Abstract
The microbial secretome, known as a pool of biomass (i.e., plant-based materials) degrading enzymes, can be utilized to discover industrial enzyme candidates for biofuel production. Proteomics approaches have been applied to discover novel enzyme candidates through comparing protein expression profiles with the whole secretome enzymatic activity under different growth conditions. However, the activity measurement of each enzyme candidate is needed for confident “active” enzyme assignments, and it remains to be elucidated. To address this challenge, we have developed an Activity-Correlated Quantitative Proteomics Platform (ACPP) that systematically correlate protein-level enzymatic activity patterns and protein elution profiles using a label-free quantitative proteomics approach. The ACPP optimized a high performance anion exchange separation for efficiently fractionating complex protein samples while preserving enzymatic activities. The detected enzymatic activity patterns in sequential fractions using microplate-based assays were cross-correlated with protein elution profiles using a customized pattern-matching algorithm with a correlation R score. The ACPP has been successfully applied to the identification of two types of “active” biomass-degrading enzymes (i.e., starch hydrolysis enzymes and cellulose hydrolysis enzymes) from Aspergillus niger secretome in a multiplexed fashion. By determining protein elution profiles of 156 proteins in A. niger secretome, we confidently identified the 1, 4-α-glucosidase as the major “active” starch hydrolysis enzyme (R=0.96) and the endoglucanase as the major “active” cellulose hydrolysis enzyme (R=0.97). The results demonstrated that the ACPP facilitated the discovery of bioactive enzymes from complex protein samples in a high-throughput, multiplexing, and untargeted fashion.
Graphical Abstract
Introduction
Bioethanol has been and is being sought as an alternative energy source due to its sustainability and renewability compared to traditional fuels. A key step in the development of bioethanol is the bioconversion of plant biomass (e.g., starch and celluloses) into fermentable sugars such as glucose. Current bioconversion technologies rely on extracellular glycosyl hydrolases (GHs) in the fungal secretome (e.g., fungal cellulases that break celluloses into glucose), which do not function effectively under the conditions required for enzymatic hydrolysis at an industrial scale. On the other hand, thousands of proteins have been predicted with GH activities based on their genome sequences, while many of them have the potentials to efficiently degrade biomass [1]. As of September 2013, around 160,000 GH family sequences have been predicted in the CAZy database. However, only 6% of the predicted CAZymes were biochemically characterized [2]. Therefore, there is a tremendous potential to screen novel and highly active biomass degrading enzymes in known industrial model fungal species (e.g., Aspergillus, Trichoderma, and Fusarium) or unknown fungal species.
Various approaches have been developed for discovering biomass-degrading enzymes in fungi. A general screening method is based on targeted gene overexpression in host cells (e.g., Escherichia coli or yeast). The overexpressed enzyme candidate was purified through several stages of liquid chromatography separations for further characterization and enzymatic activity analysis [3–5]. This approach can be readily incorporated into protein structure analysis and provide insights in protein activities. However, it is relatively low throughput because it only allows one protein to be analyzed per screening. On the other hand, fast protein liquid chromatography (FPLC) approaches have been applied to purify “active” enzymes from fungal secretome through substantial fractionations. For example, with nine different FPLC separations as well as activity measurements on all collected fractions, Olsson’s group successfully purified 5 cellulases from Penicillium brasilianum against different hydrolysis substrates (microcrystalline cellulose, carboxymethyl cellulose, galactomannan, and xylan) [6]. This method is very labor-intensive and low throughput, which often requires large amounts of starting materials.
Mass spectrometry (MS)-based proteomics is a core technique for high-throughput protein characterization in complex samples. Proteomics approaches have been applied to discover novel biomass degrading enzymes in the fungal secretome. For example, the Sze group has applied an iTRAQ based quantitative proteomics approach for analyzing the expression levels of putative GH enzymes under different growth conditions, providing key insights on designing the enzyme mixture for optimal biomass hydrolysis [7]. However, the enzyme candidates were identified through comparing their expression levels with the whole secretome enzymatic activity. Therefore, it may not provide sufficient resolution of enzyme mixtures because the expression level of each enzyme cannot be correlated to its own activity. Recently, the Activity-Based Protein Profiling (ABPP) approach has been adopted to profile the biomass degrading enzymes in the fungal secretome [8–10]. It utilized the activity-based probes (ABPs) to covalently label and affinity-purify the GHs from fungal secretome for proteomics screening. The ABPP approach has been applied to detect active biomass degrading enzymes in different microbial organisms. Currently, this method is limited by the probe availability. In addition, the optimization of GH-ABPs is required to increase the enzyme selectivity. Therefore, there is a critical need to develop a high-throughput untargeted proteomics approach to directly correlate the protein-level enzymatic activity.
In this study, we developed an Activity-Correlated quantitative Proteomics Platform (ACPP) that cross-correlated the protein-level enzymatic activity pattern and the protein elution profiles in a high throughput and untargeted fashion. The ACPP began with a high performance anion exchange separation for efficiently fractionating complex protein samples while preserving enzymatic activities. We then optimized the microplate-based enzymatic activity assays using small volume aliquots (e.g., 5–100 µl), which is capable of multiple enzyme activity characterizations. The fractions were also analyzed by means of LC-MS/MS. The elution profiles of all identified proteins were generated using peptide counting, a simple and robust label-free quantitative approach that also correlates with the enzymatic activities [11]. The detected enzymatic activity patterns in sequential fractions were cross-correlated with the proteome elution profiles of the detected proteins using a customized pattern-matching algorithm with a correlation R score range from +1 to −1. Because the enzyme concentrations are proportional to their activities, the theoretical correlation between enzyme elution profile and its activity pattern should have a “perfect” R score of 1. This platform has been successfully applied to the identification of “active” biomass degrading enzymes from the Aspergillus niger secretome. We demonstrate that the ACPP is a powerful untargeted approach for hunting unknown bioactive enzymes through the cross-correlation of activity pattern and protein elution pattern in a high-throughput fashion.
Materials and Methods
Cultivation of A. niger and secretome extraction
Chemicals were purchased from Sigma–Aldrich (Milwaukee, WI) unless otherwise noted. A. niger (ATCC11414) was reserved on a PDA plate, which is widely used for fungal growth. The cultivation was performed as described previously [12]. After 24 h of growth, the supernatant was collected by filtering the culture through two layers of sterile miracloth, and the filtered culture medium was then centrifuged at 15,000g and 4 °C for 10 min to remove the remaining cell debris. The collected secretome was further filtered through a 0.2 µm filter before being concentrated to 20 ml in a stirred cell at 4 °C (Millipore, Billerica, MA) with a 3 KDa ultrafiltration membrane. Two additional buffer exchange steps with Milli-Q water (i.e., from 250 ml to 20 ml) were applied in the same stirred cell. The remaining secretomes were further concentrated down to 3–5 ml using a centrifugal ultrafiltration with 3 kDa cut-off (Millipore, Billerica, MA). The prepared secretome was stored at −20 °C until fractionation.
Anion exchange LC separation and fractionation
The LC separation and fractionation were performed at 4 °C using an AKTA pure protein purification system (GE, Pittsburgh, PA) with a high performance mono-Q column (4.6×100 mm, GE, Pittsburgh, PA). All the samples were exchanged to buffer A (20 mM Tris, pH=8.0) before separation. Gradient elution (the flow rate at 0.5 ml/min) was performed from 0–100% buffer B (20 mM Tris, 1 M NaCl, pH=8.0) in minutes. Fractions (0.83 ml per fraction for the spiked-in sample, 1 ml per fraction for the A. niger secretome, and 1.66 ml per fraction for A. niger cellulose and cellubiose activity assays) were collected every 2 minutes for further activity assays and proteomics studies. Three aliquots were taken from each fraction: (1) one for the biomass hydrolysis activity assay (i.e., 5–100 µl), (2) one for the cellulose hydrolysis activity assay (i.e., 20–100 µl), and (3) one for digestion by trypsin and LC/MS/MS based bottom-up proteomic characterization (i.e., 100–200 µl). The remaining fractions were stored at −20 °C.
Microplate-based enzymatic activity assays
A two-step 96-well microplate-based enzymatic assay approach (e.g., substrate hydrolysis and glucose detection) was developed based on the traditional assay method [13, 14] for higher throughput and low sample consumption. First, a small aliquot of fractions (5–100 µl) was mixed with 5% (w/v) cellulose, cellubiose, or starch substrates at the ratio of 1:4. The mixture was incubated for 2 hours at 37°C and 900 rpm for substrate hydrolysis. The hydrolysis reaction was quenched on ice for 10 minutes. The supernatant was collected after centrifugation at 13kg, 4°C, 30 minutes. The released glucose concentrations in reaction supernatants were measured using the glucose assay reagent (Sigma, Milwaukee, WI). Briefly, the supernatant of reaction mixtures (i.e., 20 µl) and glucose working reagent were mixed at the ratio of 1:5 in a 96-well plate. After incubation at 25°C for 30 minutes, the absorbance of the reaction mixture was measured at 340 nm. The activities of fractions were represented by the detected glucose concentration.
LC-MS/MS analysis
A set of fraction aliquots (i.e., 100–200 µl) were subjected to in-solution trypsin digestion following previously published protocol [15]. Peptide samples were desalted, dried in SpeedVac, and resuspended in water. 1/5 of the digested sample was injected on a custom-packed C18 RPLC column (75 um i.d., 150 mm length, 2 µm C18 resin) using a Waters nano-Acquity UPLC system. A 100-min gradient from 8% buffer A (0.1% formic acid in water) to 35% buffer B (0.1% formic acid in acetonitrile) was applied for peptide separation. The eluent was analyzed online using a LTQ Orbitrap Velos Pro mass spectrometer (Thermo Fisher Scientific) with a custom nano-ESI interface. The heated capillary temperature was set to 300°C with a spray voltage of 2.4 kV. Full MS spectra were acquired at a resolution of 30K (m/z range between 350 and 2000). Low resolution collisional induced dissociation (CID) MS/MS with a normalized collision energy setting of 35% was performed in the data-dependent mode using the ten most abundant parent ions.
Peptide and protein identification
Peptides were identified using MSGF+ (Kim S, Pevzner PA. MS-GF+ makes progress towards a universal database search tool for proteomics. Nat Commun. 2014;5:5277. doi: 10.1038/ncomms6277. pmid:25358478) to search the mass spectra from LC-MS/MS analysis against the annotated A. niger database and its decoy database [16]. Peptide identifications were filtered using an MS-GF cut-off value of 1E−10 (i.e., the calculated FDR<1% at the unique peptide level. Spectral count was used for label free quantitation, and the normalized quantitation of same protein in different fractions was plotted against the fraction number to obtain the protein elution profile.
Cross-Correlation of protein profiles and activity pattern
Using the home-written Pearson correlation based bioinformatics tool with MATLAB (available upon request), the protein LC elution pattern for each protein was correlated with its measured activity pattern. The correlation efficiency was evaluated by the R score, from −1 to 1. In addition, the protein elution profiles were filtered based on its peak quality (i.e., the total identified peptide number for the most abundant elution fraction was larger than or equal to 10 for distinguish a good elution profile from random noises). For each set of data, the match significance was estimated based on the correlations between every two random elution profiles. The putatively identified proteins were manually interpreted to confidently identify the active enzyme from complicated protein mixture.
Results and Discussion
ACPP development and proof-of-concept
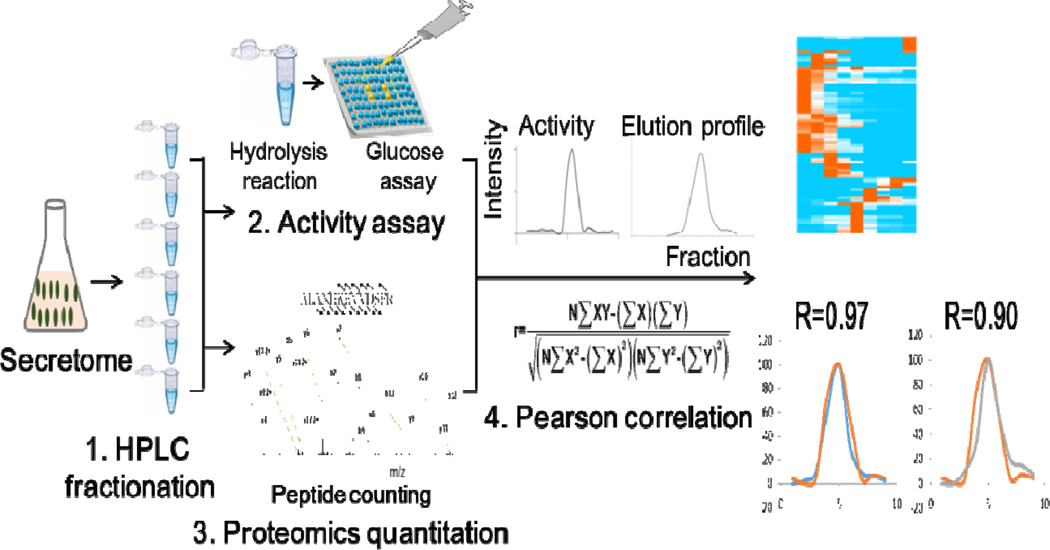
The overall ACPP strategy is demonstrated in Figure 1. In general, the ACPP strategy is composed of four steps: (1) HPLC fractionation for efficiently fractionating complex protein samples while preserving enzymatic activities; (2) enzymatic activity pattern determination using microplate based bioactivity assays, (3) protein elution profile generation using quantitative proteome analysis, and (4) correlations between enzymatic activity patterns and proteome elution profiles of the detected proteins. As a proof-of-principle, we examined the commercially available standard starch hydrolysis enzyme 1, 4-α-glucosidase (AAG) from A. niger. We evaluated the performance and reproducibility for each steps.
Figure 1.
Schematic representation of the ACPP workflow.
HPLC fractionation
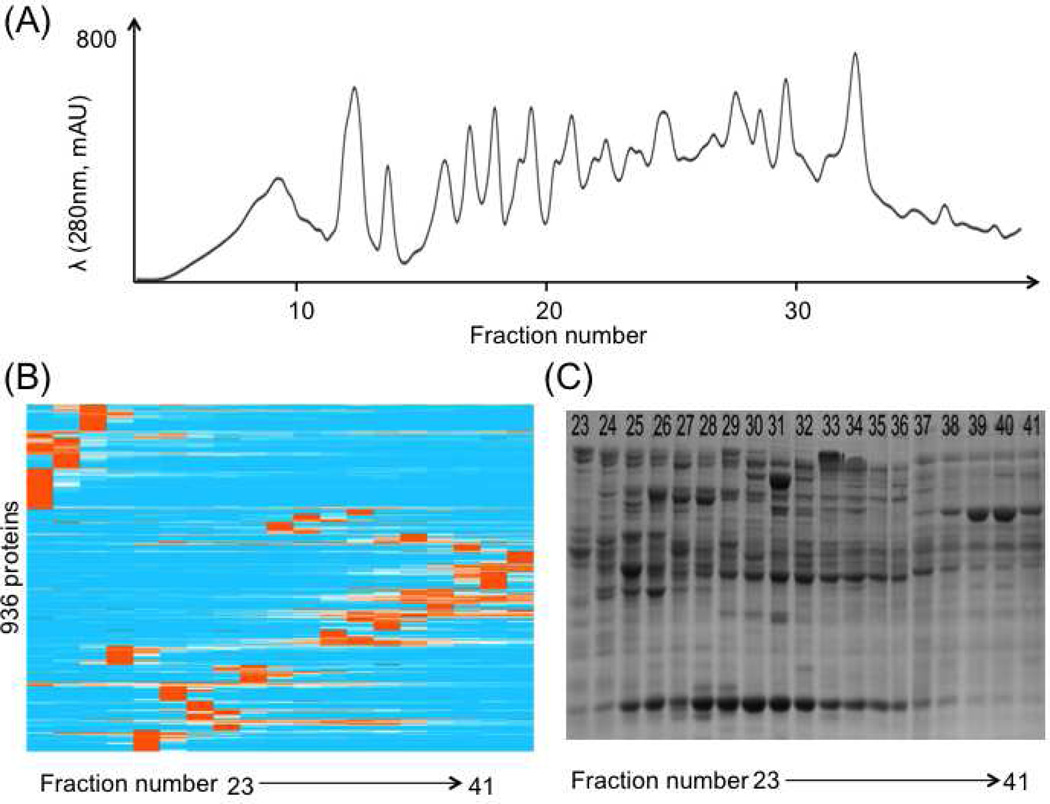
The ACPP depends on a high-resolution and reproducible LC separation that can preserve enzyme function and activities. Several formats of LC separation have been applied in enzyme purification, including size exclusive chromatography and ion exchange chromatography [17]. Here we applied a high performance anion exchange LC separation using a GE Mono Q™ column for protein fractionation. The UV traces of four replicated standard AAG analyses nicely overlapped (Supplementary figure 1), demonstrating the good reproducibility of the selected column. Three baseline-separated peaks were detected, and the proteomics results confirmed that all these peaks are protein isoforms of AAG. We also evaluated the separation of complex protein samples such as the AAG spiked E. coli cell lysate (Figure 2). The separation performance of the spiked-in E coli sample was evaluated using SDS-PAGE and proteomics results. The results confirmed that the anion exchange chromatography using the GE Mono Q™ column can efficiently separate and fractionate complex protein samples in ACPP. The following bioactivity assays on the collected fractions further confirmed that the selected anion exchange chromatography can preserve enzyme function and activities.
Figure 2.
Protein elution profiling of ACPP on the E coli. cell lysate with the spiked-in standard AAG. (A) the UV chromatogram from the mono Q based LC separation; (B) Hierarchical clustering of protein elution profiles on the basis of their similarities; (C) SDS PAGE gel image of collected LC separation fractions.
Bioactivity assay
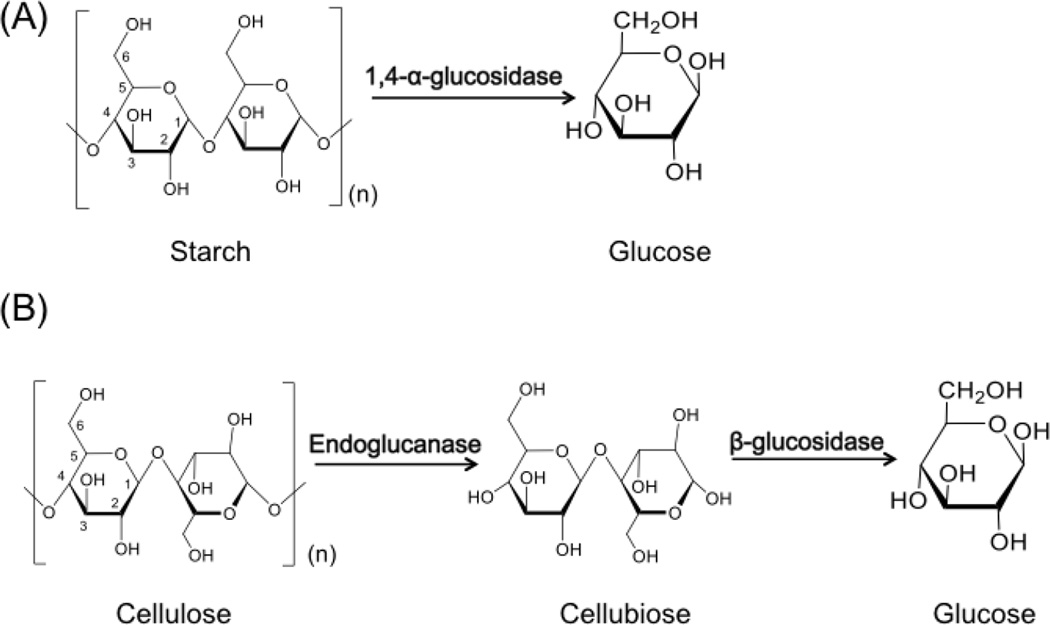
Two biomass substrates were selected in this study: starch and cellulose. Starch is a linear polymer of glucose mainly linked with (1→4) bonds, and cellulose is a polymer with repeated beta glucose units (Figure 3) [18]. The starch hydrolysis enzymes such as AAG convert starch into glucose, while cellulase catalyzes the decomposition of cellulose into glucose. Here we developed a two-step biomass degrading enzyme activity assay: (1) enzymatic hydrolysis reaction; (2) glucose assay. An aliquot of the fractions (i.e., 5–100 µl) were incubated with the substrate solution under optimized pH and temperature conditions to ensure the completion of hydrolysis. The supernatants of the reaction solution (~20 µl) in each fraction were transferred into a 96-well micro-plate for high throughput micro-scale glucose assay. A commercial glucose assay kit (Sigma) was applied, and a micro-scale assay protocol was developed based on manufactory’s instructions. For each analysis, a standard glucose calibration curve was plotted for glucose concentration calculation and enzymatic activity calculation. We evaluated the quantification performance, sensitivity, linear range, and reproducibility of the proposed activity assay using the serial dilution of standard starch hydrolysis enzyme AAG proteins (Supplementary figure 2A). We further applied our developed activity assay on the LC separation of the E. coli cell lysate with the spiked-in standard AAG (i.e., 150 µg AAG in 1.85 mg cell lysate). The assay was performed using 5µl of each fraction (i.e., 1/200 of each fraction), and a reproducible activity peak was observed for the major AAG isoform (Supplementary figure 2B). The results also supported our hypothesis that the anion exchange chromatography can preserve the biomass degrading enzyme activities. Overall, we confirmed that our high-throughput micro-plate based assay is sensitive and reproducible, which is suitable for generating quantitative enzymatic activity patterns in ACPP.
Figure 3.
Two biomass hydrolysis reactions are catalyzed by different enzymes. (A) 1, 4-α-glucosidase (AAG) is a typical enzyme that can directly convert starch into glucose; (B) Cellulose is converted into glucose in a two-step reaction, and the endo-cellulase (i.e., endoglucanase) activity can be measured in the presence of β-glucosidase.
Proteome elution profiling
An LC-MS/MS analysis was applied to each fraction for protein and peptide identifications. To generate the protein elution profiles, we used a peptide counting based label-free quantitation approach. Peptide counting has been applied to characterize commercial cellulase cocktails, demonstrating good correlations with enzyme activities [11]. For the E. coli lysate with the spiked-in AAG, we performed the LC-MS/MS analysis for fractions 23 to 41 (i.e., the fractions with starch activity, supplementary material). We acquired quantitative protein elution profiles for 934 proteins from these fractions, and the results are presented as a heat map after hierarchical clustering (Figure 3). We also compared the protein elution profiles with the SDS-PAGE gel separation. The AAG were barely observed on the gel. Since the mono-Q anion exchange LC provides a complimentary protein-level separation for peptide level LC-MS/MS analysis, the ACPP is capable of detecting and profiling low abundant active enzymes.
Correlations between activity patterns and protein elution profiles
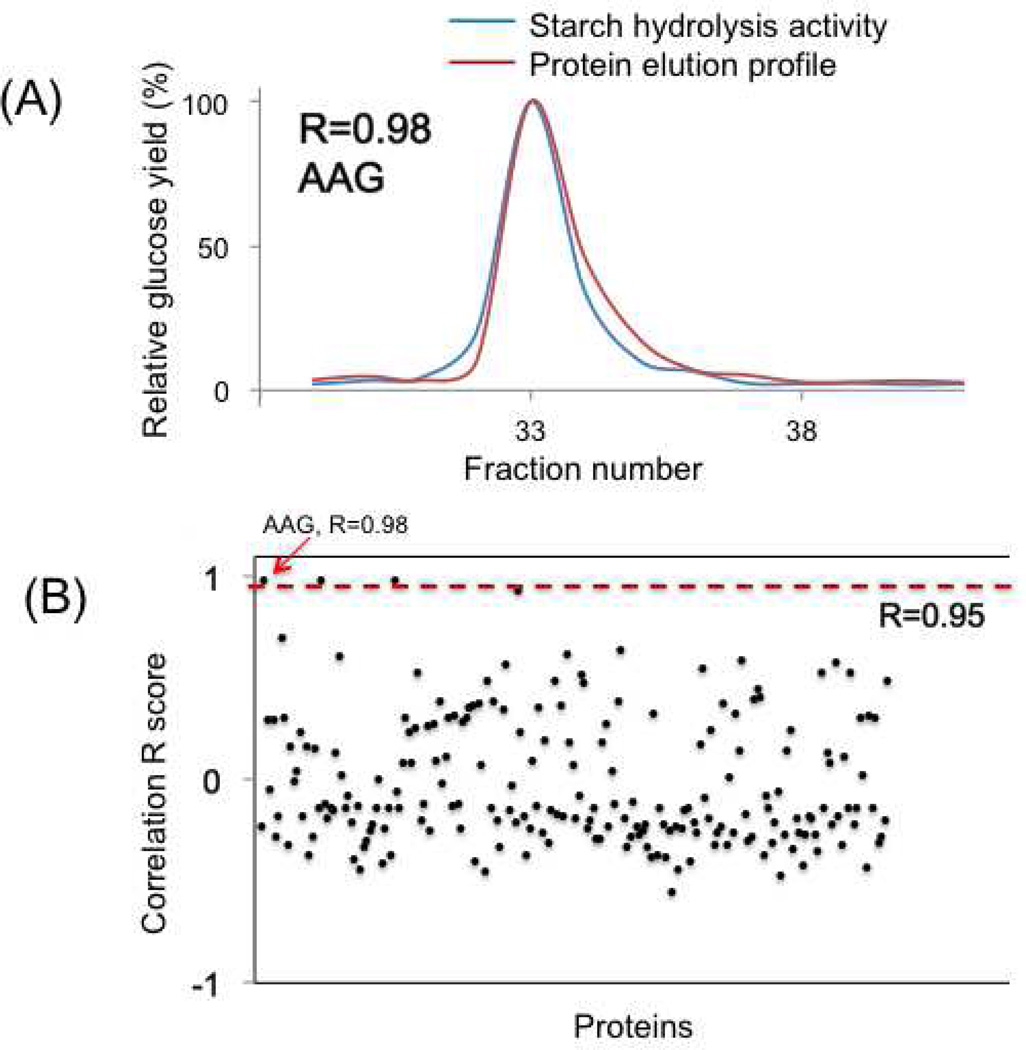
In ACPP, bioactive enzymes were detected through the correlations between the activity patterns and protein elution profiles. The detected enzymatic activity patterns in sequential fractions were cross-correlated with the label-free quantitative proteome results using a customized pattern-matching algorithm with a correlation R score range from +1 to −1. Because the enzyme concentrations are proportional to their activities, the theoretical correlation between enzyme elution profile and its activity pattern should have a “perfect” R score of 1. The correlations between co-eluted proteins can also give relatively high R scores. However, because the proteins have different pI values, with high performance LC separation, the chance of randomly co-eluted proteins with “perfect” overlapped profiles (i.e., R=1) should be relatively low. Therefore, R scores can provide good evidence for finding unknown “active” proteins. As a proof-of-principle, we plotted the R scores between the activity patterns and protein elution profiles for the AAG spiked-in experiment (Figure 4B).
Figure 4.
The ACPP on starch hydrolysis enzyme characterization from the E coli. cell lysate with the spiked-in standard AAG. (A) The overlaps between the measured starch hydrolysis activities and the AAG protein elution profiles; (B) Correlations between detected activity pattern and the filtered elution profiles from all the identified proteins (total peptide count cutoff of 10).
We evaluated the match significance of the R scores through plotting the histogram counts of the correlation scores between any two of the detected protein elution profiles and the correlation scores between the activity patterns and protein elution profiles (Supplementary figure 3). Our hypothesis is that majority of the correlations between eluted proteins are random, which can be applied for the false discovery estimation. We calculated the Z-scores for each correlation, and the Z-score of 3 was chosen as our first cutoff to reduce random correlation. The correlation R score cutoff value was 0.4 with Z-scores larger than 3. Still we noticed many randomly correlated proteins after Z-score cutoff. We manually evaluated all of the hits with Z-scores larger than 3, and selected R=0.95 to ensure good correlations and low false discovery rates. For E coli spiked-in sample, we noticed that only 3 proteins have R scores larger than 0.95 with the total peptide count cutoff of 10 (i.e., distinguish from noise peaks), while only 4 proteins have R score larger than 0.7. The elution profile of AAG protein correlates the best with the detected activities (R=0.98, Figure 4A).
Overall, the correlation R scores can be utilized for confidently assign the active protein candidates through quantifying the correlation between activity patterns and protein elution profiles. However, the basis hypothesis of the Z-score approach is that the correlation scores are normally distributed. We noticed that the actual distribution was slightly off from the normal distribution curve, which may be related to low distinguish power with relatively low fraction numbers (Supplementary figure 3). Therefore, our Z score estimation should be improved by optimizing the fractionation schemes, including two separated distributions for partially co-eluted proteins and non-co-eluted proteins, as well as incorporating manual validations.
ACPP for detecting biomass degrading enzymes from A. niger secretome
We then applied the ACPP to lab cultured fungal secretome to identify the biomass hydrolysis enzymes from A. niger. Two biomass substrates were selected in this study: starch and cellulose. We demonstrated that ACPP can be applied to detect both starch hydrolysis enzyme and cellulose hydrolysis enzyme in a multiplexed fashion with the same set of proteomics data. We measured activities for both enzymes (Figure 5A), and our results clearly demonstrated that these two enzymes have different elution patterns.
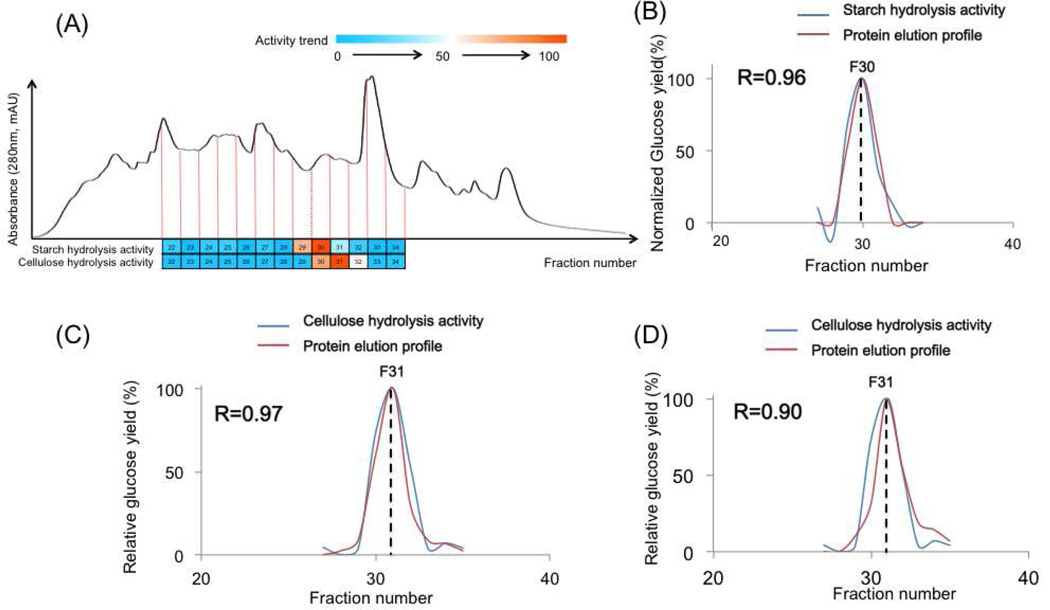
Figure 5.
The ACPP on detecting biomass degrading enzymes from A. niger secretome. (A) the UV chromatogram from the LC separation; (B) the overlaps between the measured starch hydrolysis activities and the best matched protein elution profile 1, 4-α-glucosidase (AAG); (C) the overlaps between the measured cellulose hydrolysis activities and the best matched protein elution profile (endoglucanase, GH5); (D) the overlaps between the measured starch hydrolysis activities and the protein elution profile of β-glucosidase.
For the starch hydrolysis enzyme, we observed a single Gaussian distributed peak from fraction 27 to 34 with a peak center at fraction 30 (Figure 5B). The corresponding fractions were applied for protein identifications and quantifications. In total, 156 proteins were identified from these 8 fractions. The ACPP correlated all protein elution profiles against the measured starch hydrolysis enzyme activity pattern. The calculated R score cutoff 0.95 was selected to ensure the match significance and only two proteins have R score higher than 0.95. The best-correlated protein, with R score equal to 0.96 (Figure 5B), was confirmed as 1, 4-α-glucosidase (AAG) protein (GH 15, EC 3.2.1.3), which is a classical starch hydrolysis enzyme [19]. We further manually went through all the detected GH proteins predicted by CAZY database, and confirmed a total of 25 proteins as GH proteins. Four GH proteins with R scores higher than 0.80 were listed in Table 1, including GH15 (i.e., AAG), GH36, GH32, and GH18 proteins. The predicted substrate for the detected GH36 protein is glycolipids, and the GH32 protein was reported as an inulinase. The GH18 protein was a hypothetical protein with unknown substrates. We did BLASTp of its sequence against the NCBI database [20], and it didn’t show any sequences homologous with known starch degrading enzymes. In addition, the R score of this GH18 protein is only 0.87, which indicate that it does not have good correlation with our detected enzyme activity pattern. Our results indicated that our method is powerful enough to identify the bioactive enzymes from real biological samples.
Table 1.
ACPP correlated putative starch hydrolysis proteins in A. niger secretome (R>0.8)*
| Total rank | 1st | 2nd | 4th | 5th |
| GHs’ rank | 1st | 2nd | 3rd | 4th |
| R score | 0.96 | 0.95 | 0.91 | 0.87 |
| Protein | gi|350633017 | gi|350639761 | gi|350631139 | gi|350640229 |
| GH family | GH15 | GH36 | GH32 | GH18 |
| Function | 1,4-α- glucosidase |
α- galactosidase |
Inulinase | Hypothetical protein |
| Substrates | Starch | Glycolipids/ glycoproteins |
Inulin | Unknown |
The calculated R score cutoff is 0.95
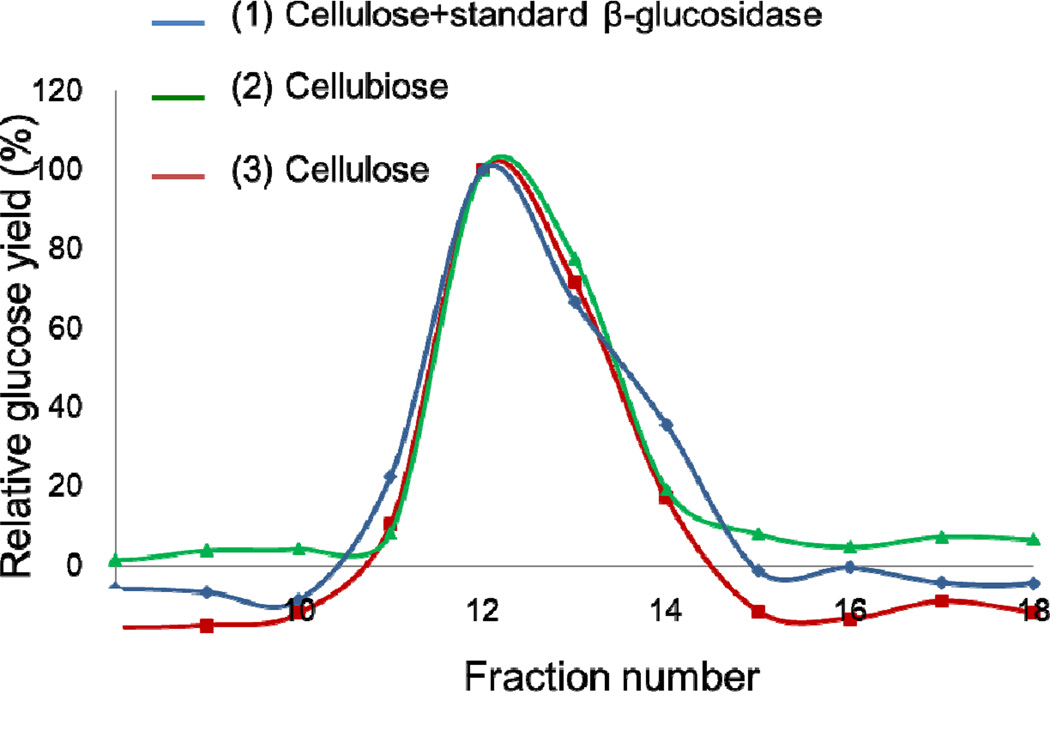
We further did cellulase screening on the A. niger secretome. The activity pattern was shown as a nice Gaussian-distributed peak centered at fraction 31 (Figure 5C). The calculated R score cutoff 0.95 was selected to ensure the match significance and only one protein has R score higher than 0.95, which is endoglucanase (GH5). It is known that two kinds of enzymes are necessary—endoglucanase and β-glucosidase to convert cellulose to glucose (Figure 3B), and endoglucanase by itself can only convert cellulose to cellubiose [21]. Our results indicated either the two enzymes were co-eluted under our LC separation conditions or it represented a novel cellulase which can directly hydrolyze cellulose to glucose. We manually evaluated our results, and the protein with second highest R score (R=0.9) is β-glucosidase, which can convert cellubiose to glucose. Because of the presence of co-eluted β-glucosidase, our glucose assay was successfully matched against the endoglucanase activity. We further confirmed that β-glucosidase cannot convert cellulose into glucose using standard proteins (Supplementary figure 4). Therefore, endoglucanase should present the best correlation under ACPP. To confirm that these two proteins are co-eluted, we performed three sets of activity assays on three separated aliquots of HPLC fractions (i.e., 5–100 µl) incubated with different substrate solutions (Figure 6): (1) cellulose and standard β-glucosidase; (2) cellubiose; (3) cellulose. The first set of assay provided the endoglucanase elution profile, the second set provided β -glucosidase elution profile, and the third set evaluates if endoglucanase and β-glucosidase are co-eluted (i.e., if they are not co-eluted, no glucose signal should be detected). Our results indicated that our method is powerful to identify the bioactive enzymes from real biological samples in an untargeted and multiplexed fashion. Moreover, it holds the potential for the analysis of co-eluted protein complexes.
Figure 6.
The cellulose activity assays on A. niger secretome. Three assays were included: Sigmacell cellulose+HPLC fractions+ standard β-glucosidase, cellubiose+HPLC fractions, and Sigmacell cellulose + HPLC fractions. The blue line represents the endoglucanase elution profile, the green line represents the β -glucosidase elution profile, and the red line evaluates if endoglucanase and β-glucosidase are co-eluted.
Conclusions
In summary, the ACPP is a powerful approach which can cross-correlate the activity patterns and protein elution profiling, in a high throughput and untargeted fashion to identify bioactive enzymes from hundreds of proteins. Due to the micro-scale activity assay and the high-sensitive proteomics, the ACP is multiplexed to identify more than one active enzymes from complicated protein mixture. Furthermore, the ACPP can be readily adapted to other applications, such as disease enzyme screening and drug target discovery. Meanwhile, we have noticed several limitations for current ACPP technique. For example, although Mono-Q based anion exchange separation provides high resolution for secretome study, it still suffers with limited resolution for more complex samples such as human cell lysate. In addition, positively charged proteins cannot bind on anion exchange columns. Therefore, a future direction to optimize current method is to add an additional dimension to the separation (i.e., size exclusion) to convert broad or overlapping peaks to individual peaks. The addition dimension to the separation can also facilitate the detection of low abundance enzymes. In addition, the protein elution profiles were generated using peptide counting based label free quantitation, which is a simple and robust approach. However, the linear dynamic range of peptide counting approach can be limited due to the saturation effects. To further improve the accuracy and sensitivity of protein elution profile generation, we will also evaluate other quantitation approaches such as intensity-based approach as well as targeted proteomics. We will also improve current bioinformatics software to accommodate sophisticated correlation situations.
Supplementary Material
Acknowledgments
We thank Dr. Da Meng for developing bioinformatics tools for ACPP. We thank Dr. Robert H. Cichewicz for helping with fungal culture setup, and Dr. Shuang Deng for providing A. niger fungal strain. Preliminary results were partially supported by the NSF national high Magnetic field laboratory user proposal with the helps from Drs. Nate Kaiser, Lissa C. Anderson, and Chris Hendrickson. We appreciate Dr. Eliza A. Ruben and OU COBRE in structure biology for the assistance with LC separation. We also thank Dr. Kenneth Smith for reviewing the manuscript. This work was partly supported by grants from NIAID CSGADP Pilot project (NIH 5U01AI101990-04, BRI No. FY15109843), NIH NIGMS R01 GM118470, and OCAST HR16-125.
References
- 1.Cantarel BL, Coutinho PM, Rancurel C, Bernard T, Lombard V, Henrissat B. The Carbohydrate-Active EnZymes database (CAZy): an expert resource for Glycogenomics. Nucleic Acids Res. 2009;37:D233–D238. doi: 10.1093/nar/gkn663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Lombard V, Ramulu HG, Drula E, Coutinho PM, Henrissat B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 2014;42:D490–D495. doi: 10.1093/nar/gkt1178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Linger JG, Adney WS, Darzins A. Heterologous Expression and Extracellular Secretion of Cellulolytic Enzymes by Zymomonas mobilis. Appl. Environ. Microbiol. 2010;76:6360–6369. doi: 10.1128/AEM.00230-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Wang J, Gao G, Li Y, Yang L, Liang Y, Jin H, Han W, Feng Y, Zhang Z. Cloning, Expression, and Characterization of a Thermophilic Endoglucanase, AcCel12B from Acidothermus cellulolyticus 11B. Int. J. Mol. Sci. 2015;16:25080–25095. doi: 10.3390/ijms161025080. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Wu WH, Hung WC, Lo KY, Chen YH, Wan HP, Cheng KC. Bioethanol production from taro waste using thermo-tolerant yeast Kluyveromyces marxianus K21. Bioresour. Technol. 2016;201:27–32. doi: 10.1016/j.biortech.2015.11.015. [DOI] [PubMed] [Google Scholar]
- 6.Jorgensen H, Eriksson T, Borjesson J, Tjerneld F, Olsson L. Purification and characterization of five cellulases and one xylanase from Penicillium brasilianum IBT 20888. Enzyme Microb. Technol. 2003;32:851–861. [Google Scholar]
- 7.Adav SS, Li AA, Manavalan A, Punt P, Sze SK. Quantitative iTRAQ secretome analysis of Aspergillus niger reveals novel hydrolytic enzymes. J. Proteome Res. 2010;9:3932–3940. doi: 10.1021/pr100148j. [DOI] [PubMed] [Google Scholar]
- 8.Chauvigne-Hines LM, Anderson LN, Weaver HM, Brown JN, Koech PK, Nicora CD, Hofstad BA, Smith RD, Wilkins MJ, Callister SJ, Wright AT. Suite of Activity-Based Probes for Cellulose-Degrading Enzymes. J. Am. Chem. Soc. 2012;134:20521–20532. doi: 10.1021/ja309790w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Li N, Overkleeft HS, Florea BI. Activity-based protein profiling: an enabling technology in chemical biology research. Curr. Opin. Chem. Biol. 2012;16:227–233. doi: 10.1016/j.cbpa.2012.01.008. [DOI] [PubMed] [Google Scholar]
- 10.Anderson LN, Culley DE, Hofstad BA, Chauvigne-Hines LM, Zink EM, Purvine SO, Smith RD, Callister SJ, Magnuson JM, Wright AT. Activity-based protein profiling of secreted cellulolytic enzyme activity dynamics in Trichoderma reesei QM6a, NG14, and RUT-C30. Mol. Biosyst. 2013;9:2992–3000. doi: 10.1039/c3mb70333a. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Chundawat SP, Lipton MS, Purvine SO, Uppugundla N, Gao D, Balan V, Dale BE. Proteomics-based compositional analysis of complex cellulase-hemicellulase mixtures. J. Proteome Res. 2011;10:4365–4372. doi: 10.1021/pr101234z. [DOI] [PubMed] [Google Scholar]
- 12.Qu Y, Feng J, Deng S, Cao L, Zhang Q, Zhao R, Zhang Z, Jiang Y, Zink EM, Baker SE, Lipton MS, Pasa-Tolic L, Hu JZ, Wu S. Structural analysis of N- and O-glycans using ZIC-HILIC/dialysis coupled to NMR detection. Fungal Genet. Biol. 2014;72:207–215. doi: 10.1016/j.fgb.2014.08.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Ghose TK. Measurement of Cellulase Activities. Pure Appl. Chem. 1987;59:257–268. [Google Scholar]
- 14.Ooshima H, Burns DS, Converse AO. Adsorption of Cellulase from Trichoderma-Reesei on Cellulose and Lignacious Residue in Wood Pretreated by Dilute Sulfuric-Acid with Explosive Decompression. Biotechnol. Bioeng. 1990;36:446–452. doi: 10.1002/bit.260360503. [DOI] [PubMed] [Google Scholar]
- 15.Zhang ZR, Wu S, Stenoien DL, Pasa-Tolic L. High-Throughput Proteomics. Annu. Rev. Anal. Chem. 2014;7:427–454. doi: 10.1146/annurev-anchem-071213-020216. [DOI] [PubMed] [Google Scholar]
- 16.Kim S, Pevzner PA. MS-GF plus makes progress towards a universal database search tool for proteomics. Nat. Commun. 2014;5 doi: 10.1038/ncomms6277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Jansen RS, Addie R, Merkx R, Fish A, Mahakena S, Bleijerveld OB, Altelaar M, IJlst L, Wanders RJ, Borst P, van de Wetering K. N-lactoyl-amino acids are ubiquitous metabolites that originate from CNDP2-mediated reverse proteolysis of lactate and amino acids. Proc. Natl. Acad. Sci. U.S.A. 2015;112:6601–6606. doi: 10.1073/pnas.1424638112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Richardson S, Gorton L. Characterisation of the substituent distribution in starch and cellulose derivatives. Anal. Chim. Acta. 2003;497:27–65. [Google Scholar]
- 19.Man JM, Yang Y, Huang J, Zhang CQ, Zhang FM, Wang YP, Gu MH, Liu QQ, Wei CX. Morphology and structural properties of high-amylose rice starch residues hydrolysed by amyloglucosidase. Food Chem. 2013;138:2089–2098. doi: 10.1016/j.foodchem.2012.12.009. [DOI] [PubMed] [Google Scholar]
- 20.Marchler-Bauer A, Derbyshire MK, Gonzales NR, Lu SN, Chitsaz F, Geer LY, Geer RC, He J, Gwadz M, Hurwitz DI, Lanczycki CJ, Lu F, Marchler GH, Song JS, Thanki N, Wang ZX, Yamashita RA, Zhang DC, Zheng CJ, Bryant SH. CDD: NCBI's conserved domain database. Nucleic Acids Res. 2015;43:D222–D226. doi: 10.1093/nar/gku1221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Jordan DB, Bowman MJ, Braker JD, Dien BS, Hector RE, Lee CC, Mertens JA, Wagschal K. Plant cell walls to ethanol. Biochem. J. 2012;442:241–252. doi: 10.1042/BJ20111922. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.