Abstract
Short interfering RNAs (siRNAs) directed against poliovirus and other viruses effectively inhibit viral replication. Although RNA interference (RNAi) may provide the basis for specific antiviral therapies, the limitations of RNAi antiviral strategies are ill defined. Here, we show that poliovirus readily escapes highly effective siRNAs through unique point mutations within the targeted regions. Competitive analysis of the escape mutants provides insights into the basis of siRNA recognition. The RNAi machinery can tolerate mismatches but is exquisitely sensitive to mutations within the central region and the 3′ end of the target sequence. Indeed, specific mutations in the target sequence resulting in G:U mismatches are sufficient for the virus to escape siRNA inhibition. However, using a pool of siRNAs to simultaneously target multiple sites in the viral genome prevents the emergence of resistant viruses. Our study uncovers the elegant precision of target recognition by the RNAi machinery and provides the basis for the development of effective RNAi-based therapies that prevent viral escape.
RNA interference (RNAi) is a eukaryotic mechanism of gene inactivation which employs double-stranded RNA (dsRNA) to produce short interfering RNAs (siRNAs) of about 21 nucleotides in length (13). A cytoplasmic RNA-induced silencing complex (RISC) binds the siRNAs and uses them as guides to direct the degradation of mRNAs containing sequences complementary to one of the strands of the siRNA (27). The efficient and sequence-specific nature of RNAi has raised the possibility of using RNAi as an antiviral therapy. Indeed, introduction of siRNAs into cells in tissue culture and in animal models can inhibit replication of several human pathogenic viruses (4, 8, 11, 12, 18, 22, 23, 28, 29, 31, 34, 37). A major problem of all antiviral therapies, however, is the emergence of resistant variants. Here, we investigate how siRNA-based antiviral strategies might be neutralized by the emergence of escape mutants. The finding that in some cases RNAi can tolerate several mismatches within the target sequence (1, 6, 7, 21, 36) suggested that RNAi might be robust enough to accommodate certain variability of the virus target RNA. But it is also possible that these mutations do not disrupt positions important for the interaction of the guide siRNA with the target RNA. Indeed, recent observations indicate that both poliovirus and human immunodeficiency virus type 1 variants resistant to RNAi can be selected carrying single point mutations in the center of the target sequence (5, 12). The issue of target recognition by the RNAi machinery has been the subject of numerous studies. In both Drosophila melanogaster and mammalian extracts, mismatches within the central position (nucleotides 9 to 11) of the target RNA lead to inefficient silencing (10, 27). In contrast, peripheral mismatches appear to have no detrimental effect. However, others have found that correct base pairing at the central position is not critical for silencing. Instead, mismatches within the 5′ half of the antisense siRNA strand disrupted effective silencing, while mismatches at the central region (positions 9 and 10) and within the 3′ half of the siRNA did not affect silencing efficiency (1, 7). Studies of gene expression profiling in siRNA-treated cells have produced additional information with respect to target sequence recognition. While some studies have found little “off-target” silencing of nearly complementary targets (6, 36), other studies reported that partial complementarity between the 5′ half of the antisense siRNA strand with a number of endogenous mRNAs leads to effective nonspecific silencing (21). Thus, the precise rules on target recognition are still controversial and poorly defined. To further examine the ability of viruses to escape RNAi inhibition, we recovered and sequenced polioviruses that escaped inhibition by two highly effective siRNAs directed against either capsid or the viral polymerase coding sequences. Analysis of these viral escape mutants shows that RNAi recognition is sensitive to subtle point mutations within the central region and the 3′ end of the target RNA. Even single transition mutations resulting in G:U mismatches are sufficient to overcome viral inhibition from defined siRNAs. However, simultaneous targeting of multiple viral sequences with a pool of siRNAs overcame resistance mechanism to RNAi and prevented measurable viral escape.
MATERIALS AND METHODS
Cells and viruses.
HeLa S3 cells were cultured as previously described (12). P19 mouse embryonic carcinoma was obtained from the University of California—San Francisco cell culture facility and maintained in minimal essential medium, alpha modification, supplemented with nucleosides (Invitrogen), 10% fetal bovine serum, 2 mM glutamine, 100 U of penicillin/ml, and 100 μg of streptomycin/ml (University of California—San Francisco cell culture facility). We used the pRib(+)XpA plasmid (17) to generate the wild-type Mahoney type 1 strain of poliovirus. All mutations were cloned back into a pRib(+)XpA backbone. For viral production, 10 μg of in vitro-transcribed viral RNA was electroporated into HeLa cells (17), and cells were left overnight until complete lysis. The plasmid encoding the Leon type 3 poliovirus strain was a kind gift of D. J. Evans; the virus was produced similarly to type 1 Mahoney. The titers of the virus were determined according to standard procedures (9). For competition assays, viruses were mixed in equal proportions unless indicated otherwise and propagated for two passages (a passage concluded with full lysis) on cells transfected with siRNAs. Transfections were carried out as previously described (12). Plaque reduction assays were carried out as previously described (12), but cells were allowed to grow for 1 day between siRNA removal and infection. P19 transfections were carried out by combining 0.5 μg of pPVR plasmid with 0.3 μl of dsRNA (1 μg/μl) or 0.3 μl of 40 μM siRNA and transfecting the mixture with 1 μl of Lipofectamine 2000 per ∼1 × 105 cells in 1 well of a 24-well plate overnight in the presence of 10% serum.
Molecular biology.
RNA oligonucleotides were bought from Dharmacon and were treated as before (12). The sequence of control siRNA (siCtrl) strands is as follows: 5′-AAUACCAGAACACCAACUGGC-3′ and 5′-CAGUUGGUGUUCUGGUAUUAC-3′. The siC region of the viral genome was amplified by isolating cytoplasmic RNA from HeLa cells at 6 h postinfection with the RNeasy kit (QIAGEN), reverse transcription with random hexamers and SuperScriptII enzyme (Invitrogen), and PCR with PfuTurbo (Strategene) and primers 5′-GCTAGACACCGTGTCTTGGA-3′ and 5′-GGACTGTGTTGTCAATCATGCT-3′. PCR products were sequenced with the primer 5′-AGATGATAGTTTCACCGAAGG-3′. For cloning of individual mutants, each PCR fragment was digested with NruI and NheI and swapped for the wild-type fragment in pRib(+)XpA. The analogous procedure for isolation of the siP region used the primers 5′-GGTGAAATCCAGTGGATGAGA-3′ and 5′-GCGAACGTGATCCTGAGTGTT-3′ for amplification, 5′-AGGAAGCAATTACATCATCACC-3′ for sequencing, and AvrII and XbaI enzyme sites for cloning the fragment into a shuttle vector. For producing point mutants in the siP region, the following primer pair was used in a QuickChange protocol (Stratagene): 5′-CAGCAGTGGGGTGCGATCCAGATTTGTTTTGGAGCAAAATTC-3′ and its reverse complement, with corresponding mutations introduced in the primers. PCRs were run from a plasmid containing the BglII-EcoRI fragment of pRib(+)XpA, and the mutant BglII-EcoRI fragments were moved back into pRib(+)XpA.
Generation of the let-7(+) virus was carried out by using 5′-TGAGGTAGTAGGTTGTATAGTTACAATTTCAACAGTTATTTCAATCAGAC-3′ and 5′-CCTCAGTGCATCAGGCAACT-3′ primers in one PCR and 5′-AACTATACAACCTACTACCTCACTTAGAGTAAACACACTCAATGG-3′ and 5′-AGAAGCCCAGTACCACCTCG-3′ primers in a parallel PCR, both of which were consequently purified, mixed, and extended again for 15 cycles with Pfu polymerase. The product was digested with BlpI and AatII and cloned into the poliovirus infectious cDNA plasmid, thus introducing the sequence identical to let-7a (UGAGGUAGUAGGUUGUAUAGUU) (12) or its complementary sequence.
The 1-kb-long regions of Mahoney and firefly luciferase were amplified with primer pairs 5′-ATGATGGAATTGGCAGAAATC-3′ and 5′-TAAGGCATGCCCATTGTTAGT-3′ and 5′-AGAACTGCCTGCGTGAGATT-3′ and 5′-TTTCTTGCGTCGAGTTTTCC-3′; one of the primers in each reaction mixture contained a T7 transcription start site. Each strand was then transcribed with T7 RNA polymerase (NEB), and the RNA was annealed and treated for 2 h at 37°C with RNase III (a gift of Dun Yang and J. M. Bishop), followed by purification on QIAGEN′s PN columns and ethanol precipitation of the flow through fraction.
RESULTS
Isolation of viruses resistant to siRNA.
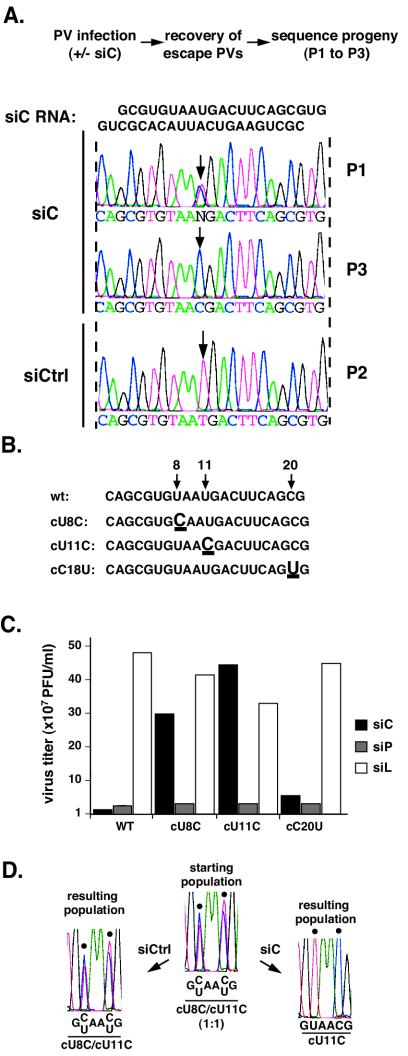
Transfection of HeLa cells with siC, an siRNA directed against a 21-nucleotide segment within the capsid-coding region (nucleotides 2417 to 2437), resulted in effective inhibition of viral replication. However, 30 to 40 h postinfection, resistant viruses emerged containing silent mutations within the siRNA-target sequence (Fig. 1A). The predominant mutation was a U-to-C transition at position 11 of the target region, but a second mutation, U to C at position 8, was also observed. By passage 3, the U-to-C mutation at nucleotide 11, but not that at position 8, prevailed in the viral population (Fig. 1A). Introduction of two of these mutations, cU8C and cU11C, into the poliovirus genome (Fig. 1B) confirmed that they were sufficient for the resistance phenotype. The most centrally located mutation cU11C conferred nearly full resistance, while a mutation at the 5′ side, cU8C, yielded partial resistance (Fig. 1C). In contrast, another mutation observed at the extreme 3′ side, cC20U, had little effect. Of note, these mutants remained fully susceptible to siRNA inhibition by siP, which targets a different region of the viral RNA. Thus, these mutations are specific for escape from siC and do not cause a general resistance to interference. We conclude that one single transition mutation within the siRNA complementary region of the target RNA is sufficient to dramatically reduce the efficiency of RNAi. To examine the relative strength of each escape mutation, we developed a competition assay in which a 1:1 mixture of escape mutants was used to infect siRNA-treated HeLa cells. After two passages in either the presence or absence of selective pressure, the resulting viral population was harvested and examined by sequencing. In siC-treated cells, cU11C became the predominant virus in the population, while in the presence of siCtrl, the initial 1:1 proportion of mutant viruses remained unchanged (Fig. 1D). Thus, the central mutation was more effective in blocking the inhibitory effect of siC. This data confirms previous studies highlighting the importance of the need for perfect base pairing at the central positions of the siRNA-target duplex (10, 14). Interestingly, the subtle difference in virus yield observed between cU8C and cU11C in single-virus cultures in the presence of siC (Fig. 1C) resulted in a dramatic advantage for cU11C in our competition assay (Fig. 1D). Thus, this sensitive assay can be employed to analyze subtle differences in the strength of different escape mutants.
FIG. 1.
Polioviruses resistant to siC. (A) Population sequencing of the viral siC region in siC-transfected cells. siCtrl, virus grown in cells transfected with control (rhinovirus-specific) siRNA after two passages; P1, a representative viral population after one passage in siC-transfected cells; P3, P1 following two additional passages under siC. Arrows point to the newly arising mutation. (B) Mutants in the siC region, engineered into the infectious clone of poliovirus. Numbers and arrows indicate the position of each mutation. (C) Titers of wild-type and mutant viruses following infection in HeLa cells transfected with siC (leftmost black bars), siP (middle gray bars), and siL (luciferase) (white bars). Infections were done at a multiplicity of infection (MOI) of 10 and produced virus collected at 8 h postinfection. (D) Competition assay between cU11C and cU8C. cU11C and cU8C were mixed at a 1:1 ratio and passaged twice with control (siCtrl)- or siC-transfected cells. The resulting population was analyzed by sequencing an RT-PCR product corresponding to the target region. Black dots indicate the positions of mutations (nucleotides 8 and 11).
Analysis of viruses resistant to an siRNA directed against the viral polymerase coding region.
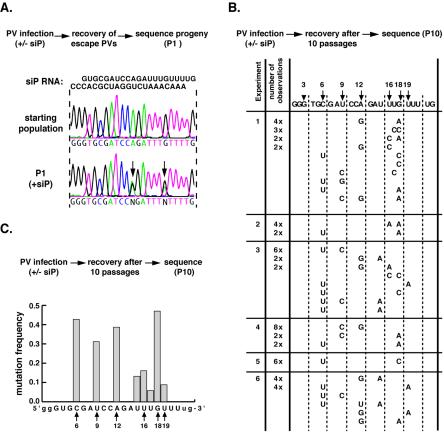
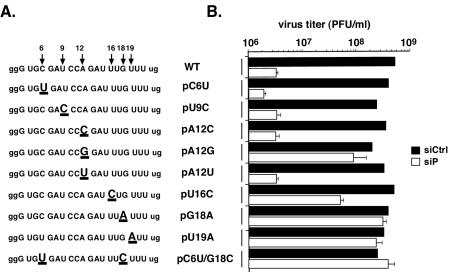
Analysis of escape mutants from another antiviral siRNA, siP (targeting nucleotides 6625 to 6645), provided insights regarding the rules that govern recognition of the target RNA by RISC. Three independent viral populations were sequenced after one passage in the presence of siP. Two major changes were observed within the target sequence at positions 12 and 18 (Fig. 2A). Unlike what was observed for siC, additional passages under siP selection did not result in one single mutant taking over the population. Instead, analysis of cDNA clones derived from resistant virus populations during six independent experiments revealed that even after 10 passages, a majority of the siP-resistant variants contained two mutations within the target region (Fig. 2B). We reasoned that several mutations in the target sequence were equally, but not entirely, effective in providing resistance to siP. Indeed, we did not isolate any single predominant mutation. The most frequent mutations occurred at position 6, at position 12 in the central region, and within a hot spot at the 3′ end of the target sequence (nucleotides 16 to 19) (Fig. 2C). Interestingly, we did not recover mutations at the extreme 5′ and 3′ ends (the first five and the last two nucleotides) of the target RNA, suggesting that these nucleotides are not critically involved in the interaction with siRNA. To evaluate the relative contribution of individual positions, we introduced single mutations in the siP target sequence of the wild-type poliovirus genome. We examined nine individual mutants carrying either single or double mutations within the target region (Fig. 3A). At position 12, three mutations were constructed, representing all three silent permutations of the central position. Other mutants carried nucleotide changes observed within the 5′ or 3′ half of the target region. In the absence of siP, none of the viruses carrying these point mutations displayed any replication defects and replicated in a manner similar to that of the wild type. Surprisingly, data based on our virus yield assay showed that mutations at positions 6 and 9, even though they were frequently isolated from the viral populations, did not provide significant resistance on their own to siP. In contrast, the central mutation (pA12G) and two mutations at the extreme 3′ end of the target sequence (pG18A and pU19A) were highly effective at neutralizing the inhibitory effect of siP. Interestingly, we find that mutations at the same position, such as pA12C and pA12U, are less robust than others (e.g., pA12G), indicating that the exact nature of the mismatch is also important in recognition.
FIG. 2.
Polioviruses resistant to siP. (A) Population sequencing of the viral siP region after one passage in siP-transfected cells. Arrows indicate newly arising mutations. (B) Sequences of individual viral genomes in the siP region appearing after 10 passages. Only nucleotides different from the wild-type sequence (shown on top) are spelled out. If a sequence was isolated more than once, then the number of isolations is given in the column second from the left. Six viral populations (numbered on the left as 1 to 6) were sequenced. Populations 1 to 3 are derived from passaging virus at an MOI of 10, and populations 4 to 6 are derived from passaging virus at an MOI of 0.1. Population pairs 1 and 4, 2 and 5, and 3 and 6 were related, as each pair was derived from one viral stock following two initial passages under siP. (C) Frequency of each mutation observed in siP-resistant viral population. Poliovirus was collected after 10 rounds of replication in siP-transfected HeLa cells. The target region was amplified by RT-PCR and cloned, and 70 clones were analyzed by sequencing. The mutation frequency was calculated as the number of times that a particular position was mutated divided by the total number of sequences analyzed.
FIG. 3.
Contribution of individual mutations to siP-resistance of poliovirus. (A) Sequences corresponding to the wild type (WT) and the siP escape mutants. (B) HeLa cells treated with either siP (white bars) or control siRNA (black bars) were infected at an MOI of 1. Virus was collected, and titers were determined after 10 h. siC infections were done in triplicate; error bars denote standard deviations.
let-7 microRNA effectively targets poliovirus genomic RNA.
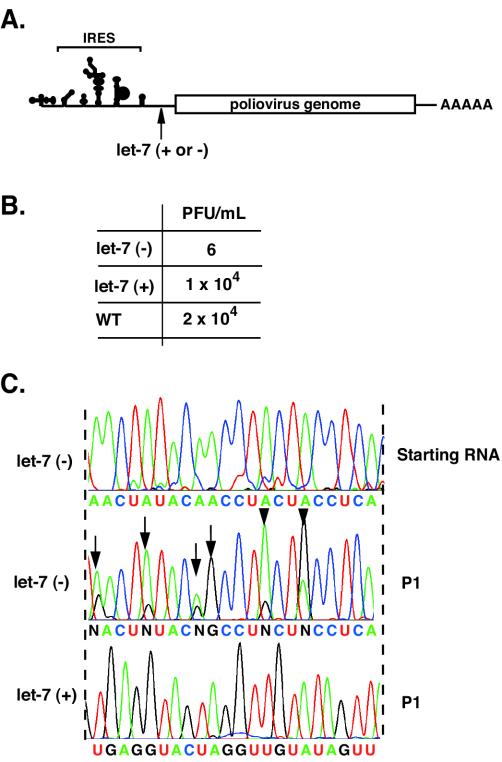
Poliovirus is a positive-stranded RNA virus that replicates through a negative-stranded intermediate. In principle, it is possible that either strand of the siRNAs can be incorporated into RISC. The rules governing the incorporation of siRNA strands into RISC are not fully defined (2), and thus either viral RNA strand can potentially be targeted by siC and siP. To understand the precise molecular consequences of the mutations leading to RNAi escape, it is important to determine which strand of the viral RNA the RNAi machinery targets. Previously, Ge et al. (11) examined this question in the context of infection by influenza virus, a negative-stranded virus. Because mutations in the antisense siRNA strand led to reduction of the RNAi effect while mutations in the sense siRNA strand did not, these authors concluded that only the viral mRNA, but not the genomic RNA, is targeted by siRNAs. A potential problem with this interpretation is that the mutations introduced in the siRNA may alter the efficiency of its incorporation into the RISC (24, 35). To circumvent this potential problem, we employed a different strategy using the particular property of microRNAs in which only one strand of the dsRNA precursor is selectively incorporated into RISC (19). Thus, by inserting either the identical or complementary microRNA sequence into the viral genome and assaying the viability of the virus in cells that express the microRNA, the susceptibility of the two strands to siRNA can be selectively evaluated. To this end, we cloned let-7 RNA [let-7(+)] or its complementary sequence [let-7(minus)] into the 5′ noncoding region of the viral genomic RNA (Fig. 4A). Following transfection into HeLa cells (which express high levels of let-7) (19, 26, 32), let-7(+) virus replicated with wild-type kinetics, while let-7(−) virus replication was impaired (Fig. 4B). Furthermore, viruses obtained from let-7(−) RNA transfections contained several mutations within the target sequence, while the target sequence of let-7(+) viruses was unchanged (Fig. 4C). Notably, all the mutations observed in let-7(−) virus populations were A-to-G transitions, which should result in G:U mismatches between let-7(+) RNA and let-7 microRNA, suggesting that these transition mutations allow for escape from let-7 RNA recognition. The most frequent position mutated was nucleotide 10 (Fig. 4C). We thus conclude that only the positive strand of the poliovirus RNA is targeted by siRNAs. In addition, this experiment also demonstrates that let-7 can act as an siRNA, directing cleavage of a perfectly complementary viral target RNA, as previously reported for nonviral systems (20).
FIG. 4.
Targeting poliovirus with let-7 microRNA. (A) Schematic representation of recombinant poliovirus carrying let-7 sequences. Either the identical let-7 RNA sequence [Let-7 (+)] or the anti-sense let-7 RNA [Let-7 (−)] was inserted at position 703 of the poliovirus genome. (B) After transfection of viral RNA into HeLa cells, viral titers were determined by infectious focus assay. (C) Population sequencing of the region in which let-7 (−) or let-7 (+) sequences were inserted. The top sequence corresponds to the starting viral RNA before replicating in HeLa cells. Let-7 (−)/P1 (middle sequence) corresponds to a representative viral population after one passage in HeLa cells. Let-7 (+)/P1 (bottom) corresponds to Let-7 (+) virus after one passage in HeLa cells. Arrows indicate an A-to-G transition mutation arising in Let-7 (−) populations.
Mapping the critical regions within the siRNA target.
Our competition assay provides a sensitive method for detecting subtle differences in the fitness of each variant carrying individual mutations. In the absence of siP, none of the mutants were outcompeted by wild-type poliovirus, indicating that these mutations do not affect viral replication. However, all the mutations, even pC6U and pU9C, which did not display a strong resistance phenotype in the virus yield assay (Fig. 3B), conferred some degree of resistance to RNAi and effectively outcompeted the wild type in siP-treated cells (data not shown). Furthermore, comparison of the respective competition trials for poliovirus resistant to either siC or siP provided a hierarchy of escape mutant strength and highlighted the importance of the different regions within the target RNA for effective siRNA recognition. In our competition assays, different results were obtained depending on the siRNA examined. For siC, the central position (cU11C) dominated over the rest of the region (Fig. 5B), while for siP, mutation both at the central position and at the extreme 3′ target region (pG18A and pU19A) could disrupt recognition (Fig. 5C). One possibility to explain these dissimilar results is that RISC may recognize the target in more than one manner. Perhaps, the local stability within the siRNA-target RNA duplex determines the weight of different mutations. Of note, the 3′ end of the siP target region is U rich; this local nucleotide characteristic may facilitate the unwinding of the siRNA-target RNA duplex. It is also possible that the degree of amino acid sequence conservation in each region may restrict the type of possible codon changes selected in escape mutants.
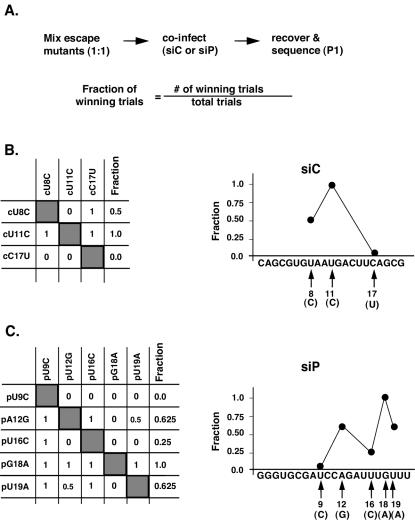
FIG. 5.
Relative escape strength of each mutation. (A) Schematic of the experiment. Two consecutive passages of a 1:1 mixture of the mutant viruses were performed with cells transfected with either siP or a control anti-rhinovirus siRNA. A change in the proportion of a given mutant after two passages was detected by RT-PCR and sequencing, as shown in Fig. 1D. Three siC mutants (B) and five siP mutants (C) were competed against each other. An increase in the proportion of a mutation in a given competition assay, defined as a winning trial, was scored as 1, a decrease was scored as 0, and no change in proportions was scored as 0.5. Each table (B and C) shows the scored mutants on the left, followed by their scores against each competing mutant (top). The fraction of winning trials (Fraction) was then calculated as the number of winning trials divided by the total number of trials. This allowed us to determine the relative escape efficiency of each mutation along the targeted region in the graphs shown on the right of panels B and C.
Targeting conserved sequences within the viral genome.
Our finding that single mutations in the viral genome consistently led to escape from siRNA inhibition uncovered a major weakness of RNAi as an antivirus therapy. One possible strategy to prevent the emergence of escape mutants is to target functional RNA sequences that, when mutated, lead to replication-deficient viruses. Strikingly, siRNAs targeting two sites in the highly conserved 5′ noncoding region of the poliovirus genome (site 1, nucleotides 445 to 465; site 2, nucleotides 536 to 556) failed to inhibit virus replication (data not shown). These regions may be inaccessible to RISC due to tertiary structure or because proteins interact with these RNA motifs, occluding the target sequences. However, it was recently reported that the highly structured 5′ noncoding region of the hepatitis C virus genome can be targeted by siRNA (39). Thus, it is also possible that the high G:C content of our siRNAs targeting the poliovirus IRES may have precluded their efficient incorporation into RISC, reducing their antiviral activity.
Simultaneous targeting with multiple siRNAs.
An alterative solution for preventing escape might be to simultaneously target multiple sequences in the viral genome. The ability of a mixed population of siRNAs to inhibit viral replication without allowing the generation of escape mutants was initially tested by transfecting P19 mouse embryonic carcinoma cells with a plasmid expressing the poliovirus receptor together with a 1-kb-long dsRNA corresponding to the poliovirus type 1 capsid region (dsC). P19 cells have the ability to initiate RNAi by converting long dsRNA into siRNAs without triggering an interferon response (3). The poliovirus titer was reduced more than 1,000 fold after treatment with dsC dsRNA (Fig. 6A), while control dsRNA (dsL, corresponding to 1,000 nucleotides of the luciferase gene) had no effect on viral production. This inhibition was sequence specific, as poliovirus type 3, which shares only 54% nucleotide identity with type 1 in the targeted capsid region, was not inhibited by dsC. Importantly, due to the emergence of escape mutants, siP only inhibited viral replication at early times postinfection (<10 h), while dsC inhibited poliovirus replication for the entire duration of the experiment (60 h), which represents at least six replication cycles (Fig. 6A). Because long dsRNA elicits a nonspecific antiviral response in a majority of eukaryotic cells, we examined whether targeting viral sequences with multiple siRNAs would prevent the emergence of escape mutants. The efficiency of RNAi was examined in viral populations obtained during serial passages in the presence of individual siRNAs or a mixture of enzymatically produced siRNAs (esiRNAs) (38). We generated esiRNAs by bacterial RNase III digestion of a 1-kb-long poliovirus capsid dsRNA dsC (esiC) or luciferase dsL (esiL) control. Transfection of esiC protected HeLa cells against type 1 poliovirus but not against type 3. In contrast, cells treated with esiL control remained susceptible to both viral strains. Most importantly, we observed no reduction of the antiviral effect with each subsequent passage, indicating that the virus remains susceptible to RNAi even after several consecutive treatments (Fig. 6B). This was directly confirmed by sequencing the 1-kb targeted region from viruses obtained after the third passage under esiC or esiL. We observed only one mutation out of six independent clones derived from esiC-treated viruses and no mutations in three clones of esiL-treated viruses (data not shown). We conclude that treatment with esiRNAs provides an efficient approach to inhibit virus replication and, strikingly, does not allow for selection of escape mutants over the limited number of generations in our experiment (approximately six to eight rounds of replication).
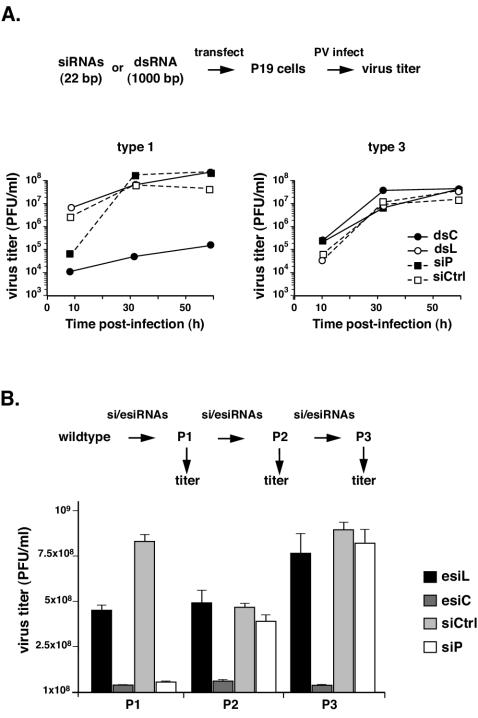
FIG. 6.
Strategy to prevent development of RNAi-resistant viruses. (A) Titers of type 1 and type 3 strains grown in P19 cells transfected with 1-kb-long dsRNA corresponding to the poliovirus type 1 genome (dsC), dsRNA to firefly luciferase (dsL), or siRNA siP or siCtrl. (B) Inhibition of poliovirus type 1 after consecutive passages in HeLa cells transfected with siRNAs siP and siCtrl or to esiRNAs prepared from dsC (esiC) and dsL (esiL). Viral titers are shown after each of three consecutive passages (P1, P2, and P3). For panels A and B, a scheme of the experiment is shown on top; see the text for details.
DISCUSSION
The rules of siRNA-target recognition have been examined employing diverse siRNAs with nucleotide substitutions (1, 7, 10, 14, 33). Here, we present a novel approach to study RNAi specificity that exploits the ability of viruses to rapidly generate a spectrum of resistant mutants. Importantly, by analyzing a single siRNA per target region, our analysis is not biased by the specificity of the siRNA processing machinery (i.e., association with RISC and unwinding) (24, 35). While observing coding sequence does have its drawbacks, our results provide insight into the rules of target recognition by siRNAs. We find that some positions, notably the center of the siRNA, are critical for recognition (Fig. 1 to 3). Mutations at either side of the central position are also effective at blocking siRNA inhibition. In addition, the 3′ end of the target sequence plays an important role in the target selection by siP (pG18A and pU19A).
Strikingly, our study also demonstrates that the RNAi machinery is exquisitely sensitive to the nature of the mismatches. For example, pA12G, which produces a G:U mismatch, abolishes RNAi targeting much more efficiently than pA12C or pA12U (Fig. 3B), which results in less stable C:U or U:U base pairing (25). Thus, it appears that the RNAi machinery does not rely solely on the thermodynamic characteristics of the duplex to select cognate target RNAs. Instead, RISC seems to discriminate based on the architectural and structural properties of the siRNA-target RNA duplex. Notably, mutations that do not confer effective resistance to siRNA by themselves (e.g., pC6U and pU9C) were repeatedly isolated in double-mutant clones (compare Fig. 2 and 3). This observation suggests that RISC can detect mismatches in a cooperative manner. This may contribute to the overall specificity of the RNAi targeting process.
The spectrum of mutants studied here is limited by the nucleotide misincorporation frequency at each given position and by the viability of the resulting virus. This constrains the mutations largely, although not exclusively to the wobble positions in the codons and does not allow interrogating each position of the target sequence. However, we find that some locations within the noncoding regions can be effectively targeted by RISC (Fig. 4). Thus, the analysis of escape mutants resistant to siRNAs that target sequences in the noncoding regions should help to understand basic rules that govern siRNA-target recognition.
Our analysis has important implications for the use of RNAi as an antiviral therapy. We find that a single substitution within the viral RNA is sufficient to render siRNAs ineffective within only a few replication cycles, even at highly conserved target regions. The very high viral mutation rates, coupled with the enormous genome heterogeneity in viral quasispecies, make it unlikely that defined, single siRNAs can be effective against viral pathogens. However, the plant RNAi machinery is very effective in combating viruses using the entire length of viral genomes a source of siRNAs. This should result in a complex mixture of siRNAs, which cannot be evaded through a limited number of mutations in the viral genome. Indeed, a 1-kb dsRNA is potentially capable of forming up to 980 different siRNAs, which should prevent the emergence of resistant viruses. Initial evidence in plant virus systems suggested that viral evasion is restricted by expression of long dsRNA in transgenic crops (15). We have now directly examined this issue in a human virus. We found that transfection of long dsRNA prevents the emergence of viral escape mutants. Thus, the use of long dsRNA may constitute an effective therapy against human and other mammalian viruses.
Unlike plants and invertebrates, mammals seem to have multiple pathways of reacting to long dsRNA, some of which may lead to a general translation shutoff (30) and possibly unwanted side effects (16). Therefore, we also examined an alternative approach to elicit silencing using dsRNA enzymatically processed into siRNAs. This method also proved effective at preventing the emergence of escape mutants, as confirmed by viral titers and genome sequencing (Fig. 6 and data not shown).
In summary, we demonstrate here that generation of escape mutants can be prevented by targeting long portions of the virus genome with dsRNA, which is an RNAi equivalent to multidrug therapy with hundreds of targets. Since the approach described here is a straightforward sequence of enzymatic reactions (reverse transcription-PCR [RT-PCR], transcription annealing, and perhaps RNase III digestion), it is extremely versatile due to its speed, low cost, and uniformity in use against different viruses. In fact, it may be useful even when the exact sequence of the viral genome is not known in advance, so long as PCR primers used for its amplification are specific. In this way, we suggest that the viral capacity to escape RNAi-based therapies can be restricted.
Acknowledgments
We are grateful to Judith Frydman, Alan Frankel, Philip Zamore, and members of the Andino lab for critical reviews of the manuscript.
This work was supported by an NIH fellowship to J.K.S. and by NIH grant AI40085 to R.A.
REFERENCES
- 1.Amarzguioui, M., T. Holen, E. Babaie, and H. Prydz. 2003. Tolerance for mutations and chemical modifications in a siRNA. Nucleic Acids Res. 31:589-595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Bartel, D. P. 2004. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281-297. [DOI] [PubMed] [Google Scholar]
- 3.Billy, E., V. Brondani, H. Zhang, U. Muller, and W. Filipowicz. 2001. Specific interference with gene expression induced by long, double-stranded RNA in mouse embryonal teratocarcinoma cell lines. Proc. Natl. Acad. Sci. USA 98:14428-14433. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bitko, V., and S. Barik. 2001. Phenotypic silencing of cytoplasmic genes using sequence-specific double-stranded short interfering RNA and its application in the reverse genetics of wild type negative-strand RNA viruses. BMC Microbiol. 1:34. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Boden, D., O. Pusch, F. Lee, L. Tucker, and B. Ramratnam. 2003. Human immunodeficiency virus type 1 escape from RNA interference. J. Virol. 77:11531-11535. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Chi, J. T., H. Y. Chang, N. N. Wang, D. S. Chang, N. Dunphy, and P. O. Brown. 2003. Genomewide view of gene silencing by small interfering RNAs. Proc. Natl. Acad. Sci. USA 100:6343-6346. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Chiu, Y. L., and T. M. Rana. 2003. siRNA function in RNAi: a chemical modification analysis. RNA 9:1034-1048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Coburn, G. A., and B. R. Cullen. 2002. Potent and specific inhibition of human immunodeficiency virus type 1 replication by RNA interference. J. Virol. 76:9225-9231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Crotty, S., B. L. Lohman, F. X. Lu, S. Tang, C. J. Miller, and R. Andino. 1999. Mucosal immunization of cynomolgus macaques with two serotypes of live poliovirus vectors expressing simian immunodeficiency virus antigens: stimulation of humoral, mucosal, and cellular immunity. J. Virol. 73:9485-9495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Elbashir, S. M., J. Martinez, A. Patkaniowska, W. Lendeckel, and T. Tuschl. 2001. Functional anatomy of siRNAs for mediating efficient RNAi in Drosophila melanogaster embryo lysate. EMBO J. 20:6877-6888. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Ge, Q., M. T. McManus, T. Nguyen, C. H. Shen, P. A. Sharp, H. N. Eisen, and J. Chen. 2003. RNA interference of influenza virus production by directly targeting mRNA for degradation and indirectly inhibiting all viral RNA transcription. Proc. Natl. Acad. Sci. USA 100:2718-2723. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Gitlin, L., S. Karelsky, and R. Andino. 2002. Short interfering RNA confers intracellular antiviral immunity in human cells. Nature 418:430-434. [DOI] [PubMed] [Google Scholar]
- 13.Hannon, G. J. 2002. RNA interference. Nature 418:244-251. [DOI] [PubMed] [Google Scholar]
- 14.Harborth, J., S. M. Elbashir, K. Vandenburgh, H. Manninga, S. A. Scaringe, K. Weber, and T. Tuschl. 2003. Sequence, chemical, and structural variation of small interfering RNAs and short hairpin RNAs and the effect on mammalian gene silencing. Antisense Nucleic Acid Drug Dev. 13:83-105. [DOI] [PubMed] [Google Scholar]
- 15.Harrison, B. D. 2002. Virus variation in relation to resistance breaking in plants. Euphytica 124:181-192. [Google Scholar]
- 16.Hendrix, C. W., J. B. Margolick, B. G. Petty, R. B. Markham, L. Nerhood, H. Farzadegan, P. O. Ts'o, and P. S. Lietman. 1993. Biologic effects after a single dose of poly(I):poly(C12U) in healthy volunteers. Antimicrob. Agents Chemother. 37:429-435. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Herold, J., and R. Andino. 2000. Poliovirus requires a precise 5′ end for efficient positive-strand RNA synthesis. J. Virol. 74:6394-6400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Hu, W. Y., C. P. Myers, J. M. Kilzer, S. L. Pfaff, and F. D. Bushman. 2002. Inhibition of retroviral pathogenesis by RNA interference. Curr. Biol. 12:1301-1311. [DOI] [PubMed] [Google Scholar]
- 19.Hutvagner, G., J. McLachlan, A. E. Pasquinelli, E. Balint, T. Tuschl, and P. D. Zamore. 2001. A cellular function for the RNA-interference enzyme Dicer in the maturation of the let-7 small temporal RNA. Science 293:834-838. [DOI] [PubMed] [Google Scholar]
- 20.Hutvagner, G., and P. D. Zamore. 2002. A microRNA in a multiple-turnover RNAi enzyme complex. Science 297:2056-2060. [DOI] [PubMed] [Google Scholar]
- 21.Jackson, A. L., S. R. Bartz, J. Schelter, S. V. Kobayashi, J. Burchard, M. Mao, B. Li, G. Cavet, and P. S. Linsley. 2003. Expression profiling reveals off-target gene regulation by RNAi. Nat. Biotechnol. 21:635-637. [DOI] [PubMed] [Google Scholar]
- 22.Jacque, J. M., K. Triques, and M. Stevenson. 2002. Modulation of HIV-1 replication by RNA interference. Nature 418:435-438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Jiang, M., and J. Milner. 2002. Selective silencing of viral gene expression in HPV-positive human cervical carcinoma cells treated with siRNA, a primer of RNA interference. Oncogene 21:6041-6048. [DOI] [PubMed] [Google Scholar]
- 24.Khvorova, A., A. Reynolds, and S. D. Jayasena. 2003. Functional siRNAs and miRNAs exhibit strand bias. Cell 115:209-216. [DOI] [PubMed] [Google Scholar]
- 25.Kierzek, R., M. E. Burkard, and D. H. Turner. 1999. Thermodynamics of single mismatches in RNA duplexes. Biochemistry 38:14214-14223. [DOI] [PubMed] [Google Scholar]
- 26.Lagos-Quintana, M., R. Rauhut, W. Lendeckel, and T. Tuschl. 2001. Identification of novel genes coding for small expressed RNAs. Science 294:853-858. [DOI] [PubMed] [Google Scholar]
- 27.Martinez, J., A. Patkaniowska, H. Urlaub, R. Luhrmann, and T. Tuschl. 2002. Single-stranded antisense siRNAs guide target RNA cleavage in RNAi. Cell 110:563. [DOI] [PubMed] [Google Scholar]
- 28.Martinez, M. A., A. Gutierrez, M. Armand-Ugon, J. Blanco, M. Parera, J. Gomez, B. Clotet, and J. A. Este. 2002. Suppression of chemokine receptor expression by RNA interference allows for inhibition of HIV-1 replication. AIDS 16:2385-2390. [DOI] [PubMed] [Google Scholar]
- 29.McCaffrey, A. P., H. Nakai, K. Pandey, Z. Huang, F. H. Salazar, H. Xu, S. F. Wieland, P. L. Marion, and M. A. Kay. 2003. Inhibition of hepatitis B virus in mice by RNA interference. Nat. Biotechnol. 21:639-644. [DOI] [PubMed] [Google Scholar]
- 30.Nicholson, A. W. 1996. Structure, reactivity, and biology of double-stranded RNA. Prog. Nucleic Acid Res. Mol. Biol. 52:1-65. [DOI] [PubMed] [Google Scholar]
- 31.Novina, C. D., M. F. Murray, D. M. Dykxhoorn, P. J. Beresford, J. Riess, S. K. Lee, R. G. Collman, J. Lieberman, P. Shankar, and P. A. Sharp. 2002. siRNA-directed inhibition of HIV-1 infection. Nat. Med. 8:681-686. [DOI] [PubMed] [Google Scholar]
- 32.Pasquinelli, A. E., B. J. Reinhart, F. Slack, M. Q. Martindale, M. I. Kuroda, B. Maller, D. C. Hayward, E. E. Ball, B. Degnan, P. Muller, J. Spring, A. Srinivasan, M. Fishman, J. Finnerty, J. Corbo, M. Levine, P. Leahy, E. Davidson, and G. Ruvkun. 2000. Conservation of the sequence and temporal expression of let-7 heterochronic regulatory RNA. Nature 408:86-89. [DOI] [PubMed] [Google Scholar]
- 33.Pusch, O., D. Boden, R. Silbermann, F. Lee, L. Tucker, and B. Ramratnam. 2003. Nucleotide sequence homology requirements of HIV-1-specific short hairpin RNA. Nucleic Acids Res. 31:6444-6449. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Qin, X. F., D. S. An, I. S. Chen, and D. Baltimore. 2003. Inhibiting HIV-1 infection in human T cells by lentiviral-mediated delivery of small interfering RNA against CCR5. Proc. Natl. Acad. Sci. USA 100:183-188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Schwarz, D. S., G. Hutvagner, T. Du, Z. Xu, N. Aronin, and P. D. Zamore. 2003. Asymmetry in the assembly of the RNAi enzyme complex. Cell 115:199-208. [DOI] [PubMed] [Google Scholar]
- 36.Semizarov, D., L. Frost, A. Sarthy, P. Kroeger, D. N. Halbert, and S. W. Fesik. 2003. Specificity of short interfering RNA determined through gene expression signatures. Proc. Natl. Acad. Sci. USA 100:6347-6352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Song, E., S. K. Lee, D. M. Dykxhoorn, C. Novina, D. Zhang, K. Crawford, J. Cerny, P. A. Sharp, J. Lieberman, N. Manjunath, and P. Shankar. 2003. Sustained small interfering RNA-mediated human immunodeficiency virus type 1 inhibition in primary macrophages. J. Virol. 77:7174-7181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Yang, D., F. Buchholz, Z. Huang, A. Goga, C. Y. Chen, F. M. Brodsky, and J. M. Bishop. 2002. Short RNA duplexes produced by hydrolysis with Escherichia coli RNase III mediate effective RNA interference in mammalian cells. Proc. Natl. Acad. Sci. USA 99:9942-9947. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Yokota, T., N. Sakamoto, N. Enomoto, Y. Tanabe, M. Miyagishi, S. Maekawa, L. Yi, M. Kurosaki, K. Taira, M. Watanabe, and H. Mizusawa. 2003. Inhibition of intracellular hepatitis C virus replication by synthetic and vector-derived small interfering RNAs. EMBO Rep. 4:602-608. [DOI] [PMC free article] [PubMed] [Google Scholar]