaTEM-1B | 15 | IncI1, IncX1 | 103,897 | 12 | B | 15 |
| N13-01319 | QC | 2012 | Food prepackaged (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2, blaTEM-1B | 15 | IncI1, IncX1 | 104,403 | 12 | B | 15 |
| N13-01349 | QC | 2012 | Food prepackaged (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1, ColpVC | 97,261 | 12 | A | 80 |
| N13-01355 | QC | 2012 | Food prepackaged (chicken) | AMC, AMP, FOX, TIO, CRO | blaCMY-2 | 15 | IncI1, IncX1 | 100,219 | 12 | C | 85 |
| N13-01301 | QC | 2012 | Food prepackaged (chicken) | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 49 |
| N13-01305 | QC | 2012 | Food prepackaged (chicken) | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 52 |
| N13-01306 | QC | 2012 | Food prepackaged (chicken) | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 53 |
| N13-01322 | QC | 2012 | Food prepackaged (chicken) | Susceptible | None | 15 | IncX1, ColpVC | NA | NA | NA | 20 |
NB, New Brunswick; ON, Ontario; QC, Québec.
AMC, amoxicillin-clavulanic acid; AMP, ampicillin; FOX, cefoxitin; TIO, ceftiofur; CRO, ceftriaxone; CHL, chloramphenicol; NAL, nalidixic acid; STR, streptomycin; SUL, sulfisoxazole; TET, tetracycline. Parentheses indicate intermediate resistance.
aadA1-aadA2, streptomycin adenylytransferase; blaCMY-2, beta-lactamase; blaTEM-1B, beta-lactamase; cmlA1, chloramphenicol efflux; strA-strB, streptomycin phosphotransferase; sul1-sul3, dihydropteroate synthase; tetB, tetracycline efflux. Determined using the Centre for Genomic Epidemiology's ResFinder program.
Determined using the Centre for Genomic Epidemiology's MLST program.
Determined using the Centre for Genomic Epidemiology's PlasmidFinder program.
Determined using Contiguator.
Determined using the Centre for Genomic Epidemiology's pMLST program.
NA, not applicable.
There were 65/113 (57.5%) isolates resistant to amoxicillin-clavulanic acid, ampicillin, cefoxitin, ceftiofur, and ceftriaxone combined. Of these, 6 (9.2%) were also resistant to at least one of the following antimicrobials: nalidixic acid, chloramphenicol, sulfisoxazole, streptomycin, and tetracycline (Table 1). All 65 cefoxitin-resistant isolates were found to harbor blaCMY-2 using PCR. Nine (13.8%) also tested positive for blaTEM, and none of the isolates had blaSHV, blaCTX-M, or blaOXA-1.
Macrorestriction analysis of S. Heidelberg.
Macrorestriction analysis using pulsed-field gel electrophoresis (PFGE) of the 113 isolates revealed that all were closely related with ≥80% similarity, with 111 (98.2%) clustering at ≥90% similarity (see Fig. S1 in the supplemental material). Overall there were a total of 16 groups, labeled A1 to A16, where A1 to A4 contained the majority of isolates (n = 99, 87.6%). The maximum band difference between A1 and the remaining groups was 6, where most groups (9/16, 56.3%) differed by only 1 to 3 bands.
WGS analysis of S. Heidelberg isolates.
Of the 113 isolates that were analyzed by whole-genome sequencing (WGS), all had an average coverage of 134 times with a range of 56 to 346 times. hqSNV analysis was performed on all isolates, which included S. Heidelberg strain 12-4374 as the reference, as this genome was fully closed in a previous study (16). A maximum of 151 hqSNVs were identified between all isolates, with 95.9% of the reference included in the core genome.
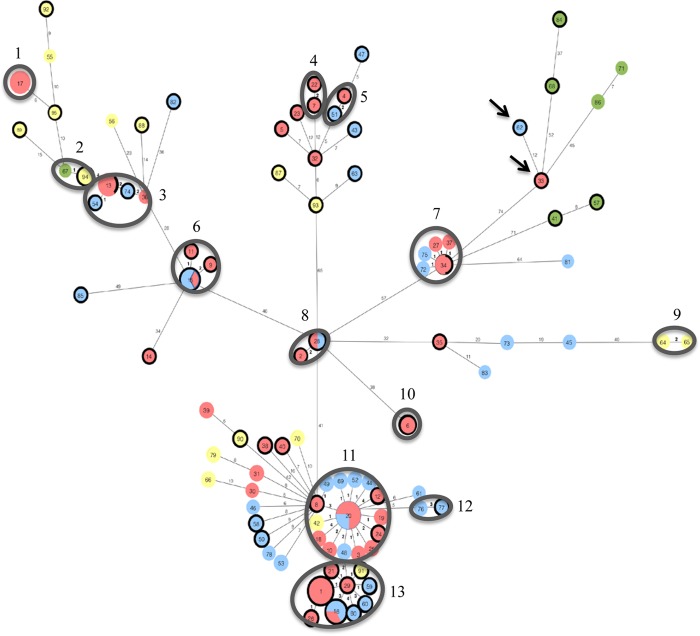
A minimum spanning tree identified 13 clusters containing 2 to 18 isolates (n = 68, 60.2%) having between 0 and 4 SNVs (Fig. 1). Two clusters (n = 32, 28.3%; clusters 11 and 13) consisted of cefoxitin-resistant and -susceptible human, abattoir chicken, and retail chicken isolates. Cluster 2 contained 2 abattoir chicken isolates clustering with a retail turkey isolate with 1 SNV difference. Among the isolates having 0 SNVs, there were 4 groups (found in clusters 6, 8, 11, and 13) with 2 to 4 isolates each of human and retail chicken origin. Two additional groups having 0 SNVs (found in clusters 3 and 7) with 3 and 2 isolates, respectively, were comprised of cefoxitin-resistant and -susceptible human isolates.
FIG 1.
Minimum spanning phylogenetic tree of the core genome of 113 sequenced Salmonella Heidelberg isolates with reference S. Heidelberg 12-4374, generated using Phyloviz. Colors for sources of isolates are as follows: red, human; yellow, abattoir chicken; blue, retail chicken; green, retail turkey. Isolates outlined in dark black are resistant to cefoxitin. Isolates in the same circle have 0 hqSNVs, and the size of each circle is proportionate to the number of isolates in the circle. Numbers on branches between 2 isolates represent the number of hqSNV differences. Clusters of isolates outlined in gray indicate those having 0 to 4 hqSNVs and are labeled 1 to 13. The 2 isolates with arrows indicate strains with a portion of a blaCMY-2 plasmid integrated into the chromosome.
All 113 isolates belonged to multilocus sequence type (MLST) ST15 (Table 1). Predicted antimicrobial resistance genes were identified using WGS data, and the majority of isolates (n = 112, 99.1%) had genes that correlated to the antimicrobial phenotypes determined by broth microdilution (Table 1). For the remaining isolate (12-5643), although initial antimicrobial susceptibility testing revealed an increased MIC for nalidixic acid, Resfinder (17) did not detect a corresponding resistance gene for this phenotype. Subsequent retesting using broth microdilution indicated that this isolate was susceptible to nalidixic acid.
CMY-2 plasmid analysis.
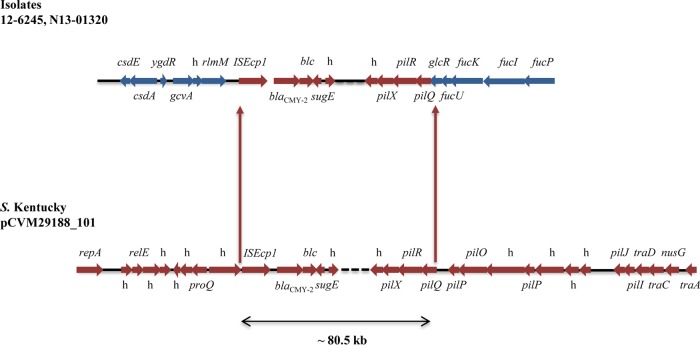
Using WGS data, 63/65 (96.9%) of the isolates containing blaCMY-2 plasmids belonged to replicon type IncI1. For the remaining 2 isolates (12-6245 and N13-01320), the blaCMY-2 gene resided on the largest contig (>747 kb) corresponding to the chromosome, suggesting that these plasmids might have integrated into the chromosome. An alignment of the previously characterized IncI1 blaCMY-2 plasmid, S. Kentucky pCVM29188_101 (GenBank accession number CP001121.1) (18), against the chromosomal contigs identified part of the blaCMY-2 plasmid (approximately 80.5 kb), which had inserted into the chromosome (Fig. 2). The plasmid inserted 386 bp downstream of the chromosomal rlmM gene and into the transcriptional repressor glcR, causing the terminal 91 bp to be deleted. No direct or inverted repeats flanking the insertion sites were found. Nucleotide sequence analysis of both 12-6245 and N13-01320 identified ISEcp1 approximately 200 bp downstream of the plasmid insertion site, followed by blaCMY-2 and a portion of the remaining plasmid. Several genes did not integrate with the plasmid into the chromosome, including those belonging to pil and tra operons, which are associated with transfer pili (Fig. 2). The plasmid replication initiation gene repA also did not insert into the chromosome, which may explain the lack of an IncI1 replicon type found in these isolates. The chromosomal insertion site for both isolates was confirmed using PCR. Using the hqSNV analysis, there were 12 SNVs found between isolates 12-6245 and N13-01320 containing the chromosomally located blaCMY-2 (Fig. 1).
FIG 2.
Schematic diagram of plasmid insertion into chromosome for strains 12-6245 and N13-01320 (top). The previously closed IncI1 blaCMY-2 Salmonella Kentucky pCVM29188_101 plasmid was used as a scaffold to identify the insertions (bottom). Blue arrows represent chromosomal open reading frames (ORFs); red arrows represent plasmid ORFs; h, hypothetical protein.
Using plasmid multilocus sequence types (pMLST) based on WGS data, the primary IncI1 plasmid sequence type of the 63 blaCMY-2 plasmids was ST12 (n = 56, 88.9%) (Table 1). Four (6.3%) plasmids were identified as belonging to ST2, ST25, ST26, and ST66, and three (4.8%) were untypeable, as only a portion (289/343 bp) of the ardA allele was present. The ST2 plasmid was isolated from a human and was resistant to only the beta-lactam antimicrobials. The plasmid with ST26 was isolated from a chicken and was resistant to beta-lactams, chloramphenicol, and sulfisoxazole. The remaining 2 plasmids with ST25 and ST66 were isolated from retail turkey and retail chicken, respectively, and were resistant to beta-lactams.
Sequence analysis of the blaCMY-2-containing plasmids revealed high homology (95 to 99%) to a previously described plasmid (pCVM29188_101) found in S. Kentucky (18). Nucleotide sequence alignments were performed to identify differences in genes compared to pCVM29188_101. All cefoxitin-resistant plasmids in this study belonged to 1 of 10 variant plasmids (Fig. 3). The majority of the 63 plasmids belonged to 1 of 3 variants, labeled as group A (n = 33, 52.4%), group B (n = 3, 4.8%), and group C (n = 20, 31.7%) (Table 1). The 7 remaining plasmids were isolated from individual strains. In comparison to S. Kentucky pCVM29188_101, all plasmids were missing the IS66 transposase. Some plasmids were missing additional genes encoding proteins that included transposases, recombinases, translational repressor protein RelE, DNA polymerase III subunit epsilon, colicin 1B immunity protein, quaternary ammonium resistance protein SugE, chromosome partitioning protein ParA, part of a membrane protein, and part of shufflon protein A.
FIG 3.
Nucleotide sequence alignments of the previously characterized S. Kentucky pCVM29188_101 plasmid against the 10 variant blaCMY-2-containing plasmids found in all S. Heidelberg isolates in this study, generated using GView Server. The reference plasmid is represented by the black boxes that denote open reading frames. For the remaining plasmids, solid colored boxes indicate that DNA is present. The gray line toward the bottom represents the GC content of the reference. Proteins listed above the alignment indicate those that are not present in certain plasmids, except for CMY-2, which was found in all plasmids. A representative sequence of a group A plasmid has been previously published (16).
Virulence data analysis.
In this study, 169 (8.4%) virulence genes were identified among the 113 strains. These genes were found to be involved in a variety of processes, including adhesion, type III secretion system (T3SS), regulation of genes, resistance to antimicrobial peptides, magnesium uptake, and regulation of stress factors. All sequenced isolates carried genes for curli, fimbriae, Salmonella pathogenicity island 1 and 2, PhoPQ, SodCl, and Mig-14. The majority of isolates (n = 97, 85.8%) did not contain stfA but contained the other genes in this operon (stfCDEFG), which is involved in fimbrial adherence of the organism. Most isolates (n = 110, 97.3%) also did not contain SeAg-B4893, which encodes a putative outer membrane protein that is involved in fimbrial adherence. Some isolates (n = 12, 10.6%) did not carry a greater percentage of virulence genes than the others. The genes missing in these isolates were all different, and they were not associated with any one characteristic, including source, antimicrobial resistance pattern, or type of human infection.
DISCUSSION
The use of antimicrobials in agriculture is of concern to public health, as overuse or misuse of these drugs can lead to resistance that may transfer to humans (19). A previous study has suggested that the use of ceftiofur in poultry in Québec has contributed to the elevated levels of cephalosporin resistance in S. Heidelberg from animals and humans (9). This current study utilized hqSNV analysis of S. Heidelberg from humans, abattoir poultry, and retail poultry to further understand the genetic relatedness between isolates from these sources in Québec in 2012.
As stated in previous studies (20), our findings support the fact that macrorestriction analysis using PFGE lacked the discriminatory power that was needed to identify potential relationships of this clonal serovar retrieved from human, abattoir poultry, and retail poultry sources, in contrast to hqSNV analysis using WGS.
Currently, there exists no defined range for the number of SNVs observed between 2 S. Heidelberg isolates that are genetically related. A study by Bekal et al. examined 3 epidemiologically defined outbreaks of S. Heidelberg in Québec and found a maximum number of 4 SNVs between isolates belonging to the same outbreak (20). Another study by Leekitcharoenphon et al. used hqSNV to characterize an outbreak of S. Heidelberg from the United States in 2011 and identified a maximum of 19 SNVs between their outbreak isolates (21). Using a maximum of 4 SNVs, our guideline to consider potential genetic linkages between isolates, we observed 13 clusters of 2 to 18 isolates each containing 0 to 4 SNVs. Four clusters contained up to 4 isolates each of human and retail chicken origin with 0 SNVs in the core genome. Identification of these clusters with 0 to 4 SNVs suggests that isolates within a cluster potentially originated from a common source.
Two groups of isolates with 0 SNVs (found in clusters 3 and 7) had isolates that were susceptible to all antimicrobials tested, and those that contained an IncI1 blaCMY-2 plasmid conferring resistance to beta-lactams. As extrachromosomal DNA was not included in the core genome used for the analysis, the occurrence of both antimicrobial-resistant and -susceptible isolates with identical core genomes may be indicative of plasmids being gained or lost, as was previously shown with the loss of an IncI1 blaCMY-2 plasmid found in Salmonella enterica serovar Bredeney within 49 days in an antimicrobial-free environment (22).
The majority of the turkey isolates (6/7, 85.7%) did not cluster with human or chicken isolates. The turkey isolates had different resistance patterns, including being resistant to beta-lactams, streptomycin, sulfisoxazole, and tetracycline (n = 2, 28.6%), resistant to only beta-lactams and either tetracycline or streptomycin (n = 2, 28.6%), or resistant to only ampicillin or tetracycline (n = 3, 42.9%). The fact that no human S. Heidelberg isolates clustered with turkey isolates suggested either that the isolates are less virulent in humans or that, possibly more likely, turkey is consumed less frequently than chicken, so humans are exposed to S. Heidelberg from turkey less frequently. In Canada, 85.6% of people reported consuming chicken in the previous 7 days compared to 11.8% who consumed turkey; and in Québec specifically, 86.9% of people reported eating chicken in the previous 7 days compared to 7.5% who consumed turkey (23).
All isolates were screened for virulence genes, which were defined as those that aid an organism to colonize a host, replicate, and cause inflammation, tissue damage, and consequently disease (24). There was no correlation seen between type of infection and virulence gene content.
In this study, two blaCMY-2 plasmids were found to have inserted into the chromosome. It has been suggested that ISEcp1 upstream of blaCMY-2 is involved in mediating such plasmid insertions (25). We identified 12 SNVs between the 2 isolates containing the plasmid integrations. It is unknown whether these integrations occurred separately at 2 different time points or whether they occurred once and the 2 isolates consequently evolved over time.
Comparison of our blaCMY-2 plasmids against previously characterized plasmids revealed homology to the S. Kentucky pCVM29188_101 plasmid and identified 10 plasmid subtypes. Three of the plasmids (groups A, B, and C) were found in multiple isolates of human, abattoir poultry, and retail poultry origins, and all plasmids in these groups belonged to ST12. The finding that the majority of blaCMY-2-containing plasmids were related suggests the dissemination of a similar resistant plasmid in variable genetic backgrounds of S. Heidelberg isolates.
In conclusion, we have compared cefoxitin-resistant and -susceptible S. Heidelberg isolates from humans, abattoir poultry, and retail poultry from a specific year and for a specific region in Canada using hqSNV analysis. Our findings suggest that although there is evidence to imply highly related isolates (0 to 4 SNVs) from human and poultry, suggesting a potential common source, it would seem that the majority of cefoxitin-resistant isolates studied occurred due to the dissemination of a plasmid between different S. Heidelberg genetic backgrounds. This suggests that the transmission of blaCMY-2 is due to the horizontal transfer of an IncI1 plasmid rather than clonal dissemination of a particular S. Heidelberg strain. Further WGS studies are under way to examine the temporal and spatial distribution of cefoxitin-resistant S. Heidelberg in Canada.
MATERIALS AND METHODS
Bacterial isolates.
Salmonella Heidelberg isolates were collected as part of CIPARS (8). A convenience sample of 113 isolates from humans (n = 51), abattoir poultry (n = 18), and retail poultry (n = 44) were studied, with the majority (n = 103) from Québec from 2012. Since no cefoxitin-resistant abattoir isolates from Québec in 2012 were available, 10 resistant isolates from chicken from Ontario and New Brunswick in 2011 and 2012 were selected for inclusion.
Initial characterization of isolates.
Antimicrobial susceptibility testing was performed using the broth microdilution Sensititre Automated Microbiology System (Trek Diagnostic Systems Ltd., Oakwood Village, OH, USA) and the CMV3AGNF panel in accordance with Clinical and Laboratory Standards Institute (CLSI) guidelines (26, 27). Multiplex PCR was performed on all cefoxitin-resistant strains (n = 65) to identify common extended-spectrum beta-lactamase (ESBL)/AmpC genes (blaSHV, blaTEM, blaCTX-M, blaOXA-1, and blaCMY-2), as described previously (28). The genetic relatedness was evaluated using pulsed-field gel electrophoresis (PFGE) according to the Centers for Disease Control and Prevention (CDC) PulseNet protocol (29). DNA fingerprints were analyzed using BioNumerics version 5.1 (Applied Maths, Saint Martens-Latem, Belgium).
Whole-genome sequencing.
DNA was extracted using the EpiCentre MasterPure Complete DNA and RNA purification kit (Illumina, Madison, WI, USA). Libraries were prepared using the Nextera XT DNA Sample Prep kit (Illumina, Madison, WI, USA). Sequencing was performed using paired-end reads to obtain an average coverage of >60 times for all isolates.
A comparative hqSNV analysis of the core genome was performed (A. Petkau, Core phylogenomics [https://github.com/apetkau/core-phylogenomics]). Horizontally acquired regions of the reference genome (S. Heidelberg 12-4374; GenBank accession CP012924.1) (16) containing prophages, genomic islands, and repeats were identified using PHAST (30), IslandViewer (31), and Nucmer v3.1 (32), respectively, and removed from the analysis. Sequencing reads were mapped to S. Heidelberg 12-4374 via SMALT v0.7.6 (33), using a k-mer size of 13 and a step size of 6. Variants were detected using FreeBayes v0.9.8 (34), with a minimum mapping quality of 30, a minimum base quality of 30, and a minimum alternate fraction of 0.75. Phylogenetic relatedness and a minimum spanning tree were constructed using Phyloviz (35).
Sequences were assembled into contigs with SPAdes v3.5.0 (36), and genomes were annotated via Prokka v1.1 (37). Assembled contigs were submitted to the Centre for Genomic Epidemiology's multilocus sequence type (MLST) (38), plasmid MLST (pMLST) (39), PlasmidFinder (39), and ResFinder (17) modules to determine whole-genome sequence types (ST), plasmid ST, existing plasmid replicon types, and resistance genes, respectively. Contiguator (40) was used to identify the contigs containing blaCMY-2 plasmids. Comparison of blaCMY-2 plasmids was performed against a previously characterized plasmid, S. Kentucky pCVM29188_101 (GenBank accession CP001121.1), using GView Server (41). Virulence genes were determined with an in-house workflow using SRST2 v0.1.4.5 (42), which maps Illumina raw reads against chromosomal and plasmid virulence genes found in the Virulence Factor Database for Salmonella (24). Currently, the database contains 2,017 genes associated with virulence in Salmonella (VFDB—current status [http://www.mgc.ac.cn/VFs/status.htm]).
Supplementary Material
ACKNOWLEDGMENTS
We thank Dave Spreitzer, Ken Fakharuddin, Stacie Langer, Danielle Daignault, Andrea Desruisseau, and Chad Gill for laboratory technical expertise and Laura Mataseje, Amrita Bharat, and Chand Mangat for assistance in sequence analysis. We also thank staff at the DNA core facility at the NML for next-generation sequencing and the Bioinformatics department at the NML for their expert technical support.
Footnotes
Supplemental material for this article may be found at https://doi.org/10.1128/AAC.01919-16.
REFERENCES
- 1.Majowicz SE, Musto J, Scallan E, Angulo FJ, Kirk M, O'Brien SJ, Jones TF, Fazil A, Hoekstra RM. 2010. The global burden of nontyphoidal Salmonella gastroenteritis. Clin Infect Dis 50:882–889. doi: 10.1086/650733. [DOI] [PubMed] [Google Scholar]
- 2.Public Health Agency of Canada. 2014. National Enteric Surveillance Program (NESP) annual report 2014. http://publications.gc.ca/collections/collection_2016/aspc-phac/HP37-15-2014-eng.pdf.
- 3.Vugia DJ, Samuel M, Farley MM, Marcus R, Shiferaw B, Shallow S, Smith K, Angulo FJ. 2004. Invasive Salmonella infections in the United States, FoodNet, 1996-1999: incidence, serotype distribution, and outcome. Clin Infect Dis 38:S149–S156. doi: 10.1086/381581. [DOI] [PubMed] [Google Scholar]
- 4.Public Health Agency of Canada. 2012. National Enteric Surveillance Program (NESP) annual report 2012. http://publications.gc.ca/collections/collection_2014/aspc-phac/HP37-15-2012-eng.pdf.
- 5.Currie A, MacDougall L, Aramini J, Gaulin C, Ahmed R, Isaacs S. 2005. Frozen chicken nuggets and strips and eggs are leading risk factors for Salmonella Heidelberg infections in Canada. Epidemiol Infect 133:809–816. doi: 10.1017/S0950268805004383. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Parmley EJ, Pintar K, Majowicz S, Avery B, Cook A, Jokinen C, Gannon V, Lapen DR, Topp E, Edge TA, Gilmour M, Pollari F, Reid-Smith R, Irwin R. 2013. A Canadian application of One Health: integration of Salmonella data from various Canadian surveillance programs (2005-2010). Foodborne Pathog Dis 10:747–756. doi: 10.1089/fpd.2012.1438. [DOI] [PubMed] [Google Scholar]
- 7.Seiffert SN, Hilty M, Perreten V, Endimiani A. 2013. Extended-spectrum cephalosporin-resistant Gram-negative organisms in livestock: an emerging problem for human health? Drug Resist Updat 16:22–45. doi: 10.1016/j.drup.2012.12.001. [DOI] [PubMed] [Google Scholar]
- 8.Public Health Agency Canada. 2013. Canadian Integrated Program for Antimicrobial Resistance Surveillance (CIPARS) annual report—design and methods. http://publications.gc.ca/collections/collection_2015/aspc-phac/HP2-4-2013-1-eng.pdf.
- 9.Dutil L, Irwin R, Finley R, Ng LK, Avery B, Boerlin P, Bourgault AM, Cole L, Daignault D, Desruisseau A, Demczuk W, Hoang L, Horsman GB, Ismail J, Jamieson F, Maki A, Pacagnella A, Pillai DR. 2010. Ceftiofur resistance in Salmonella enterica serovar Heidelberg from chicken meat and humans, Canada. Emerg Infect Dis 16:48–54. doi: 10.3201/eid1601.090729. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Federal Drug Administration. 2003. Merck's Mefoxin (cefoxitin for injection). http://www.fda.gov/ohrms/dockets/dailys/03/jun03/060403/03p-0227-cp00001-03-exhibit-b-vol1.pdf.
- 11.Hawkey PM. 1998. The origins and molecular basis of antibiotic resistance. BMJ 317:657–660. doi: 10.1136/bmj.317.7159.657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Jacoby GA, Munoz-Price LS. 2005. The new beta-lactamases. N Engl J Med 352:380–391. doi: 10.1056/NEJMra041359. [DOI] [PubMed] [Google Scholar]
- 13.Miriagou V, Tassios PT, Legakis NJ, Tzouvelekis LS. 2004. Expanded-spectrum cephalosporin resistance in non-typhoid Salmonella. Int J Antimicrob Agents 23:547–555. doi: 10.1016/j.ijantimicag.2004.03.006. [DOI] [PubMed] [Google Scholar]
- 14.Jacoby GA. 2009. AmpC beta-lactamases. Clin Microbiol Rev 22:161–182. doi: 10.1128/CMR.00036-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Mataseje LF, Xiao J, Kost S, Ng LK, Doré K, Mulvey MR. 2009. Characterization of Canadian cefoxitin-resistant non-typhoidal Salmonella isolates, 2005-06. J Antimicrob Chemother 64:723–730. doi: 10.1093/jac/dkp249. [DOI] [PubMed] [Google Scholar]
- 16.Labbé G, Edirmanasinghe R, Ziebell K, Nash JHE, Bekal S, Parmley EJ, Mulvey MR, Johnson RP. 2016. Complete genome and plasmid sequences of three Canadian isolates of Salmonella enterica subsp. enterica serovar Heidelberg from human and food sources. Genome Announc 4:e01526–15. doi: 10.1128/genomeA.01526-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Zankari E, Hasman H, Cosentino S, Vestergaard M, Rasmussen S, Lund O, Aarestrup FM, Larsen MV. 2012. Identification of acquired antimicrobial resistance genes. J Antimicrob Chemother 67:2640–2644. doi: 10.1093/jac/dks261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Fricke WF, McDermott PF, Mammel MK, Zhao S, Johnson TJ, Rasko DA, Fedorka-Cray PJ, Pedroso A, Whichard JM, LeClerc JE, White DG, Cebula TA, Ravel J. 2009. Antimicrobial resistance-conferring plasmids with similarity to virulence plasmids from avian pathogenic Escherichia coli strains in Salmonella enterica serovar Kentucky isolates from poultry. Appl Environ Microbiol 75:5963–5971. doi: 10.1128/AEM.00786-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Laxminarayan R, Duse A, Wattal C, Zaidi AKM, Wertheim HFL, Sumpradit N, Vlieghe E, Hara GL, Gould IM, Goossens H, Greko C, So AD, Bigdeli M, Tomson G, Woodhouse W, Ombaka E, Peralta AQ, Qamar FN, Mir F, Kariuki S, Bhutta ZA, Coates A, Bergstrom R, Wright GD, Brown ED, Cars O. 2013. Antibiotic resistance—the need for global solutions. Lancet Infect Dis 13:1057–1098. doi: 10.1016/S1473-3099(13)70318-9. [DOI] [PubMed] [Google Scholar]
- 20.Bekal S, Berry C, Reimer AR, Van Domselaar G, Beaudry G, Fournier E, Doualla-Bell F, Levac E, Gaulin C, Ramsay D, Huot C, Walker M, Sieffert C, Tremblay C. 2016. Usefulness of high-quality core genome single-nucleotide variant analysis for subtyping the highly clonal and the most prevalent Salmonella enterica serovar Heidelberg clone in the context of outbreak investigations. J Clin Microbiol 54:289–295. doi: 10.1128/JCM.02200-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Leekitcharoenphon P, Nielsen EM, Kaas RS, Lund O, Aarestrup FM. 2014. Evaluation of whole genome sequencing for outbreak detection of Salmonella enterica. PLoS One 9:e87991. doi: 10.1371/journal.pone.0087991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.de Toro M, García P, Rodríguez I, Rojo-Bezares B, Helmuth R, Sáenz Y, Rodicio MR, Guerra B, Torres C. 2013. Characterisation of plasmids implicated in the mobilisation of extended-spectrum and AmpC beta-lactamase genes in clinical Salmonella enterica isolates and temporal stability of the resistance genotype. Int J Antimicrob Agents 42:167–172. doi: 10.1016/j.ijantimicag.2013.04.016. [DOI] [PubMed] [Google Scholar]
- 23.Public Health Agency of Canada. 2015. Foodbook report. http://healthycanadians.gc.ca/publications/eating-nutrition/foodbook-2015/index-eng.php.
- 24.Chen L, Yang J, Yu J, Yao Z, Sun L, Shen Y, Jin Q. 2005. VFDB: a reference database for bacterial virulence factors. Nucleic Acids Res 33(Database issue):D325–D328. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Naseer U, Haldorsen B, Simonsen GS, Sundsfjord A. 2010. Sporadic occurrence of CMY-2-producing multidrug-resistant Escherichia coli of ST-complexes 38 and 448, and ST131 in Norway. Clin Microbiol Infect 16:171–178. doi: 10.1111/j.1469-0691.2009.02861.x. [DOI] [PubMed] [Google Scholar]
- 26.Clinical and Laboratory Standards Institute. 2012. Performance standards for antimicrobial susceptibility testing; twenty-second informational supplement. M100-S22. Clinical and Laboratory Standards Institute, Wayne, PA. [Google Scholar]
- 27.Clinical and Laboratory Standards Institute. 2012. Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically; approved standard—9th ed M07-A9. Clinical and Laboratory Standards Institute, Wayne, PA. [Google Scholar]
- 28.Mataseje LF, Boyd DA, Willey BM, Prayitno N, Kreiswirth N, Gelosia A, Poutanen SM, Low DE, Jenkins SG, Katz K, Mulvey MR. 2011. Plasmid comparison and molecular analysis of Klebsiella pneumoniae harbouring bla(KPC) from New York City and Toronto. J Antimicrob Chemother 66:1273–1277. doi: 10.1093/jac/dkr092. [DOI] [PubMed] [Google Scholar]
- 29.Centers for Disease Control and Prevention. 2013. Pulsed-field gel electrophoresis (PFGE). http://www.cdc.gov/pulsenet/pathogens/pfge.html.
- 30.Zhou Y, Liang Y, Lynch KH, Dennis JJ, Wishart DS. 2011. PHAST: a fast phage search tool. Nucleic Acids Res 39(Web Server issue):W347–W352. doi: 10.1093/nar/gkr485. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Dhillon BK, Laird MR, Shay JA, Winsor GL, Lo R, Nizam F, Pereira SK, Waglechner N, McArthur AG, Langille MGI, Brinkman FSL. 2015. IslandViewer 3: more flexible, interactive genomic island discovery, visualization and analysis. Nucleic Acids Res 43:W104–W108. doi: 10.1093/nar/gkv401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Delcher AL, Phillippy A, Carlton J, Salzberg SL. 2002. Fast algorithms for large-scale genome alignment and comparison. Nucleic Acids Res 30:2478–2483. doi: 10.1093/nar/30.11.2478. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Sanger Institute. SMALT. http://www.sanger.ac.uk/science/tools/smalt-0. [Google Scholar]
- 34.Garrison E, Marth G. 2012. Haplotype-based variant detection from short-read sequencing. 1207.3907. https://arxiv.org/pdf/1207.3907.pdf.
- 35.Francisco AP, Vaz C, Monteiro PT, Melo-Cristino J, Ramirez M, Carrico JA. 2012. PHYLOViZ: phylogenetic inference and data visualization for sequence based typing methods. BMC Bioinformatics 13:87. doi: 10.1186/1471-2105-13-87. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, Lesin VM, Nikolenko SI, Pham S, Prjibelski AD, Pyshkin AV, Sirotkin AV, Vyahhi N, Tesler G, Alekseyev MA, Pevzner PA. 2012. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19:455–477. doi: 10.1089/cmb.2012.0021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Seemann T. 2014. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30:2068–2069. doi: 10.1093/bioinformatics/btu153. [DOI] [PubMed] [Google Scholar]
- 38.Larsen MV, Cosentino S, Rasmussen S, Friis C, Hasman H, Marvig RL, Jelsbak L, Sicheritz-Pontén T, Ussery DW, Aarestrup FM, Lund O. 2012. Multilocus sequence typing of total-genome-sequenced bacteria. J Clin Microbiol 50:1355–1361. doi: 10.1128/JCM.06094-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Carattoli A, Zankari E, García-Fernández A, Larsen MV, Lund O, Villa L, Aarestrup FM, Hasman H. 2014. In silico detection and typing of plasmids using PlasmidFinder and plasmid multilocus sequence typing. Antimicrob Agents Chemother 58:3895–3903. doi: 10.1128/AAC.02412-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Galardini M, Biondi EG, Bazzicalupo M, Mengoni A. 2011. CONTIGuator: a bacterial genomes finishing tool for structural insights on draft genomes. Source Code Biol Med 6:11. doi: 10.1186/1751-0473-6-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Petkau A, Stuart-Edwards M, Stothard P, Van Domselaar G. 2010. Interactive microbial genome visualization with GView. Bioinformatics 26:3125–3126. doi: 10.1093/bioinformatics/btq588. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Inouye M, Dashnow H, Raven LA, Schultz MB, Pope BJ, Tomita T, Zobel J, Holt KE. 2014. SRST2: rapid genomic surveillance for public health and hospital microbiology labs. Genome Med 6:90. doi: 10.1186/s13073-014-0090-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.