Abstract
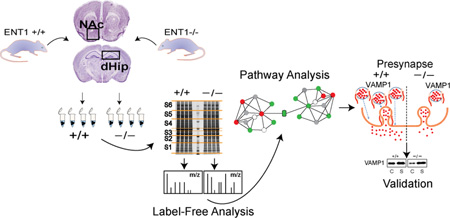
The neural circuit of the dorsal hippocampus (dHip) and nucleus accumbens (NAc) contributes to cue-induced learning and addictive behaviors, as demonstrated by the escalation of ethanol-seeking behaviors observed following deletion of the adenosine equilibrative nucleoside transporter 1 (ENT1−/−) in mice. Here we perform quantitative LC–MS/ MS neuroproteomics in the dHip and NAc of ENT1−/− mice. Using Ingenuity Pathway Analysis, we identified proteins associated with increased long-term potentiation, ARP2/3-mediated actin cytoskeleton signaling and protein expression patterns suggesting deficits in glutamate degradation, GABAergic signaling, as well as significant changes in bioenergetics and energy homeostasis (oxidative phosphorylation, TCA cycle, and glycolysis). These pathways are consistent with previously reported behavioral and biochemical phenotypes that typify mice lacking ENT1. Moreover, we validated decreased expression of the SNARE complex protein VAMP1 (synaptobrevin-1) in the dHip as well as decreased expression of pro-dynorphin (PDYN), neuroendocrine convertase (PCSK1), and Leu-Enkephalin (dynorphin-A) in the NAc. Taken together, our proteomic approach provides novel pathways indicating that ENT1-regulated signaling is essential for neurotransmitter release and neuropeptide processing, both of which underlie learning and reward-seeking behaviors.
Keywords: adenosine, ENT1, hippocampus, nucleus accumbens, neuroproteomics
Graphical Abstract
INTRODUCTION
The hippocampus (Hip) comprises part of a neural circuit that projects and forms a synaptic network in basal ganglia structures, including the nucleus accumbens (NAc). The Hip-NAc circuit orchestrates behaviors involved in assessment of reward value associated with various affective states, as well as consolidation and storage of information related to novel environmental stimuli. However, dysfunction of this circuit contributes to the development of addiction and anxiety.1,2 More specifically, the neural circuit connecting the dorsal Hip (dHip) and NAc has been shown to regulate natural and drug rewards, thereby contributing to the development of alcohol use disorder (AUD) as well as3,4 cocaine- and amphetamine-seeking behaviors.5–7 Interestingly, the co-occurrence of drug-induced positive reinforcement and associated environmental cues potentiates the dHip-NAc circuit, strengthening contexual drug-seeking8,9 and relapse, utilizing glutamatergic, GABAergic, and adenosinergic systems.10,11
Adenosine signaling plays an essential role in fine-tuning glutamatergic and GABAergic neurotransmission in cortical, hippocampal, and striatal circuits involved in learning, motivation, and addiction.12–14 Previously, we demonstrated that deletion of the ethanol-sensitive adenosine transporter (ENT1−/−) decreases extracellular adenosine and presynaptic adenosine A1 receptor function, producing a hyper-glutamatergic state in the NAc.15,16 These effects are associated with tolerance to the intoxicating effects of alcohol and increased alcohol drinking in mice.
To elucidate the proteomic signatures of the dHip and NAc in mice lacking ENT1, we employed label-free LC–MS/MS. Advances in proteomic methodologies has become a powerful instrument for hypothesis generation and discovery of novel targets for the treatment of alcoholism.17,18 Label-free neuroproteomics, when used concomitantly with well-curated bioinformatic algorithms, provide a platform that can facilitate understanding of neurobiological pathology and disease mechanisms. Moreover, label-free technology permits a cost-effective increase in proteome coverage of relative comparisons of protein expression derived from technical and biological replicates19
In this study, we highlight a pivotal role of ENT1 in the regulation of proteins involved in mitochondrial bioenergetics, actin-cytoskeleton reorganization, synaptic plasticity, and amino acid degradation–biosynthesis within the neural circuit that incorporates the dHip and NAc. Moreover, our validation of altered proteins critically involved in neurotransmitter release and neuropeptide processing exemplifies how discovery neuroproteomics can aid understanding adenosine’s role in the development of AUD and other maladaptive neuro-psychiatric conditions.
MATERIALS AND METHODS
Detailed materials and methods are provided in the Supporting Information.
Animals
ENT1−/− and WT mice were cared for under a 12 h/12 h light-dark cycle with lights on at 6am and off at 6 pm, and generated as previously described.16,20 All mice selected for this study were littermates that were group housed and age matched at 3-to-5 months of age. Mice were permitted ad libitum access to rodent chow and water. All experimental procedures were approved by the Mayo Clinic Institutional Animal Care and Use Committee in accordance with NIH guidelines.
Proteomic Analysis Cohorts
To examine the neuroproteome of the dHip (cytosolic fraction),we used equal numbers of ENT1−/− and WT mice (n = 5). For neuroproteomic analysis of the NAc (cytosolic fraction), a separate cohort of ENT1−/− (n = 3) and WT mice (n = 4) was utilized.
Validation Cohorts
We validated our neuroproteomic findings from the dHip with Western blot (WB) assessment of the cytosolic fraction from a separate cohort of ENT1−/− (n = 7) and WT (n = 6) mice. Of these, equal numbers of each genotype (n = 3) were assessed for protein expression in synaptosomes. For the NAc, we validated our neuroproteomic findings via WB of the cytosolic fraction of a separate cohort of ENT1−/− (n = 4) and WT (n = 3) mice.
Tissue and Gel Preparation for Neuroproteomic Analysis
The dHip and NAc from both hemispheres of WT and ENT1−/− mice were hand dissected with the aid of a surgical microscope21 and subsequently processed to isolate the cytosolic fractions for SDS-PAGE (Figure 1). The processed fractions from individual biological replicates of the dHip and NAc were assayed for protein concentration, loaded into polyacrylamide gels at 30 µg/lane and separated via electrophoresis using standard methods (see Supporting Information for details). Gels were fixed and stained with Bio-Safe Coomasie G-250 and destained with water. Stained gel photographs from the dHip (Supplementary Figure 1A) and NAc (Supplementary Figure 1B) were taken with the Gel Doc XR+ Imager (Bio-Rad).
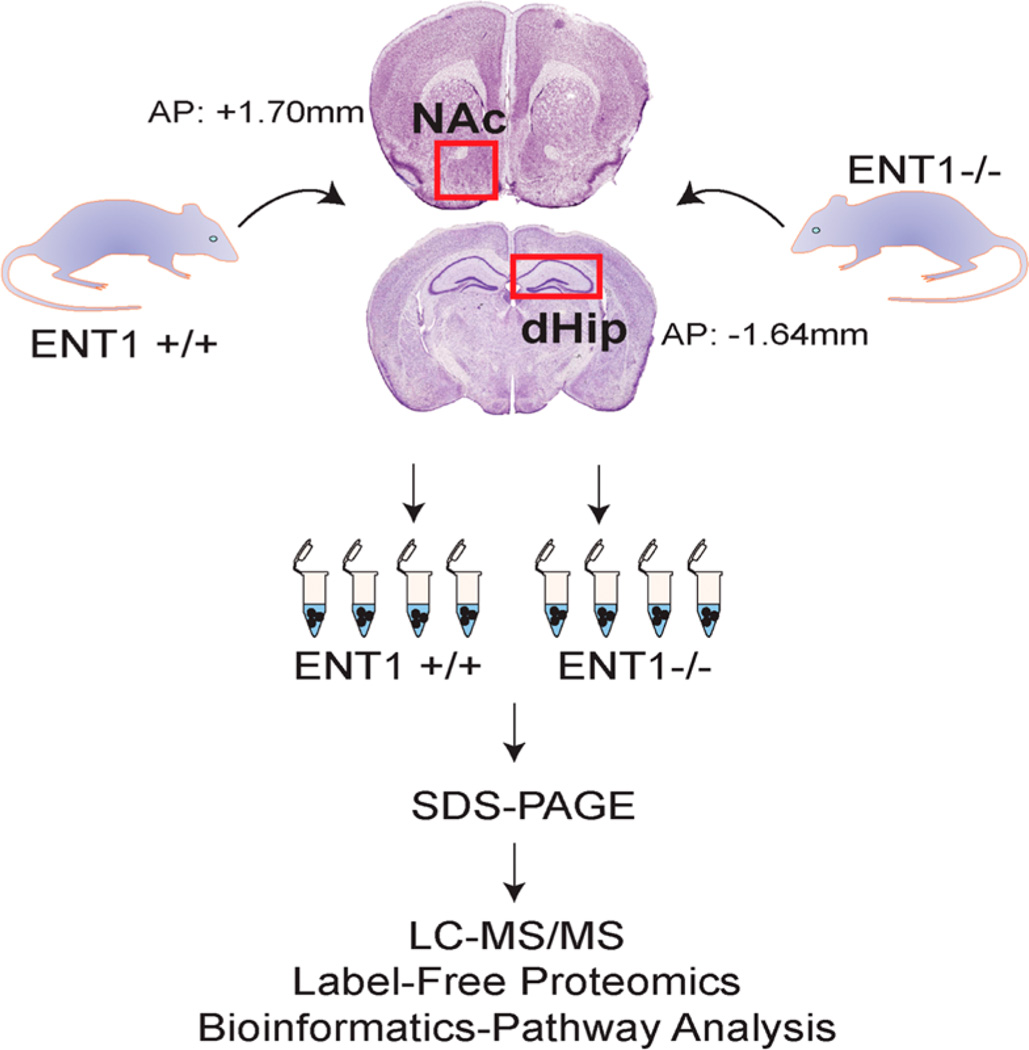
Figure 1.
Label-free neuroproteomic workflow for the dorsal hippocampus and nucleus accumbens of WT (ENT1+/+) and ENT1−/− mice. AP, anterior-posterior brain atlas coordinates relative to Bregma; dHip, dorsal hippocampus; NAc, nucleus accumbens.
Label-Free Proteomics
Gel Sectioning
To examine the effect of ENT1 deletion on relative basal protein expression in the dHip and NAc, we employed a label-free proteomics method. The gel containing the biological replicates from the dHip of WT and ENT1−/− mice was divided into four sections (Supplementary Figure 1A). In an attempt to gain deeper coverage of the NAc proteome, the gel containing the biological replicates from the NAc of WT and ENT1−/− mice was divided into six sections (Supplementary Figure 1B).
Sample Preparation for Proteomics of the dHip
The resulting gel fractions from the dHip underwent destaining, dehydration, alkylation, and rehydration steps prior to trypsin digestion (see Supporting Information for details). Gel segments were digested with trypsin (Promega, Fitchburg, WI) overnight, and peptides were extracted by adding 2% trifluoroacetic acid (TFA). The supernatant was removed and saved, and the peptide-containing supernatants were dried under vacuum and stored at −20 °C until LC–MS/MS analysis.
LC–MS/MS of the dHip
Peptides were analyzed by LC– S/MS on an LTQ Orbitrap (Thermo Fisher Scientific) interfaced to an Eksigent Nano Liquid Chromatography-2D system (Eksigent, Dublin, CA). Survey scans were acquired in the Orbitrap at 60 000 resolving power using a target ion population of 1 × 106 charges. Data-dependent MS/MS was conducted on the top 5 doubly or triply charged precursor masses using an isolation width setting of 2.0, normalized collision energy of 35%, and a linear ion trap target ion population of 8 × 103 charges.22 Precursor masses selected for MS/MS experiments were placed on an exclusion list for 45 s.
Bioinformatic Analysis of the dHip
Data files from our MS/MS analysis were imported into Rosetta Elucidator software (Seattle, WA).23,24 Features were detected such that m/z, retention time, and peak intensity alignment of MS1 data were extracted across samples. Database interrogation was initiated within Elucidator using Mascot (v2.2, Matrix Science, Boston, MA) with mouse UniProt/Swiss-Prot database. Annotation was performed using the Peptide Prophet implementation in Elucidator.
Sample Preparation for Proteomics of the NAc
Preparation of the NAc for label-free proteomics was performed as described for the dHip. Gel sections were destained, digested with trypsin, reduced with dithiothreitol, and alkylated with iodoacetamide as previously described.25 Peptides were extracted from gel section fragments, followed by evaporation to dryness on a vacuum concentrator and stored at −80 °C until LC–MS/MS analysis (see Supporting Information for complete methods).
LC–MS/MS of the NAc
Peptides were analyzed by LC-MS/MS on a QExactive mass spectrometer (Thermo-Fisher Scientific) interfaced with a Dionex UltiMate 3000 RSLC liquid chromatography system (Thermo-Fisher Scientific). The instrument was operated in data-dependent mode by collecting MS1 data at 70 000 resolving power, and precursors were fragmented with normalized collision energy of 27, fragments measured at 17 500 resolving power, and a fixed first mass of 140. MS/MS spectra were collected on the top 15 precursor masses present in each MS1 using an AGC value of 1 × 105, max ion fill time of 100 ms, an isolation window of 3.0 Da, isolation offset of 0.5 Da, and a dynamic exclusion time of 60s (see Supporting Information for complete methods).
Bioinformatic Analysis of the NAc
We utilized MaxQuant (version 1.5.1) software to process the raw data files to produce a list of protein groups and their corresponding intensities in each sample.26 MaxQuant was configured to use a composite mouse protein sequence database containing UniProt mouse reference proteome. The software filtered peptide and protein identifications at 2% false discovery rate (FDR), grouped protein identifications into groups, and reported protein group intensities.
Pathway Analysis and Validation
Pathway Analysis
Proteins identified in the dHip with Rosetta Elucidator and in the NAc with MaxQuant were subsequently uploaded into Ingenuity Pathway Analysis (IPA) for stratification and categorization of direct and indirect network interactions using an IPA (Qiagen, Redwood City, CA) functional analysis algorithm and the curated IPA Ingenuity Knowledge Base (IPAIKB). A CORE analysis for the dHip and NAc was performed using default IPA settings and excluding cancer cell lines. Significance expression value threshold filters were set to a 1.4 fold change (FC) ratio between WT and ENT1−/− mice and a confidence value of p ≤ 0.05.
Synaptosome Enrichment
For examination of enriched proteins from synaptosomes in the dHip of WT and ENT1−/− mice, we homogenized the tissue for 2 min at a speed setting of 2 (Storm 24 Bullet Blender, 0.5 mm ZrO2 beads) in combination with Syn-PER synaptic protein extraction reagent (1 mg tissue: 5 µL of buffer) containing protease and phosphatase inhibitor cocktails (Thermo Scientific). The homogenized tissue (including beads) was centrifuged at 1200g (4 °C) for 10 min. Following centrifugation, the resulting main homogenate (MH) supernatants were transferred to a precooled 1.5 mL tube and the pellet (including beads) was discarded. A 10 µL aliquot of the MH was saved from each replicate for subsequent WB analysis. The MH was then centrifuged for 15 min at 15 000g (4 °C), and the resulting supernatant (cytosolic fraction) was saved for WB analysis. The resulting synaptosome pellet was resuspended at a ratio of 1 g starting tissue: 2000 µL of Syn-PER reagent. Protein concentrations were determined from each replicate of the MH, cytosolic fraction, and synaptosome pellet, as described above. Because of the low quantity of starting material, technical replicates were combined from each step of the enrichment process (MH, cytosolic fraction, synaptosomes) for each genotype by aliquoting equal protein concentrations (the lowest concentration) from each replicate into a single pooled master suspension. Protein concentration for each master suspension was determined as described above; 20 µg was loaded in triplicate for SDS-PAGE, which was followed by immunoblotting as described below. The MH fraction was loaded into a single lane and not used for statistical analysis.
Western Blot Validation
For cytosolic protein expression analysis from the dHip and NAc of WT and ENT1−/− mice, tissue extraction and protein concentrations were quantified using standard methods (for details, see the Supporting Information). In brief, all replicate brain samples were loaded at 20 µg and separated on a 4–12% NuPage Bis-Tris gel (Invitrogen), followed by transfer onto a PVDF membrane. Membranes were then immunoblotted overnight at 4 °C with primary antibodies specific for synaptobrevin (VAMP1; 1:500), pro-dynorphin (PDYN; 1:500), neuro-endocrine convertase (PCSK1; 1:1000), Leu-Enkephalin (Leu-ENK; 1:1000), GAPDH (1:1000), and appropriate secondary antibodies. Images were developed on a Kodak Image Station 4000R scanner (New Haven, CT), and band optical density quantification was performed using NIH ImageJ software.
Statistical Analysis
Statistical analysis resulting from associations between our reference and focus protein data sets with canonical pathways or upstream regulators was performed by the IPA functional analysis algorithm using a right-tailed Fisher’s Exact Test, where p ≤ 0.05 was considered statistically significant.
For WB analysis of cytosolic VAMP1, PDYN, PCSK1, and Leu-ENK, optical densities for all replicates were normalized to GAPDH protein expression. Protein expression differences were determined with a two-tailed unpaired Student’s t test with a significance cutoff of p ≤ 0.05. For analysis of VAMP1 enrichment in synaptosomes, protein expression optical densities for all sample replicates were normalized to their respective GAPDH protein expression. These values were then normalized to the average expression observed in the cytosolic compartment. The resulting values for each sample replicate in the WB experiments were subsequently analyzed with a two-tailed unpaired Student’s t test or a one-way ANOVA, followed by Bonferroni’s multiple comparisons where appropriate. Results were considered statistically significant when p ≤ 0.05. All statistical analyses were done using Prism (Version 5, GraphPad Software, La Jolla, CA).
RESULTS
Proteomic Analysis of the Dorsal Hippocampus and Nucleus Accumbens in Mice Lacking ENT1
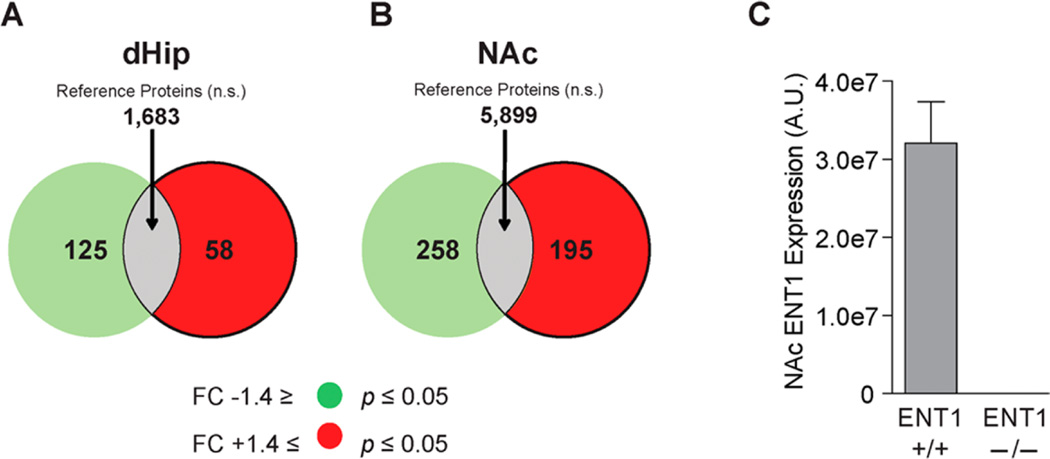
LC–MS/MS protein expression analysis was utilized to investigate the proteome of the dHip and NAc. We utilized the following criteria to identify significantly altered proteins: (1) An FC expression ratio of greater than ±1.4 and (2) a corresponding significance value of p ≤ 0.05. We detected significantly downregulated (green) and upregulated (red) proteins in the dHip (Figure 2A) and NAc (Figure 2B) as well as reference data set proteins that did not meet these criteria (gray). As shown in the Venn diagram (Figure 2A), we identified a total of 1866 proteins in the dHip (Supplementary Table 1). Of these, 58 were significantly decreased, while 125 were significantly increased in ENT1−/− mice. In the NAc, we identified a total of 6352 proteins, of which 258 were significantly decreased and 195 were significantly increased in ENT1−/− mice (Figure 2B and Supplementary Table 2). Moreover, we confirmed that ENT1 was absent in the NAc of ENT1−/− mice (Figure 2C and Supplementary Table 2). These data indicate that label-free proteomics can elucidate the profound effects of ENT1 deletion on protein expression profiles of the dHip and NAc neural circuits.
Figure 2.
Differential protein expression in the dorsal hippocampus and nucleus accumbens of ENT1−/− mice. Venn diagram showing significantly decreased focus proteins (green), significantly increased proteins (red) in the (A) dHip and (B) NAc of ENT1−/− relative to WT (ENT1+/+) mice. The gray area in the center of the Venn diagram (overlap) indicates the number of identified proteins that did not meet focus protein criteria. Focus protein threshold criteria was a fold chance of ±1.4 and relative comparison significance of p ≤ 0.05. (C) Label-free analysis in the NAc of ENT1−/− confirmed deletion of ENT1 in comparison to expression levels in ENT1+/+ mice. dHip, dorsal hippocampus; NAc, nucleus accumbens.
Pathway Analysis of Protein Profile
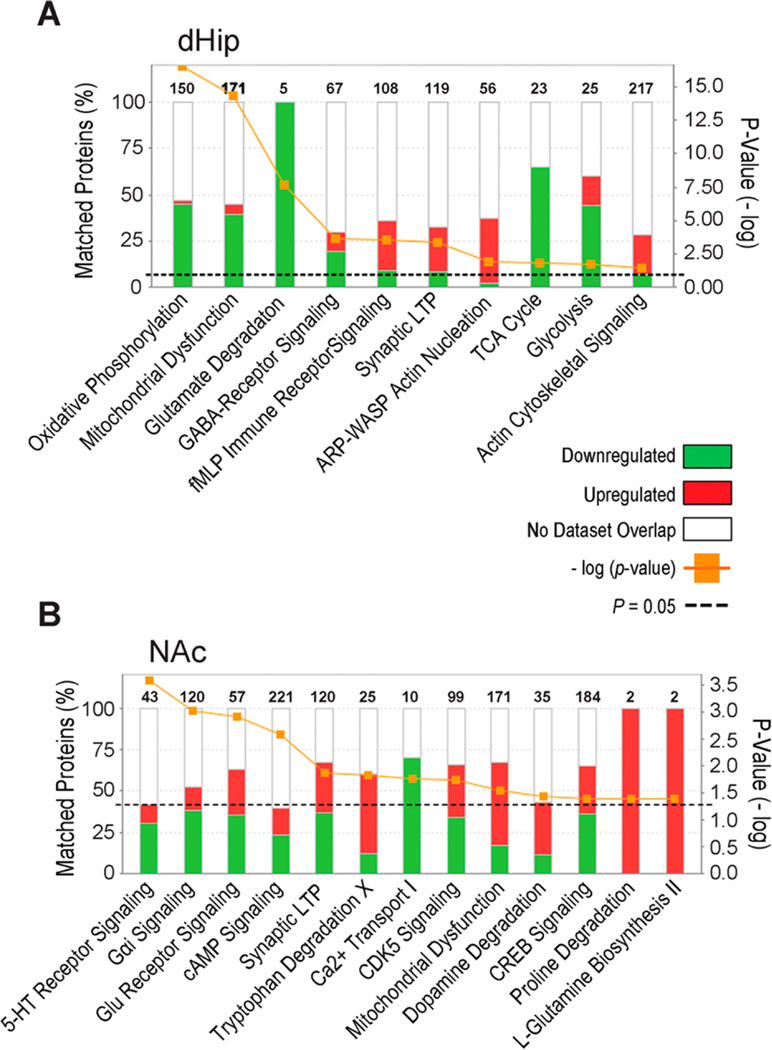
To gain more insight into the differential effects of ENT1 deletion on the dHip and NAc neuroproteome, IPA mapped reference and focus proteins from both brain regions to specific biological canonical pathways. As shown in Figure 3A and Table 1, glutamate degradation (p ≤ 0.05) was a top canonical pathway associated with significantly downregulated proteins in the dHip (5/5 down; GAD2, SUCLG2, GABA-T, ALDH5A1). Likewise, proteins involved in GABA-receptor signaling (12/47 down, 6/47 up) were significantly downregulated (PVALB, CALB2, SLC6A11, SLC32A1), suggesting a possible deficit in GABA-mediated regulation of glutamatergic function (p ≤ 0.05). Similarly, proteins involved in bioenergetic processes such as oxidative phosphorylation (Ox-Phos) and mitochondrial dysfunction (Complex I-to-Complex V; down 59/170, up 11/150) as well as glycolysis (ALDOA) and the TCA cycle (IDH3, SDHB) also underwent significant decreases in protein expression (Figure 3A and Table 1) in the dHip (all p ≤ 0.05). In contrast, Figure 3A and Table 1 show that ENT1 deletion induces significantly increased (PKC-β, PKC-γ, GRIA2, CAMKV, MEK1) and decreased (VAMP1, VAPB, VAT1L, PPP1R14A) in expression of proteins (29/119 up, 10/119 down) associated with synaptic long-term potentiation (LTP), suggesting the augmentation of this pathway in ENT1−/− mice (p ≤ 0.05). Not surprisingly, related pathways involved in regulation of synaptic plasticity, including actin nucleation and cytoskeletal signaling (all p ≤ 0.05) associated with the ARP2/ 3-WASP proteins ARPC2, ARPC5, WASF1, WASL, and WIPF3, which were also significantly upregulated in ENT1−/− mice (22/66 up, 1/66 down). Additionally, our analysis detected significant changes (p ≤ 0.05) in calcineurin-induced (PPP3CA, PPP3CB, PPP3R1) activation of inflammatory signaling mediated by the N-formyl-Met-Leu-Phe (fMLP) receptor (29/108 up, 10/108 down).
Figure 3.
Ingenuity Pathway Analysis identifies canonical pathways associated with expression of proteins in the dorsal hippocampus and nucleus accumbens of ENT1−/− mice relative to WT expression. (A) Top canonical pathways associated with significantly altered focus proteins in the dHip of ENT1−/− mice. (B) Top canonical pathways associated with significantly altered focus proteins in the NAc of ENT1−/− mice. Orange line indicates the p value of association between reference and focus proteins for each pathway. Black dashed line indicates threshold of significance for Fisher’s right tailed test *p < 0.05. Numbers on top of pathways indicate the total number of proteins associated with that pathway. Red indicates % of focus proteins upregulated that matched each pathway. Green indicates % of focus proteins downregulated that matched each pathway. dHip, dorsal hippocampus; NAc, nucleus accumbens.
Table 1.
Top Focus Proteins and Associated Canonical Pathways Identified in the Dorsal Hippocampus of ENT1−/− Mice
| UniProt KB Swiss Prot ID | protein name | protein description | canonical pathway | fold change | P value |
|---|---|---|---|---|---|
| P63328 | PPP3CA | protein phosphatase 3, catalytic alpha | fMLP neutrophil signaling | 1.42 | 2.2 × 10−5 |
| P48453 | PPP3CB | protein phosphatase 3, catalytic beta | fMLP neutrophil signaling | 1.42 | 9.3 × 10−7 |
| Q63810 | PPP3R1 | protein phosphatase 3, regulatory B alpha | fMLP neutrophil signaling | 1.51 | 5.5 × 10−5 |
| P63318 | PRKCG | protein kinase C, gamma | synaptic LTP | 1.55 | 2.1 × 10−4 |
| P68404 | PRKCB | protein kinase C, beta | synaptic LTP | 1.38 | 8.0 × 10−3 |
| P31938 | MAP2K1 | MEK1 | synaptic LTP | 1.57 | 8.7 × 10−4 |
| P63085 | MAPK1 | ERK2 | synaptic LTP | 1.24 | 4.0 × 10−3 |
| Q62442 | VAMP1 | synaptobrevin 1 | synaptic LTP | −1.82 | 2.0 × 10−3 |
| Q9QY76 | VAPB | VAMP-associated protein B | synaptic LTP | −1.42 | 7.9 × 10−5 |
| Q80TB8 | VAT1L | vesicle amine transport protein 1 homologue | synaptic LTP | −3.52 | 5.0 × 10−4 |
| Q91VC7 | PPP1R14A | protein phosphatase 1, regulatory 14A | synaptic LTP | −1.61 | 6.0 × 10−3 |
| P48320 | GAD2 | glutamate decarboxylase 2 | Glu degradation | −1.58 | 6.0 × 10−3 |
| Q9Z2I8 | SUCLG2 | succinate-CoA ligase beta | Glu degradation | −1.58 | 4.8 × 10−4 |
| Q8BWF0 | ALDH5A1 | aldehyde dehydrogenase 5A1 | Glu degradation | −1.51 | 9.0 × 10−6 |
| P61922 | GABA-T | GABA transaminase | Glu degradation | −1.73 | 1.1 × 10−2 |
| Q8BLE7 | SLC17A6 | VGLUT2 | Glu receptor signaling | −1.66 | 1.5 × 10−5 |
| P23819 | GRIA2 | AMPA2, glutamate lonotropic receptor | Glu receptor signaling | 1.41 | 5.0 × 10−3 |
| P03995 | GFAP | glial fibrillary acidic protein | Glu receptor signaling | −1.25 | 1.5 × 10−1 |
| P32848 | PVALB | parvalbumin | GABA receptor signaling | −2.04 | 6.0 × 10−3 |
| Q08331 | CALB2 | calbindin 2 | GABA receptor signaling | −3.02 | 2.3 × 10−6 |
| O35633 | SLC32A1 | GABA vesicular transporter | GABA receptor signaling | −1.93 | 4.7 × 10−4 |
| P31650 | SLC6A11 | GABA transporter | GABA receptor signaling | −3.51 | 1.5 × 10−2 |
| Q03265 | ATP5A1 | ATP synthase, F1 complex, alpha 1 | mitochondrial dysfunction | −1.90 | 4.0 × 10−3 |
| P56382 | ATP5E | ATP synthase, F1 subunit E | mitochondrial dysfunction | −1.42 | 1.7 × 10−2 |
| Q06185 | ATP5I | ATP synthase, Fo complex, subunit E | mitochondrial dysfunction | −1.53 | 8.3 × 10−5 |
| Q9CPQ8 | ATP5L | ATP synthase, subunit G | mitochondrial dysfunction | −1.58 | 3.3 × 10−7 |
| Q9CPQ1 | COX6C | cytochrome C oxidase subunit Vic | mitochondrial dysfunction | −1.79 | 2.1 × 10−2 |
| Q9CQ75 | NDUFA2 | NADH dehydrogenase-complex I | mitochondrial dysfunction | −1.89 | 4.2 × 10−2 |
| Q99LC3 | NDUFA10 | NADH dehydrogenase-complex I | mitochondrial dysfunction | −1.53 | 7.7 × 10−2 |
| Q9CQA3 | SDHB | succinate dehydrogenase-complex II | TCA cycle | −1.56 | 2.0 × 10−3 |
| Q9CQ69 | UQCRQ | Ubiq-Cyt C reductase-complex III | mitochondrial dysfunction | −1.77 | 1.6 × 10−2 |
| Q9D855 | UQCRB | Ubiq-Cyt C reductase-complex III | mitochondrial dysfunction | −1.80 | 2.9 × 10−2 |
| Q9D6R2 | IDH3A | isocitrate dehydrogenase 3 alpha | TCA cycle | −1.46 | 3.8 × 10−4 |
| Q05920 | PC | pyruvate carboxylase | mitochondrial dysfunction | −1.48 | 1.2 × 10−5 |
| Q64669 | NQO1 | NAD(P)H dehydrogenase, quinone 1 | mitochondrial dysfunction | −1.53 | 4.0 × 10−4 |
| Q9QXV0 | PCSK1N | proSAAS | neuropeptide signaling | −2.14 | 1.0 × 10−3 |
| P05064 | ALDOA | aldolase A, fructose-bisphosphate | glycolysis | −1.49 | 2.0 × 10−3 |
| Q9EQ20 | ALDH6A1 | aldehyde dehydrogenase 6A1 | Val and β-ala degradation | −1.76 | 3.7 × 10−2 |
| Q9JLZ3 | AUH | AU RNA bind protein/enoyl-CoA hydratase | β-ala degradation | −1.46 | 4.0 × 10−3 |
| Q99L13 | HIBADH | 3-hydroxyisobutyrate dehydrogenase | β-ala degradation | −1.48 | 6.6 × 10−6 |
| P28650 | ADSSL1 | adenylosuccinate synthase like 1 | purine synthesis | −1.81 | 7.1 × 10−7 |
| Q920P5 | AK5 | adenylate kinase 5 | purine synthesis | 1.57 | 4.4 × 10−5 |
| Q8R5H6 | WASF1 | Wiskott-Aldrich syndrome protein 1 | actin nucleation ARP-WASP | 1.54 | 1.5 × 10−7 |
| P0C7L0 | WIPF3 | WAS/WASL interacting protein 3 | actin nucleation ARP-WASP | 1.62 | 3.6 × 10−5 |
| Q91YD9 | WASL | Wiskott-Aldrich syndrome-like | actin nucleation ARP-WASP | 1.42 | 5.6 × 10−10 |
| Q9CVB6 | ARPC2 | actin protein 2/3 complex, subunit 2 | actin nucleation ARP-WASP | 1.52 | 3.3 × 10−18 |
| Q9CPW4 | ARPC5 | actin protein 2/3 complex, subunit 5 | actin nucleation ARP-WASP | 1.63 | 7.8 × 10−4 |
| P61148 | FGF1 | fibroblast growth factor 1 | Rac-RhoA signaling | −1.79 | 9.5 × 10−4 |
| Q9QVP9 | PTK2B | PTK2B protein tyrosine kinase 2 beta | Rac-RhoA signaling | 1.43 | 9.1 × 10−4 |
| Q3UQ44 | IQGAP2 | IQ motif GTPase activating protein 2 | Rac-RhoA signaling | 1.49 | 1.2 × 10−2 |
| P62814 | ATP6V1B2 | V- ATPase subunit B2, brain | axoskeleton maintenance | −1.52 | 3.5 × 10−2 |
| P46660 | INA | alpha-internexin | axoskeleton maintenance | −1.59 | 1.1 × 10−2 |
| P04370 | MBP | myelin basic protein | axoskeleton maintenance | −1.54 | 1.7 × 10−4 |
| P08551 | NEFL | neurofilament, light polypeptide | axoskeleton maintenance | −1.77 | 1.0 × 10−2 |
| P19246 | NEFH | neurofilament, heavy polypeptide | axoskeleton maintenance | −2.36 | 8.0 × 10−3 |
| P08553 | NEFM | neurofilament, medium polypeptide | axoskeleton maintenance | −1.76 | 1.0 × 10−3 |
| P26645 | MARCKS | myristoylated ala-protein kinase C | plasma membrane anchoring | −1.46 | 6.0 × 10−3 |
| Q91XV3 | BASP1 | brain abundant signal protein 1 | plasma membrane anchoring | 4.98 | 4.0 × 10−3 |
| Q60625 | ICAM5 | telencephalin | cell-cell adhesion | 1.53 | 4.5 × 10−4 |
| P11627 | L1CAM | neural cell adhesion molecule-L1 | cell-cell adhesion | −1.43 | 2.0 × 10−4 |
| O35136 | NCAM2 | neural cell adhesion molecule 2 | cell-cell adhesion | −1.50 | 8.5 × 10−4 |
|
Control P16858 |
GAPDH | glyceraldehyde-3-phos dehydrogenase | −1.00 | 7.5 × 10−2 |
Next, we examined the top canonical pathways associated with altered proteins identified in the NAc of ENT1−/− mice. As shown in Figure 3B and Table 2, our analysis revealed that differentially expressed proteins in the NAc were significantly associated with glutamate (Glu) receptor signaling (16/57 up, 20/57 down), GαI signaling (17/120 up, 46/120 down), mitochondrial dysfunction (86/171 up, 29/171 down), cAMP signaling (51/221 down, 36/221 up), synaptic LTP (44/120 down, 37/120 up), Ca2+ transport (7/10 down, 0/10 up), CREB signaling (66/184 down, 55/181 down), 5-HT receptor signaling (13/43 down, 5/43 up), CDK5 signaling (34/99 down, 31/99 up), tryptophan degradation (3/25 down, 12/25 up), dopamine degradation (4/35 down, 11/35 up), praline degradation (0/2 down, 2/2 up), and l-glutamine biosynthesis (0/2 down, 2/2 up). As displayed in Table 2, we observed significant decreases in the expression of the metabotropic glutamate receptors (GRM3, GRM7, GRM8), and the glutamate transporters EAAT2 (SLC1A2), GLAST (SLC1A3), all of which significantly associated with synaptic LTP and glutamate receptor signaling. Our analysis also identified significant increases in expression of enzymes involved in 5-HT receptor signaling s well as tryptophan and dopamine degradation (MAOA, MAOB). We detected several proteins involved in SNARE complex function including synaptotagmin (SYT7, SYT13) and SNAP 25. An examination of the overall changes in protein expression shared between the dHip and the NAc showed that several altered proteins exert similar effects on well-characterized canonical pathways. Finally, we detected the potassium channel protein TREK-1 (KCKN2), which is involved in ion transport in astrocytes as well as the c-Myc-responsive protein Rcl (DNPH1), which has been implicated in cell death and survival (Table 2). Notably, our proteomic analysis did not detect expression of these proteins in the NAc of ENT1−/− mice.
Table 2.
Top Focus Proteins and Associated Canonical Pathways Identified in the Nucleus Accumbens of ENT1−/− Mice
| UniProt KB Swiss Prot ID |
protein name |
protein description | canonical pathway | fold change | P valuea |
|---|---|---|---|---|---|
| Q80VJ3 | DNPH1 | 2-deoxynucleoside 5-phosp N-hydrol 1 | cell death and survival | −2.04 × 1011 | – |
| P97438-2 | KCNK2 | K+ Chan subfam K2 (TREK-1) | ion transport in astrocytes | −7.22 × 109 | – |
| P56564 | SLC1A3 | EAAT1 (GLAST) | GluR signaling, Syn-LTP | −3.75 | 8.7 × 10−3 |
| P28571-1 | SLC6A9 | solute carrier fam 6 member 9-Gly trans | GluR signaling, Syn-LTP | −2.37 | 3.8 × 10−2 |
| P43006 | SLC1A2 | EAAT2 (GLT-1) | GluR signaling, Syn-LTP | −2.91 | 9.1 × 10−3 |
| Q9QYS2 | GRM3 | Glu-R, metabotropic 3 | GluR and CREB signaling, Syn-LTP | −2.43 | 6.5 × 10−4 |
| Q05BD6 | GRM8 | Glu-R, metabotropic 8 | GluR and CREB signaling, Syn-LTP | −2.36 | 1.1 × 10−4 |
| O35874 | SLC1A4 | solute carrier family 1 member 4 | GluR signaling, Syn-LTP | −2.12 | 1.6 × 10−2 |
| Q68ED2 | GRM7 | Glu-R, metabotropic 7 | GluR and CREB signaling, Syn-LTP | −1.63 | 4.7 × 10−4 |
| Q543V3 | IRS1 | insulin receptor substrate 1 | CREB signaling | −1.68 | 3.5 × 10−2 |
| Q9WU79 | PRODH | proline dehydrogenase 1 | proline degradation, GluR signaling | 1.62 | 1.3 × 10−3 |
| Q99M46 | POLR2C | polymerase (RNA) II subunit C | CREB signaling | 1.57 | 7.6 × 10−3 |
| O08530 | S1PR1 | sphingosine-1-phosphate receptor 1 | Gal signaling, cAMP signaling | −4.78 | 2.5 × 10−6 |
| P49817 | CAV1 | caveolin 1 | Gal signaling, cAMP signaling | −2.01 | 2.6 × 10−4 |
| P97490 | ADCY8 | adenylate cyclase 8 (brain) | Gal signaling, cAMP signaling | −1.56 | 5.3 × 10−3 |
| A5A4Y9 | PPP1R11 | prot phsphtse 1 reg inhbt 11 | CDK5 signaling | −2.03 | 5.5 × 10−4 |
| Q6ZWR4 | PPP2R2B | prot phsphtse 2 reg subunit B, β | CDK5 signaling | 2.05 | 6.1 × 10−3 |
| P63085 | MAPK1 | ERK2 | CDK5 signaling | 1.61 | 3.6 × 10−2 |
| Q8BW96-2 | CAMK1D | Iso 2 of Ca2+/Calm-kinase type 1D | neuropath pain/CREB signaling | 8.35 | 1.9 × 10−2 |
| S4R1C4 | ATP2B2 | ATPase Ca2+ transport 2 | Ca2+ transport I | −2.20 | 5.9 × 10−3 |
| A2ALL9 | ATP2B3 | ATPase Ca2+ transport 3 | Ca2+ transport I | −1.45 | 4.5 × 10−9 |
| Q05915 | GCH1 | GTP cyclohydrolase 1 | 5-HT-R-signaling | −1.61 | 1.1 × 10−2 |
| Q3TYJ1 | SLC18A3 | solute carrier family 18 - A3 | 5-HT-R-signaling | −2.01 | 2.7 × 10−5 |
| Q80VQ0 | ALDH3B1 | aldehyde dehydrogenase 3 - B1 | DA, Trp degradation | −1.48 | 1.6 × 10−2 |
| Q8BW75 | MAOB | monoamine oxidase B | 5-HT-R-signaling, DA, Trp degradation | 1.63 | 7.1 × 10−3 |
| Q64133 | MAOA | monoamine oxidase A | 5-HT-R-signaling, DA, Trp degradation | 1.76 | 3.8 × 10−4 |
| Q80T41 | GABBR2 | GABA-B receptor subunit 2 | GABA-R-signaling | −1.75 | 5.7 × 10−3 |
| Q923×4-2 | GLRX2 | glutaredoxin 2 | mitochondrial dysfunction | −2.13 | 2.3 × 10−3 |
| P57716 | NCSTN | nicastrin | mitochondrial dysfunction | −1.46 | 1.2 × 10−2 |
| Q00519 | XDH | xanthine dehydrogenase | mitochondrial dysfunction | 1.88 | 4.5 × 10−4 |
| Q61387 | COX7A2L | cyt-C oxidase subunit 7A2 like | mitochondrial dysfunction | 2.4 × 101 | 1.0 × 10−9 |
| E9PZA8 | SYT7 | synaptotagmin 7 | SNARE complex | −1.82 | 3.7 × 10−2 |
| P60879-2 | SNAP25 | iso 2 of synaptosomal-assoc. prot 25 | SNARE complex | −1.45 | 2.8 × 10−3 |
| Q8BGN8-2 | SYNPR | synaptoporin | SNARE complex | −1.65 | 1.9 × 10−5 |
| Q9EQT6 | SYT13 | synaptotagmin 13 | SNARE complex | 3.55 | 1.5 × 10−2 |
| Q99JT1 | GATB | glutamyl-tRNA(Gln)amidotransf B, mito | l-glutamine biosynthesis | 1.42 | 4.8 × 10−7 |
| E9PXM6 | SLC29A1 | equilibrative nucleoside transporter 1 (ENT1) |
−2.04 × 1011 | – | |
| P16858 | GAPDH | glyceraldehyde-3-phospho dehydrogenase | 1.09 | 1.4 × 10−1 |
−, statistics not available due to undetectable expression in ENT1−/−- mice.
Network Interactome Analysis of Protein Profile
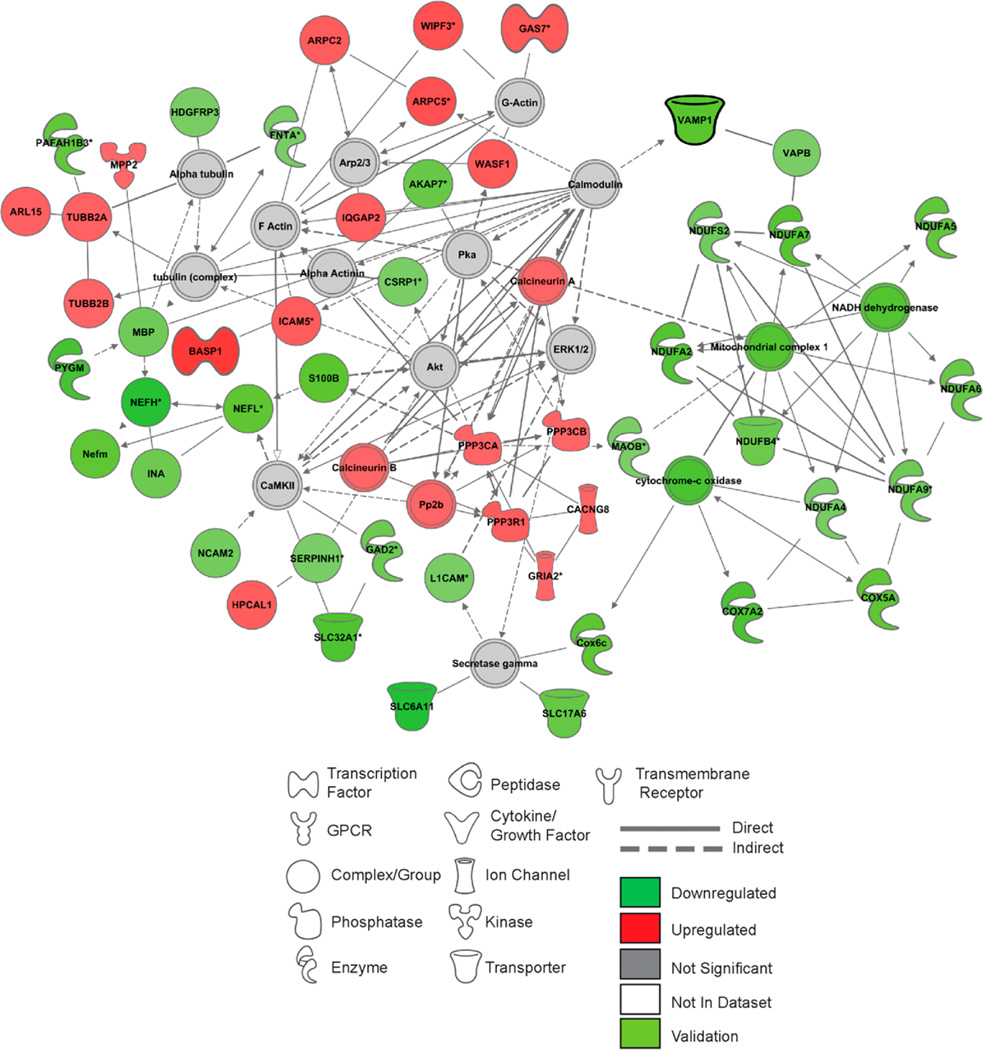
To determine how ENT1 deletion affected existing and inferred molecular interactions between focus proteins and reference proteins specific to the dHip, we combined the top two networks implicated by IPA to generate a more complete molecular model underlying the defects observed in these mice. Implicated pathways included significantly downregulated Ox-Phos complex proteins associated with deficits in Ox-Phos and mitochondrial dysfunction (Figure 4). We also detected an indirect molecular interaction between the SNARE proteins VAPB and VAMP1 and the calcium messenger protein calmodulin, providing a potential unification between down-regulated mitochondrial proteins and the upregulated ARP2/ 3–WASP complex proteins (ARPC2, ARPC5, WASF1) associated with increased cytoskeletal signaling (Figure 4).
Figure 4.
Differential expression of proteins in the dorsal hippocampus generates specific network interactomes in ENT1−/− mice. (A) Top protein network interactome generated from focus and reference proteins identified in the dHip of ENT1−/− relative to WT mice. (B) Top protein network interactome generated from focus and reference proteins identified in the NAc of ENT1−/− relative to ENT1+/+ mice. Red and green intensities indicate upregulated and downregulated protein expression, respectively. Gray indicates protein was detected but did not meet focus protein threshold criteria. Light green indicates proteins that were underwent validation analysis in this brain region. White proteins are imputed by Ingenuity Pathway Analysis to interact with proteins in network. Solid line and dashed line indicate a direct or indirect interaction, respectively. dHip, dorsal hippocampus; NAc, nucleus accumbens.
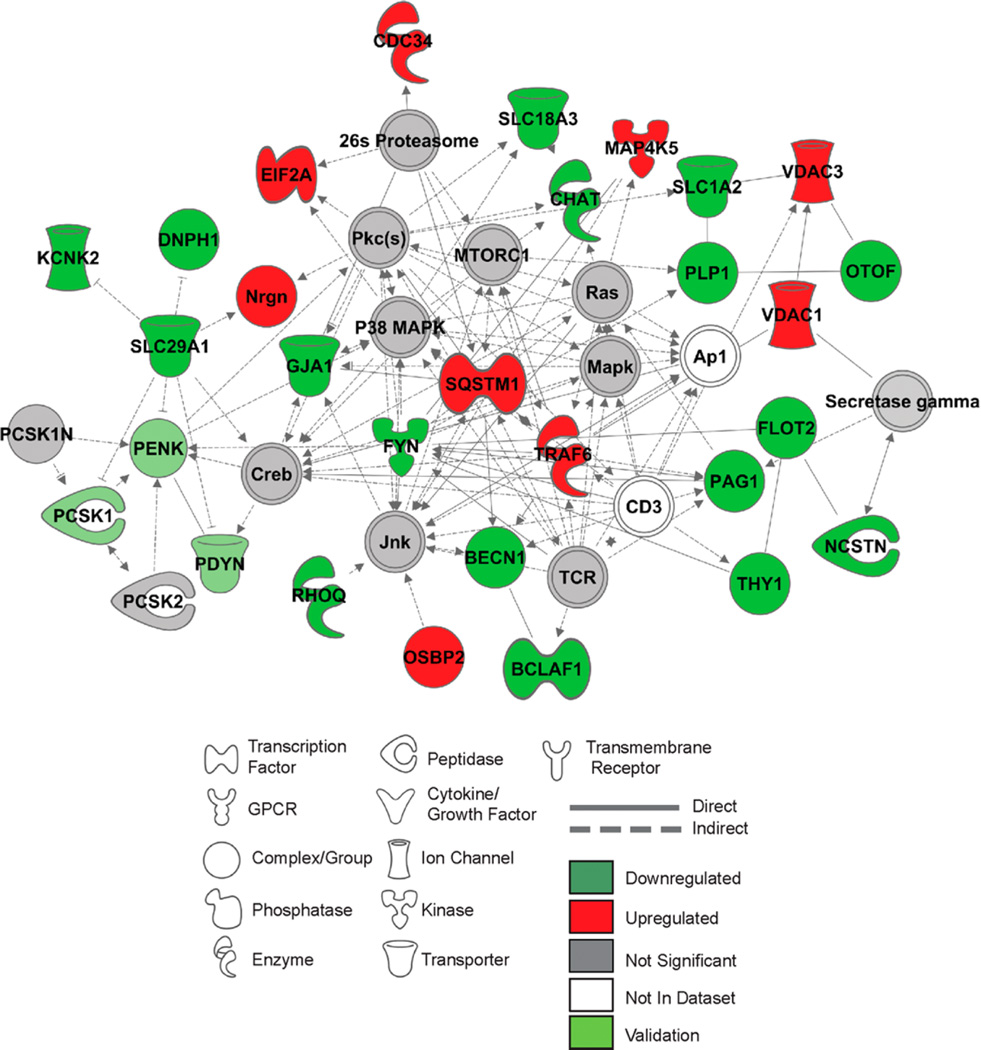
Advanced network analysis in the NAc of ENT1−/− mice revealed interactions between proteins involved in regulation of glutamate signaling, including a significant increase in neuro-granin (NGRN). Additionally, there was an interaction between SLC29A1 (ENT1), DNPH1, and TREK-1 (KCNK2), all of which were significantly downregulated (Figure 5). In addition we identified network interactions between ENT1 and CREB-mediated pro-dynorphin (PDYN) generation, including the neuro-endocrine convertases PCSK1, PCSK2, and an inhibitor of these convertases (PCSK1N), suggesting possible deficits in regulation of neuro-endocrine peptide processing as this cluster of proteins were decreased in expression (Figure 5 and Supplementary Table 2). Although decreased in expression but not meeting proteomic focus protein criteria, these results were nonetheless interesting given that we have previously reported that chronic ethanol exposure and withdrawal in mice can differentially affect PDYN expression in the striatum.25 Interestingly, this network identified a significant upregulation of proteins belonging to TNFα superfamily (TRAF6 and SQSTM1), which regulate activation of NFκB and T-cell activation (PAG1).
Figure 5.
Differential expression of proteins in the nucleus accumbens generates specific network interactomes in ENT1−/− mice. (A) Top protein network interactome generated from focus and reference proteins identified in the NAc of ENT1−/− relative to WT mice. (B) Top protein network interactome generated from focus and reference proteins identified in the NAc of ENT1−/− relative to ENT1+/+ mice. Red and green intensities indicate upregulated and downregulated protein expression, respectively. Gray indicates protein was detected but did not meet focus protein threshold criteria. Light green indicates proteins that were underwent validation analysis in this brain region. White proteins are imputed by Ingenuity Pathway Analysis to interact with proteins in network. Solid line and dashed line indicates a direct or indirect interaction, respectively. dHip, dorsal hippocampus; NAc, nucleus accumbens.
Protein Expression Validation of Hippocampal-Accumbal Neuroproteomics
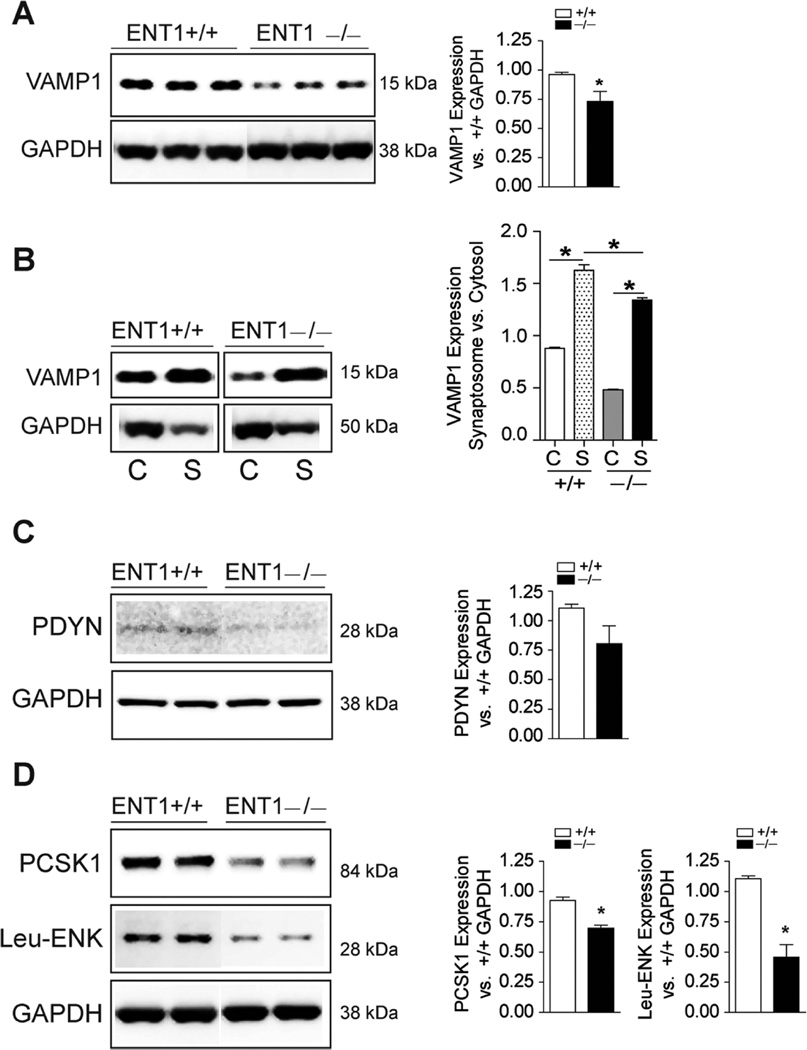
Although postsynaptic ENT1 signaling in the NAc has been shown to play a critical role in regulating Ca2+-calmodulin signaling, we sought to determine whether ENT1 deletion could also affect presynaptic SNARE VAMP1 protein expression in the dHip, given that we have previously reported a hyperglutamatergic state in ENT1−/− mice.12,27 As shown in Figure 6A and Supplementary Figure 2A,B, WB analysis detected a significant decrease in expression of VAMP1 in the cytosolic compartment of the dHip in ENT1−/− mice (t11 = 2.40, *p < 0.05). To determine whether this decrease in VAMP1 expression was evident in presynaptic vesicles, we examined VAMP1 enrichment in synaptosomes from the dHip of ENT1−/− and WT mice. As evidenced in Figure 6B and Supplementary Figure 2B, we detected enrichment of synaptosomal VAMP1 protein in both genotypes relative to their respective cytosolic fraction [F(3, 11) = 297.2, p < 0.0001]. However, multiple comparisons indicated a significant decrease (all *p < 0.001) in cytosolic and synaptosomal expression of VAMP1 in ENT1−/− mice relative to WT mice (Figure 6B).
Figure 6.
Validation of neuroproteomics from the dorsal hippocampus and nucleus accumbens of ENT1−/− mice in comparison with WT (ENT1+/+) mice. (A) Western blot and densitometry quantitation in the dHip of ENT1−/− mice shows significantly decreased expression of VAMP1 in the cytosolic compartment. n = 6–7/genotype, *p ≤ 0.05 by unpaired two-tailed t-test. (B) Synaptosomal fraction examination in the dHip shows decreased expression of VAMP1 in ENT1−/− mice. n = 3/genotype, *p ≤ 0.05 by Bonferroni’s multiple comparisons. (C) Western blot and densitometry quantitation in the NAc of ENT1−/− mice shows decreased expression of PDYN. n = 3-4/genotype. (D) Significantly decreased protein expression of PCSK1 and Leu-Enkephalin (Leu-ENK) in the NAc of ENT1−/− mice. n = 3-4/genotype, *p ≤ 0.01 by unpaired two-tailed t-test. Protein expression data is expressed as mean ± SEM. dHip, dorsal hippocampus; NAc, nucleus accumbens.
Our proteomic analysis in the NAc of ENT1−/− mice revealed a network interaction between CREB signaling and neuropeptide processing (PDYN–PCSK1–PCSK2–PCSK1N). This, in conjunction with the observed alterations of PDYN in the NAc as a result of ethanol withdrawal,25 prompted further investigation of expression of proteins in this pathway. As shown in Figure 6C and Supplementary Figure 3A, this analysis revealed a decrease in expression of PDYN in ENT1−/− mice relative to WT mice (t5 = 1.67, p = 0.15). Likewise, examination of the pro-hormone-converting enzyme PCSK1 (t5 = 6.28, *p < 0.01) and Leu-Enkephalin, the product of this enzymatic reaction (t5 = 5.22, *p < 0.01), was also appropriately decreased in ENT1−/− mice (Figure 6D and Supplementary Figure 3B). Taken together, validation of our proteomic analysis of the hippocampal-accumbal circuit in ENT1−/− mice suggests that deficits in adenosinergic regulation induced by ENT1 deletion have profound effects on key proteins involved in presynaptic vesicular neurotransmitter release as well as neuropeptide processing/signaling. These processes are essential regulators of learning, memory, and reward seeking behaviors.
Biological Significance and Disease State Analysis
Diseases and Disorders
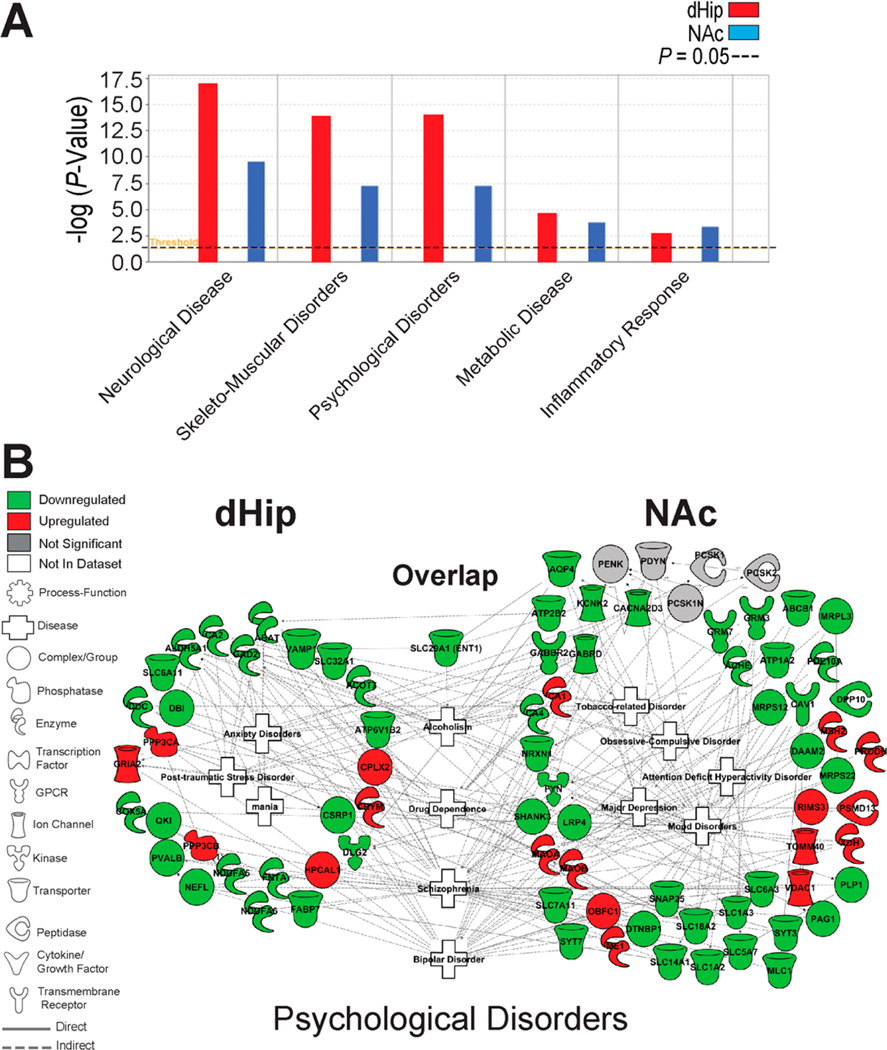
To explore the biological and pathophysiological significance of the pathway alterations induced by ENT1 deletion within the dHip-NAc circuit, we employed a comparative analysis of the two regions with IPA. Multiple pathologies were associated with differentially expressed proteins in the dHip and NAc of ENT1 mice, including neurological disease, skeletomuscular disorder, metabolic disorder, and inflammatory responses (all p < 0.05) (Figure 7A and Supplementary Table 3). Next, we examined the degree of overlap between differentially expressed proteins their associated biological functions to generate a connectivity network relating molecular alterations in the dHip-NAc circuit. This analysis revealed an overlap between significantly altered proteins in the dHip and NAc and alcoholism, drug dependence, bipolar disorder, and schizophrenia (Figure 7B). VAMP1 and PCSK1 associated with schizophrenia, while PDYN associated with major depression, bipolar disorder, and mood disorders (Figure 7B). Furthermore, our data set of differentially expressed proteins in the dHip and NAc also associated with neurological diseases including Alzheimer’s disease, movement disorders, cognitive impairment, dystonia, Parkinson’s disease, paralysis, and progressive motor neuropathy (Supplementary Figure S4 and Supplementary Table 3). Interestingly, VAMP1 formed a network connection with paralysis and dystonia (Supplementary Figure S4).
Figure 7.
Bioinformatic comparison analysis of the top diseases and disorders associated with the dorsal hippocampus and nucleus accumbens of ENT1−/− mice relative to WT mice. (A) Comparison analysis of the top diseases and disorders associated with focus proteins in the dHip and NAc of ENT1−/− mice. Left red bars, dHip; Right blue bars, NAc. (B) Representative diseases-disorders (center of connectome) indicate how differentially expressed proteins associate with psychological disorders. Red and green intensities indicate upregulated and downregulated protein expression, respectively. Gray indicates protein was detected but did not meet focus protein criteria. White functions are imputed by Ingenuity Pathway Analysis. Solid line and dashed line indicates a direct or indirect interaction, respectively. Functions or proteins below “overlap” indicate proteins or functions that were associated in both the dHip and NAc. dHip, dorsal hippocampus; NAc, nucleus accumbens.
Physiological and Developmental Functions
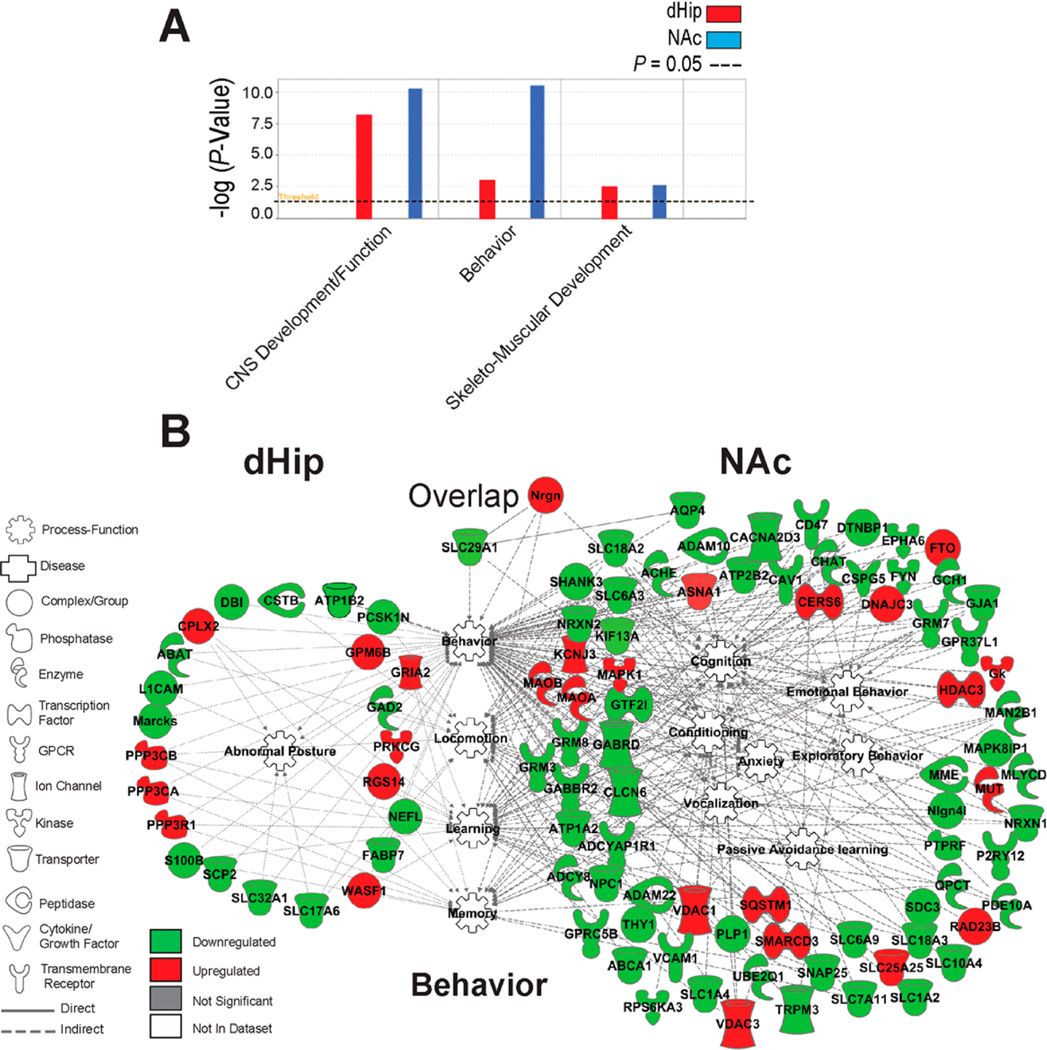
Next, we examined physiological and developmental processes affected by ENT1 deletion in the dHip and NAc. As shown in Figure 8A and Supplementary Table 5, we detected a significant number of differentially expressed proteins involved in central nervous system (CNS) development/function, behavior, and muscu-loskeletal system development/function in the dHip and NAc of ENT1−/− mice (all p < 0.05) (Supplementary Table 4 and Supplementary Figure S5). More specifically, significantly altered proteins in the dHip and NAc of ENT1 null mice were associated with behavioral development of anxiety, cognition, learning, memory, conditioning, passive avoidance learning, abnormal posture, and locomotion (Figure 8B). In particular, ENT1 (SLC29A1) associated with conditioning and behavior (Figure 8B). In summary, it is evident from our neuroproteomic results that ENT1 deletion has widespread effects on critical proteins in the dHip-NAc neural circuit regulating neuropsychiatric conditions, neurological disorders, and CNS physiological functions.
Figure 8.
Comparison bioinformatic analysis of the top physiological development-functions associated with the dorsal hippocampus and nucleus accumbens of ENT1−/− mice. (A) Comparison analysis of the top physiological-developmental functions associated with focus proteins in the dHip and NAc of ENT1−/− mice. Left red bars, dHip; Right blue bars, NAc. (B) Representative physiological-developmental functions (center of connectome) indicate how differentially expressed proteins associate with Behavior. Red and green intensities indicate upregulated and downregulated protein expression, respectively. Gray indicates protein was detected but did not meet focus protein criteria. White functions are imputed by Ingenuity Pathway Analysis. Solid line and dashed line indicate a direct or indirect interaction, respectively. Functions or proteins below “overlap” indicate proteins or functions that were associated in both the dHip and NAc. dHip, dorsal hippocampus; NAc, nucleus accumbens.
DISCUSSION
The present study executes a neuroproteome comparative analysis of the dHip and NAc in ENT1−/− mice, revealing vast changes in protein expression in both of these brain regions. More importantly, the results provided testable hypotheses elucidating how adenosinergic signaling in the dHip and NAc may affect learning and reward seeking behaviors.
Recent approaches in neuroproteomics have permitted a greater understanding of the underlying pathophysiology of complex neuropsychiatric disorders, which likely involve differential dysfunction in numerous brain regions.28 However, the immense amount of data generated during large-scale proteomic techniques makes these methodologies vulnerable to experimental and investigator bias. Therefore, individual biochemical validation of significant findings is prudent to ensure the accuracy of proteomic findings.29 Consequently, we utilized the proteomic library obtained during a previous iTRAQ proteome investigation of the NAc as a reference to confirm our current label-free findings, which identified EAAT2, neurogranin (NRGN), and PKCγ and GFAP, as differentially altered in ENT1−/− mice.27,30
Our IPA bioinformatic analysis detected a significant decrease in expression of VAMP1 in the dHip of ENT1−/− mice. This result was confirmed by WB protein expression analysis, demonstrating that VAMP1, a SNARE complex protein essential in vesicular presynaptic neurotransmitter release,31 was significantly decreased in ENT1−/− mice. Moreover, our analysis of the NAc also detected significant decreases in the SNARE complex proteins SNAP25 (isoform) and synaptotagmin, which have previously been demonstrated to be affected by ethanol.32,33 Consequently, synaptosomal fractionation revealed that VAMP1 seems decreased in presynaptic terminals, which is in agreement with our observed decreased expression within the cytosolic compartment. Given our results, we cannot conclusively determine whether the observed depletion of cytosolic and synaptosomal VAMP1 expression is occurring at glutamatergic or GABAergic neurons. However, physiological need for purinergic regulation of SNARE complex may be necessary, as large clusters of the adenosine triphosphate (ATP) P2×1 purinoceptor have been located opposite of SNARE complex proteins (including VAMP1) in abnormally dilated blood vessels.34
Our proteomic findings identified a decrease in expression of proteins critical for inhibitory regulation of presynaptic glutamate release (metabotropic glutamate receptors subtypes 3, 7, and 8) as well as for removal of synaptic glutamate (EAAT1 and EAAT2). Taken together, these results may explain the increased levels of synaptic glutamate previously observed as result of ENT1 deletion.12,27 The neuroproteomic profile of the NAc in our study identified decreases prepro hormone processing in ENT1−/− mice and we have previously reported differential expression of PDYN in the NAc.25 This prompted a closer investigation of PDYN neuropeptide processing, which results in the production of Leu-Enkephalin by the neuro-endocrine convertase PCSK1.35 WB analysis in the NAc of ENT1−/− mice revealed decreases in protein expression of PDYN, PCSK1, and Leu-Enkephalin, thus validating our neuroproteomic observations. Previously, we have reported decreases in CREB activity in the NAc of ENT1−/− mice.27 It is well known that dynorphin-enkephalin neuropeptide processing is regulated by CREB in response to alcohol and other drugs of abuse, in particular, drug-induced aversive effects.36 It is likely that deficits in dynorphin-enkephalin neuropeptide processing in the NAc of ENT1−/− mice may play a role in dampening any ethanol-induced aversive effects, thus partly explaining the ability for this mouse model to excessively consume ethanol. More importantly, these findings provide further evidence supporting a relationship between deficits in ENT1 and adenosinergic signaling via the A2A receptor, resulting in decreased enkephalin production in the striatum.37,38 Other studies further support the adenosine-enkephalin relationship as adenosine A1 and A2A knockout mouse models of Parkinson’s and Huntington’s disease result in decreased striatal expression of pro-enkephalin and enkephalin mRNA.39,40 Additional evidence strengthens the purinergic-enkephalin relationship in terms of nociception, given that Leu-Enkephalin inhibits P2X3-associated ion currents in rat dorsal root ganglion neurons.41 Thus, through our neuroproteomic investigation we are the first to report that alterations in adenosine signaling induced by deletion of ENT1 may mimic deficits in adenosine A2A receptor function and produce deficits in PDYN–PCSK1–enkephalin signaling.20 Notably, human patients42 and animal models exhibiting deficient PDYN activity and decreased enkephalin production demonstrate increased ethanol consumption43 and cocaine-seeking behav-ior.44
These results suggest that ENT1 may regulate ethanol consumption and goal-directed behaviors by modulating the release of excitatory or inhibitory neurotransmitters such as glutamate or GABA via VAMP1. However, a more mechanistic and biochemical approach is required to determine precisely how deficits in ENT1-mediated adenosine signaling affect vesicular neurotransmitter release and synaptic processing in the hippocampal-striatal circuit. Interestingly, recent reports examining ethanol’s effect on the cardio-vasculature revealed a direct ENT1–calmodulin interaction45,46 that is regulated by NMDA receptor function. Therefore, this confirms the importance of this transporter in cardio-protection, albeit at the expense of potential aberrations in CNS purinergic signaling, which may lead to addictive behaviors.
This extensive protein expression analysis provides unique insight into the signaling pathways, molecular functions, and physiology affected by altered ENT1 function and adenosine signaling. In accordance with the known ethanol drinking phenotype of ENT1−/− mice, we detected significant correlations between proteins in the dHip and NAc in relation to substance-use disorders, schizophrenia, and bipolar disorder. Our bioinformatic analysis identified proteins and pathways implicated in neurological movement disorders, paralysis, as well as Alzheimer’s disease, Parkinson’s disease, and Hunting-ton’s disease. Indeed ENT1 may affect these neurological conditions, which is particularly interesting given the newly established role of ENT1 and purinergic signaling in musculoskeletal function as well as the association of ENT1 dysfunction with neurological abnormalities.47–49
Our neuroproteomic analysis suggested enhancements in actin cytoskeletal reorganization resulting from upregulation of proteins involved in actin nucleation by the ARP2/3 complex in the dHip. In particular, these proteins include the Wiskott–Aldrich-syndrome family proteins (WASP) WASF1, WIPF1, and WASL. The WASP–ARP2/3 protein complex is regulated by the Rho-Rac GTPase, and Cdc42, which work in concert to activate actin polymerization and facilitate PKA substrate targeting.50–52 Recent studies have shown that deficits in protein expression of WASF1 lead to deficits in learning and memory as well as diminished depolarization-induced mito-chondrial movement into dendritic spines.53,54 Therefore, it is conceivable that increased expression of WASP–ARP2/3 complex proteins in our system promotes synaptic reorganization in response to augmented glutamatergic signaling, providing possible insights into the behavioral alterations observed in ENT1−/− mice. Alternatively, it is possible that the observed alterations of WASP–ARP2/3 expression are simply adaptive compensatory responses to the hyperglutama-tergic state observed in ENT1−/− mice. Our results are specially intriguing due to recent findings highlighting the importance of proteins involved in cytoskeletal reorganization and neuronal morphogenesis in neuropsychiatric diseases such as bipolar disorder, stress-induced anxiety, and addiction.8,55–58
An additional pathway that emerged from our neuro-proteomic analysis of the dHip and NAc was mitochondrial dysfunction. There is a growing body of literature that examines the effects of alcohol on the function of CNS mitochondria.59 In support, our neuroproteomic analysis of the dHip of ENT1−/− mice identified deficits in expression of proteins involved in bioenergetics and cellular energy homeostasis, including glycolysis and the TCA cycle. Properly functioning mitochondrial Ox-Phos and Ca2+ buffering against mitochon-drial depolarization, reactive oxygen species, and superoxide radical generation is a process that may be uniquely affected in ENT1−/− mice.60 Further investigation is necessary to determine the involvement of our identified proteins in regards to their contribution toward excitoxicity and abnormal synaptic plasticity associated with addiction and other neuropsychiatric disorders.
CONCLUSIONS
Our neuroproteomic and functional analysis suggests that the behavioral alterations resulting from ENT1 deletion may result in part from adenosine-mediated malfunctions in neuropeptide processing and control of neurotransmitter release. Taken together, our neuroproteomic findings indicate that disrupted adenosine-mediated neurotransmission may promote the aberrant alcohol seeking behavior displayed by ENT1−/− mice. Given that the neural circuit involving the dHip and NAc may be instrumental for the anticipation and perception of reward,7,61–63 our results provide insight into several novel targets, which may modulate adenosine signaling and thereby aid in the treatment of AUD and other mood disorders.
Supplementary Material
Acknowledgments
This work was supported by the Samuel C. Johnson for Genomics of Addiction Program at Mayo Clinic, the Ulm Foundation, the Godby Foundation, David Lehr Research Award from American Society for Pharmacology and Experimental Therapeutics, Mayo Graduate School, and the National Institute on Alcohol Abuse and Alcoholism (AA018779).
ABBREVIATIONS
- dHip
dorsal hippocampus
- NAc
nucleus accumbens
Footnotes
ASSOCIATED CONTENT
Supporting Information
Supplementary Figure S1. SDS-PAGE analysis in the dorsal hippocampus and nucleus accumbens shows gels that were fixed and stained with Coomasie G-250 prior to label-free proteomic analysis. Supplementary Figure S2. Western blot validation of neuroproteomics from the dorsal hippocampus shows complete blots of cytosolic VAMP1 expression and synaptosomal VAMP1 expression in WT mice relative to ENT1−/− mice. Supplementary Figure S3. Western blot validation of neuro-proteomics from nucleus accumbens shows complete blots of PDYN, PCSK1, and Leu-Enkephalin (Leu-ENK) expression of WT mice relative to ENT1−/− mice. Supplementary Figure S4. Comparison bioinformatic analysis of the neurological disease function associated with the dorsal hippocampus (dHip) and nucleus accumbens (NAc) of ENT1−/− mice relative to WT mice. Supplementary Figure S5. Comparison bioinformatic analysis of skeleto-muscular development associated with the dorsal hippocampus (dHip) and nucleus accumbens (NAc) of ENT1−/− mice relative to WT mice. Supplementary Table S3. Ingenuity Pathway Analysis identified diseases and disorder functions altered due to ENT1 deletion. Supplementary Table S4. Ingenuity Pathway Analysis identified top physiological-developmental functions altered due to ENT1 deletion. Supporting Information and methods. Detailed materials and methods. (PDF)Supplementary Table S1. Group expression data of proteins identified in the dorsal hippocampus (dHip) of ENT1−/− mice relative to WT mice. (XLSX)Supplementary Table S2. Group expression data of proteins identified in the nucleus accumbens (NAc) of ENT1−/− mice relative to WT mice. (XLSX)
The authors declare no competing financial interest.
REFERENCES
- 1.Everitt BJ, Robbins TW. Neural systems of reinforcement for drug addiction: from actions to habits to compulsion. Nat. Neurosci. 2005;8:1481–1489. doi: 10.1038/nn1579. [DOI] [PubMed] [Google Scholar]
- 2.Pennartz CM, Ito R, Verschure PF, Battaglia FP, Robbins TW. The hippocampal-striatal axis in learning, prediction and goal-directed behavior. Trends Neurosci. 2011;34:548–559. doi: 10.1016/j.tins.2011.08.001. [DOI] [PubMed] [Google Scholar]
- 3.Dayas CV, Liu X, Simms JA, Weiss F. Distinct patterns of neural activation associated with ethanol seeking: effects of naltrexone. Biol. Psychiatry. 2007;61:979–989. doi: 10.1016/j.biopsych.2006.07.034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Marinelli PW, Funk D, Juzytsch W, Li Z, Le AD. Effects of opioid receptor blockade on the renewal of alcohol seeking induced by context: relationship to c-fos mRNA expression. Eur. J. Neurosci. 2007;26:2815–2823. doi: 10.1111/j.1460-9568.2007.05898.x. [DOI] [PubMed] [Google Scholar]
- 5.Ito R, Hayen A. Opposing roles of nucleus accumbens core and shell dopamine in the modulation of limbic information processing. J. Neurosci. 2011;31:6001–6007. doi: 10.1523/JNEUROSCI.6588-10.2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Ito R, Robbins TW, Pennartz CM, Everitt BJ. Functional interaction between the hippocampus and nucleus accumbens shell is necessary for the acquisition of appetitive spatial context conditioning. J. Neurosci. 2008;28:6950–6959. doi: 10.1523/JNEUROSCI.1615-08.2008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Luo AH, Tahsili-Fahadan P, Wise RA, Lupica CR, Aston-Jones G. Linking context with reward: a functional circuit from hippocampal CA3 to ventral tegmental area. Science. 2011;333:353–357. doi: 10.1126/science.1204622. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Barak S, Liu F, Ben Hamida S, Yowell QV, Neasta J, Kharazia V, Janak PH, Ron D. Disruption of alcohol-related memories by mTORC1 inhibition prevents relapse. Nat. Neurosci. 2013;16:1111–1117. doi: 10.1038/nn.3439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Di Ciano P, Everitt BJ. Neuropsychopharmacology of drug seeking: Insights from studies with second-order schedules of drug reinforcement. Eur. J. Pharmacol. 2005;526:186–198. doi: 10.1016/j.ejphar.2005.09.024. [DOI] [PubMed] [Google Scholar]
- 10.Bannerman DM, Rawlins JN, McHugh SB, Deacon RM, Yee BK, Bast T, Zhang WN, Pothuizen HH, Feldon J. Regional dissociations within the hippocampus-memory and anxiety. Neurosci. Biobehav. Rev. 2004;28:273–283. doi: 10.1016/j.neubiorev.2004.03.004. [DOI] [PubMed] [Google Scholar]
- 11.Britt JP, Benaliouad F, McDevitt RA, Stuber GD, Wise RA, Bonci A. Synaptic and behavioral profile of multiple glutamatergic inputs to the nucleus accumbens. Neuron. 2012;76:790–803. doi: 10.1016/j.neuron.2012.09.040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Chen J, Nam HW, Lee MR, Hinton DJ, Choi S, Kim T, Kawamura T, Janak PH, Choi DS. Altered glutamatergic neurotransmission in the striatum regulates ethanol sensitivity and intake in mice lacking ENT1. Behav. Brain Res. 2010;208:636–642. doi: 10.1016/j.bbr.2010.01.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Ferre S, Fredholm BB, Morelli M, Popoli P, Fuxe K. Adenosine-dopamine receptor-receptor interactions as an integrative mechanism in the basal ganglia. Trends Neurosci. 1997;20:482–487. doi: 10.1016/s0166-2236(97)01096-5. [DOI] [PubMed] [Google Scholar]
- 14.Goto Y, Grace AA. Dopamine modulation of hippocampal-prefrontal cortical interaction drives memory-guided behavior. Cereb Cortex. 2008;18:1407–1414. doi: 10.1093/cercor/bhm172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Asatryan L, Nam HW, Lee MR, Thakkar MM, Saeed Dar M, Davies DL, Choi DS. Implication of the purinergic system in alcohol use disorders. Alcohol.: Clin. Exp. Res. 2011;35:584–594. doi: 10.1111/j.1530-0277.2010.01379.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Choi DS, Cascini MG, Mailliard W, Young H, Paredes P, McMahon T, Diamond I, Bonci A, Messing RO. The type 1 equilibrative nucleoside transporter regulates ethanol intoxication and preference. Nat. Neurosci. 2004;7:855–861. doi: 10.1038/nn1288. [DOI] [PubMed] [Google Scholar]
- 17.Gorini G, Adron Harris R, Dayne Mayfield R. Proteomic approaches and identification of novel therapeutic targets for alcoholism. Neuropsychopharmacology. 2014;39:104–130. doi: 10.1038/npp.2013.182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Sari Y, Zhang M, Mechref Y. Differential expression of proteins in fetal brains of alcohol-treated prenatally C57BL/6 mice: a proteomic investigation. Electrophoresis. 2010;31:483–496. doi: 10.1002/elps.200900385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Domon B, Aebersold R. Options and considerations when selecting a quantitative proteomics strategy. Nat. Biotechnol. 2010;28:710–721. doi: 10.1038/nbt.1661. [DOI] [PubMed] [Google Scholar]
- 20.Nam HW, Hinton DJ, Kang NY, Kim T, Lee MR, Oliveros A, Adams C, Ruby CL, Choi DS. Adenosine transporter ENT1 regulates the acquisition of goal-directed behavior and ethanol drinking through A2A receptor in the dorsomedial striatum. J. Neurosci. 2013;33:4329–4338. doi: 10.1523/JNEUROSCI.3094-12.2013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Franklin KBJ, Paxinos G. The Mouse Brain in Stereotaxic Coordinates. 3rd. San Diego, CA: Elsevier Academic; 2007. [Google Scholar]
- 22.Olsen JV, de Godoy LM, Li G, Macek B, Mortensen P, Pesch R, Makarov A, Lange O, Horning S, Mann M. Parts per million mass accuracy on an Orbitrap mass spectrometer via lock mass injection into a C-trap. Mol. Cell. Proteomics. 2005;4:2010–2021. doi: 10.1074/mcp.T500030-MCP200. [DOI] [PubMed] [Google Scholar]
- 23.Neubert H, Bonnert TP, Rumpel K, Hunt BT, Henle ES, James IT. Label-free detection of differential protein expression by LC/MALDI mass spectrometry. J. Proteome Res. 2008;7:2270–2279. doi: 10.1021/pr700705u. [DOI] [PubMed] [Google Scholar]
- 24.Wiener MC, Sachs JR, Deyanova EG, Yates NA. Differential mass spectrometry: a label-free LC-MS method for finding significant differences in complex peptide and protein mixtures. Anal. Chem. 2004;76:6085–6096. doi: 10.1021/ac0493875. [DOI] [PubMed] [Google Scholar]
- 25.Ayers-Ringler JR, Oliveros A, Qiu Y, Lindberg DM, Hinton DJ, Moore RM, Dasari S, Choi DS. Label-Free Proteomic Analysis of Protein Changes in the Striatum during Chronic Ethanol Use and Early Withdrawal. Front. Behav. Neurosci. 2016;10:46. doi: 10.3389/fnbeh.2016.00046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Cox J, Hein MY, Luber CA, Paron I, Nagaraj N, Mann M. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Mol. Cell. Proteomics. 2014;13:2513–2526. doi: 10.1074/mcp.M113.031591. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Nam HW, Lee MR, Zhu Y, Wu J, Hinton DJ, Choi S, Kim T, Hammack N, Yin JC, Choi DS. Type 1 equilibrative nucleoside transporter regulates ethanol drinking through accumbal N-methyl-D-aspartate receptor signaling. Biol. Psychiatry. 2011;69:1043–1051. doi: 10.1016/j.biopsych.2011.02.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Craft GE, Chen A, Nairn AC. Recent advances in quantitative neuroproteomics. Methods. 2013;61:186–218. doi: 10.1016/j.ymeth.2013.04.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Zhu W, Smith JW, Huang CM. Mass spectrometry-based label-free quantitative proteomics. J. Biomed. Biotechnol. 2010;2010:840518. doi: 10.1155/2010/840518. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Hinton DJ, Lee MR, Jang JS, Choi DS. Type 1 equilibrative nucleoside transporter regulates astrocyte-specific glial fibrillary acidic protein expression in the striatum. Brain Behav. 2014;4:903–914. doi: 10.1002/brb3.283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Zimmermann J, Trimbuch T, Rosenmund C. Synaptobrevin 1 mediates vesicle priming and evoked release in a subpopulation of hippocampal neurons. J. Neurophysiol. 2014;112:1559–1565. doi: 10.1152/jn.00340.2014. [DOI] [PubMed] [Google Scholar]
- 32.Pignataro L, Miller AN, Ma L, Midha S, Protiva P, Herrera DG, Harrison NL. Alcohol regulates gene expression in neurons via activation of heat shock factor 1. J. Neurosci. 2007;27:12957–12966. doi: 10.1523/JNEUROSCI.4142-07.2007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Varodayan FP, Pignataro L, Harrison NL. Alcohol induces synaptotagmin 1 expression in neurons via activation of heat shock factor 1. Neuroscience. 2011;193:63–71. doi: 10.1016/j.neuroscience.2011.07.035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Barden JA, Cottee LJ, Bennett MR. Vesicle-associated proteins and P2X receptor clusters at single sympathetic varicosities in mouse vas deferens. J. Neurocytol. 1999;28:469–480. doi: 10.1023/a:1007053004771. [DOI] [PubMed] [Google Scholar]
- 35.Seidah NG. What lies ahead for the proprotein convertases? Ann. N. Y. Acad. Sci. 2011;1220:149–161. doi: 10.1111/j.1749-6632.2010.05883.x. [DOI] [PubMed] [Google Scholar]
- 36.Muschamp JW, Carlezon WA., Jr Roles of nucleus accumbens CREB and dynorphin in dysregulation of motivation. Cold Spring Harbor Perspect. Med. 2013;3:a012005. doi: 10.1101/cshperspect.a012005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Tanimura Y, Vaziri S, Lewis MH. Indirect basal ganglia pathway mediation of repetitive behavior: attenuation by adenosine receptor agonists. Behav. Brain Res. 2010;210:116–122. doi: 10.1016/j.bbr.2010.02.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Wardas J, Pietraszek M, Dziedzicka-Wasylewska M. SCH 58261, a selective adenosine A2A receptor antagonist, decreases the haloperidol-enhanced proenkephalin mRNA expression in the rat striatum. Brain Res. 2003;977:270–277. doi: 10.1016/s0006-8993(03)02759-8. [DOI] [PubMed] [Google Scholar]
- 39.Dassesse D, Massie A, Ferrari R, Ledent C, Parmentier M, Arckens L, Zoli M, Schiffmann SN. Functional striatal hypodopaminergic activity in mice lacking adenosine A(2A) receptors. J. Neurochem. 2001;78:183–198. doi: 10.1046/j.1471-4159.2001.00389.x. [DOI] [PubMed] [Google Scholar]
- 40.Xiao D, Cassin JJ, Healy B, Burdett TC, Chen JF, Fredholm BB, Schwarzschild MA. Deletion of adenosine A(1) or A((2)A) receptors reduces L-3,4-dihydroxyphenylalanine-induced dyskinesia in a model of Parkinson’s disease. Brain Res. 2011;1367:310–318. doi: 10.1016/j.brainres.2010.08.099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Chizhmakov I, Kulyk V, Khasabova I, Khasabov S, Simone D, Bakalkin G, Gordienko D, Verkhratsky A, Krishtal O. Molecular mechanism for opioid dichotomy: bidirectional effect of mu-opioid receptors on P2X(3) receptor currents in rat sensory neurones. Purinergic Signalling. 2015;11:171–181. doi: 10.1007/s11302-015-9443-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Sarkisyan D, Hussain MZ, Watanabe H, Kononenko O, Bazov I, Zhou X, Yamskova O, Krishtal O, Karpyak VM, Yakovleva T, Bakalkin G. Downregulation of the endogenous opioid peptides in the dorsal striatum of human alcoholics. Front. Cell. Neurosci. 2015;9:187. doi: 10.3389/fncel.2015.00187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Femenia T, Manzanares J. Increased ethanol intake in prodynorphin knockout mice is associated to changes in opioid receptor function and dopamine transmission. Addict. Biol. 2012;17:322–337. doi: 10.1111/j.1369-1600.2011.00378.x. [DOI] [PubMed] [Google Scholar]
- 44.Gutierrez-Cuesta J, Burokas A, Mancino S, Kummer S, Martin-Garcia E, Maldonado R. Effects of genetic deletion of endogenous opioid system components on the reinstatement of cocaine-seeking behavior in mice. Neuropsychopharmacology. 2014;39:2974–2988. doi: 10.1038/npp.2014.149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Bicket A, Mehrabi P, Naydenova Z, Wong V, Donaldson L, Stagljar I, Coe IR. Novel regulation of equlibrative nucleoside transporter 1 (ENT1) by receptor-stimulated Ca2+-dependent calmodulin binding. Am. J. Physiol. Cell Physiol. 2016;310:C808–C820. doi: 10.1152/ajpcell.00243.2015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Ramadan A, Naydenova Z, Stevanovic K, Rose JB, Coe IR. The adenosine transporter, ENT1, in cardiomyocytes is sensitive to inhibition by ethanol in a kinase-dependent manner: implications for ethanol-dependent cardioprotection and nucleoside analog drug cytotoxicity. Purinergic Signalling. 2014;10:305–312. doi: 10.1007/s11302-013-9391-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Daniels G, Ballif BA, Helias V, Saison C, Grimsley S, Mannessier L, Hustinx H, Lee E, Cartron JP, Peyrard T, Arnaud L. Lack of the nucleoside transporter ENT1 results in the Augustine-null blood type and ectopic mineralization. Blood. 2015;125:3651–3654. doi: 10.1182/blood-2015-03-631598. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Hinton DJ, McGee-Lawrence ME, Lee MR, Kwong HK, Westendorf JJ, Choi DS. Aberrant bone density in aging mice lacking the adenosine transporter ENT1. PLoS One. 2014;9:e88818. doi: 10.1371/journal.pone.0088818. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Warraich S, Bone DB, Quinonez D, Ii H, Choi DS, Holdsworth DW, Drangova M, Dixon SJ, Seguin CA, Hammond JR. Loss of equilibrative nucleoside transporter 1 in mice leads to progressive ectopic mineralization of spinal tissues resembling diffuse idiopathic skeletal hyperostosis in humans. J. Bone Miner. Res. 2013;28:1135–1149. doi: 10.1002/jbmr.1826. [DOI] [PubMed] [Google Scholar]
- 50.Miki H, Yamaguchi H, Suetsugu S, Takenawa T. IRSp53 is an essential intermediate between Rac and WAVE in the regulation of membrane ruffling. Nature. 2000;408:732–735. doi: 10.1038/35047107. [DOI] [PubMed] [Google Scholar]
- 51.Rohatgi R, Ma L, Miki H, Lopez M, Kirchhausen T, Takenawa T, Kirschner MW. The interaction between N-WASP and the Arp2/3 complex links Cdc42-dependent signals to actin assembly. Cell. 1999;97:221–231. doi: 10.1016/s0092-8674(00)80732-1. [DOI] [PubMed] [Google Scholar]
- 52.Westphal RS, Soderling SH, Alto NM, Langeberg LK, Scott JD. Scar/WAVE-1, a Wiskott-Aldrich syndrome protein, assembles an actin-associated multi-kinase scaffold. EMBO J. 2000;19:4589–4600. doi: 10.1093/emboj/19.17.4589. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Soderling SH, Langeberg LK, Soderling JA, Davee SM, Simerly R, Raber J, Scott JD. Loss of WAVE-1 causes sensorimotor retardation and reduced learning and memory in mice. Proc. Natl. Acad. Sci. U. S. A. 2003;100:1723–1728. doi: 10.1073/pnas.0438033100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Sung JY, Engmann O, Teylan MA, Nairn AC, Greengard P, Kim Y. WAVE1 controls neuronal activity-induced mitochondrial distribution in dendritic spines. Proc. Natl. Acad. Sci. U. S. A. 2008;105:3112–3116. doi: 10.1073/pnas.0712180105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Dietz DM, Sun H, Lobo MK, Cahill ME, Chadwick B, Gao V, Koo JW, Mazei-Robison MS, Dias C, Maze I, Damez-Werno D, Dietz KC, Scobie KN, Ferguson D, Christoffel D, Ohnishi Y, Hodes GE, Zheng Y, Neve RL, Hahn KM, Russo SJ, Nestler EJ. Rac1 is essential in cocaine-induced structural plasticity of nucleus accumbens neurons. Nat. Neurosci. 2012;15:891–896. doi: 10.1038/nn.3094. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Fumagalli F, Di Pasquale L, Caffino L, Racagni G, Riva MA. Stress and cocaine interact to modulate basic fibroblast growth factor (FGF-2) expression in rat brain. Psychopharmacology (Berl) 2008;196:357–364. doi: 10.1007/s00213-007-0966-x. [DOI] [PubMed] [Google Scholar]
- 57.Golden SA, Christoffel DJ, Heshmati M, Hodes GE, Magida J, Davis K, Cahill ME, Dias C, Ribeiro E, Ables JL, Kennedy PJ, Robison AJ, Gonzalez-Maeso J, Neve RL, Turecki G, Ghose S, Tamminga CA, Russo SJ. Epigenetic regulation of RAC1 induces synaptic remodeling in stress disorders and depression. Nat. Med. 2013;19:337–344. doi: 10.1038/nm.3090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Klengel T, Mehta D, Anacker C, Rex-Haffner M, Pruessner JC, Pariante CM, Pace TW, Mercer KB, Mayberg HS, Bradley B, Nemeroff CB, Holsboer F, Heim CM, Ressler KJ, Rein T, Binder EB. Allele-specific FKBP5 demethylation mediates gene-childhood trauma interactions. Nat. Neurosci. 2012;16:33–41. doi: 10.1038/nn.3275. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Manzo-Avalos S, Saavedra-Molina A. Cellular and mitochon-drial effects of alcohol consumption. Int. J. Environ. Res. Public Health. 2010;7:4281–4304. doi: 10.3390/ijerph7124281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Bone DB, Antic M, Quinonez D, Hammond JR. Hypoxanthine uptake by skeletal muscle microvascular endothelial cells from equilibrative nucleoside transporter 1 (ENT1)-null mice: effect of oxidative stress. Microvasc. Res. 2015;98:16–22. doi: 10.1016/j.mvr.2014.11.005. [DOI] [PubMed] [Google Scholar]
- 61.Floresco SB, Blaha CD, Yang CR, Phillips AG. Modulation of hippocampal and amygdalar-evoked activity of nucleus accumbens neurons by dopamine: cellular mechanisms of input selection. J. Neurosci. 2001;21:2851–2860. doi: 10.1523/JNEUROSCI.21-08-02851.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Floresco SB, Todd CL, Grace AA. Glutamatergic afferents from the hippocampus to the nucleus accumbens regulate activity of ventral tegmental area dopamine neurons. J. Neurosci. 2001;21:4915–4922. doi: 10.1523/JNEUROSCI.21-13-04915.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Legault M, Rompre PP, Wise RA. Chemical stimulation of the ventral hippocampus elevates nucleus accumbens dopamine by activating dopaminergic neurons of the ventral tegmental area. J. Neurosci. 2000;20:1635–1642. doi: 10.1523/JNEUROSCI.20-04-01635.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.