Abstract
Cancer cells prefer aerobic glycolysis, but little is known about the underlying mechanism. Recent studies showed that the rate-limiting glycolytic enzymes, pyruvate kinase M2 (PKM2) directly phosphorylates H3 at threonine 11 (H3T11) to regulate gene expression and cell proliferation, revealing its non-metabolic functions in connecting glycolysis and histone modifications. We have reported that the yeast homolog of PKM2, Pyk1 phosphorylates H3T11 to regulate gene expression and oxidative stress resistance. But how glycolysis regulates H3T11 phosphorylation remains unclear. Here, using a series of glycolytic enzyme mutants and commercial available metabolites, we investigated the role of glycolytic enzymes and metabolites on H3T11 phosphorylation. Mutation of glycolytic genes including phosphoglucose isomerase (PGI1), enolase (ENO2), triosephosphate isomerase (TPI1), or folate biosynthesis enzyme (FOL3) significantly reduced H3T11 phosphorylation. Further study demonstrated that glycolysis regulates H3T11 phosphorylation by fueling the substrate, phosphoenonylpyruvate and the coactivator, FBP to Pyk1. Thus, our results provide a comprehensive view of how glycolysis modulates H3T11 phosphorylation.
Introduction
Glycolysis is the fundamental metabolism highly conserved in most organisms, which comprises a series of enzymatic steps that sequentially convert glucose to pyruvate. In the presence of oxygen, most pyruvate undergoes oxidative phosphorylation to generate ATP in mitochondria; while in the absence of oxygen, pyruvate is converted to lactate with few ATP produced [1,2]. However, cancer cells preferentially convert pyruvate to lactate even in the presence of oxygen, a phenomenon known as “Warburg effect” or aerobic glycolysis [1]. Aerobic glycolysis enables cells to accumulate a large amount of glycolytic intermediates, which serve as building blocks to meet cell rapid growth and division [1–4]. Nevertheless, it remains poorly understood about why tumor cells prefer accelerated glycolysis and reduced oxidative phosphorylation.
Emerging evidence showed that most glycolytic enzymes are deregulated in cancer cells and plays important roles in tumorigenesis [2,5]. All essential glycolytic enzymes can be translocated into nucleus where they participate in tumor progression independent of their canonical metabolic roles [6]. One such non-metabolic function is catalyzing and/or modulating histone modifications. The typical example is tumor specific pyruvate kinase M2 (PKM2), which plays important roles in cancer metabolism rewiring [7]. Yang et al. reported that in human glioblastoma multiforme cells, PKM2 translocates into nucleus upon epidermal growth factor (EGF) receptor activation, where it phosphorylates histone H3 at threonine 11 (H3T11), which is required for dissociation of histone deacetylase 3 (HDAC3) from the promoter regions of CCDN1(encoding cyclin D1) and MYC, leading to their activation, tumor cell proliferation, cell-cycle progression, and brain tumorigenesis [8]. Previously, we have reported that Pyk1, the yeast homologue of PKM2 also has some non-metabolic functions [9]. Similar to PKM2, Pyk1 can phosphorylate H3T11 in vivo and in vitro and this protein kinase activity is regulated by serine metabolic pathway [9]. Specifically, H3T11 phosphorylation is regulated by enzymes involved in serine metabolism including Ser1, Ser2, Ser33, Shm2, Met6 and Met13. Moreover, by combining protein purification technique with mass spectrometry, we found that Pyk1 forms a novel complex, SESAME (serine responsive SAM-containing metabolic enzyme complex) with other metabolic enzymes, including Sam1, Sam2, Ser33, Shm2 and Acs2 [9]. Further studies showed that SESAME interacts with Set1 complex, which methylates H3K4. By supplying the cofactor SAM for Set1 complex, SESAME regulates both H3K4me3 and H3T11 phosphorylation. As a consequence, SESAME regulates gene expression and cell resistance to oxidative stress [9].
Cellular metabolism regulates histone modifications and many metabolites serve as essential cofactors for chromatin-modifying enzymes to control the transcription or translation processes [2,10,11]. For example, about 5% glucose is used for hexosamine biosynthetic pathways to produce GlcNAc, which is the donor for histone glycosylation [12]. Through glycolysis, glucose can be converted to acetyl CoA, along with decreased NAD+/NADH, which in turn regulate the activity of histone acetyltransferases and histone deacetylases as well as the chromatin structure [10,12–14]. We have previously shown that glucose is required for SESAME to phosphorylate H3T11 [9]; however, the pathways and metabolites critical for H3T11 phosphorylation remain poorly defined. Here, we analyzed the function of glycolytic metabolic enzymes and metabolites on H3T11 phosphorylation.
Materials and methods
Cells and growth conditions
All yeast strains used in this study are described in Table 1. All yeast cells were grown in YPD (2% yeast extract, 1% peptone, 2% glucose) medium unless otherwise indicated. For doxycycline treatment, WT Tet and mutants were grown in YPD to an OD600 of 0.7 and then treated with 0, 12.5, 25 and 50 μg/ml of doxycycline.
Table 1. List of strains used in this study.
| Name | Genotype | Source |
|---|---|---|
| BY4741 | MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0 | |
| sam1Δ | MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0 sam1Δ::KAN | Open Biosystems |
| sam2Δ | MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0 sam2Δ::KAN | Open Biosystems |
| ser33Δ | MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0 ser33Δ::KAN | Open Biosystems |
| acs1Δ | MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0 acs1Δ::KAN | Open Biosystems |
| shm2Δ | MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0 shm2Δ::KAN | Open Biosystems |
| eno1Δ | MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0 eno1Δ::KAN | Open Biosystems |
| pdc1Δ | MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0 pdc1Δ::KAN | Open Biosystems |
| WT Tet (R1158) | MATa his3-1 leu2-0 met15-0 URA3::CMV-tTA | Yeast Tet-promoters Hughes Collection |
| TetO7-ENO2 | MATa his3-1 leu2-0 met15-0 pENO2::kanR-tet07-TATA URA3::CMV-tTA | Yeast Tet-promoters Hughes Collection |
| TetO7-TPI1 | MATa his3-1 leu2-0 met15-0 pTPI1::kanR-tet07-TATA URA3::CMV-tTA | Yeast Tet-promoters Hughes Collection |
| TetO7-PGI1 | MATa his3-1 leu2-0 met15-0 pPGI1::kanR-tet07-TATA URA3::CMV-tTA | Yeast Tet-promoters Hughes Collection |
| TetO7-FOL3 | MATa his3-1 leu2-0 met15-0 pFOL3::kanR-tet07-TATA URA3::CMV-tTA | Yeast Tet-promoters Hughes Collection |
Histone extraction
Histones were extracted from yeast cells as described previously [9,15]. Briefly, cells grown in 5 ml culture was harvested and lysed in 2M NaOH with 8% β-mercaptoethanol. Cell lysate was centrifuged at 13,000 rpm for 2 min and the pellet was washed three times with TAP extraction buffer (40 mM HEPES-KOH pH7.5, 10% glycerol, 350 mM NaCl, 0.1% Tween-20). Cell pellets were resuspended in 1× SDS-sample buffer.
Western blots analysis
Protein samples were separated by 15% SDS-PAGE and transferred to Immobilon-P PVDF membrane (Merck Millipore). The blots were probed with antibodies against H3 (Abclonal Biotechnology) and H3pT11 (Abcam, ab5168) followed by incubation with horseradish peroxidase-labeled IgG secondary antibodies (Abclonal Biotechnology). The specific proteins were visualized by using the ECL Chemiluminescence Detection Kit (Amersham Biosciences). Western blots were quantified with ImageJ software.
Quantitative reverse-transcription PCR (qRT-PCR)
10 ml cultures were grown in YPD to an OD600 of 0.6–0.8 and treated with doxycycline for 0.5 hour. Total RNA was isolated and Real-Time RT-PCR was performed with SYBR green and gene specific primers as described previously [9]. All transcripts quantities were normalized against the amount of ACT1 transcript. Primers used were listed in S1 Table.
Analysis of fructose 1, 6-biphosphate (FBP)
50 ml cells were grown in YPD to an OD600 of ~1.0 and treated with 50 μg/μl doxycycline for 3 hours. Cells were harvested and lysed with glass beads. The intracellular FBP concentrations were determined using FBP analysis kit according to the protocol recommended by the manufacturer (Comin Biotechnology Co., Ltd, Suzhou).
Results
Glucose is required for SESAME-mediated H3T11 phosphorylation
Previously, we have used a temperature-sensitive (ts) strain defective in Pyk1 (cdc19-1) when grown at 37°C to demonstrated that Pyk1 phosphorylates H3T11 in vivo. Here, we employed the strains that display different PYK1 expression and activity by expressing PYK1 under the control of either a strong (TEF1) or a weak (CYC1) constitutive promoter [16] (S1A Fig). We grew these two strains (TEF1pr-PYK1, CYC1pr-PYK1) in rich media and examined the global levels of H3T11 phosphorylation by western blot analysis. As shown in Fig 1A, H3T11 phosphorylation was significantly reduced in CYC1pr-PYK1 strain (Fig 1A), which has reduced PYK1 expression and lower pyruvate kinase activity (S1A Fig) [16], confirming our previous results that Pyk1 phosphorylates H3T11 in vivo [9]. H3T11 phosphorylation has been shown to confer cells the resistance to oxidative stress [9] and the expression of genes involved in oxidative energy metabolism (CIT1, COX1) was significantly reduced in H3T11A mutant (S2 Fig). We thus examined the effect of Pyk1 in oxidative stress resistance. In the absence of oxidative stress, TEF1pr-PYK1 strain grew much better than CYC1pr-PYK1 in glucose-containing medium due to higher pyruvate kinase activity; however, CYC1pr-PYK1 grew similar to TEF1pr-PYK1 when cells were grown in the presence of oxidative stress (Fig 1B), implying that lower expression of PYK1 could confer cells the resistance to oxidative stress, which is consistant with published results [16]. As H3T11 phosphorylation acts as a feedback mechanism to repress PYK1 expression [9], it is possible that H3T11 phosphorylation combats oxidative stress by suppressing PYK1 expression.
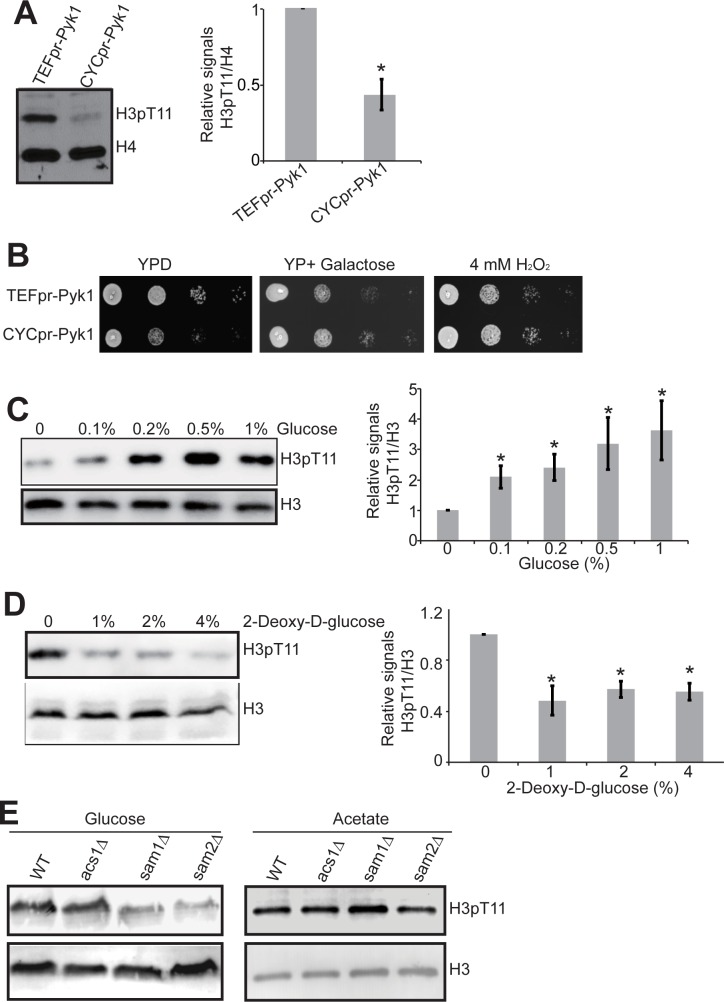
Fig 1. Glucose metabolism regulates H3T11 phosphorylation.
(A) Effect of Pyk1 on H3T11 phosphorylation. Left panel: TEFpr-PYK1 and CYCpr-PYK1 cells were cultured in YPD medium until OD600 of 0.7. Cells were harvested and extracted histones were analyzed by western blots with indicated antibodies. Right panel: Quantitation of western blots in left panel. Shown is the relative intensities of H3pT11/H4 quantified with standard error (SE) (n = 3). *, P<0.05. (B) Lower PYK1 expression confers oxidative stress resistance. Serial diluted TEFpr-PYK1 and CYCpr-PYK1 cells were spotted on YPD, YP+2% galactose or YPD+4mM H2O2. Shown is the typical example of three independent experiments. (C) Glucose is required for H3T11 phosphorylation. Left panel: Cells were cultured in YP medium, and 0, 0.1%, 0.2%, 0.5% or 1% glucose were then supplied to the medium 3 hours before harvest. Right panel: Quantitation of western blots in left panel. Shown is the relative intensities of H3pT11/H3 quantified with standard error (SE) (n = 3). *, P<0.05. (D) Inhibition of glycolysis by 2-Deoxy-D-glucose reduced H3T11 phosphorylation. Cells were grown in YPD medium with addition of 0, 1%, 2% or 4% 2-Deoxy-D-glucose for 3 hours before harvest. Right panel: Quantitation of western blots in left panel. Shown is the relative intensities of H3pT11/H3 quantified with standard error (SE) (n = 3). *, P<0.05. (E) Glucose was required for SESAME to regulate H3pT11. WT, acs1Δ, sam1Δ, and sam2Δ were grown in YP + 2% glucose or YP + 0.1M potassium acetate. Histones were extracted and analyzed by western blots with indicated antibodies. Histone H3 was a loading control.
It is noteworthy that TEF1pr-PYK1 and CYC1pr-PYK1 grew differently when glucose or galactose was used as the sole carbon source (Fig 1B). Hence, we examined the impact of glucose on H3T11 phosphorylation. First, we treated cells with different concentrations of glucose and found that glucose significantly stimulated H3T11 phosphorylation (Fig 1C), indicating that glucose is required for H3T11 phosphorylation. To investigate whether glycolysis is required for H3T11 phosphorylation, we treated cells with glucose analog, 2-Deoxy-D-glucose, which inhibited the activity of hexokinase to phosphorylate glucose and hence suppressed glycolysis. As shown in Fig 1D, the global level of H3T11 phosphorylation was significantly inhibited by 2-Deoxy-D-glucose (Fig 1D, P<0.05), indicating that glycolysis is required for carbohydrate-induced H3T11 phosphorylation.
To further confirm that glycolysis is required for SESAME activity, we grew SESAME mutants (sam1Δ, sam2Δ) with different carbon sources (glucose and potassium acetate) and then examined their effects on global H3T11 phosphorylation. In contrast to glucose as the sole carbon source, the global levels of H3pT11 were comparable between SESAME mutants and its parental wild type strain when potassium acetate was used as the carbon source (Fig 1E). Together, these data indicate that glycolysis is required for SESAME to phosphorylate H3T11.
Effect of phosphoglucose isomerase and fructose 1, 6-biphosphate (FBP) on H3T11 phosphorylation
Next, we explored the functions of the glycolysis downstream metabolic enzymes and metabolites in Pyk1-catalyzed H3T11 phosphorylation. Fructose 1, 6-biphosphate (FBP) is an important cofactor for the pyruvate kinase activity of both PKM2 and Pyk1 and depletion of glucose immediately reduces the intracellular level of FBP [17–20]. As an intermediate of glycolysis, FBP has been shown to stimulate the protein kinase activity of PKM2 to phosphorylate H3T11 [21]. To explore the impact of FBP on Pyk1-catalyzed H3T11 phosphorylation, we blocked FBP biosynthesis via down-regulating the expression of PGI1, which encodes a phosphoglucose isomerase that catalyzes the conversion of glucose-6-phosphate to fructose-6-phosphate, a precursor for FBP (Fig 2A). We employed the promoter-shutoff strain, TetO7-PGI1, in which the PGI1 promoter was replaced with TetO7, whose transcription can be shutoff by adding doxycycline [22]. We treated the TetO7-PGI1 mutant with doxycycline to down-regulate PGI1 transcription from the TetO7 promoter (S1B Fig). We also found that down-regulation of PGI1 expression reduced the intracellular FBP level in TetO7-PGI1 mutant upon doxycycline treatment (S3 Fig, P<0.05). In addition, treatment of TetO7-PGI1 mutant with doxycycline reduced the global H3T11 phosphorylation levels; however, doxycycline has no effect on H3T11 phosphorylation in WT Tet cells (Fig 2B and 2C), indicating that Pgi1 is required for Pyk1-mediated H3T11 phosphorylation.
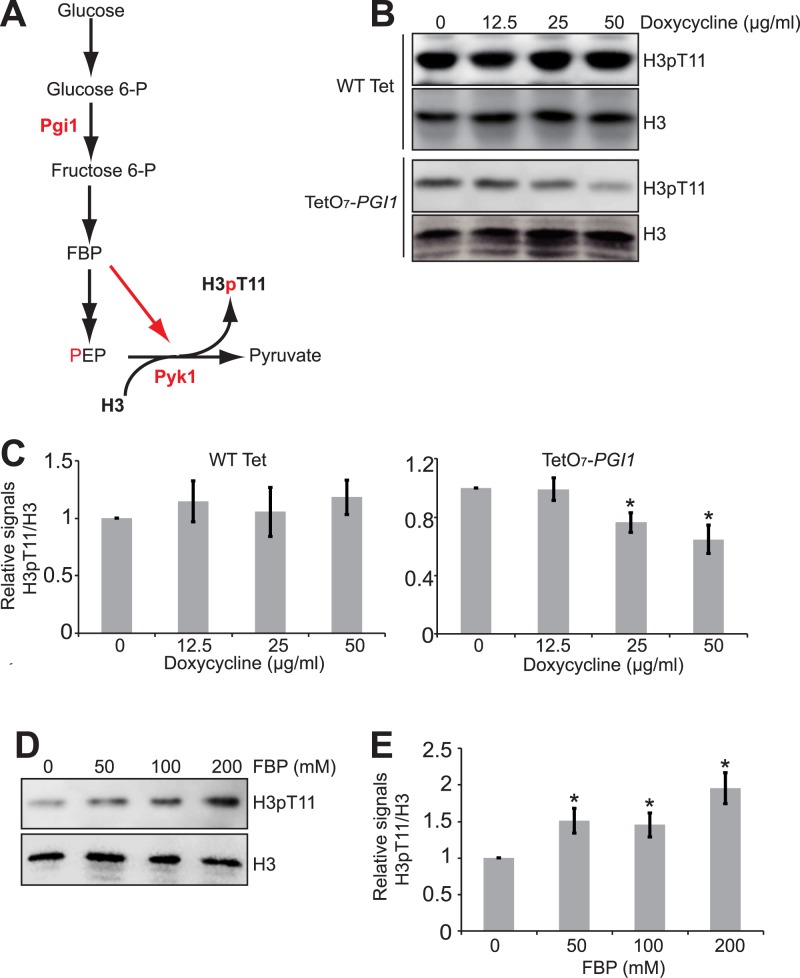
Fig 2. Pgi1 and FBP are required for SESAME-catalyzed H3T11 phosphorylation.
(A) Diagram displaying the functions of Pgi1 in glycolysis. (B) Down-regulated PGI1 leads to reduced H3pT11. Wild type and TetO7-PGI1 mutant was treated with 0, 12.5, 25, and 50 μg/ml doxycycline for 3 hours. Extracted histones were analyzed by western blots with indicated antibodies. (C) Quantitation of western blots in 2B. Shown is the relative intensities of H3pT11/H3 quantified with standard error (SE) (n = 3). *, P<0.05. (D) FBP addition increased H3T11 phosphorylation. Cells were grown in YPD medium with addition of 0, 50, 100, 200 mM FBP for 3 hours before harvest. Extracted histones were analyzed by western blots with indicated antibodies. (E) Quantitation of western blots in (D). Shown is the relative intensities of H3pT11/H3 quantified with standard error (SE) (n = 3). *, P<0.05.
To directly investigate whether FBP stimulates the protein kinase activity of Pyk1, we treated cells with different amounts of FBP and examined the global levels of H3T11 phosphorylation. FBP significantly stimulated Pyk1-catalyzed H3T11 phosphorylation (Fig 2D and 2E). Thus, like serine and SAICAR (succinyl-5-aminoimidazole-4-carboxamide-1-ribose-5’-phosphate) [9], FBP not only stimulates the pyruvate kinase activity of Pyk1 but also its protein kinase activity to phosphorylate H3T11.
Effect of enolase and phosphoenonylpyruvate (PEP) on SESAME-mediated H3T11 phosphorylation
In glycolysis, the presence of glucose promotes the conversion of the PEP to pyruvate by Pyk1 [17–19] (Fig 3A). Since Pyk1 utilized PEP as the donor to phosphorylate H3, we therefore examined the effect of blocking PEP synthesis on H3T11 phosphorylation. We down-regulated the expression of ENO1, ENO2, which encode enolase converting 2-phosphoglycerate to PEP. The expression of ENO2 was down-regulated in TetO7-ENO2 mutant upon doxycycline treatment (S1C Fig) [22]. The levels of H3pT11 were reduced upon doxycycline treatment in the TetO7-ENO2 strain but not in WT Tet cells (Fig 3B and 3C). When its paralog, ENO1 was deleted (S1D Fig), there was no significant change of H3T11 phosphorylation (Fig 3D), implying that Eno2 plays a major role in PEP biosynthesis. As enolase is required for PEP synthesis, it is conceivable that Eno2 regulates H3T11 phosphorylation via PEP.
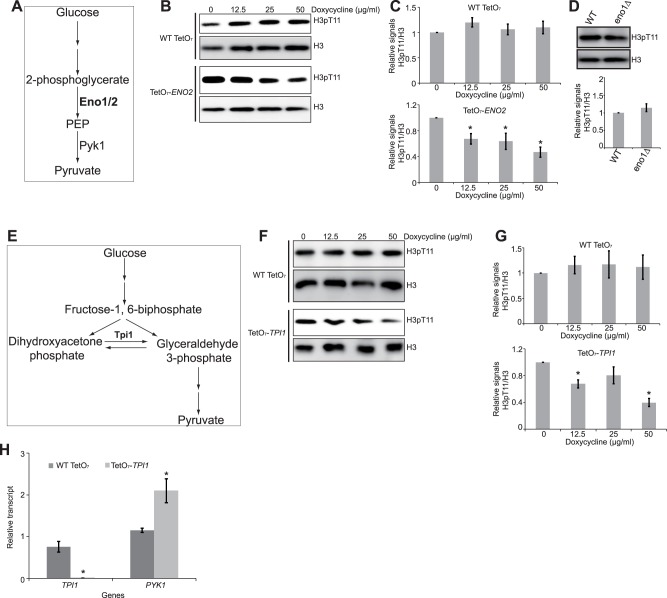
Fig 3. Eno2 and Tpi1 are required for SESAME-mediated H3T11 phosphorylation.
(A) Diagram displaying the functions of Eno1/2 in glycolysis. (B) Down-regulated ENO2 leads to reduced H3pT11. WT Tet and TetO7-ENO2 mutant were treated with 0, 12.5, and 25 μg/ml doxycycline for 3 hours. Histones were extracted and analyzed by western blots with indicated antibodies. Histone H3 was a loading control. (C) Quantitation of western blots in 3B. Shown is the relative intensities of H3pT11/H3 quantified with standard error (SE) (n = 3). *, P<0.05. (D) Deletion of ENO1 has no effect on H3T11 phosphorylation. Top panel: Analysis of H3T11 phosphorylation in WT and eno1∆ by western blots. Extracted histones were analyzed by western blots with indicated antibodies. Bottom panel: Quantitation of western blots in top panel. Shown is the relative intensities of H3pT11/H3 quantified with standard error (SE) (n = 4). P>0.05. (E) Diagram displaying the functions of Tpi1 in glycolysis. (F) Down-regulated TPI1 leads to reduced H3pT11. WT Tet and TetO7-TPI1 mutant were treated with 0, 12.5, and 25 μg/ml doxycycline for 3 hours. (G) Quantitation of western blots in 3F. Shown is the relative intensities of H3pT11/H3 quantified with standard error (SE) (n = 3). *, P<0.05. (H) PYK1 transcription was higher in TetO7-TPI1 mutant than wild type. Wild type and TetO7-TPI1 mutant were treated with doxycycline and the expression of TPI1 and PYK1 was measured by qRT-PCR. Actin was used as an internal control. Data represent the mean ± S.E. (n = 3). *, P<0.05.
Gruning et al. showed that reduced Pyk1 activity leads to accumulation of PEP, which in turn inhibits the upper glycolysis enzyme, triosephosphate isomerase (Tpi1) [16]. Tpi1 catalyzes the inter-conversion between glyceraldehyde 3-phosphate and dihydroxyacetone phosphate and its inhibition diverts glycolysis towards pentose phosphate pathway (Fig 3E) [16]. We treated TetO7-TPI1 mutant with doxycycline to reduce TPI1 transcription from the Tet promoter (S1E Fig). The levels of H3pT11 were reduced upon doxycycline treatment in the TetO7-TPI1 strain but not in WT Tet cells (Fig 3F and 3G), suggesting that Tpi1 is required for H3T11 phosphorylation. As reduced PYK1 expression confers oxidative stress resistance (Fig 1B), we thus examined the impact of Tpi1 on PYK1 expression. Our data showed that PYK1 expression was increased in TPI1 mutant (Fig 3H), suggesting that Tpi1 could mediate oxidative stress resistance.
Effect of pyruvate decarboxylase and pyruvate on SESAME-mediated H3T11 phosphorylation
Pyruvate kinase catalyzes the last step of glycolysis, which is also the rate-limiting and irreversible step. We next investigated whether pyruvate as the product of pyruvate kinase (Fig 4A), can feedback inhibit the protein kinase activity to phosphorylate H3T11. We treated cells with different amounts of pyruvate and examined the global levels of H3T11 phosphorylation. As shown in Fig 4B, pyruvate has no inhibitory effect on H3T11 phosphorylation (Fig 4B), consistent with the result that pyruvate kinase catalyzes the irreversible reaction.
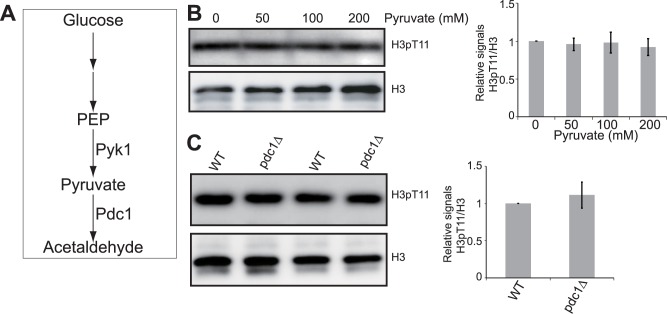
Fig 4. Pyruvate has no effect on SESAME-mediated H3T11 phosphorylation.
(A) Diagram displaying the functions of Pdc1 in glycolysis. (B) Addition of pyruvate did not significantly reduce H3T11 phosphorylation. Left panel: Cells were grown in YPD medium with addition of 0, 50, 100, 200 mM pyruvate for 3 hours before harvest. Histones were extracted and analyzed by western blots with indicated antibodies. Histone H3 was a loading control. Right panel: Quantitation of western blots in left panel. Shown is the relative intensities of H3pT11/H3 quantified with standard error (SE) (n = 3). (C) Deletion of PDC1 did not affect H3T11 phosphorylation. Left panel: Analysis of H3T11 phosphorylation in WT and pdc1∆ mutant by western blots. Right panel: Quantitation of western blots in left panel. Shown is the relative intensities of H3pT11/H3 quantified with standard error (SE) (n = 3).
We also examined the impact of in vivo accumulation of pyruvate by deleting genes encoding pyruvate decarboxylase, which encodes pyruvate decarboxylase that converts pyruvate to acetaldehyde. H3T11 phosphorylation was not significantly affected in either pdc1∆ mutant (Fig 4C) or in pdc5∆ mutant (data not shown). Together, these data indicate that pyruvate decarboxylase and pyruvate do not regulate the activity of SESAME to phosphorylate H3T11.
Effect of folate biosynthesis pathway on SESAME-mediated H3T11 phosphorylation
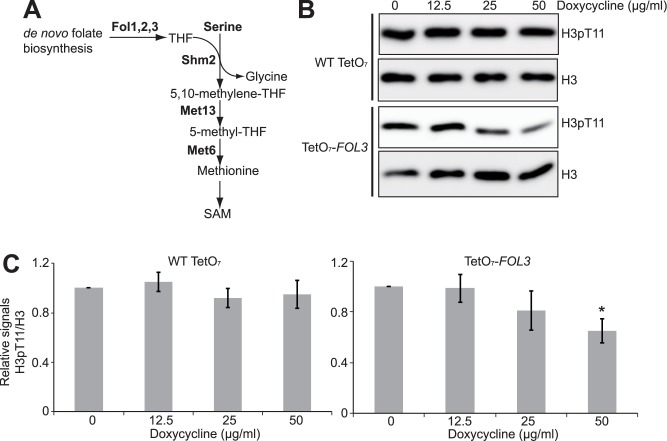
We have previously reported that histone methyltransferase Set1 stimulates SESAME-catalyzed H3T11 phosphorylation in a SAM-dependent manner and SAM increased global H3K4me3 and H3T11 phosphorylation in a dose dependent manner [9]. Glycolysis-derived serine provides an important methyl source for methionine and SAM synthesis. We have reported that blocking methionine biosynthesis by deletion of MET6 and MET13 specifically reduced both H3K4me3 and H3pT11 [9] (Fig 5A). In addition to serine, another critical source for methionine and SAM synthesis is folate and its derivatives tetrahydrofolate (THF) (Fig 5A). Sadhu et al. reported that preventing folate biosynthesis by deleting FOL3 specifically reduced global H3K4me3 [23]. Since H3K4me3 is directly related to H3T11 phosphorylation, we thus examined the effect of folate metabolism on H3T11 phosphorylation. We used a TetO7-FOL3 mutant and treated it with doxycycline to down-regulate FOL3 transcription from the Tet promoter (S1F Fig). The levels of H3pT11 were reduced upon doxycycline treatment in the TetO7-FOL3 strain (Fig 5B and 5C), indicating that folate biosynthesis pathway is required for H3T11 phosphorylation.
Fig 5. Folate metabolism is required for H3T11 phosphorylation.
(A) Diagram displaying de novo folate biosynthesis and methionine, SAM biosynthesis. (B) Down-regulated FOL3 leads to reduced H3pT11. TetO7-FOL3 mutant was treated with 0, 12.5, 25, and 50 μg/ml doxycycline for 3 hours. Extracted histones were analyzed by western blots with indicated antibodies. (C) Quantitation of western blots in 5B. Shown is the relative intensities of H3pT11/H3 quantified with standard error (SE) (n = 3). *, P<0.05.
Discussion
Glycolysis is required for global histone acetylation and mono-ubiquitination of H2B at K123 [13,24]. Our work showed that glucose and its metabolism regulates histone H3T11 phosphorylation (Fig 6). Combined with our previous study, we have shown that metabolic enzymes (Hxk1/2, Ser1, Ser2, Ser33, Sam1, Sam2, Pgi1, Tpi1, Eno2, Fol3) and metabolites (FBP, PEP, SAM) regulate Pyk1-mediated H3T11 phosphorylation, providing an intricate connection among glycolysis, histone modification and probably gene expression. Since glycolytic enzymes and metabolites are highly conserved from yeast to mammalian cells, it is conceivable that glycolysis regulates PKM2-mediated H3T11 phosphorylation via its metabolic enzymes and generated metabolites. Hence, our study provides insights into the connection between glycolysis and histone modifications and most importantly, provides one plausible explanation of the “Warburg effect”.
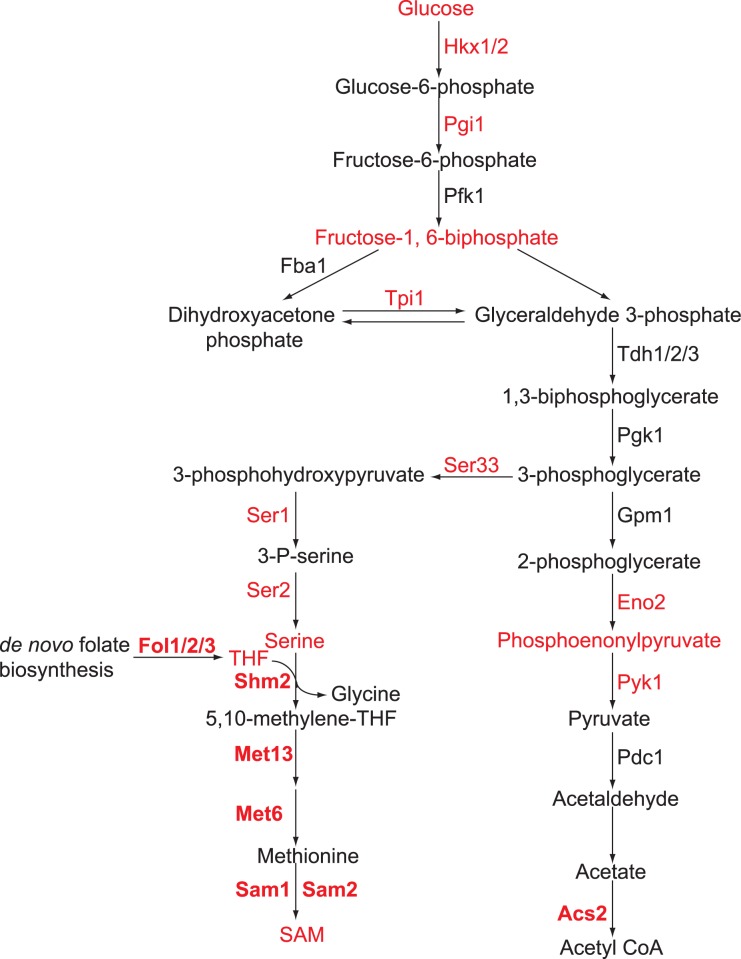
Fig 6. Diagram showing regulation of H3T11 phosphorylation by glycolysis.
Metabolic enzymes and metabolites highlighted in red color are required for H3T11 phosphorylation.
Gruning et al. have shown that lower Pyk1 activity increases antioxidative capacity [16] and our data confirmed this conclusion (Fig 1B). We have previously reported that H3T11 phosphorylation inhibits PYK1 expression and confers cells the resistance to oxidative stress [9]. Based on this information, we proposed that nucleus Pyk1-catalyzed H3T11 phosphorylation represses PYK1 expression, which in turn stimulates flux towards pentose phosphate pathway. As a consequence, redox metabolism is enhanced to prevent the accumulation of reactive oxygen species (ROS) (S4 Fig). Nevertheless, much work has to be done to prove this model.
Glucose metabolism is required for SESAME-mediated H3T11 phosphorylation. Depletion of glucose or inhibition of glycolysis by 2-Deoxy-D-glucose significantly reduced H3T11 phosphorylation (Fig 1C and 1D). When cells were grown with acetate as the carbon source, H3T11 phosphorylation was not significantly altered in SESAME mutants, implying that H3T11 phosphorylation was not regulated by SESAME under these circumstances. SESAME required glucose-derived metabolites to catalyze enzymatic reactions.
One contribution of glycolysis to H3T11 phosphorylation is providing the substrate, PEP for Pyk1. Down-regulating the expression of ENO2 reduced the global levels of H3T11 phosphorylation (Fig 3C). Although there are two genes encoding enolase (ENO1, ENO2), only ENO2 is required for H3T11 phosphorylation. The role of Eno2 in regulating H3T11 phosphorylation is probably related to their abundance with 20-fold higher Eno2 than Eno1 when glucose is used as the carbon source [25]. However, adding PEP into YPD rich media failed to stimulate Pyk1-mediated H3T11 phosphorylation (data not shown). This is probably caused by the dissociation constant (Km) of Pyk1 to PEP is very low, approximately 0.3 mM [20].
Another contribution of glycolysis to H3T11 phosphorylation is supplying the cofactor FBP for Pyk1. It is well-known that FBP stimulates Pyk1 to convert PEP to pyruvate [20]. FBP has been shown to activate both the pyruvate kinase and protein kinase activity of PKM2 [21]. Here, we found that FBP also activates the protein kinase activity of Pyk1 to phosphorylate H3T11 (Fig 2E). Hence, the stimulatory effect of FBP on pyruvate kinase-catalyzed H3T11 phosphorylation is quite conserved from yeast to tumor cells.
Together, we identified three major contribution of glucose metabolism to H3T11 phosphorylation: 1. Glycolysis provides the substrate PEP; 2. Glycolysis supplies cofactor FBP; 3. Glycolysis promotes the de novo synthesis of serine. On one hand, serine derived from glycolysis contributes to H3T11 phosphorylation by acting as a coactivator for Pyk1; on the other hand, serine can be fueled to SAM synthesis and facilitate H3K4me3, which then enhanced the ability of Pyk1 to phosphorylate H3T11 via a cross-talk between Set1 and SESAME [9]. Given the fact that yeast and cancer cells prefer aerobic glycolysis and PKM2 and H3T11 phosphorylation play important role in regulating “Warburg effect” and tumor progression [26,27], understanding how glycolysis modulates SESAME activity is important in understanding the Warburg effect and the connection between glycolysis and gene expression.
Supporting information
(A) PYK1 was expressed at a lower level in TEFpr-PYK1 than CYC1pr-PYK1. Actin was used as an internal control. Data represent the mean ± SE (n = 3). *, P<0.05. (B) The expression of PGI1 was significantly down-regulated in TetO7-PGI1 mutant by doxycycline. (C) The expression of ENO2 was significantly down-regulated in TetO7-PGI1 mutant by doxycycline. (D) ENO1 was deleted in eno1∆ mutant. The genome of WT and eno1∆ mutant were extracted. The deletion of ENO1 was confirmed by PCR using ENO1 specific primers. ACT1 primers were used as an internal control. (E) The expression of PGI1 was significantly down-regulated in TetO7-PGI1 mutant by doxycycline. (F) The expression of PGI1 was significantly down-regulated in TetO7-PGI1 mutant by doxycycline. All experiments except S1D Fig were measured by qRT-PCR. Data represent the mean ± SE of three independent experiments. *, P<0.05; **, P<0.01; ***, P<0.001.
(EPS)
Actin was used as an internal control. Data represent the mean ± SE (n = 3). *, P<0.05.
(EPS)
(A) The expression of PGI1 was significantly down-regulated in TetO7-PGI1 mutant by doxycycline treatment. (B) Down-regulated PGI1 leads to reduced intracellular FBP concentrations. Data represent the mean ± SE (n = 3). *, P<0.05; ***, P<0.001.
(EPS)
(A) Around 1.9% Pyk1 is localized in nucleus and this nucleus Pyk1 catalyzed H3T11 phosphorylation [9]. H3T11 phosphorylation in turn repressed PYK1 expression. Reduced PYK1 confers cells resistance to oxidative stress by stimulating pentose phosphate pathway, which increased antioxidative metabolism and prevents ROS accumulation. (B) Model explains oxidative stress resistance in CYC1pr-PYK1 mutant. In CYC1pr-PYK1 mutant, the protein level of Pyk1 was low, which increased flux towards pentose phosphate pathway to gain resistance to oxidative stress. (C) Model explains oxidative stress resistance in H3T11A mutant. In H3T11A mutant, PYK1 expression was up-regulated, which attenuates flux towards pentose phosphate pathway and reduces resistance to oxidative stress.
(EPS)
(DOC)
Acknowledgments
We would like to thank Professor Markus Ralser (Max Planck Institute for Molecular Genetics, Germany) for providing strains (TEF1pr-PYK1, CYC1pr-PYK1). We thank Professor Zhang Lab in Hubei University for technique support. This project was supported by funding from Hubei Collaborative Innovation Center for Green Transformation of Bio-resources (Shanshan Li), Hubei University start grant (080–170035 to Xilan Yu), Natural Science Foundation of Hubei Province (2016CFB103 to Xilan Yu), National Natural Science Foundation of China (No. 31671335 to Shanshan Li; No. 31600046 to Xilan Yu).
Data Availability
All relevant data are within the paper and its Supporting Information files.
Funding Statement
This project was supported by funding from Hubei Collaborative Innovation Center for Green Transformation of Bio-resources (S. Li), Hubei University start grant (080-170035 to Xilan Yu), National Natural Science Foundation of China (no. 31671335 to S. Li; no. 31600046 to Xilan Yu), and the Natural Science Foundation of Hubei Province (2016CFB103 to Xilan Yu). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1.Vander Heiden MG, Cantley LC, Thompson CB (2009) Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324: 1029–1033. doi: 10.1126/science.1160809 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Yu X, Li S (2016) Non-metabolic functions of glycolytic enzymes in tumorigenesis. Oncogene 10.1038/onc.2016.410. [DOI] [PubMed] [Google Scholar]
- 3.Jiang L, Deberardinis RJ (2012) Cancer metabolism: When more is less. Nature 489: 511–512. doi: 10.1038/489511a [DOI] [PubMed] [Google Scholar]
- 4.Yu X, Mao W, Zhai Y, Tong C, Liu M, Ma L, et al. (2017) Anti-tumor activity of metformin: from metabolic and epigenetic perspectives. Oncotarget 8: 5619–5628. doi: 10.18632/oncotarget.13639 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Kim JW, Dang CV (2005) Multifaceted roles of glycolytic enzymes. Trends Biochem Sci 30: 142–150. doi: 10.1016/j.tibs.2005.01.005 [DOI] [PubMed] [Google Scholar]
- 6.Boukouris AE, Zervopoulos SD, Michelakis ED (2016) Metabolic Enzymes Moonlighting in the Nucleus: Metabolic Regulation of Gene Transcription. Trends Biochem Sci 41: 712–730. doi: 10.1016/j.tibs.2016.05.013 [DOI] [PubMed] [Google Scholar]
- 7.Christofk HR, Vander Heiden MG, Harris MH, Ramanathan A, Gerszten RE, et al. (2008) The M2 splice isoform of pyruvate kinase is important for cancer metabolism and tumour growth. Nature 452: 230–233. doi: 10.1038/nature06734 [DOI] [PubMed] [Google Scholar]
- 8.Yang W, Xia Y, Hawke D, Li X, Liang J, Xing D, et al. (2012) PKM2 phosphorylates histone H3 and promotes gene transcription and tumorigenesis. Cell 150: 685–696. doi: 10.1016/j.cell.2012.07.018 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Li S, Swanson SK, Gogol M, Florens L, Washburn MP, Workman JL, et al. (2015) Serine and SAM responsive complex SESAME regulates histone modification crosstalk by sensing cellular metabolism. Mol Cell 60: 408–421. doi: 10.1016/j.molcel.2015.09.024 [DOI] [PubMed] [Google Scholar]
- 10.Xu W, Wang F, Yu Z, Xin F (2016) Epigenetics and Cellular Metabolism. Genet Epigenet 8: 43–51. doi: 10.4137/GEG.S32160 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Li S, Shogren-Knaak MA (2008) Cross-talk between histone H3 tails produces cooperative nucleosome acetylation. Proc Natl Acad Sci U S A 105: 18243–18248. doi: 10.1073/pnas.0804530105 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Lu C, Thompson CB (2012) Metabolic regulation of epigenetics. Cell Metab 16: 9–17. doi: 10.1016/j.cmet.2012.06.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Liu XS, Little JB, Yuan ZM (2015) Glycolytic metabolism influences global chromatin structure. Oncotarget 6: 4214–4225. doi: 10.18632/oncotarget.2929 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Li S, Shogren-Knaak MA (2009) The Gcn5 bromodomain of the SAGA complex facilitates cooperative and cross-tail acetylation of nucleosomes. J Biol Chem 284: 9411–9417. doi: 10.1074/jbc.M809617200 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Fingerman IM, Wu CL, Wilson BD, Briggs SD (2005) Global loss of Set1-mediated H3 Lys4 trimethylation is associated with silencing defects in Saccharomyces cerevisiae. J Biol Chem 280: 28761–28765. doi: 10.1074/jbc.C500097200 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Gruning NM, Rinnerthaler M, Bluemlein K, Mulleder M, Wamelink MM, Lehrach H, et al. (2011) Pyruvate kinase triggers a metabolic feedback loop that controls redox metabolism in respiring cells. Cell Metab 14: 415–427. doi: 10.1016/j.cmet.2011.06.017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Xu YF, Zhao X, Glass DS, Absalan F, Perlman DH, Broach JR, et al. (2012) Regulation of yeast pyruvate kinase by ultrasensitive allostery independent of phosphorylation. Mol Cell 48: 52–62. doi: 10.1016/j.molcel.2012.07.013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Chaneton B, Hillmann P, Zheng L, Martin AC, Maddocks OD, Chokkathukalam A, et al. (2012) Serine is a natural ligand and allosteric activator of pyruvate kinase M2. Nature 491: 458–462. doi: 10.1038/nature11540 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Ye J, Mancuso A, Tong X, Ward PS, Fan J, Rabinowitz JD, et al. (2012) Pyruvate kinase M2 promotes de novo serine synthesis to sustain mTORC1 activity and cell proliferation. Proc Natl Acad Sci U S A 109: 6904–6909. doi: 10.1073/pnas.1204176109 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Boles E, Schulte F, Miosga T, Freidel K, Schluter E, Zimmermann FK, et al. (1997) Characterization of a glucose-repressed pyruvate kinase (Pyk2p) in Saccharomyces cerevisiae that is catalytically insensitive to fructose-1,6-bisphosphate. J Bacteriol 179: 2987–2993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Keller KE, Doctor ZM, Dwyer ZW, Lee YS (2014) SAICAR induces protein kinase activity of PKM2 that is necessary for sustained proliferative signaling of cancer cells. Mol Cell 53: 700–709. doi: 10.1016/j.molcel.2014.02.015 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Mnaimneh S, Davierwala AP, Haynes J, Moffat J, Peng WT, Zhang W, et al. (2004) Exploration of essential gene functions via titratable promoter alleles. Cell 118: 31–44. doi: 10.1016/j.cell.2004.06.013 [DOI] [PubMed] [Google Scholar]
- 23.Sadhu MJ, Guan Q, Li F, Sales-Lee J, Iavarone AT, Hammond MC, et al. (2013) Nutritional control of epigenetic processes in yeast and human cells. Genetics 195: 831–844. doi: 10.1534/genetics.113.153981 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Dong L, Xu CW (2004) Carbohydrates induce mono-ubiquitination of H2B in yeast. J Biol Chem 279: 1577–1580. doi: 10.1074/jbc.C300505200 [DOI] [PubMed] [Google Scholar]
- 25.McAlister L, Holland MJ (1982) Targeted deletion of a yeast enolase structural gene. Identification and isolation of yeast enolase isozymes. J Biol Chem 257: 7181–7188. [PubMed] [Google Scholar]
- 26.Diaz-Ruiz R, Rigoulet M, Devin A (2011) The Warburg and Crabtree effects: On the origin of cancer cell energy metabolism and of yeast glucose repression. Biochim Biophys Acta 1807: 568–576. doi: 10.1016/j.bbabio.2010.08.010 [DOI] [PubMed] [Google Scholar]
- 27.Diaz-Ruiz R, Uribe-Carvajal S, Devin A, Rigoulet M (2009) Tumor cell energy metabolism and its common features with yeast metabolism. Biochim Biophys Acta 1796: 252–265. doi: 10.1016/j.bbcan.2009.07.003 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
(A) PYK1 was expressed at a lower level in TEFpr-PYK1 than CYC1pr-PYK1. Actin was used as an internal control. Data represent the mean ± SE (n = 3). *, P<0.05. (B) The expression of PGI1 was significantly down-regulated in TetO7-PGI1 mutant by doxycycline. (C) The expression of ENO2 was significantly down-regulated in TetO7-PGI1 mutant by doxycycline. (D) ENO1 was deleted in eno1∆ mutant. The genome of WT and eno1∆ mutant were extracted. The deletion of ENO1 was confirmed by PCR using ENO1 specific primers. ACT1 primers were used as an internal control. (E) The expression of PGI1 was significantly down-regulated in TetO7-PGI1 mutant by doxycycline. (F) The expression of PGI1 was significantly down-regulated in TetO7-PGI1 mutant by doxycycline. All experiments except S1D Fig were measured by qRT-PCR. Data represent the mean ± SE of three independent experiments. *, P<0.05; **, P<0.01; ***, P<0.001.
(EPS)
Actin was used as an internal control. Data represent the mean ± SE (n = 3). *, P<0.05.
(EPS)
(A) The expression of PGI1 was significantly down-regulated in TetO7-PGI1 mutant by doxycycline treatment. (B) Down-regulated PGI1 leads to reduced intracellular FBP concentrations. Data represent the mean ± SE (n = 3). *, P<0.05; ***, P<0.001.
(EPS)
(A) Around 1.9% Pyk1 is localized in nucleus and this nucleus Pyk1 catalyzed H3T11 phosphorylation [9]. H3T11 phosphorylation in turn repressed PYK1 expression. Reduced PYK1 confers cells resistance to oxidative stress by stimulating pentose phosphate pathway, which increased antioxidative metabolism and prevents ROS accumulation. (B) Model explains oxidative stress resistance in CYC1pr-PYK1 mutant. In CYC1pr-PYK1 mutant, the protein level of Pyk1 was low, which increased flux towards pentose phosphate pathway to gain resistance to oxidative stress. (C) Model explains oxidative stress resistance in H3T11A mutant. In H3T11A mutant, PYK1 expression was up-regulated, which attenuates flux towards pentose phosphate pathway and reduces resistance to oxidative stress.
(EPS)
(DOC)
Data Availability Statement
All relevant data are within the paper and its Supporting Information files.