Abstract
Gaussian Bayesian networks have become a widely used framework to estimate directed associations between joint Gaussian variables, where the network structure encodes the decomposition of multivariate normal density into local terms. However, the resulting estimates can be inaccurate when the normality assumption is moderately or severely violated, making it unsuitable for dealing with recent genomic data such as the Cancer Genome Atlas data. In the present paper, we propose a mixture copula Bayesian network model which provides great flexibility in modeling non-Gaussian and multimodal data for causal inference. The parameters in mixture copula functions can be efficiently estimated by a routine expectation–maximization algorithm. A heuristic search algorithm based on Bayesian information criterion is developed to estimate the network structure, and prediction can be further improved by the best-scoring network out of multiple predictions from random initial values. Our method outperforms Gaussian Bayesian networks and regular copula Bayesian networks in terms of modeling flexibility and prediction accuracy, as demonstrated using a cell signaling data set. We apply the proposed methods to the Cancer Genome Atlas data to study the genetic and epigenetic pathways that underlie serous ovarian cancer.
Keywords: Bayesian network, copula function, the Cancer Genome Atlas, systems biology, serous ovarian cancer
Introduction
In recent years, there has been considerable interest in estimating causal relationships between random variables in a graphical framework. Among several types of graphical models, Bayesian networks (BNs) or, equivalently, probability-weighted directed acyclic graphs (DAGs) have received the most attention due to their simplicity and flexibility in modeling directed associations in the domain.1–4 The associations between d random variables can be summarized by a graph in which V = {Xi | i = 1,2,…, d} represents the set of variables and E ⊂ V × V represents the dependency between variables. Under the acyclicity and Markov assumptions, the joint likelihood function of (X1,…,Xd) in a BN has the following simple form based on the conditional densities:
 |
(1) |
where Πi denotes the parent set of Xi, i.e. ∏i ={Xj | Xj → Xi, Xj ∈V \ {Xi}} (Πi can be empty).
The two most popular BN models are the Gaussian Bayesian network (GBN) model1 and multinomial Bayesian network (MBN) model,5 for continuous variables and discrete variables, respectively. MBN models suffer from a super-exponentially increasing number of parameters, therefore they can only estimate small-scale networks in practice.5 To deal with networks with a relatively large number of nodes, GBN models have been commonly used due to their simple setup and efficient estimation. However, GBN models may fail to identify the true causalities when the joint distribution of interest is far from the multivariate normal, for example, when the underlying distribution is bimodal or multimodal. To tackle the problem of non-normality, several new BN models have been developed, for instance, the logistic BN by Zhang et al.4 which discretizes all the continuous variables to fit a multi-category logit model. Considerable work has also been done in nonparametric and semiparametric estimation of the BN structure. For instance, Voorman et al.6 proposed the following nonparametric model to deal with non-normality issue:
 |
where the fik(·) lies in some function space ℱ. The model by Voorman et al. focuses on estimating the conditional mean E(Xi | Πi). It is essentially a generalized additive model without assuming the independence between εi and fik(·). However, this method relies on a known causal ordering of the true network which is unavailable in most cases.
In 2010, Elidan7 introduced an innovative copula Bayesian network (CBN), a marriage between copula functions and graphical models, which extends conventional BN models to a more flexible framework. A CBN model constructs multivariate distribution with univariate marginals and a copula function C that links these marginals. In general, one can estimate marginals using a parametric or non-parametric approach, and then use a small number of parameters to capture the dependence structure. However, as we shall see in a real data set (Section 4), the regular copula functions such as Gaussian copula may not be able to accurately depict multimodal joint distributions. In addition, the CBN model is subject to the choice of copula function for each local term. Motivated by Elidan’s work, we extend the regular CBNs to a mixture copula Bayesian network (MCBN) using finite mixture models, to better deal with non-normality, multimodality, and heavy tails that are commonly seen in current massive genomic data. The parameters in an MCBN model can be efficiently estimated by a routine EM algorithm. As demonstrated by the real data, the performance of a two-component Gaussian MCBN is generally promising, and our model achieves reasonable accuracy in identifying the true edges in a sparse causal network.
The rest of this paper is organized as follows. In Section 2, we review Elidan’s CBN model, and introduce the proposed MCBN model using a two-component Gaussian mixture for illustration. In Section 3, we present a heuristic local search approach combined with a routine EM algorithm for graph structure estimation, as well as the best-scoring network out of multiple predictions with random initial values. The comparison of three BN models is carried out over a cell signaling data set in Section 4. The new model is applied to the Cancer Genome Atlas (TCGA) data for serous ovarian cancer in Section 5. We discuss and conclude this paper in Sections 6 and 7.
Method
Copula and Elidan’s CBN
Unless otherwise stated, we use , as the marginals, and similarly for multivariate density f (x) ≡ f X (x). The formal definition of a copula function is as follows.
Definition 1. Let (X1, X2,…, Xd) be a vector of continuous random variables and (F(x1), F(x2),… F(xd)) be the marginal distribution functions. The copula function of (X1, X2,…, Xd), C: [0,1]d → [0,1], is defined as the cumulative distribution function (CDF) of (F(X1), F(X2),…, F(Xd)):
 |
(2) |
By definition, a copula function is a multivariate distribution function where the marginals are uniform. By choosing an appropriate copula, one can generate multivariate distribution of any complex form. In practice, one can completely separate the choice of marginals and the choice of dependency patterns between random variables. Sklar’s theorem below guarantees that any multivariate distribution can be expressed with univariate marginals and a copula function which links these variables.
Theorem 1. Let F(x1, x2,…, xd) be a multivariate distribution over real-valued d-dimension random vectors, then there exists a copula function that satisfies
 |
(3) |
Furthermore, the copula function C is unique when the marginal distribution F(xi) is continuous for i ∈{1,2,…, d}.
By taking the first derivative for both sides of Equation (3), we can derive the copula density function defined as . The copula density is simply a ratio between the joint density and the product of all the marginals:
 |
(4) |
An immediate consequence of Equation (4) is that c(F(x1), F(x2),…, F(xd)) = 1 if and only if X1,…, Xd are independent. For a subset of variables (Y, X1,…, Xp), as , the conditional density f (y | x1,…, xp) can be expressed as follows:
 |
(5) |
Motivated by Equations (1) and (5), Elidan proposed a CBN based on the following local density:
 |
(6) |
where .
By Equation (6), we have the following decomposition for the joint density of variables in a BN.
Theorem 2. Let (X1,…, Xd) be d random variables (nodes) in a BN, and πi = {xj | Xj ∈ Πi}. The joint density can be represented as follows:
 |
(7) |
Although the construction of local copulas can significantly reduce the complexity of the structure learning, choosing an appropriate copula for each local term Gc(xi | πi) is essential. Elidan suggested a small set of pre-selected copula functions (or copula families) such as Gaussian copula, Frank’s copula, Ali–Mikhail–Haq (AMH) copula and Gumbel–Barnett (GB) copula. However, as we discuss in Section 4, these regular copula functions might be inadequate to model the complex dependence structure. To this end, we extend the CBN to a more flexible framework using a finite mixture model.
An MCBN
For illustration purposes, we limit ourselves to Gaussian MCBN, but other mixture models such as Gamma mixture and Beta mixture models can be adapted similarly. The K-component Gaussian mixture copula for variables (Y, X1, …, Xp) can be formulated as follows:
 |
where α(k) and denote the weight and CDF of the kth Gaussian component, respectively, and Φ−1 (·) represents the quantile function of N (0,1). The corresponding copula density can be obtained immediately:
 |
where ϕ(·) represents the standard normal density function.
The Gaussian MCBN model above takes advantage of a finite mixture model to better fit the bimodal and multimodal distributions. Similar to Elidan’s CBN, the marginals should be estimated prior to fitting the mixture copula, with either parametric or nonparametric method. We can, for example, fit the marginals using parametric or non-parametric method, then transform (y,x1,…, xp) to (F(y), F(x1),…, F(xp)) using the fitted CDF functions. The transformed values will be used for estimating the copula function. Based on the estimated mixture copula for each local term in BN, we can calculate the joint likelihood by Equation (7).
Graph estimation using EM and local search algorithms
EM algorithm for a finite Gaussian mixture
In this section, we introduce the EM algorithm to estimate the mixture copula for each local term Gc(xi | πi).
For a given variable Xi and its parent set Π, the regular k-means algorithm can provide warm starts for the mean vector µk (of dimension | Πi | +1) and the covariance matrix Σk (of dimension (| Πi | +1)×(| Πi | +1)) for each mixture component, as well as the mixing rate α(k). Let and uj = {uhj}, where xhj is the observed value for variable Xh and sample j, Xh ∈ {Xi, Πi}, j = 1,2,…, N. Let z = (z1,…, zN) be the vector of indicators for the membership of each sample (mutually exclusive and exhaustive), i.e. α(k) = P(zj=k), j = 1,…, N and . Denote Θk = (µk, ∑k) and Θ = {Θk}, the EM algorithm with missing information can be implemented as follows.
- E step. Given the current estimate of all the parameters (α(k), Θ), we compute the weighted membership as follows:

- M step. Use data uj and membership weights to update all the parameters:

Given an estimate of the graph structure and the parameters , the log-likelihood can be written as
 |
where the denominator of Gc (xi | πi), i.e. must be evaluated. Here we use notation pi as the number of parents of Xi, i.e. pi =| Πi |. A simple idea for estimating Gc (xi | πi) is to generate a list of Monte Carlo samples from f (xi), and by the law of large numbers:
 |
where . However, it is noteworthy that drawing samples from f (xi) might be complicated and time-consuming when marginals were estimated with non-parametric method. Further, the likelihood may fail to converge due to the randomness of Gc (xi | πi) estimation. Therefore, for practical consideration, one can directly use all the observations as samples so that the convergence is guaranteed.
Score-based local search for learning MCBN
In this part, we introduce an efficient heuristic search algorithm based on the Bayesian information criterion (BIC) to learn the structure of the underlying network . The BIC score can be evaluated by the following formula:
 |
where represents the log-likelihood function, is the set of all the parameters including the mixing rates, mean vectors, and covariance matrices of Gaussian components, and denotes the total number of free parameters in . We start from a randomly generated network or empty network, and greedily advance through basic edge operations including addition, deletion, and reversal, until the BIC score reaches the minimum.7 Unfortunately, this local search algorithm may easily become trapped in a local maximum due to the high dimensionality and non-convexity of the likelihood function, making it impractical to find the global maximum. Enlightened by one of the reviewers, we conducted the heuristic search algorithm multiple times, each with a random initial value, and the best-scoring network (with minimum BIC score) was returned as the best predicted network.
Comparison with existing models
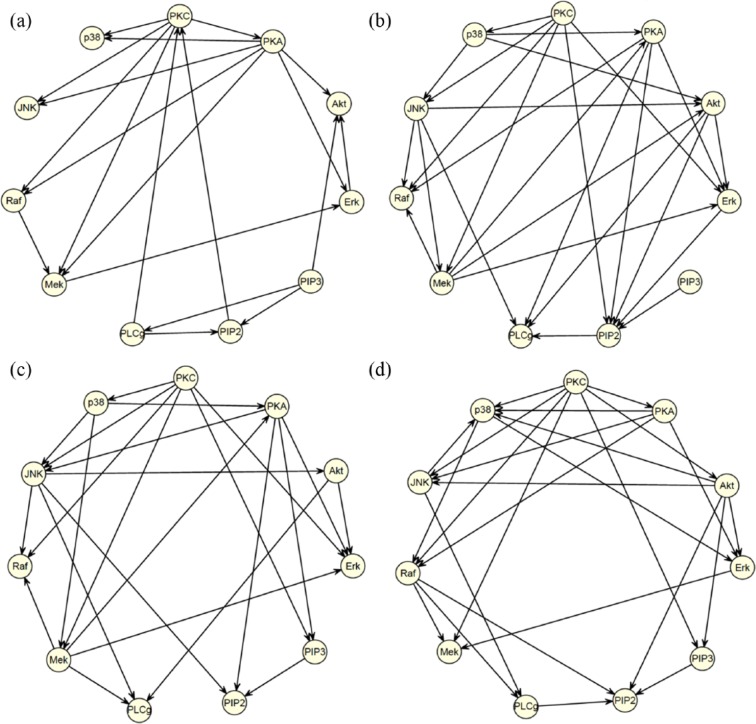
In this section, we compare the proposed MCBN model with two existing BN models: the GBN model and Elidan’s CBN model. We tested the three models using a flow cytometry data set generated by Sachs et al.8 Sachs et al.’s data contains simultaneous measurement on 11 protein and phospholipid components, which was used for elucidating the signaling pathway structure in the cells of the human immune system. The known network shown in Figure 1(a) is a BN containing 11 nodes and 20 causal relations. Each causal edge in the network was well validated by experimental intervention, therefore this network structure is often used as the benchmark to assess the accuracy of different directed or undirected graphical models.
Figure 1.
Comparison of three Bayesian network models on Sachs et al.’s data: (a) the benchmark network; (b) network predicted by the GBN model; (c) network predicted by the Gaussian CBN model; (d) network predicted by the two-component Gaussian MCBN model.
Sachs et al.’s data has both continuous and discrete versions. In our analysis, we used the continuous data which was log-transformed and normalized by subtracting the mean and dividing by standard deviation. Three BN models were then applied to the preprocessed data for network structure learning, with detailed implementation as follows.
GBN: We considered the linear regression setting, , where the graph structure and parameters were estimated by a blockwise coordinate descent (BCD) algorithm proposed by Fu and Zhou.1 It has been shown that the BCD algorithm outperforms the popular PC algorithm9 under regular settings. The intervention information was also incorporated in the modeling and a geometric sequence of 100 candidate tuning parameters (λ1,…, λ100) were predefined (λ1 = 0.001, λ100 = 1). All the calculations were done using the source code provided by the authors (personal communication).
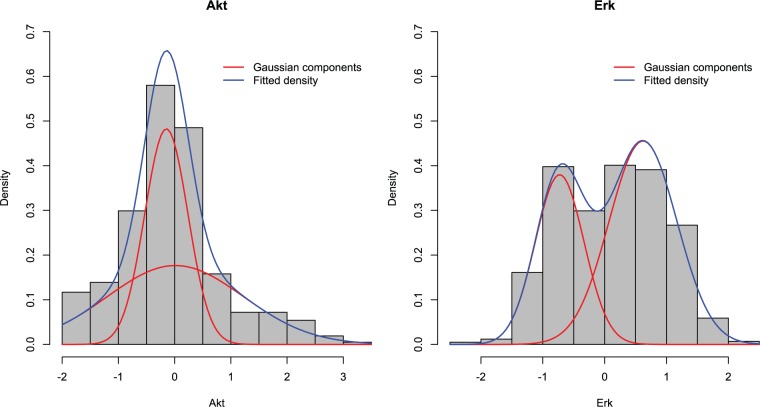
MCBN: For simplicity of calculation, we considered a two-component Gaussian MCBN. The two-component Gaussian mixture model was also applied to the univariate marginals. Figure 2 shows two examples of fitted marginals for proteins Art and Erk.We set the maximum number of parental nodes at 5, i.e. maxi | Πi | ⩽5. The local search algorithm with BIC criterion was applied to BN structure learning, starting from an empty network. In the EM estimation of the copula function, we used k-means (K = 2) to obtain initial values for all the parameters, and used threshold for convergence, where and represent the resulting mixing rates in two consecutive EM runs.
CBN: Elidan’s CBN model can be treated as a special case of MCBN model when the copula density function has only one component (Gaussian copula). For the sake of comparison, all the marginals were also fitted using a two-component Gaussian mixture. The same threshold as in MCBN was used as the convergence criterion of the EM algorithm.
Figure 2.
Fitted marginals by a two-component Gaussian mixture for the abundance of proteins Akt (left) and Erk (right).
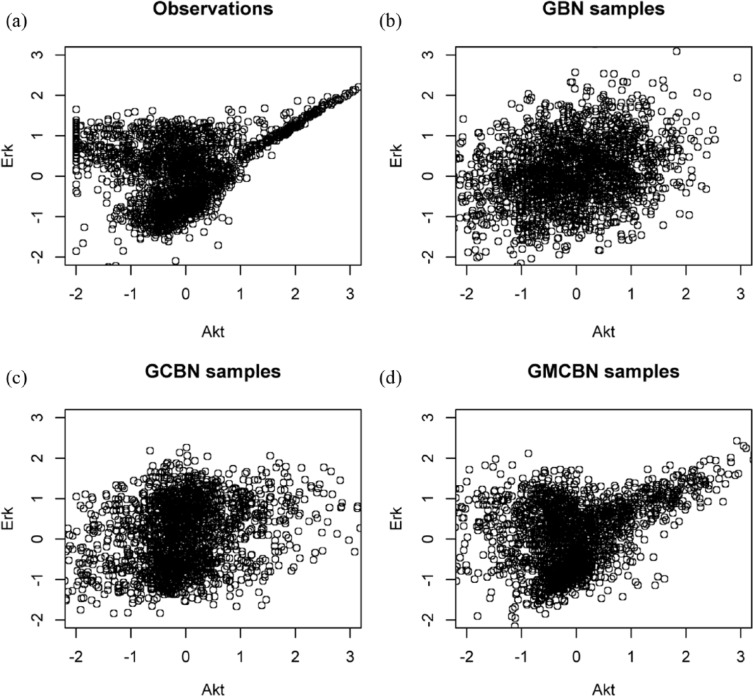
The estimated graphs from the three different models are shown in Figure 1(b)–(d). Table 1 summarizes the true positive rate (TPR), false discovery rate (FDR) as well as running times by the three models (all timing were carried out on a Intel Xeon 3.2 GHz processor). In this comparison, a predicted edge is considered correct if both connection and direction are correct. It can be seen that the proposed MCBN model achieves significantly higher accuracy than the two existing models in terms of TPR and FDR, but it is more computationally expensive than the two simpler BNs. To further improve prediction, we conducted 100 predictions using random initial networks and obtained the best-scoring network, which contained 25 predicted edges. Out of 20 true edges, 13 were correctly identified in the best-scoring network. Furthermore, we compared different models in capturing the dependency pattern between variables. Figure 3 shows the scatterplot of Art and Erk, and the plots of simulated samples from three generative models. Compared with other models, the two-component Gaussian MCBN better depicted the multimodal dependency between Akt and Erk.
Table 1.
Comparison of three different BN models.
Figure 3.
Dependence between proteins Art and Erk: (a) observations; (b) simulated samples from the GBN; (c) simulated samples from the Gaussian CBN; (d) simulated samples from two-component Gaussian MCBN.
To select the most confident edges, we calculated the log-likelihood decrease by removing one edge from the network. We found that an edge giving more likelihood increase has higher probability to be a true edge in the network. For instance, we selected the 10 most confident edges based on the likelihood change, and seven of them turned out to be true edges including Akt→Erk, PKC→P38, PIP3→PIP2, PKA→Raf, PKC→JNK, PKC→Raf, and PLCg→PIP2. In addition, we evaluated the performance of our model in predicting the network skeleton (undirected edges). The proposed MCBN was compared with two simple alternatives including Pearson’s correlation and Spearman’s correlation. In this comparison, a predicted edge is considered correct as long as the connection is correct. Figure 4 shows the undirected networks by three approaches, and the TPR/FDR are summarized in Table 2.
Figure 4.
Comparison of three undirected networks: (a) skeleton of the known network presented in Figure 1(a); (b) network consisting of the top 25 edges based on Pearson’s correlation coefficient; (c) network consisting of the top 25 edges based on Spearman’s correlation coefficient; (d) skeleton of the network predicted by the MCBN model presented in Figure 1(d).
Table 2.
Comparison with Pearson’s and Spearman’s methods.
Application to TCGA ovarian cancer data
In this section, we applied the proposed MCBN to TCGA data,10 to study the interactions between oncomarkers that are associated with serous ovarian cancer. The TCGA data is one of the most comprehensive cancer genomic data sets, with more than 30 cancer types and subtypes which include but not limited to ovarian cancer, breast cancer, lung cancer, brain cancer, and liver cancer. The sample sizes range from 50 to 1200 for different cancer types, and each sample is represented by both the molecular profile and clinical information. The molecular profile contains measurements for various types of (epi)genetic factors including gene expression quantification (both microarray and RNA-Seq), DNA methylation, single nucleotide polymorphism (SNP), copy number variation (CNV), somatic mutation, microRNA, etc. The clinical data provide information such as race, gender, tumor stage, outcome of surgery, and resistance to chemotherapy.
The TCGA ovarian cancer data collected 567 tumor samples and 8 organ-specific normal controls. We incorporated three data types into our model including gene expression level, DNA methylation level (in gene promoter region), and CNV. The data were normalized using a quantile normalization method by Bolstad et al.11 and Mai and Zhang12 to correct the bias due to non-biological causes. In addition, we applied an effective method by Hsu et al.13 to remove age and batch effects (three age groups are defined as < 40, [40,70], and > 70 year old). Hsu et al.’s method is essentially a median-matching and variance-matching strategy. For example, the batch-effect-adjusted gene expression values can be obtained as follows:
 |
where gijk represents the expression level of gene i from batch j and sample k, Mij denotes the median of gij = (gij1,…, gijn), Mi denotes the median of gi = (gi1,…, giJ), and are the standard deviation of gi and gij, respectively.
The set of biomarkers was identified by a stepwise correlation-based feature selector (SCBS) by Zhang et al.,4 which mimics the hierarchy of the underlying causal network. The SCBS algorithm starts by selecting the nodes that are strongly associated with the phenotype node and progressively selects the nodes that are associated with the nodes selected in the previous step. This algorithm is more effective in identifying phenotype-associated nodes, especially those nodes that are indirectly associated with the phenotype. By three runs of SCBS, we identified 73 oncomarkers including the expression level of 50 genes, CNV at 15 sites and methylation level at 8 sites. Among the 73 oncomarkers, many were reported previously in the literature including BRCA1,10 BRCA2,10 RB1,14 PTEN,15 and OPCML.16
We then fit an MCBN model to study the regulatory relationships between these oncomarkers. The marginals were fitted by a two-component Gaussian mixture (other mixture models can also be used, e.g. Beta-mixture for DNA methylation). Figures 5 and 6 show several examples of the fitted marginals for TP53 (expression level), SPARC (expression level), BRCA1 (methylation level), and NOTCH3 (methylation level).
Figure 5.
Fitted marginals by a two-component Gaussian mixture for the expression level of gene TP53 (left) and SPARC (right).
Figure 6.
Fitted marginals by a two-component Gaussian mixture for the promoter methylation level of gene BRCA1 (left) and NOTCH3 (right).
In the biological network, we assumed that the genetic or epigenetic change (CNV and DNA methylation) cannot be induced by gene expression, and imposed this constraint into our modeling (note that this assumption is completely from a biological point of view and it can be dropped without affecting our modeling and computation). The predicted graph (in Figure 7, the best-scoring network from 100 predictions) contains 73 nodes connected by 124 directed edges. Many of the edges in the graph can be confirmed in the literature. To name a few, the edge between AURKA and BRCA2 may be due to the fact that a negative regulatory loop exists between AURKA and BRCA2 expression in ovarian cancer.17. The connection between STAT3 and ETV6 was suggested previously that ETV6 is a negative regulator of STAT3 activity.18 The edges between RAB25 (methylation) and RAB25 (expression) and between CSNK2A1 (CNV) and CSNK2A1 (expression) had been reported in several studies.10, 19, 20 Other highly ranked edges (based on likelihood increase) include but are not limited to: STAT3→DLEC1, PTEN→EGFR, RIMBP2→BRCA2, and ARID1A→ERD, which can be confirmed in the literature of cancer biology.10, 21–23 These findings demonstrate the effectiveness of the MCBN model. In addition, as illustrated in Figure 8, the two-component Gaussian MCBN is accurate in depicting the dependency between the gene expression level and methylation level.
Figure 7.
Predicted network by a two-component Gaussian MCBN model, containing the expression level of 50 genes (in light yellow), methylation level at 8 sites (in light green), and CNV at 15 sites (in light blue).
Figure 8.
Dependence between the methylation level and expression level of gene C19orf53: (a) observations; (b) simulated samples from the two-component Gaussian MCBN.
Discussion
In this paper, we have proposed a novel BN model to analyze recent cancer genomic data at the system level. The major innovation of our model is explicitly modeling the multimodal dependency structure between variables through a copula function and more accurately estimating the causal network structure. The parameters in the mixture copula were efficiently estimated by a routine EM algorithm, and the directed network structure was estimated by minimizing the BIC score.
The proposed BN model allows strict probabilistic inference of biological pathways, however, it also has several limitations. First, it lacks flexibility to model the cyclic mechanism due to the acyclicity constraint, for instance, A→B→A, which may exist in a gene regulatory network. Second, the parameter estimation assumes sparsity of the network for computational feasibility. If the true network is dense or locally dense, the weak causations may fail to be detected. Third, due to the model complexity, the implementation of MCBN is more computationally expensive than simpler BN models such as the GBN model and regular copula BN model. For large data sets, one needs to reduce the number of variables by filtering out irrelevant and redundant variables, and then feed the selected variables into the network model for causal inference.
It is noteworthy that the Gaussian MCBN used in the two illustrative examples can be generally adapted to other mixture models such as Gamma mixture and Beta mixture. The number of mixture components can be further increased depending on the complexity of the underlying dependency structure. For a relatively small data set, it is also possible to conduct statistical testing to select the best number of mixture components for each local term, however, this will significantly increase the computational complexity.
Conclusions
Understanding the biological mechanism of cancers has significant practical importance for clinical diagnosis and treatment. In this paper, we developed an MCBN model for causal inference using complex cancer genomic data. The proposed model is based on finite mixture models and copula functions, and it explicitly models multimodality in the data. The graph structure and model parameters can be efficiently estimated by a routine EM approach, embedded in a heuristic search algorithm based on BIC. The prediction could be further improved by selecting the best-scoring model from multiple predictions with random initial values. In addition, we proposed a likelihood-based approach to select the most confident edges. The proposed MCBN model was applied to a flow cytometry data and the TCGA ovarian cancer data for inferring the causal relationships between different biological features. Compared with existing BN models, MCBN better depicts the complex dependency structure between variables, therefore may better predict the underlying causal network.
Abbreviations
- TCGA
The Cancer Genome Atlas
- BN
Bayesian network
- GBN
Gaussian Bayesian network
- CBN
copula Bayesian network
- MCBN
mixture copula Bayesian network
- EM
expectation–maximization
- BIC
Bayesian information criterion
Footnotes
PEER REVIEW: 4 peer reviewers contributed to the peer review report. Reviewers reports totaled 1460 words, excluding any confidential comments to the academic editor.
FUNDING: The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work has been supported in part by the Arkansas Biosciences Institute, the major research component of the Arkansas Tobacco Settlement Proceeds Act of 2000.
DECLARATION OF CONFLICTING INTERESTS: The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author contribution
QZ conceived the study. QZ and XS analyzed the data. QZ wrote the manuscript. Both authors read and approved the final manuscript.
Data Availability
The flow cytometry data by Sachs et al. can be downloaded from http://science.sciencemag.org/content/suppl/2005/04/21/308.5721.523.DC1. TCGA ovarian cancer data can be downloaded via TCGA data portal https://tcga-data.nci.nih.gov.
REFERENCES
- 1.Fu F, Zhou Q. Learning sparse causal Gaussian networks with experimental intervention: Regularization and coordinate descent. J Amer Stat Assoc. 2013;108(501):288–300. [Google Scholar]
- 2.Friedman N, Linial M, Nachman I, et al. Using Bayesian networks to analyze expression data. J Computat Biol. 2000;7(3):601–20. doi: 10.1089/106652700750050961. [DOI] [PubMed] [Google Scholar]
- 3.Xu Y, Zhang J, Yuan Y, et al. A Bayesian graphical model for integrative analysis of TCGA data; 2012 IEEE International Workshop on Genomic Signal Processing and Statistics; 2012. p. 31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Zhang Q, Burdette J, Wang JP. Integrative network analysis of TCGA data for ovarian cancer. BMC Syst Biol. 2014;8(1338):1–18. doi: 10.1186/s12918-014-0136-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Ellis B, Wong WH. Learning causal Bayesian network structures from experimental data. J Amer Stat Assoc. 2008;103(482):778–789. [Google Scholar]
- 6.Voorman A, Shojaie A, Witten D. Graph estimation with joint additive models. Biometrika. 2014;101(1):85–101. doi: 10.1093/biomet/ast053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Elidan G. Copula Bayesian networks. Adv Neur Inform Process Syst. 2010;23:559–567. [Google Scholar]
- 8.Sachs K, Perez O, Pe’er D, et al. Causal protein-signaling networks derived from multiparameter single-cell data. Science. 2005;308(5721):523–529. doi: 10.1126/science.1105809. [DOI] [PubMed] [Google Scholar]
- 9.Kalisch M, Buhlmann P. Estimating high-dimensional directed acyclic graphs with the pc-algorithm. J Machine Learn Res. 2007;8:613–636. [Google Scholar]
- 10.The Cancer Genome Atlas Research Network Integrated genomic analyses of ovarian carcinoma. Nature. 2011;474:609–15. doi: 10.1038/nature10166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Bolstad B, Irizarry R, Astrand M, et al. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics. 2002;19(2) doi: 10.1093/bioinformatics/19.2.185. [DOI] [PubMed] [Google Scholar]
- 12.Mai K, Zhang Q. Identification of biomarkers for predicting the overall survival of ovarian cancer patients: a sparse group lasso approach. Int J Statist Prob. 2016;5(6) [Google Scholar]
- 13.Hsu F, Serpedin E, Hsiao T, et al. Reducing confounding and suppression effects in TCGA data: an integrated analysis of chemotherapy response in ovarian cancer. BMC Genomics. 2012;13 doi: 10.1186/1471-2164-13-S6-S13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Song H, Ramus S, Shadforth D, et al. Common variants in rb1 gene and risk of invasive ovarian cancer. Cancer Res. 2006;66(20):10220–6. doi: 10.1158/0008-5472.CAN-06-2222. [DOI] [PubMed] [Google Scholar]
- 15.Takei Y, Saga Y, Mizukami H, et al. Overexpression of PTEN in ovarian cancer cells suppresses i.p. dissemination and extends survival in mice. Mol Cancer Therapeut. 2008;7(3):704–11. doi: 10.1158/1535-7163.MCT-06-0724. [DOI] [PubMed] [Google Scholar]
- 16.Mckie A, Vaughan S, Zanini E, et al. The opcml tumor suppressor functions as a cell surface repressor-adaptor, negatively regulating receptor tyrosine kinases in epithelial ovarian cancer. Cancer Discov. 2012;2(2):156–71. doi: 10.1158/2159-8290.CD-11-0256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Yang F, Guo X, Yang G, et al. Aurka and brca2 expression highly correlate with prognosis of endometrioid ovarian carcinoma. Mod Pathol. 2011;24(6):836–845. doi: 10.1038/modpathol.2011.44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Schick N, Oakeley E, Hynes N, et al. Tel/etv6 is a signal transducer and activator of transcription 3 (stat3)-induced repressor of stat3 activity. J Biol Chem. 2004;279(37):38787–38796. doi: 10.1074/jbc.M312581200. [DOI] [PubMed] [Google Scholar]
- 19.Wrzeszczynski K, Varadan V, Byrnes J, et al. Identification of tumor suppressors and oncogenes from genomic and epigenetic features in ovarian cancer. PLoS One. 2011;6(12):0028503. doi: 10.1371/journal.pone.0028503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Liu Y, Tao X, Jia L, et al. Knockdown of rab25 promotes autophagy and inhibits cell growth in ovarian cancer cells. Mol Med Rep. 2012;6(5):1006–1012. doi: 10.3892/mmr.2012.1052. [DOI] [PubMed] [Google Scholar]
- 21.Yuan J, Zhang F, Niu R. Multiple regulation pathways and pivotal biological functions of stat3 in cancer. Sci Rep. 2015;5:17663. doi: 10.1038/srep17663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Carracedo A, Pandolfi P. The pten-pi3k pathway: of feedbacks and cross-talks. Oncogene. 2008;27:5527–5541. doi: 10.1038/onc.2008.247. [DOI] [PubMed] [Google Scholar]
- 23.Wu J, Roberts C. Arid1a mutation in cancer: Another epigenetic tumor suppressor. Cancer Discov. 2013;3(1):35–43. doi: 10.1158/2159-8290.CD-12-0361. [DOI] [PMC free article] [PubMed] [Google Scholar]