Abstract
Background
Oxidative stress can induce cell injury in vascular endothelial cells, which is the initial event in the development of atherosclerosis. Although quantitative real-time polymerase chain reaction (qRT-PCR) has been widely used in gene expression studies in oxidative stress injuries, using carefully validated reference genes has not received sufficient attention in related studies. The objective of this study, therefore, was to select a set of stably expressed reference genes for use in qRT-PCR normalization in oxidative stress injuries in human umbilical vein endothelial cells (HUVECs) induced by hydrogen peroxide (H2O2).
Results
Using geNorm analysis, we found that five stably expressed reference genes were sufficient for normalization in qRT-PCR analysis in HUVECs treated with H2O2. Genes with the most stable expression according to geNorm were U6, TFRC, RPLP0, GAPDH, and ACTB, and according to NormFinder were ALAS1, TFRC, U6, GAPDH, and ACTB.
Conclusion
Taken together, our study demonstrated that the expression stability of reference genes may differ according to the statistical program used. U6, TFRC, RPLP0, GAPDH, and ACTB was the optimal set of reference genes for studies on gene expression performed by qRT-PCR assays in HUVECs under oxidative stress study.
Keywords: Hydrogen peraoxide, Human umbilical vein endothelial cells, qRT-PCR, Reference genes, Normalization
Background
Atherosclerosis (AS) is the leading cause of coronary heart disease, and is associated with high morbidity and mortality [1]. The initial event in its development is vascular endothelial injury induced by oxidative stress, which is associated with changes in gene expression [2, 3]. Gene expression studies are therefore of great importance to oxidative stress injury research. Under pathological conditions, such as ischemia–reperfusion and inflammation, reactive oxygen species (ROS) are generated and lead to vascular endothelial injury [4]. As one of the most common ROS, hydrogen peroxide (H2O2) causes cell and tissue damage through producing the highly reactive radical OH [5, 6]. Thus, H2O2 has been extensively used as an oxidative stimulus to induce oxidative stress in in vitro models [7].
Analysis of gene expression under different physiological and pathological conditions, including oxidative stress, often uses quantitative real-time polymerase chain reaction (qRT-PCR) because of its low template input requirement, high sensitivity, and high specificity [8, 9]. Given that the expression of target genes is normalized to one or more reference genes in this approach, it is of great importance to use an optimal normalizer for improving the accuracy and reliability of expression measurements [10]. However, this assumes that expression of the reference gene remains constant in all cell/tissue types under specific experimental conditions. Unfortunately, increasing data have shown that no single gene is expressed constantly across all cell types or under all physiological/pathological conditions [11–13]. Therefore, to obtain accurate gene expression information, it is imperative that stable reference genes be chosen for the specific type of tissue and experimental condition [14]. GeNorm [13] and NormFinder [15] are the most commonly used methods to evaluate reference genes, but different statistical algorithms are known to cause inconsistent rankings. Candidate genes can be used as reference genes for the normalization of qRT-PCR results if they demonstrate stable expression under different experimental conditions and statistical algorithms [16].
In this study, 15 common reference genes were identified in HUVECs exposed to different concentrations of H2O2. GeNorm and NormFinder software was used to calculate the variability of candidate gene expression and to obtain the most suitable reference genes. This study provides a basis for the selection of reference genes and useful guidelines for future gene expression studies in human umbilical vein endothelial cells HUVECs exposed to H2O2.
Methods
Cell culture and H2O2 studies
HUVECs were purchased from the China Center for Type Culture Collection (Wuhan, China) and cultured in a humidified 5% CO2, 37 °C incubator. The vascular cell basal medium (ATCC, USA) added with the endothelial cell growth kit-BBE (ATCC, USA) was used as the complete growth medium for this cell line, and contained the following components: 0.2% bovine brain extract; 5 ng/mL rh EGF; 10 mmol/L l-glutamine; 0.75 units/mL heparin sulfate; 1 µg/mL hydrocortisone hemisuccinate; 2% fetal bovine serum, and 50 µg/mL ascorbic acid. HUVECs were cultured with different concentrations of H2O2 (0, 500, 1000, 2000, 3000, 4000, 5000, or 6000 µmol/L) for 24 h. Each experiment was performed in triplicate.
Total RNA extraction
Total RNA from HUVECs was extracted using the Eastep® Super Total RNA Extraction Kit (Promega, USA) following the manufacturer’s instructions. Genomic DNA was eliminated by on-column treatment with RNase-free DNase I. The concentration and purity of RNA were measured using a NanoDrop 2000 spectrophotometer (Thermo, USA).
Reverse transcription
Purified RNA was reverse transcribed immediately after extraction with the TranScript One-Step gDNA Removal and cDNA Synthesis SuperMix Kit (Transgen Biotech, China) according to the manufacturer’s instructions. For each sample, cDNA was synthesized from 300 ng total RNA in a final volume of 20 µL and stored at −20 °C until further use.
Quantitative real-time PCR
All primers were purchased from Sangon Biotech, China. Primer sequences are listed in Table 1. qRT-PCR was performed in 96-well plates using the Light Cycler 480 system (Roche, Swiss). Each 20-µL reaction contained 10 µL of TransStart Green qPCR SuperMix (Transgen Biotech, China), 0.5 µL of each primer (10 μmmol/L), 8 µL of ddH2O, and 1 µL of cDNA. PCR conditions were as follows: 95 °C for 1 min, followed by 40 cycles at 95 °C for 20 s, and 61 °C for 31 s.
Table 1.
Name, primer sequences, and product size of candidate reference genes
| Symbol | Gene name | Primer sequences (forward/reverse) | Product length (bp) |
|---|---|---|---|
| 18S | 18S ribosomal RNA | CGGCTACCACATCCAAGGAA/GCTGGAATTACCGCGGCT | 186 |
| GAPDH | Glyceraldehyde-3-phosphate dehydrogenase | GACAGTCAGCCGCATCTTCT/TTAAAAGCAGCCCTGGTGAC | 120 |
| U6 | U6 snRNA | AACGCTTCACGAATTTGCGT/CTCGCTTCGGCAGCACA | 109 |
| ALAS1 | 5′-Aminolevulinate synthase 1 | GGCAGCACAGATGAATCAGA/CCTCCATCGGTTTTCACACT | 150 |
| ACTB | Actin, beta | AGAAAATCTGGCACCACACC/TAGCACAGCCTGGATAGCAA | 173 |
| TFRC | Transferrin receptor | GTCGCTGGTCAGTTCGTGATT/AGCAGTTGGCTGTTGTACCTCTC | 80 |
| PPIA | Peptidylprolyl isomerase A | AGACAAGGTCCCAAAGAC/ACCACCCTGACACATAAA | 118 |
| RPLP0 | Ribosomal protein lateral stalk subunit P0 | CCATTCTATCATCAACGGGTACAA/TCAGCAAGTGGGAAGGTGTAAT | 75 |
| PBGD | Hydroxymethylbilane synthase | AGTGTGGTGGGAACCAGC/CAGGATGATGGCACTGAACTC | 144 |
| GUSB | Glucuronidase beta | AGCCAGTTCCTCATCAATGG/GGTAGTGGCTGGTACGGAAA | 160 |
| B2M | Beta-2-microglobulin | AGCGTACTCCAAAGATTCAGGTT/ATGATGCTGCTTACATGTCTCGAT | 206 |
| HPRT1 | Hypoxanthine phosphoribosyl transferase 1 | GACCAGTCAACAGGGGACAT/CCTGACCAAGGAAAGCAAAG | 132 |
| RPL29 | Ribosomal protein L29 | GGCGTTGTTGACCCTATTTC/GTGTGTGGTGTGGTTCTTGG | 120 |
| PUM1 | Pumilio RNA binding family member 1 | CAGGCTGCCTACCAACTCAT/GTTCCCGAACCATCTCATTC | 211 |
| TBP | TATA-box binding protein | TGCACAGGAGCCAAGAGTGAA/CACATCACAGCTCCCCACCA | 132 |
Statistical analysis
Two versions of Excel-based software, geNorm and NormFinder, were used to evaluate the stability of the candidate reference genes. For both versions, Ct values were converted to relative quantities for analysis according to the formula: [17].
GeNorm software analyzes gene stability based on the average pairwise variation of a particular gene against all other control genes as the M value. Candidate reference genes are ranked according to their expression stability by stepwise exclusion of genes with the highest M-value. Genes with the lowest M value have the most stable expression [13]. To determine the possible need or utility of control genes for normalization, the pairwise variation Vn/n+1 was calculated between the two sequential normalization factors NFn and NFn+1. For the pairwise variation Vn/n+1, 0.2 was taken as a cut-off value, below which the inclusion of an additional control gene is not required [13, 18].
NormFinder software, based on an ANOVA mathematical model, estimates both intra- and inter-group expression variation and calculates a candidate gene stability value. A lower stability value indicates a more stable reference gene expression [15].
Results
Expression profiles of candidate reference genes
Fifteen candidate reference genes in HUVECs were analyzed by qRT-PCR (Table 1), and their Ct values are shown in Fig. 1. The Ct values ranged from 9.225 to 36.19, representing a wide variation, although most were in the range of 22–27. The most highly expressed gene was 18S, which exhibited a median Ct value of 10.66. All other genes had median Ct values larger than 20, while PUM1 presented with the lowest expression level with a median Ct value of 31.2. 18S had the widest range of 14.83 cycles, whereas ALAS1 had the narrowest range of 2.665 cycles. Mean Ct, STDEVP (STD), and coefficient of variation (CV) were calculated as shown in Table 2. CV values for candidate reference genes ranged from 2.89 to 36.64%. ALAS1 had the lowest CV, at 2.89%, indicating the lowest variation in gene expression. By contrast, the CV of 18S was the highest at 36.64%, indicating the highest variation in gene expression.
Fig. 1.
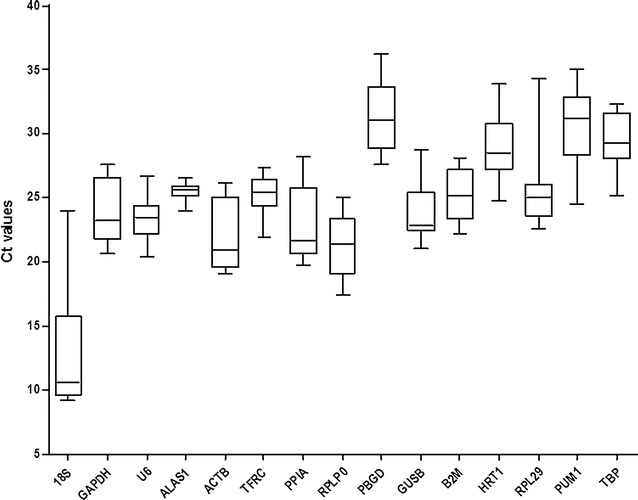
Ct values of 15 candidate reference genes in all samples. Raw Ct values of eight samples, under normal conditions and different concentrations of H2O2, are described using a box and whisker plot. The outer box is determined from the 25th to the 75th percentiles, and the line across the box is the median
Table 2.
Mean Ct, STD, and coefficient of variation of candidate reference genes
| Gene name | Mean Ct | STD | CV (%) |
|---|---|---|---|
| 18S | 13.01 | 4.77 | 36.64 |
| GAPDH | 23.88 | 2.36 | 9.89 |
| U6 | 23.4 | 1.76 | 7.53 |
| ALAS1 | 25.57 | 0.74 | 2.89 |
| ACTB | 21.9 | 2.62 | 11.96 |
| TFRC | 25.24 | 1.58 | 6.27 |
| PPIA | 22.87 | 2.85 | 12.46 |
| RPLP0 | 21.38 | 2.37 | 11.06 |
| PBGD | 31.29 | 2.68 | 8.56 |
| GUSB | 23.86 | 2.3 | 9.62 |
| B2M | 25.2 | 1.95 | 7.75 |
| HRT1 | 28.97 | 2.54 | 8.78 |
| RPL29 | 25.81 | 3.37 | 13.04 |
| PUM1 | 30.52 | 3.05 | 10 |
| TBP | 29.38 | 2.14 | 7.28 |
Expression stability of candidate reference genes
GeNorm analysis
GeNorm software was used to evaluate the stability of candidate reference genes, and the calculated M values are shown in Fig. 2a. A lower M value indicates a higher stability. U6 and TFRC had the lowest M values of 0.97, whereas 18S had the highest M value of 3.35. The three reference genes of highest stability were U6, TFRC, and RPLP0, while 18S, RPL29, and PUM1 showed the lowest stability. Figure 2b shows the pairwise variation for all data. V5/V6 was found to be lower than 0.2, suggesting that the top five reference genes were adequate for normalization, and that an additional 16 reference gene was not necessary.
Fig. 2.

Expression stability values of reference genes analyzed by geNorm software. a Average expression stability measures (M) of reference genes. The x-axis from left to right indicates the ranking of the genes according to their stability; higher M values indicate lower stabilities. b Determination of the suitable number of reference genes required for normalizing. The software calculates the normalization factor from at least two genes, and the V value defines the pair-wise variation between two sequential normalization factors
NormFinder analysis
The expression stability of candidate reference genes was also calculated using NormFinder software. Similar to the GeNorm software, genes with the lowest stability value are the most stable expressed ones. As shown in Fig. 3, the most stable reference gene was ALAS1, following TFRC and U6. The least stable reference genes were 18S, RPL29, and PUM1. A list of candidate reference genes ranked according to stability by the two versions of software is shown in Table 3.
Fig. 3.

Stability values of each reference gene from the NormFinder algorithm. Ranking of candidate reference genes based on stability values calculated by NormFinder
Table 3.
Ranking of candidate reference genes evaluated by geNorm and NormFinder statistical algorithms
| Gene name | geNorm | NormFinder | ||
|---|---|---|---|---|
| Stability value | Rank | Stability value | Rank | |
| U6 | 0.97 | 1 | 0.81 | 3 |
| TFRC | 0.97 | 2 | 0.71 | 2 |
| RPLP0 | 1.08 | 3 | 1.33 | 6 |
| GAPDH | 1.16 | 4 | 1.11 | 4 |
| ACTB | 1.28 | 5 | 1.32 | 5 |
| PBGD | 1.32 | 6 | 1.52 | 8 |
| PPIA | 1.39 | 7 | 1.52 | 7 |
| ALAS1 | 1.54 | 8 | 0.4 | 1 |
| B2M | 2 | 9 | 1.7 | 9 |
| TBP | 2.32 | 10 | 1.79 | 10 |
| HRT1 | 2.56 | 11 | 2.05 | 11 |
| GUSB | 2.75 | 12 | 2.14 | 12 |
| PUM1 | 2.94 | 13 | 2.55 | 13 |
| RPL29 | 3.11 | 14 | 2.69 | 14 |
| 18S | 3.35 | 15 | 3.21 | 15 |
Discussion
ROS are involved in the pathophysiology of cardiovascular diseases such as hyperlipidemia, hypertension, ischemic heart disease, and chronic heart failure [19, 20]. They also cause changes in gene expression, which can be accurately and sensitively measured by qRT-PCR [17, 20]. This technique normalizes the gene of interest against an endogenous control whose expression remains unaltered in the samples under analysis [21]. The concept of validating reference genes used for normalization in qRT-PCR analysis before use was initially suggested in 2002 [22], and has been realized in various scientific disciplines such as plant sciences [23, 24], cancer [25, 26], stem cells [27, 28], and cardiovascular research [14, 29, 30]. Considering that an algorithm is one-sided for evaluating the expression stability of reference genes, many statistical approaches are usually integrated to determine the optimal reference genes under different experimental conditions [17].
The Ct value is used to evaluate gene expression in qRT-PCR analysis. At the same RNA concentration, gene expression levels are negatively associated with Ct values [31, 32]. Generally, neither a very high (threshold cycle Ct > 30) Ct value of a reference gene nor a very low (Ct < 15) is suitable for qRT-PCR [32]. In the present study, the Ct values of the 15 candidate reference genes tested showed large variations across all of the tested samples. The Ct values of PBGD, HRT1, RPL29, PUM1, and TBP in some samples were higher than the threshold value, whereas that of 18S in some samples was lower than the threshold value. Therefore, these six genes should not be used as reference genes in HUVECs treated with H2O2. The Ct value of the remaining candidate reference genes, GAPDH, ALAS1, U6, TFRC, ACTB, PPIA, RPLP0, GUSB, and B2M, ranged from 15 to 30. This analysis indicates that the most suitable reference genes in HUVECs treated with H2O2 should be selected from this list.
CV values can represent the variability of candidate reference genes and reflect their stability to some extent. However, analysis according to CV alone is not sufficiently reliable. In our study, the CV value of TBP is relatively low, indicating a low variation in gene expression. However, according to geNorm and NormFinder software analysis, TBP stability is relatively low. This finding demonstrates the importance of evaluating the stability of reference genes for the normalization of gene expression under different experimental treatments.
Notably, the stability of some reference genes may vary under different conditions. 18S and GAPDH have been widely used as the reference for gene analysis in qRT-PCR [33]. However, our data showed that 18S was the least stable reference gene and GAPDH was not the best choice for gene analysis in HUVECs under H2O2 treatment. ACTB was reported to be unstable in HUVECs in response to hypoxia [34], but was stably expressed in HUVECs treated with H2O2 in the present study. However, RPLP0 and TFRC were reported to be the most stably expressed reference genes in HUVECs treated with hypoxia [34], which was confirmed by our present findings. These findings demonstrated that studying the expression stability of reference genes under different conditions was important for gene expression research.
The ranking for expression stability of reference genes may differ according to the statistical program used. In the present study, we employed two different statistical programs, geNorm and NormFinder, to evaluate gene expression stability in HUVECs treated with H2O2. The majority of the results from both versions of software were the same. For example, following both analyses, the rank of TFRC, GAPDH, and ACTB was shown to be relatively high, while the least stable genes were 18S, RPL29, PUM1, GUSB, HRT1, TBP, and B2M. However, the results from the two software versions showed some differences, notably the ranks of U6 and RPLP0 were different though both relatively high. Considering this fact, it appears that U6 and RPLP0 are relatively stable in HUVECs exposed to H2O2. However, though NormFinder analysis suggested that ALAS1 is the most stable reference gene, this was not confirmed by geNorm software which calculated M > 1.5. In this case, we considered that ALAS1 is not a reliable reference gene in HUVECs treated with H2O2. Some previous reports have indicated that a single gene is not a reliable reference for normalization [13]. For this reason, we propose using a combination of five stably expressed reference genes (U6, TFRC, RPLP0, GAPDH, and ACTB).
In addition, we used extracellular H2O2 as an oxidative stress model, which has been employed worldwide. Besides extracellular H2O2, there are some other models for oxidative stress induction, including extracellular O2 − and normobaric hyperoxia [35]. The toxic effects of extracellular O2 − and H2O2 are almost similar; still, there are some differences between these two models. On the one hand, H2O2 crosses cellular membrane easily, which is an advantage over extracellular O2 −; on the other hand, the effect of H2O2 depends on the cell density. Hyperoxia is a relative measure, defined as oxygen concentration higher than normal. Reactive oxygen species in hyperoxia model are generated intracellular, which is different from extracellular H2O2 and O2 −.
Actually, measurement of a suitable positive control could help to validate these reference genes. However, despite validation, the positive control has no significant influence on the stability evaluation by the calculation software. Thus, many studies [14, 36–39] evaluated suitable reference genes by calculation software without the positive control.
Conclusions
Our study demonstrates that a combination of U6, TFRC, RPLP0, GAPDH, and ACTB is the optimal reference gene set for HUVECs treated with H2O2. These will be useful for studies on gene expression in response to oxidative stress induced by ROS in HUVECs.
Authors’ contributions
TL and BL designed research; TL, HD, LZ, YX and JZ conducted research; NL, YY, XT and BL analyzed data; TL, WS and BL wrote the paper. All authors read and approved the final manuscript.
Acknowledgements
We would like to thank Editage [http://www.editage.cn] for English language editing.
Competing interests
The authors declare that they have no competing interests.
Availability of data and materials
The datasets supporting the conclusions of this article are available upon request.
Funding
This study was supported by grants from the National Natural Science Foundation of China (No. 81570250), Jilin Province Department of Finance, China (No. 2014520) and the National Clinical Key Specialty Project.
Abbreviations
- qRT-PCR
quantitative real-time polymerase chain reaction
- HUVEC
human umbilical vein endothelial cell
- H2O2
hydrogen peroxide
- AS
atherosclerosis
- ROS
reactive oxygen species
- STD
STDEVP
- CV
coefficient of variation
- 18S
18S ribosomal RNA
- GAPDH
glyceraldehyde-3-phosphate dehydrogenase
- U6
U6 snRNA
- ALAS1
5′-aminolevulinate synthase 1
- ACTB
actin, beta
- TFRC
transferrin receptor
- PPIA
peptidylprolyl isomerase A
- RPLP0
ribosomal protein lateral stalk subunit P0
- PBGD
hydroxymethylbilane synthase
- GUSB
glucuronidase beta
- B2M
beta-2-microglobulin
- HPRT1
hypoxanthine phosphoribosyl transferase 1
- RPL29
ribosomal protein L29
- PUM1
pumilio RNA binding family member 1
- TBP
TATA-box binding protein
Contributor Information
Tianyi Li, Email: litianyi_1993@163.com.
Hongying Diao, Email: hydiao70@sina.com.
Lei Zhao, Email: zhaolei0518@aliyun.com.
Yue Xing, Email: xingyuesir@163.com.
Jichang Zhang, Email: jichang@jlu.edu.cn.
Ning Liu, Email: ningliu08@163.com.
Youyou Yan, Email: 420979165@qq.com.
Xin Tian, Email: 479966163@qq.com.
Wei Sun, Email: 546158374@qq.com.
Bin Liu, Phone: 86-431-88796739, Email: liubin3333@vip.sina.com.
References
- 1.Mozaffarian D, Benjamin EJ, Go AS, Arnett DK, Blaha MJ, Cushman M, Das SR, de Ferranti S, Després J-P, Fullerton HJ. Executive Summary: Heart Disease and Stroke Statistics—2016 Update A Report From the American Heart Association. Circulation. 2016;133(4):447–454. doi: 10.1161/CIR.0000000000000366. [DOI] [PubMed] [Google Scholar]
- 2.Kao C-L, Chen L-K, Chang Y-L, Yung M-C, Hsu C-C, Chen Y-C, Lo W-L, Chen S-J, Ku H-H, Hwang S-J. Resveratrol protects human endothelium from H2O2-induced oxidative stress and senescence via SirT1 activation. J Atheroscler Thromb. 2010;17(9):970–979. doi: 10.5551/jat.4333. [DOI] [PubMed] [Google Scholar]
- 3.Ma L, Guo X, Chen W. Inhibitory effects of oleoylethanolamide (OEA) on H2O2-induced human umbilical vein endothelial cell (HUVEC) injury and apolipoprotein E knockout (ApoE−/−) atherosclerotic mice. Int J Clin Exp Pathol. 2015;8(6):6301. [PMC free article] [PubMed] [Google Scholar]
- 4.Yang B, Oo TN, Rizzo V. Lipid rafts mediate H2O2 prosurvival effects in cultured endothelial cells. FASEB J Off Publ Fed Am Soc for Exp Biol. 2006;20(9):1501–1503. doi: 10.1096/fj.05-5359fje. [DOI] [PubMed] [Google Scholar]
- 5.Kamata H, Hirata H. Redox regulation of cellular signalling. Cell Signal. 1999;11(1):1–14. doi: 10.1016/S0898-6568(98)00037-0. [DOI] [PubMed] [Google Scholar]
- 6.Liu L, Gu L, Ma Q, Zhu D, Huang X. Resveratrol attenuates hydrogen peroxide-induced apoptosis in human umbilical vein endothelial cells. Eur Rev Med Pharmacol Sci. 2013;17(1):88–94. [PubMed] [Google Scholar]
- 7.Liu S, Luo R, Xiang Q, Xu X, Qiu L, Pang J. Design and synthesis of novel xyloketal derivatives and their protective activities against H2O2-induced HUVEC injury. Marine Drugs. 2015;13(2):948–973. doi: 10.3390/md13020948. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Bustin S. Quantification of mRNA using real-time reverse transcription PCR (RT-PCR): trends and problems. J Mol Endocrinol. 2002;29(1):23–39. doi: 10.1677/jme.0.0290023. [DOI] [PubMed] [Google Scholar]
- 9.Bustin SA, Benes V, Nolan T, Pfaffl MW. Quantitative real-time RT-PCR—a perspective. J Mol Endocrinol. 2005;34(3):597–601. doi: 10.1677/jme.1.01755. [DOI] [PubMed] [Google Scholar]
- 10.Sun R, He Q, Zhang B, Wang Q. Selection and validation of reliable reference genes in Gossypium raimondii. Biotech Lett. 2015;37(7):1483–1493. doi: 10.1007/s10529-015-1810-8. [DOI] [PubMed] [Google Scholar]
- 11.Bustin SA. Real-time, fluorescence-based quantitative PCR: a snapshot of current procedures and preferences. Expert Rev Molecular Diagn. 2005;5(4):493–498. doi: 10.1586/14737159.5.4.493. [DOI] [PubMed] [Google Scholar]
- 12.Huggett J, Dheda K, Bustin S, Zumla A. Real-time RT-PCR normalisation; strategies and considerations. Genes Immun. 2005;6(4):279–284. doi: 10.1038/sj.gene.6364190. [DOI] [PubMed] [Google Scholar]
- 13.Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A, Speleman F. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002;3(7):1–12. doi: 10.1186/gb-2002-3-7-research0034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Li B, Matter EK, Hoppert HT, Grayson BE, Seeley RJ, Sandoval DA. Identification of optimal reference genes for RT-qPCR in the rat hypothalamus and intestine for the study of obesity. Int J Obes. 2014;38(2):192–197. doi: 10.1038/ijo.2013.86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Andersen CL, Jensen JL, Ørntoft TF. Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res. 2004;64(15):5245–5250. doi: 10.1158/0008-5472.CAN-04-0496. [DOI] [PubMed] [Google Scholar]
- 16.Wei Y, Liu Q, Dong H, Zhou Z, Hao Y, Chen X, Xu L. Selection of reference genes for real-time quantitative PCR in pinus massoniana post nematode inoculation. PLoS ONE. 2016;11(1):e0147224. doi: 10.1371/journal.pone.0147224. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Wu ZJ, Tian C, Jiang Q, Li XH, Zhuang J. Selection of suitable reference genes for qRT-PCR normalization during leaf development and hormonal stimuli in tea plant (Camellia sinensis) Sci Rep. 2016;6:19748. doi: 10.1038/srep19748. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Chang K, Mestdagh P, Vandesompele J, Kerin MJ, Miller N. MicroRNA expression profiling to identify and validate reference genes for relative quantification in colorectal cancer. BMC Cancer. 2010;10(1):173. doi: 10.1186/1471-2407-10-173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Rao GN, Berk BC. Active oxygen species stimulate vascular smooth muscle cell growth and proto-oncogene expression. Circ Res. 1992;70(3):593–599. doi: 10.1161/01.RES.70.3.593. [DOI] [PubMed] [Google Scholar]
- 20.Wang Z, Wang Y, Chen Y, Lv J. The IL-24 gene protects human umbilical vein endothelial cells against H2O2-induced injury and may be useful as a treatment for cardiovascular disease. Int J Mol Med. 2016;37(3):581–592. doi: 10.3892/ijmm.2016.2466. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Egidi MG, Cochetti G, Guelfi G, Zampini D, Diverio S, Poli G, Mearini E. Stability assessment of candidate reference genes in urine sediment of prostate cancer patients for miRNA applications. Dis Mark. 2015;2015:973597. doi: 10.1155/2015/973597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Derveaux S, Vandesompele J, Hellemans J. How to do successful gene expression analysis using real-time PCR. Methods. 2010;50(4):227–230. doi: 10.1016/j.ymeth.2009.11.001. [DOI] [PubMed] [Google Scholar]
- 23.Chen D, Pan X, Xiao P, Farwell MA, Zhang B. Evaluation and identification of reliable reference genes for pharmacogenomics, toxicogenomics, and small RNA expression analysis. J Cell Physiol. 2011;226(10):2469–2477. doi: 10.1002/jcp.22725. [DOI] [PubMed] [Google Scholar]
- 24.Le DT, Aldrich DL, Valliyodan B, Watanabe Y, Van Ha C, Nishiyama R, Guttikonda SK, Quach TN, Gutierrez-Gonzalez JJ. Tran L-SP: Evaluation of candidate reference genes for normalization of quantitative RT-PCR in soybean tissues under various abiotic stress conditions. PLoS ONE. 2012;7(9):e46487. doi: 10.1371/journal.pone.0046487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Rienzo M, Schiano C, Casamassimi A, Grimaldi V, Infante T, Napoli C. Identification of valid reference housekeeping genes for gene expression analysis in tumor neovascularization studies. Clin Transl Oncol. 2013;15(3):211–218. doi: 10.1007/s12094-012-0904-1. [DOI] [PubMed] [Google Scholar]
- 26.Ma Y, Dai H, Kong X, Wang L. Impact of thawing on reference gene expression stability in renal cell carcinoma samples. Diagn Mol Pathol. 2012;21(3):157–163. doi: 10.1097/PDM.0b013e31824d3435. [DOI] [PubMed] [Google Scholar]
- 27.Farrokhi A, Eslaminejad M, Nazarian H, Moradmand A, Samadian A, Akhlaghi A. Appropriate reference gene selection for real-time PCR data normalization during rat mesenchymal stem cell differentiation. Cell Mol Biol (Noisy-le-Grand) 2012;58:OL1660–1670. [PubMed] [Google Scholar]
- 28.Chooi WH, Zhou R, Yeo SS, Zhang F, Wang D-A. Determination and validation of reference gene stability for qPCR analysis in polysaccharide hydrogel-based 3D chondrocytes and mesenchymal stem cell cultural models. Mol Biotechnol. 2013;54(2):623–633. doi: 10.1007/s12033-012-9604-x. [DOI] [PubMed] [Google Scholar]
- 29.Ellefsen S, Bliksøen M, Rutkovskiy A, Johansen IB, Kaljusto M-L, Nilsson GE, Vaage JI, Stensløkken K-O. Per-unit-living tissue normalization of real-time RT-PCR data in ischemic rat hearts. Physiol Genom. 2012;44(12):651–656. doi: 10.1152/physiolgenomics.00004.2012. [DOI] [PubMed] [Google Scholar]
- 30.Tan SC, Carr CA, Yeoh KK, Schofield CJ, Davies KE, Clarke K. Identification of valid housekeeping genes for quantitative RT-PCR analysis of cardiosphere-derived cells preconditioned under hypoxia or with prolyl-4-hydroxylase inhibitors. Mol Biol Rep. 2012;39(4):4857–4867. doi: 10.1007/s11033-011-1281-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Ribeiro MA, dos Reis MB, de Moraes LN, Briton-Jones C, Rainho CA, Scarano WR. Defining suitable reference genes for RT-qPCR analysis on human sertoli cells after 2,3,7,8-tetrachlorodibenzo-p-dioxin (TCDD) exposure. Mol Biol Rep. 2014;41(11):7063–7066. doi: 10.1007/s11033-014-3637-0. [DOI] [PubMed] [Google Scholar]
- 32.Xu X, Liu X, Chen S, Li B, Wang X, Fan C, Wang G, Ni H. Selection of relatively exact reference genes for gene expression studies in flixweed (Descurainia sophia) by quantitative real-time polymerase chain reaction. Pestic Biochem Physiol. 2016;127:59–66. doi: 10.1016/j.pestbp.2015.09.007. [DOI] [PubMed] [Google Scholar]
- 33.Mehta A, Dobersch S, Dammann RH, Bellusci S, Ilinskaya ON, Braun T, Barreto G. Validation of Tuba1a as appropriate internal control for normalization of gene expression analysis during mouse lung development. Int J Mol Sci. 2015;16(3):4492–4511. doi: 10.3390/ijms16034492. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Bakhashab S, Lary S, Ahmed F, Schulten HJ, Bashir A, Ahmed FW, Al-Malki AL, Jamal HS, Gari MA, Weaver JU. Reference genes for expression studies in hypoxia and hyperglycemia models in human umbilical vein endothelial cells. G3. 2014;4(11):2159–2165. doi: 10.1534/g3.114.013102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Gille JJ, Joenje H. Cell culture models for oxidative stress: superoxide and hydrogen peroxide versus normobaric hyperoxia. Mutat Res DNAging. 1992;275(3):405–414. doi: 10.1016/0921-8734(92)90043-O. [DOI] [PubMed] [Google Scholar]
- 36.Tashiro RM, Philips JG, Winefield CS. Identification of suitable grapevine reference genes for qRT-PCR derived from heterologous species. Mol Genet Genom. 2016;291(1):483–492. doi: 10.1007/s00438-015-1081-z. [DOI] [PubMed] [Google Scholar]
- 37.Żyżyńska-Granica B, Koziak K. Identification of suitable reference genes for real-time PCR analysis of statin-treated human umbilical vein endothelial cells. PLoS ONE. 2012;7(12):e51547. doi: 10.1371/journal.pone.0051547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Gharbi S, Shamsara M, Khateri S, Soroush MR, Ghorbanmehr N, Tavallaei M, Nourani MR, Mowla SJ. Identification of reliable reference genes for quantification of microRNAs in serum samples of sulfur mustard-exposed veterans. Cell J (Yakhteh) 2015;17(3):494. doi: 10.22074/cellj.2015.9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Wang H, Wang J, Sun S, Wang Y, Guo J, Ning C, Yang K, Liu JF. Identification of reference microRNAs for quantitative expression analysis in porcine peripheral blood mononuclear cells treated with polyinosinic-polycytidylic acid. Int J Immunogenet. 2015;42(3):217–225. doi: 10.1111/iji.12198. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The datasets supporting the conclusions of this article are available upon request.


