Abstract
Abiotic stresses adversely affect cellular homeostasis, impairing overall growth and development of plants. These initial stress signals activate downstream signalling processes, which, subsequently, activate stress-responsive mechanisms to re-establish homeostasis. Dehydrins (DHNs) play an important role in combating dehydration stress. Rice (Oryza sativa L.), which is a paddy crop, is susceptible to drought stress. As drought survival in rice might be viewed as a trait with strong evolutionary selection pressure, we observed DHNs in the light of domestication during the course of evolution. Overall, 65 DHNs were identified by a genome-wide survey of 11 rice species, and 3 DHNs were found to be highly conserved. The correlation of a conserved pattern of DHNs with domestication and diversification of wild to cultivated rice was validated by synonymous substitution rates, indicating that Oryza rufipogon and Oryza sativa ssp. japonica follow an adaptive evolutionary pattern; whereas Oryza nivara and Oryza sativa ssp. indica demonstrate a conserved evolutionary pattern. A comprehensive analysis of tissue-specific expression of DHN genes in japonica and their expression profiles in normal and PEG (poly ethylene glycol)-induced dehydration stress exhibited a spatiotemporal expression pattern. Their interaction network reflects the cross-talk between gene expression and the physiological processes mediating adaptation to dehydration stress. The results obtained strongly indicated the importance of DHNs, as they are conserved during the course of domestication.
Introduction
Plants effectively survive diverse and variable environments due to elasticity in their cellular metabolism, physiology, and development. However, extreme environmental conditions, such as the increased incidence of droughts, floods, and rising temperatures trigger extensive crop losses in most part of the world [1]. Abiotic stresses adversely affect cellular homeostasis, which, ultimately, impairs overall growth and fitness of plants. A plant’s behavioural responses to drought are complex and involve varied adaptive mechanisms. One such mechanism works via utilizing cellular mechanisms that produce an array of proteins that help with cellular adaptations to drought stress. Proteins encoded by the Late Embryogenesis Abundant (LEA) genes are expressed in response to water deficit [2]. LEA proteins have been identified in many plant species and are classified into six [2] or seven [3] classes. DHNs are a class of hydrophilic, thermostable stress proteins with a high percentage of charged amino acids, and they belong to Group II of the LEA family [4]. They supposedly carry out their function through membrane stabilization by acting as chaperones to prevent the aggregation and/or inactivation of proteins under dehydration [5]. DHNs are also called as RAB proteins because they are usually responsive to abscisic acid (RAB). It is already reported that when plants are exposed to various stresses related to cellular dehydration (e.g., drought, osmotic stress, salinity, and temperature), DHNs accumulate to high amounts in all vegetative tissues [6, 7, 8]. Several transgenic studies revealed that dehydrin gene expression has a positive effect on plant stress tolerance, including cold, drought and salinity. Kumar et al., [9] reported that OsDHN1 gene overexpressed in rice (Oryza sativa L.) confers high tolerance to drought and salt stress. Similarly, Liu et al., [10] reported a positive correlation between ShDHN gene expression and cold and drought tolerance in tomato (Solanum habrochaites L.). Citrus (Citrus unshiu Marcov.) dehydrin gene, CuCOR19 and maize (Zea mays L.) ZmDHN2b when overexpressed in tobacco showed tolerance to cold stress [11, 12]. In an another study, overexpressing dehydrin genes such as ERD10, LTI30, RcDhn5, and DHN-5 in Arabidopsis plants showed tolerance to various abiotic stresses including freezing and salt [13, 14, 7]. Xu et al., [15] reported that overexpression of Brassica juncea BjDHN2 and BjDHN3 in tobacco resulted in higher tolerance to heavy metal stress. Therefore, it is well known that these proteins play an important protective role during cellular dehydration but their precise function remains unclear.
Rice (Oryza sativa L.) is susceptible to drought-induced stress as compared to other cereals [16] resulting in poor seedling vigor [17], fertility and adversely affecting the crop yield. The history of rice domestication has long been a subject of debate. The cultivated rice differs from wild-type progenitors in several ways and has lost many traits found in wild species, for example, loss of grain shattering and transition from perennial to annual crops [18]. Conscious selection by early farmers and unconscious selection due to agricultural practices or environments during long term domestication resulted in diversification of the cultivated rice gene pool across the geographic and climatic boundaries, leading to emergence of independent species, where its wild relatives could not grow at all [19]. Drought survival in rice might be considered a trait with strong evolutionary selection pressure. Therefore it would be interesting to examine the evolution of the drought-responsive protein (DHN) in rice in the light of domestication. Given the significance of DHNs in dehydration stress responses, the present study was performed in 9 species and 2 sub-species of Oryza to identify the presence and importance of DHNs during the course of domestication of rice. This study includes the six wild relatives of rice (Oryza rufipogon, O. nivara, O. longstaminata, O. barthii, O. glumaepatula, and O. meridionalis) and the cultivated O. glaberrima (African rice); the two subspecies of Oryza sativa (Asian), that is, indica and japonica with the AA genome [20]; and a wild variety from BB (O. punctata) [20] and FF (O. brachyantha) [21] genome to elucidate an almost precise picture of diversification, domestication, and evolution with respect to DHN. Of these, O. rufipogon is the species that is the closest to O. sativa ssp. japonica (hereafter mentioned as japonica) and is generally believed to be its progenitor, and O. nivara is believed to be the progenitor of O. sativa ssp. indica (hereafter mentioned as indica) [22].
Materials and methods
Identification of dehydrins and domain detection
To identify DHNs from 9 species and 2 sub-species of rice, a three- step approach was applied. Firstly, Oryza sativa ssp. japonica DHNs from database of Rice Genome Annotation Project (Release 7 of the MSU Rice Genome Annotation Project) were used as query sequences to search and retrieve DHNs from using BLASTP program[23]; applying e-value of 5 and considering the top hits with highest identity value for further analysis. Secondly, all possible sequence architecture for DHNs was searched using HMM profile of DHNs [24]. Finally, DHN sequences were searched in Plant Ensembl Database (www.plants.ensembl.org/) in each rice specie and sub-specie and then by name/function search. The protein sequences of the identified DHN members were then used as queries in multiple databases to ensure that no additional related genes were missed from the database. All sequences that satisfied the requirements were analysed using SMART database (http://smart.embl-heidelberg.de/), and the Conserved Domain Database of the NCBI (https://www.ncbi.nlm.nih.gov/Structure/cdd/cdd.shtml) to eliminate those genes that did not contain the conserved domains and motifs of the DHN family members.
Multiple sequence alignment and phylogenetic analysis
Multiple sequence alignments were done on the amino acid sequences of identified DHNs using ClustelX version 2.0 [25] with default settings. Subsequently, MEGA7.0 software (http://www.megasoftware.net) [26] was used to construct an unrooted phylogenetic tree using the Maximum likelihood method with the bootstrap value of 1000.
Chromosomal localization, whole genome duplication, gene structure and evolutionary analysis
Information about the physical locations of all identified DHN genes on chromosomes was mapped using CIRCOS tool (www.circos.ca/) [27]. Whole genome duplication (WGD) events are crucial for gene families and DHN genes were tested for their synteny conservation status, and the collinear blocks of identified DHNs were plotted using MCScanX tool [28]. Gene structure of DHN genes, showing their exon-intron boundaries and UTR regions, was generated using GSDS server (http://gsds.cbi.pku.edu.cn/) [29]. Interspecies evolutionary pattern was analyzed using PAL2NAL (www.bork.embl.de/pal2nal/) online tool which provided the rate of substitutions between the rice groups. An evolutionary time scale of rice species was generated using an online tool (www.timetree.org).
Gene expression analysis
The DHN genes expression profile was retrieved from Rice Expression Profile Database (http://ricexpro.dna.affrc.go.jp), which provides expression from microarray analysis of tissues/organs encompassing the entire growth of the rice plant under natural field conditions and after treatment with various phytohormones [30].
Gene motif and protein structural analysis
Promoter region sequences of all the retrieved DHN gene sequences were checked for the presence of important cis-regulatory elements using PLANTCARE (http://bioinformatics.psb.ugent.be/webtools/plantcare/html/). MEME de-novo motif detection tool (http://meme-suite.org) was used to find out novel sequence motifs in putative promoter region of DHNs. To check similarities and differences between the evolutionary connected rice groups, in-silico protein structural analysis was performed. Protein structures were modelled using PHYRE2 (www.sbg.bio.ic.ac.uk/~phyre) online server. Structures modelled by the server were then analysed for their structural properties using Chimera (https://www.cgl.ucsf.edu/chimera) 3D structure viewing tool.
Plant growth, treatment and sampling
The rice cv. Nipponbare (Oryza sativa ssp. japonica) was germinated and allowed to grow for 5 day (D) at 26°C in dark and then transferred to Hewitt media for growth. After 10 D of growth, seedlings of uniform size and growth were treated with 10% PEG and 20% PEG under standard physiological conditions of 16 h light (115 μM quanta m-2 s-1) and 8 h dark photoperiod at 25 ± 2°C. Shoot and root samples were collected after 1, 3 and 7 D of treatment, and untreated samples were also collected on each day as control. All the samples were ground in liquid N2 and stored at -80°C for expression analysis.
For development specific expression patterns in various rice tissues/organs, i.e. seedling root (R), mature leaf (ML) and young/immature leaf (IL) and stages of reproductive development; young panicle (YP), anther (A) and gynaecium (G) were used for RNA extraction. Stages of seed development have been categorized according to days after pollination (DAP), middle and late globular embryo/ middle flowering stage (MF) after 5–10 DAP, and dormancy and desiccation tolerance embryo/after flowering stage (AF) after 21–25 DAP.
RNA extraction, quantitative real-time PCR and network analyses
Total RNA was extracted from above samples by using RNeasy plant Mini Kit (QIAGEN, MD) and treated with RNase free DNaseI (QIAGEN, MD) according to manufacturer’s instructions. RNA integrity was checked by electrophoresis and quantified by using a NanoDrop® ND-1000 UV-Vis spectrophotometer (Thermo Fisher Scientific). 2 μg RNA was reverse transcribed using oligo (dT) primers and Superscript II RT (Invitrogen, Rockville, MD, USA) into first- strand c-DNA in a 20-μL reaction as per manual instructions. Quantitative real-time PCR [31] was performed by Power SYBR Green PCR Master Mix (ABI, USA) using the ABI 7500 Fast Real-Time PCR Detection System (Applied Biosystems). The list of eight selected genes and oligonucleotide primers (Eurofins, India) for each gene used for development and drought related expression patterns are listed in S1 Table. Oligonucleotide primers for rice actin gene were used as the internal control for establishing equal amounts of cDNA in all reactions. The reactions were performed using the following cycle conditions: an initial 94°C for 2 min, followed by 30 cycles of 94°C for 30 s, 60°C for 30 s, and 72°C for 30 s, and the final 5 min extension at 72°C. The comparative ΔΔCt method was used to calculate the relative expression levels of different genes by using actin as the reference gene. All the experiments were repeated using three biological replicates. The data were analyzed by two way analysis of variance to confirm the variability and validity of results, and Duncan's multiple range test (DMRT) was performed for multiple comparisons of means. Significance levels were compared at p < 0.05 and the analyses were performed by SPSS 20.0 software. All DHNs from japonica were checked for their interaction network using STRING tool (http://string-db.org/).
Results
Identification, domain detection, multiple sequence alignment and phylogenetic analyses of dehydrins in Oryza species
Dehydrin proteins (DHNs) in Oryza sativa ssp. japonica were reported earlier as class 2 LEA proteins[4]. In the present study a total of 65 dehydrin genes (DHN) were identified from 11 Oryza groups (Table 1). The nomenclature of each DHN is made by following “Gene nomenclature system for rice” by McCouch and Cooperative [32]. We adopted the acronyms of DHN1 to DHN8, in increasing order of their presence on their chromosome number.
Table 1. Structural features of identified DHNs.
| Rice Cultivars | Protein Length (aa) | Name | Mwt (kDa) | pI |
|---|---|---|---|---|
| Oryza barthii (4) | ||||
| OBART02G27280.1 | 290 | ObaDHN1 | 30.9 | 5.9236 |
| OBART11G12460.1 | 326 | ObaDHN2 | 31.3 | 8.202 |
| OBART11G12590.1 | 151 | ObaDHN3 | 15.6 | 10.1527 |
| OBART11G12600.1 | 447 | ObaDHN4 | 45.2 | 9.7825 |
| Oryza brachyantha (5) | ||||
| OB02G34640.1 | 366 | ObrDHN1 | 40.2 | 5.3197 |
| OB11G19890.1 | 330 | ObrDHN2 | 31.3 | 9.2766 |
| OB11G19900.1 | 132 | ObrDHN3 | 13.5 | 10.1673 |
| OB11G19910.1 | 157 | ObrDHN4 | 15.9 | 9.7211 |
| OB11G19920.1 | 169 | ObrDHN5 | 17.0 | 10.0721 |
| Oryza glaberrima (6) | ||||
| ORGLA01G0229900.1 | 654 | OglaDHN1 | 71.2 | 7.3869 |
| ORGLA02G0235300.1 | 290 | OglaDHN2 | 30.9 | 5.9236 |
| ORGLA11G0106700.1 | 326 | OglaDHN3 | 31.3 | 8.202 |
| ORGLA11G0107700.1 | 151 | OglaDHN4 | 15.6 | 10.1527 |
| ORGLA11G0107800.1 | 164 | OglaDHN5 | 16.8 | 9.9226 |
| ORGLA11G0107900.1 | 164 | OglaDHN6 | 16.5 | 9.7765 |
| Oryza glumaepatula (6) | ||||
| OGLUM01G31840.1 | 612 | OgluDHN1 | 66.3 | 6.7087 |
| OGLUM11G12270.1 | 330 | OgluDHN3 | 31.5 | 9.305 |
| OGLUM11G12330.1 | 153 | OgluDHN4 | 15.9 | 9.5319 |
| OGLUM11G12340.1 | 164 | OgluDHN5 | 16.7 | 9.7618 |
| OGLUM11G12350.1 | 164 | OgluDHN6 | 16.6 | 9.7765 |
| OGLUM11G12360.1 | 185 | OgluDHN7 | 18.7 | 10.0477 |
| Oryza meridionalis (1) | ||||
| OMERI02G26520.1 | 290 | OmDHN1 | 30.9 | 5.9236 |
| Oryza nivara (7) | ||||
| ONIVA01G31820.1 | 654 | OnDHN1 | 71.2 | 7.5921 |
| ONIVA02G29800.1 | 292 | OnDHN2 | 31.1 | 5.9236 |
| ONIVA11G12140.1 | 326 | OnDHN3 | 31.2 | 9.0859 |
| ONIVA05G12830.1 | 151 | OnDHN4 | 15.5 | 9.8049 |
| ONIVA05G12790.1 | 176 | OnDHN5 | 17.9 | 10.0476 |
| ONIVA05G12840.1 | 84 | OnDHN6 | 16.7 | 9.7618 |
| ONIVA05G12780.1 | 164 | OnDHN7 | 16.6 | 9.9399 |
| Oryza punctata (7) | ||||
| OPUNC01G27760.1 | 629 | OpDHN1 | 68.3 | 7.7635 |
| OPUNC02G24940.1 | 290 | OpDHN2 | 30.8 | 6.0337 |
| OPUNC11G10660.1 | 327 | OpDHN3 | 31.4 | 9.0614 |
| OPUNC11G10740.1 | 142 | OpDHN4 | 14.9 | 10.0156 |
| OPUNC11G10760.1 | 235 | OpDHN5 | 23.8 | 10.2266 |
| OPUNC11G10750.1 | 160 | OpDHN6 | 16.5 | 9.776 |
| OPUNC11G10780.1 | 168 | OpDHN7 | 16.9 | 9.8538 |
| Oryza rufipogon (7) | ||||
| ORUFI01G30930.1 | 648 | OrDHN1 | 68.3 | 7.7635 |
| ORUFI02G28750.1 | 290 | OrDHN2 | 30.9 | 5.9236 |
| ORUFI11G13160.1 | 349 | OrDHN3 | 33.8 | 9.7286 |
| ORUFI11G13260.1 | 151 | OrDHN4 | 15.6 | 9.805 |
| ORUFI11G13290.1 | 160 | OrDHN5 | 16.2 | 9.8538 |
| ORUFI11G13280.1 | 276 | OrDHN6 | 28.5 | 10.3513 |
| ORUFI11G13270.1 | 59 | OrDHN7 | 6.2 | 10.2178 |
| Oryza longistaminata (7) | ||||
| KN538697.1_FGP002 | 1279 | OlDHN1 | 141.8 | 6.26 |
| KN541601.1_FGP001 | 225 | OlDHN2 | 24.4 | 5.5 |
| KN539530.1_FGP003 | 142 | OlDHN3 | 13.3 | 9.01 |
| KN539940.1_FGP003 | 151 | OlDHN4 | 15.6 | 9.4 |
| KN539940.1_FGP004 | 119 | OlDHN5 | 12.4 | 10.35 |
| KN539940.1_FGP001 | 49 | OlDHN6 | 4.9 | 9.77 |
| KN542973.1_FGP002 | 109 | OlDHN7 | 11.3 | 9.5 |
| Oryza sativa ssp. japonica (8) | ||||
| LOC_Os01g50700.1 | 653 | OsjDHN1 | 71.0 | 7.07 |
| LOC_Os02g44870.1 | 291 | OsjDHN2 | 30.9 | 5.68 |
| LOC_Os03g45280.1 | 93 | OsjDHN3 | 10.4 | 6.7 |
| LOC_Os11g26570.1 | 327 | OsjDHN4 | 31.3 | 8.95 |
| LOC_Os11g26750.1 | 152 | OsjDHN5 | 15.5 | 9.13 |
| LOC_Os11g26760.1 | 165 | OsjDHN6 | 16.7 | 9.25 |
| LOC_Os11g26780.1 | 165 | OsjDHN7 | 16.5 | 9.27 |
| LOC_Os11g26790.1 | 173 | OsjDHN8 | 17.3 | 9.19 |
| Oryza sativa ssp. indica (7) | ||||
| BGIOSGA004279-PA | 639 | OsiDHN1 | 69.4 | 7.2237 |
| BGIOSGA005869-PA | 292 | OsiDHN2 | 31.1 | 5.9236 |
| BGIOSGA018448-PA | 326 | OsiDHN3 | 31.2 | 9.0859 |
| BGIOSGA034054-PA | 151 | OsiDHN4 | 15.6 | 9.8049 |
| BGIOSGA034053-PA | 164 | OsiDHN5 | 16.7 | 9.7618 |
| BGIOSGA034051-PA | 164 | OsiDHN6 | 16.5 | 9.7921 |
| BGIOSGA034052-PA | 489 | OsiDHN7 | 54.8 | 6.93 |
The identified genes were further checked for the presence of DHN domain. There are 15 reported architectures for DHN domain in plants, including monocots and dicots (http://pfam.xfam.org/family/PF00257#tabview=tab1). In our analysis we found only 4 domain architecture among 11 Oryza groups, viz. single small DHN, single large DHN, DHN-DNAJ, and DHN-DNAJ and DNAJ-X domains (S1 Fig). Exceptionally, OsiDHN7 protein was observed with presence of FAR-1 domain.
The DHNs were further aligned by Clustal X software to confirm their sequence similarity. Interestingly, multiple sequence alignment of DHNs revealed the absence of K-seg in few rice DHN such as OnDHN6, OrDHN7, OlDHN3 and OlDHN6. During dehydration, the K-segments adopt α-helical conformation through the formation of class A2 amphipathic α-helix thus enhancing their amphipathic character in protein-protein or protein-biomembrane interactions [33]. It has been suggested by Koag et al [34] that these interactions between partly dehydrated surfaces of DHN molecules and other proteins and/or biomembranes, present the basis of dehydrin protective functions. OlDHN6 and OlDHN7 showed absence of S-seg which indicates the lack of phosphorylation of the S-segment which is associated with its transport to the nucleus and/or cation binding [33] (S2 Fig). DHNs varied in their length from 59 aa to 654 aa, molecular weight in the range of 13 to 71 kDa, and pI in the range of 5.6 to 10.3 (Table 1). Three DHN genes showed remarkable conservation in 7, 8 and 9 species, respectively, and were localised on chr1, 2 and 11, respectively. The DHN localised on chr 1 (DHN1) encoded a protein of 600 aa and had a representative single DHN domain with DNAJ and DNAJ-X domains. The DHN localised on chr 2 encoded for a protein of 290-293aa, and contained a single small DHN domain. The DHN localised on chr11 encoded a protein of 300–350 aa and had a single large DHN domain.
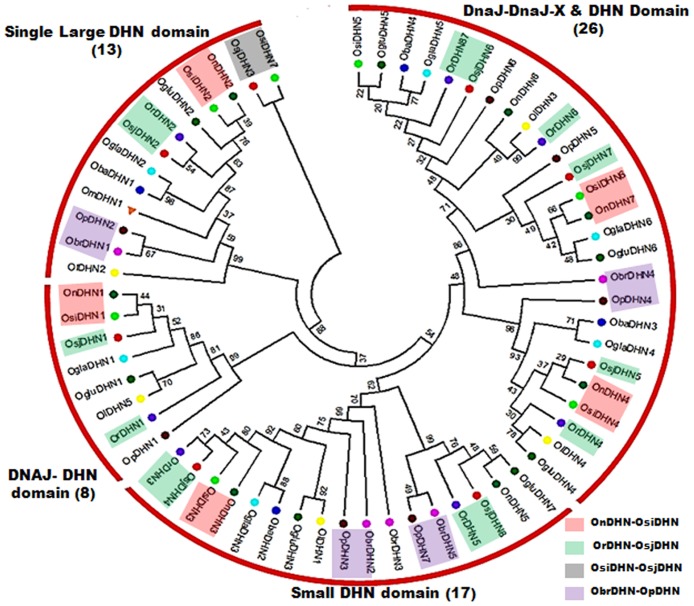
To study the phylogenetic relationships among rice species we constructed an unrooted phylogenetic tree. Based on our phylogenetic results, the DHN’s could be divided into four groups, corresponding to four obtained DHN domain architecture in rice (Fig 1). Eight DHNs with DHNJ and DHN domain get clubbed into one clade, while 13 members containing single large DHN domain assembled in a single pool creating another clade. Similarly, 17 DHNs with single DHN domain forms another clade. 26 DHNs with DnaJ-DnaJ-X and DHN domain formed a clade bifurcating into two sub-clades. OnDHN2-OsiDHN2, OnDHN1-OsiDHN1, OnDHN7-OsiDHN6, OrDHN3-OsjDHN4 and OrDHN2-OsjDHN2 pairs were found to be proximally placed and had similar protein length, shared same clade with similar domain architecture pairs. Interestingly, the DHN’s of African domesticated variety O. glaberrima were also found proximal to both japonica and indica.
Fig 1. Phylogenetic analysis of the DHN gene in 11 rice species.
The tree was calculated with MEGA 7.0 software.
Chromosomal localization, whole genome duplication, gene structure and evolutionary analysis
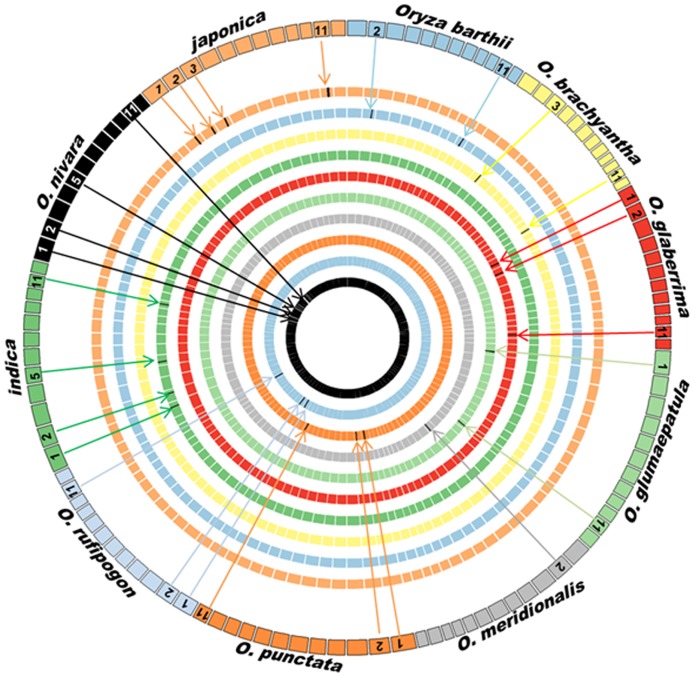
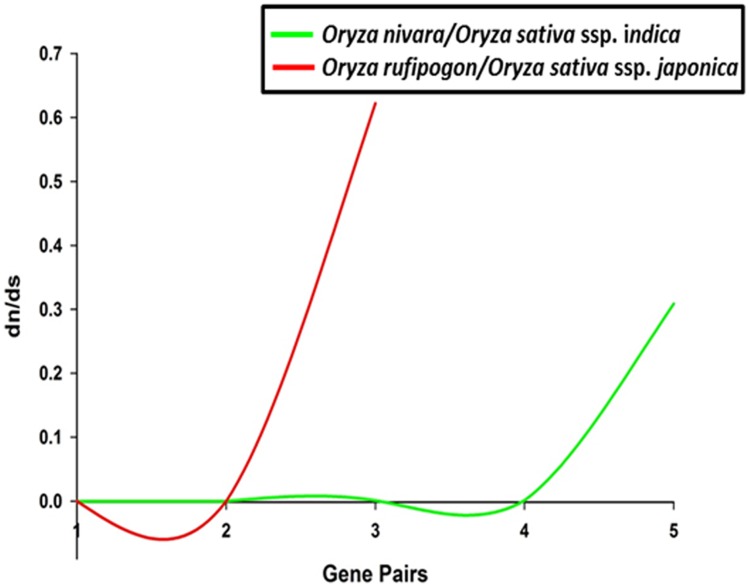
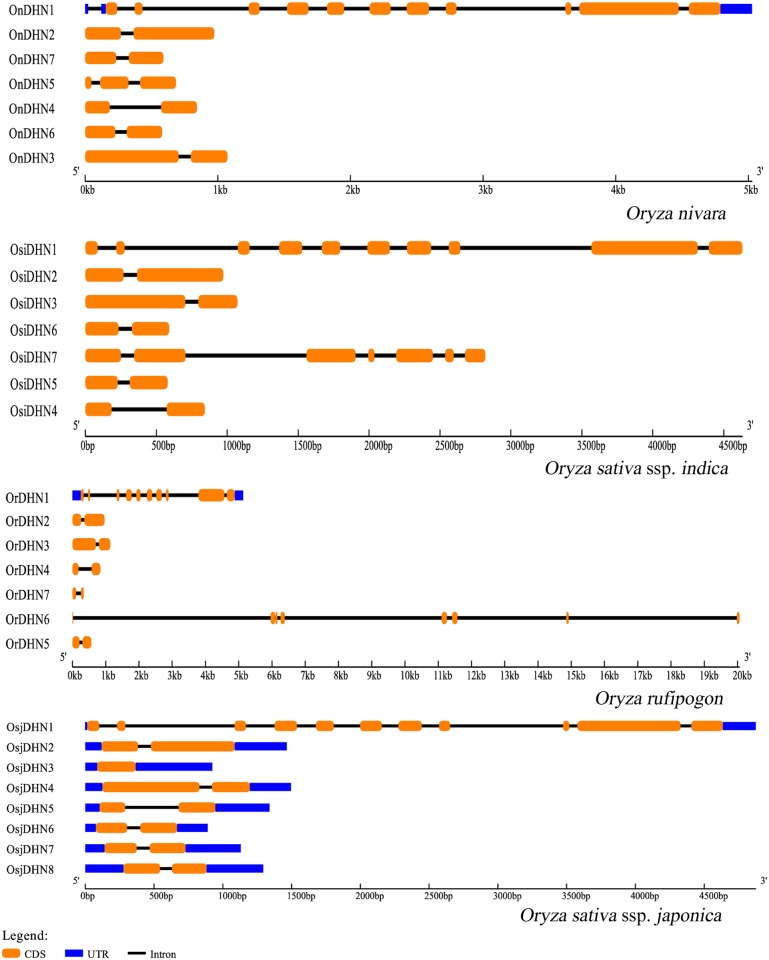
DHN genes were located on chromosomes 1, 2, 3, 5, and 11. The largest cluster was found on chromosome 11 while the others were majorly distributed on chromosome 1 and 2 (Fig 2). Gene duplication is recognized as an important requirement for the diversification of gene function. The identified DHN genes were checked for their synteny conservation status and the collinear blocks of DHNs were plotted. Overall, one duplication event was observed between chr 1–5 in O. glumaepatula, O. barthii and O. nivara. Two duplication events were observed in O. glaberrima, O. brachyantha and O. punctata, one between chr 1–5 and another between chr 8–9 (S3 Fig). The time tree was constructed for the approximate estimation of divergence in rice species. The time tree (S4 Fig) reflected that the Asian rice group diverted around 0.825 MYA (million years ago) whereas the African rice group diverted around 2.23 MYA. To check the evolutionary pressure, rate of substitution (synonymous and non-synonymous) was checked for all DHN sequences. We observed similar values for ds in Asian rice group, while their dn values varied (S2 Table). To further check evolutionary pressure on gene pairs we applied dn/ds ratio between orthologous gene pairs that has been calculated to analyse the synonymous and non-synonymous substitution rate for O. nivara-indica (hereafter mentioned as N-I) and O. rufipogon-japonica (hereafter mentioned as R-J) as they are evolutionary connected. Selection of ortholog gene pairs of rice groups was based on best hits of reciprocal blast. It could be inferred from 3 gene pairs that N-I followed evolutionary pattern to conserve the ancestral state, although 2 gene pair tends to follow adaptive pattern (Fig 3). The comparison of gene structure of DHN among N-I and R-J also support their independent evolutionary descent. In the case of R-J the CDS stretches are similar with the exception of absence of UTR region in japonica. Although N-I also shows similar gene structure for their orthologous DHN genes (Fig 4). The gene structures for the remaining rice species are given in S5 Fig.
Fig 2. Chromosomal localisation of DHN genes in rice species; each colour and square representing specie and a chromosome respectively, and they are numbered clockwise.
Fig 3. Divergence of DHN genes between orthologous gene pairs of N-I and R-J showing evolutionary substitution pressure on gene pairs.
Fig 4. Gene structure of O. nivara-indica and O.rufipogon-japonica rice species showing their CDS, intron boundaries and UTR regions in orange black and blue colour respectively.
Gene expression
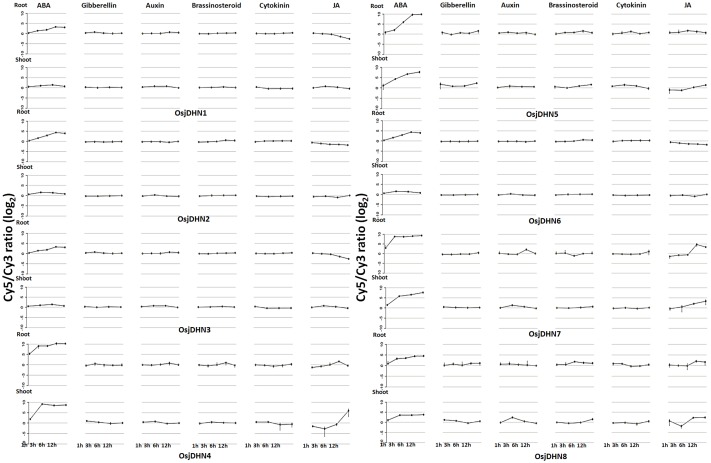
Expression profile of DHN genes has been retrieved and analysed under different hormone treatments which provided significant information regarding their involvement in hormonal signalling cascade. Expression data of DHNs under different hormonal conditions is taken from RiceXpro database (http://ricexpro.dna.affrc.go.jp/). The analysis revealed that, apart from phytohormone abscisic acid (ABA) and jasmonic acid (JA), other hormones do not alter expression of DHN gene. However, root expression data of OsjDHN1 demonstrates the only instance of upregulation by cytokinin treatment. Dehydrin proteins play key roles in defences against stress, in either an ABA independent or -dependent manner. In our analysis, all genes were up-regulated by abscisic acid. In our study, the expression of DHN genes in response to JA demonstrated up-regulation in most of the genes for both root and shoot, though few genes did not show induction, while OsjDHN1, OsjDHN2, OsjDHN3 in root showed down-regulation (Fig 5).
Fig 5. Expression profile of DHN genes in different hormonal conditions.
Conserved motif search & structural analysis of DHN proteins
cis-regulatory elements are located upstream of genes and act as binding sites for transcription factors (TFs) and they have essential functions in determining the tissue-specificity or stress-responsive expression patterns of genes [35]. In order to understand potential functions of these DHN genes, 1000 bp regions upstream of the transcriptional start site were extracted and applied to identify cis-regulatory elements and other important motifs. Essential regulatory elements such as TATA box and CAAT box are present along with some other regulatory elements. A number of biotic stress-related cis-elements like EIRE, BOX W1, 5UTR Py-rich [33] were identified. Abiotic stress-related cis-elements were found in the putative promoters of DHN genes in rice, as for example: dehydration sensitive element (DRE), induction of CBF exspression region 2 (ICEr2), LTR motif (low temperature), Myb binding site (MBS) and sulfur-responsive element (SURE) [36]. Hormone-related cis-elements like- Abscisic acid responsive (CE-3, Motif IIb and ABRE)[37], auxin responsive (Aux RR), gibberellin-responsive (GARE, P Box) and Methyl jasmonate responsive (MeJA-RE, CGTCA) were found in the putative promoters of DHN genes in rice. The salicylic acid (SA) response is reflected by the presence of TCA elements [2]. These cis-elements are counted and classified in Table 2.
Table 2. Comparison of regulatory elements in 11 rice DHN promoter regions.
| cis-element | Number of cis-elements | Function | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Oba | Obr | Ogla | Oglu | Ol | Om | On | Op | Or | Osj | Osi | ||
| ABRE | 1–3 | 3 | 0 | 3 | 1–3 | 0 | 3 | 3 | 0 | 3 | 3 | Abscisic acid responsiveness |
| (GACACGTACGT) | ||||||||||||
| DRE | 1 | 0 | 1 | 0 | 1 | 1 | 1–2 | 0 | 1 | 1 | 1 | Drought and cold responsiveness |
| (ACCGAC) | ||||||||||||
| LTR | 1 | 1–2 | 0 | 0 | 1 | 0 | 1–2 | 0 | 0 | 1–2 | 1–2 | Low-temperature responsiveness |
| (CCGAAA) | ||||||||||||
| TC-rich repeats | 0 | 0 | 1 | 1 | 0 | 1 | 0 | 1 | 1 | 0 | 0 | Stress and defense responsiveness |
| (ATTCTCTAAC) | ||||||||||||
| MeJA-RE | 2 | 1–3 | 1 | 1 | 2 | 1 | 1–3 | 1 | 1 | 1–3 | 1–3 | MeJA-responsiveness |
| (CGTCA) | ||||||||||||
| TCA element | 1 | 1–2 | 1 | 1 | 1 | 1 | 1–2 | 1 | 1 | 1–2 | 1–2 | Salicylic acid responsiveness |
| (CAGAAAAGGA) | ||||||||||||
| MBS | 0 | 0 | 1 | 1 | 0 | 1 | 0 | 1 | 1 | 1–3 | 1–3 | MYB-binding sites |
| (CGGTCA) | ||||||||||||
| Box-W1 | 0 | 0 | 1 | 0 | 0 | 1 | 0 | 0 | 1 | 0 | 0 | Fungal elicitor responsive |
| (TTGACC) | ||||||||||||
| CGTCA | 2 | 3 | 1 | 1 | 2 | 1 | 3 | 1 | 1 | 3 | 3 | Me-JA responsive |
| TGACG | 3 | 1 | 1 | 1 | 3 | 1 | 1 | 1 | 1 | 1 | 1 | Me-JA responsive |
| GARE | 1 | 0 | 1 | 0 | 1 | 1 | 0 | 0 | 1 | 0 | 0 | gibberellin-responsive |
| (TCTGTTG) | ||||||||||||
| Aux-RR | 0 | 2 | 0 | 0 | 0 | 0 | 2 | 0 | 0 | 2 | 2 | Auxin responsive |
| (GGTCCAT) | ||||||||||||
| CE-3 | 1 | 1 | 0 | 0 | 1 | 0 | 1 | 0 | 0 | 1 | 1 | ABA & VP1 responsive |
| (GACGCGTG) | ||||||||||||
| EIRE | 0 | 2 | 0 | 0 | 0 | 0 | 2 | 0 | 0 | 2 | 2 | Elicitor responsive |
| (TTCGACC) | ||||||||||||
| P-Box | 0 | 2 | 0 | 0 | 0 | 0 | 2 | 0 | 0 | 2 | 2 | Gibberellin responsive |
| (CCTTTTG) | ||||||||||||
| SURE | 0 | 0 | 1 | 1 | 1 | 0 | 1 | 0 | 0 | 1 | 1 | Sulphur responsive element |
| (GAGAC) | ||||||||||||
| PHR1-binding site | 1 | 0 | 1 | 0 | 0 | 0 | 1 | 0 | 1 | 1 | 1 | Phosphorus starvation responsive |
| (GNATATNC) | ||||||||||||
| 5UTR Py-rich | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | Biotic stress responsive |
| (TTTCTTCTCT) | ||||||||||||
| Motif IIb | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | ABA responsive |
| (CCGCCGCGCT) | ||||||||||||
| GC motif | 1 | 0 | 1 | 0 | 0 | 0 | 1 | 0 | 1 | 1 | 1 | Enhancer like, anoxic specific inducibility |
| (AGCGCGCCG) | ||||||||||||
| ARE | 1 | 0 | 1 | 1 | 1 | 0 | 0 | 1 | 1 | 0 | 0 | Essential for anaerobic induction |
| (TGGTTT) | ||||||||||||
| MYCR | 1 | 1 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | Water deficit, ABA responsive |
| (CACATC) | ||||||||||||
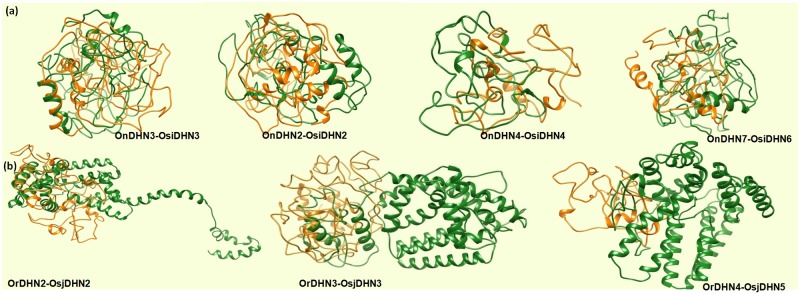
The structure of each DHN protein was analysed and modelled using PHYRE2 web server showed abundance of loops in the secondary structure of DHNs in japonica, indicative of divergence, whereas the secondary structure of O. rufipogon comprised mainly of α helices (Fig 6a). Interestingly, in the case of N-I, abundance of loops was observed in all orthologous pairs (Fig 6b).
Fig 6. Structural prediction of DHN genes, in-silico structural analysis of orthologous gene pairs of N-I and R-J.
Proteins in green from (a) O. rufipogon and (b) O. nivara and proteins in orange indicate the structure of DHN proteins from (a) japonica and (b) indica.
Expression profiles of DHN genes in various tissues, organs, and developmental stages
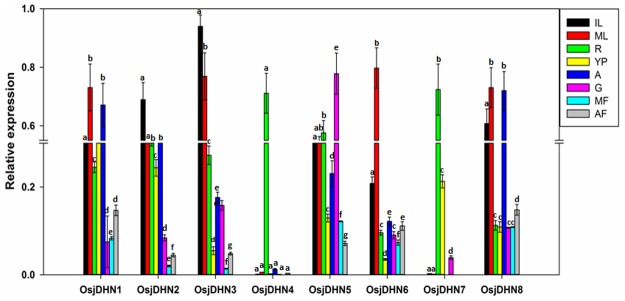
qRT-PCR analysis of various tissues, organs and developmental stages indicated high variability in transcript abundance of DHN genes in rice (Fig 7). Majority of these genes, except OsjDHN4 and OsjDHN7, showed relatively high expression in vegetative organs like root (R), mature leaf (ML), immature/young leaf (IL) and reproductive organ like androecium (A). However, the gene showed lower transcripts in gynoecium (G), young panicle (YP), middle stage of flowering and after stage of flowering. These findings indicated spatio-temporal expression characteristics of DHNs. Two genes viz. OsjDHN4 and OsjDHN7 showed high expression only in root, indicating their possible function in dehydration tolerance. All DHNs except OsjDHN1 and OsjDHN4 accumulated more in leaves (IL and ML). Roots also accumulated higher amount of DHNs, however OsjDHN6 and OsjDHN8 expression was lesser in comparison to all the DHNs in root tissue. OsjDHN4 specifically accumulates in roots in comparison to all other DHNs and all other tissues.
Fig 7. Relative expression analysis of tissue- or organ-specific DHN gene expression profiles in rice species.
R, roots, S, mature leaf (ML), immature/young (IL) androecium (A) gynoecium (G), young panicle (YP) middle flowering stage (MF) and after flowering stage (AF.) The expression levels of genes are presented using fold-change values transformed to Log2 format.
Expression profiles of DHN genes under PEG treatment in japonica rice and gene interaction network
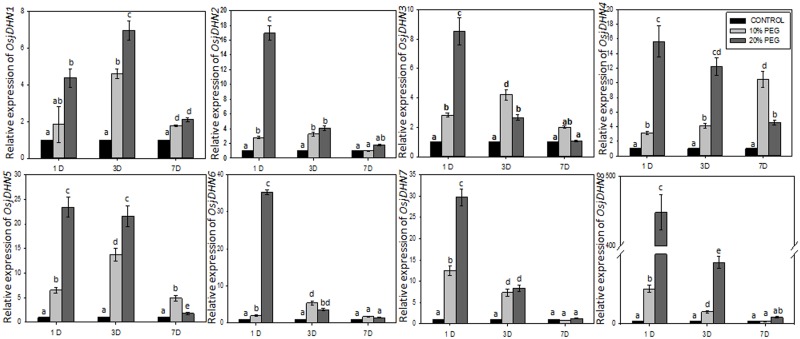
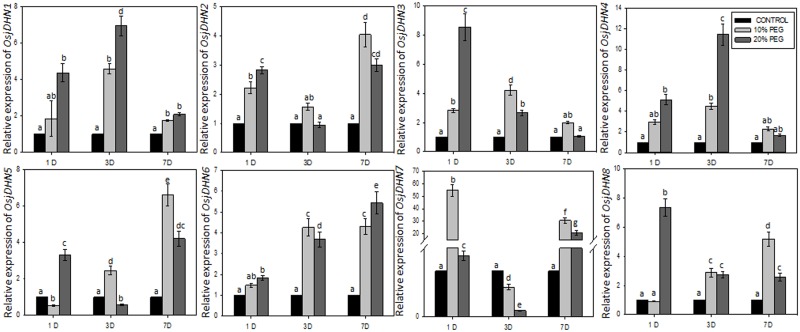
The expression analysis of 8 genes in shoots, on day (D) 1 and moderate (10%PEG) stress condition resulted in nearly 3 fold increase in expression of OsjDHN2, OsjDHN3, and OsjDHN4. However, OsjDHN7 and OsjDHN8 demonstrated nearly 12 fold increase in transcript abundance under same condition. During severe (20%PEG) drought stress, all DHN genes were upregulated with OsjDHN8 showing 448 fold increase in expression pattern. OsjDHN5, OsjDHN6 and OsjDHN7 demonstrated almost 30 fold increase in their transcript abundance. Thus, in shoots, on D1 more DHNs expressed significantly during 20% PEG induced drought stress. On D3, OsjDHN4, OsjDHN5 and OsjDHN7 showed 10 fold greater expressions during 20% PEG treatment as compared to 10% PEG treatment. However, other DHN genes expressions were almost similar under moderate and severe dehydration stress. However, on D7, only OsjDHN4 and OsjDHN5 showed transcript abundance during moderate stress conditions; whereas only OsjDHN4 was upregulated during 20% PEG induced dehydration stress (Fig 8).
Fig 8. Relative expression analysis of DHN genes in shoots.
The expression levels of genes are presented using fold-change values transformed to Log2 format compared with control. All the values are means of triplicate ± SD. Values marked with similar letters are not significantly (Duncan’s test: p < 0.05) different in a particular tissue/treatment.
In roots, on D1, only OsjDHN7 exhibited around 55 fold increase in expression level during moderate stress conditions; whereas OsjDHN8 expression levels were high during 20% PEG treatment. On D3, expression level of OsjDHN4 was around 6 fold greater during 20% PEG treatment as compared to 10% PEG treatment. However, other DHN genes expressions were almost similar under moderate and severe dehydration stress. On D7, almost all DHNs were upregulated, expect for OsjDHN1 and OsjDHN4; interestingly, these two genes were upregulated on D3 (Fig 9). Unexpectedly, the transcript abundance on D3 and 7 was reversed in roots and shoots, thus signifying the spatiotemporal expression pattern of DHNs during drought conditions. On D7, OsjDHN2, OsjDHN5, OsjDHN6, and OsjDHN8 showed nearly 4~5 fold higher transcript abundance; OsjDHN7 demonstrated 30 fold higher expression during 10% PEG treatment. During 20% PEG treatment OsjDHN4 and OsjDHN7 showed 11 and 21 fold greater expression, while OsjDHN5 and OsjDHN6 showed nearly 4~5 fold higher transcript abundance (Fig 9).
Fig 9. Relative expression analysis of DHN genes in roots for the response patterns to moderate (10% PEG) and severe drought (20% PEG) conditions.
The expression levels of genes are presented using fold-change values transformed to Log2 format compared with control. All the values are means of triplicate ± SD. Values marked with similar letters are not significantly (Duncan’s test: p < 0.05) different in a particular tissue/treatment.
OsjDHN7 had ABRE, DRE, LTR rich regulatory elements in their promoter region. cis-element of OsjDHN5 was exclusively rich in ABA- responsive elements. However, both ABRE and DRE were absent in OsjDHN1, other regulatory elements like MYB, TC rich repeats and 5’ UTR Py-rich were present (S3 Table).
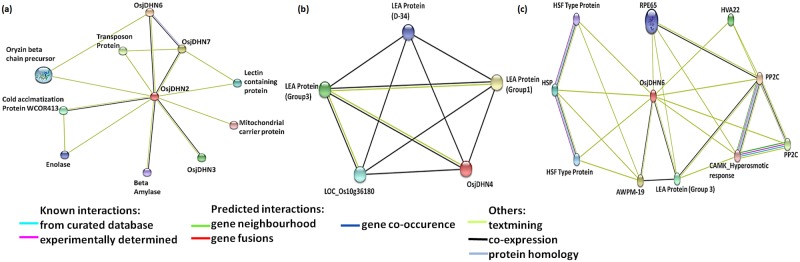
The gene interaction network of OsjDHN2, OsjDHN4, OsjDHN6, and OsjDHN7 further demonstrates the interaction pattern where the network involves other DHNs. OsjDHN2 networks also involve OsjDHN3, OsjDHN6, OsjDHN7 and cold acclimatization protein (WCOR413) and other metabolic pathways (Fig 10a). On the other hand, OsjDHN4 requires the presence of LEA genes in their interactional network (Fig 10b). In the case of OsjDHN6, we found the presence of OsjDHN7, which further interacts with LEA proteins (Fig 10c). OsjDHN6, showed an interaction with HSPs, an abscisic acid-induced protein (HVA22) and cold shock-resistant genes (AWPM-19).
Fig 10. Gene interaction network of OsjDHN7 (a), OsjDHN4 (b), and OsjDHN6 (c).
Discussion
Dehydrins are protective and ubiquitous proteins that are found in plants undergoing osmotic stress, for example, cold, high salinity, and drought. Domestication, being a unique form of biological evolution, may decrease the overall fitness of plants in the wild but increase adaptability when completely dependent on humans for their survival, leading to a loss of certain traits. Drought stress is posed by adverse environmental conditions, and surviving drought could be viewed as a trait that is more related to wild species and diversification. Complete and accurate identification of DHN genes in wild and domesticated species is an essential step towards further evolutionary and functional studies of the gene family. By definition, all DHNs contain, at least one copy of a highly conserved sequence, the K-seg, and may or may not include other conserved sequences, the S-seg and the Y-seg [38]. On the basis of the presence of the conserved sequences, we identified 65 DHNs in eight wild, one domesticated species and two rice sub-species. The absence of K-seg in DHN such as OnDHN6, OrDHN7, OlDHN3 and OlDHN6 was observed. One such DHN, lacking complete K-segment has been also been reported by Perdiguero et al [39]However, these four proteins show high overall similarity in domain architecture and homology with typical dehydrins, especially in N-terminal region, S-segment and Y-segment, which leads us to still consider them as DHNs.
The presence of DHNs in all these diverse rice species is indicative of their significance, as they have not been excluded during the course of domestication. The protein domain architecture is related to evolutionary dynamics [40], and the presence of only a4-domain architecture among Oryza species and sub-species indicates their functional specificity for rice groups. Three DHN genes with conserved chromosomal localization in 7, 8, and 9 species signifies the importance of DHNs in domestication as well as evolution of rice, and thereby emphasizing their significance in the dehydration cascade in different Oryza species.
The phylogenetic analysis indicated protein pattern similarity between O. rufipogon with japonica and O. nivara with indica. The phylogenetic tree supports the independent origin of japonica and indica from O. rufipogon and O. nivara respectively, as the DHN of these species are paired and share the same clade with their diverged wild progenitors. OrDHN1-OsjDHN1 is in one clade but is distantly placed in the phylogenetic tree, signifying the adaptive gain during the course of evolution. However, the proximity of African domesticated rice- O. glaberrima, with Asian domesticated rice—japonica and indica, indicates convergent yet independent selection of common genes during two geographically distinct domestication processes [41]. Few OnDHN-OsiDHN and OrDHN-OsjDHN pairs were found to be in one clade but not paired, as they must have probably lost one of their conserved stretches (K, Y, or S) but were able to maintain their functionality. This again supports the evolutionary adaptation of DHNs during the course of domestication. Evolutionary proximity was observed between the BB genome representative- O. punctata and the FF genome representative- O. brachyantha, where the DHN representatives of both species were present in one clade, indicating the evolutionary relationship between BB and FF genomes. Gene duplication is recognized as one of the important requirements for diversification of gene function [42]. Expansion of the DHN family has generally occurred through tandem and whole-genome duplication (WGD) events. For example, 3 genes each from tandem duplications (TD) and WGD were reported in Arabidopsis [43], resulting in an increase of 6 DHN genes. Similarly in poplar, WGD and TD events were responsible for an increase of 3 and 2 DHN genes [44], respectively. Three TD events have been reported in japonica rice [4]. Overall, two duplication events were observed, one between chr 1–5 and another between chr 8–9, except for O. nivara, no DHN was found to be present on either chr 5, 8 or 9. As mentioned previously, DHN1 encodes a protein with single DHN domain with DNAJ and DNAJ-X domains. The duplicate genes found on chr 5, 8 and 9 were cross-checked for domain detection and we found that collinearity on chr 5, 8 and 9 was observed due to the presence of larger stretch of DNAJ and DNAJ-X domain and not on the basis of DHN domain. Thus it could be inferred that, despite of the presence of segmental duplication event, no DHN exhibit synteny conservation. The result of time of divergence of Asian rice group and their substitution analysis implies that the evolution of DHN proteins of indica is connected to O. nivara and demonstrates a conserved evolutionary pattern, which is also supported by the gene structure of the DHN genes in both species. On the other hand, the substitution pattern of R-J DHN pairs shows an adaptive evolutionary pattern. It has been already reported that japonica grows in a much varied environment as compared to indica, and the former might have gained divergent features as compared to O. rufipogon during the course of domestication. The gene structure analysis of DHNs in N-I and R-J in our study is also in agreement with the independent evolutionary descent of japonica and indica.
Dehydrins are major key players in providing defences against stress. However, whether they are utilized in a hormone-independent or -dependent manner is still unclear. For example, Arabidopsis At1g54410 is induced by cold but not by abscisic acid [43]. In our analysis, all DHN genes were upregulated by this hormone, indicating their possible involvement in ABA-dependent pathways. It has also been reported that many genes involved in jasmonic acid (JA) biosynthesis and signalling are induced by drought and cold treatment [45]. Similar results have been observed when the overexpression of DHN5 in wheat resulted in down-regulation of genes encoding 3 members of JAZ (jasmonate- ZIM domain) proteins, which are negative regulators of JA signalling. Thus, differential expression of JA might depend on the complex inter-regulation of the signalling cascade. cis-regulatory elements, being located upstream of genes and acting as binding sites for TFs, are instrumental in determining the tissue specificity or stress-responsive expression patterns of genes [35]. When taken together, all of the putative regulatory elements identified within rice species and sub-species, the promoters had the most diverse collection of putative cis-regulatory elements, including several involved in stress response, like MBS involved in drought inducibility in OsGATLs in rice [46]; drought and salinity together or alone lead to enrichment of 2 cis elements ABRE and DRE [47]; and hormone signalling like CE-3 and Motif IIb [37]. Presence of biotic stress responsive elements like Box-W1 and EIRE also indicates the diverse activity of DHNs.
DHNs being intrinsically disordered proteins (IDPs) with ~ 0.7–5% of α-helices [48], these unstructured states assist the protective functions of DHN proteins, resulting in their functional versatility [49]. Proteins belonging to the same family may contain a similar active domain but due to their conformational changes, they may show different behaviours. Our results demonstrated the change in folding pattern of DHNs from O. rufipogon (comprised mainly of α-helices) have evolved into IDPs (as seen in japonica), providing functional benefits, such as an increased interaction surface area and conformational flexibility to interact with several targets. However, indica and O. nivara exhibited a similar structural prediction. This is in corroboration with the results obtained by synonymous and non-synonymous substitution rate analysis.
The expression analysis indicated a contrasting expression pattern between vegetative and reproductive organs. OsjDHN3 showed high expression only in leaves (IL and ML). One possibility is that the high osmotic potential is generated in the cytoplasm of guard cells of leaves during stomatal opening which probably promote accumulation of DHNs in these tissues. The high osmotic potential in the cytoplasm of open guard cells promotes water stress on the nucleus. Thus, DHNs may have a role in protecting this compartment from dehydration stress. This is in agreement with the postulated protective function for these proteins during osmotic stress caused by either drought, salinity or freezing [50]. Two genes, that is, OsjDHN4 and OsjDHN7, showed high expression only in roots; thus, possibly involved in the water level balance in the roots. As roots are the primary target for dehydration stress DHN accumulation in roots are in accordance with their protective role during dehydration stress. OsjDHN5 was the only gene exhibiting transcript abundance in gynoecium. However all genes except OsDHN4 showed transcript abundance in reproductive tissue. These DHNs may have protective role against the desiccation or dehydration stress during the development of seeds. Roots are generally believed to be the main organs involved in drought stress, although our study showed a greater induction rate of DHN genes in shoots. Interestingly, a unique pattern was observed in gene induction by dehydration in both the tissues. In shoots, higher expression levels of OsjDHNs were observed during initial phase (D1) with moderate stress. However, OsjDHN8 was found to be upregulated by 448 fold during severe stress conditions. Contrarily, during severe and prolonged dehydration stress only OsjDHN4 was upregulated by 10 fold. In roots, almost all OsjDHN’s were upregulated during moderate stress. OsjDHN7 demonstrated 30 and 20 fold higher expression during moderate and severe stress, respectively. OsjDHN7 was the only gene with 12 fold higher transcript abundance on D3 during severe stress. The transcript abundance on D3 and 7 was reversed in roots and shoots, thus signifying the spatiotemporal expression pattern of DHNs during dehydration stress. The expression analysis exhibits the importance of OsjDHN1, OsjDHN4 and OsjDHN7 in imparting PEG induced dehydration tolerance. Interestingly, OsjDHN1 and OsjDHN4 are among 3 DHNs that were found to be conserved on the basis of their chromosomal localization, protein length and domain architecture. Also, the OsjDHN1 expression pattern was similar in both tissues. Similar results were reported by Kumar et al. [9], stating that OsDhn1-overexpression in transgenic rice plants (OsDhn1-OXs) shows enhanced drought and salt stress tolerance. The dehydration tolerance could be attributed to presence of abiotic stress related cis-regulatory elements, especially ABRE and DRE motifs. Due to the presence of ABRE motif, it could be concluded that all OsjDHNs (except OsjDHN1) expression is probably regulated by ABA dependent signalling pathways. Various other hormone related cis elements like CGTCA and TCA elements were found and they are in corroboration with our gene expression analysis. The gene interactions of OsjDHN2 involve OsjDHN3, OsjDHN6, and OsjDHN7; thus, it could be possible that these three genes require OsjDHN2 as a signal for their response. The following network is also reflected by the expression profiles of PEG-treated, roots and shoots, where the expression pattern of OsjDHN3, OsjDHN6, and OsjDHN7 is similar to that of OsjDHN2. On the other hand, OsjDHN4 requires the presence of LEA genes in their interactional network. The expression profile of OsjDHN6 also indicated their role in signalling the expression of OsjDHN7, which further interacts with LEA proteins and illustrates a specific pattern in the shoot expression profile in PEG-treated plants. OsjDHN6, which was highly expressed during PEG treatment, showed an interaction with HSPs and cold shock-resistant genes (AWPM-19), also indicating its specific role in combatting dehydration caused due to low temperature stress.
Conclusion
In summary, a total 65 DHNs were identified in Oryza, 9 species and 2 sub-species. This study provides an insight into the evolutionary association of DHNs. The phylogeny, domain detection, and MSA exhibited useful information about conserved DHNs, indicative of their functional conservation in varied rice species. Structural analysis and evolutionary pattern supported the independent origin of japonica and indica rice varieties. We also focused on the response patterns of DHNs genes in drought conditions, and their interaction network revealed their mode of action and their signalling properties. Our result advocates DHNs as conserved proteins during the course of domestication/evolution of rice, and they play an important role in combating dehydration stress. Also, the results provide a base for further functional and evolutionary studies of the DHN gene family in rice and other plants.
Supporting information
(TIF)
The alignment was performed using ClustalX. The conserved K Y and S segments are highlighted.
(TIF)
(TIF)
Each red line indicates one duplication event.
(TIF)
(TIF)
(DOC)
(DOC)
(DOC)
Acknowledgments
Authors sincerely thank Dr. K. N. Nair for his role in extensive correction of the manuscript. This research was supported by the Science and Engineering Research Board, New Delhi, India (project file no. SB/YS/LS-74/2014) and also by the network project (BSC-204) funded by the Council of Scientific and Industrial Research, New Delhi. The authors are also thankful to Director, CSIR-National Botanical Research Institute, Lucknow for the facilities and the financial support. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Data Availability
All relevant data are within the paper and its Supporting Information files.
Funding Statement
This research was supported by the Science and Engineering Research Board, New Delhi, India (project file no. SB/YS/LS-74/2014) and also by the network project (BSC-204) funded by the Council of Scientific and Industrial Research, New Delhi. The authors are also thankful to Director, CSIR-National Botanical Research Institute, Lucknow, for the facilities and the financial support. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1.Bailey-Serres J, Lee SC, Brinton E. Waterproofing crops: effective flooding survival strategies. Plant Physiol. 2012;160: 1698–1709. 10.1104/pp.112.208173 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Yang YZ, He MY, Zhu ZG, Li SX, Xu Y, Zhang CH, et al. Identification of the dehydrin gene family from grapevine species and analysis of their responsiveness to various forms of abiotic and biotic stress. Bmc Plant Biol. 2012;12: 140 10.1186/1471-2229-12-140 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Battaglia M, Olvera-Carrillo Y, Garciarrubio A, Campos F, Covarrubias AA. The enigmatic LEA proteins and other hydrophilins. Plant Physiol. 2008;148: 6–24. 10.1104/pp.108.120725 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Wang XS, Zhu HB, Jin GL, Liu HL, Wu WR, Zhu J. Genome-scale identification and analysis of LEA genes in rice (Oryza sativa L.). Plant Sci. 2007;172: 414–420. [Google Scholar]
- 5.Graether SP, Boddington KF. Disorder and function: a review of the dehydrin protein family. Front Plant Sci. 2014;5: 576 10.3389/fpls.2014.00576 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Yin Z, Rorat T, Szabala BM, Ziółkowska A, Malepszy S. Expression of a Solanum sogarandinum SK 3-type dehydrin enhances cold tolerance in transgenic cucumber seedlings. Plant Sci. 2006;170: 1164–1172. [Google Scholar]
- 7.Peng Y, Reyes JL, Wei H, Yang Y, Karlson D, Covarrubias AA, et al. RcDhn5, a cold acclimation-responsive dehydrin from Rhododendron catawbiense rescues enzyme activity from dehydration effects in vitro and enhances freezing tolerance in RcDhn5-overexpressing Arabidopsis plants. Physiol Plant. 2008;134: 583–597. 10.1111/j.1399-3054.2008.01164.x [DOI] [PubMed] [Google Scholar]
- 8.Shekhawat UKS, Srinivas L, Ganapathi TR. MusaDHN-1, a novel multiple stress-inducible SK3-type dehydrin gene, contributes affirmatively to drought-and salt-stress tolerance in banana. Planta. 2011;234: 915 10.1007/s00425-011-1455-3 [DOI] [PubMed] [Google Scholar]
- 9.Kumar M, Lee SC, Kim JY, Kim SJ, Aye SS, Kim SR. Over-expression of dehydrin gene, OsDhn1, improves drought and salt stress tolerance through scavenging of reactive oxygen species in rice (Oryza sativa L.). Journal of Plant Biol. 2014;57: 383–393. [Google Scholar]
- 10.Liu H, Yu C, Li H, Ouyang B, Wang T, Zhang J, et al. Overexpression of ShDHN, a dehydrin gene from Solanum habrochaites enhances tolerance to multiple abiotic stresses in tomato. Plant Sci. 2015;231: 198–211. 10.1016/j.plantsci.2014.12.006 [DOI] [PubMed] [Google Scholar]
- 11.Hara M, Terashima S, Fukaya T, Kuboi T. Enhancement of cold tolerance and inhibition of lipid peroxidation by citrus dehydrin in transgenic tobacco. Planta. 2003;217: 290–298. 10.1007/s00425-003-0986-7 [DOI] [PubMed] [Google Scholar]
- 12.Xing X, Liu Y, Kong X, Liu Y, Li D. Overexpression of a maize dehydrin gene, ZmDHN2b, in tobacco enhances tolerance to low temperature. Plant Growth Regul. 2011;65: 109–118. [Google Scholar]
- 13.Puhakainen T, Hess MW, Mäkelä P, Svensson J, Heino P, Palva ET. Overexpression of multiple dehydrin genes enhances tolerance to freezing stress in Arabidopsis. Plant Mol Biol. 2004;54: 743–753. 10.1023/B:PLAN.0000040903.66496.a4 [DOI] [PubMed] [Google Scholar]
- 14.Brini F, Hanin M, Lumbreras V, Amara I, Khoudi H, Hassairi A, et al. Overexpression of wheat dehydrin DHN-5 enhances tolerance to salt and osmotic stress in Arabidopsis thaliana. Plant Cell Rep. 2007;26: 2017–2026. 10.1007/s00299-007-0412-x [DOI] [PubMed] [Google Scholar]
- 15.Xu J, Zhang YX, Wei W, Han L, Guan ZQ, Wang Z, et al. BjDHNs confer heavy-metal tolerance in plants. Mol Biotechnol. 2008;38: 91–98. 10.1007/s12033-007-9005-8 [DOI] [PubMed] [Google Scholar]
- 16.Bouman BAM, Peng S, Castaneda AR, Visperas RM. Yield and water use of irrigated tropical aerobic rice, systems. Agric Water Manag. 2005;74: 87–105. [Google Scholar]
- 17.Mostajeran A, Rahimi-Eichi V Effects of drought stress on growth and yield of rice (Oryza sativa L.) cultivars and accumulation of proline and soluble sugars in sheath and blades of their different ages leaves. American-Eurasian J. Agric. & Environ. Sci. 2009;5: 264–272. [Google Scholar]
- 18.Sang T, Ge S. Understanding rice domestication and implications for cultivar improvement. Current Opin Plant Biol. 2013;16: 139–146. [DOI] [PubMed] [Google Scholar]
- 19.Vaughan DA, Morishima H, Kadowaki K. Diversity in the Oryza genus. Curr Opin Plant Biol. 2003;6: 139–146. [DOI] [PubMed] [Google Scholar]
- 20.Vaughan D, Kadowaki K, Kaga A, Tomooka N. Ecogentic diversification in the genus Oryza: implications for sustainable rice production. In:Toriyama K, Heong KL, Hardy B, editors. Rice is Life Scientific Perspectives for the 21st Century: Proceedings of the World Rice Research Conference, Tsukuba, Japan. 2005; pp. 44–46.
- 21.Ge S, Sang T, Lu BR, Hong DY. Phylogeny of rice genomes with emphasis on origins of allotetraploid species. Proc. Natl. Acad. Sci. U.S.A. 1999;96: 14400–14405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Vaughan DA, Lu BR, Tomooka N. The evolving story of rice evolution. Plant Sci. 2008;174: 394–408. [Google Scholar]
- 23.Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215: 403–410. 10.1016/S0022-2836(05)80360-2 [DOI] [PubMed] [Google Scholar]
- 24.Finn RD, Clements J, Eddy SR. HMMER web server: interactive sequence similarity searching. Nucleic Acids Res. 2011;gkr367. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, et al. Clustal W and Clustal X version 2.0. Bioinformatics. 2007;23: 2947–2948. 10.1093/bioinformatics/btm404 [DOI] [PubMed] [Google Scholar]
- 26.Kumar S, Stecher G, Tamura K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol Biol Evol. 2016; msw054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D, et al. Circos: an information aesthetic for comparative genomics. Genome Res. 2009;19:1639–1645. 10.1101/gr.092759.109 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Wang Y, Tang H, DeBarry JD, Tan X, Li J, Wang X, et al. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012;407: e49. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Guo AY, Zhu QH, Chen X, Luo JC GSDS: a gene structure display server. Yi Chuan. 2007;29(8):1023–1026. [PubMed] [Google Scholar]
- 30.Sato Y, Takehisa H, Kamatsuki K, Minami H, Namiki N, Ikawa H, et al. RiceXPro version 3.0: expanding the informatics resource for rice transcriptome. Nucleic Acids Res. 2012;gks1125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Wu SB, Rodgers N, Choct M. Real-Time PCR Assay for Clostridium perfringens in Broiler Chickens in a Challenge Model of Necrotic Enteritis. Appl Environ Microbiol. 2011;77: 1135–1139. 10.1128/AEM.01803-10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Mccouch SR, Cooperative RG. Gene Nomenclature System for Rice. Rice. 2008;1: 72–84. [Google Scholar]
- 33.Hanin M, Brini F, Ebel C, Toda Y, Takeda S, Masmoudi K. Plant dehydrins and stress tolerance: versatile proteins for complex mechanisms. Plant Signal Behav 2011;6: 1503–1509. 10.4161/psb.6.10.17088 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Koag MC, Wilkens S, Fenton RD, Resnik J, Vo E, Close TJ. The K-segment of maize DHN1 mediates binding to anionic phospholipid vesicles and concomitant structural changes. Plant Physiol. 2009;150: 1503–1514. 10.1104/pp.109.136697 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Yamaguchi-Shinozaki K, Shinozaki K. Organization of cis-acting regulatory elements in osmotic- and cold-stress-responsive promoters. Trends in Plant Sci. 2005;10: 88–94. [DOI] [PubMed] [Google Scholar]
- 36.Yue R, Lu C, Sun T, Peng T, Han X, Qi J, et al. Identification and expression profiling analysis of calmodulin-binding transcription activator genes in maize (Zea mays L.) under abiotic and biotic stresses. Front Plant Sci. 2015;6: 576 10.3389/fpls.2015.00576 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Passricha N, Saifi S, Ansari MW, Tuteja N. (2016) Prediction and validation of cis-regulatory elements in 5′ upstream regulatory regions of lectin receptor-like kinase gene family in rice. Protoplasma. 2016;1: 16. [DOI] [PubMed] [Google Scholar]
- 38.Close TJ. Dehydrins: a commonalty in the response of plants to dehydration and low temperature. Physiol Plant. 1997;100: 291–296. [Google Scholar]
- 39.Perdiguero P, Collada C, Soto Á. Novel dehydrins lacking complete K-segments in Pinaceae. The exception rather than the rule. Front Plant Sci. 2014;5: 682 10.3389/fpls.2014.00682 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Zhang Y, Li J, Yu F, Cong L, Wang L, Burkard G, et al. Cloning and expression analysis of SKn-type dehydrin gene from bean in response to heavy metals. Mol Biotechnol. 2006;32: 205–218. 10.1385/MB:32:3:205 [DOI] [PubMed] [Google Scholar]
- 41.Wang M, Yu Y, Haberer G, Marri PR, Fan C, Goicoechea JL, et al. The genome sequence of African rice (Oryza glaberrima) and evidence for independent domestication. Nature Genet. 2014;46: 982–988. 10.1038/ng.3044 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Flagel LE, Wendel JF. Gene duplication and evolutionary novelty in plants. New Phytol. 2009;183: 557–564. 10.1111/j.1469-8137.2009.02923.x [DOI] [PubMed] [Google Scholar]
- 43.Hundertmark M, Hincha DK. LEA (late embryogenesis abundant) proteins and their encoding genes in Arabidopsis thaliana. BMC genomics. 2008;9: 1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Liu CC, Li CM, Liu BG, Ge SJ, Dong XM, Li W, et al. Genome-wide identification and characterization of a dehydrin gene family in poplar (Populus trichocarpa). Plant Mol Biol Report. 2012;30: 848–859. [Google Scholar]
- 45.Du H, Liu H, Xiong L. Endogenous auxin and jasmonic acid levels are differentially modulated by abiotic stresses in rice. Front Plant Sci. 2013;4: 397 10.3389/fpls.2013.00397 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Liu J, Luo M, Yan X, Yu C, Li S. Characterization of genes coding for galacturonosyltransferase-like (GATL) proteins in rice. Genes Genom. 2016;1: 13. [Google Scholar]
- 47.Zhou J, Wang X, Jiao Y, Qin Y, Liu X, He K, et al. Global genome expression analysis of rice in response to drought and high-salinity stresses in shoot, flag leaf, and panicle. Plant Mol Biol. 2007;63: 591–608. 10.1007/s11103-006-9111-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Gsponer J, Futschik ME, Teichmann SA, Babu MM. Tight Regulation of Unstructured Proteins: From Transcript Synthesis to Protein Degradation. Science. 2008;322: 1365–1368. 10.1126/science.1163581 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Mouillon JM, Eriksson SK, Harryson P. Mimicking the Plant Cell Interior under Water Stress by Macromolecular Crowding: Disordered Dehydrin Proteins Are Highly Resistant to Structural Collapse. Plant Physiol. 2008;148: 1925–1937. 10.1104/pp.108.124099 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Nylander M, Svensson J, Palva ET, Welin BV. Stress-induced accumulation and tissue-specific localization of dehydrins in Arabidopsis thaliana. Plant Mol Biol. 2001;45: 263–279. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
(TIF)
The alignment was performed using ClustalX. The conserved K Y and S segments are highlighted.
(TIF)
(TIF)
Each red line indicates one duplication event.
(TIF)
(TIF)
(DOC)
(DOC)
(DOC)
Data Availability Statement
All relevant data are within the paper and its Supporting Information files.