Abstract
IDS is responsible for the lysosomal degradation of heparan sulfate and dermatan sulfate and linked to an X-linked lysosomal storage disease, mucopolysaccharidosis 2 (MPS2), resulting in neurological damage and early death. Comparative IDS amino acid sequences and structures and IDS gene locations were examined using data from several vertebrate genome projects. Vertebrate IDS sequences shared 60–99% identities with each other. Human IDS showed 47% sequence identity with fruit fly (Drosophila melanogaster) IDS. Sequence alignments, key amino acid residues, N-glycosylation sites and conserved predicted secondary and tertiary structures were also studied, including signal peptide, propeptide and active site residues. Mammalian IDS genes usually contained 9 coding exons. The human IDS gene promoter contained a large CpG island (CpG46) and 5 transcription factor binding sites, whereas the 3′-UTR region contained 5 miRNA target sites. These may contribute to IDS gene regulation of expression in the brain and other neural tissues of the body. An IDS pseudogene (IDSP1) was located proximally to the IDS gene on the X-chromosome in primate genomes. Phylogenetic analyses examined the relationships and potential evolutionary origins of the vertebrate IDS gene. These suggested that IDS has originated in an invertebrate ancestral genome and retained throughout vertebrate evolution and conserved on marsupial and eutherian X-chromosomes, with the exception of rat Ids on chromosome 8.
Electronic supplementary material
The online version of this article (doi:10.1007/s13205-016-0595-3) contains supplementary material, which is available to authorized users.
Keywords: Vertebrates, Iduronate 2-sulfatase, Amino acid sequence, IDS, X-chromosome, IDS gene regulation, Evolution
Introduction
Iduronate 2-sulfatase (IDS; EC 3.1.6.13) is responsible for the lysosomal degradation of the glycoaminoglycans, heparan sulfate and dermatan sulfate (Bielicki et al. 1990), and is one of the 19 members of human sulfatase gene families and 17 members of the mouse sulfatase gene families which catalyze the hydrolysis of sulfate esters in the body derived from several catabolic pathways (Ratzka et al. 2010). Many IDS gene mutations and IDS deficiencies have been studied in human populations which result in the lysosomal storage of glycoaminoglycans and Hunter syndrome, an X-linked chromosome disease, referred to as mucopolysaccharidosis type 2 (MPS2) (Wilson et al. 1990; Rathmann et al. 1996; Chistiakov et al. 2014; Kosuga et al. 2016). Major clinical features for this rare genetic disease (1:100,000 births) include obstructive and restrictive airway disease, skeletal deformations, cardiac disease, joint contractures and mental retardation (Beck 2011; Tylki-Szymańska 2014; Anekar et al. 2015). Mouse and zebra fish animal models have been used to study the disease in more detail, including studies of Ids − /Ids − knock out mice which have shown that IDS-deficiency generates many of the defects reported for human MPS2 (Garcia et al. 2007). In addition, possible treatments for the disease by enzyme replacement therapy have been investigated (Garcia et al. 2007; Moro et al. 2010; Fusar Poli et al. 2013; Cho et al. 2015; Parini et al. 2015) and a phase I/II clinical trial of intrathecal IDS replacement therapy in children with severe MPS2 has been recently reported (Muenzer et al. 2016).
The gene encoding IDS (IDS in primates; Ids in rodents) is expressed at high levels in neural tissues, particularly in the cortex, hippocampus, other brain and eye tissues; and is also widely expressed throughout the body (Smith et al. 2014). The enzyme catalyzes the first step in the degradation of glycoaminoglycans, dermatan sulfate and heparan sulfate (Bielicki et al. 1990). Human IDS is expressed as three major isoforms which have distinct C-terminal sequences: IDSa encoding a 550 amino acid protein, expressed in brain tissues and with a wide tissue distribution; IDSb, 460 amino acids also expressed in brain tissues; and IDSc, encoding a 446 amino acid enzyme expressed in ductal carcinoma cells and pancreas (Thierry-Mieg and Thierry-Mieg 2006). The genomic organization of the human and mouse IDS/Ids genes have been reported with 9 exons observed for 24 kb and 22 kbs of DNA, respectively (Wilson et al. 1993; Thierry-Mieg and Thierry-Mieg 2006).
Biochemical and predictive structural studies of human IDS have shown that it comprises several domains: an N-terminus signal peptide (residues 1–25); a propeptide sequence (residues 26–33); five Ca2+ binding sites (1 Ca2+ per subunit); two active site residues (334Asp and 335His); and seven N-glycosylation sites (Bielicki et al. 1990; Wilson et al. 1990; Kosuga et al. 2016). A predicted tertiary structure has been reported for human IDS (Sáenz et al. 2007), which shows strong similarities with other human sulfatases: GALNS (Rivera-Colón et al. (2012)); ARSA (Chruszcz et al. 2003) and STS (Hernandez-Guzman et al. 2003).
This paper reports the predicted gene structures and amino acid sequences for several vertebrate IDS genes and proteins, the predicted structures for vertebrate IDS proteins, a number of potential sites for regulating human IDS gene expression and the structural, phylogenetic and evolutionary relationships for these genes and enzymes.
Methods
Vertebrate IDS gene and protein identification
BLAST studies were undertaken using web tools from NCBI (http://www.ncbi.nlm.nih.gov/) (Camacho et al. 2009). Protein BLAST analyses used human and mouse IDS amino acid sequences previously described (Bielicki et al. 1990; Garcia et al. 2007) (Table 1). Protein sequence databases for several vertebrate genomes were examined using the blastp algorithm (see Holmes 2016). Predicted IDS protein sequences were obtained in each case and subjected to analyses of predicted protein and gene structures.
Table 1.
Vertebrate IDS Proteins
| IDS Protein | Species | UNIPROT ID | Amino acids | Subunit MW | pI | N-Glycosylation sites | Signal peptide | % Identity human IDS |
|---|---|---|---|---|---|---|---|---|
| Human | Homo sapiens | P22304 | 550 | 61,873 | 5.2 | 115, 144, 246, 280, 325, 513, 537 | 1..25 | 100 |
| Chimpanzee | Pan troglodytes | na | 550 | 61,861 | 5.2 | 115, 144, 246, 280, 325, 513, 537 | 1..25 | 99 |
| Orangutan | Pongo abelii | H2PX10 | 550 | 62,083 | 5.4 | 115, 144, 246, 280, 325, 513, 537 | 1..25 | 96 |
| Baboon | Papio anubis | na | 550 | 61,885 | 5.1 | 115, 144, 246, 280, 325, 513, 537 | 1..25 | 96 |
| Marmoset | Callithrix jacchus | F7EJG2 | 550 | 61,812 | 5.4 | 115, 144, 246, 280, 325, 513, 537 | 1..25 | 94 |
| Mouse | Mus musculus | Q08890 | 552 | 62,186 | 5.5 | 117, 146, 248, 282, 515, 539 | 1..29 | 86 |
| Rat | Rattus norvegicus | Q32KJ4 | 543 | 62,370 | 5.5 | 117, 146, 248, 181, 515, 539 | 1..20 | 85 |
| Cow | Bos taurus | F1N2D5 | 547 | 61,389 | 5.8 | 112, 141, 243, 277, 509, 533 | 1..20 | 82 |
| Sheep | Ovis aries | W5PI67 | 547 | 61,019 | 5.6 | 112, 141, 243, 277, 510, 534 | 1..20 | 82 |
| Opossum | Monodelphis domestica | F7DJA1 | 558 | 63,374 | 5.3 | 129, 260, 294, 339, 457, 524, 552 | 1..23 | 75 |
| Tasmanian devil | Sarcophilus harrisii | na | 539 | 61,392 | 5.3 | 111, 140, 242, 276, 321, 505, 509, 533 | 1..22 | 74 |
| Chicken | Gallus gallus | F1NFI0 | 601 | 68,047 | 6.6 | 156, 185, 287, 321, 366, 584 | na | 67 |
| Lizard | Anolis carolinensis | H9GGQ8 | 524 | 59,239 | 5.8 | 92, 121, 223, 257, 478, 507 | na | 63 |
| Frog | Xenopus tropicalis | A8WGX6 | 542 | 61,858 | 6.1 | 112, 141, 243, 277, 322 | 1..18 | 66 |
| Zebra fish | Danio rerio | A1A5V0 | 561 | 63,771 | 7.7 | 109, 138, 181, 240, 274, 499 | 1..25 | 60 |
| Fruit Fly | Drosophila melanogaster | na | 502 | 57,760 | 7.3 | 93, 12, 22, 22, 22, 82, 60, 400 | na | 47 |
UNIPROT refers to UniprotKB/Swiss-Prot IDs for individual IDS proteins (see http://kr.expasy.org); pI refers to theoretical isoelectric points
BLAT analyses were subsequently undertaken for each of the predicted IDS amino acid sequences using the UC Santa Cruz (UCSC) Genome Browser with the default settings to obtain the predicted locations for each of the vertebrate IDS genes, including predicted exon boundary locations and gene sizes (Kent et al. 2002). BLAT analyses were similarly undertaken for other vertebrate IDS genes using previously reported sequences in each case (Table 2). Structures for human isoforms (splicing variants) were obtained using the AceView website to examine predicted gene and protein structures (Thierry-Mieg and Thierry-Mieg 2006).
Table 2.
Vertebrate IDS Genes
| IDS Gene | Species | RefSeq ID | GenBank ID | Chromosome location | Coding exons (strand) | Gene size (bps) |
|---|---|---|---|---|---|---|
| Human | Homo sapiens | NM_000202 | BC006170 | X:149,482,749–149,505,137 | 9 (−ve) | 22,389 |
| IDSP1 | Homo sapiens | na | na | X:149,525,002–149,525,923 | na | 922 |
| Chimpanzee | Pan troglodytes | XP_016799854 | na | X:150,217,197–150,239,595 | 9 (−ve) | 22,399 |
| Orangutan | Pongo abelii | XP_002832265 | na | X:149,468,629–149,491,886 | 9 (−ve) | 23,258 |
| Baboon | Papio anubis | XP_003918436 | na | X:137,241,259–137,263,520 | 9 (−ve) | 22,262 |
| Marmoset | Callithrix jacchus | XP_002763402 | na | X:136,661,096–136,690,421 | 9 (−ve) | 29,326 |
| Mouse | Mus musculus | NM_010498 | BN000750 | X:70,346,204–70,364,903 | 9 (−ve) | 18,700 |
| Rat | Rattus norvegicus | XP_017451660 | BN000743 | 8:69,158,393–69,174,447 | 9 (−ve) | 16,055 |
| Cow | Bos taurus | NM_001192851 | na | X:32,309,006–32,324,359 | 9 (−ve) | 15,354 |
| Sheep | Ovis aries | XP_012016345 | na | X:81,295,118–81,310,976 | 9 (+ve) | 15,859 |
| Opossum | Monodelphis domestica | XP_007507328 | na | X:38,769,936–38,797,831 | 9 (−ve) | 27,896 |
| Tasmanian devil | Sarcophilus harrisii | XP_012408735 | na | X_GL867598:1,290,327–1,307,074 | 9 (−ve) | 16,748 |
| Chicken | Gallus gallus | XP_015133789 | na | 4:18,031,638–18,046,283 | 9 (+ve) | 14,646 |
| Lizard | Anolis carolinensis | XP_016851828 | na | GL343310:1,066,926–1,092,995 | 8 (+ve) | 26,070 |
| Frog | Xenopus tropicalis | NM_001197132 | BC154891 | KB021658:33,136,298–33,145,211 | 9 (+ve) | 8914 |
| Zebra fish | Danio rerio | NM_001080068 | BC128823 | 14:20,572,602–20,594,434 | 8 (−ve) | 21,833 |
| Fruit Fly | Drosophila melanogaster | NM_139557 | AAY55004 | 3L:3,378,315–3,380,000 | 4 (+ve) | 1686 |
GenBank IDs are derived from NCBI http://www.ncbi.nlm.nih.gov/genbank/; GL and KB refer to a scaffold; bps refers to base pairs of nucleotide sequences; the number of coding exons are listed
RefSeq The reference sequence, XP predicted sequence, na not available
Predicted structures and properties of vertebrate IDS
Predicted secondary and tertiary structures for vertebrate IDS proteins were obtained using the SWISS-MODEL web-server (http://swissmodel.expasy.org/) (Schwede et al. 2003) using the reported tertiary structure for human arylsulfatase A (ARSA) (Lukatela et al. 1998; Chrusczcs et al. 2003) (PDB:1n2kA) with a modeling range of 35–549 for human IDS. Molecular weights, N-glycosylation sites and signal peptide cleavage sites for vertebrate IDS proteins were obtained using Expasy web tools (http://au.expasy.org/tools/pi_tool.html). The identification of conserved domains for IDS was conducted using NCBI web tools (Marchler-Bauer et al. 2011).
Human IDS tissue expression
RNA-seq gene expression profiles across 53 selected tissues (or tissue segments) were examined from the public database for human IDS, based on expression levels for 175 individuals (GTEx Consortium 2015) (Data Source: GTEx Analysis Release V6p (dbGaP Accession phs000424.v6.p1) (http://www.gtex.org).
Amino acid sequence alignments and phylogenetic analyses
Alignments of vertebrate and Drosophila melanogaster IDS sequences were undertaken using Clustal Omega, a multiple sequence alignment program (Sievers and Higgins 2014) (Table 1). Percentage identities were derived from the results of these alignments (Table 1). Phylogenetic analyses used several bioinformatic programs, coordinated using the http://www.phylogeny.fr/ bioinformatic portal, to enable alignment (MUSCLE), curation (Gblocks), phylogeny (PhyML) and tree rendering (TreeDyn), to reconstruct phylogenetic relationships (Dereeper et al. 2008). Sequences were identified as vertebrate IDS members and a proposed primordial Drosophila melanogaster IDS gene and protein (Tables 1, 2).
Results and discussion
Alignments of vertebrate IDS amino acid sequences
The deduced amino acid sequences for frog (Xenopus tropicalis) and zebrafish (Danio rerio) IDS are shown in Fig. 1 together with previously reported sequences for human (Bielicki et al. 1990) and mouse IDS (Garcia et al. 2007) (Table 1). Alignments of human with other vertebrate IDS sequences examined were between 60 and 99% identical, suggesting that these are products of the same family of genes, whereas comparisons of sequence identities of vertebrate IDS proteins with other human ARS proteins exhibited ≥27% identities, indicating that these are members of distinct ARS-like gene families (Table 1; Supplementary Table 1).
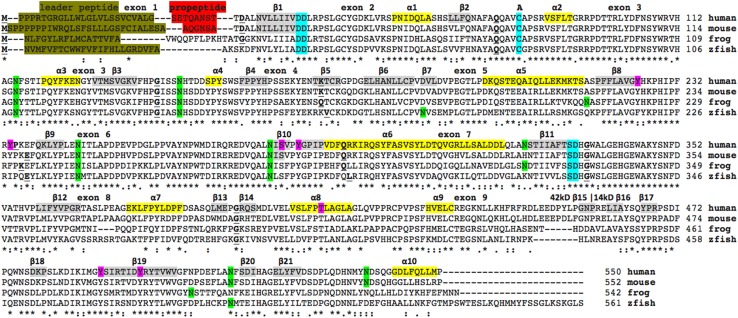
Fig. 1.
Amino acid sequence alignments for vertebrate IDS sequences. See Table 1 for sources of IDS sequences; asterisk shows identical residues for IDS subunits; colon similar alternate residues; dot dissimilar alternate residues; predicted phosphoresidues are in pink; predicted N-glycosylated Asn sites are in green; the active site residues (for human IDS) are shown in blue; active site residue subject to modification is shown as A; predicted α-helices for human IDS is in shaded yellow and numbered in sequence; predicted β-sheets are in shaded gray and also numbered in sequence from the N-terminus; bold underlined font shows residues corresponding to known or predicted exon start sites; exon numbers refer to human IDS gene exons; leader peptide is in brown; propeptide in red
The amino acid sequences for vertebrate IDS proteins contained 550–561 amino acids (Fig. 1; Table 1). Previous studies have reported several key regions and residues for human and mouse IDS proteins (human IDS amino acid residues were identified in each case) (Bielicki et al. 1990). These included an N-terminus leader peptide (24 residues excluding the N-terminus methionine) followed by a propeptide 8-residue segment (residues 25–33) (Wilson et al. 1990). A comparison of 10 mammalian IDS sequences for these N-terminal exon 1 regions revealed species specific variability in these sequences, with the signal peptides containing multiple proline and hydrophobic residues, and the propeptides exhibiting distinct mammalian sequences (see Figs. 1, 2). In contrast, amino acid sequences located further upstream within exon 2, nearer to the active site catalytic residues (Asp45; Asp46), were predominantly invariant among the mammalian and other vertebrate sequences examined (Figs. 1, 2). One of the conserved active site residues observed for these mammalian and other vertebrate IDS sequences, included an active site catalytic residue (Cys84) which undergoes post-translational modification by sulfatase modifying factor 1 (SUMF1) to form C(alpha)-formylglycine (Fgly), required at the active site of many sulfatases (Sardiello et al. 2005). Other invariant active site residues included 334Asp/335His, which are likely to be involved in Ca2+ binding, based on predictions derived from 3D structures from other human sulfatases (Bond et al. 1997; Hernandez-Guzman et al. 2003). An internal proteolytic cleavage has been proposed for this enzyme as a result of the presence of 42- and 14-kD polypeptides in enzyme preparations derived from human liver, kidney, lung and placenta extracts (Wilson et al. 1990) (Fig. 1). It should be noted that the 42kD polypeptide contains the N-terminal sequence with all of the active site regions, whereas the 14kD polypeptide contained the catalytically inactive C-terminus region of human IDS.
Fig. 2.
Amino acid sequence alignments for mammalian IDS N-terminus sequences. See Table 1 for sources of IDS sequences; asterisk shows identical residues for IDS subunits; colon similar alternate residues; dot dissimilar alternate residues; the active site residues (for human IDS) are shown in blue; leader peptide is in brown; propeptide in red; bold underlined font shows residues corresponding to known or predicted exon start sites; exon numbers refer to human IDS gene exons
Five N-glycosylation sites were consistently found for vertebrate IDS sequences (human IDS amino acid sequences identified in each case): Asn115-Phe116-Ser117 (site 1); Asn144-His145-Thr173 (site 2); Asn246-Ile247-Thr248 (site 3); Asn280-Ile281-Ser282 (site 4); and Asn513-Phe514-Ser515 (site 5). Two other N-glycosylation sites were observed for human IDS which were not commonly shared with other vertebrate IDS sequences, including Asn325-Ser326-Ser327 (site 6) and Asn537-Asp538-Ser539 (site 7), the latter restricted to mammalian IDS sequences (Fig. 1; Table 1). Mutation analysis of the human IDS gene has shown that amino acid substitution of Asn115 (Asn→Tyr) (for site 1) resulted in Hunter’s disease, reflecting the key role of this N-glycosylation site in supporting the structure of this enzyme (Vafiadaki et al. 1998). Figure 1 also shows predicted phosphosites sites that may contribute to regulating downstream cellular processes, molecular functions and protein–protein interactions (Hornbeck et al. 2015). Five of these were strictly conserved among the vertebrate IDS sequences examined (human IDS residues: Ser282; Try285; Thr409; Tyr490; and Tyr497) supporting a role for these residues, as yet unknown.
Predicted secondary and tertiary structures for vertebrate IDS
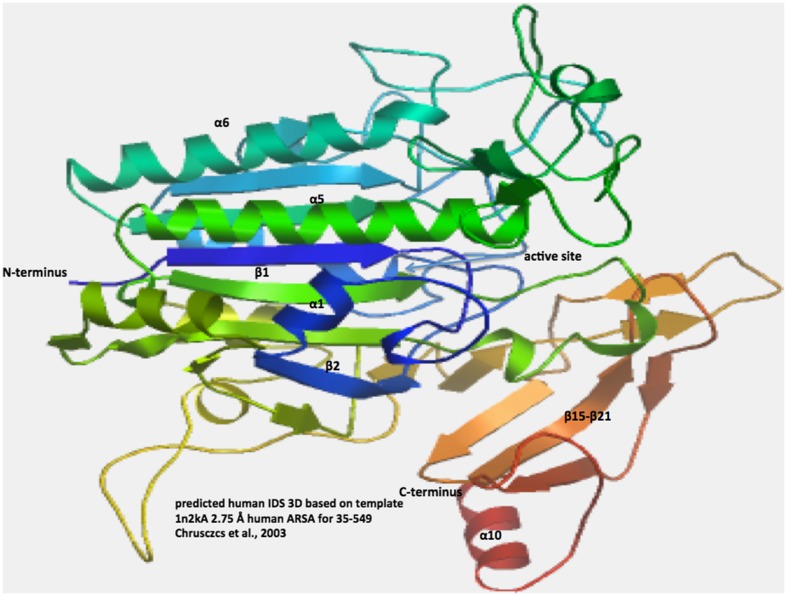
A predicted secondary structure for the human IDS sequence was examined (Fig. 1) using the known structure reported for human ARSA (Lukatela et al. 1998). Ten predicted α-helix and 21 β-sheet structures were observed for human IDS. Of particular interest were β-sheet structures (β1 and β11) and α-helix (α2) which were located proximate to the predicted active site residues for human IDS. The C-terminal end of human IDS contained a sequence of β-sheet structures (β15–β21), in addition to the α-helix (α10) located at the C-terminus. A predicted tertiary structure for human IDS is shown in Fig. 3. Two major domains for this enzyme were observed, that enclose a large cavity previously shown to contain the enzyme’s active site. The more N-terminal of these domains contained the active site residues and comprised the bulk of the 42kD polypeptide chain previously reported (Wilson et al. 1990), whereas the other domain comprised most of the 14kD polypeptide, including the β-sheet structures (β15–β21) and the C-terminal α-helix (α10).
Fig. 3.
Predicted tertiary structure for human IDS. The predicted structure for human IDS is based on the reported structure for human ARSA (Chrusczcz et al. 2003) and obtained using the SWISS MODEL web site based on PDB 1N2KA http://swissmodel.expasy.org/workspace/. The rainbow color code describes the 3-D structures from the N- (blue) to C-termini (red color) for residues 35–549 for human IDS; predicted α-helices, β-sheets, proposed active site cleft, and N- and C-termini are shown
Comparative human IDS tissue expression
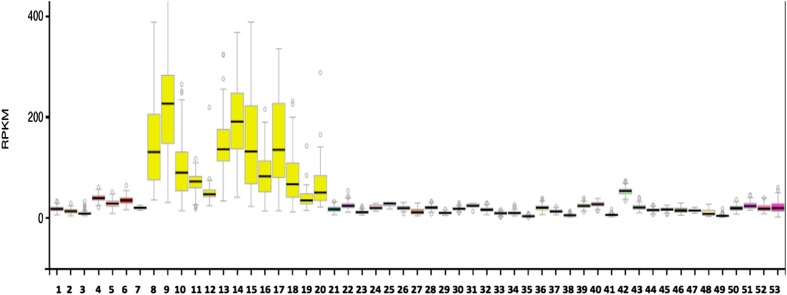
Figure 4 shows comparative gene expression for various human tissues obtained from RNA-seq gene expression profiles for the human IDS gene obtained for 53 selected tissues or tissue segments for 175 individuals (GTEx Consortium 2015) (Data Source: GTEx Analysis Release V6p (dbGaP Accession phs000424.v6.p1) (http://www.gtex.org). These data supported high levels of gene expression for human IDS in regions of the brain, particularly within the cortex, amygdala, hippocampus, hypothalamus and basal ganglia, but with lower levels in the brain cerebellum and spinal cord. IDS activity was also widely distributed at low levels among all other tissues examined. It is readily apparent that IDS is predominantly expressed in brain and nerve tissues of the body, which may reflect a specific role for IDS in neural glycoaminoglycan (GAG) metabolism, involving the efficient clearance of GAG sulfate residues within the extracellular matrix of nervous tissue.
Fig. 4.
Tissue expression for human IDS. RNA-seq gene expression profiles across 53 selected tissues (or tissue segments) were examined from the public database for human IDS, based on expression levels for 175 individuals (Data Source: GTEx Analysis Release V6p (dbGaP Accession phs000424.v6.p1) (http://www.gtex.org). Tissues: 1. Adipose-Subcutaneous; 2. Adipose-Visceral (Omentum); 3. Adrenal gland; 4. Artery-Aorta; 5. Artery-Coronary; 6. Artery-Tibial; 7. Bladder; 8. Brain-Amygdala; 9. Brain-Anterior cingulate Cortex (BA24); 10. Brain-Caudate (basal ganglia); 11. Brain-Cerebellar Hemisphere; 12. Brain-Cerebellum; 13. Brain-Cortex; 14. Brain-Frontal Cortex; 15. Brain-Hippocampus; 16. Brain-Hypothalamus; 17. Brain-Nucleus accumbens (basal ganglia); 18. Brain-Putamen (basal ganglia); 19. Brain-Spinal Cord (cervical c-1); 20. Brain-Substantia nigra; 21. Breast-Mammary Tissue; 22. Cells-EBV-transformed lymphocytes; 23. Cells-Transformed fibroblasts; 24. Cervix-Ectocervix; 25. Cervix-Endocervix; 26. Colon-Sigmoid; 27. Colon-Transverse; 28. Esophagus-Gastroesophageal Junction; 29. Esophagus- Mucosa; 30. Esophagus-Muscularis; 31. Fallopian Tube; 32. Heart-Atrial Appendage; 33. Heart-Left Ventricle; 34. Kidney-Cortex; 35. Liver; 36. Lung; 37. Minor Salivary Gland; 38. Muscle-Skeletal; 39. Nerve-Tibial; 40. Ovary; 41. Pancreas; 42. Pituitary; 43. Prostate; 44. Skin-Not Sun Exposed (Suprapubic); 45. Skin-Sun Exposed (Lower leg); 46. Small Intestine-Terminal Ileum; 47. Spleen; 48. Stomach; 49. Testis; 50. Thyroid; 51. Uterus; 52. Vagina; 53. Whole Blood
Gene locations, exonic structures and regulatory sequences for vertebrate IDS genes
Table 2 summarizes the predicted locations for vertebrate and fruit fly (Drosophila melanogaster) IDS genes based upon BLAT interrogations of several genomes using the reported sequence for human IDS (Bielicki et al. 1990; Wilson et al. 1990) and the predicted sequences for other IDS enzymes and the UCSC genome browser (Kent et al. (2002)). The predicted vertebrate IDS genes were transcribed on both the negative strand (primates, mouse, rat, cow, marsupial and zebra fish genomes) and the positive strand (sheep, chicken, lizard and frog genomes). Of particular interest is the X-chromosome location for IDS for all eutherian and marsupial mammals examined with the exception of rat Ids gene, which is located on an autosome (chromosome 8). This is indicative of a chromosomal transfer between the common ancestral X-chromosome and chromosome 8 during rat evolution. An IDS pseudogene (designated as IDSP1) was also observed for human and other primate genomes. Figure 1 summarizes the predicted exonic start sites for human, mouse, frog and zebra fish IDS genes with each having 9 coding exons, in identical or similar positions to those predicted for the human IDS gene. In each case, exon 1 encoded the leader peptide and propeptide with exons 2, 3 and 7 encoding the predicted active site regions for this enzyme.
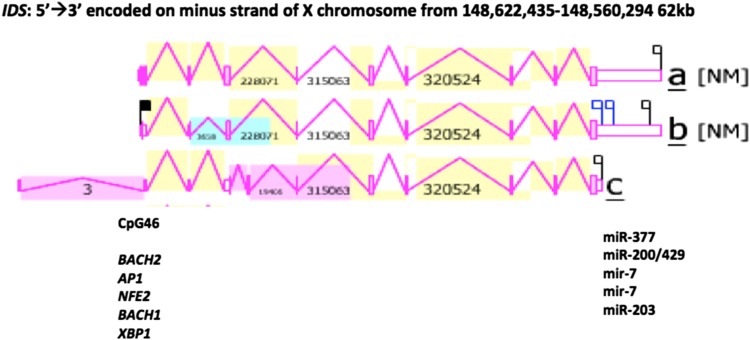
Figure 5 shows the predicted structures for the three major human IDS transcripts (IDSa; IDSb; and IDSc) together with CpG46 and several transcription factor binding sites (TFBS), which are located at the 5′ end of the gene, consistent with roles in regulating the transcription of this gene and forming part of the IDS gene promoter. The human IDSa transcript was 6088 bps in length with an extended 3′-untranslated region (UTR) containing 5 microRNA target sites; the human IDSb transcript was 5808 bps in length, also containing 5 microRNA target sites; whereas the IDSc transcript was much shorter in length (2213 bps), comprising only 8 coding exons and with no microRNA target sites present. The presence of miR-200 within the 3′-UTR of the human IDS gene was of special interest due to this miR family being induced and having a specific role during the late stages of neuronal differentiation (Beclin et al. 2016). In addition, the presence of miR-7 in this region may also be significant given that miR-7 inhibits neuronal apoptosis in a cellular Parkinson’s disease model (Li et al. 2016) and contributes to the alteration of neuronal morphology and function (Zhang et al. 2015). Moreover, miR-203 has a proposed role as a stemness inhibitor of glioblastoma stem cells and may contribute to the increased expression of glial and neuronal differentiation markers (Deng et al. 2016).
Fig. 5.
Gene structure and major gene transcripts for the human IDS gene. Derived from the AceView website http://www.ncbi.nlm.nih.gov/IEB/Research/Acembly/ (Thierry-Mieg and Thierry-Mieg 2006); shown with capped 5′- and 3′- ends for the predicted mRNA sequences; NM refers to the NCBI reference sequence; coding exons are in pink; the direction for transcription is shown as 5′ → 3′; a large CpG46 island at the gene promoter is shown (see Table 4 for details of CpG islands for human and other vertebrate IDS gene promoters); 5 predicted transcription factor binding sites (TFBS) for human IDS are shown (see Table 1s for details); 5 predicted miRNA target sites were identified within the extended 3′-UTR region of human IDSa and IDSb transcripts
The human IDS genome sequence also contained several predicted transcription factor binding sites (TFBS) and a large CpG island (CpG46) located in the 5′-untranslated promoter region of human IDS on the X-chromosome. CpG46 contained 432 bps with a C plus G count of 279 bps, a C or G content of 65% and showed a ratio of observed to expected CpG of 1.02. Similar CpG islands were observed in the IDS gene promoters for other primate, eutherian mammal, marsupial (opossum) and bird (chicken) genomes (Table 3). It is likely therefore that these IDS CpG islands play a key role in regulating this gene and may contribute to the very high level of gene expression observed in neural tissues (Fig. 4) (Saxanov et al. 2006). At least 5 TFBS sites were colocated with CpG46 in the human IDS promoter region which may contribute to the high expression of this gene in human nerve and brain tissues (Table 4). Of special interest among these transcription factor binding sites were the following: BACH1 and BACH2 have been recognized as members of the BTB-basic region leucine zipper transcription factor family which downregulate cell proliferation of neuroblastoma cells (Shim et al. 2006); AP1 is constitutively upregulated in activated microglia and during the pathogenesis of Parkinson’s disease (Pal et al. 2016); NFE2 has been shown to participate in the developmental regulation of the brain in zebrafish embryos (Williams et al. 2013); and XBP1 has been identified as a risk factor for Alzheimer’s disease and bipolar disorders, contributing to impairment of contextual memory formation (Martinez et al. 2016).
Table 3.
Vertebrate IDS CpG Islands
| Vertebrate | CpG Island ID | Chromosomal position | CpG size | C count plus G count | % C or G | Ratio of observed to expected CpG |
|---|---|---|---|---|---|---|
| Human | CpG 46 | ChrX:148,586,553–148,586,984 | 432 | 279 | 65 | 1.02 |
| Baboon | CpG 50 | ChrX:137,263,406–137,263,837 | 432 | 306 | 71 | .92 |
| Rhesus | CpG53 | ChrX:143,222,778–143,223,221 | 444 | 318 | 72 | .93 |
| Mouse | CpG 26 | ChrX:70,364,872–70,365,161 | 290 | 159 | 55 | 1.2 |
| Rat | CpG 26 | Chr8:69,175,527–69,175,735 | 209 | 138 | 66 | 1.14 |
| Cow | CpG 53 | ChrX:32,324,232–32,324,656 | 425 | 317 | 75 | .9 |
| Dog | CpG 51 | ChrX;117,515,293–117,515,743 | 451 | 293 | 65 | 1.1 |
| Opossum | CpG2 29 | ChrX:38,797,675–38,797,993 | 319 | 189 | 59 | 1.05 |
| Chicken | CpG 54 | Chr4:18,031,448–18,032,009 | 562 | 333 | 59 | 1.09 |
The identification of IDS CpG islands, sequences and properties was undertaken using various vertebrate genome browsers (http://genome.ucsc.edu)
Table 4.
Identification of transcription factor binding sites (TFBS) within the human IDS gene promoter
| TFBS | Strand | Chr 1 Position | Function/role | Sequence | UNIPROT ID |
|---|---|---|---|---|---|
| BACH2 | (+ve) | X:148,585,129–139 | Binds to Maf recognition elemants |
GCTGAGTCATG |
Q9BYV9 |
| AP1 | (−ve) | X:148,585,128–140 | Regulating cells forming the skeleton |
GCATGACTCAGCT |
P01101 |
| NFE2 | (+ve) | X:148,585,128–138 | Regulating erythroid maturation | AGCTGAGTCAT | Q16621 |
| BACH1 | (+ve) | X:148,585,127–141 | Coordinates transcription by MAFK | TAGCTGAGTCATGCA | O14867 |
| XBP1 | (+ve) | X:148,584,868–884 | Regulation during ER stress | ATGGTCACATAGCCATT | P17861 |
The identification of TFBS within the IDS promoter region was undertaken using the human genome browser (http://genome.ucsc.edu); UNIPROT refers to UniprotKB/Swiss-Prot IDs for individual TFBS sequences (see http://kr.expasy.org); ER refers to endoplasmic reticulum
Phylogeny and divergence of vertebrate IDS
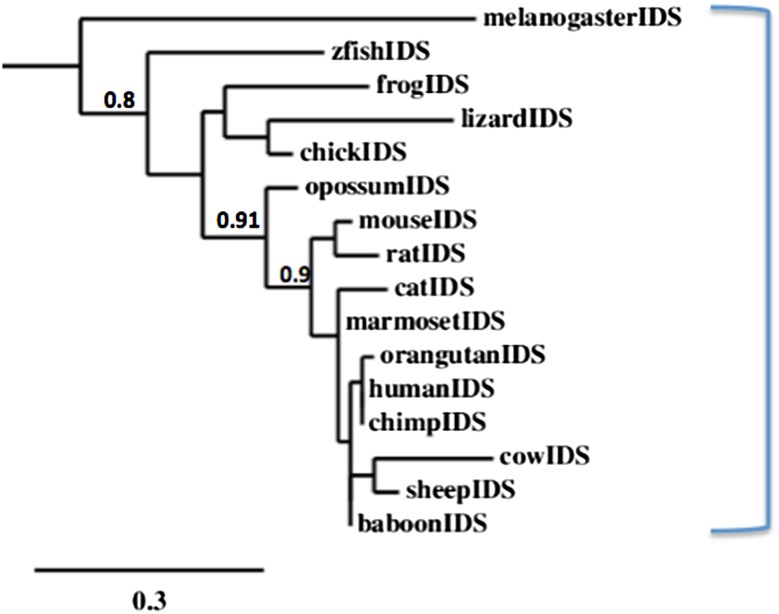
A phylogenetic tree (Fig. 6) was calculated by the progressive alignment of 15 vertebrate IDS amino acid sequences with several other human ARS-like sequences (see Table 3). The IDS phylogram was ‘rooted’ with the fruit fly (Drosophila melanogaster) IDS sequence (see Table 1). The phylogram showed clustering of the IDS sequences into a single group which is represented throughout vertebrate evolution and has apparently evolved from an invertebrate IDS gene ancestor.
Fig. 6.
Phylogenetic tree of vertebrate IDS amino acid sequences. The tree is labeled with the vertebrate and fruit fly IDS. A genetic distance scale is shown. The number of times a clade (sequences common to a node or branch) occurred in the bootstrap replicates are shown. Replicate values of .9 or more which are highly significant (values of .9 or more), are shown with 100 bootstrap replicates performed in each case
Conclusions
The current results indicate that vertebrate IDS genes and encoded proteins represent a distinct gene and protein family of ARS-like proteins. IDS has a distinct property among human arylsulfatases in being responsible for the lysosomal degradation of the glycoaminoglycans, heparan sulfate and dermatan sulfate, by hydrolysing 2-sulfate groups of the l-iduronate 2-sulfate units (Bielicki et al. 1990). IDS is encoded by a single gene among the vertebrate genomes examined and is highly expressed in human brain and other nerve tissues, and contained 9 coding exons on the negative strand of the human genome. Primate genomes contained an IDS pseudogene (IDSP1) located in a proximal position on the X-chromosome. The promoter region of the human IDS gene contained a large CpG island together with at least 5 TFBS, which may contribute to the high level of gene expression in the brain. In addition, 5 microRNA target sites were observed within the extended 3′-UTR of the human IDS gene which may be implicated in regulating gene expression during brain development. Predicted secondary and tertiary structures for human IDS showed strong similarities with other ARS-like proteins. Several major structural domains were apparent for mammalian IDS, including the N-terminal leader peptide and propeptide regions; the active site (including a calcium binding site), which is responsible for arylsulfatase activity; and five conserved N-glycosylation sites. Phylogenetic studies using 15 vertebrate and one invertebrate (Drosophila melanogaster) IDS sequences indicated that the IDS gene has appeared early in evolution, prior to the appearance of bony fish.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Table 1s: Percentage Amino Acid Sequence Identities for Human Arylsulfatase Proteins. Amino acid sequences for the human ARS enzymes were derived from the following: IDS (iduronate 2-sulfatase: UNIPROT: P22304); ARSA (arylsulfatase A: UNIPROT P15289); ARSB (arylsulfatase B: UNIPROT: P15848); STS (sterylsulfatase: UNIPROT P08842); ARSK (arylsulfatase K: UNIPROT: Q6UWY0); SGSH (heparan N-sulfatase: UNIPROT: P51688); SULF1 (extracellular Sulfatase 1: UNIPROT: Q8IWU6); SUMF1 (sulfatase modifying factor 1: UNIPROT: Q8NBK3). (XLSX 45 kb)
Abbreviations
- IDS
Iduronate 2-sulfatase
- GAG
Glycoaminoglycan
- ARS
Arylsulfatase
- ARSA
Arylsulfatase A
- kbps
Kilobase pairs
- CpG island
Multiple C (cytosine)-G (guanine) dinucleotide region
- miRNA
MicroRNA binding region
- MPS
Mucopolysaccharidosis
- BLAST
Basic local alignment search tool
- BLAT
Blast-like alignment tool
- NCBI
National Center for Biotechnology Information
- SWISS-MODEL
Automated protein structure homology-modeling server
Compliance with ethical standards
Conflict of interest
The author declares that he has no conflicts of interest.
Footnotes
Electronic supplementary material
The online version of this article (doi:10.1007/s13205-016-0595-3) contains supplementary material, which is available to authorized users.
References
- Anekar J, Deepa Narayanan C, Raj AC, Sandeepa NC, Nappalli D. A rare case of mucopolysaccharidosis: Hunter syndrome. J Clin Diagn Res. 2015;9:ZD23-6. doi: 10.7860/JCDR/2015/13251.5858. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beck M. Mucopolysaccharidosis Type II (Hunter Syndrome): clinical picture and treatment. Curr Pharm Biotechnol. 2011;12:861–866. doi: 10.2174/138920111795542714. [DOI] [PubMed] [Google Scholar]
- Beclin C, Follert P, Stappers E, Barral S, Nathalie C, de Chevigny A, Magnone V, Lebrigand K, Bissels U, Huylebroeck D, Bosio A, Barbry P, Seuntjens E, Cremer H. miR-200 family controls late steps of postnatal forebrain neurogenesis via Zeb2 inhibition. Sci Rep. 2016;6:35729. doi: 10.1038/srep35729. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bielicki J, Freeman C, Clements PR, Hopwood JJ. Human liver iduronate-2-sulfatase. Purification, characterization and catalytic properties. Biochem J. 1990;271:75–86. doi: 10.1042/bj2710075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bond CS, Clements PR, Ashby SJ, Collyer CA, Harrop SJ, Hopwood JJ, Guss JM. Structure of a human lysosomal sulfatase. Structure. 1997;5:277–289. doi: 10.1016/S0969-2126(97)00185-8. [DOI] [PubMed] [Google Scholar]
- Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL. BLAST+: architesture and applications. BMC Bioinform. 2009;10:421. doi: 10.1186/1471-2105-10-421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chistiakov DA, Kuzenkova LM, Savost’anov KV, Gevorkyan AK, Pushkov AA, Nikitin AG, Vashakmadze ND, Zhurkova NV, Podkletnova TV, Namazova-Baranova LS, Baranov AA. Genetic analysis of 17 children with Hunter syndrome: identification and functional characterization of four novel mutations in the iduronate-2-sulfatase gene. J Genet Genom. 2014;41:197–203. doi: 10.1016/j.jgg.2014.01.007. [DOI] [PubMed] [Google Scholar]
- Cho SY, Lee J, Ko AR, Kwak MJ, Kim S, Sohn YB, Park SW, Jin DK. Effect of systemic high dose enzyme replacement therapy on the improvement of CNS defects in a mouse model of mucopolysaccharidosis type II. Orphanet J Rare Dis. 2015;10:141. doi: 10.1186/s13023-015-0356-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chruszcz M, Laidler P, Monkiewicz M, Ortlund E, Lebioda L, Lewinski K. Crystal structure of a covalent intermediate of endogenous human arylsulfatase A. J Inorg Biochem. 2003;96:386–392. doi: 10.1016/S0162-0134(03)00176-4. [DOI] [PubMed] [Google Scholar]
- Deng Y, Zhu G, Luo H, Zhao S. MicroRNA-203 As a Stemness Inhibitor of Glioblastoma Stem Cells. Mol Cells. 2016;39:619–624. doi: 10.14348/molcells.2016.0118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dereeper A, Guignon V, Blanc G, Audic S, Buffet S, Chevenet F, Dufayard JF, Guindon S, Lefort V, Lescot M, Claverie JM, Gascuel O. Phylogeny.fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res. 2008;36:W465–W469. doi: 10.1093/nar/gkn180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fusar Poli E, Zalfa C, D’Avanzo F, Tomanin R, Carlessi L, Bossi M, Nodari LR, Binda E, Marmiroli P, Scarpa M, Delia D, Vescovi AL, De Filippis L. Murine neural stem cells model Hunter disease in vitro: glial cell-mediated neurodegeneration as a possible mechanism involved. Cell Death Dis. 2013;4:e906. doi: 10.1038/cddis.2013.430. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garcia AR, Pan J, Lamsa JC, Muenzer J. The characterization of a murine model of mucopolysaccharidosis II (Hunter syndrome) J Inherit Metab Dis. 2007;30:924–934. doi: 10.1007/s10545-007-0641-8. [DOI] [PubMed] [Google Scholar]
- GTEx Consortium Human genomics. The genotype-tissue expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science. 2015;348:648–660. doi: 10.1126/science.1262110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hernandez-Guzman FG, Higashiyama T, Pangborn W, Osawa Y, Ghosh D. Structure of human estrone sulfatase suggests functional roles of membrane association. J Biol Chem. 2003;278:22989–22997. doi: 10.1074/jbc.M211497200. [DOI] [PubMed] [Google Scholar]
- Holmes RS. Comparative and evolutionary studies of vertebrate arylsulfatase B, arylsulfatase I and arylsulfatase J genes and proteins: evidence for an ARSB-like sub-family. J Prot Bioinform. 2016;9:11. [Google Scholar]
- Hornbeck PV, Zhang B, Murray B, Kornhauser JM, Latham V, Skrzypek E. PhosphoSitePlus, 2014: mutations, PTMs and recalibrations. Nucleic Acids Res. 2015;43:D512–D520. doi: 10.1093/nar/gku1267. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM, Haussler D. The human genome browser at UCSC. Genome Res. 2002;12:994–1006. doi: 10.1101/gr.229202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kosuga M, Mashima R, Hirakiyama A, Fuji N, Kumagai T, Seo JH, Nikaido M, Saito S, Ohno K, Sakuraba H, Okuyama T. Molecular diagnosis of 65 families with mucopolysaccharidosis type II (Hunter syndrome) characterized by 16 novel mutations in the IDS gene: genetic, pathological, and structural studies on iduronate-2-sulfatase. Mol Genet Metab. 2016;118:190–197. doi: 10.1016/j.ymgme.2016.05.003. [DOI] [PubMed] [Google Scholar]
- Li S, Lv X, Zhai K, Xu R, Zhang Y, Zhao S, Qin X, Yin L, Lou J. MicroRNA-7 inhibits neuronal apoptosis in a cellular Parkinson’s disease model by targeting Bax and Sirt2. Am J Transl Res. 2016;8:993–1004. [PMC free article] [PubMed] [Google Scholar]
- Lukatela G, Krauss N, Theis K, Selmer T, Gieselmann V, von Figura K, Saenger W. Crystal structure of human arylsulfatase A: the aldehyde function and the metal ion at the active site suggest a novel mechanism for sulfate ester hydrolysis. Biochemistry. 1998;37:3654–3664. doi: 10.1021/bi9714924. [DOI] [PubMed] [Google Scholar]
- Marchler-Bauer A, Lu S, Anderson JB, Chitsaz F, Derbyshire MK, DeWeese-Scott C, Fong JH, Geer LY, Geer RC, Gonzales NR, Gwadz M, Hurwitz DI, Jackson JD, Ke Z, Lanczycki CJ, Lu F, Marchler GH, Mullokandov M, Omelchenko MV, Robertson CL, Song JS, Thanki N, Yamashita RA, Zhang D, Zhang N, Zheng C, Bryant SH. CDD: a conserved domain database for the functional annotation of proteins. Nucleic Acid Res. 2011;39:D225–D229. doi: 10.1093/nar/gkq1189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martínez G, Vidal RL, Mardones P, Serrano FG, Ardiles AO, Wirth C, Valdés P, Thielen P, Schneider BL, Kerr B, Valdés JL, Palacios AG, Inestrosa NC, Glimcher LH, Hetz C. Regulation of memory formation by the transcription factor XBP1. Cell Rep. 2016;14:1382–1394. doi: 10.1016/j.celrep.2016.01.028. [DOI] [PubMed] [Google Scholar]
- Moro E, Tomanin R, Friso A, Modena N, Tiso N, Scarpa M, Argenton F. A novel functional role of iduronate-2-sulfatase in zebrafish early development. Matrix Biol. 2010;29:43–50. doi: 10.1016/j.matbio.2009.09.001. [DOI] [PubMed] [Google Scholar]
- Muenzer J, Hendriksz CJ, Fan Z, Vijayaraghavan S, Perry V, Santra S, Solanki GA, Mascelli MA, Pan L, Wang N, Sciarappa K, Barbier AJ. A phase I/II study of intrathecal idursulfase-IT in children with severe mucopolysaccharidosis II. Genet Med. 2016;18:73–81. doi: 10.1038/gim.2015.36. [DOI] [PubMed] [Google Scholar]
- Pal R, Tiwari PC, Nath R, Pant KK. Role of neuroinflammation and latent transcription factors in pathogenesis of Parkinson’s disease. Neurol Res. 2016;3:1–12. doi: 10.1080/01616412.2016.1249997. [DOI] [PubMed] [Google Scholar]
- Parini R, Rigoldi M, Tedesco L, Boffi L, Brambilla A, Bertoletti S, Boncimino A, Del Longo A, De Lorenzo P, Gaini R, Gallone D, Gasperini S, Giussani C, Grimaldi M, Grioni D, Meregalli P, Messinesi G, Nichelli F, Romagnoli M, Russo P, Sganzerla E, Valsecchi G, Biondi A. Enzymatic replacement therapy for Hunter disease: up to 9 years experience with 17 patients. Mol Genet Metab Rep. 2015;3:65–74. doi: 10.1016/j.ymgmr.2015.03.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rathmann M, Bunge S, Beck M, Kresse H, Tylki-Szymanska A, Gal A. Mucopolysaccharidosis type II (Hunter syndrome): mutation “hot spots” in the iduronate-2-sulfatase gene. Am J Hum Genet. 1996;59:1202–1209. [PMC free article] [PubMed] [Google Scholar]
- Ratzka A, Mundlos S, Vortkamp A. Expression patterns of sulfatase genes in the developing mouse embryo. Dev Dyn. 2010;239:1779–1788. doi: 10.1002/dvdy.22294. [DOI] [PubMed] [Google Scholar]
- Rivera-Colón Y, Schutsky EK, Kita AZ, Garman SC. The structure of human GALNS reveals the molecular basis for mucopolysaccharidosis IV A. J Mol Biol. 2012;423:736–751. doi: 10.1016/j.jmb.2012.08.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sáenz H, Lareo L, Poutou RA, Sosa AC, Barrera LA. Computational prediction of the tertiary structure of the human iduronate 2-sulfate sulfatase. Biomedica. 2007;27:7–20. doi: 10.7705/biomedica.v27i1.229. [DOI] [PubMed] [Google Scholar]
- Sardiello M, Annunziata I, Roma G, Ballabio A. Sulfatases and sulfatase modifying factors: an exclusive and promiscuous relationship. Hum Mol Genet. 2005;14:3203–3217. doi: 10.1093/hmg/ddi351. [DOI] [PubMed] [Google Scholar]
- Saxonov S, Berg P, Brutlag DL. A genome-wide analysis of CpG dinucleotides in the human genome distinguishes two distinct classes of promoters. Proc Natl Acad Sci USA. 2006;103:1412–1417. doi: 10.1073/pnas.0510310103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schwede T, Kopp J, Guex N, Pietsch MC. SWISS-MODEL: an automated protein homology-modelling server. Nucleic Acids Res. 2003;31:3381–3385. doi: 10.1093/nar/gkg520. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shim KS, Rosner M, Freilinger A, Lubec G, Hengstschläger M. Bach2 is involved in neuronal differentiation of N1E−115 neuroblastoma cells. Exp Cell Res. 2006;312:2264–2278. doi: 10.1016/j.yexcr.2006.03.018. [DOI] [PubMed] [Google Scholar]
- Sievers F, Higgins DG. Clustal omega. Curr Protoc Bioinform. 2014;2014(48):1–16. doi: 10.1002/0471250953.bi0313s48. [DOI] [PubMed] [Google Scholar]
- Smith CM, Finger JH, Hayamizu TF, McCright IJ, Xu J, Berghout J, Campbell J, Corbani LE, Forthofer KL, Frost PJ, Miers D, Shaw DR, Stone KR, Eppig JT, Kadin JA, Richardson JE, Ringwald M. The mouse gene expression database (GXD): 2014 update. Nucleic Acids Res. 2014;42(D1):D818–D824. doi: 10.1093/nar/gkt954. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thierry-Mieg D, Thierry-Mieg J (2006) AceView: a comprehensive cDNA-supported gene and transcripts annotation. Genome Biol. 7(Suppl 1) S12:1-14 [DOI] [PMC free article] [PubMed]
- Tylki-Szymańska A. Mucopolysaccharidosis type II, Hunter’s syndrome. Pediatr Endocrinol Rev. 2014;12(Suppl 1):107–113. [PubMed] [Google Scholar]
- Vafiadaki E, Cooper A, Heptinstall LE, Hatton CE, Thornley M, Wraith JE. Mutation analysis in 57 unrelated patients with MPS II (Hunter’s disease) Arch Dis Child. 1998;79:237–241. doi: 10.1136/adc.79.3.237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williams LM, Timme-Laragy AR, Goldstone JV, McArthur AG, Stegeman JJ, Smolowitz RM, Hahn ME. Developmental expression of the Nfe2-related factor (Nrf) transcription factor family in the zebrafish, Danio rerio. PLoS One. 2013;8:e79574. doi: 10.1371/journal.pone.0079574. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilson PJ, Morris CP, Anson DS, Occhiodoro T, Bielicki J, Clements PR, Hopwood JJ. Hunter syndrome: isolation of an iduronate-2-sulfatase cDNA clone and analysis of patient DNA. Proc Natl Acad Sci USA. 1990;87:8531–8535. doi: 10.1073/pnas.87.21.8531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilson PJ, Meaney CA, Hopwood JJ, Morris CP. Sequence of the human iduronate 2-sulfatase (IDS) gene. Genomics. 1993;17(3):773–775. doi: 10.1006/geno.1993.1406. [DOI] [PubMed] [Google Scholar]
- Zhang J, Sun XY, Zhang LY. MicroRNA-7/Shank3 axis involved in schizophrenia pathogenesis. J Clin Neurosci. 2015;22:1254–1257. doi: 10.1016/j.jocn.2015.01.031. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary Table 1s: Percentage Amino Acid Sequence Identities for Human Arylsulfatase Proteins. Amino acid sequences for the human ARS enzymes were derived from the following: IDS (iduronate 2-sulfatase: UNIPROT: P22304); ARSA (arylsulfatase A: UNIPROT P15289); ARSB (arylsulfatase B: UNIPROT: P15848); STS (sterylsulfatase: UNIPROT P08842); ARSK (arylsulfatase K: UNIPROT: Q6UWY0); SGSH (heparan N-sulfatase: UNIPROT: P51688); SULF1 (extracellular Sulfatase 1: UNIPROT: Q8IWU6); SUMF1 (sulfatase modifying factor 1: UNIPROT: Q8NBK3). (XLSX 45 kb)