Significance
Here, we demonstrate that the cytokine activin-A instructs the differentiation of human IL-10–producing type 1 regulatory T (Tr1)-like cells that exhibit strongly suppressive functions against allergen-induced naive and effector CD4+ T-cell responses. In addition, we show that activin-A induces the activation of interferon regulatory factor (IRF4), which, along with aryl hydrocarbon receptor (AhR) and its binding partner, AhR nuclear translocator, forms a tripartite transcription factor complex that is essential for the differentiation and effector functions of human Tr1 cells. Importantly, administration of human activin-A–induced Tr1 cells in a humanized model of asthma confers protection against cardinal disease manifestations in preventive and therapeutic regimes. Collectively, our studies unravel a biological function for activin-A in the generation of suppressive human Tr1 cells that may be exploited for the control of allergic diseases.
Keywords: cytokines, allergic inflammation, immune regulation, activin-A, human T regulatory cells
Abstract
Type 1 regulatory T (Tr1) cells play a pivotal role in restraining human T-cell responses toward environmental allergens and protecting against allergic diseases. Still, the precise molecular cues that underlie their transcriptional and functional specification remain elusive. Here, we show that the cytokine activin-A instructs the generation of CD4+ T cells that express the Tr1-cell–associated molecules IL-10, inducible T-Cell costimulator (ICOS), lymphocyte activation gene 3 protein (LAG-3), and CD49b, and exert strongly suppressive functions toward allergic responses induced by naive and in vivo-primed human T helper 2 cells. Moreover, mechanistic studies reveal that activin-A signaling induces the activation of the transcription factor interferon regulatory factor (IRF4), which, along with the environmental sensor aryl hydrocarbon receptor, forms a multipartite transcriptional complex that binds in IL-10 and ICOS promoter elements and controls gene expression in human CD4+ T cells. In fact, IRF4 silencing abrogates activin-A–driven IL10 and ICOS up-regulation and impairs the suppressive functions of human activin-A–induced Tr1-like (act-A–iTr1) cells. Importantly, using a humanized mouse model of allergic asthma, we demonstrate that adoptive transfer of human act-A–iTr1 cells, both in preventive and therapeutic protocols, confers significant protection against cardinal asthma manifestations, including pulmonary inflammation. Overall, our findings uncover an activin-A–induced IRF4-aryl hydrocarbon receptor (AhR)–dependent transcriptional network, which generates suppressive human Tr1 cells that may be harnessed for the control of allergic diseases.
Overactive T helper 2 (Th2) cell-driven responses against inhaled allergens contribute to the development of airway inflammation and airway hyperresponsiveness (AHR), and are associated with allergic asthma, a rising health care problem worldwide (1). Regulatory T (Treg) cells and, specifically, IL-10–producing type 1 Treg (Tr1) cells exert crucial nonredundant functions in restraining exuberant Th2 responses and maintaining tolerance in mouse models of experimental asthma (2–5). Moreover, in humans, the balance between allergen-specific Tr1 and Th2 cells is considered decisive for the outcome of allergic responses (6). In fact, the tolerogenic effects of allergen-specific immunotherapies are associated with the generation of Tr1 cells (7). Hence, the identification of factors that can induce and/or expand human Tr1 cells represents an attractive therapeutic approach for the control of allergic responses and linked asthmatic disease.
Despite recent advances in deciphering the human Tr1-cell transcriptional network, the molecular cues that control the differentiation of Tr1 cells remain largely unknown (8, 9). Seminal studies have demonstrated that aryl hydrocarbon receptor (AhR), along with the c-Maf proto-oncogene transcription factor (c-Maf), participate in human Tr1-cell generation; still, these transcription factors also contribute to the differentiation of other Th subsets, and thus cannot be considered as Tr1-cell lineage specification factors (8–10). Moreover, the precise molecular pathways underlying the suppressive functions of human Tr1 cells remain incompletely defined.
Activin-A is a member of the TGF-β superfamily, initially described as a gonadal protein stimulating the biosynthesis and secretion of follicle-stimulating hormone (11). Since then, a growing body of evidence has uncovered critical functions for activin-A as a cytokine in fundamental biological processes, including development, hematopoiesis, tissue repair, stem cell maintenance, and differentiation (12). In line with this evidence, mice lacking the βA subunit of activin-A die after birth due to severe craniofacial defects, whereas mice lacking the βB subunit exhibit abnormal development and reduced reproductive capacity (13, 14). Activin-A signals through a heterotetrameric receptor complex consisting of two types of activin receptors: the type I activin-like kinase (ALK2, ALK4, and ALK7) receptors and the type II activin (ActRIIA and ActRIIB) receptors, which, upon activation, phosphorylate receptor-regulated Smads (R-Smads) that translocate in the cell nucleus and regulate gene transcription (15). Activin-A exerts both beneficial and detrimental effects on allergic inflammation and linked diseases, depending on the route of administration and the spatiotemporal context (16, 17). Our previous studies, using an ovalbumin-induced mouse model of allergic airway inflammation, have revealed that activin-A protects against experimental asthma partly through the generation of Foxp3−IL-10+ Treg cells (18). Moreover, we and others have shown that activin-A and its signaling components are induced in T cells infiltrating the lungs of asthmatics and become further up-regulated following in vivo inhalational allergen challenge (19, 20). Still, the effects of activin-A on the generation of human Treg cells suppressive against allergic responses, and the molecular mechanisms involved, remain unexplored.
Here, using an experimental approach that combines in vitro priming of human naive T cells with ex vivo analysis of T-effector cells, we demonstrate that activin-A instructs the differentiation of human IL-10–producing Tr1-like cells that inhibit allergen-induced T-cell responses. Mechanistic studies show that activin-A signaling induces the activation of IRF4 in human T cells, which binds in IL10 and ICOS promoter elements and controls gene expression. Moreover, our findings reveal that IRF4, along with AhR and its binding partner, AhR nuclear translocator (ARNT), form a transcription factor complex that is essential for effector molecule expression by human Tr1 cells. We further demonstrate that in vivo administration of human activin-A–induced Tr1-like (act-A–iTr1) cells can prevent, and even reverse, established allergic airway inflammation and confer protection against asthma manifestations in an IL-10–dependent manner. Collectively, our studies unravel activin-A as an inducer of human Tr1 cells that may be exploited for the suppression of allergic inflammation and the reestablishment of tolerance.
Results
Activin-A Drives the Differentiation of IL-10–Producing Human Tr1-Like Cells.
To delineate the effects of activin-A on the regulation of human T-cell–driven allergic responses, we used our well-established in vitro T-cell priming model (21, 22). Briefly, we sorted naive CD4+ T cells from the peripheral blood of atopic individuals and stimulated them with allergen-loaded, mitomycin-treated, CD3-depleted antigen-presenting cells (APCs) in the presence of activin-A or control (PBS) (Fig. S1). This protocol has been shown to skew T-cell responses toward Th2 (23). Indeed, control-treated CD4+ T cells (hereafter referred to as Tconv cells) exhibited marked proliferation, concomitant with high percentages of IL-4+ and IL-13+ cells and moderate to low IFN-γ+ (IFN-γ) and IL-17+ cells following allergen exposure, whereas T cells stimulated in the absence of allergen did not exhibit discernible proliferation (Fig. 1 A and B and Fig. S2 A and B). In contrast, activin-A–stimulated T cells displayed a reduced proliferative capacity and significantly decreased percentages of cytokine-producing cells (Fig. 1 A and B and Fig. S2B). Remarkably, activin-A significantly increased the frequencies of CD4+IL-10+ T cells (Fig. 1A and Fig. S2B). In accordance with the flow cytometry data, activin-A–treated human T cells expressed low levels of mRNAs encoding IL-4, IL-13, IL-5, IFN-γ, and IL-17, whereas IL-10 levels were increased (Fig. S2C). Moreover, IL-4, IL-13, and IFN-γ release in culture supernatants was decreased, whereas IL-10 production was significantly elevated in the presence of activin-A (Fig. 1C). Blockade of TGF-β in primary induction cultures did not affect the activin-A–mediated increase in the percentages of CD4+IL-10+ T cells (Fig. S2D).
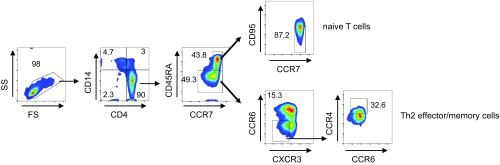
Fig. S1.
Experimental protocol used for the isolation of human CD4+ T-cell subsets. Naive and Th2-effector/memory cells were sorted from the PBMC fraction according to the gating strategy indicated. Representative FACS plots are shown. FS, forward scatter; SS, side scatter.
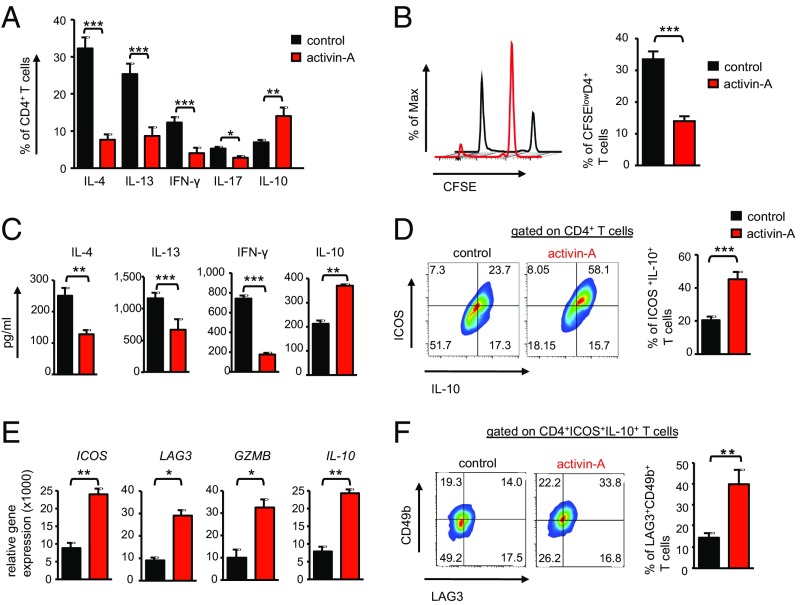
Fig. 1.
Activin-A drives the differentiation of human Tr1-like cells. (A) Human naive CD4+ T cells were sorted from the peripheral blood of atopics; labeled with CFSE; and stimulated with allergen (mixed grasses extract)-loaded, mitomycin-treated, CD3-depleted APCs in the presence of PBS (control) or activin-A for 7 d (or 9 d for IL-10) and analyzed by flow cytometry. The percentages of cytokine-producing T cells are shown. Data are expressed as mean ± SEM and are pooled from n = 8 independent experiments (n = 8 donors). (B, Left) Representative FACS plots showing T-cell proliferation gated on CD4+ T cells. Max, maximum. (B, Right) Cumulative data pooled from n = 6–8 independent experiments (n = 8 donors). (C) Cytokines in culture supernatants are shown. Data are expressed as mean ± SEM of triplicate wells and are pooled from n = 8 independent experiments (n = 8 donors). (D) Representative FACS plots showing ICOS and IL-10 expression gated on CD4+ T cells. Cumulative data are pooled from n = 6–8 independent experiments (n = 8 donors). (E) Real-time PCR analysis of ICOS, LAG3, GZMB, and IL10 mRNA levels in CD4+ T cells stimulated as above for 3 d. Results are presented relative to GAPDH and are pooled from n = 5 separate experiments (n = 5 donors). (F) Representative FACS plots (Left) and percentages (Right) of LAG3+CD49b+ cells among CD4+IL-10+ICOS+ T cells are depicted. Data are pooled from n = 4–6 separate experiments (n = 6 donors). Statistical significance was obtained by the Student’s t test (*P < 0.05; **P < 0.01; ***P < 0.001).
Fig. S2.
Activin-A inhibits allergen-driven effector cytokine release by human CD4+ T cells. Human naive CD4+ T cells were labeled with CFSE and cultured with mitomycin-treated, allergen-loaded, CD3-depleted APCs in the presence of control (PBS) or activin-A for 7 or 9 d (for IL-10). Representative FACS plots show T-cell proliferation in the absence of allergen (A) and cytokine-expressing CD4+ T cells (B) and are representative of n = 8 independent experiments (n = 8 donors). Max, maximum. (C) Real-time PCR analysis of cytokine mRNA expression in CD4+ T cells stimulated as above for 3 d. Results are presented relative to GAPDH and are representative of n = 3 separate experiments (n = 3 donors). (D) Human naive CD4+ T cells were stimulated as in A in the presence or absence of an anti–TGF-β antibody. FACS plots show IL-10–expressing cells and gated CD4+ T cells and represent n = 3 separate experiments (n = 3 donors).
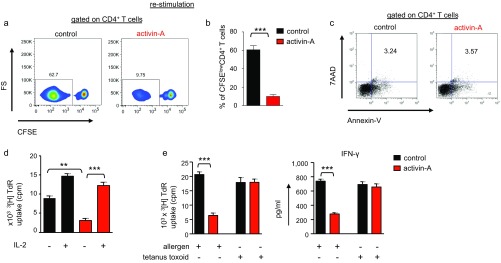
Interestingly, activin-A–treated T cells remained unresponsive upon secondary restimulation with fresh allergen-loaded APCs (Fig. S3 A and B). This hyporesponsiveness was not associated with induction of T-cell apoptosis/necrosis, as reflected by similar frequencies of 7AAD+Annexin-V+CD4+ T cells among activin-A– and control-treated T cells (Fig. S3C). In fact, addition of IL-2 reversed activin-A antiproliferative effects, similar to recent observations on human follicular helper T (Tfh) cells (24) (Fig. S3D). Notably, activin-A–treated CD4+ T cells exhibited Ag specificity because allergen-primed T cells retained their ability to respond upon subsequent exposure to tetanus toxoid, an irrelevant antigen (Fig. S3E). Activin-A significantly decreased T-cell proliferation and enhanced the frequencies of CD4+IL-10+ T cells in the context of anti-CD3/CD28 stimulation in the absence of APCs, pointing to direct effects on CD4+ T cells that are not associated with regulation of APC functions (Fig. S4 A and B).
Fig. S3.
Human act-A–iTr1 cells are hyporesponsive to restimulation in vitro. (A) Tconv or act-A–iTr1 cells were isolated from primary stimulation cultures as above; labeled with CFSE; and restimulated with fresh allergen-loaded, mitomycin-treated APCs for another 7 d. Representative FACS plots are shown. (B) Cumulative data, shown as mean ± SEM, from n = 3 independent experiments (n = 3 donors) are depicted. (C) FACS plots showing annexin-V and 7-AAD expressing Tconv or act-A–iTr1 cells. Numbers in plots indicate percentages of annexin-V+7AAD+CD4+ T cells (data represent n = 2 donors). (D) Tconv or act-A–iTr1 cells were restimulated with allergen-loaded, mitomycin-treated APCs and recombinant human IL-2, as indicated. 3[H]Thymidine incorporation is depicted. Data shown are mean ± SEM of triplicate wells and are representative of n = 4 independent experiments (n = 4 donors). TdR, tritiated thymidine. (E) Naive CD4+ T cells were cultured with mitomycin-treated, allergen-loaded APCs in the presence of control (PBS) or activin-A for 14 d, and CD4+ T cells were then isolated and cultured with fresh APCs in the presence of tetanus toxoid for another 6 d. 3[H]Thymidine incorporation and IFN-γ release in secondary cultures are depicted. Data shown are mean ± SEM of triplicate wells and are pooled from n = 2 independent experiments (n = 2 donors). Statistical significance was obtained by the Student’s t test (**P < 0.01; ***P < 0.001).
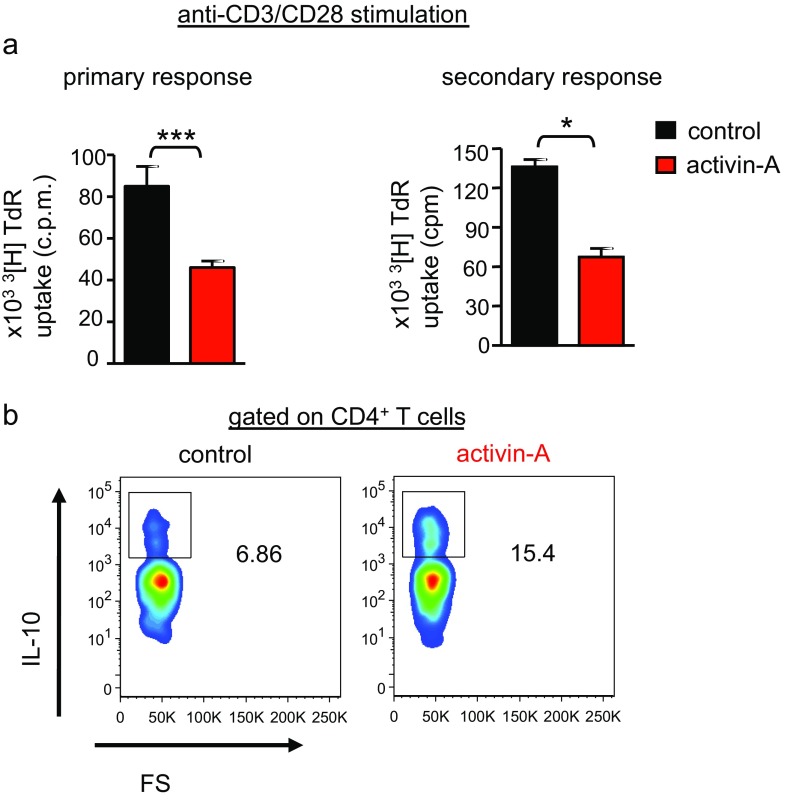
Fig. S4.
Activin-A suppresses anti-CD3–driven human T-cell responses in an APC-free system. (A, Left) Human naive CD4+ T cells were stimulated with anti-CD3/CD28 antibodies in the presence of activin-A (act-A–iTr1 cells) or PBS (Tconv cells) for 7 d. (A, Right) Tconv or act-A–iTr1 cells were isolated from primary induction cultures and restimulated with anti-CD3/CD28 antibodies for another 7 d. 3[H]Thymidine incorporation is depicted. Data shown are mean ± SEM of triplicate wells and are pooled from n = 4 independent experiments (n = 4 donors). (B) Representative FACS plots showing IL-10–expressing cells, gated on CD4+ T cells, stimulated as above. Data represent n = 4 independent experiments (n = 4 donors). Statistical significance was obtained by the Student’s t test (*P < 0.05; ***P < 0.001).
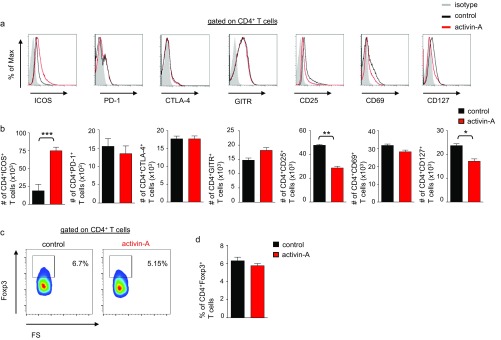
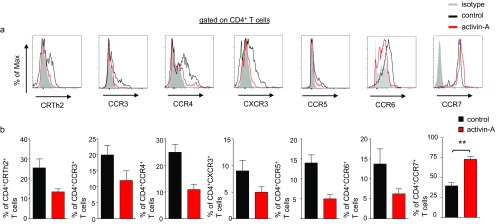
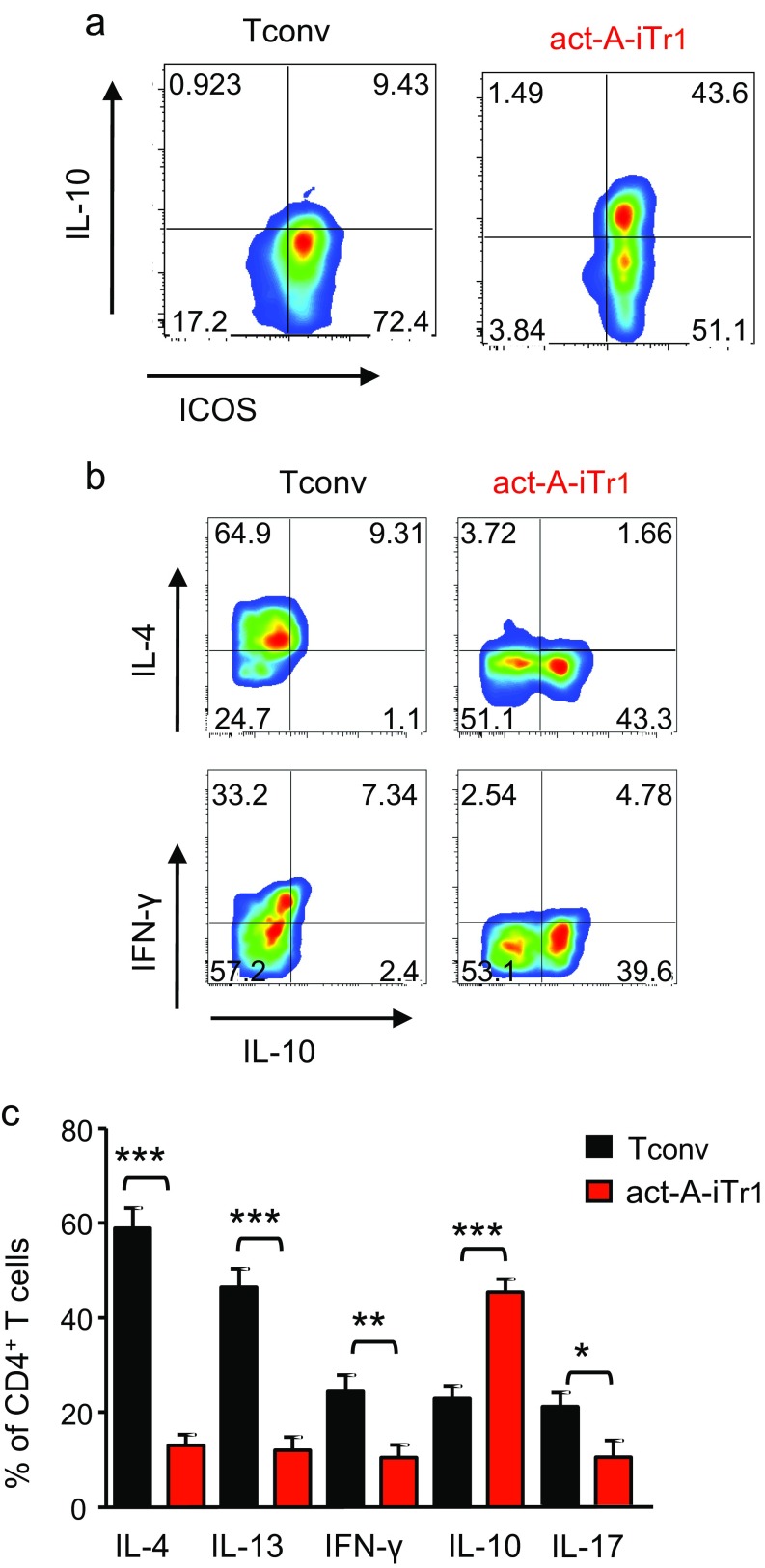
To characterize the immunophenotype of activin-A–treated human T cells further, we investigated the expression of Treg-cell–associated markers. PD-1, GITR, and CTLA-4 were expressed at equivalent levels between activin-A– and control-treated T cells (Fig. S5 A and B). Moreover, Foxp3 expression did not differ between the two groups (Fig. S5 C and D). Still, ICOS levels were markedly increased upon stimulation with activin-A, whereas the T-cell activation markers CD69, CD25, and CD127 were decreased (Fig. S5 A and B). CRTh2, CCR3, CCR4, CCR5, CCR6, and CXCR3 were dampened in activin-A–treated T cells, whereas CCR7 was increased (Fig. S6 A and B). Interestingly, we observed a striking increase in the percentages of ICOS+IL-10+CD4+ T cells among activin-A–treated T cells, accompanied by markedly elevated IL10 and ICOS mRNA levels (Fig. 1 D and E). Moreover, a large proportion of CD4+ICOS+IL-10+ T cells among activin-A–treated T cells expressed CD49b and LAG3, recently identified as human Tr1-cell–associated markers (25), concomitant with a significant increase in the mRNAs encoding LAG3 and GZMB (Fig. 1 E and F).
Fig. S5.
Act-A–iTr1 cells express the Treg-cell–associated marker ICOS. Representative FACS plots show surface marker expression (A) or intracellular Foxp3 levels (C) in human CD4+ T cells following stimulation with activin-A or control (PBS) in the presence of allergen-loaded APCs. Cumulative data depict the numbers of Treg-cell markers expressing CD4+ T cells (B) or the percentages of Foxp3+CD4+ T cells (D). Data shown are mean ± SEM and are pooled from n = 8 independent experiments (n = 8 donors). Statistical analysis was performed by the Student's t test (*P < 0.05; **P < 0.01; ***P < 0.001).
Fig. S6.
Act-A–iTr1 cells express the chemokine receptor CCR7. (A) Representative FACS plots showing chemokine receptor levels on human CD4+ T cells following stimulation with activin-A or control (PBS) in the presence of allergen-loaded APCs. (B) Cumulative data depict the percentages of chemokine receptor-expressing CD4+ T cells. Data are shown as mean ± SEM and are pooled from n = 8 independent experiments (n = 8 donors). Statistical significance was obtained by the Student's t test (**P < 0.01).
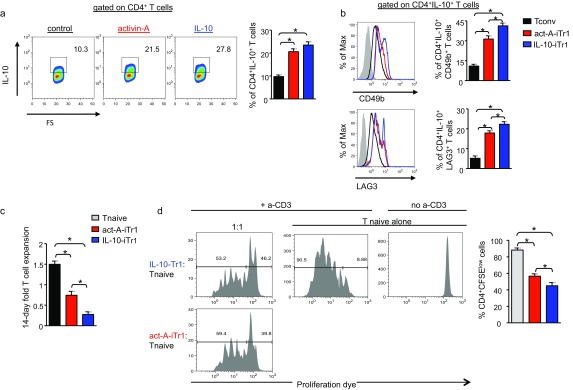
IL-10–induced human Tr1 cells, generated as previously described (25), contained similar percentages of CD4+IL-10+ T cells, compared with activin-A–treated CD4+ T cells, whereas LAG-3 and CD49b expression was increased in IL-10–induced Tr1 cells (Fig. S7 A and B). The expansion of activin-A–treated CD4+ T cells in culture was significantly increased compared with the expansion of IL-10–induced Tr1 cells, which exhibited poor expansion in agreement with previous studies (25) (Fig. S7C).
Fig. S7.
Functional and phenotypic similarities between act-A–iTr1 cells and IL-10–induced human Tr1 cells. (A) Human naive CD4+ T cells were stimulated with anti-CD3/CD28 antibodies in the presence of mitomycin-treated, CD3-depleted APCs and activin-A (act-A–iTr1), PBS (Tconv), or rhIL-10/rhIFN-α (IL-10–iTr1) cells for 7 d. Tconv, act-A–iTr1, or IL-10–Tr1 cells were isolated from primary induction cultures and restimulated as above for another 7 d. (Left) Representative FACS plots show IL-10–expressing cells gated on CD4+ T cells. (Right) Cumulative data are pooled from n = 4 independent experiments (n = 4 donors). (B) Representative FACS plots showing CD49b and LAG3 expression, gated on CD4+IL-10+ T cells. (Right) Cumulative data are pooled from n = 4 independent experiments (n = 4 donors). (C) Fold expansion of cells over 14 d of stimulation. Data are pooled from n = 4 independent experiments (n = 4 donors). (D) Human naive CD4+ T cells were stimulated as in A, and T cells were then isolated and cocultured with CellTrace-CFSE–labeled autologous CD4+ T responders in the presence of anti-CD3/CD28 antibodies for 5 d. (Left) Representative FACS plots show T-responder proliferation. (Right) Cumulative data, shown as mean ± SEM, are pooled from n = 4 independent experiments (n = 4 donors). Statistical analysis was performed by a Mann–Whitney unpaired Student’s t test (*P < 0.05).
Notably, even after prolonged (4 wk) T-cell receptor stimulation, CD4+ T cells stimulated with activin-A did not express Th-effector cytokines, although they maintained high frequencies of CD4+IL-10+ICOS+ T cells (Fig. S8 A–C). Collectively, these findings reveal that activin-A stimulation restrains allergen-driven human T-cell responses and instructs the differentiation of IL-10–producing Tr1-like cells.
Fig. S8.
Act-A–iTr1 cells retain their phenotype following prolonged stimulation in vitro. Human Tconv or act-A–iTr1 cells were restimulated (once per week) with allergen-loaded APCs and IL-2 for 4 wk. Representative FACS plots show ICOS and IL-10 expression gated on CD4+ T cells (A) and intracellular cytokine expression (B). Data represent n = 5–6 separate experiments (n = 6 donors). (C) Cumulative data showing the percentages of cytokine-producing CD4+ T cells. Data shown are mean ± SEM and are pooled from n = 5–6 independent experiments (n = 6 donors). Statistical analysis was performed by the Student’s t test (*P < 0.05; **P < 0.01; ***P < 0.001).
Activin-A–Induced Human Tr1-Like Cells Display Strong Suppressive Functions.
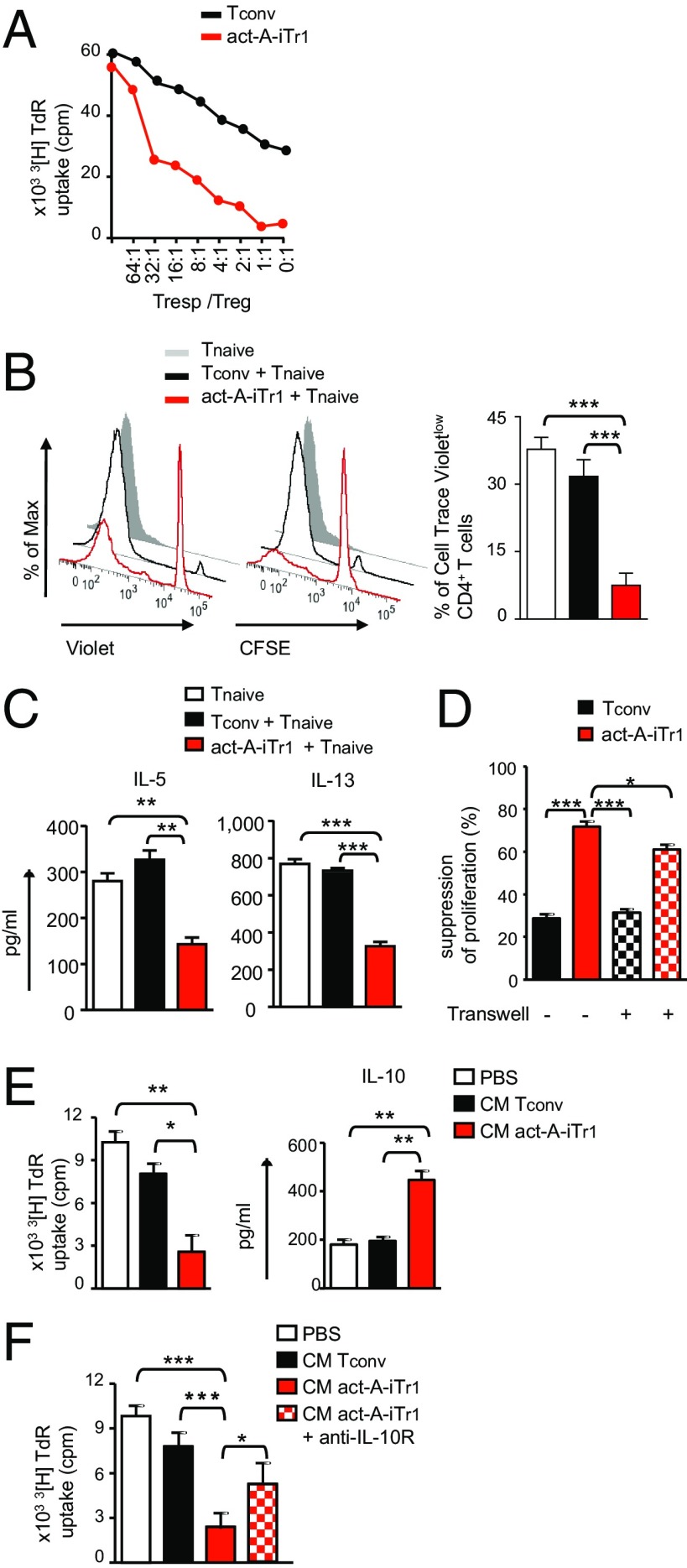
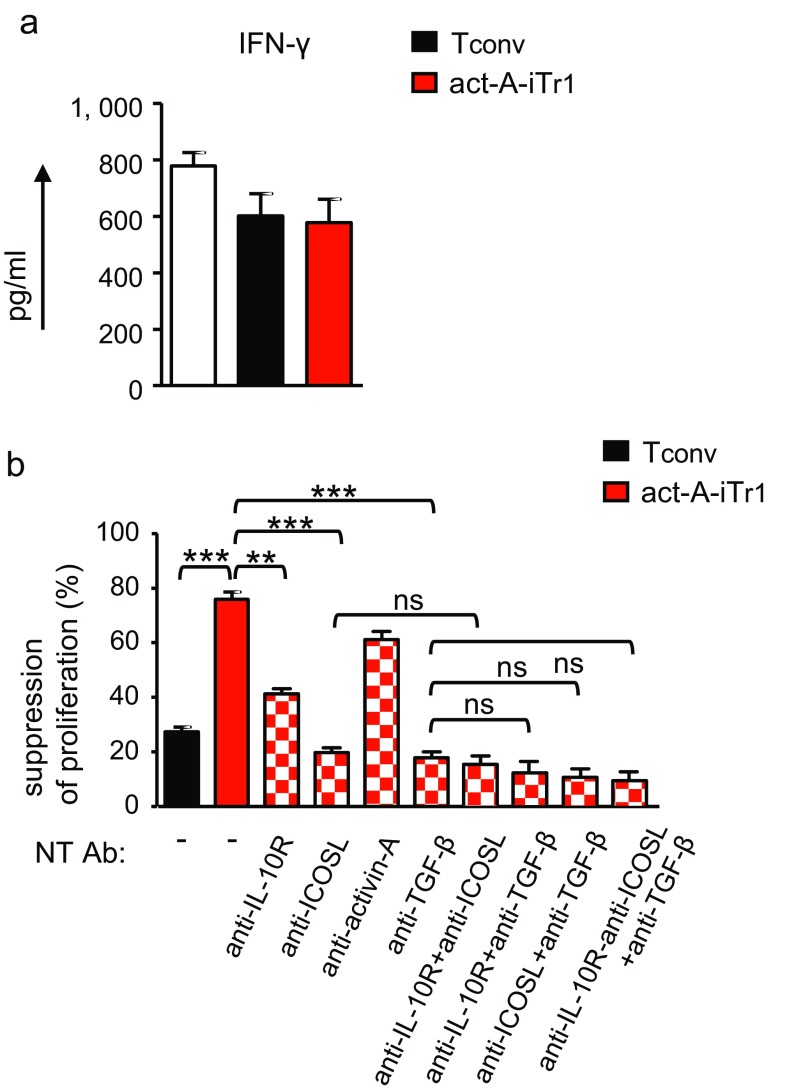
On the basis of the aforementioned findings, we next explored whether stimulation with activin-A endows human T cells with immunoregulatory functions. To address this issue, we cocultured carboxyfluorescein succinimidyl ester (CFSE)-labeled, activin-A–treated, or control-treated T cells, as potential suppressors, with autologous Cell Trace Violet-labeled naive CD4+ T responders and allergen-primed APCs. Our data showed that activin-A–treated T cells greatly suppressed the proliferation of T responders, even at high T-responder/T-suppressor ratios (Fig. 2 A and B). Moreover, act-A–iTr1 cells significantly decreased IL-5 and IL-13 release in culture supernatants (Fig. 2C). Still, IFN-γ levels remained unaltered, possibly due to activation of a counterbalancing mechanism used by act-A–iTr1 cells to inhibit Th2 cytokine secretion (Fig. S9A). In contrast, control-treated T cells did not exhibit discernible suppressor functions (Fig. 2 A–C). The proliferation of act-A–iTr1 cells remained low in suppressive cultures (Fig. 2B). Naive T responders cultured with allergen-pulsed APCs alone displayed enhanced proliferative capacity, compared with the other groups (Fig. 2B). IL-10–induced human Tr1 cells demonstrated an enhanced suppressive capacity toward T-responder cell proliferation, compared with act-A–iTr1 cells (Fig. S7D).
Fig. 2.
Activin-A–induced human Tr1-like cells display strong suppressive functions. (A) Human naive CD4+ T cells were stimulated with allergen-loaded, mitomycin-treated APCs in the presence of PBS (Tconv) or activin-A (act-A-iTr1) cells, and T cells were then isolated and cocultured with autologous naive CD4+ T responders (at the indicated ratios) in the presence of allergen-loaded APCs for 4 d. 3[H]Thymidine incorporation is shown. Data are representative of n = 3 independent experiments (n = 3 donors). TdR, tritiated thymidine. (B) Tconv or act-A–iTr1 cells were generated as in A, CFSE-labeled, and cocultured with Cell Trace Violet-labeled autologous naive CD4+ T responders and allergen-loaded, mitomycin-treated APCs. Representative FACS plots showing T-cell proliferation (Left) and cumulative data (Right), shown as mean ± SEM, are pooled from n = 6–8 separate experiments (n = 8 donors). (C) Th2 cytokines in culture supernatants are shown. Data are expressed as mean ± SEM of triplicate wells and are pooled from n = 6–8 separate experiments (n = 8 donors). (D) Tconv or act-A–iTr1 cells and allergen-loaded APCs were cultured in a transwell system across from cocultures of autologous naive CD4+ T cells and APCs. Suppression of T-responder proliferation is shown. Data are expressed as mean ± SEM of triplicate wells and are pooled from n = 3 independent experiments (n = 3 donors). (E) CM from Tconv or act-A–iTr1 culture supernatants was added to naive CD4+ T-cell/APC cocultures. 3[H]Thymidine incorporation and IL-10 release are depicted. Data are expressed as mean ± SEM of triplicate wells and are pooled from n = 4–6 separate experiments (n = 4 donors). (F) CM from Tconv or act-A–iTr1 culture supernatants was added to naive CD4+ T-cell/APC cocultures, as indicated. 3[H]Thymidine incorporation is depicted, and data are expressed as mean ± SEM of triplicate wells and pooled from n = 3–4 experiments (n = 4 donors). Statistical analysis was performed by the Student’s t test (A–C and E) and a nonparametric (Mann–Whitney) unpaired Student's t test (D and F) (*P < 0.05; **P < 0.01; ***P < 0.001).
Fig. S9.
TGF-β, IL-10, and ICOS are involved in the suppressive functions of human act-A–iTr1 cells. (A) Human naive CD4+ T cells were stimulated with allergen-loaded, mitomycin-treated APCs in the presence of PBS (Tconv cells) or activin-A (act-A–iTr1 cells), and T cells were then isolated and cocultured with autologous naive CD4+ T responders in the presence of allergen-loaded APCs for 4 d. IFN-γ in culture supernatants is shown. Data are expressed as mean ± SEM of triplicate wells and are pooled from n = 6–8 separate experiments (n = 8 donors). (B) Tconv or act-A–iTr1 cells were cocultured with autologous T responders and allergen-loaded, mitomycin-treated APCs, as indicated. Suppression of T-cell proliferation is shown. Data are shown as mean ± SEM of triplicate wells and summarize n = 5 separate experiments (n = 5 donors). Statistical analysis was performed by the Student’s t test **P < 0.01; ***P < 0.001). ns, not significant; NT Ab, neutralizing antibody.
To elucidate the mechanisms involved in the suppressive functions of human act-A–iTr1 cells, we initially used a transwell system. The suppressive capacity of activin-A–treated CD4+ T cells across a permeable membrane was decreased, but not abolished, in this setting, pointing to the involvement of both cell contact-dependent and independent mechanisms (Fig. 2D). Indeed, transfer of the conditioned medium (CM) obtained from the supernatants of act-A–iTr1 cells to fresh CD4+ T-cell/APC cocultures suppressed T-responder proliferation (Fig. 2E). Interestingly, IL-10 production was significantly increased in secondary cultures (Fig. 2E). Disruption of IL-10 signaling, using an antibody against the IL-10 receptor (IL-10R) in secondary cultures, partially reversed the inhibitory effects of the CM obtained from act-A–iTr1 cells, emphasizing the involvement of IL-10 in their suppressive functions (Fig. 2F). Blockade of IL-10, TGF-β, or ICOS signaling in secondary suppressive cultures partly impaired the suppressive capacity of act-A–iTr1 cells, whereas blockade of activin-A did not significantly affect their functions (Fig. S9B). Concomitant blocking of IL-10, TGF-β, and ICOS signaling further reduced act-A–iTr1 cell inhibitory functions, compared with blocking each pathway alone or combinations of two; however, the data did not reach statistical significance (Fig. S9B). Overall, these findings reveal that human act-A–iTr1 cells exhibit robust suppressive functions against naive and effector T-cell responses to allergen through IL-10–, TGF-β–, and ICOS-dependent pathways.
IRF4 Is Activated in Human Act-A–iTr1 Cells and Forms Multipartite Complexes with AhR in the ICOS Promoter.
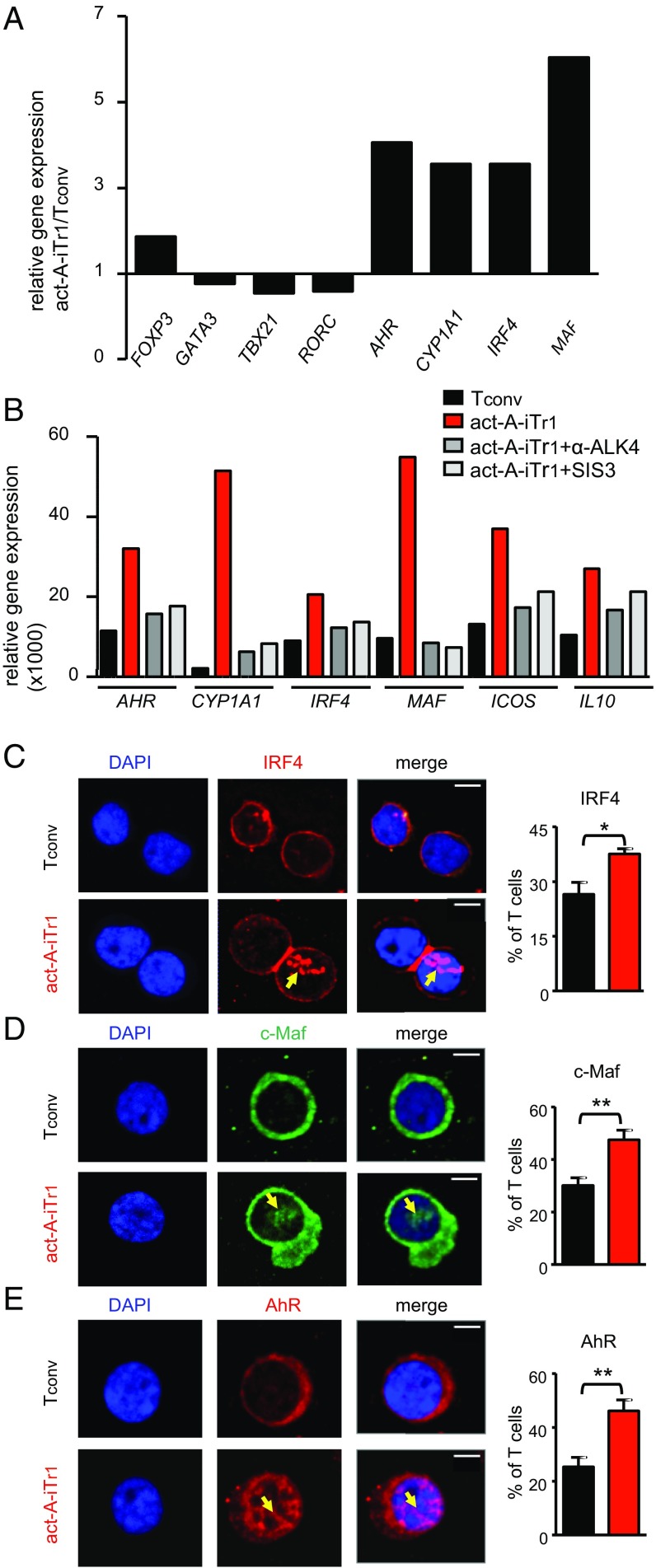
We next attempted to decipher the transcriptional pathways contributing to activin-A–induced human Tr1-cell differentiation. Investigation of the expression of the major Th-cell lineage–associated transcription factors TBX21, GATA3, and RORC at an earlier time point (day 3) during T-cell differentiation demonstrated that mRNA levels were similar or slightly decreased in act-A–iTr1 cells, whereas FOXP3 was moderately increased, compared with Tconv cells (Fig. 3A). Interestingly, act-A–iTr1 cells expressed increased transcripts of the Tr1-cell–associated transcription factors MAF (encoding c-Maf) and AHR, along with its major gene target, xenobiotic metabolizing enzyme cytochrome P450, encoded by CYP1A1 (8) (Fig. 3A). Moreover, activin-A stimulation induced a marked up-regulation in the mRNA levels of IRF4, a transcription factor not previously associated with human or mouse Tr1 cells (Fig. 3A). Inhibition of activin-A signaling, using an anti-ALK4 antibody or the Smad3 inhibitor (SIS3), decreased activin-A–induced gene transcription, pointing to the involvement of its canonical signaling pathways (Fig. 3B). Confocal microscopy studies demonstrated that activin-A stimulation induced an enhanced translocation of IRF4, c-Maf, and AhR into the nucleus of human CD4+ T cells (Fig. 3C).
Fig. 3.
IRF4, AhR, and c-Maf are activated upon stimulation of human CD4+ T cells with activin-A. (A) Human naive CD4+ T cells were stimulated with allergen-loaded, mitomycin-treated APCs in the presence of PBS or activin-A for 3 d. Real-time PCR analysis of the indicated genes is shown. Results are representative of n = 4 separate experiments (n = 4 donors) and were normalized to GAPDH. (B) Tconv or act-A–iTr1 cells were generated as above with an anti-ALK4 blocking antibody or the Smad3 inhibitor, SIS3. Real-time PCR results are shown relative to GAPDH. Data are representative of n = 4 independent experiments (n = 4 donors). (C–E) Confocal microscopy images of Tconv or act-A–iTr1 cells stained with fluorescently conjugated anti-AhR, anti-IRF4, and anti–c-Maf antibodies. Nuclei are stained blue with DAPI. (Scale bars: IRF4 images, 4 μm; AhR and c-Maf images, 2 μm). Cumulative data showing the percentages of AhR+, IRF4+, or c-Maf+ T cells were pooled from n = 4 independent experiments (n = 4 donors). Statistical analysis was performed by a nonparametric (Mann–Whitney) unpaired Student’s t test (*P < 0.05; **P < 0.01).
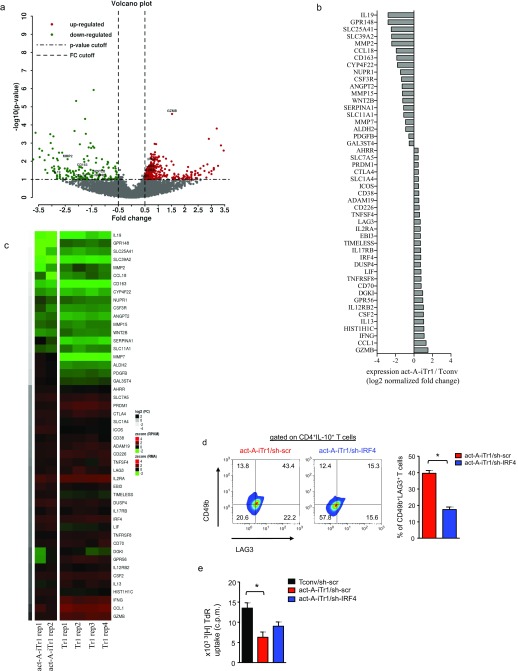
Comparison of the global gene expression profile of human act-A–iTr1 cells with the global gene expression profile of Tconv cells, using RNA-sequencing (RNA-seq) analysis, revealed notable differences in their transcriptome profile, with 513 genes being significantly differentially regulated, 200 of which were down-regulated and 313 of which were up-regulated (Fig. S10 A and B). Volcano plot visualization demonstrated that the transcriptional signature of act-A–iTr1 cells was characterized by substantial changes compared with Tconv cells, with nearly 50% of the total differentially expressed genes showing a difference in expression greater than twofold [log2 (fold change [FC]) < −1 or > 1] (Fig. S10A). These genes belonged to biologically important gene families, including chemokines/cytokines or their receptors, transcription factors, surface receptors, and Treg-cell markers (Fig. S10B). Interestingly, several genes, including LAG3, IRF4, GZMB, CCL1, CSF2, DGKI, CD70, and EBI3, were enriched in both human act-A–iTr1 cells and IL-10–induced Tr1 cells (25), whereas others, such as CCL18, WNT2B, MMP15, and CD163, were down-regulated in both groups (Fig. S10C).
Fig. S10.
Transcriptional signature of human act-A–iTr1 cells. (A) Changes in gene expression (x axis) and statistical significance (y axis) in human act-A–iTr1 cells relative to Tconv cells, presented as a volcano plot [red: P < 0.1 and log2 (FC) > 0.5, green: P < 0.1 and log2 (FC) < −0.5]. (B) Differential expression of 48 immune response-related genes in act-A–iTr1 cells relative to Tconv cells. (C) Comparison of the transcriptional profile of act-A–iTr1 cells with the transcriptional profile of human Tr1 cells (25). The genes shown in B, which were differentially expressed in act-A–iTr1 cells, were used for the comparison. Genes were sorted by increasing RNA-seq log2 (FC), revealing similar patterns of expression between the two platforms. Regression analysis showed a strong correlation between the average reads per kilobase per million mapped reads (RPKM) values (RNA-seq) and the average RMA values (microarrays) (P < 0.001). All expression values were scaled to have a mean of zero and SD of 1. (D) Tconv or act-A–iTr1 cells were stimulated and transduced, as in Fig. 5. Representative FACS plots (Left) and cumulative data (Right) show LAG3 and CD49b expression, gated on GFP+CD4+ T cells representative of n = 4 separate experiments (n = 4 donors). (E) 3[H]Thymidine incorporation is depicted. Data shown are mean ± SEM of triplicate wells and are pooled from n = 4 independent experiments (n = 4 donors). Statistical analysis was performed by a nonparametric (Mann–Whitney) unpaired Student’s t test (*P < 0.05).
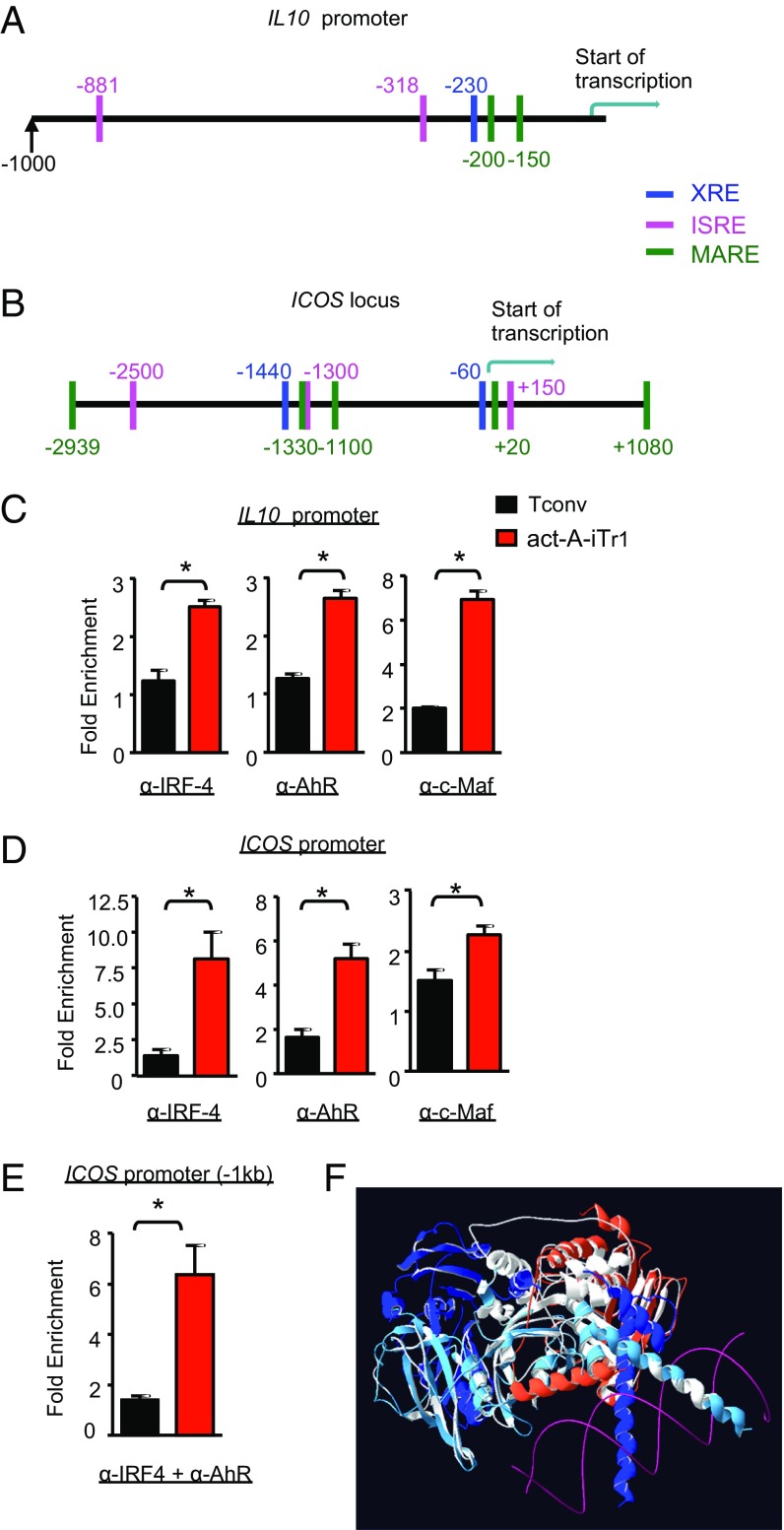
In view of the aforementioned findings, we hypothesized that activin-A up-regulates IL10 and ICOS expression in human CD4+ T cells through IRF4-, AhR-, and c-Maf–dependent pathways. To examine this hypothesis, we initially performed bioinformatics analysis to decipher whether these transcription factors have putative binding sites in human IL10 and ICOS gene loci. Indeed, we identified two IRF4-binding sites [IFN-stimulated response elements (ISREs)] in the IL10 promoter and three ISREs in the ICOS locus (Fig. 4 A and B). In agreement with previous studies, we found a putative AhR-binding site [xenobiotic response element (XRE-1)] and a c-Maf site [MAF-recognition element (MARE-1)] within the IL10 promoter, which were in high proximity and partially overlapping (8) (Fig. 4A). Moreover, we identified a c-Maf–binding site (MARE-2, −150 bp) in the IL10 promoter and two XRE and five MARE sites in the ICOS locus (Fig. 4 A and B).
Fig. 4.
IRF4 assembles into a complex with AhR in the ICOS promoter in human act-A–iTr1 cells. Schematic representation of the human IL10 promoter (A) and ICOS locus (B); the c-Maf, AhR, and IRF4 putative binding sites are shown. Positions are given in base pairs relative to the transcription start site. (C and D) Tconv or act-A–iTr1 cells were generated following stimulation with anti-CD3/CD28 antibodies in the presence of PBS or activin-A, respectively. ChIP analysis of IRF4, AhR, and c-Maf binding in the IL10 (C) and ICOS (D) promoters is shown. Data are presented as ± SEM and are pooled from two independent experiments (n = 4 donors, two donors per experiment). (E) Tconv or act-A–iTr1 cells were generated as in C. Sequential ChIP analysis demonstrates IRF4 and AhR cobinding in the ICOS promoter. Data are presented as mean ± SEM and are pooled from two independent experiments (n = 4 donors, two per experiment). (F) Computational model of the putative protein–protein interactions of IRF4 and AHR is shown in white superimposed with the solved structures used for the homology modeling (PDB ID codes 3DSH and 4ZPR). The template chain of the IRF4 model is shown in orange. In the 4ZPR structure, the ARNT chain is shown in blue, the AHR template chain is shown in light blue, and the DNA is shown in magenta. Statistical analysis was performed by a nonparametric (Mann–Whitney) unpaired Student’s t test (*P < 0.05).
We next focused on investigating whether activin-A signaling in CD4+ T cells induces the binding of IRF4, AhR, and c-Maf to their response elements in IL10 and ICOS loci. Chromatin immunoprecipitation (ChIP) assays revealed a marked enrichment of IRF4 binding to the ISRE (−318 bp) site in the IL10 promoter in act-A–iTr1 cells, compared with Tconv cells (Fig. 4C). In addition, AhR and c-Maf bound on the XRE-1 (−230 bp) and MARE-1 (−200 bp) sites, respectively, in the IL10 promoter in human act-A–iTr1 cells (Fig. 4C). In contrast, AhR showed no discernible binding to the XRE-1 element in the IL10 promoter in Tconv cells, whereas c-Maf bound moderately to its MARE-1 site (Fig. 4C). Pertinent to the ICOS locus, our data showed that IRF4 exhibited great enrichment to its ISRE (−1.3 kb) site, whereas AhR bound to its respective XRE site (−1,440 bp) in act-A–iTr1 cells (Fig. 4D). Modest binding of c-Maf to the MARE site (−1,330 bp) in the ICOS locus was also observed (Fig. 4D).
The ability of IRF4 and AhR to bind to the ICOS promoter in human T cells in response to activin-A stimulation, and the proximity of their binding sites, prompted us to explore whether they form transcriptional complexes at the ICOS locus. To address this issue, we performed sequential ChIP analysis of activin-A– or Tconv cells. Remarkably, our findings identified the presence of IRF4/AhR complexes in the ICOS promoter in act-A–iTr1 cells, but not in Tconv cells (Fig. 4E).
To dissect the molecular interactions of IRF4 with AhR further, we performed in silico analysis of putative protein–protein interactions between the two transcription factors. Because a description of the solved structures of the full-length sequences of these transcription factors is currently lacking, reliable models were constructed and submitted for molecular docking. Interestingly, our studies revealed that IRF4 and AhR can form a tripartite molecular complex together with the AhR-binding partner, ARNT (26) (Fig. 4F). This complex was highly stable thermodynamically because after the minimization of its energy, it was characterized by an energy value of −260685.0 KJ/mol. Moreover, this structure remained stable throughout the duration (10 ns) of the molecular dynamics simulation studies. Additionally, when this energetically minimized complex was superimposed with the initial docking output, the rmsd values between the carbon α-atoms were extremely low (Cα rmsd values at 0.52 Å). Together, this evidence supports the notion that the proposed docking result of IRF4/AhR is highly reliable and exhibits high thermodynamic stability.
These results collectively demonstrate that activin-A induces the activation of IRF4, AhR, and c-Maf in human CD4+ T cells, which bind in their response elements in IL-10 and ICOS promoters through the formation of multipartite transcriptional complexes.
IRF4 and AhR Signaling Is Essential for Activin-A–Induced Human Tr1-Cell Differentiation.
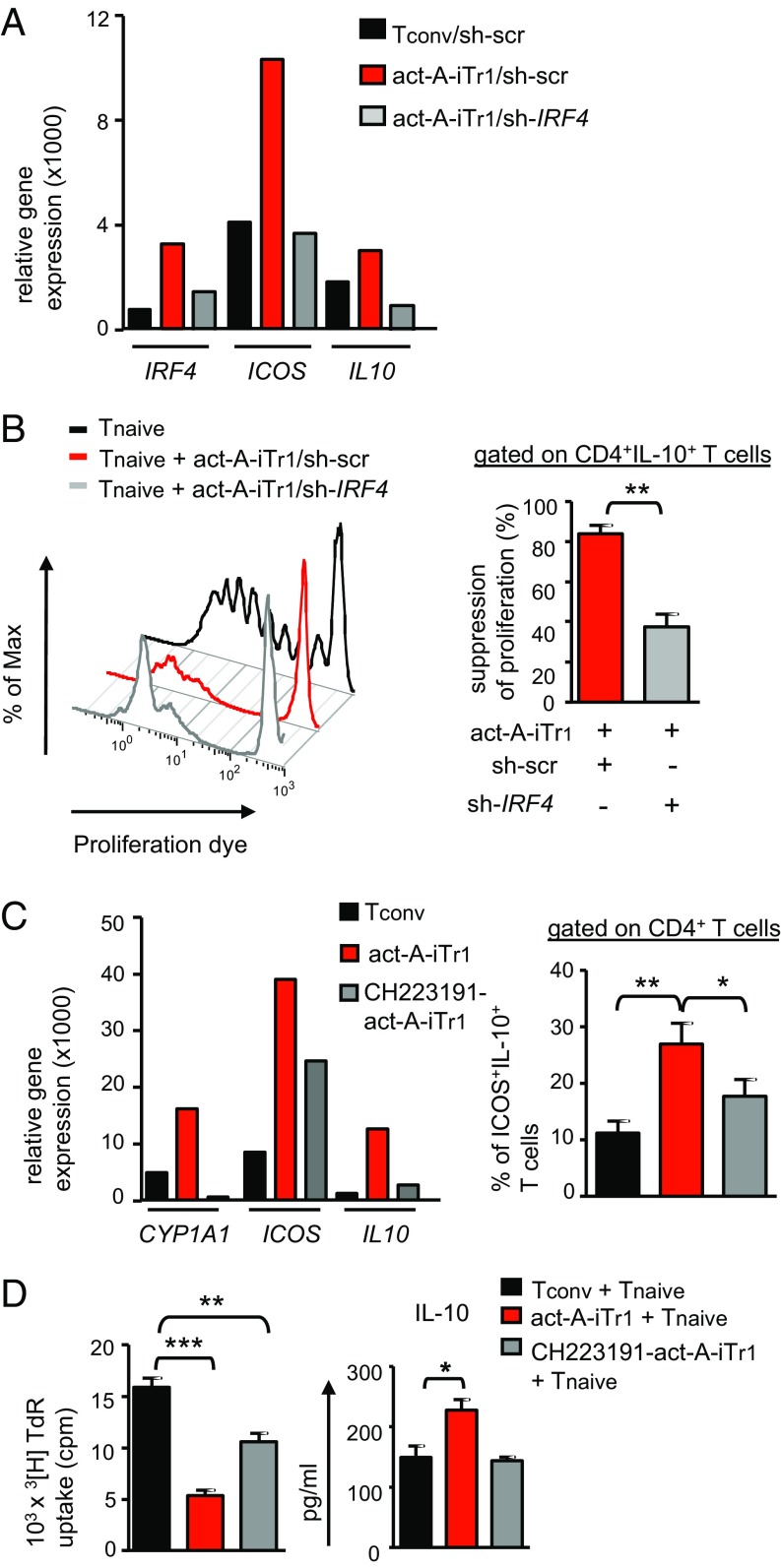
To obtain functional evidence that IRF4 is involved in activin-A–mediated differentiation of human Tr1-like cells, we used IRF4-specific shRNA (shIRF4) to target IRF4 expression in T cells directly. Briefly, CD4+ T cells transduced with lentiviral particles containing constructs of GFP-shRNA for IRF4 were sorted based on viability and GFP expression, and used in in vitro suppression assays. Indeed, knockdown of IRF4 decreased IRF4, ICOS, and IL10 mRNA levels in CD4+ T cells in response to activin-A (Fig. 5A). The percentages of LAG-3+CD49b+ T cells were also decreased, compared with sh-scrambled–treated, activin-A–stimulated T cells (Fig. S10D). IRF4 silencing also compromised the suppressive functions of act-A–iTr1 cells and partly reversed their hypoproliferative state (Fig. 5B and Fig. S10E).
Fig. 5.
IRF4 and AhR drive activin-A–induced human Tr1 cell differentiation. (A) Tconv or act-A–iTr1 cells were stimulated with APCs and allergen, and transduced with shRNA against human IRF4 (sh-IRF4) or a scrambled nontargeting control RNA sequence (sh-scr). Real-time PCR analysis is shown. Data are relative to GAPDH and represent n = 3 independent experiments (n = 3 donors). (B) Transduced act-A–iTr1 cells were cocultured with naive Cell Trace Violet-labeled CD4+ T cells and allergen-loaded APCs. Representative FACS plots of T-responder cell proliferation are shown. Cumulative data, shown as mean ± SEM, depict suppression of proliferation (%) and are pooled from n = 4 separate experiments (n = 4 donors). (C) Tconv or act-A–iTr1 cells were generated following stimulation with allergen-loaded APCs in the presence or absence of the AhR antagonist CH223191. (Left) Real-time PCR analysis is shown. Results are relative to GAPDH and represent n = 4 separate experiments (n = 4 donors). (Right) Cumulative data showing the percentages of CD4+ICOS+IL-10+ T cells summarize four independent experiments (n = 4 donors). (D) Tconv or act-A–iTr1 cells, generated as in C, were isolated and cocultured with CD4+ T cells and allergen-loaded APCs. 3[H]Thymidine incorporation and IL-10 release are shown. Data are mean± SEM of triplicate wells and are pooled from n = 4 independent experiments (n = 2 donors). Statistical significance was obtained by the Student's t test (*P < 0.05; **P < 0.01; ***P < 0.001).
Because IRF4 assembles into a complex with AhR in human T cells in response to activin-A, we next explored the role of AhR in activin-A–induced responses using CH-223191, a potent AhR antagonist (8). Pharmacological blockade of AhR nearly abrogated activin-A–induced AhR activation, as shown by a marked down-regulation of CYP1A1 mRNA (Fig. 5C). Interestingly, inhibition of AhR signaling partly reversed activin-A–induced IL10, ICOS, and MAF up-regulation and significantly decreased the frequencies of CD4+ICOS+IL-10+ T cells (Fig. 5C). In addition, disruption of AhR signaling impaired the suppressive capacities of act-A–iTr1 cells, accompanied by a significant reduction in IL-10 release (Fig. 5D). In summary, these results provide compelling evidence that IRF4 and AhR are critically involved in activin-A–induced differentiation of human Tr1-like cells.
In Vivo Administration of Human Act-A–iTr1 Cells Prevents the Development of Allergic Airway Disease.
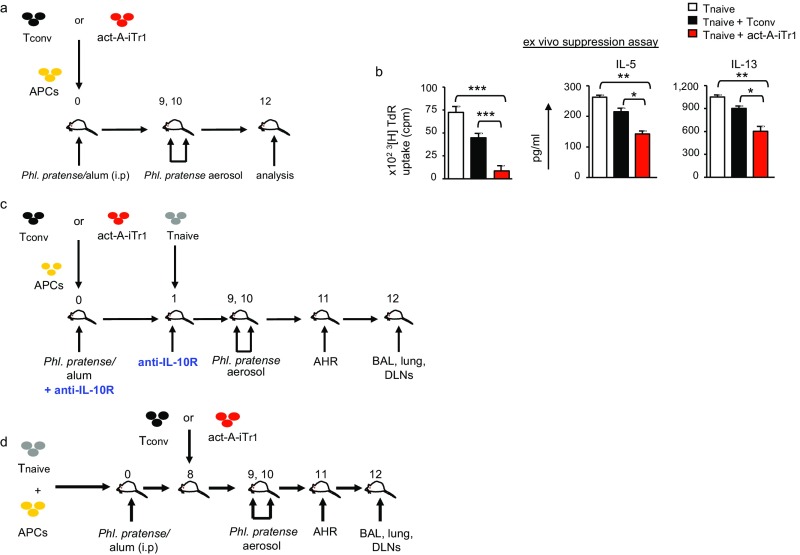
Considering ongoing efforts to use adoptive Treg-cell transfers as a therapy for immune-mediated human diseases, the issue of their in vivo stability is of critical clinical importance. To explore whether act-A–iTr1 cells can maintain their suppressive functions upon transfer in vivo, we used a well-established humanized mouse model of allergic airway inflammation (23, 27, 28). Briefly, we transferred allergen-primed human act-A–iTr1 or Tconv cells, along with autologous APCs, into immunodeficient NOD-SCID mice and examined their responses upon immunization and aerosol challenge with allergen (Fig. S11A). Indeed, human CD4+ T cells isolated from the lymph nodes (LNs) of NOD-SCID mice reconstituted with act-A–iTr1 cells were able to suppress the proliferation and Th2 cytokine release by cocultured naive T responders in ex vivo suppression assays (Fig. S11B). In contrast, CD4+ T cells obtained from the LNs of Tconv-cell–reconstituted mice did not exhibit discernible suppressor functions (Fig. S11B). These data indicate that act-A–iTr1 cells can retain their potent inhibitory functions against other T-cell responses even following in vivo transfer into a highly inflammatory milieu.
Fig. S11.
Schematic representation of the humanized mouse models of allergic asthma. (A) Experimental protocol used. (B) Human CD4+ T cells were isolated from the LNs and spleens of reconstituted mice at the time of euthanasia and cocultured with freshly isolated autologous naive CD4+ T responders, APCs, and Phleum pratense. 3[H]Thymidine incorporation and cytokine release are depicted. Data shown are mean ± SEM of triplicate wells and are representative of two to three independent experiments (n = 4–6 mice per group). Statistical analysis was performed by the Student's t test (* P < 0.05; **P < 0.01; ***P < 0.001). (C) In the prevention protocol, NOD-SCID mice were reconstituted on day 0 with 2 × 106 Tconv or act-A–iTr1 cells and 2 × 106 autologous APCs, and were immunized (i.p.) with P. pratense. On day 1, mice received autologous CD4+ T cells (106 cells). On days 9 and 10, mice were challenged with aerosolized P. pratense; AHR was measured the following day, and mice were killed. In other experiments, mice were administered (i.p.) anti-human IL-10Rα or Ig control 1 d prior to and following act-A–iTr1 cell (or Tconv) cell transfer. (D) In the therapeutic protocol, NOD-SCID mice were reconstituted on day 0 with naive CD4+ T cells (106 cells) and autologous APCs (106 cells), and immunized with P. pratense/alum. On day 8, mice received autologous Tconv or act-A–iTr1 cells (2 × 106 cells).
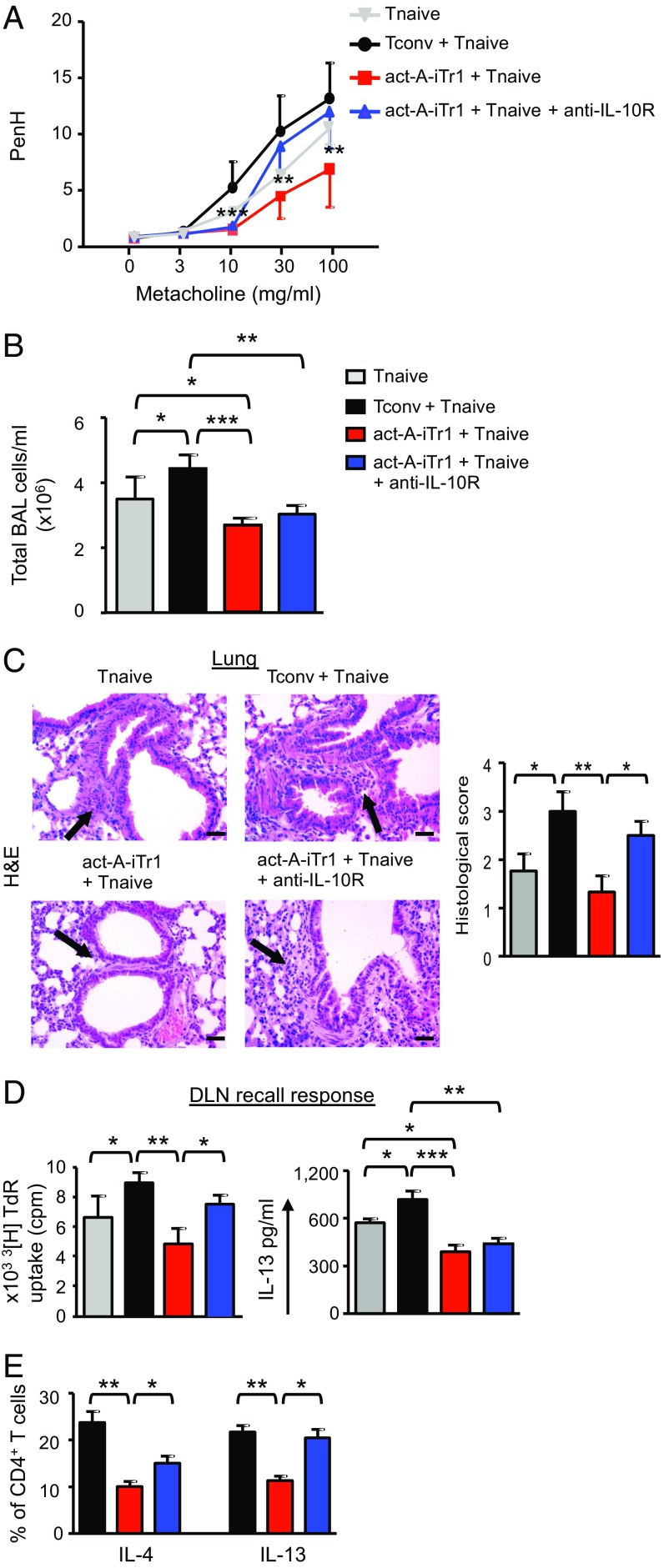
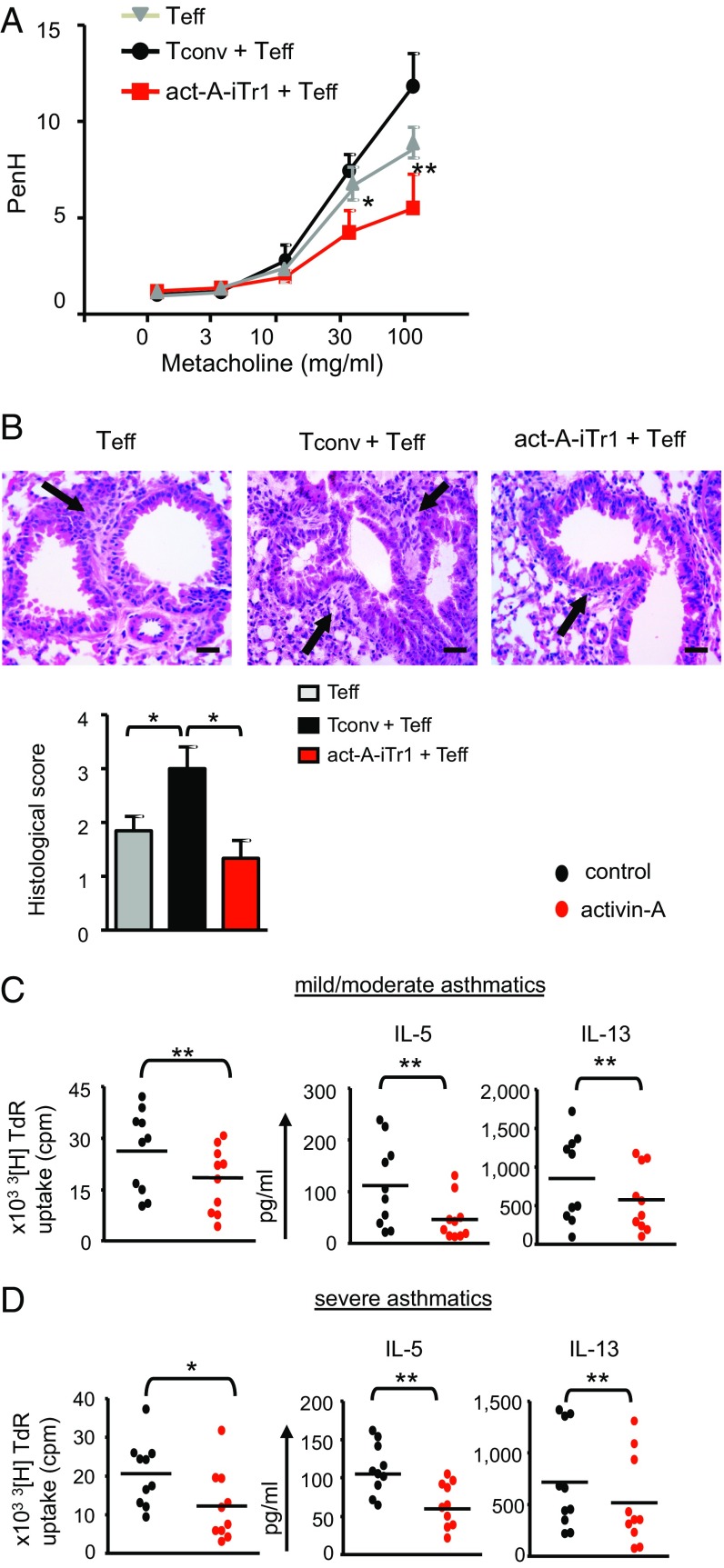
Subsequently, we investigated whether in vivo administration of human act-A–iTr1 cells in a preventive protocol could confer protection against features of experimental asthma (Fig. S11C). Indeed, adoptive transfer of act-A–iTr1 cells significantly decreased AHR to metacholine, a cardinal asthma manifestation, induced by cotransferred human T effectors (Fig. 6A). Importantly, administration of act-A–iTr1 cells significantly dampened inflammatory cell influx in the bronchoalveolar lavage (BAL) and prevented the development of pulmonary inflammation (Fig. 6 B and C). Tconv cells did not suppress airway inflammation or AHR; in fact, allergic responses were enhanced compared with recipients of T responders alone (Fig. 6 A–C). Considering the key role of IL-10 in the suppressive functions of human act-A–iTr1 cells in vitro, we examined its effects in vivo through administration of an anti–IL-10R antibody (or an isotype-matched control) (Fig. S11C). Disruption of IL-10 signaling partly reversed the suppressive effects of act-A–iTr1 cells on AHR and lung inflammation, but did not affect inflammatory cell responses in the BAL, compared with mice that received act-A–iTr1 cells but were treated with the isotype-matched control antibody (Fig. 6 A–C).
Fig. 6.
In vivo administration of human act-A–iTr1 cells prevents allergic airway disease. (A) AHR is depicted. Enhanced pause (PenH) results are expressed as mean ± SEM. Data were analyzed by two-way ANOVA for repeated measures, followed by the Student’s t test, and are pooled from two to three independent experiments (n = 4–6 mice per group). (B) Total BAL cell numbers are shown. (C) Representative photomicrographs and histological scores of H&E-stained lung sections. (Scale bars: 50 μm). (D) DLN cells were harvested at the time of euthanasia and restimulated with P. pratense. 3[H]Thymidine incorporation and IL-13 release in culture supernatants are presented. Data are mean ± SEM of triplicate wells. (E) Percentages of cytokine-producing human CD4+ T cells in the DLNs are shown. Data are representative of two to three independent experiments (n = 4–6 mice per group). Statistical analysis was performed by the Student’s t test (*P < 0.05; **P < 0.01; ***P < 0.001).
To explore the effects of act-A–iTr1 cells on the regulation of human T-effector responses in vivo, we isolated lung-draining lymph node (DLN) cells and restimulated them with allergen ex vivo. In line with the attenuated allergic airway disease severity, the capacity of DLN cells to proliferate and secrete IL-13 was significantly decreased in mice cotransferred with act-A–iTr1 cells, an effect that was partly reversed upon inhibition of IL-10 signaling (Fig. 6D). Moreover, the frequencies of IL-4+ and IL-13+ CD4+ T-effector cells in the DLNs of mice administered act-A–iTr1 cells were significantly decreased, and this effect was also reversed upon IL-10R blockade (Fig. 6E).
Altogether, these findings reveal that human act-A–iTr1 cells retain their immunosuppressive properties upon transfer in vivo and confer significant protection against asthma manifestations, partly through an IL-10–dependent mechanism.
Therapeutic Administration of Human Act-A–iTr1 Cells Reverses Established Allergic Responses and Ameliorates Asthma Severity.
To explore whether human act-A–iTr1 cells could reverse ongoing allergic airway inflammation, we adoptively transferred act-A–iTr1 cells, right before aerosol challenge, in mice reconstituted with T cells and sensitized to allergen (Fig. S11D). Indeed, administration of act-A–iTr1 cells induced a significant decrease in AHR induced by human Th2-effector cells and attenuated inflammatory cell infiltration in the lungs (Fig. 7 A and B), suggesting that this regime may have applications in ameliorating disease manifestations.
Fig. 7.
Therapeutic administration of human act-A–iTr1 cells reverses established allergic airway disease. (A) AHR is depicted. Results are expressed as means ± SEM. Data were analyzed as in Fig. 6A. (B) Representative photomicrographs and histological scores of H&E-stained lung sections. (Scale bars: 50 μm.) (C and D) CD4+ T cells were isolated from the peripheral blood of asthmatics and stimulated with mitomycin-treated, CD3-depleted, mixed grasses extract-loaded APCs, as indicated. 3[H]Thymidine incorporation and cytokine release in culture supernatants are depicted. Data are mean ± SEM of triplicate wells (n = 10 donors). Statistical analysis was performed by a nonparametric (Mann–Whitney) unpaired Student’s t test (*P < 0.05; **P < 0.01). Teff, T effector.
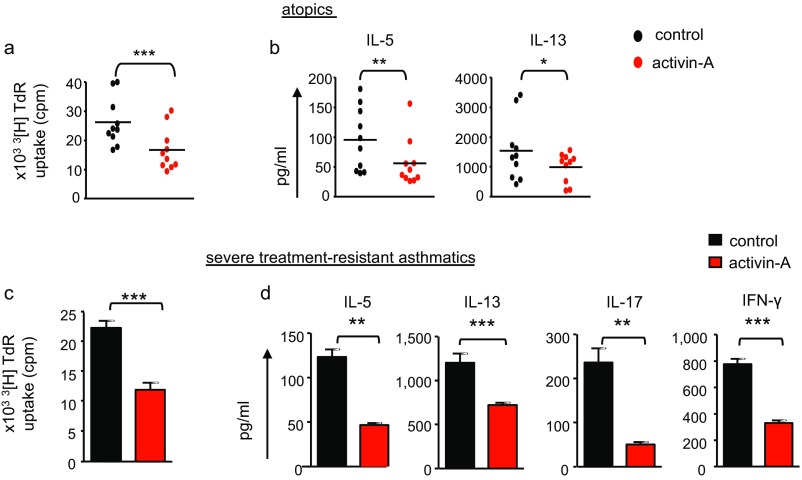
Finally, to investigate whether our findings had potential clinical relevance, we examined the effects of activin-A on the suppression of T-cell responses to mixed grasses extract, a clinically relevant allergen, from individuals with atopy as well as distinct asthma severities. Interestingly, activin-A induced a significant decrease in robust proliferative responses of CD4+ T cells to allergen in atopic individuals and in subjects with mild/moderate and severe asthma (Fig. 7 C and D and Fig. S12 A and B). This decrease was accompanied by a substantial down-regulation in the release of IL-5 and IL-13 (Fig. 7 C and D and Fig. S12 A and B). Notably, activin-A also restrained T-cell responses, including IL-17 production, in a small group of patients with severe, steroid-refractory asthma (Fig. S12 C and D).
Fig. S12.
Activin-A restrains allergen-driven T-cell responses from asthmatic individuals with distinct disease severities. (A and B) CD4+ T cells were isolated from the peripheral blood of atopics and stimulated with mitomycin-treated, CD3-depleted APCs and mixed grasses extract. 3[H]Thymidine incorporation (A) and cytokine release (B) are depicted. Data are mean ± SEM of triplicate wells and are pooled from n = 10 donors. (C and D) CD4+ T cells were isolated from the peripheral blood of subjects with severe, steroid-resistant asthma and stimulated as above. 3[H]Thymidine incorporation (C) and cytokine release (D) are depicted. Data are mean ± SEM of triplicate wells and are pooled from n = 5–6 donors. Statistical significance was obtained using the Mann–Whitney test (*P < 0.05; **P < 0.01; ***P < 0.001).
Discussion
In the present studies, we uncover activin-A as an inducer of human Tr1-like cells that exhibit highly suppressive functions against allergen-induced naive and effector Th2 responses. Moreover, our data underscore a crucial role for IRF4 in human Tr1 cell biology, showing that in response to activin-A, IRF4 assembles into a multipartite molecular complex with AhR and controls Tr1-cell differentiation and effector functions. Of translational importance, we demonstrate that upon therapeutic transfer in a humanized model of allergic asthma, act-A–iTr1 cells confer protection against cardinal disease manifestations. Overall, our studies highlight human act-A–iTr1 cells as potent therapeutic targets that can be used for the design of novel cell-based immunotherapies aiming to control allergic diseases.
Using our established in vitro human T-cell priming model that closely resembles the allergen sensitization phase, we demonstrated that activin-A significantly inhibits T-cell proliferation and Th2-effector cytokine release and renders human T cells hyporesponsive following reexposure to allergen in vitro and adoptive transfer into humanized mice in vivo. In addition, stimulation with activin-A endowed human naive T cells with key Tr1-cell–associated characteristics, such as expression of ICOS, LAG3, CD49b, and granzyme-B as well as production of copious amounts of IL-10 (25, 29–31). Moreover, act-A–iTr1 cells exhibited potent suppressive functions accompanied by enhanced expansion in culture, compared with IL-10–induced Tr1 cells. Interestingly, RNA-seq analysis of the global gene expression profile of human act-A–iTr1 cells, compared with the recently identified LAG3+CD49b+ Tr1 cells, uncovered several similarities in their transcriptional signatures. Seminal studies have documented a critical role for activin-A in instructing the differentiation of Tfh cells in humans and of Th9 and induced Foxp3+ Treg cells in mice (17, 24, 32). Considering that several cytokines induce the differentiation of Th cells into distinct subsets depending on the context and the presence of other cytokines in the micromilieu (33), our findings demonstrating that activin-A can drive the differentiation of human Tr1-like cells extend the findings of previous reports.
An important finding of the present studies is the newly identified role of IRF4 as an essential component of the transcriptional network through which activin-A instructs human Tr1-cell differentiation. IRF4 contributes to the generation of several Th subsets, such as Th1, Th2, Th17, and T follicular helper cells; still, its involvement in human Tr1 cell biology has not been previously described (34, 35). Our findings demonstrate that IRF4 knockdown impairs activin-A–induced IL10 and ICOS up-regulation, decreases the percentages of LAG3+CD49b+ T cells, and compromises the suppressive functions of human act-A–iTr1 cells. Moreover, using sequential ChIP experiments and computational studies, we provide evidence that IRF4 forms a transcriptional complex with AhR and ARNT that seems to be important for activin-A–induced ICOS up-regulation in Tr1 cells. Considering that AhR and ARNT activate target gene expression through direct interactions with other transcription factors (26), our data illuminate IRF4 as a key partner in AhR/ARNT-driven ICOS regulation. Interestingly, ablation of a conditional IRF4 allele in mouse Foxp3+ Treg cells reduces ICOS, Gzmb, and IL-10 expression and decreases their inhibitory capacity upon adoptive transfer in vivo, also implicating IRF4 in ICOS regulation and Treg-cell functions (36). Taken together, it becomes evident that IRF4 represents a central nodule of the transcriptional machinery that instructs effector molecule expression and suppressive functions in both Foxp3+ Treg and Foxp3− Tr1 cells.
Our results also showed that inhibition of AhR signaling abrogates activin-A–induced IL10 and ICOS up-regulation and impairs the inhibitory capacities of human act-A–iTr1 cells. Seminal studies have demonstrated that, depending on the stimuli, signaling through AhR drives the differentiation of human naive T cells into either Foxp3+ Treg cells or Foxp3− Tr1 cells (8, 9). Our studies strengthen this notion, adding activin-A to the team of factors that use AhR-induced signals for the differentiation of human Tr1 cells. Moreover, because in the presence of TGF-β, AhR activation leads to the generation of Foxp3+ Treg cells, our findings reinforce the concept that although belonging to the same superfamily, activin-A and TGF-β exert disparate nonredundant functions on human T-cell responses, instructing the differentiation of distinct Treg-cell subsets (18). In agreement, neutralization of TGF-β did not affect an activin-A–mediated increase in the percentages of CD4+IL-10+ T cells. In addition, given that AhR interacts with c-Maf to control IL-27–driven IL-10 expression in mouse CD4+ T cells, interactions between these transcription factors may also operate in our settings and will be investigated in future studies (9).
A striking finding of our studies was that in vivo administration of act-A–iTr1 cells prevented the development of allergen-driven airway inflammation and ameliorated disease severity in a humanized model of allergic asthma. Of high relevance to clinical settings, we also showed that mice receiving act-A–iTr1 cells during allergen inhalational challenge and after the establishment of allergic inflammation exhibited dampened Th2-effector responses and attenuated pulmonary inflammation, suggesting that this regime may have therapeutic applications. Notably, the inhibitory functions of act-A–iTr1 cells largely depended on IL-10 signaling in vivo, further emphasizing the crucial role of IL-10 in the establishment of tolerance in the airway (4, 37). In severe asthma, T-effector cells are overactivated and often become resistant to mainstay immunosuppressive therapies, including corticosteroids. Given that activin-A was able to restrain allergen-driven responses by T-effector cells from asthmatic individuals across the spectrum of disease severity, our studies may mark this cytokine as a potentially attractive therapeutic target for asthma.
In conclusion, we have identified an activin-A–induced, IRF4/AhR-dependent transcriptional network that instructs the differentiation of human Tr1 cells suppressive against Th2-cell–mediated allergic responses. Considering that adoptive T-cell therapies are underway to control immune-mediated diseases, our findings illuminate act-A–iTr1 cells as targets that can be exploited for the control of asthma and other human diseases associated with dysregulated immunity.
Materials and Methods
Study Population.
Peripheral blood samples were obtained from 13 atopic individuals without asthma and 22 individuals with atopic asthma (mild/moderate and severe) at the “Sotiria” Athens Chest Hospital, the Allergology Department of the General Military Hospital of Athens, and the Lugano and Milano University Hospitals. Asthma was defined according to the latest guidelines from the Global Strategy for Asthma Management and Prevention, Global Initiative for Asthma 2015 (ginasthma.org/2017-gina-report-global-strategy-for-asthma-management-and-prevention). Severe asthma was defined according to the latest American Thoracic Society (ATS)/European Respiratory Society (ERS) guidelines (38). Subjects did not receive treatment with anti-IgE or anti–IL-5 monoclonal antibodies throughout the study. Atopic and asthmatic individuals had a positive skin prick test result (wheal size ≥ 3 mm) to LPS-free mixed grasses extract, and/or increased allergen-specific IgE and blood samples were taken out of season. On entering the study, subjects were stable. Individuals with no clinical features of infection for at least 4 wk before the start of the study and none throughout the study period were included. Patients with autoimmune disorders, cancer, and/or other respiratory diseases were excluded. The study was conducted according to the principles described in the Declaration of Helsinki, and participants signed a written informed consent form before inclusion in the study approved by the ethics committee of “Sotiria” Athens Chest Hospital, Cantonal Ethical Committee of Cantone Ticino, ethics committee of “401 General Military Hospital of Athens”, and Lugano and Milano University Hospitals.
Mice.
Female (6–8 wk old) NOD-SCID mice were obtained from The Jackson Laboratories, maintained at the Biomedical Research Foundation of the Academy of Athens Animal Facility, and regularly checked for the absence of serum Igs. Animal handling and procedures were in accordance with the US NIH Statement of Compliance (Assurance) with Standards for Humane Care and Use of Laboratory Animals (no. A5736-01) and with the European Union Directive 86/609/European Economic Community(EEC) on protection of animals used for experimental purposes.
Isolation and Culture of Peripheral Blood T Cells.
Peripheral blood mononuclear cells (PBMCs) were obtained by Lymphoprep (Axis-Shield Diagnostics) and/or Histopaque (Sigma–Aldrich) density-gradient centrifugation. Naive CD4+ T cells were isolated using magnetic microbeads (CD4+CD25−CD45RO−; Miltenyi Biotec) or by sorting with a FACS Aria cell sorter (BD Biosciences). Cell viability staining was performed, and dying cells were excluded from analyses. CD3-depleted PBMCs (Miltenyi Biotec) or CD14+ monocytes (Miltenyi Biotec) were treated with mitomycin-C (Sigma–Aldrich) and used as APCs. Similar results were obtained using either APC population, and results were pooled for statistical analysis. Naive CD4+ T cells (2 × 104 cells) were stimulated with autologous APCs (2 × 104 cells) loaded with LPS-free mixed grasses extract [ALK-Abello, 1,000 standardized quality-units (SQ-U)/mL; kindly provided by P. Wurtzen, ALK-Abello, Hørsholm, Denmark] in the presence of PBS (Tconv cells) or recombinant activin-A (act-A-iTr1 cells, 50 ng/mL; R&D Systems) for 7–9 d. In all in vitro stimulations, mixed grasses extract was used. In certain experiments, naive CD4+ T cells were stimulated with plate-bound anti-human CD3 (1 μg/mL, OKT3; BD Biosciences) and soluble anti-human CD28 (1 μg/mL, CD28.2; BD Biosciences). Cells were cultured in RPMI 1640/GlutaMAX supplement medium with 10% FBS and 100 units/mL penicillin and 100 μg/mL streptomycin (all from Gibco) in 96-well, U-bottomed Nunc MicroWell Plates with Nunclon Delta Surface (Thermo Scientific). In studies examining IL-10 production, cells were stimulated for an additional 24–48 h. Dose–response studies were performed to determine the optimal doses of activin-A, allergen, and anti-CD3/CD28 antibodies. Human recombinant IL-2 (20 international units/mL; R&D Systems); the Smad3 inhibitor SIS3 (10 nM; Sigma–Aldrich); CH-223191 (3 μM; Calbiochem); and neutralizing antibodies against ALK4 (20 μg/mL; R&D Systems), IL-10R (20 μg/mL; R&D Systems), and TGF-β (5 μg/mL; R&D Systems), or the respective Ig controls, were added in cell cultures. To obtain CM, CD4+ T cells were isolated from activin-A– or control-treated cultures, washed, and stimulated overnight with anti-CD3/CD28 antibodies. CM was then added in cocultures of naive CD4+ T cells and allergen-loaded APCs.
In other experiments, total CD4+ T cells (5 × 104 cells) obtained from atopic and asthmatic individuals were isolated using magnetic microbeads (Miltenyi Biotec) or the Dynal CD4 Positive Isolation Kit (Dynabeads; Invitrogen) and stimulated with autologous APCs (2 × 104 cells) loaded with mixed grasses extract for 4–5 d.
In experiments investigating T-cell hyporesponsiveness, CD4+ T cells were isolated from primary induction cultures, labeled using a CellTrace CFSE Proliferation Kit (Molecular Probes), and restimulated either with fresh allergen-loaded APCs or with anti-CD3/CD28 antibodies for another 5–6 d.
In experiments investigating in vitro stability during prolonged culture, CD4+ T cells were isolated from primary cultures as above and stimulated every week with freshly isolated, allergen-loaded APCs for a period of 4 wk and in the presence of IL-2.
For evaluation of Ag specificity, naive CD4+ T cells were cultured with allergen-loaded, mitomycin-treated APCs in the presence of control (PBS) or activin-A for two rounds of stimulation (7 d each); then, CD4+ T cells were isolated and cultured with fresh APCs in the presence of tetanus toxoid (5 μg/mL; Alexis Biochemicals) for another 6 d.
For in vitro human Tr1-cell generation, naive CD4+ T cells were stimulated with plate-bound anti-CD3 and soluble anti-CD28 antibodies in the presence of mitomycin-treated, CD3-depleted PBMCs and recombinant human (rh) IL-10 (100 ng/mL; R&D Systems) and rhIFN-α2b (5 ng/mL; Merck). After 7 d, T cells were restimulated under identical conditions for an additional 7 d. At the end of the 14 d of culture, IL-10, CD49b, and LAG-3 expression was analyzed by FACS. The fold expansion of T cells was calculated by dividing the absolute number of cells after 14 d of in vitro culture by the absolute number of cells at the start of the culture.
In Vitro Suppression Assays.
Tconv or act-A–iTr1 cells (2 × 104 cells) were stained with CFSE (Molecular Probes, Life Technologies) and cocultured with Cell Trace Violet-labeled (Molecular Probes, Life Technologies) autologous naive or Th2-effector/memory cells (at a 1:1 ratio) and allergen-loaded, autologous, mitomycin-treated APCs for 4 d. A Transwell system (R&D Systems) was used, wherein naive CD4+ T cells, APCs, and allergen were placed in the bottom chambers and act-A–iTr1 or Tconv cells, APCs, and allergen were added in the upper chambers. T-cell proliferation was evaluated by 3[H]thymidine incorporation or CFSE and Cell Trace Violet staining and analyzed by flow cytometry.
Neutralizing antibodies against activin-A (20 μg/mL; R&D Systems), IL-10R (30 μg/mL; R&D Systems), ICOS ligand (ICOSL) (25 μg/mL; eBioscience), and TGF-β (7.5 μg/mL; R&D Systems) were added in secondary suppressive cultures. In other experiments, Tconv, act-A–iTr1, or IL-10–Tr1 cells were cocultured with naive CFSE-labeled CD4+ T responders in the presence of anti-human CD3 (1 μg/mL) and soluble anti-human CD28 (1 μg/mL) for 5 d. The suppression of cell proliferation was calculated by measuring the proliferation [3H]thymidine incorporation or CellTrace proliferation dye dilution-based division index (39) of T responders cultured across Tconv or act-A–iTr1 cells in comparison to T responders alone, according to the formula:
Knockdown with shRNA.
Naive CD4+ T cells (2 × 104 cells) were activated with allergen-loaded APCs (2 × 104 cells) or anti-CD3/CD28 antibodies with or without activin-A for 2.5 d. Cells were spun with lentivirus containing GFP-expressing shRNA against human AHR, IRF4, or a scrambled RNA sequence (Dharmacon) and 8 μg/mL polybrene (Sigma–Aldrich). Following 12 h of incubation (37 °C), T cells were washed and differentiated for 3 d, as described above. Transduced cells were selected by sorting of live GFP+ T cells, using a FACS Aria cell sorter.
Cytokine Analysis.
Cytokines were measured in culture supernatants and mouse BAL using commercially available ELISA kits for human IL-10, IL-5, IL-4, IL-13, IL-17, and IFN-γ (R&D Systems).
Flow Cytometry Analysis.
Cells were stained with fluorescently labeled antibodies to CD4, ICOS, PD-1, CTLA-4, GITR, CD25, CD69, CD127, LAG3, CD49b, CCR3, CCR5, and CRTH2 (all from eBiosciences); CCR4 (BD Pharmingen); and CCR6, CCR7, CXCR3, CD14, CD95, and CD45RA (all from Biolegend). The staining for CD49b and LAG3 was performed at 37 °C for 20 min (25). For intracellular cytokine staining, T cells were restimulated with phorbol 12-myristate 13-acetate (PMA)/ionomycin (Sigma–Aldrich) and Golgi-Stop (BD Biosciences), and stained with antibodies against IL-4, IL-5, and IL-13 (all from eBiosciences); IL-17 (Beckman Coulter); and IFN-γ (Beckman Coulter) according to the manufacturers’ instructions. Foxp3 (eBiosciences) staining was performed according to the manufacturer’s instructions. For IL-10 detection, T cells were restimulated with anti-CD3/CD28 antibodies overnight. FACS acquisition was performed with a Cytomics FC500 cytometer (Beckman Coulter) and an LSRFortessa cytometer (BD Biosciences), and data were analyzed using FlowJo software.
Immunofluorescence Microscopy.
T-cell cytospins (5 × 104 cells per slide) were prepared in poly-l-lysine slides (Thermo Scientific), fixed in 4% (wt/vol) paraformaldehyde (at 4 °C), and incubated in 10% sucrose. Slides were washed twice and blocked in 10% donkey serum, and then diluted in 0.1% Triton-X for 30 min. Slides were incubated overnight at 4 °C with primary biotinylated antibodies against human AhR (Santa Cruz Biotechnology), c-Maf (Santa Cruz Biotechnology), IRF4 (R&D Systems), or control IgG (R&D Systems). Slides were then incubated with fluorescently labeled secondary antibodies (R&D Systems). Nuclear staining and mounting of the slides were carried out using ProLong Gold Anti-Fade Reagent with DAPI (Life Technologies). Image acquisition was performed by a confocal laser scanning microscope (Leica TCS SP5), and images were acquired by differential interference contrast optics. AhR+, c-Maf+, and IRF4+ T cells were counted in each cytospin by two independent observers and expressed as the percentage of total cells (40).
Reconstitution of NOD-SCID Mice.
In the prevention protocol, NOD-SCID mice were reconstituted on day 0 with 2 × 106 Tconv or act-A–iTr1 cells and 2 × 106 autologous APCs, and were immunized (i.p.) with Phleum pratense (5 μg per mouse in 0.2 mL of alum; kindly provided by P. Wurtzen) (28, 41). On day 1, mice received autologous, BrdU-labeled, CD4+ T cells (106 cells). On days 9 and 10, mice were challenged with aerosolized P. pratense; AHR was measured the following day, and mice were killed. In other experiments, mice were administered (i.p.) anti-human IL-10Rα (30 μg per mouse; Biolegend) or Ig control prior to and 1 d following act-A–iTr1 cell (or Tconv cell) transfer. CD4+ T-responder cells were incubated with anti–IL-10Rα antibody or Ig control before transfer. In the therapeutic protocol, NOD-SCID mice were reconstituted on day 0 with naive CD4+ T cells (106 cells) and autologous APCs (106 cells), and were immunized with P. pratense/alum. On day 8, mice received autologous Tconv or act-A–Tr1 cells (2 × 106 cells). Mice were challenged and killed as described above.
AHR.
AHR was measured by whole-body plethysmography (Buxco Research Systems) to calculate enhanced pause (PenH) as described (18, 42).
BAL Analysis.
BAL harvesting was performed as described and kept at −80 °C until use (18, 42).
Lung Histology.
Paraffin-embedded 4-μm lung sections were stained to evaluate cell infiltration, and histological scores were obtained as described (18, 42).
Mouse Cell Culture, Oroliferation, and ex Vivo Suppression Assays.
DLN cells were isolated at the time of euthanasia and cultured at 106 cells per well with 2.5 μg/mL P. pratense. T-cell proliferation was measured by 3[H]thymidine incorporation or BrdU staining, followed by flow cytometry.
Statistics.
Data distribution was assessed with the D’Agostino–Pearson normality test. Analyses were performed using the Student’s t test and the Mann–Whitney test as appropriate, using GraphPad Prism Version 5. AHR was analyzed by two-way ANOVA for repeated measures, followed by the Student’s t test. Any difference with a P value of 0.05 or less was considered as statistically significant (*P < 0.05; **P < 0.01; ***P < 0.001). Additional information regarding the quantitative real-time PCR, RNA-seq and bioinformatics analysis, ChIP, in silico regulatory region analyses of selected genes, and in silico structural study of AhR/IRF4/ARNT interactions is provided in SI Materials and Methods.
SI Materials and Methods
Quantitative Real-Time PCR.
Human naive CD4+ T cells were stimulated with allergen-loaded, mitomycin-treated APCs or anti-CD3/CD28 antibodies in the presence of PBS (Tconv) or activin-A (act-A–iTr1) cells for 3 d. Total RNA was isolated in TRI Reagent (Molecular Research Center) and reverse-transcribed using SuperScript II (Invitrogen) according to the manufacturer’s recommendations. The relative gene expression over the expression of GAPDH was calculated as described (43). Primer sequences are presented in Table S1.
Table S1.
Primer pairs that were used for the real-time PCR analyses
| Genes | Forward primer | Reverse primer |
| AHR | CCGTGTCGATGTATCAGTGC | GCCTGGCAGTACTGGATTGT |
| CYP1A1 | CAGCTGACTTCATCCCTATTC | AGCTGGACATTGGCGTTCTCA |
| MAF | ACGAGAAGTTGGTGAGCAGC | TTCCAAAATGTGGCGTATCC |
| IRF4 | CATGACAACGCCTTACCCTT | AAACGTCATGGGACATTGGT |
| ICOS | GGACCATTCTCATGCCAACT | AAAGGCTGCACATCCTATGG |
| IL-10 | GGCGCTGTCATCGATTTCTT | TTGGAGCTTATTAAAGGCATTCTTC |
| GZMB | GTGCAGGAAGATCGAAAGTGC | GGGCCACCTTGTTACACACA |
| LAG3 | GGCAATCATCACAGTGACTCC | GAGCTCCACACAAAGCGTTC |
| FOXP3 | GAAACAGCACATTCCCAGAGTTC | ATGGCCCAGCGGATGAG |
| GATA3 | GCGGGCTCTATCACAAAATGA | CTCTCCTGGCTGCAGACAGC |
| TBX21 | GATGCGCCAGGAAGTTTCAT | GCACAATCATCTGGGTCACATT |
| RORC | GCAGCGCTCCAACATCTTCTC | GCACACCGTTCCCACATCTC |
| IL-4 | CTTTGCTGCCTCCAAGAACAC | GCGAGTGTCCTTCTCATGGT |
| IL-5 | TACGTGTATGCCATCCCCAC | TTCAGTGCACAGTTGGTGATTT |
| IL-13 | TGACAGCTGGCATGTACTGTG | ATGCAAGCTGGAAAACTGCC |
| IL-17A | CCCGGACTGTGATGGTCAAC | TCTCTTGCTGGATGGGGACA |
| IFNG | GAGTGTGGAGACCATCAAGGA | CTGTTTTAGCTGCTGGCGAC |
| GAPDH | GCAAATTCCATGGCACCGT | TCGCCCCACTTGATTTTGG |
RNA-Seq and Bioinformatics Analysis.
RNA-seq libraries were generated using Illumina’s TruSeq RNA Library Preparation Kit v2 (Illumina), and the sequencing was implemented with a HiSeq 2000 system (Illumina) by the European Molecular Biology Laboratory. Two biological replicates were used for each condition. For analyzing the sequencing files of Tconv and act-A–iTr1 human cells, the following pipeline was performed: For each biological replicate, fastq quality control was assessed; when required, Illumina adapters were clipped, and bases with quality less than 10 were trimmed from the 3′ end of the reads, allowing reads of a minimum length of 20 bases. The high-quality reads were then aligned against the UCSC Genome Browser hg19 human reference genome build, using TopHat2 (44) with default settings and keeping only uniquely aligned reads. Alignments were then processed for differential expression analysis using the metaseqR (45) Bioconductor package. Read counts were calculated on the Ensembl exon set version 69, and effective library sizes were estimated to normalize the counts table [DESeq method (46)]. For differential expression statistical testing, DESeq and edgeR (47) bioconductor packages were combined to calculate a “meta P-value,” resulting in 513 significantly differentially regulated genes. To compare gene expression patterns between human act-A–iTr1 cells and Tr1 cells (25), reads per kilobase per million mapped reads (RPKM) and robust multi-array average (RMA) signal intensity values were scaled to have a mean of 0 and an SD of 1. Expression values of the two platforms were plotted in a heat map for 48 RNA-seq differentially expressed genes also present in the microarray study and sorted increasingly with respect to the RNA-seq comparison log2 (FC) value. Bioinformatics analysis and visualizations were applied through custom Perl and R scripts. Microarray data were downloaded from the Gene Expression Omnibus (accession no. GSM1295189).
CHiP.
Naive CD4+ T cells were differentiated for 6 d with anti-human CD3/CD28 antibodies in the presence or absence of activin-A. T cells were fixed with 1% formaldehyde, quenched with 0.125 M glycine, and lysed to release chromatin, which was sheared to a mean size of 150- to 300-bp fragments by sonication. Samples were incubated overnight with anti-human IRF4, AhR, and c-Maf antibodies (Santa Cruz Biotechnology). Normal rabbit or goat IgG (Santa Cruz Biotechnology) served as a negative control. Magnetic Dynal G beads (Invitrogen) were incubated with the immunocomplexes. For sequential ChIPs, the initial ChIP was performed with the anti-IRF4 antibody, chromatin was eluted with DTT, and a second immunoprecipitation was performed using the anti-AhR or anti–c-Maf antibody. Immunoprecipitated DNA was extracted by reverse cross-linking at 65 °C and proteinase K treatment, followed by AMPure magnetic beads purification. The relative enrichment of ChIP versus IgG (relative to input DNA) was determined and calculated by quantitative RT-PCR for the regions of interest. The following primer pairs were used: IL10, 5′-GTCTTGGGTATTCATCCCAGGTTGGGG-3′ (forward) and 5′-CTGTGGGTTCTCATTCGCGTGTTCCTA-3′ (reverse); ICOS (−2 kb), 5′- AACCAACTTTCCATGCCCCA-3′ (forward) and 5′-CCATGCCTCGCCTTCTCAAA-3′ (reverse); ICOS (−1 kb), 5′-GCCTCCTCCTTACCTTGCAG-3′ (forward) and 5′-TCTGAACGCGCTTCCAAGAT-3′ (reverse); and ICOS [transcription start site (TSS)], 5′-AACATGAAGTCAGGCCTCTGG-3′ (forward) and 5′-ACATGAGTGACATGGTGCCC-3′ (reverse).
In Silico Regulatory Region Analyses of Selected Genes.
The analyses aimed at the identification of the consensus binding sequences of c-Maf, AhR, and IRF4 in human ICOS and IL10. The ICOS and IL10 sequences, with the accession numbers ENSG00000163600 and ENSG00000136634, respectively, were extracted from the Ensembl database (useast.ensembl.org/index.html). The retrieved regulatory regions extending 3,000 bp upstream to 3,000 bp downstream of every known TSS of each gene were submitted to the MatInspector platform in the Genomatix database (www.genomatix.de/en/index.html) to identify putative transcription factor-binding sites (TFBSs) for all of the transcription factors of Matrix Family Library Version 9.0, which contains 1,381 weight matrices. To filter the output of the algorithm, the experimentally verified TFBSs of the known regulators in IL10 (8) were considered as evidence for the true-positive results. Any predicted TFBSs with P values and scores beyond the values of the experimentally verified ones were excluded from the results.
In Silico Structural Study of AhR/IRF4/ARNT Interactions.
AhR and IRF4 were requested from the Protein Data Bank (PDB) (www.rcsb.org/pdb/home/home.do), and the most relevant solved structures (with PDB ID codes 4ZRB and 3DSH, respectively) sharing sequence similarity greater than 30% with these factors were retrieved. Briefly, we used the solved structure of the heterodimeric HIF-1a/ARNT complex with HRE DNA (PDB ID code 4ZPR) as a template to construct the structure of AhR, based on previous studies showing that AhR can functionally interact with ARNT (26). Homology modeling was carried out through the SWISS-MODEL platform (https://swissmodel.expasy.org/), and the models were subsequently evaluated with Anolea, QMEAN6, Gromos, DFire, and Procheck (swissmodel.expasy.org/workspace/?func=tools_structureassessment1). The models of AhR and IRF4 were used for molecular docking via the ZDOCK server (zdock.umassmed.edu/), whereas Yet Another Scientific Artificial Reality Application (YASARA; www.yasara.org/) was used to perform the molecular dynamics simulations and the energy minimization of the ZDOCK output complex, as well as the recalculation of its energy. The duration of the molecular dynamics simulation studies was 10 ns. Superimpositions, rmsd calculations, and visualization of the structures were performed using the DeepView-Swiss-PdbViewer.
Acknowledgments
We thank Professor Clare Lloyd and Dr. P. Verginis for critical evaluation of the manuscript. G.X. is supported by Academia-Industry Co-Operation Program Grant 09-12-1074, by “Research Excellence” Grant 5035 funded by the General Secretariat for Research and Technology, and by a European Respiratory Society “Romain Pauwels” Research Excellence Award. S.T. is supported by “Fondation Sante” Biomedical Research Grants, a European Academy of Allergy and Clinical Immunology Fellowship, and a European Respiratory Society Fellowship. M.S. is supported by “Research Excellence” Grant 5035 funded by the General Secretariat for Research and Technology and by a European Respiratory Society Fellowship. I.M. is supported by Academia-Industry Co-Operation Program Grant 09-12-1074 funded by the General Secretariat for Research and Technology. K.S. is supported by a Hellenic Thoracic Society Fellowship. A.I.T. is supported by the “Theoharis” Foundation, Biomedical Research Foundation of the Academy of Athens.
Footnotes
The authors declare no conflict of interest.
This article is a PNAS Direct Submission.
Data deposition: We have generated and analyzed RNA-sequencing data of activin-A–induced Tr1-like cells, which we have deposited in the Gene Expression Omnibus (GEO) database, www.ncbi.nlm.nih.gov/geo (accession no. GSE95820).
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1616942114/-/DCSupplemental.
References
- 1.Barnes PJ. Pathophysiology of allergic inflammation. Immunol Rev. 2011;242(1):31–50. doi: 10.1111/j.1600-065X.2011.01020.x. [DOI] [PubMed] [Google Scholar]
- 2.Shevach EM, Thornton AM. tTregs, pTregs, and iTregs: Similarities and differences. Immunol Rev. 2014;259(1):88–102. doi: 10.1111/imr.12160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Curotto de Lafaille MA, et al. Adaptive Foxp3+ regulatory T cell-dependent and -independent control of allergic inflammation. Immunity. 2008;29(1):114–126. doi: 10.1016/j.immuni.2008.05.010. [DOI] [PubMed] [Google Scholar]
- 4.Roncarolo MG, Gregori S, Bacchetta R, Battaglia M. Tr1 cells and the counter-regulation of immunity: Natural mechanisms and therapeutic applications. Curr Top Microbiol Immunol. 2014;380:39–68. doi: 10.1007/978-3-662-43492-5_3. [DOI] [PubMed] [Google Scholar]
- 5.Josefowicz SZ, et al. Extrathymically generated regulatory T cells control mucosal TH2 inflammation. Nature. 2012;482(7385):395–399. doi: 10.1038/nature10772. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Akdis M, et al. Immune responses in healthy and allergic individuals are characterized by a fine balance between allergen-specific T regulatory 1 and T helper 2 cells. J Exp Med. 2004;199(11):1567–1575. doi: 10.1084/jem.20032058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Campbell JD, et al. Peptide immunotherapy in allergic asthma generates IL-10-dependent immunological tolerance associated with linked epitope suppression. J Exp Med. 2009;206(7):1535–1547. doi: 10.1084/jem.20082901. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Gandhi R, et al. Activation of the aryl hydrocarbon receptor induces human type 1 regulatory T cell-like and Foxp3(+) regulatory T cells. Nat Immunol. 2010;11(9):846–853. doi: 10.1038/ni.1915. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Apetoh L, et al. The aryl hydrocarbon receptor interacts with c-Maf to promote the differentiation of type 1 regulatory T cells induced by IL-27. Nat Immunol. 2010;11(9):854–861. doi: 10.1038/ni.1912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Veldhoen M, Hirota K, Christensen J, O’Garra A, Stockinger B. Natural agonists for aryl hydrocarbon receptor in culture medium are essential for optimal differentiation of Th17 T cells. J Exp Med. 2009;206(1):43–49. doi: 10.1084/jem.20081438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Hedger MP, de Kretser DM. The activins and their binding protein, follistatin-Diagnostic and therapeutic targets in inflammatory disease and fibrosis. Cytokine Growth Factor Rev. 2013;24(3):285–295. doi: 10.1016/j.cytogfr.2013.03.003. [DOI] [PubMed] [Google Scholar]
- 12.Kariyawasam HH, Semitekolou M, Robinson DS, Xanthou G. Activin-A: A novel critical regulator of allergic asthma. Clin Exp Allergy. 2011;41(11):1505–1514. doi: 10.1111/j.1365-2222.2011.03784.x. [DOI] [PubMed] [Google Scholar]
- 13.Matzuk MM, et al. Functional analysis of activins during mammalian development. Nature. 1995;374(6520):354–356. doi: 10.1038/374354a0. [DOI] [PubMed] [Google Scholar]
- 14.Matzuk MM, Kumar TR, Bradley A. Different phenotypes for mice deficient in either activins or activin receptor type II. Nature. 1995;374(6520):356–360. doi: 10.1038/374356a0. [DOI] [PubMed] [Google Scholar]
- 15.Chng Z, Vallier L, Pedersen R. Activin/nodal signaling and pluripotency. Vitam Horm. 2011;85:39–58. doi: 10.1016/B978-0-12-385961-7.00003-2. [DOI] [PubMed] [Google Scholar]
- 16.Hardy CL, Rolland JM, O’Hehir RE. The immunoregulatory and fibrotic roles of activin A in allergic asthma. Clin Exp Allergy. 2015;45(10):1510–1522. doi: 10.1111/cea.12561. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Huber S, et al. Activin a promotes the TGF-beta-induced conversion of CD4+CD25- T cells into Foxp3+ induced regulatory T cells. J Immunol. 2009;182(8):4633–4640. doi: 10.4049/jimmunol.0803143. [DOI] [PubMed] [Google Scholar]
- 18.Semitekolou M, et al. Activin-A induces regulatory T cells that suppress T helper cell immune responses and protect from allergic airway disease. J Exp Med. 2009;206(8):1769–1785. doi: 10.1084/jem.20082603. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Kariyawasam HH, et al. Activin and transforming growth factor-beta signaling pathways are activated after allergen challenge in mild asthma. J Allergy Clin Immunol. 2009;124(3):454–462. doi: 10.1016/j.jaci.2009.06.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Karagiannidis C, et al. Activin A is an acute allergen-responsive cytokine and provides a link to TGF-beta-mediated airway remodeling in asthma. J Allergy Clin Immunol. 2006;117(1):111–118. doi: 10.1016/j.jaci.2005.09.017. [DOI] [PubMed] [Google Scholar]
- 21.Zielinski CE, et al. Pathogen-induced human TH17 cells produce IFN-γ or IL-10 and are regulated by IL-1β. Nature. 2012;484(7395):514–518. doi: 10.1038/nature10957. [DOI] [PubMed] [Google Scholar]
- 22.Becattini S, et al. T cell immunity. Functional heterogeneity of human memory CD4+ T cell clones primed by pathogens or vaccines. Science. 2015;347(6220):400–406. doi: 10.1126/science.1260668. [DOI] [PubMed] [Google Scholar]
- 23.Hammad H, et al. Monocyte-derived dendritic cells induce a house dust mite-specific Th2 allergic inflammation in the lung of humanized SCID mice: Involvement of CCR7. J Immunol. 2002;169(3):1524–1534. doi: 10.4049/jimmunol.169.3.1524. [DOI] [PubMed] [Google Scholar]
- 24.Locci M, et al. Activin A programs the differentiation of human TFH cells. Nat Immunol. 2016;17(8):976–984. doi: 10.1038/ni.3494. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Gagliani N, et al. Coexpression of CD49b and LAG-3 identifies human and mouse T regulatory type 1 cells. Nat Med. 2013;19(6):739–746. doi: 10.1038/nm.3179. [DOI] [PubMed] [Google Scholar]
- 26.Rowlands JC, McEwan IJ, Gustafsson JA. Trans-activation by the human aryl hydrocarbon receptor and aryl hydrocarbon receptor nuclear translocator proteins: Direct interactions with basal transcription factors. Mol Pharmacol. 1996;50(3):538–548. [PubMed] [Google Scholar]
- 27.Shultz LD, Ishikawa F, Greiner DL. Humanized mice in translational biomedical research. Nat Rev Immunol. 2007;7(2):118–130. doi: 10.1038/nri2017. [DOI] [PubMed] [Google Scholar]
- 28.Perros F, Hoogsteden HC, Coyle AJ, Lambrecht BN, Hammad H. Blockade of CCR4 in a humanized model of asthma reveals a critical role for DC-derived CCL17 and CCL22 in attracting Th2 cells and inducing airway inflammation. Allergy. 2009;64(7):995–1002. doi: 10.1111/j.1398-9995.2009.02095.x. [DOI] [PubMed] [Google Scholar]
- 29.Pot C, Apetoh L, Awasthi A, Kuchroo VK. Induction of regulatory Tr1 cells and inhibition of T(H)17 cells by IL-27. Semin Immunol. 2011;23(6):438–445. doi: 10.1016/j.smim.2011.08.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Pot C, et al. Cutting edge: IL-27 induces the transcription factor c-Maf, cytokine IL-21, and the costimulatory receptor ICOS that coordinately act together to promote differentiation of IL-10-producing Tr1 cells. J Immunol. 2009;183(2):797–801. doi: 10.4049/jimmunol.0901233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Roncarolo MG, et al. Interleukin-10-secreting type 1 regulatory T cells in rodents and humans. Immunol Rev. 2006;212:28–50. doi: 10.1111/j.0105-2896.2006.00420.x. [DOI] [PubMed] [Google Scholar]
- 32.Jones CP, Gregory LG, Causton B, Campbell GA, Lloyd CM. Activin A and TGF-β promote T(H)9 cell-mediated pulmonary allergic pathology. J Allergy Clin Immunol. 2012;129(4):1000–10.e3. doi: 10.1016/j.jaci.2011.12.965. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Zhou L, Chong MM, Littman DR. Plasticity of CD4+ T cell lineage differentiation. Immunity. 2009;30(5):646–655. doi: 10.1016/j.immuni.2009.05.001. [DOI] [PubMed] [Google Scholar]
- 34.Taniguchi T, Ogasawara K, Takaoka A, Tanaka N. IRF family of transcription factors as regulators of host defense. Annu Rev Immunol. 2001;19:623–655. doi: 10.1146/annurev.immunol.19.1.623. [DOI] [PubMed] [Google Scholar]
- 35.Rengarajan J, et al. Interferon regulatory factor 4 (IRF4) interacts with NFATc2 to modulate interleukin 4 gene expression. J Exp Med. 2002;195(8):1003–1012. doi: 10.1084/jem.20011128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Zheng Y, et al. Regulatory T-cell suppressor program co-opts transcription factor IRF4 to control T(H)2 responses. Nature. 2009;458(7236):351–356. doi: 10.1038/nature07674. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Zeng H, Zhang R, Jin B, Chen L. Type 1 regulatory T cells: A new mechanism of peripheral immune tolerance. Cell Mol Immunol. 2015;12(5):566–571. doi: 10.1038/cmi.2015.44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Chung KF, et al. International ERS/ATS guidelines on definition, evaluation and treatment of severe asthma. Eur Respir J. 2014;43(2):343–373. doi: 10.1183/09031936.00202013. [DOI] [PubMed] [Google Scholar]
- 39.Yao Y, et al. Tr1 Cells, but not Foxp3+ regulatory T cells, suppress NLRP3 inflammasome activation via an IL-10-dependent mechanism. J Immunol. 2015;195(2):488–497. doi: 10.4049/jimmunol.1403225. [DOI] [PubMed] [Google Scholar]
- 40.Samitas K, et al. Activin-A is overexpressed in severe asthma and is implicated in angiogenic processes. Eur Respir J. 2016;47(3):769–782. doi: 10.1183/13993003.00437-2015. [DOI] [PubMed] [Google Scholar]
- 41.Sonar SS, et al. Antagonism of TIM-1 blocks the development of disease in a humanized mouse model of allergic asthma. J Clin Invest. 2010;120(8):2767–2781. doi: 10.1172/JCI39543. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Xanthou G, et al. Osteopontin has a crucial role in allergic airway disease through regulation of dendritic cell subsets. Nat Med. 2007;13(5):570–578. doi: 10.1038/nm1580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods. 2001;25(4):402–408. doi: 10.1006/meth.2001.1262. [DOI] [PubMed] [Google Scholar]
- 44.Kim D, et al. TopHat2: Accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013;14(4):R36. doi: 10.1186/gb-2013-14-4-r36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Moulos P, Hatzis P. Systematic integration of RNA-Seq statistical algorithms for accurate detection of differential gene expression patterns. Nucleic Acids Res. 2015;43(4):e25. doi: 10.1093/nar/gku1273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15(12):550. doi: 10.1186/s13059-014-0550-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Robinson MD, McCarthy DJ, Smyth GK. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010;26(1):139–40. doi: 10.1093/bioinformatics/btp616. [DOI] [PMC free article] [PubMed] [Google Scholar]