Abstract
The Saccharomyces cerevisiae Srs2 DNA helicase has important roles in DNA replication, recombination and repair. In replication, Srs2 aids in repair of gaps by repair synthesis by preventing gaps from being used to initiate recombination. This is considered to be an anti-recombination role. In recombination, Srs2 plays both prorecombination and anti-recombination roles to promote the synthesis-dependent strand annealing recombination pathway and to inhibit gaps from initiating homologous recombination. In repair, the Srs2 helicase actively promotes gap repair through an interaction with the Exo1 nuclease to enlarge a gap for repair and to prevent Rad51 protein from accumulating on single-stranded DNA. Finally, Srs2 helicase can unwind hairpin-forming repeat sequences to promote replication and prevent repeat instability. The Srs2 activities can be controlled by phosphorylation, SUMO modification and interaction with key partners at DNA damage or lesions sites, which include PCNA and Rad51. These interactions can also limit DNA polymerase function during recombinational repair independent of the Srs2 translocase or helicase activity, further highlighting the importance of the Srs2 protein in regulating recombination. Here we review the myriad roles of Srs2 that have been documented in genome maintenance and distinguish between the translocase, helicase and additional functions of the Srs2 protein.
Keywords: DNA helicase; translocase; anti-recombinase; Srs2, toxic recombination; DNA repair, recombination
Srs2 is a multifunctional protein that acts during cell replication to insure faithful and accurate copying of genetic information.
INTRODUCTION
The SRS2 gene was first identified in screens for suppressors of yeast rad6 trimethoprim and UV sensitivities (Lawrence and Christensen 1979). Subsequently, it was realized that the RADH gene, identified as a suppressor of rad18 (Aboussekhra et al.1989), and HPR5, identified as a hyper-rec mutant and a suppressor of rad18 (Rong et al.1991), were alleles of SRS2. Genetics studies showed that the SRS2 gene functions in the post-replication repair (PRR) pathway to channel DNA lesions such as those induced by UV, ionizing radiation or MMS treatment into a gap repair pathway and that when it is non-functional, these lesions are channeled into a recombination repair pathway (Aboussekhra et al.1989; Schiestl, Prakash and Prakash 1990; Rong et al.1991) (Fig. 1). This was the first demonstration of an anti-recombinase phenotype in eukaryotes. Suppressors of srs2 mutants were found in the RAD51 gene (Aboussekhra et al.1992), and this suggested that anti-recombinase function of Srs2 was achieved through removal of Rad51 from single-stranded DNA, a prediction that was verified in vitro (Krejci et al.2003; Veaute et al.2003). The activity of Srs2 in removing Rad51 nucleofilaments was subsequently called a ‘strippase’ activity, to distinguish this from a DNA helicase activity. Both the strippase and DNA helicase activities require the translocase activity of Srs2. With the demonstration of the strippase activity, many of the phenotypes of srs2 mutants could be interpreted as a failure to prevent homologous recombination (HR), during both repair and rescue of stalled replication forks (Barbour and Xiao 2003; Watts 2006; Lambert et al.2010) and in other instances of mitotic HR (Robert et al.2006; Le Breton et al.2008; Burgess et al.2009; Kerrest et al.2009; Urulangodi et al.2015). The DNA helicase unwinding activity in vivo was demonstrated to be necessary to counteract the deleterious consequences of trinucleotide repeat sequences that could form stable hairpin structures during replication (Bhattacharyya and Lahue 2004, 2005; Kerrest et al.2009) and in unwinding at lesion sites produced by cleavage at rNMP residues in DNA to limit mutagenesis (Potenski et al.2014). The many roles of Srs2 in genome stability are centered on these two key activities: the Rad51 strippase activity to control the use of HR and the DNA helicase activity. We review these roles below with a focus on the regulation of Srs2 activity under different conditions, and finally summarize the functions of Srs2 and propose how similar roles are achieved and by what enzymes in mammalian cells.
Figure 1.
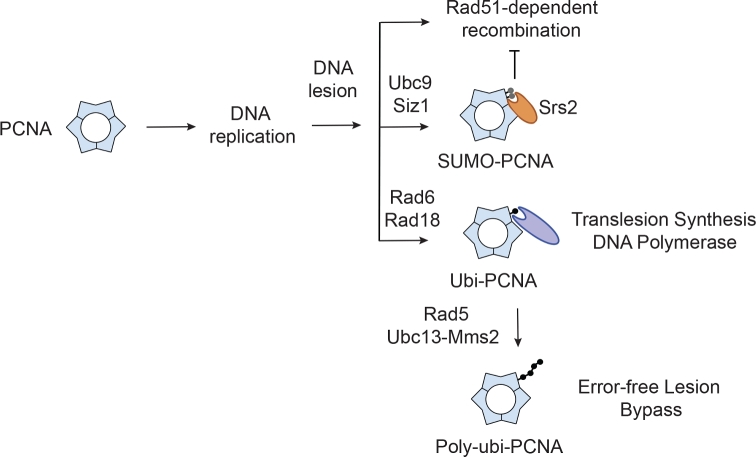
Role of Srs2 in the promotion of PRR. PCNA interacts with DNA polymerases to facilitate processive leading and lagging strand DNA synthesis. DNA replication can stall when a DNA lesion is encountered, and exposure of excess single-stranded DNA may lead to Rad51-dependent salvage recombination events. Sumoylation of PCNA at K164 and K127 by Ubc9 and Siz1 recruits Srs2 helicase, which actively dismantles Rad51 filament on the single-stranded DNA and antagonizes HR. Replication fork stalling also triggers monoubiquitylation of PCNA in a manner that depends on Rad6 and Rad18. Monoubiquitylated PCNA recruits translesion DNA polymerases to bypass the DNA lesion through translesion synthesis pathway (TLS). The Ubc13-Mms2-Rad5 E2 and E3 ensemble can further polyubiquitylate PCNA, which activates template switching and error-free lesion bypass.
Srs2 PROTEIN
Srs2, a SF1A family (Super Family 1A) helicase, contains 1174 amino acids and has multiple domains (Fig. 2A). The helicase domain of Srs2, which resides at the amino terminus (aa 1–776), shares homology with the UvrD helicase of bacteria (Aboussekhra et al.1989; Rong et al.1991). Following the helicase domain, Srs2 contains a flexible C-terminal region of nearly 400 amino acids in length that mediates most of the protein–protein interactions and contains the regions of the post-translation modifications (Chiolo et al.2005). In a two-hybrid analysis of Srs2, more than 70 proteins including major HR-related factors, such as the Mre11 nuclease and the Sgs1 and Mph1 helicases, were identified as Srs2 interactors (Chiolo et al.2005). Further analysis suggested that Srs2 forms complexes with Mre11 and Sgs1, which are subjected to regulation by CDK and checkpoint kinases (Mec1 and Tel1) upon DNA damage (Chiolo et al.2005; Saponaro et al.2010). The functional importance of the interaction between Srs2 and Mre11, as well as many other proteins, remains to be defined.
Figure 2.
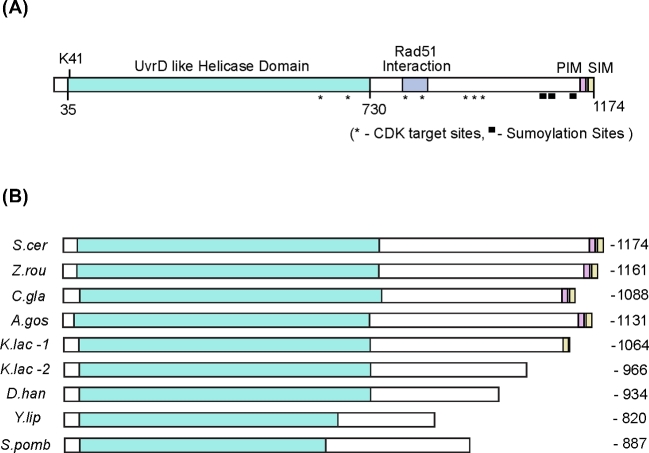
Scheme of Srs2 domain organization. (A) The UvrD like helicase domain is marked in green. The motifs mediating interaction with Rad51, PCNA and SUMO are marked in purple, pink and yellow, respectively. K41 is a key residue in the Walker A motif and is required for ATP binding. Mutation of K41 to alanine inactivates the helicase and translocase activities of Srs2. The seven Cdk1 phosphorylation sites are labeled with a star, and the locations of three lysine residues that are sumoylated in vivo were marked with solid square. (B) A comparison of Srs2 homologs from multiple fungal species shows high conservation in the N-terminal helicase domain, but greater variation at the C-terminus. Helicase domains are colored in green. PCNA and SUMO-binding motifs are colored in pink and yellow, respectively. Notably, Kluyveromyces lactis contains two Srs2 homologs with C-terminus in difference length. (S. cer: Saccharomyces cerevisiae; Z. rou: Zygosaccharomyces rouxii; C. gla: Candida glabrata; A. gos: Eremothecium gossypii; K. lac: Kluyveromyces lactis; D. han: Debaryomyces hansenii; Y. lip: Yarrowia lipolytica; S. pomb: Schizosaccharomyces pombe.
Srs2 interacts with both Rad51 and PCNA to fulfill its anti-recombination role during DNA replication. Both interactions have been mapped to the C-terminus of Srs2. While the C-terminal truncated Srs2-1-910 is still able to interact with Rad51 in vitro, Srs2-1-875 retains minimal Rad51 binding. Further deletion of residues between 875 and 910 in the full-length Srs2 largely abolished its interaction with Rad51 and attenuated its capability of Rad51 removal in vitro (Fig. 2A) (Colavito et al. 2009). At the very end of the Srs2 C-terminus, there is a degenerate PCNA interaction motif (PIM domain) and a conserved SUMO interaction motif (SIM domain). Both motifs contribute to optimal binding of SUMO-modified PCNA by Srs2, which targets Srs2 to stalled replication forks to suppress HR (Papouli et al.2005; Pfander et al.2005; Armstrong, Mohideen and Lima 2012). An Srs2 C-terminal fragment binding to SUMO-PCNA has been crystallized and has revealed a bipartite interaction between Srs2 and SUMO-PCNA within a single SUMO-PCNA protomer (Armstrong, Mohideen and Lima 2012). In addition to the interaction with PCNA, a more recent study suggests that the SIM domain of Srs2 also mediates its interaction with other recombination factors, e.g. Mre11, Rad51 and Rad52, and likely helps to fulfill a prorecombination role (Kolesar et al.2016).
Similar to the UvrD helicase, Srs2 unwinds duplex DNA with a 3΄-5΄ polarity and in a manner that requires ATP binding and hydrolysis (Rong and Klein 1993). As a consequence, mutation of lysine 41 located in the Walker A motif of Srs2 helicase to alanine abolishes ATPase and helicase activities (Krejci et al.2004). Srs2 is capable of unwinding duplex DNA with 3΄-overhanging, 5΄-overhanging or blunt ends (Van Komen et al.2003) with a preference for 3΄-overhangings containing a minimum of 10 nucleotides (Van Komen et al.2003). A recent study with a truncated Srs2 protein (aa 1–898) that harbors the intact helicase domain showed that Srs2 helicase can unwind variety of DNA structures that associate with DNA replication and repair, including Flap, fork and Holliday junctions (Marini and Krejci 2012).
Srs2 STRIPPASE ACTIVITY
The most well-characterized activity of the Srs2 protein is the dismantling of a Rad51-ssDNA nucleoprotein filament. This strippase activity is critical to antagonize HR and to remove unproductive recombination intermediates (Fig. 3). The optimal strippase activity of Srs2 requires both the translocase activity and an interaction with Rad51. Srs2 mutants that fail to interact with Rad51 are also defective in dismantling Rad51 filaments (Antony et al.2009; Colavito et al.2009; Islam et al.2012). The strippase activity of Srs2 is limited to Rad51 and does not affect the nucleoprotein filament formed by RecA (Qiu et al.2013). Indeed, a RecA filament can be turned over by the UvrD helicase (Veaute et al.2005), which suggests a species-specific regulation of RecA/Rad51 filaments by the Srs2/UvrD helicases. Although the molecular underpinnings of antagonizing Rad51-filament formation by Srs2 are not fully defined, several activities of Srs2 that may be involved have been described. One study has discovered that physical interaction between Srs2 and Rad51 stimulates the Rad51 ATPase activity so as to facilitate Rad51 turnover since efficient DNA binding by Rad51 requires its binding to ATP (Antony et al.2009). Notably, the Rad51 paralogs Rad55 and Rad57 were shown to counteract the strippase activity of Srs2 (Fig. 3B) (Liu et al.2011), likely due to the role for a Rad55-Rad57 complex in stabilizing the Rad51 filament (Sung 1997; Liu et al.2011). Multiple studies also implicate Rad52 in a similar manner (Burgess et al.2009; Seong et al.2009).
Figure 3.
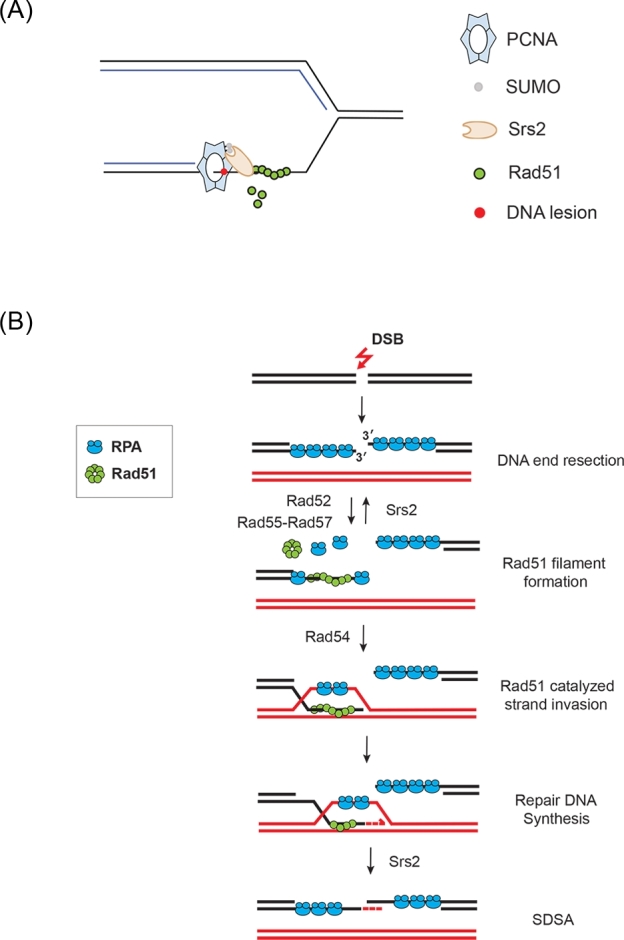
Anti-recombination and anti-crossover activities of Srs2 during replication and HR mediated DSB repair. (A) Accumulation of single-stranded DNA at stalled replication forks can lead to Rad51 filament assembling and Rad51-catalyzed recombination events. PCNA, when sumoylated, actively recruits Srs2 helicase to turn over Rad51 filaments and suppress salvage recombination events. (B) Double-strand breaks (DSBs) can be repaired by HR, which is initiated by the generation of 3΄ single-stranded DNA through DNA end resection. RPA, the single-stranded DNA-binding protein loaded during DNA end resection, is replaced by Rad51, a reaction facilitated by mediator activities from Rad52 and Rad55-Rad57. Rad51 is able to catalyzed strand invasion when Rad54 is present. In its absence, the Rad51 filament may become toxic and requires strippase activity from Srs2 for its turnover, so that alternative repair mechanisms can be employed. Upon successful strand invasion, the invaded 3΄ end is extended by repair DNA synthesis to recover the missing genetic information based on the available homologous template. The resulting intermediates may be processed through SDSA pathway where the invading strand is displaced and annealed to the opposite 3΄ end of the break followed by gap filling. Multiple lines of genetic evidence suggest Srs2 plays a key role in SDSA pathway.
Srs2 IN VIVO
In vivo, the srs2 null allele exhibits synthetic lethality or slow growth with a series of mutants, most of which are related to double-strand break repair or DNA replication, for example, CTF4, CTF8, MRC1, TOF1, MMS4, MUS81, RAD50, MRE11, XRS2, RAD27, POL32, SGS1, RRM3 among many others (Gangloff, Soustelle and Fabre 2000; Klein 2001; Ooi, Shoemaker and Boeke 2003; Tong et al.2004; Xu, Boone and Klein 2004; Pan et al.2006). Many of the synthetic lethalities arise from the formation of lethal recombination intermediates (Aboussekhra et al.1989; Rong et al.1991; Gangloff, Soustelle and Fabre 2000; Klein 2001) as deletion of RAD51 can rescue the lethality. Of note is the srs2 rad54 double mutant lethality, which is due to inappropriate recombination. Since Rad54 protein is needed for many steps during HR, the double mutant lethality, which is rescued by rad51 mutation, suggests that Srs2 has a prorecombination role that overlaps with Rad54 (Klein 2001; Ira et al.2003; Kolesar et al.2016). In some cases, lethality is not rescued by RAD51 deletion as recombination repair is essential to collapsed replication fork restart (Debrauwere et al.2001).
The Srs2 multifunctional DNA helicase motor protein is conserved in fungal species although some species such as Schizosaccharomyces pombe have an additional related helicase called Fbh1 (Morishita et al.2005). In some fungal species, the Srs2 protein does not have as extensive a C-terminal region as in the Saccharomyces cerevisiae protein (Richard et al.2005) (Fig. 2B). However, the Srs2 proteins appear to function as anti-recombinases and in PRR in most organisms that contain Srs2 homologs.
Initial genetic studies placed the SRS2 gene in the PRR pathway (Lawrence and Christensen 1979; Aboussekhra et al.1989; Schiestl, Prakash and Prakash 1990; Rong et al.1991; Friedl et al.2001) through mutant suppression of strong UV and MMS sensitivities of PRR mutants rad6 and rad18. Further analysis placed SRS2 within the error-free gap repair branch of the PRR pathway (Ulrich 2001; Broomfield and Xiao 2002), which was in keeping with the idea that Srs2 acts to channel DNA gaps into an error-free gap repair pathway thorough inhibition of recombination at DNA gaps. Once it was understood that Srs2 protein had Rad51 strippase activity, there has been an added focus on regulation of recombination outcomes by Srs2. Srs2 promotes non-crossover synthesis-dependent strand annealing (SDSA) recombination at DNA breaks (Fig. 3B), which results in reduced crossover formation. Consequently, loss of Srs2 reduces only non-crossover SDSA events, with an outcome of an increase in the proportion of crossovers. Another function of Srs2 was revealed though rescue of deleterious mutant combinations by eliminating Rad51 or by providing Srs2 protein. This has led to the concept of lethal recombination intermediates accumulating in the absence of Srs2 and another recombination factor such as Rad54 or Sgs1 (Fig. 3B) (Klein 2001; Fabre et al.2002).
Srs2 regulates repair at stalled replication forks in part though recruitment to SUMO-modified PCNA (Papouli et al.2005; Pfander et al.2005); however, Srs2 can remove lethal recombination intermediates independent of this interaction (Le Breton et al.2008; Kolesar et al.2016). In addition to regulating recombination at stalled forks, the Srs2 protein regulates recombination by limiting crossovers and promoting SDSA (Ira et al.2003; Robert et al.2006; Miura, Shibata and Kusano 2013), thus limiting loss of heterozygosity events during mitotic repair of lesions through HR. Recombination between sequences that are highly related but not identical, called homeologous recombination, remains limited by the mismatch repair system in a srs2 mutant background, implying that Srs2 does not play an active role in limiting recombination at the step of strand invasion and base pairing (Spell and Jinks-Robertson 2004).
Srs2 indirectly affects DNA damage signaling as persistent Rad51 filaments impact the formation and reversal of a DNA damage checkpoint signal. srs2 mutants fail to remove Rad51 filaments formed after DNA damage, and as a consequence, are unable to recover from DNA damage and reverse the checkpoint damage signal emanating from persistent single-stranded DNA (Vaze et al.2002; Yeung and Durocher 2011).
Srs2 MUTANTS
srs2 mutants have been isolated based on a DNA repair or recombination phenotype, or through site-directed mutagenesis. The first srs2 mutants were isolated as suppressors of the PRR mutants rad6 and rad18, using suppression of extreme trimethoprim, UV sensitivity or MMS sensitivity as a selection (Lawrence and Christensen 1979; Aboussekhra et al.1989; Schiestl, Prakash and Prakash 1990). The hpr5 mutant was isolated in a screen for hyper-rec mutants (Aguilera and Klein 1988) and soon it was realized that the hpr5 mutant was an allele of srs2 on the basis of phenotype and cloning of the SRS2 gene (Rong et al.1991). The srs2 mutants that suppressed rad18 were semidominant, in that heterozygous diploids had a slight DNA damage sensitive phenotype. Moreover, all srs2 phenotypes were exacerbated in diploid strains, which allowed the recovery of suppressors of srs2 that turned out to be defects in RAD51 (Aboussekhra et al.1992). Additionally, the original hpr5-1 allele and the null allele have a slight meiotic phenotype with reduced crossovers and spore viability (Palladino and Klein 1992).
Site-directed mutagenesis has yielded mutants in the ATPase Walker A box motif (Krejci et al.2004), the seven phosphorylation sites of Srs2 (Saponaro et al.2010) and the sumoylation sites of Srs2 (Saponaro et al.2010; Kolesar et al.2012) (Fig. 2A). While the phosphorylation and sumoylation modifications are thought to play some role in regulating recombination and repair in vivo, loss of these modifications does not grossly affect spontaneous recombination rates. However, Srs2 phosphorylation modification does regulate Rad51-dependent DNA double-strand break repair, as opposed to gap repair (Saponaro et al.2010).
ATP hydrolysis is needed for both the strippase and helicase activities and hence ATPase-defective alleles of SRS2 are both hyper-rec and DNA damage sensitive and exhibit reduced growth. Curiously, while the null allele is viable in diploids, the ATPase-defective allele is diploid lethal in either the homozygous or heterozygous state over the null allele (Keyamura, Arai and Hishida 2016). This suggests that in diploids Srs2 protein performs functions that become essential if the protein becomes bound to DNA and cannot translocate.
Although most of the Srs2 modifications occur in the C-terminal region of the protein, which also includes the Rad51 and PCNA interaction domains (Fig. 2A), this domain is not essential. Truncation alleles that retain the DNA helicase activity are viable, hyper-rec and suppress rad18 (Colavito et al.2009). In particular, the srs2-1-860 allele, which contains the DNA helicase domain but lacks the majority of Srs2 C-terminus, still retains the anti-crossover activity (Mitchel, Lehner and Jinks-Robertson 2013) and is functional in preventing mutagenesis promoted by Top1 cleavage in embedded genomic ribonucleotides (Potenski et al.2014). Moreover, Srs2-Δ875-902, the aforementioned Srs2 mutant defective in Rad51 binding, is still capable of Rad51 removal in cells (Sasanuma et al.2013). Thus, the motor activity of Srs2 provides the vital force for fulfilling its multifaceted role in vivo.
Srs2 IN RECOMBINATION
Once it was realized that many srs2 mutants were hyper-rec, could be suppressed by loss of Rad51 or other HR-initiating mutations and had a synthetic growth defect with rad54, the SRS2 gene was thought to be involved in negatively regulating recombination. The realization that the anti-recombinase activity arose from the removal of Rad51 filaments (Burgess et al.2009) and prevented toxic recombination intermediates from formation (Bernstein et al.2011) explained many of the double mutant lethalities with srs2 as well as the suppression of mutants within the PRR pathway without affecting induced mutagenesis.
However, neither all of the double mutant genetic interactions of srs2 nor all of the mutant phenotypes are completely explained by the anti-recombinase model. It became apparent that Srs2 could also promote certain types of HR while at the same time preventing deleterious recombination. Srs2 regulates the outcome of HR, once an intermediate is established that is not toxic, and here Srs2 functions to promote non-crossover recombination and reduce crossovers (Robert et al.2006; Saponaro et al.2010; Miura, Shibata and Kusano 2013). Indeed, there is a delicate balance between anti-recombination and prorecombination activities in vivo, part of which is achieved through making a Rad51 filament that is resistant to Srs2 strippase activity (Liu et al.2011).
The prorecombination function of Srs2 in connected to its DNA helicase function and recombination intermediates that have Rad51 on double-stranded DNA, a substrate formed after strand invasion (Dupaigne et al.2008). Regulation of Srs2 in this situation does not involve interaction with PCNA but is stimulated by SUMO and the SUMO-like protein Esc2 (Urulangodi et al.2015; Kolesar et al.2016).
Srs2 IN REPLICATION
As discussed above, Srs2 functions in replication as an anti-recombinase at stalled forks to prevent gaps from being used to initiate HR events. Srs2 helicase appears to have additional roles at the replication checkpoint that may be independent of HR (Yasuhira 2009). srs2 mutants are defective in activation of the DNA damage response (Liberi et al.2000), allowing lethal recombination intermediates to form. This Srs2 activity is modulated by phosphorylation of the Srs2 protein and is mediated by the Cdk1 kinase. The effect of this modification is to target the Srs2 protein to DNA breaks and promote SDSA recombinational repair, using the DNA helicase activity to disassemble any recombination intermediates that may be destined for crossover formation and allow promote SDSA (Saponaro et al.2010).
Recovery from checkpoint arrest also requires Srs2 protein (Vaze et al.2002; Yeung and Durocher 2011) and this appears to result from the strippase activity, which is needed to remove Rad51 from single-stranded DNA after repair. In another study, the interaction of Mrc1 with Srs2 was related to replication and the formation of inappropriate recombination intermediates but was independent of checkpoint signaling (Xu, Boone and Klein 2004). In this case, the important factor seemed to be the establishment of sister chromatid cohesion to promote gap repair.
A study examining the effects of Srs2 overexpression found that the ability of Srs2 to bind to sumoylated PCNA independent of its helicase and ATPase activity was critical in regulating recruitment of other factors to PCNA during replication (Leon Ortiz et al.2011). A further wrinkle in the complex dance of factors binding to sumoylated PCNA has been revealed by studies of the Esc2 protein, which can bind to stalled replication forks and counteract the ability of Srs2 to remove Rad51 from single-stranded regions at stalled forks (Urulangodi et al.2015). The balance between Rad51 filament formation and recombination repair versus gap repair at stalled or damaged forks is regulated by Srs2 removal and degradation and through Srs2 modifications.
Srs2 also has a role in the unwinding of unusual nucleic acid structures at the replication fork (Fig. 4). Both in vivo and in vitro Srs2 can unwind triplet repeat hairpins (Bhattacharyya and Lahue 2004; Dhar and Lahue 2008; Anand et al.2012), permitting replication without expansion or contraction of repeats. In vitro, compared to a random 3΄-overhanging duplex DNA substrate, purified Srs2 helicase displays higher efficiency unwinding or a similar 3΄-overhanging DNA with a triplet-repeat hairpin embedded (Bhattacharyya and Lahue 2005). Whether the Srs2-catalyzed hairpin unfolding is restricted to triplet repeats or is also applied to other hairpin structures in cells remains to be determined.
Figure 4.
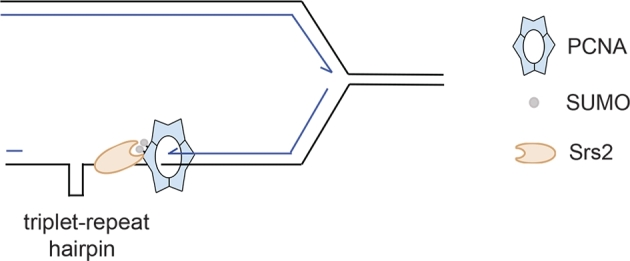
Srs2 catalyzed triplet-repeat hairpin unwinding during replication. Triplet-repeat hairpins form when single-stranded DNA is exposed during replication and may cause expansion or contraction of the repetitive sequences. Srs2 is recruited by PCNA to actively unfold the triplet-repeat hairpin so as to maintain the stability of the repetitive sequences.
Srs2 also has a novel role in preventing mutations from occurring at Top-processed ribonucleotide residues that can arise in DNA during replication. If these residues are not removed by RNase H2-dependent ribonucleotide excision repair pathway (Fig. 5), they can be cleaved by Top1 to produce unligatable DNA nicks bearing a 2΄, 3΄-cyclic phosphate terminated 3΄ end and an –OH group at the 5΄ end. Recent data have shown that Srs2 helicase can unwind from the 5΄-end of the rNMP-derived nicks, and then recruits the Exo1 nuclease to remove the resulting 5΄-Flap to create a gapped duplex DNA for further repair (Potenski et al.2014). Knowing that a secondary processing of 2΄, 3΄-cyclic phosphate terminated nicks by Top1 may cause slippage mutations at the site containing short repetitive DNA sequence (Sparks and Burgers 2015), the Srs2-Exo1 catalyzed gap formation may serve as a mechanism to avoid secondary processing by Top1 and to channel the repair through error-free pathways (Potenski et al.2014) (Fig. 5).
Figure 5.
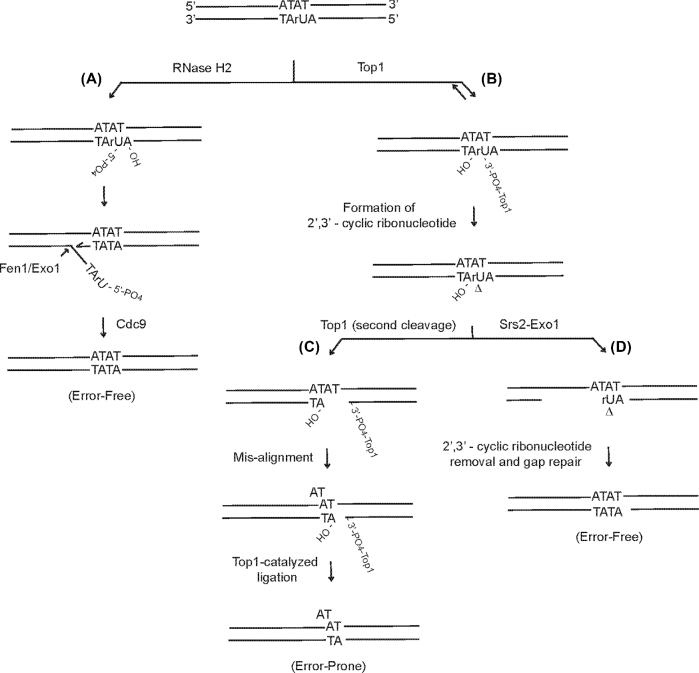
Pathways for removing misincorporated ribonucleotides during replication. Ribonucleotides are incorporated into the genome by replicative DNA polymerases, primarily on the leading strand. A majority of the embedded rNMPs are removed by ribonucleotide excision repair (RER), a coupled reaction of RNase H2, DNA polymerase, Fen1/Exo1 endonuclease and DNA ligase I (Cdc9) as depicted (A). The misincorporated ribonucleotides that escaped from RER pathway are cleaved by Top1 to create nicks with a 3΄ end that is terminated by a 2΄, 3΄-cyclic phosphate and a 5΄-end terminated by a 5΄-hydroxyl group (B). The 2΄, 3΄-cyclic phosphate terminated nicks, although unligatible, may be processed by Top1 again and cause formation of slippage mutations when the misincorporation occurs in short repetitive DNA regions (C). Srs2 helicase and Exo1 nuclease together are able to generate a gap at the 5΄ end of the nick, which prevents Top1-mediated error-prone repair and facilitates error-free gap repair (D).
NON-MOTOR ROLES FOR Srs2
As a multitasking repair factor, Srs2 has also been shown to regulate replication and repair in a manner that is independent of its translocase activity. A biochemical study demonstrated that Srs2 physically interacts with Mus81/Mms4 complex and stimulates its 3΄-flap endonuclease activity even in the absence of ATP. This direct interaction is bridged by Mus81 protein and a domain located between amino acid 783 and 860 of the Srs2 protein (Chavdarova et al.2015). Interestingly, the presence of Mus81-Mms4 complex inhibits the helicase activity of Srs2 (Chavdarova et al.2015). Knowing that Mus81-Mms4 and Srs2 promote crossover and non-crossover pathways, respectively, the functional crosstalk between Mus81-Mms4 and Srs2 may be regulated to balance the repair pathway choice. In a separate study, binding of Srs2 to SUMO-PCNA has been found to attenuate repair synthesis by occluding Polδ and Polη from binding to PCNA (Burkovics et al.2013). Again, this activity does not require the helicase activity of Srs2 (Burkovics et al.2013).
TEMPORAL AND SPATIAL REGULATION OF Srs2
In cells, the functions of Srs2 are tightly regulated to ensure that Srs2 acts on the right substrate and at the right time. Besides the aforementioned CDK-mediated regulation of Srs2 activities, SRS2 is also transcriptionally regulated by cell cycle. Consistent with its role in DNA replication and repair, the expression of SRS2 gene is at a low level and is increased concomitantly with DNA replication and in G2 phase upon exposure to DNA damaging agents (Heude, Chanet and Fabre 1995). Srs2 is actively recruited to the replication fork by binding to sumoylated PCNA, which depends on the E3 ligase activity of Siz1 (Papouli et al.2005; Pfander et al.2005). Binding to PCNA is indispensible for Srs2 to suppress recombination and maintain triplet repeat stability during replication (Papouli et al.2005; Pfander et al.2005). Interestingly, the defects of srs2Δ in repair of 2΄, 3΄-cyclic phosphate terminated nicks can be fully complemented by srs2-1-860 (Potenski et al.2014), a mutant allele that lacks the SUMO and PCNA binding motif, suggesting that Srs2 is recruited to 2΄, 3΄-cyclic phosphate terminated nicks by alterative mechanisms.
SUMMARY AND OUTLOOK
Srs2 is a multifunction protein that acts in replication, recombination and repair. It has a DNA helicase activity and a strippase activity, which is directed against Rad51 nucleofilaments. These activities allow Srs2 to antagonize and promote recombination, dependent on the context and step in the recombination reaction. Some of the regulation occurs at the level of Srs2 modifications. Despite the many roles of Srs2 the protein is not essential, suggesting that potentially lethal intermediates do not form in its absence or they can be successfully processed by other enzymes.
In mammals, a clear Srs2 ortholog is missing. However, the biochemical activities of Srs2 are substituted by several other helicases. Studies have suggested that RecQL5, one of the five mammalian RecQ family helicases, supplies the strippase activity. Deficiency of RecQL5 leads to elevated spontaneous Rad51 foci formation and increase recombination rate. Similar to Srs2 helicase, RecQL5 interacts with both RAD51 and PCNA and the purified RecQL5 helicase displays strippase activity in vitro (Hu et al.2007). Recently, hFBH1 has also been reported to disrupt RAD51 filaments on DNA and is required to limit recombination (Simandlova et al.2013). In addition, PARI, a PCNA and RAD51-interacting protein that contains a degenerate UvrD like helicase domain, was identified to also limit Rad51 foci formation and suppress recombination in cells (Moldovan et al.2012; Burkovics et al.2016). The purified PARI protein, although lacking an apparent ATPase activity, destabilized RAD51 filaments when present at stoichiometric levels (Moldovan et al.2012). The relationship among these strippase activities remains to be determined. Since the mechanism of Srs2΄s function in SDSA is unclear, it is difficult to locate a similar activity in mammals. RTEL1, a helicase first identified as a telomere-length regulator in mammals, was found to be able to disrupt a D-loop structure and promote the SDSA pathway during HR (Barber et al.2008). Interestingly, similar to Srs2 helicase in yeast, RTEL1 was recently found to efficiently unwind triplet repeat hairpins in vitro and suppress repeat expansion in vivo (Frizzell et al.2014). The helicase that substitutes Srs2 for repairing Top1-derived nicks in mammals remains to be determined.
FUNDING
This work was supported by National Institutes of Health grants R00 ES021441 (HN) and CA146940 (HLK).
Conflict of interest. None declared.
REFERENCES
- Aboussekhra A, Chanet R, Adjiri A et al. Semidominant suppressors of Srs2 helicase mutations of Saccharomyces cerevisiae map in the RAD51 gene, whose sequence predicts a protein with similarities to procaryotic RecA proteins. Mol Cell Biol 1992;12:3224–34. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aboussekhra A, Chanet R, Zgaga Z et al. RADH, a gene of Saccharomyces cerevisiae encoding a putative DNA helicase involved in DNA repair. Characteristics of radH mutants and sequence of the gene. Nucleic Acids Res 1989;17:7211–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aguilera A, Klein HL. Genetic control of intrachromosomal recombination in Saccharomyces cerevisiae. I. Isolation and genetic characterization of hyper-recombination mutations. Genetics 1988;119:779–90. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anand RP, Shah KA, Niu H et al. Overcoming natural replication barriers: differential helicase requirements. Nucleic Acids Res 2012;40:1091–105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Antony E, Tomko EJ, Xiao Q et al. Srs2 disassembles Rad51 filaments by a protein-protein interaction triggering ATP turnover and dissociation of Rad51 from DNA. Mol Cell 2009;35:105–15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Armstrong AA, Mohideen F, Lima CD. Recognition of SUMO-modified PCNA requires tandem receptor motifs in Srs2. Nature 2012;483:59–63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barber LJ, Youds JL, Ward JD et al. RTEL1 maintains genomic stability by suppressing homologous recombination. Cell 2008;135:261–71. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barbour L, Xiao W. Regulation of alternative replication bypass pathways at stalled replication forks and its effects on genome stability: a yeast model. Mutat Res 2003;532:137–55. [DOI] [PubMed] [Google Scholar]
- Bernstein KA, Reid RJ, Sunjevaric I et al. The Shu complex, which contains Rad51 paralogues, promotes DNA repair through inhibition of the Srs2 anti-recombinase. Mol Biol Cell 2011;22:1599–607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bhattacharyya S, Lahue RS. Saccharomyces cerevisiae Srs2 DNA helicase selectively blocks expansions of trinucleotide repeats. Mol Cell Biol 2004;24:7324–30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bhattacharyya S, Lahue RS. Srs2 helicase of Saccharomyces cerevisiae selectively unwinds triplet repeat DNA. J Biol Chem 2005;280:33311–7. [DOI] [PubMed] [Google Scholar]
- Broomfield S, Xiao W. Suppression of genetic defects within the RAD6 pathway by srs2 is specific for error-free post-replication repair but not for damage-induced mutagenesis. Nucleic Acids Res 2002;30:732–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burgess RC, Lisby M, Altmannova V et al. Localization of recombination proteins and Srs2 reveals anti-recombinase function in vivo. J Cell Biol 2009;185:969–81. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burkovics P, Dome L, Juhasz S et al. The PCNA-associated protein PARI negatively regulates homologous recombination via the inhibition of DNA repair synthesis. Nucleic Acids Res 2016;44:3176–89. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burkovics P, Sebesta M, Sisakova A et al. Srs2 mediates PCNA-SUMO-dependent inhibition of DNA repair synthesis. EMBO J 2013;32:742–55. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chavdarova M, Marini V, Sisakova A et al. Srs2 promotes Mus81-Mms4-mediated resolution of recombination intermediates. Nucleic Acids Res 2015;43:3626–42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chiolo I, Carotenuto W, Maffioletti G et al. Srs2 and Sgs1 DNA helicases associate with Mre11 in different subcomplexes following checkpoint activation and CDK1-mediated Srs2 phosphorylation. Mol Cell Biol 2005;25:5738–51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Colavito S, Macris-Kiss M, Seong C et al. Functional significance of the Rad51-Srs2 complex in Rad51 presynaptic filament disruption. Nucleic Acids Res 2009;37:6754–64. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Debrauwere H, Loeillet S, Lin W et al. Links between replication and recombination in Saccharomyces cerevisiae: a hypersensitive requirement for homologous recombination in the absence of Rad27 activity. P Natl Acad Sci USA 2001;98:8263–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dhar A, Lahue RS. Rapid unwinding of triplet repeat hairpins by Srs2 helicase of Saccharomyces cerevisiae. Nucleic Acids Res 2008;36:3366–73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dupaigne P, Le Breton C, Fabre F et al. The Srs2 helicase activity is stimulated by Rad51 filaments on dsDNA: implications for crossover incidence during mitotic recombination. Mol Cell 2008;29:243–54. [DOI] [PubMed] [Google Scholar]
- Fabre F, Chan A, Heyer WD et al. Alternate pathways involving Sgs1/Top3, Mus81/Mms4, and Srs2 prevent formation of toxic recombination intermediates from single-stranded gaps created by DNA replication. P Natl Acad Sci USA 2002;99:16887–92. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Friedl AA, Liefshitz B, Steinlauf R et al. Deletion of the SRS2 gene suppresses elevated recombination and DNA damage sensitivity in rad5 and rad18 mutants of Saccharomyces cerevisiae. Mutat Res 2001;486:137–46. [DOI] [PubMed] [Google Scholar]
- Frizzell A, Nguyen JH, Petalcorin MI et al. RTEL1 inhibits trinucleotide repeat expansions and fragility. Cell Rep 2014;6:827–35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gangloff S, Soustelle C, Fabre F. Homologous recombination is responsible for cell death in the absence of the Sgs1 and Srs2 helicases. Nat Genet 2000;25:192–4. [DOI] [PubMed] [Google Scholar]
- Heude M, Chanet R, Fabre F. Regulation of the Saccharomyces cerevisiae Srs2 helicase during the mitotic cell cycle, meiosis and after irradiation. Mol Gen Genet 1995;248:59–68. [DOI] [PubMed] [Google Scholar]
- Hu Y, Raynard S, Sehorn MG et al. RECQL5/Recql5 helicase regulates homologous recombination and suppresses tumor formation via disruption of Rad51 presynaptic filaments. Gene Dev 2007;21:3073–84. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ira G, Malkova A, Liberi G et al. Srs2 and Sgs1-Top3 suppress crossovers during double-strand break repair in yeast. Cell 2003;115:401–11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Islam MN, Paquet N, Fox D et al. A variant of the breast cancer type 2 susceptibility protein (BRC) repeat is essential for the RECQL5 helicase to interact with RAD51 recombinase for genome stabilization. J Biol Chem 2012;287:23808–18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kerrest A, Anand RP, Sundararajan R et al. SRS2 and SGS1 prevent chromosomal breaks and stabilize triplet repeats by restraining recombination. Nat Struct Mol Biol 2009;16:159–67. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keyamura K, Arai K, Hishida T. Srs2 and Mus81-Mms4 prevent accumulation of toxic inter-homolog recombinaiton intermediates. PLoS Genet 2016;12:e1006136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klein HL. Mutations in recombinational repair and in checkpoint control genes suppress the lethal combination of srs2Delta with other DNA repair genes in Saccharomyces cerevisiae. Genetics 2001;157:557–65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kolesar P, Altmannova V, Silva S et al. Pro-recombination role of Srs2 protein requires SUMO (small ubiquitin-like modifier) but is independent of PCNA (proliferating cell nuclear antigen) Interaction. J Biol Chem 2016;291:7594–607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kolesar P, Sarangi P, Altmannova V et al. Dual roles of the SUMO-interacting motif in the regulation of Srs2 sumoylation. Nucleic Acids Res 2012;40:7831–43. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krejci L, Macris M, Li Y et al. Role of ATP hydrolysis in the antirecombinase function of Saccharomyces cerevisiae Srs2 protein. J Biol Chem 2004;279:23193–9. [DOI] [PubMed] [Google Scholar]
- Krejci L, Van Komen S, Li Y et al. DNA helicase Srs2 disrupts the Rad51 presynaptic filament. Nature 2003;423:305–9. [DOI] [PubMed] [Google Scholar]
- Lambert S, Mizuno K, Blaisonneau J et al. Homologous recombination restarts blocked replication forks at the expense of genome rearrangements by template exchange. Mol Cell 2010;39:346–59. [DOI] [PubMed] [Google Scholar]
- Lawrence CW, Christensen RB. Metabolic suppressors of trimethoprim and ultraviolet light sensitivities of Saccharomyces cerevisiae rad6 mutants. J Bacteriol 1979;139:866–76. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Le Breton C, Dupaigne P, Robert T et al. Srs2 removes deadly recombination intermediates independently of its interaction with SUMO-modified PCNA. Nucleic Acids Res 2008;36:4964–74. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leon Ortiz AM, Reid RJ, Dittmar JC et al. Srs2 overexpression reveals a helicase-independent role at replication forks that requires diverse cell functions. DNA Repair (Amst) 2011;10:506–17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liberi G, Chiolo I, Pellicioli A et al. Srs2 DNA helicase is involved in checkpoint response and its regulation requires a functional Mec1-dependent pathway and Cdk1 activity. EMBO J 2000;19:5027–38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu J, Renault L, Veaute X et al. Rad51 paralogues Rad55-Rad57 balance the antirecombinase Srs2 in Rad51 filament formation. Nature 2011;479:245–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marini V, Krejci L. Unwinding of synthetic replication and recombination substrates by Srs2. DNA Repair (Amst) 2012;11:789–98. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mitchel K, Lehner K, Jinks-Robertson S. Heteroduplex DNA position defines the roles of the Sgs1, Srs2, and Mph1 helicases in promoting distinct recombination outcomes. PLoS Genet 2013;9:e1003340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miura T, Shibata T, Kusano K. Putative antirecombinase Srs2 DNA helicase promotes noncrossover homologous recombination avoiding loss of heterozygosity. P Natl Acad Sci USA 2013;110:16067–72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moldovan GL, Dejsuphong D, Petalcorin MI et al. Inhibition of homologous recombination by the PCNA-interacting protein PARI. Mol Cell 2012;45:75–86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morishita T, Furukawa F, Sakaguchi C et al. Role of the Schizosaccharomyces pombe F-Box DNA helicase in processing recombination intermediates. Mol Cell Biol 2005;25:8074–83. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ooi SL, Shoemaker DD, Boeke JD. DNA helicase gene interaction network defined using synthetic lethality analyzed by microarray. Nat Genet 2003;35:277–86. [DOI] [PubMed] [Google Scholar]
- Palladino F, Klein HL. Analysis of mitotic and meiotic defects in Saccharomyces cerevisiae SRS2 DNA helicase mutants. Genetics 1992;132:23–37. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pan X, Ye P, Yuan DS et al. A DNA integrity network in the yeast Saccharomyces cerevisiae. Cell 2006;124:1069–81. [DOI] [PubMed] [Google Scholar]
- Papouli E, Chen S, Davies AA et al. Crosstalk between SUMO and ubiquitin on PCNA is mediated by recruitment of the helicase Srs2p. Mol Cell 2005;19:123–33. [DOI] [PubMed] [Google Scholar]
- Pfander B, Moldovan GL, Sacher M et al. SUMO-modified PCNA recruits Srs2 to prevent recombination during S phase. Nature 2005;436:428–33. [DOI] [PubMed] [Google Scholar]
- Potenski CJ, Niu H, Sung P et al. Avoidance of ribonucleotide-induced mutations by RNase H2 and Srs2-Exo1 mechanisms. Nature 2014;511:251–4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qiu Y, Antony E, Doganay S et al. Srs2 prevents Rad51 filament formation by repetitive motion on DNA. Nat Commun 2013;4:2281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Richard GF, Kerrest A, Lafontaine I et al. Comparative genomics of hemiascomycete yeasts: genes involved in DNA replication, repair, and recombination. Mol Biol Evol 2005;22:1011–23. [DOI] [PubMed] [Google Scholar]
- Robert T, Dervins D, Fabre F et al. Mrc1 and Srs2 are major actors in the regulation of spontaneous crossover. EMBO J 2006;25:2837–46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rong L, Klein HL. Purification and characterization of the SRS2 DNA helicase of the yeast Saccharomyces cerevisiae. J Biol Chem 1993;268:1252–9. [PubMed] [Google Scholar]
- Rong L, Palladino F, Aguilera A et al. The hyper-gene conversion hpr5–1 mutation of Saccharomyces cerevisiae is an allele of the SRS2/RADH gene. Genetics 1991;127:75–85. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saponaro M, Callahan D, Zheng X et al. Cdk1 targets Srs2 to complete synthesis-dependent strand annealing and to promote recombinational repair. PLoS Genet 2010;6:e1000858. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sasanuma H, Furihata Y, Shinohara M et al. Remodeling of the Rad51 DNA strand-exchange protein by the Srs2 helicase. Genetics 2013;194:859–72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schiestl RH, Prakash S, Prakash L. The SRS2 suppressor of rad6 mutations of Saccharomyces cerevisiae acts by channeling DNA lesions into the RAD52 DNA repair pathway. Genetics 1990;124:817–31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Seong C, Colavito S, Kwon Y et al. Regulation of Rad51 recombinase presynaptic filament assembly via interactions with the Rad52 mediator and the Srs2 anti-recombinase. J Biol Chem 2009;284:24363–71. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simandlova J, Zagelbaum J, Payne MJ et al. FBH1 helicase disrupts RAD51 filaments in vitro and modulates homologous recombination in mammalian cells. J Biol Chem 2013;288:34168–80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sparks JL, Burgers PM. Error-free and mutagenic processing of topoisomerase 1-provoked damage at genomic ribonucleotides. EMBO J 2015;34:1259–69. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spell RM, Jinks-Robertson S. Examination of the roles of Sgs1 and Srs2 helicases in the enforcement of recombination fidelity in Saccharomyces cerevisiae. Genetics 2004;168:1855–65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sung P. Yeast Rad55 and Rad57 proteins form a heterodimer that functions with replication protein A to promote DNA strand exchange by Rad51 recombinase. Gene Dev 1997;11:1111–21. [DOI] [PubMed] [Google Scholar]
- Tong AH, Lesage G, Bader GD et al. Global mapping of the yeast genetic interaction network. Science 2004;303:808–13. [DOI] [PubMed] [Google Scholar]
- Ulrich HD. The srs2 suppressor of UV sensitivity acts specifically on the RAD5– and MMS2-dependent branch of the RAD6 pathway. Nucleic Acids Res 2001;29:3487–94. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Urulangodi M, Sebesta M, Menolfi D et al. Local regulation of the Srs2 helicase by the SUMO-like domain protein Esc2 promotes recombination at sites of stalled replication. Gene Dev 2015;29:2067–80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Komen S, Reddy MS, Krejci L et al. ATPase and DNA helicase activities of the Saccharomyces cerevisiae anti-recombinase Srs2. J Biol Chem 2003;278:44331–7. [DOI] [PubMed] [Google Scholar]
- Vaze MB, Pellicioli A, Lee SE et al. Recovery from checkpoint-mediated arrest after repair of a double-strand break requires Srs2 helicase. Mol Cell 2002;10:373–85. [DOI] [PubMed] [Google Scholar]
- Veaute X, Delmas S, Selva M et al. UvrD helicase, unlike Rep helicase, dismantles RecA nucleoprotein filaments in Escherichia coli. EMBO J 2005;24:180–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Veaute X, Jeusset J, Soustelle C et al. The Srs2 helicase prevents recombination by disrupting Rad51 nucleoprotein filaments. Nature 2003;423:309–12. [DOI] [PubMed] [Google Scholar]
- Watts FZ. Sumoylation of PCNA: Wrestling with recombination at stalled replication forks. DNA Repair (Amst) 2006;5:399–403. [DOI] [PubMed] [Google Scholar]
- Xu H, Boone C, Klein HL. Mrc1 is required for sister chromatid cohesion to aid in recombination repair of spontaneous damage. Mol Cell Biol 2004;24:7082–90. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yasuhira S. Redundant roles of Srs2 helicase and replication checkpoint in survival and rDNA maintenance in Schizosaccharomyces pombe. Mol Genet Genomics 2009;281:497–509. [DOI] [PubMed] [Google Scholar]
- Yeung M, Durocher D. Srs2 enables checkpoint recovery by promoting disassembly of DNA damage foci from chromatin. DNA Repair (Amst) 2011;10:1213–22. [DOI] [PubMed] [Google Scholar]