Abstract
The mRNA encoding Nor-1/NR4A3 is rapidly and strikingly induced by β2-adrenergic signaling in glycolytic and oxidative skeletal muscle. In skeletal muscle cells, Nor-1 expression is important for the regulation of oxidative metabolism. Transgenic skeletal muscle-specific expression of activated Nor-1 resulted in the acquisition of an endurance phenotype, an increase in type IIA/X oxidative muscle fibers, and increased numbers of mitochondria. In the current study, we used dual-energy x-ray absorptiometry and magnetic resonance imaging analysis to demonstrate decreased adiposity in transgenic (Tg) Nor-1 mice relative to that in wild-type littermates. Furthermore, the Tg-Nor-1 mice were resistant to diet-induced weight gain and maintained fasting glucose at normoglycemic levels. Expression profiling and RT-quantitative PCR analysis revealed significant increases in genes involved in glycolysis, the tricarboxylic acid cycle, oxidative phosphorylation, fatty acid oxidation, and glycogen synthesis, in concordance with the lean phenotype. Moreover, expression profiling identified several Z-disc and sarcomeric binding proteins that modulate fiber type phenotype and endurance, eg, α-actinin-3. In addition, we demonstrated that the Tg-Nor-1 mouse line has significantly higher glycogen content in skeletal muscle relative to that in wild-type littermates. Finally, we identified a decreased NAD+/NADH ratio with a concordant increase in peroxisome proliferator-activated receptor γ coactivator-1α1 protein/mRNA expression. Increased NADH was associated with an induction of the genes involved in the malate-aspartate shuttle and a decrease in the glycerol 3-phosphate shuttle, which maximizes aerobic ATP production. In conclusion, skeletal muscle-specific Nor-1 expression regulates genes and pathways that regulate adiposity, muscle fiber type metabolic capacity, and endurance.
In the context of whole-body metabolism, skeletal muscle accounts for up to 40% of the total body mass and energy expenditure and is the predominant site for fatty acid oxidation and glycolysis (1, 2). Skeletal muscle uses complex networks of metabolic machinery to regulate (1) energy homeostasis and (2) anaerobic and oxidative substrate utilization in response to environmental loads and signals. Consequently, the regulation of skeletal muscle metabolic networks can have a significant impact on many physiological outcomes such as the regulation of body composition, energy balance, insulin sensitivity, and the blood lipid profile.
Nuclear receptors (NRs) in the nuclear hormone receptor superfamily are structurally related transcription factors that bind DNA and translate endocrine, metabolic, and pathophysiological signals into gene regulation. Many NRs have well-defined roles in the regulation of biological networks within skeletal muscle, in particular metabolism, and this has been extended to 2 members of the orphan nuclear receptor 4A (NR4A) subgroup in recent years (3–6). The NR4A subgroup consists of 3 closely related orphan NRs: neuron-derived clone 77 (Nur77/NR4A1), nuclear receptor related 1 (Nurr1/NR4A2), and neuron-derived orphan receptor 1 (Nor-1/NR4A3). The expression of NR4A receptors is highly inducible by a range of metabolites, metabolic signaling molecules, and environmental stimuli, ie, β-adrenoceptor agonists (7–9), cold (10, 11), fatty acids (12), glucose (13), insulin (14), cholesterol (15), melanocortins (16), and thiazolidinediones (17). No bona fide endogenous or native Nor-1 ligands have been identified, and thus it is designated as an “orphan” NR; however, several agonists that modulate the activity of this NR have been described (18, 19). Gain- and loss-of-function studies (in vitro and in vivo) in metabolic tissues have associated Nor-1 and Nur77 with specific aspects of lipid, carbohydrate, and energy homeostasis (3–6, 8, 9, 20).
Nur77-null mice exhibited limited metabolic changes when fed a normal diet but displayed an impaired capacity to adapt to the metabolic challenges presented by high-fat feeding. These mice displayed increased weight gain, decreased energy usage, insulin resistance in skeletal muscle, and impaired blood glucose clearance (4). Previous studies have also demonstrated that Nor-1 expression is important for the regulation of oxidative metabolism in vitro (7). Targeted silencing of Nor-1 in skeletal muscle cells resulted in increased lactate production and decreased fatty acid oxidation associated with the attenuation of gene expression involved in the aerobic metabolism of carbohydrate and fat metabolism (7). To investigate the role of this NR in vivo, we developed a transgenic mouse model with skeletal muscle–specific expression of activated Nor-1 (3). The expression of activated Nor-1 in skeletal muscle resulted in a striking transition toward a more oxidative phenotype with increased myoglobin expression, mitochondrial DNA/number, oxidative enzyme staining, and oxidative myosin heavy chain IIA/IIX expression. Furthermore, we observed systemic changes associated with an improved oxidative capacity in the skeletal muscle such as increased oxygen consumption and enhanced endurance (but reduced sprint performance). Interestingly, compared with wild-type (WT) mice, the transgenic (Tg) Nor-1 mice exhibited increased GLUT4 protein expression and improved glucose clearance. Recently, selective overexpression of Nur77 in skeletal muscle enhanced mitochondrial number and function, with accordant increases in oxidative metabolism in transgenic mice (5). Furthermore, decreased adiposity has been demonstrated in other transgenic models that also display increased oxygen consumption and enhanced mitochondrial function in skeletal muscle (21–23). These converging lines of evidence led us to conduct a comprehensive study to investigate the relationship between Nor-1-dependent changes in skeletal muscle gene expression and (the dynamics of) energy utilization and storage in both muscle and adipose depots.
In the present study, we revealed a significant reduction in adiposity on normal chow and high-fat diet in the Tg-Nor-1 mice (relative to that in WT littermates). This occurs in the context of normal activity levels and food intake. The fasting blood glucose levels of the transgenic mice are significantly lower than those of WT mice fed a high-fat diet, commensurate with resistance to obesity, although this protection does not extend to glucose tolerance, which isc similarly impaired in the Tg-Nor-1 mice. We employ extensive gene expression profiling and pathway analysis to identify gene changes in glycolysis, the tricarboxylic acid (TCA) cycle, fatty acid oxidation, oxidative phosphorylation, and glycogen deposition. We also identified changes in the expression of numerous genes encoding Z-disc and sarcomeric binding proteins that may be involved in fiber type reprogramming and the endurance phenotype of the Tg-Nor-1 mice. A dramatic increase in skeletal muscle glycogen content is described, demonstrating that in addition to increased oxidative metabolism, surplus glucose is being stored in the skeletal muscle. Finally, we report a decreased NAD+/NADH ratio, an activation of the malate-aspartate shuttle (relative to the glycerol 3-phosphate shuttle) that maximizes the production of aerobic ATP, and elevated peroxisome proliferator-activated receptor γ coactivator-1α1 (PGC-1α1) mRNA and protein expression in skeletal muscle of Tg-Nor-1 mice.
Results
Transgenic skeletal muscle–specific activated Nor-1 expression leads to decreased adiposity
Our previous study investigated the function of Nor-1 by the targeted skeletal muscle-specific expression of an activated form of Nor-1 in transgenic mice. This induced an endurance phenotype associated with increased (1) myoglobin expression, (2) mitochondrial density, and (3) expression of key mitochondrial mRNA/proteins. Moreover, we observed a transition from glycolytic type IIB to oxidative type IIA and IIX skeletal muscle fibers. The fatigue-resistant phenotype is associated with decreased sprint performance (3).
Although the Tg-Nor-1 mice showed no significant difference in body weight when fed a normal chow diet, the body weight of Tg-Nor-1 mice trends below that of WT mice (3). Therefore, we were particularly interested in investigating whether this phenotype was associated with changes in whole-animal adiposity. We examined the white subcutaneous (inguinal) and visceral (epididymal) fat pad depots and the interscapular brown adipose tissue from normal chow-fed Tg-Nor-1 mice relative to those of WT littermates. Tissue weights (as a percentage of total body weight) indicated significant decreases in the weights of the white and brown adipose depots from fasted Tg-Nor-1 mice relative to those of WT littermates (Figure 1A) and to those of other nonadipose organs such as liver. The decrease in adiposity was further highlighted by the frontal view after dissection (Figure 1B) and the direct side-by-side comparison of isolated fat depots (Figure 1C). To extend these observations on individual fat pads, we performed DXA to measure “whole-body” compositions of mice unaffected by fasting (Figure 1D). The DXA analysis indicated that (fed) Tg-Nor-1 mice displayed a significant reduction in total fat mass (Figure 1D) relative to that of WT littermates (P < .001). This was against a background of no significant changes in total body weight and lean mass or bone area (Figure 1D). Moreover, these observations were confirmed by MRI, in which the differences in adiposity between the normal chow-fed transgenic and WT mice are clearly and statistically substantiated (Figure 1, E and F). The decrease in adiposity is associated with decreased adipocyte cell size (Figure 1G) and significantly reduced fasting plasma leptin (Figure 1H). In summary, the Tg-Nor-1 mice display a lean phenotype. The trend toward lower body weight is associated with decreased white and brown adipose depots and not reduced body frame size.
Figure 1.
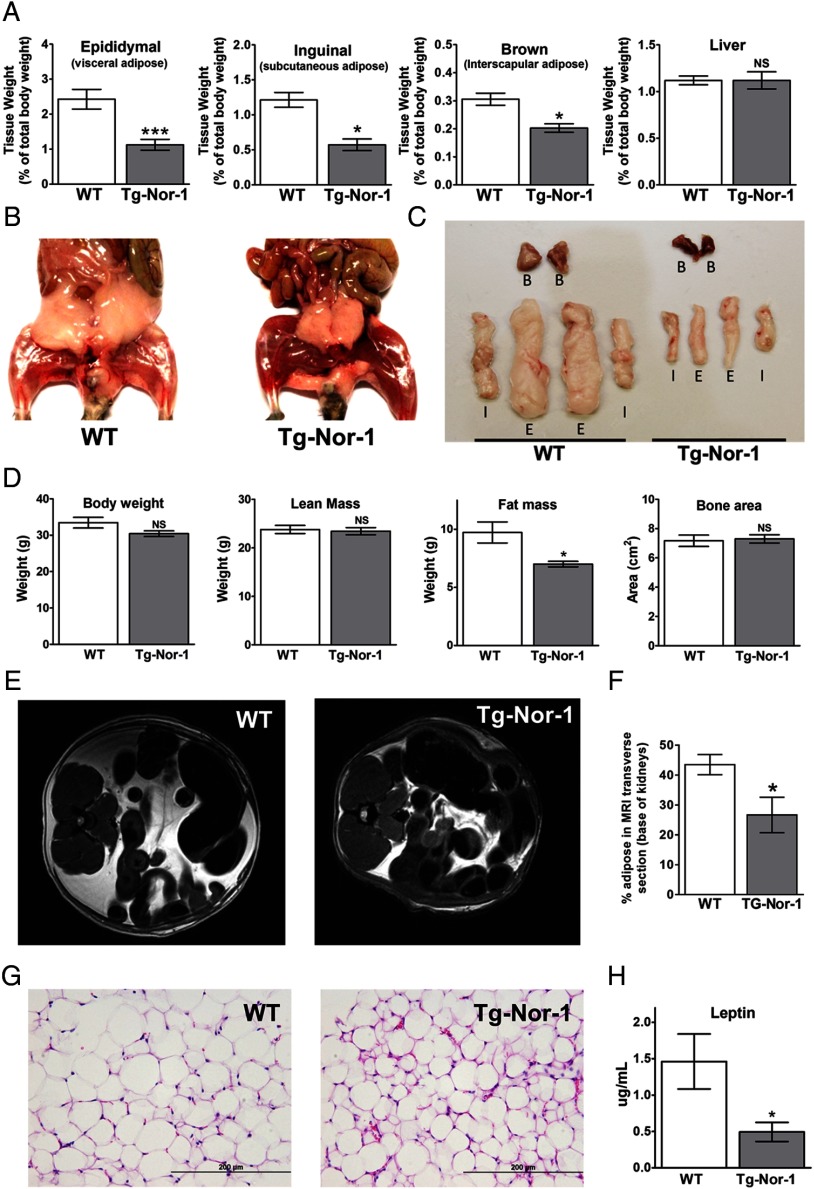
Transgenic skeletal muscle-specific Nor-1 expression leads to decreased adiposity. A, Tissue weights (as a percentage of total body weight) were determined for epididymal adipose tissue, inguinal adipose tissue, brown adipose tissue, and liver for WT and Tg-Nor-1 mice (n = 8–11). B, Photographs of representative WT and Tg-Nor-1 mice displaying decreased adiposity. C, Samples of isolated adipose tissue (B. brown; I, inguinal; E, epididymal) from WT and Tg-Nor-1 mice. D, DXA body composition analysis showing body weight, lean mass, fat mass, and bone area (n = 4). E, MRI scans displaying a transverse section at the base of the kidney with (F) percentage of the total area of the slice occupied by adipose (n = 4). G, Representative hematoxylin and eosin–stained epididymal adipose tissue sections. H, plasma leptin concentration (n = 8–11). Statistical calculation was performed using a Student t test, and P values are indicated on graphs as follows: *, P < .05; **, P < .01; ***, P < .001.
Metabolic profiling indicated reduced fat deposition with a normal chow diet is associated with an increased respiratory exchange ratio (RER) during feeding
We have previously used metabolic profiling to demonstrate that the Tg-Nor-1 mice display increased oxygen consumption and energy expenditure with a normal chow diet (3); however, we had not analyzed the relative food consumption, activity, and respiratory exchange ratio in the Tg-Nor-1 mice (relative to those of WT littermates). Mice were acclimated for 24 hours in the Columbus Instruments Comprehensive Lab Animal Monitoring System (CLAMS), before comprehensive monitoring and measurement over the next 24-hour night/day cycle followed by an evening fasting cycle. We demonstrate that Tg-Nor-1 mice have increased carbon dioxide production (Figure 2A). The analysis also identified a significant increase in the RER during the nighttime feeding cycle in the Tg-Nor-1 mice fed the normal chow diet (Figure 2B). This occurred in a background of no significant changes in food consumption and total activity between the Tg-Nor-1 mice and WT littermates (Figure 2, C and D). The increased RER during nighttime feeding is suggestive of enhanced carbohydrate oxidation or lipolysis during this period.
Figure 2.
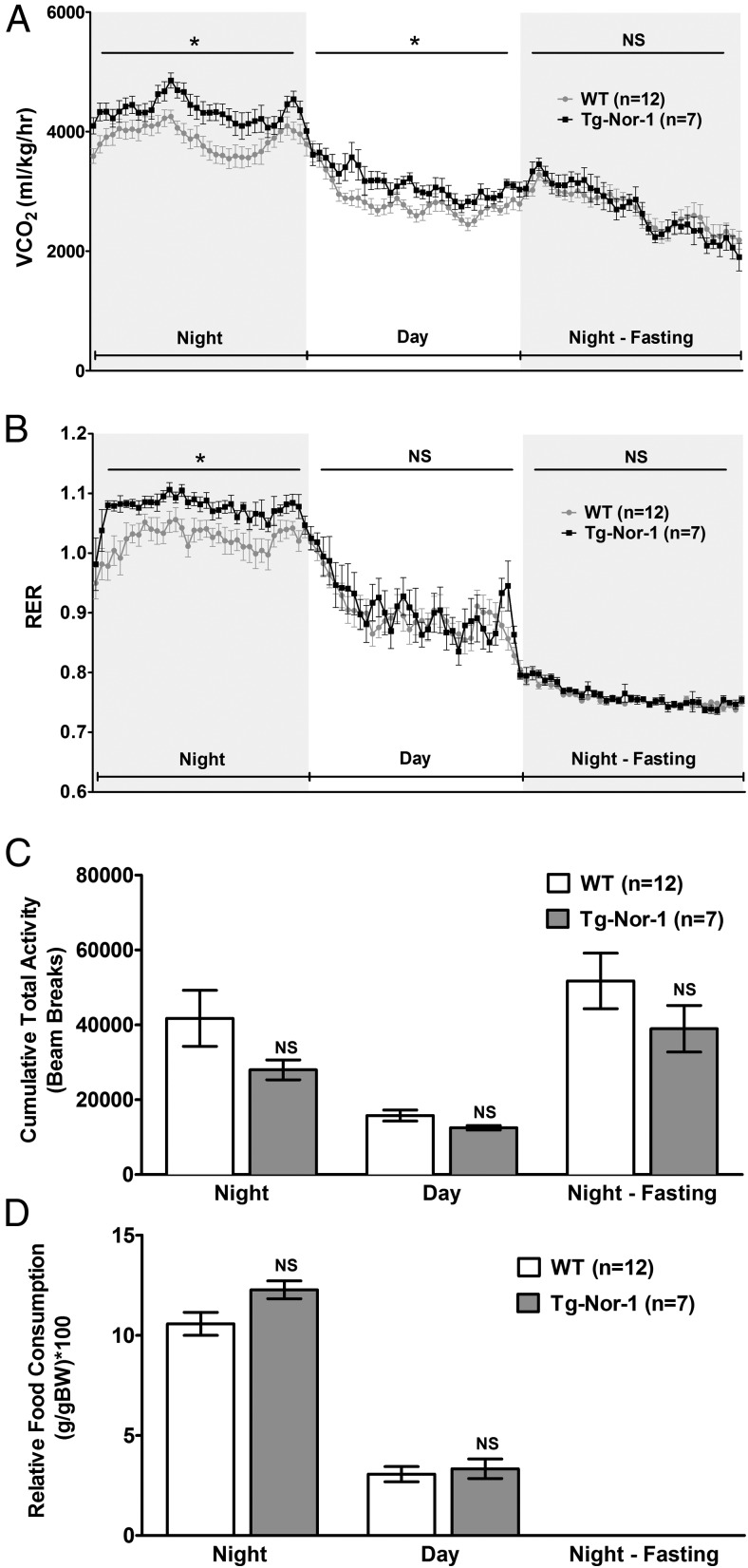
Metabolic profiling of Tg-Nor-1 mice. A and B, CO2 production (corrected for body weight) (A) and RER in WT and Tg-Nor-1 mice (n = 7–12) (B). Data were analyzed using repeated-measures two-way ANOVA. C, Cumulative total activity of the mice measured by horizontal beam breaks. D, Relative food consumption (corrected for body weight) analyzed by one-way ANOVA with column selected Bonferroni post hoc analysis. *, P < .05.
Tg-Nor-1 mice are resistant to high-fat diet-induced weight gain and hepatic triglyceride accumulation
We subsequently investigated the physiological response of Tg-Nor-1 mice to a high-fat diet. WT and Tg-Nor-1 mice were fed a high-fat diet (containing 36% fat) for a 12-week period at weaning (with digestible energy of 19.9 and 22.8 MJ/kg, respectively). In our previous study with a normal chow diet (3), no significant differences in the weight of Tg-Nor-1 mice, relative to that of WT littermates were observed over a 12-week period with a regular chow diet (<10% of total calories from fat). In contrast, with a high-fat diet (∼60% of energy from fat), the WT mice experienced statistically significant increases of ∼15% to 20% in weight, relative to that of the Tg-Nor-1 mice after 12 weeks of consuming a high-fat diet (Figure 3A). This was also evident from visually larger WT mice (Figure 3B), and this effect was independent of body length (Figure 3C).
Figure 3.
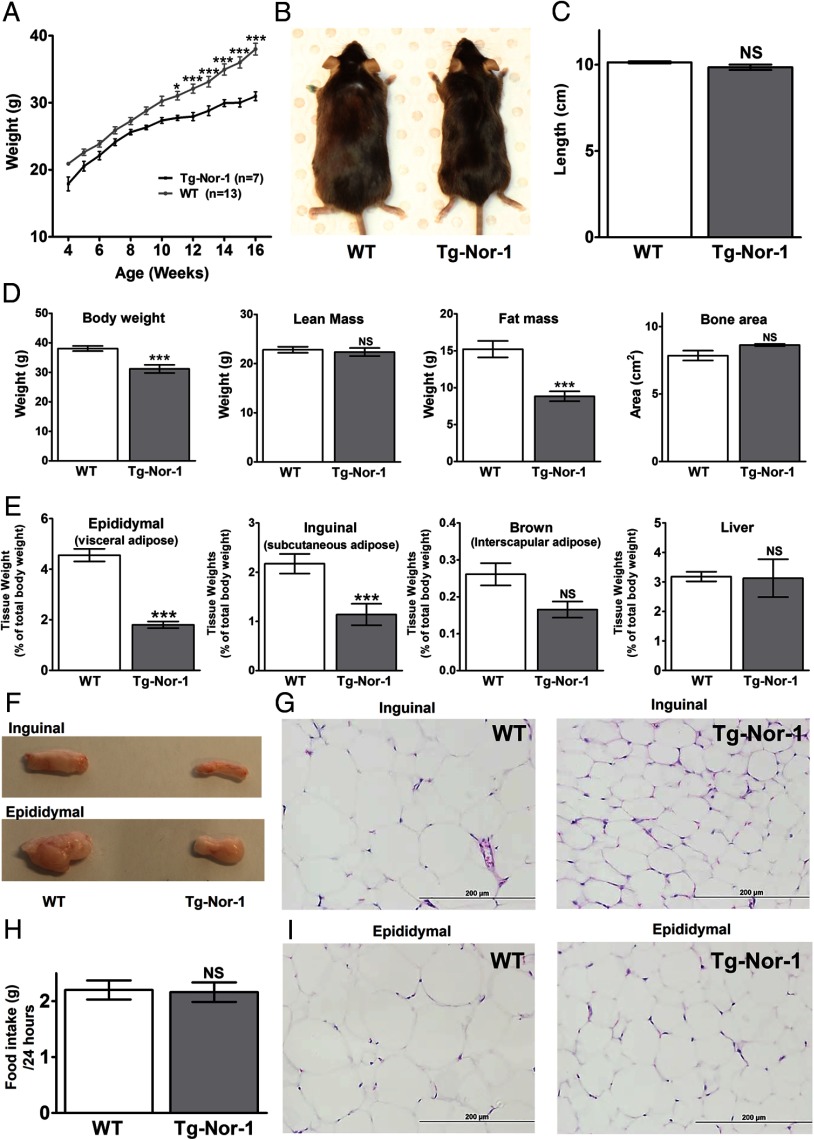
Skeletal muscle–specific Tg-Nor-1 mice are protected against high-fat diet–induced weight gain. A, Growth curve measured over 12 weeks with the high-fat diet (n = 7–13) analyzed by one-way ANOVA with column selected Bonferroni post hoc analysis. B, Representative image of 17-week-old WT and Tg-Nor-1 mice. C, Length of mice measured at 17 weeks (n = 6–8). D, DXA body composition analysis showing body weight, lean mass, fat mass, and bone area. E, Tissue weights (corrected for total body weight) were determined for epididymal adipose tissue, inguinal adipose tissue, brown adipose tissue, and liver for WT and Tg-Nor-1 mice (n = 6–8). F, Photographs showing representative samples of inguinal and epididymal adipose tissue. Hematoxylin and eosin stained inguinal (G) and epididymal (I) adipose tissue sections. H, Total high-fat diet consumption over a 24-hour period (n = 5–8). Statistical calculation was performed using a Student t test, and P values are indicated on graphs as follows: *, P < .05; **, P < .01; ***, P < .001; NS, not significant.
We subsequently used DXA for body composition analysis (Figure 3D). The DXA analysis identified that (fed) Tg-Nor-1 mice displayed a significant reduction in total body weight and fat mass relative to that of WT littermates (P < .001). This was against a background of no significant differences in lean mass or bone area (Figure 3D). This finding was underscored by significantly decreased amounts of epididymal (visceral) and inguinal (subcutaneous) white adipose tissue in the Tg-Nor-1 mice (Figure 3E) relative to those of WT male mice (fed the high-fat diet) (P < .001). In contrast, no significant differences in corrected brown adipose and liver tissue weights (from Tg-Nor-1 and WT mice) were observed (Figure 3E) with the high-fat diet.
Direct side-by-side comparison of isolated fat depots shows the size difference (Figure 3F). Furthermore, the decrease in adiposity with a high-fat diet is associated with decreased adipocyte cell size in inguinal (Figure 3G) and epididymal (Figure 3I) adipose depots. These changes are independent of food intake as measured over a 24-hour period (Figure 3H).
Analysis of the liver triglycerides from fasted animals fed the high-fat diet revealed decreased amounts in the Tg-Nor-1 mice (Supplemental Figure 1A published on The Endocrine Society's Journals Online web site at http://mend.endojournals.org.). In concordance, the hematoxylin and eosin staining of the fasted livers from the high-fat diet study revealed decreased numbers of lipid droplets in the Tg-Nor-1 mice, consistent with decreased triglycerides in the livers from the Tg-Nor-1 animals fed the high-fat diet (Supplemental Figure 1B).
We examined whether the lack of fat accumulation in Tg-Nor-1 mice (fed the high-fat diet) relative to that of WT littermates involved changes in oxygen consumption, energy expenditure, and RER. The Tg-Nor-1 mice and WT littermates were acclimated in the CLAMS. The mice were comprehensively monitored over the next 24-hour night/day cycle followed by an evening fasting period. The indirect calorimetry studies revealed no significant differences between the Tg-Nor-1 mice and the WT littermates in the context of oxygen consumption (Figure 4A), carbon dioxide production (Figure 4B), and RER (Figure 4C) with the high-fat diet; however, in all cases, the Tg-Nor-1 mice trended higher for both oxygen consumption and carbon dioxide production.
Figure 4.
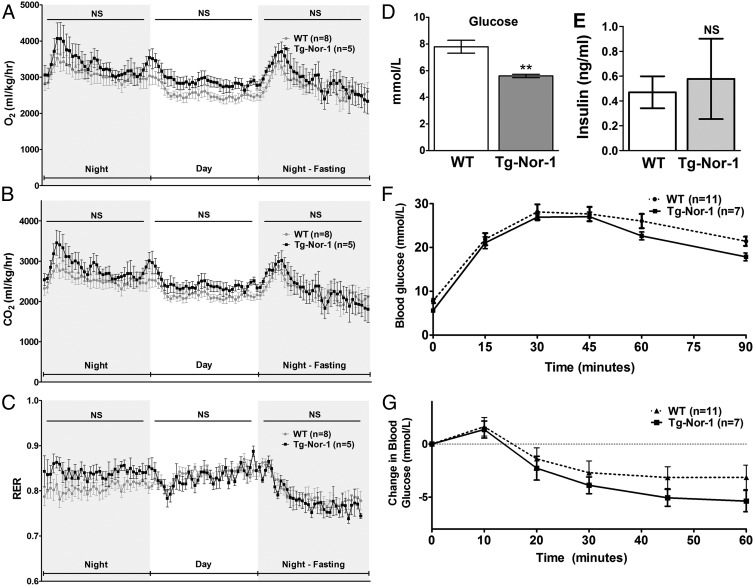
Gas exchange and glucose tolerance with the high-fat diet. A and B, Oxygen consumption (A) and CO2 production (B) relative to body weight. C, RER for WT and Tg-Nor-1 mice fed the high-fat diet (n = 5–8). Data were analyzed using repeated-measures two-way ANOVA. D and E, Blood glucose levels (D) and plasma insulin levels (E) after a 12-hour fast. Statistical calculation was performed using a Student t test. **, P < .01; NS, not significant. F and G, Intraperitoneal glucose tolerance test (F) and intraperitoneal insulin tolerance test (G) performed on high-fat diet–fed WT and Tg-Nor-1 mice. Data were analyzed as nonsignificant by one-way ANOVA with column selected Bonferroni post hoc analysis.
In summary, the decreased adiposity in the Tg-Nor-1 mice is maintained with the high-fat diet and the distinct and relative lack of fat accumulation in Tg-Nor-1 mice (fed the high-fat diet) relative to that in WT littermates did not involve underlying changes in oxygen consumption, energy expenditure, RER, feeding behavior, and food intake.
Muscle-specific Tg-Nor-1 mice maintain fasting blood glucose at normoglycemic levels with a high-fat diet
We have previously demonstrated that the skeletal muscle-specific Tg-Nor-1 mice display normal fasting glucose and significantly improved glucose tolerance with a normal chow diet. In addition, insulin tolerance tests demonstrated a trend toward increased insulin sensitivity, but this did not attain significance (3). We have subsequently performed pyruvate tolerance tests in mice fed the normal chow diet and observed that the Tg-Nor-1 mice displayed normal glucose production relative to that of WT littermates (Supplemental Figure 2A), indicative of normal liver glucose output and intact insulin signaling (with the normal chow diet). In concordance, we observed no significant differences in AKT and pAKT (serine 473 and threonine 308) expression in liver from WT and Tg-Nor-1 littermates fed the normal chow diet (Supplemental Figure 2B)
We subsequently examined WT and Tg-Nor-1 mice fed the high-fat diet until 16 to 17 weeks. We observed normal blood glucose levels in the Tg-Nor-1 mice after a 12-hour fast (Figure 4D); in contrast, the WT mice were hyperglycemic. However, there were no significant differences in (12-hour) fasting plasma insulin levels in the WT and Tg-Nor-1 mice (Figure 4E). We further examined systemic glucose metabolism and performed ip glucose tolerance tests. Both WT and Tg-Nor-1 mice (after 12 weeks of the high-fat diet) displayed impaired glucose clearance (ie, glucose intolerance) after a glucose challenge (Figure 4F). The WT and Tg-Nor-1 mice both displayed compromised responsiveness to the administration of ip insulin (indicating insulin insensitivity, ie, impaired whole-body insulin action) after 12 weeks of the high-fat diet, although the decrease in blood glucose after insulin did trend lower in the Tg-Nor-1 mouse line (Figure 4G). In concordance, we observed no significant differences in AKT and pAKT (serine 473) expression in skeletal muscle of WT and Tg-Nor-1 littermates fed the high-fat diet (Supplemental Figure 2C). Furthermore, the analysis of insulin-stimulated phosphorylation of Thr308 and Ser473 in WT and Tg-Nor-1 mice revealed no significant differences in insulin signaling between the WT and transgenic mice fed the high-fat diet in skeletal muscle (Supplemental Figure 2D) and liver (Supplemental Figure 2E).
In summary, we observed that the Tg-Nor-1 mouse line maintained normal fasting blood glucose levels in contrast to hyperglycemia in WT mice fed the high-fat diet. However, they are not protected from glucose intolerance and insulin insensitivity with the high-fat diet (despite reduced weight gain). It is important to note that with a normal chow diet the transgenic mice display improved glucose tolerance on a background of normal insulin sensitivity (3).
Expression profiling revealed a generalized increased expression of genes associated with enhanced substrate utilization (glycolysis, the TCA cycle, mitochondrial oxidation, and pyruvate/fatty acid metabolism) in skeletal muscle
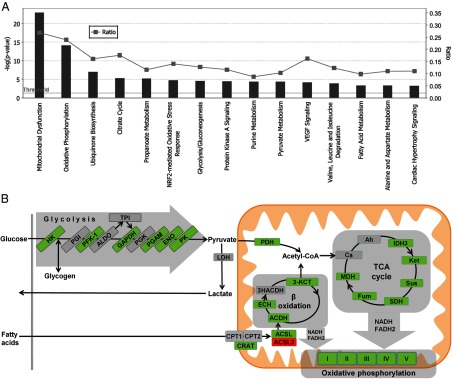
We performed expression profiling on quadriceps femoris skeletal muscle RNA from 4 littermate pairs of fasted WT and Tg-Nor-1 mice (normal chow diet) on the Illumina BeadArray platform. The raw data were analyzed using GeneSpring GX software. The analysis identified >1400 genes/mRNAs that were differentially expressed in a significant manner. The top 50 up- and down-regulated genes are listed in Tables 1 and 2. Ingenuity pathway analysis was performed to identify canonical pathways with a statistically significant enrichment of differentially expressed genes from the Illumina BeadArray expression data and the top 15 results are shown in Figure 5A. More than 25% of the genes in the mitochondrial dysfunction canonical pathway (P < 10−26) encompassing key mitochondrial genes including oxidative phosphorylation were significantly differentially expressed in the Tg-Nor-1 mice relative to expression in WT littermates (Figure 5A and Supplemental Figure 3). Moreover, in the context of metabolism, Ingenuity analysis also identified significantly differential gene expression associated with the pathways involved in oxidative phosphorylation (>20% of genes), the TCA cycle (>15% of genes), glycolysis/gluconeogenesis (>10% of genes), pyruvate metabolism (>10% of genes), and fatty acid metabolism (9% of genes) (Figure 5A). Moreover, increased oxidative metabolism of glucose and fatty acids in skeletal muscle is a hallmark of decreased adiposity and resistance to diet-induced obesity (21–23).
Table 1.
The Top 50 Up-Regulated Genes from Illumina Expression Profiling in Quadriceps Femoris
| Regulation | Fold Change | Symbol | P Value | Corrected P Value | Entrez_Gene_ID | Probe ID |
|---|---|---|---|---|---|---|
| Up | 9.6792555 | Myh8 | 4.64E − 04 | .013100724 | 17885 | 2900131 |
| Up | 7.919811 | Slc47a1 | 6.24E − 05 | .006003437 | 67473 | 6520022 |
| Up | 7.3127294 | Fhl1 | 1.23E − 04 | .007900368 | 14199 | 5550309 |
| Up | 7.270418 | Fhl1 | 1.23E − 04 | .007900368 | 14199 | 10639 |
| Up | 6.8818226 | Abra | 2.71E − 05 | .004622437 | 223513 | 7550066 |
| Up | 6.770568 | Mb | 4.25E − 04 | .012674389 | 17189 | 1710341 |
| Up | 6.687135 | Atp1b4 | 1.99E − 05 | .004599521 | 67821 | 6370632 |
| Up | 6.1377153 | Nr4a3 | 4.68E − 06 | .002363166 | 18124 | 1990255 |
| Up | 5.700996 | Car14 | 1.56E − 04 | .008914627 | 23831 | 5870220 |
| Up | 5.4764333 | Nr4a3 | 9.85E − 08 | 8.17E − 04 | 18124 | 4260731 |
| Up | 5.231374 | Myl2 | .00124067 | .019644732 | 17906 | 2120356 |
| Up | 5.205544 | Csrp3 | .002208978 | .024711978 | 13009 | 2470133 |
| Up | 5.18676 | Hr | 1.02E − 04 | .007475142 | 15460 | 6980671 |
| Up | 5.1608715 | Cul7 | 1.43E − 05 | .004219131 | 66515 | 5260309 |
| Up | 5.121954 | Myoz2 | .002081666 | .023822872 | 59006 | 5050181 |
| Up | 4.7647805 | Csrp3 | .001039745 | .018017199 | 13009 | 3140202 |
| Up | 4.729777 | Rab15 | 4.19E − 05 | .005433246 | 104886 | 6590343 |
| Up | 4.720416 | Smyd2 | 1.34E − 06 | .001393425 | 226830 | 2600086 |
| Up | 4.7062006 | Slc47a1 | 1.60E − 05 | .004278829 | 67473 | 1260129 |
| Up | 4.686325 | Actn2 | 1.10E − 05 | .004219131 | 11472 | 1050678 |
| Up | 4.61831 | 2310042D19Rik | .001507363 | .020879112 | 74183 | 5130593 |
| Up | 4.5696363 | Aqp4 | 1.15E − 04 | .007770291 | Aqp4 | 7160193 |
| Up | 4.566424 | Slc47a1 | 1.44E − 04 | .008713499 | 67473 | 2030278 |
| Up | 4.43305 | Arhgap24 | 6.22E − 05 | .006003437 | 231532 | 6550470 |
| Up | 4.339516 | Ankrd2 | .007291165 | .04572547 | 56642 | 6060201 |
| Up | 4.28916 | Hr | 5.32E − 05 | .006003437 | 15460 | 6330605 |
| Up | 4.227805 | Csrp3 | .003147146 | .029207906 | 13009 | 5340722 |
| Up | 4.2260685 | Bcl2 | 8.41E − 05 | .006518213 | 12043 | 6420612 |
| Up | 4.1306534 | Myh1 | .001700432 | .021839757 | 17879 | 5080725 |
| Up | 4.0597634 | Stbd1 | 1.73E − 04 | .009111944 | 52331 | 1430129 |
| Up | 3.8280413 | Slc7a2 | .00401804 | .03363351 | 11988 | 5960221 |
| Up | 3.7488759 | Myh8 | .002434061 | .025693897 | 17885 | 6650435 |
| Up | 3.6844172 | Myh1 | .002344499 | .025527965 | 17879 | 6860228 |
| Up | 3.6547031 | Hspb7 | .001493377 | .020819558 | 29818 | 6480612 |
| Up | 3.4966555 | Ctnnbl1 | 4.22E − 04 | .01262911 | 66642 | 6130300 |
| Up | 3.3678412 | Smyd2 | 1.96E − 06 | .001527346 | 226830 | 5720050 |
| Up | 3.11052 | Car3 | .001585771 | .021162586 | 12350 | 1450242 |
| Up | 3.0825765 | Bcl2 | 7.21E − 05 | .00615364 | 12043 | 1400168 |
| Up | 3.068402 | Tbc1d1 | 1.53E − 05 | .004219131 | 57915 | 3940142 |
| Up | 3.016272 | Lmcd1 | .001059903 | .018169448 | 30937 | 5720164 |
| Up | 2.9962347 | Hspa2 | 7.95E − 05 | .006439419 | 15512 | 1070484 |
| Up | 2.9834642 | Lmod2 | .001405266 | .020559967 | 93677 | 2750475 |
| Up | 2.8527691 | Mb | .002019617 | .023433225 | 17189 | 1470154 |
| Up | 2.805313 | Dnaja4 | 4.98E − 04 | .013371915 | 58233 | 5690181 |
| Up | 2.7607124 | Fam171b | .005425021 | .03903851 | 241520 | 7150026 |
| Up | 2.7307756 | Mustn1 | 6.30E − 04 | .014816274 | 66175 | 2570193 |
| Up | 2.714702 | Ppap2a | 1.95E − 04 | .009111944 | 19012 | 2480706 |
| Up | 2.6870883 | Gm4841 | .003623456 | .03154976 | 225594 | 7550161 |
| Up | 2.676914 | Rom1 | .001309442 | .02000817 | 19881 | 6280377 |
| Up | 2.6764731 | Fxyd6 | .002522926 | .025938926 | 59095 | 6550309 |
Table 2.
The Top 50 Down-Regulated Genes from Illumina Expression Profiling in Quadriceps Femoris
| Regulation | Symbol | Fold Change | P Value | Corrected P Value | Entrez_Gene_ID | Probe ID |
|---|---|---|---|---|---|---|
| Down | Dkk3 | −13.161947 | 4.65E − 05 | .005760497 | 50781 | 3180682 |
| Down | Dkk3 | −11.689332 | 2.31E − 05 | .004622437 | 50781 | 2680189 |
| Down | Gdap1 | −6.014221 | 8.66E − 05 | .006653792 | 14545 | 7400356 |
| Down | Odc1 | −4.722649 | 4.26E − 05 | .005433246 | 18263 | 6840121 |
| Down | Ppp1r1a | −4.372698 | 1.85E − 04 | .009111944 | 58200 | 1300288 |
| Down | Adrb2 | −4.0957026 | 5.40E − 04 | .013978633 | 11555 | 3450440 |
| Down | Amd2 | −3.8233154 | 8.65E − 04 | .016820148 | 11703 | 4830768 |
| Down | Slc2a3 | −3.5210745 | 3.74E − 05 | .005037385 | 20527 | 460327 |
| Down | Tmem100 | −3.5197556 | 1.90E − 04 | .009111944 | 67888 | 1410168 |
| Down | Gpt2 | −3.4827254 | 5.72E − 04 | .014148096 | 108682 | 1340026 |
| Down | Ces1d | −3.4255352 | .002538188 | .026031334 | 104158 | 670603 |
| Down | Cbr2 | −3.399576 | 2.62E − 04 | .010703774 | 12409 | 2490672 |
| Down | Pde4b | −3.3287776 | 1.14E − 04 | .007770291 | 18578 | 1400360 |
| Down | Gdap1 | −3.2977467 | 8.75E − 04 | .016820148 | 14545 | 6550341 |
| Down | Acsl3 | −3.292784 | 7.00E − 04 | .015164451 | 74205 | 5560615 |
| Down | Aebp1 | −3.225082 | 6.94E − 05 | .006121588 | 11568 | 4480364 |
| Down | Amd1 | −3.1573136 | 3.75E − 04 | .011781803 | 11702 | 4610059 |
| Down | Ppp1r27 | −3.0609093 | 1.03E − 04 | .007475142 | 68701 | 4220575 |
| Down | Cbfb | −3.0067728 | 5.37E − 05 | .006003437 | 12400 | 5960672 |
| Down | Cbfb | −2.979785 | 3.73E − 05 | .005037385 | 12400 | 4860692 |
| Down | Gpcpd1 | −2.8454146 | 2.09E − 04 | .009310676 | 74182 | 7100608 |
| Down | Mmd | −2.8296688 | 1.75E − 04 | .009111944 | 67468 | 5670082 |
| Down | Cacng1 | −2.8092854 | 7.42E − 05 | .00615364 | 12299 | 3780347 |
| Down | Tpra1 | −2.8016882 | 9.65E − 04 | .017562693 | 24100 | 6130193 |
| Down | Gpcpd1 | −2.779834 | 5.74E − 05 | .006003437 | 74182 | 1110538 |
| Down | Chac1 | −2.747746 | 2.01E − 04 | .009111944 | 69065 | 540300 |
| Down | Kcnab1 | −2.7270942 | 6.84E − 05 | .006121588 | 16497 | 6130471 |
| Down | Gpx3 | −2.6830268 | 1.48E − 04 | .008731124 | 14778 | 7320053 |
| Down | Prei4 | −2.65897 | 3.23E − 05 | .004624852 | 74182 | 3290068 |
| Down | Ociad2 | −2.6388347 | 1.85E − 04 | .009111944 | 433904 | 6350133 |
| Down | Exoc7 | −2.6352344 | 1.25E − 04 | .007959393 | 53413 | 290167 |
| Down | Cova1 | −2.5704007 | .002921225 | .02811764 | 209224 | 5870450 |
| Down | Gadd45g | −2.5554438 | .006729802 | .043691054 | 23882 | 6330377 |
| Down | Sub1 | −2.4991143 | 6.05E − 07 | 9.72E − 04 | 20024 | 610278 |
| Down | Actn3 | −2.4937787 | 4.90E − 04 | .013334262 | 11474 | 3140164 |
| Down | Kcnc4 | −2.472529 | 1.53E − 04 | .008804545 | 99738 | 4010465 |
| Down | Clip1 | −2.47217 | .00112276 | .018825145 | 56430 | 6900091 |
| Down | Cbfb | −2.4476533 | .001793815 | .022247061 | 12400 | 2710452 |
| Down | Sepx1 | −2.419562 | .001182965 | .019145174 | 27361 | 3800136 |
| Down | 4933407P14Rik | −2.4178221 | 2.00E − 05 | .004599521 | 237958 | 5910156 |
| Down | Vldlr | −2.4066355 | 3.44E − 04 | .011415064 | 22359 | 4070201 |
| Down | Tceal5 | −2.3569448 | 6.81E − 04 | .015085584 | 331532 | 7610767 |
| Down | Pde4a | −2.310972 | .001935201 | .023069492 | 1857 | 1940072 |
| Down | Camk2a | −2.308476 | 6.46E − 06 | .002821202 | 12322 | 2630600 |
| Down | Tiam1 | −2.307557 | .002809688 | .027523002 | 21844 | 5270452 |
| Down | Ampd1 | −2.2858315 | 5.37E − 04 | .013969612 | 229665 | 610671 |
| Down | Rtn4 | −2.2635634 | 7.63E − 04 | .015763158 | 68585 | 7040554 |
| Down | Ampd1 | −2.2286847 | .001012626 | .017930113 | 229665 | 4290273 |
| Down | Cyp27a1 | −2.2272053 | 2.72E − 04 | .010916186 | 104086 | 4010020 |
| Down | Slc15a4 | −2.2066624 | .003429194 | .030725729 | 100561 | 3780167 |
Figure 5.
Ingenuity pathway analysis. A, Differentially expressed genes from the Illumina BeadArray analysis comparing WT and Tg-Nor-1 quadriceps femoris muscle were analyzed via Ingenuity pathway analysis to reveal the top 15 canonical pathways associated with differentially expressed genes. B, Schematic summary diagram of significant metabolic pathways identified by ingenuity pathway analysis as being significantly regulated. Green denotes that 1 or more subunits within a protein are significantly increased in Tg-Nor-1 quadriceps femoris muscle compared to WT. Conversely, red denotes significant repression.
The differentially expressed and significantly induced genes (highlighted in green) from Illumina profiling were primarily associated with the oxidative utilization of glucose and fatty acids (Figure 5B) and in concordance with the increased oxidative capacity and mitochondrial biogenesis reported in the previous study (3).
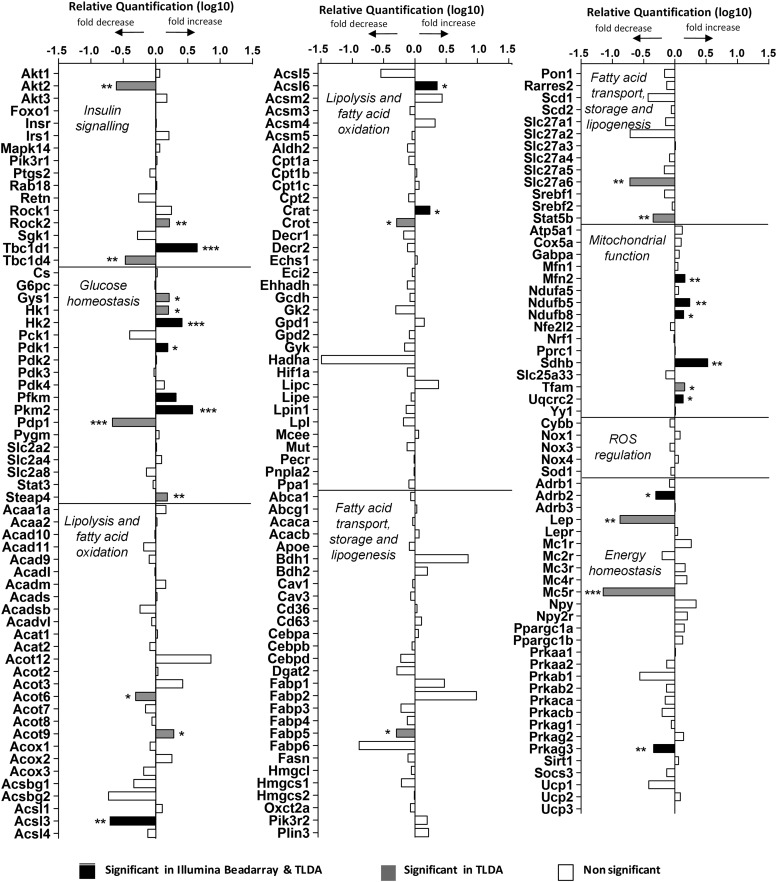
In parallel, we used custom-designed TaqMan low-density arrays (TLDAs) to examine the expression of ∼200 critical genes involved in lipid, carbohydrate, and energy homeostasis and insulin signaling by quantitative PCR (qPCR) in skeletal muscle from the normal chow diet study. This approach validated many hits identified by Illumina profiling and identified additional target genes in those pathways (Figure 6). Similar genes were identified in the qPCR-TLDA profiling of skeletal muscle RNA from the high-fat diet study (Supplemental Figure 4).
Figure 6.
Categorized gene expression from custom TLDA analysis of quadriceps femoris muscle. Significance from Illumina BeadArray expression of quadriceps femoris muscle was also included. *, P < .05; **, P < .01; ***, P < .001.
Interestingly, we identified the significant and differential expression of several other genes relevant to other phenotypic attributes of the Tg-Nor-1 mice. In the context of the endurance phenotype (and decreased sprint performance), we identified decreased expression of the mRNA encoding α-actinin-3 (Actn3) in quadriceps, gastrocnemius, and soleus muscles (Figure 7, A, B, and C, respectively). The α-actinins are sarcomeric binding proteins (interacting with the thick and thin filaments) and important constituents of the Z-disc line regulating contractile activity. ACTN3 polymorphisms in humans affect metabolism and exercise performance. The frequency of the null polymorphism is significantly increased in endurance competitors (24, 25). Furthermore, α-actinin-3-deficient rodents display increased endurance, reduced power, a phenotypic shift in the characteristics of type II muscle to type I and the elevated activity of oxidative mitochondrial enzymes (26–28). This phenotype bears a marked resemblance to the Tg-Nor-1 phenotype.
Figure 7.
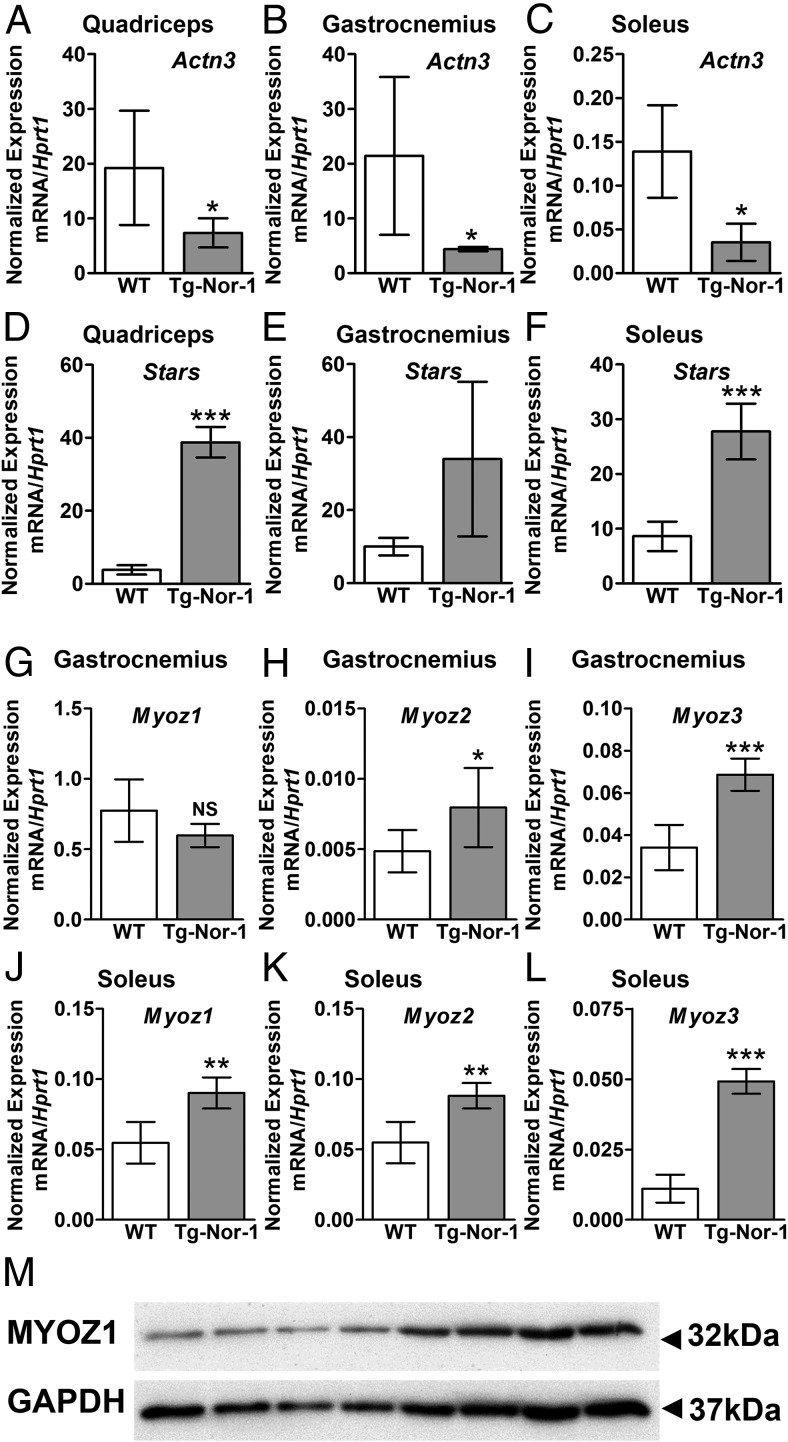
Expression of sarcomeric genes by Tg-Nor-1 mice. A–C, Expression of Actn3 in quadriceps femoris (A), gastrocnemius (B), and soleus (C) via RT-qPCR (n = 4). D–F, Expression of Stars in quadriceps femoris (F), gastrocnemius (E), and soleus (F) via RT-qPCR (n = 4). G–I, Expression of Myoz1 (G), Myoz2 (H), and Myoz3 (I) in gastrocnemius via RT-qPCR (n = 4–8). J–L, Expression of Myoz1 (J), Myoz2 (K), and Myoz3 (L) in soleus via RT-qPCR (n = 4–8). Statistical calculation was performed using a Student t test, and P values are indicated on graphs as follows: *, P < .05; **, P < .01; ***, P < .001. M, Western blot analysis of MyOZ1 on quadriceps femoris muscle (n = 4).
Moreover, we identified significantly induced expression of actin-binding Rho-activating protein (ABRA)/striated muscle activator of Rho signaling (STARS) in the Tg-Nor-1 mice (Figure 7, D, E, and F). STARS is a skeletal muscle actin-binding protein that responds to endurance, environmental, and mechanical signals influencing muscle reprogramming, growth, and contraction (29–31). For example, ABRA/STARS overexpression induces myosin heavy chain (MHC) IIA and IIX, in agreement with the contractile protein transitions observed in our Tg-Nor-1 mice (32). We also observed increased expression of serum response factor (Srf), myocardin-related transcription factors A and B (Mrtf-a and Mrtf-b), and Cox4 (Supplemental Figure 5), these genes have been associated with ABRA/STARS overexpression in cell culture and exercise in human skeletal muscle (29–31).
Finally, we observed significant differential expression of genes regulating calcineurin signaling and fiber type reprogramming in the Tg-Nor-1 mice, eg, Myoz1–3 genes encoding the calsarcin muscle–specific calcineurin-binding proteins that interact with many Z-band interacting proteins including α-actinin (Table 1 and Figure 7, G–L). Specifically, we observed significantly increased expression of the mRNA encoding Myoz1/calsarcin-2 in type I (soleus) muscle (Figure 7J) and the Myoz1/calsarcin-2 protein in quadriceps femoris muscle tissue (Figure 7M) from the Tg-Nor-1 mice relative to that in WT littermates. In addition, we observed significantly increased expression of Myoz2 mRNA in type II (gastrocnemius) and type I (soleus) muscle tissue (Table 1 and Figure 7, H and K). Finally, we observed significantly increased expression of Myoz3 mRNA in type II (gastrocnemius) and type I (soleus) muscle tissue (Figure 7, I and L). As reported in previous studies (33, 34), Myoz2 and Myoz3 are preferentially expressed in type I and type II skeletal muscle, respectively (Figure 7, K and I). This finding is consistent with the decrease in type I MHC and increase in type IIA/IIX MHC in the soleus tissue identified in the previous article (3).
In summary, the profiling identified differential expression of genes involved in oxidative metabolism. Furthermore, the profiling identified several critical genes encoding sarcomeric and calcineurin-binding proteins that regulate fatigue resistance, respond to mechanical loads, and reprogram skeletal muscle.
Transgenic muscle-specific expression of activated Nor-1 increases the glycogen levels of skeletal muscle
Elevated skeletal muscle glycogen is a hallmark of endurance; furthermore, glycogen content is increased in α-actinin-3-deficient humans and mice (35). Moreover, increasing glycogen synthase activity by small molecule inhibition of glycogen synthase kinase prevents diet-induced obesity in rodents (36), and exercise enhances glycogen synthase activity in obese patients (37). Interestingly, in this context, the array and TLDA analysis (Tables 1 and 2 and Figure 6) demonstrated significantly increased expression of the mRNAs encoding glycogen synthase (Gys1) and starch binding domain-containing protein 1 (Stbd1) and significantly decreased expression of protein phosphatase 1 regulatory (inhibitor) subunit 1a (Ppp1r1a).
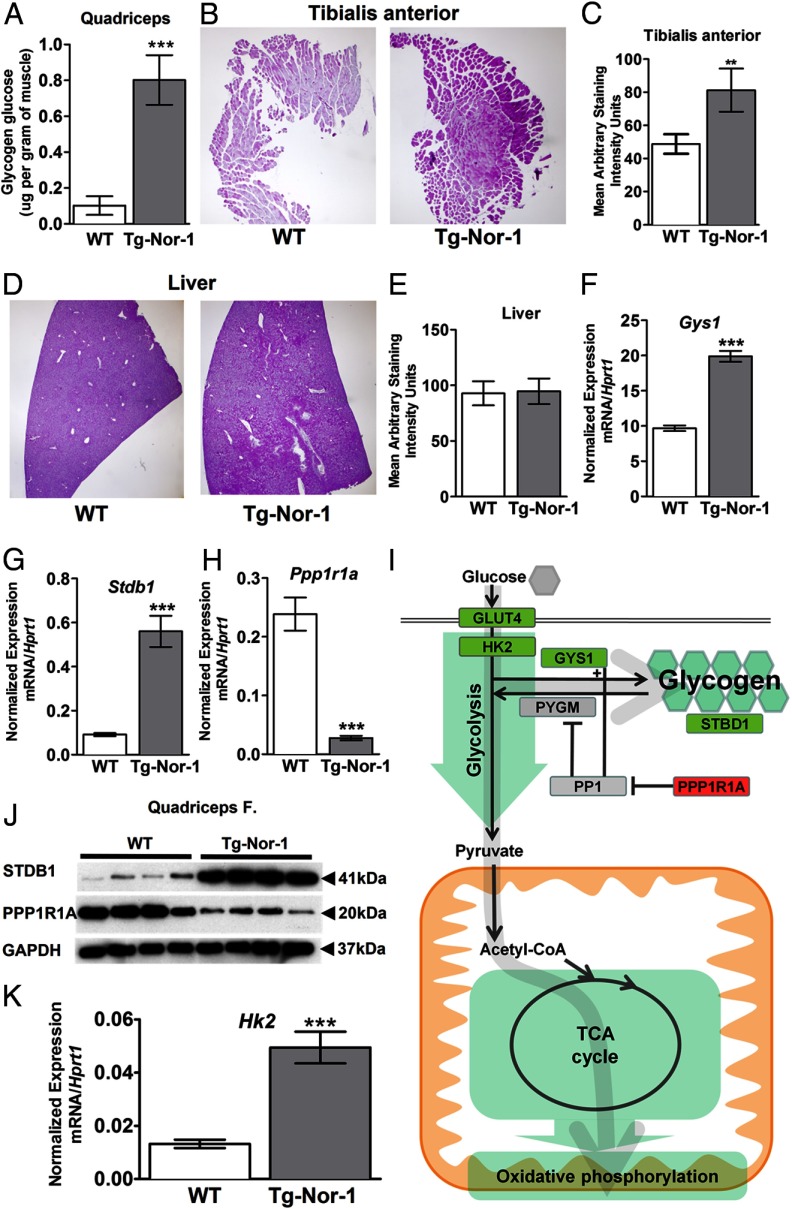
Consequently, we used the BioVision glycogen assay kit (utilizing glucoamylase hydrolysis) to directly measure tissue glycogen, which identified a very significant increase in glycogen levels in quadriceps muscle tissue from Tg-Nor-1 mice relative to that in WT littermates (n = 4) (Figure 8A). This result was further validated by the use of periodic acid-Schiff (PAS) staining, which confirmed the significant increase in glycogen in another muscle, tibialis anterior, from the Tg-Nor-1 mice relative to that in WT littermates (Figure 8, B and C), against the background of no differences in liver glycogen levels in WT and Tg-Nor-1 mice (Figure 8, D and E).
Figure 8.
Tg-Nor-1 mice display increased glycogen content and expression of glycogenic genes. A, Glycogen content in quadriceps femoris muscle samples from nonfasted WT and Tg-Nor-1 mice (n = 8). B, PAS staining of tibialis anterior muscle sections to visualize glycogen stores (representative photographs, n = 5 mice/group), C, Quantification of the area of tibialis anterior muscle sections positively stained with Schiff reagent. D, PAS staining of liver sections (n = 5 mice/group). E, Quantification of the area of liver sections positively stained with Schiff reagent. F–H, RT-qPCR examining quadriceps femoris mRNA expression of Gys1 (F), Stbd1 (G), and Ppp1r1a (H) (n = 8). I, Diagram illustrating significant gene changes in glycolysis, glycogen, the TCA cycle, and the oxidative phosphorylation pathway. J, Western blot analysis of STBD1 and PPP1R1A on quadriceps femoris muscle (n = 4). K, RT-qPCR examining quadriceps femoris mRNA expression of Hk2 (n = 8). Statistical calculation was performed using a Student t test, and P values are indicated on graphs as follows: *, P < .05; ***, P < .001.
We observed significantly increased expression of the mRNAs encoding Gys1 and Stbd1 and significantly decreased expression of Ppp1r1a in our array analysis, which was validated by qPCR (Tables 1 and 2 and Figure 8, F–H). Using commercially available antibodies and Western blot analysis, we validated the significant repression of Ppp1r1a that suppresses protein phosphatase 1 activity, an inhibitor of glycogen phosphorylase/glycogenolysis, and the increase in Stbd1 (Figure 8J). The increase in Gys1 mRNA and validation (by qPCR and Western blot) of repression of Ppp1r1a that suppresses protein phosphatase 1 activity (coupled to the increase in Stbd1) are consistent with the observed net increase in glycogen in the Tg-Nor-1 mice. Decreased Ppp1r1a expression would presumably lead to activated protein phosphatase 1 activity and the inhibition of glycogen phosphorylase and glycogenolysis (Figure 8I).
Finally, we observed elevated hexokinase 2 (Hk2) mRNA expression by qPCR (and on the Illumina array) in the Tg-Nor-1 mice relative to that in WT littermates fed the normal chow diet (Figure 8K and Supplemental Figure 6A). Glucose is phosphorylated by HK2 in skeletal muscle (38), and this is a key mechanism controlling glucose uptake and glycogen synthesis in skeletal muscle (39–41). Therefore, this finding is in concordance with the endurance phenotype and elevated glycogen levels (42, 43). Furthermore, we also observed elevated protein expression of the glucose transporter GLUT4 in the Tg-Nor-1 mice fed the high-fat diet (Supplemental Figure 6B).
In summary, the increased level of glycogen in skeletal muscle from Tg-Nor-1 mice is in concordance with decreased adiposity, increased endurance phenotype, and the gene expression profile.
Decreased NAD+/NADH ratio in skeletal muscle and the induction of the malate-aspartate shuttle
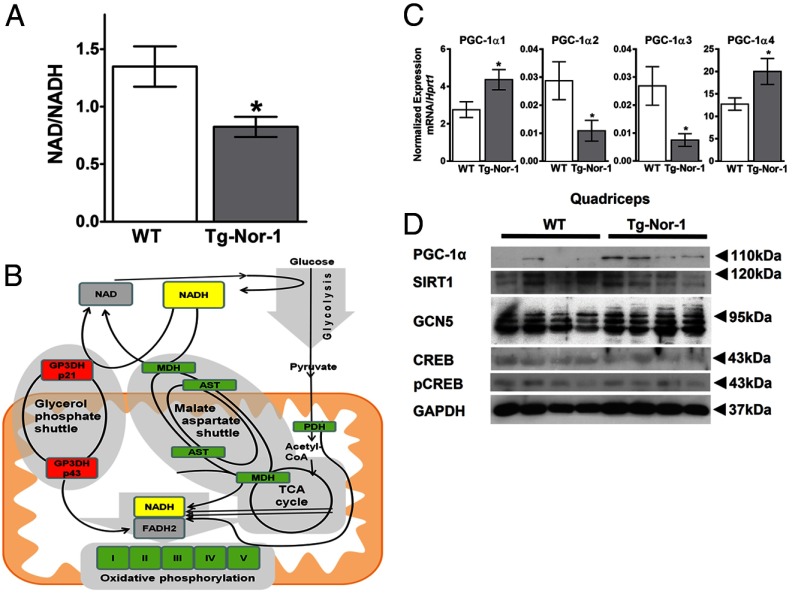
The redox state (NAD+/NADH) reflects the oxidative state and capacity for the generation of ATP in mitochondria. In skeletal muscle, metabolism and exercise modulate the ratio of these coenzymes. We investigated the NAD+ and NADH levels in gastrocnemius muscle from Tg-Nor-1 mice, relative to those in WT littermates. NAD and NADH levels were measured using an NAD/NADH Assay Kit that specifically detects NADH and NAD, but not NADP nor NADPH. We observed a decreased NAD+/NADH ratio in skeletal muscle (Figure 9A) that suggests an increase in oxidative ATP production (44).
Figure 9.
NAD+/NADH and PGC-1α splice variants are modulated by Nor-1 expression. A, NAD+/NADH ratio assay performed on gastrocnemius muscle tissue. B, Diagram illustrating glycolysis, NAD+, NADH, and the malate-aspartate shuttle, highlighting the significant hits from Illumina mRNA expression profiling. C, RT-qPCR analysis of quadriceps femoris muscle examining the PGC-1α splice variants of WT and Tg-Nor-1 mice fed thea normal chow diet (n = 4). D and E, High-fat diet-fed quadriceps femoris PGC-1α (D) and Western blot analysis on quadriceps femoris muscle (E) (n = 4). Statistical calculation was performed using a Student t test, and P values are indicated on graphs as follows: *, P < .05.
In this context, we note increases in the expression of genes involved in the malate-aspartate shuttle and a decrease in the glycerol 3-phosphate shuttle, the pathways responsible for translocation of electrons (generated from glycolysis) across the mitochondrial inner membrane for oxidative phosphorylation (Figure 9B). The induction of genes in the malate-aspartate shuttle is consistent with maximal and most efficient ATP production. For example, the malate-aspartate shuttle boosts the number of ATPs formed in glycolysis relative to those for the glycerol 3-phosphate shuttle.
Transgenic skeletal muscle-specific Nor-1 expression leads to increased PGC-1α (mRNA and protein) in skeletal muscle
We have previously reported increased mitochondrial numbers/DNA and significantly increased mRNA/protein expression of key mitochondrial mRNA/proteins (3). This finding coupled to increased glycogen, the endurance phenotype, increased MEF2C expression and the acquisition of more oxidative type II muscle fibers in the Tg-Nor-1 mice is indicative of increased PGC-1α expression or function (3). Using RT-qPCR analysis in our previous study (3), we did observe increased expression of the mRNAs encoding Pgc-1α and Pgc-1β in soleus/type I (soleus) and type II (gastrocnemius/quadriceps femoris) skeletal muscle (that did not obtain significance). However, the RT-qPCR analysis was conducted with primers that did not discriminate the newly characterized (45) splice variants of Pgc-1α (1–4). We conducted RT-qPCR analysis on RNA extracted from quadriceps muscle tissue isolated from normal chow-fed WT and Tg-Nor-1 mice. We observed very abundant and significantly increased expression of the PGC-1α1 splice variant in the skeletal muscle from Tg-Nor-1 mice relative to that in WT littermates (Figure 9C). The expression of the PGC-1a2–4 splice variants was approximately 100-fold lower. This in concordance with the endurance phenotype (45).
It has been recently demonstrated that the 20S gatekeeper molecule NQO1 binds and guards PGC-1α protein from 20S proteosomal decay in an NADH-dependent manner (46). Given the enhanced NADH results above, we examined PGC-1α protein expression by Western blot analysis in Tg-Nor-1 mice relative to that in WT littermates (Figure 9D). A single band at approximately 110 to 120 kDa was observed to be increased in Tg-Nor-1 mice. This is consistent with the 113-kDa estimated size of PGC-1α1 (45). No other bands were visible. We also observed no changes in the protein expression levels of the enzymes that acetylate (GCN5) (47) and deacetylate (SIRT1) PGC-1α (48). Furthermore, we did not observe any changes in the transcriptional activator of PGC-1α, CREB, or phospho-CREB at Ser133 (49).
In summary, the increased expression of PGC-1α protein and specific mRNA splice variants is in concordance with the oxidative phenotype, the endurance phenotype, and reduced adiposity.
Discussion
In skeletal muscle, adrenergic activation regulates lipolysis, fatty acid oxidation, and energy expenditure (50–53). The mRNA encoding Nor-1 and the other members of the NR4A subgroup are dramatically and significantly induced by β2-adrenergic signaling in skeletal muscle cells in vitro (10- to 30-fold) (54) and in glycolytic and oxidative skeletal muscle (50- to 100-fold) (7, 9, 54). The β-adrenergic-dependent response, coupled to a selective activation of this subgroup of orphan NRs in skeletal muscle and other major mass tissues (55) suggested a role in metabolism. In vitro analysis in muscle culture models validated this hypothesis and has demonstrated that Nor-1 expression correlates with oxidative metabolism (7, 54). For example, Nor-1 small interfering RNA (that resulted in a 3-fold decrease in Nor-1 mRNA expression) increased anaerobic metabolism and lactate accumulation (7). Several lines of evidence have also indicated the importance of Nor-1 in the regulation of insulin responsiveness. Expression of Nor-1 mRNA is (1) induced by insulin (14) and thiazolidinediones in adipocytes (17) and (2) decreased in muscle tissue in multiple models of insulin resistance (17, 56). In vitro, overexpression of Nor-1 in adipocyte and muscle cells increased (1) the ability of insulin to augment glucose transport and (2) insulin-dependent phosphorylation of insulin receptor substrate-1 (IRS-1) and AKT (17, 56). The expression of NR4A family member Nur77 is decreased in muscle biopsy samples from obese men relative to those from lean controls and inversely correlated with body fat content and insulin sensitivity (6). Global genetic ablation of NR4A1/Nur77 displayed an amplified predisposition to diet-induced obesity and insulin resistance (4). In humans, NR4A3/Nor-1 mRNA expression is increased in obese adipose tissue (20); however, information on the expression and function of skeletal muscle Nor-1 in an obesogenic environment has been lacking.
Previously, we have demonstrated that transgenic skeletal muscle–specific expression of activated NR4A3/Nor-1 (that increased total Nor-1 mRNA expression 3- to 15-fold in type I and II fibers) resulted in the acquisition of an endurance phenotype, an increase in type IIA/X oxidative muscle fibers, myoglobin, numbers of mitochondria, and glucose tolerance (3). In our current study, we have demonstrated that muscle-specific Tg-Nor-1 mice (relative to WT littermates) displayed reduced adiposity (with the normal chow diet) and resistance to high-fat diet–induced obesity. Moreover, the Tg-Nor-1 mice maintained fasting glucose at normoglycemic levels with the energy-dense diet. Expression profiling on the Illumina array platform, extensive qPCR validation on the TLDA platform coupled to manual qPCR (∼200 genes) and Ingenuity analysis indicated significant increases in several canonical pathways involved in glycolysis, the TCA cycle, oxidative phosphorylation, fatty acid oxidation, and glycogen synthesis. Increased oxidative metabolism of glucose and fatty acids in skeletal muscle is a hallmark of decreased adiposity and resistance to diet-induced obesity. Indeed, some features of the lean phenotype displayed by Tg-Nor-1 mice show a resemblance to other NR mouse models that also produce enhanced oxidative glycolytic fiber types (IIX/IIA), improved endurance, and increased mitochondria, eg, gain of function of peroxisome proliferator-activated receptor-δ (21) and estrogen-related receptor-γ transgenic models (57).
The decrease in type I skeletal muscle fibers and reciprocal increase in type IIA/X fibers in the Tg-Nor-1 mice suggests increased oxidative capacity. Our hypothesis that the lean phenotype involves increased utilization of carbohydrates in this mouse model has multiple lines of support. First, we report patterns of increased gene expression in pathways that direct glycolysis, the TCA cycle, pyruvate dehydrogenase, and oxidative phosphorylation. This suggests increased aerobic utilization of glucose as shown in Figures 5C and 8I. Second, we observed an elevated RER during nighttime feeding. Typically this can be attributed to either enhanced carbohydrate oxidation/utilization or increased carbohydrate-dependent lipogenesis. Given that the Tg-Nor-1 mice have a lean body composition, it appears likely that carbohydrate oxidation is favored over lipogenesis. Third, we observed that the Tg-Nor-1 mice maintained normal fasting glucose with the high-fat diet. This was associated with significantly increased GLUT4 (protein) and Hk2 mRNA expression in the Tg-Nor-1 mice fed the normal chow and high-fat diets. Hk2 commits glucose to either being metabolized or stored (as glycogen) in skeletal muscle, whereas GLUT4 controls the flow of glucose in response to demand. Increased Hk2 mRNA expression is also consistent with increased storage and utilization of glucose observed in this mouse model.
In the context of the endurance phenotype (associated with decreased sprint performance), we identified significantly reduced expression of the mRNA encoding Actn3 in the Tg-Nor-1 mice. The α-actinins are sarcomeric binding proteins that are preferentially expressed in type II fibers and interact with the thick and thin filaments. As constituents of the Z-disc line, they are important for regulating contractile activity. In humans, polymorphisms in the Actn3 gene affect both metabolism and exercise performance: the null polymorphism is present in ∼15% of the population. The frequency is significantly enriched in endurance and road cycling competitors and is strongly linked with decreased performance in sprint and power sporting activities (24, 25). Similarly, α-actinin-3–deficient rodents display increased endurance, reduced power, a phenotypic shift in the characteristics of type II muscle to type I, and elevated expression of genes (eg, Hk2) in glycolysis and oxidative mitochondrial enzymes in aerobic metabolism (26–28). This phenotype is almost identical to the Tg-Nor-1 phenotype described previously (3) and in the current article. Interestingly, we also observed the compensatory increase in actinin-2 mRNA expression (Table 3), the actinin/actin binding protein that is ubiquitously expressed in skeletal and cardiac muscle. This response to actinin-3 deficiency is also observed in the actinin-3-deficient mouse models and in humans with the null polymorphism in this gene (35).
Table 3.
Sarcomeric and Z-Disk Binding Proteins
| Name | Symbol | Fold Change |
|---|---|---|
| Actin-binding ρ activating protein | ABRA | 6.882 |
| Cysteine and glycine-rich protein 3 | CSRP3 | 5.206 |
| Myozenin 2 | MYOZ2 | 5.122 |
| Actinin, alpha 2 | ACTN2 | 4.686 |
| LIM and cysteine-rich domains 1 | LMCD1/Dyxin | 3.016 |
| Musculoskeletal, embryonic nuclear protein 1 | MUSTN1 | 2.731 |
| PDZ and LIM domain 1 | PDLIM1 | 2.058 |
| Dystrobrevin, alpha | DTNA | 2.024 |
| Dysbindin (dystrobrevin binding protein 1) domain containing 2 | DBNDD2 | 2.014 |
| Myosin binding protein C, slow type | MYBPC1 | 1.957 |
| Myotilin | MYOT | 1.705 |
| Regulator of calcineurin | RCAN | 1.646 |
| Vinculin | VCL | 1.644 |
| Calcium/calmodulin-dependent protein kinase II δ | CAMK2D | 1.564 |
| Myosin VIIA and Rab interacting protein | MYRIP | 1.529 |
| PDZ and LIM domain 4 | PDLIM4 | 1.448 |
| Myomesin (M protein) 2 | MYOM2 | 1.444 |
| LIM domain binding 3 | LDB3 | 1.44 |
| Actinin alpha 3 | ACTN3 | −2.494 |
Moreover, we identified significantly induced expression of ABRA/STARS in the Tg-Nor-1 mice. STARS is a skeletal muscle actin-binding protein that influences muscle reprogramming, growth, and contraction in response to environmental and mechanical signals. For example, ABRA/STARS overexpression induces MHCIIA and IIX, in agreement with the contractile protein transitions observed in the Tg-Nor-1 mice and is a PGC-1α-dependent gene. STARS overexpression is associated with increased markers of regeneration, such as the myosin heavy chain 8 (perinatal) isoform (32) and is one of the most significantly induced genes in our study (Table 1). It is interesting to note that actinin-3-deficient mice display increased susceptibility to muscle damage and remodeling and differential expression of Z-disk-binding proteins (58). In the context of our current study, we have observed significantly differential expression of genes encoding several Z-disk interacting proteins and regulators of calcineurin signaling (Table 3). Together these genes/proteins control muscle: reprogramming fiber type, regeneration, and repair. We identified many genes encoding LIM proteins (including, but not limited to, CSRP3, LMCD1, and Fhl1), MUSTN1, myotilin, vinculin, the Myoz/calsarcin genes, RCAN2, and several other sarcomeric interacting proteins (Table 3). MUSTN1 is abundantly expressed in regenerating muscle (59) and along with CSRP3 is induced during the physical adaptations to exercise in humans (60). LIM proteins have been associated with the promotion of myogenesis, and mutations in LIM proteins and other sarcomeric binding protein have been implicated in a plethora of cardiac and skeletal myofibrillar myopathies (61). Finally, the differential expression of the Myoz1-3 genes encoding the calsarcin proteins (Table 1 and Figure 7, G–L) is consistent with the fiber type remodeling observed in the Tg-Nor-1 mice. The calsarcins are muscle-specific calcineurin-binding proteins that interact with many Z-band-interacting proteins including α-actinin-3. Various calsarcins are preferentially expressed in type I or II fiber types and modulate skeletal muscle reprogramming. Specifically, we observed significantly increased expression of the mRNAs encoding Myoz1 and Myoz2 in type I (soleus) muscle and Myoz2 and Myoz3 in type II fibers. Interestingly, Myoz1/calsarcin 2 directly interacts with α-actinin-3 (62). As expected, the relative expression of Myoz3 in gastrocnemius (type II/fast) muscle is elevated in the WT mice, whereas the Tg-Nor-1 mice display comparably high relative expression of Myoz3 in both gastrocnemius (type II/fast) and soleus (type I/slow) muscles. The global knockout of calsarcins 1 (Myoz2) and 2 (Myoz1) results in a shift to type I fibers (33, 34), whereas increased expression of Myoz1–3 in type I and II muscle fibers, reported here, is consistent with the shift to type IIA/X fibers and the down-regulation of type I fibers in Tg-Nor-1 muscles described previously (3). Interestingly, we also observed an increase in Rcan2 mRNA expression (a regulator of calcineurin) that is affected by differential expression of calsarcin (34).
The dramatic increase in skeletal muscle glycogen content observed in the Tg-Nor-1 mice has been underscored by profiling, qPCR, Western blot experiments, and enzymatic glycogen analysis that collectively have provided a substantial foundation for understanding the underlying mechanism. This is an important aspect of the Tg-Nor-1 phenotype for several reasons. First, elevated skeletal muscle glycogen is a hallmark of endurance, and glycogen content is increased in α-actinin-3-deficient humans and mice (35). Second, increasing glycogen synthase activity by small molecule inhibition of glycogen synthase kinase prevents diet-induced obesity in rodents (36), and exercise enhances glycogen synthase activity in obese patients (37). Third, we observed increased hexokinase 2 expression in the skeletal muscle from the Tg-Nor-1 mice. Endurance phenotypes and elevated glycogen are associated with amplified hexokinase 2 mRNA expression (42, 43). Finally, increased GLUT4 expression is consistent with increased glycogen synthesis (39–41) and higher hexokinase 2 expression (that phosphorylates glucose) in skeletal muscle (38). Moreover, studies have demonstrated that increased storage of glucose as glycogen is associated with elevated glucose uptake in trained athletes (63).
Taken together, our results show that multiple aspects of this phenotype converge with respect to carbohydrate utilization. We have reported improved glucose tolerance (with normal chow), the lack of hyperglycemia (with a high-fat diet), and increased GLUT4 expression in the Tg-Nor-1 mice. These findings are consistent with other studies suggesting that nonoxidative glucose storage, ie, glycogen synthesis, is a major pathway of nonoxidative glucose utilization (64, 65). We hypothesize that the both of the pathways controlling the fate of glucose, ie, storage (glycogen) and utilization (via glycolysis, the TCA cycle, and oxidative phosphorylation), are operating at maximal capacity, which would account for the elevated NADH levels. Furthermore, we observed increases in the expression of genes involved in the malate-aspartate shuttle and a concurrent decrease in the glycerol 3-phosphate shuttle, the pathways responsible for translocation of redox potentials (generated from glycolysis) across the mitochondrial inner membrane for oxidative phosphorylation (Figure 9B). The induction of genes in the malate-aspartate shuttle is consistent with the maximal and most efficient aerobic ATP production. For example, the malate-aspartate shuttle boosts the number of ATPs formed in glycolysis relative to that with the glycerol 3-phosphate shuttle. It has been demonstrated that endurance training increased levels of malate-aspartate shuttle enzymes in human skeletal muscle (66). We suggest that the enhanced metabolic efficiency of the Tg-Nor-1 skeletal muscle may provide the sustained energy resources required for improved endurance performance.
We have previously reported increased mitochondrial numbers and DNA and significantly increased mRNA and protein of oxidative phosphorylation complexes I to V in skeletal muscle from Tg-Nor-1. This finding, coupled to increased glycogen, the endurance phenotype, increased MEF2C expression, increased STARS expression (which is a PGC-1α-dependent target gene increased by endurance and resistance training) (31), and the acquisition of more oxidative type II muscle fibers in the Tg-Nor-1 mice is indicative of increased PGC-1α expression. Although initially we did not find increased Pgc-1α mRNA expression (3), in this study we have identified enhanced PGC-1α protein expression and a decreased NAD+/NADH ratio in skeletal muscle. At the protein level, the 20S gatekeeper molecule NQO1 binds and guards PGC-1α from 20S proteosomal decay in an NADH-dependent manner, hence facilitating PGC-1α protein stability (46). Furthermore, individual analysis of the PGC-1α splice variants has revealed that the PGC-1α1 mRNA species is very abundant and significantly increased in Tg-Nor-1 mice relative to that in WT littermates (fed both the normal chow and high-fat diets), which is in concordance with the endurance phenotype found previously (45, 67). The expression of the PGC-1α2–4 splice variants was approximately 100-fold lower.
In conclusion, muscle-specific Nor-1 expression regulates many genes and pathways that are responsible for adiposity. Furthermore, Nor-1 regulates genes and pathways controlling (1) the metabolic and oxidative capacity of skeletal muscle, (2) glucose utilization and storage, (3) calcineurin signaling and fiber type reprogramming, and (4) endurance. In liver, intact insulin signaling is required for lipogenesis and subsequent hepatosteatosis and hypertriglyceridemia under hyperinsulinemia conditions, which constitutes one arm of the vicious cycle 2 and 3. In the meantime, hepatic insulin signaling is also required for suppression of gluconeogenesis and subsequent glucose output. Therefore, defects in the upstream hepatic insulin signaling cause co-occurrence of glucose intolerance and reduced triglyceride levels in liver and blood, in other words, dissociation of hepatosteatosis from insulin resistance.
The physiological significance of our mouse model is supported by several lines of evidence. First, in a line of rats selected for high running ability, Nor-1 expression is increased by 20% to 30% (68). Second, in humans in response to acute endurance exercise, Nor-1/NR4A3 is induced by 4.5-fold (69). Third, our unpublished data demonstrate that 1 hour of exercise in WT C57BL/6J mice increases Nor-1 mRNA expression by 3- to 4-fold. Finally, the clear parallels between our mouse model, and the human phenotype with the actinin-3-null polymorphism underscore the potential utility of targeting this NR selectively in skeletal muscle.
Materials and Methods
Generation of transgenic mice
All animal-related procedures were approved by the animal experimentation ethics committees of the University of Queensland and conformed to the Guidelines for the Care and Use of Experimental Animals described by the National Health and Medical Research Council of Australia. The VP16-activated Nor-1 mouse line (Tg-Nor-1) has been described previously (3). Mice were maintained on a 12-hour light/dark cycle, with standard mouse chow (Meat Free Rat and Mouse; Specialty Feeds) and water provided ad libitum at the University of Queensland Biological Resources animal house facility. All experiments were performed on male mice of 12 to 17 weeks of age. Experimental mice were backcrossed to the C57BL/6J background for >7 generations, and experiments consisted of heterozygous transgenic mice with WT siblings.
High-fat diet
After weaning at 4 weeks, mice were fed a high-fat diet (SF03–002 36% Fat Modified Rodent Diet; Specialty Feeds). This diet is a very high-fat modification of AIN93G. Mice were weighed until they were 16 weeks old, and experimental work was performed at 16 to 17 weeks of age.
Food intake
Food was measured before and after a 24-hour period with isolated mice to determine intake with the high-fat diet over 24 hours.
Dual-energy X-ray absorptiometry (DXA)
Lean mass (percentage) and fat mass (percentage) were determined by DXA using a PIXImus Densitometer (GE Lunar). Mice were killed and frozen with their limbs splayed apart in a prone position on the imaging tray.
Magnetic resonance imaging (MRI)
Mice were scanned with a 16.4-T Bruker AV700 MRI system using the fast spin echo (rapid acquisition with relaxation enhancement) sequence with the following parameters: slice thickness, 0.8 mm; field of view, 30 × 30 mm; image matrix, 256 × 256; rapid acquisition with relaxation enhancement factor, 4; echo time, 30; repetition time, 9 seconds (60 slices) or 7 seconds (50 slices); and averages, 2. For quantification, transverse sections at the base of the kidney for reference were viewed by a MicroDicom viewer and exported to Adobe Photoshop. Adipose tissue was manually selected and colorized by hand. ImageJ (70) was used to determine the quantity of colorized adipose tissue as a percentage of total body cross-sectional area.
Leptin concentration
The leptin concentration was measured in plasma from overnight fasted mice as part of the MILLIPLEX Mouse Serum Adipokine Kit (MADPK-71K; Millipore).
RNA extraction, cDNA synthesis, and RT-qPCR
WT and Tg-Nor-1 mice were killed with carbon dioxide or cervical fracture with organs and tissues surgically excised. All samples were snap-frozen in liquid nitrogen. RNA and cDNA synthesis was performed as described previously (71, 72). Target cDNA were compared by RT-qPCR in 25-μL reactions as described previously (71). Expression levels were normalized to control genes as stated. A TLDA platform was utilized as described previously (3) in a custom designed format.
Primers
Primers for analysis of cDNA populations using SYBR Green were described in detail for Stars, Cox4, Cpt1b, Srf, Mrtf-1a, and Mrtf-1b (31, 32).
Western blot analysis
Western Blot analysis was performed as described previously (7). Blots were probed with anti-GAPDH (2275-PC-100, 1:20 000; Trevigen), anti-GLUT4 (Ab62375, 1:5000; Abcam), anti-AKT (9272, 1:2000; Cell Signaling), anti-pAKT (Ser473, 4058S, 1:2000; Cell Signaling), anti-pAKT (Thr308, 4056S, 1:2000; Cell Signaling), anti-Stbd1 (11842–1-AP, 1:1000; Proteintech), anti-AMPKα (2532, 1:1000; Cell Signaling), anti-pAMPKα (Thr172, 2535S, 1:1000; Cell Signaling), anti-ACC (3662, 1:1000; Cell Signaling), anti-pACC (Ser79, 3661S, 1:1000; Cell Signaling), anti-CREB (9192, 1:1000; Cell Signaling); anti-pCREB (Ser133, 9198S, 1:1000; Cell Signaling); GCN5 (Ab18381, 1:500; Abcam), anti-PPP1R1A (clone EP902Y) (TA300665, 1:100 000; OriGene), anti-SIRT1 (3931S, 1:1000; Cell Signaling), and anti-PGC-1α (Ab54481, 1:1000; Abcam) in Tris-buffered saline with 5% skim milk powder (or 1% BSA for anti-phosphoproteins). Secondary anti-rabbit/mouse horseradish peroxidase (Pierce Biotechnology) was used at 1:5000 to 1:10 000 in 5% skim milk powder. Densitometry was performed using ImageJ (70).
Insulin injection for Western blot analysis
Sixteen- to 17-week-old mice fed a high-fat diet were fasted for 6 hours and given an ip injection of insulin (Actrapid; Novo Nordisk) at 1 IU/kg of body weight.
Histological analyses
Tissues were extracted and placed in formaldehyde overnight. Samples were then processed by a Leica tissue processor and embedded in wax. Samples were cut in 7-μm sections, deparaffinized, and rehydrated. Samples were then stained accordingly following standard hematoxylin and eosin or PAS staining procedures. Samples were dehydrated and mounted on slides with DPX mounting media. Images were taken with an Olympus BX-51 bright field/fluorescence microscope.
NAD/NADH measurements
NAD and NADH levels were measured using an NAD/NADH assay kit (ab65348; Abcam) as per the manufacturer's protocol. Cell lysates were filtered using a Millipore Amicon Ultra-0.5 3K centrifugal filter at 4°C to remove enzymes that degrade NADH.
Glycogen measurements
A glycogen colorimetric/fluorometric assay kit (BIO-K646–100; BioVision, Inc) was used as per the manufacturer's protocol.
Insulin measurements
Blood was extracted via cardiac puncture, and then the insulin concentration was analyzed using a mouse ultrasensitive insulin ELISA (80-INSMSU-E01; Alpco) as per the manufacturer's protocol.
Oxygen consumption and heat production
Oxygen consumption was measured using the CLAMS (Columbus Instruments). Mice were acclimated for 1 day and than were measured over the following night and day cycle.
Statistical analysis
Statistical analyses for all non-TLDA RT-qPCR data were performed using Prism 6 (GraphPad Software Inc). Non-TLDA data were analyzed using a Student t test unless otherwise stated. All results are expressed as means ± SEM. Significant changes in expression of TLDA data were analyzed using the StatMiner software package (ABI/Integromics) as described previously (16, 55, 72). Differentially expressed genes were identified by linear models (contained in the limma package for Bioconductor R embedded in StatMiner). Significance was assigned by application of the empirical Bayes statistic, described as equivalent to shrinkage of the estimated sample variances toward a pooled estimate, resulting in a far more stable inference when the number of arrays is small (73). It returns the empirical Bayes log odds of differential expression (ie, the probability) that a gene is differentially expressed (a higher score represents a more significant result). For example, a B statistic of 0 indicates a 50:50 chance of differential expression, B scores >0 indicate a >50:50 chance of differential expression, B scores <0 (negative scores) reflect odds that a gene is more than likely not differentially expressed. The B statistic considers and ranks a proportion of differentially expressed genes (P < .01). Analysis also includes t, the empirical Bayes moderated t statistic (a variant t test), an empirically moderated estimate of SE. Data are presented as relative quantification, ie, the calculated fold differences normalized to one of the selected controls (for example, 18S rRNA) between the transgenic and the WT littermates. Relative quantification (RQ) was determined using the equation RQ = 2−ΔΔCt (74).
Illumina BeadArray hybridization and statistical analysis
For BeadArray analysis, 8 mice in total were analyzed with 4 WT and 4 Tg-Nor-1 mice. Total RNA (500 ng) was amplified using the Illumina TotalPrep RNA amplification kit (Ambion) with an in vitro transcription reaction period of 12 hours. Biotinylated, amplified cRNA was assessed for quantity and quality using the a 2100 Bioanalyzer (Agilent). Then 1500 ng of amplified cRNA per array was hybridized to Illumina MouseRef-6 BeadChip arrays according to the manufacturer's directions. Hybridized BeadChip arrays were stained with Amersham fluorolink streptavidin-Cy3 (GE Healthcare). BeadChip arrays were scanned with an Illumina BeadStation scanner, and data values with detection scores were compiled using BeadStudio v2.3.41 (Illumina) and imported into GeneSpring GX (Agilent) for data analysis. A mouse Illumina probe set was defined in the GeneSpring Workgroup using the Illumina TargetIDs as the unique identifiers and annotated according to array content files supplied by Illumina. Normalized data were produced using GeneSpring GX via normalization to control genes, where control genes were represented by all genes with an Illumina detection score equal to 1 in at least 4 of the 8 samples. All probes except for probes that were determined to have an Illumina detection score equal to 1 in at least 4 of the 8 samples were filtered out to remove probes without adequate expression levels. A parametric Welch t test (where variances were not assumed equal) was performed on the probes with expression with a P value cutoff of .05. Multiple testing correction (Benjamini and Hochberg false discovery rate) was then applied to genes that had passed the parametric Welch t test. About 5.0% of the identified probes would be expected to pass the restriction by chance.
Ingenuity pathway analysis
The canonical pathways and functions functional analyses were generated through the use of IPA (Ingenuity Systems) from Illumina gene expression data.
Glucose and insulin tolerance testing
These procedures were performed as described previously (75) on 16- to 17-week-old mice fed the high-fat diet as described above. Mice were fasted overnight for glucose tolerance tests and 6 hours for insulin tolerance tests. Glucose was used at 2 g/kg of body weight, whereas insulin was used at a standard dose of 0.0321 IU per mouse.
Pyruvate tolerance testing
After overnight fasting, 14- to 16-week-old mice were administered 1.5 g/kg sodium pyruvate. Glucose was measured as described previously (75).
Measurement of hepatic triglyceride concentration
To obtain the hepatic triglyceride concentration, frozen liver tissue was saponified by digestion with ethanolic KOH, neutralized, and then assayed using Free Glycerol Reagent (Sigma-Aldrich).
Acknowledgments
This work was supported by a research project grant from the National Health and Medical Research Council (NHMRC) of Australia. G.E.O.M. is a University of Queensland, Vice Chancellor's Senior Research Fellow of the NHMRC, and Natalie Eriksson and Joel Goode are recipients of an Australian Postgraduate Awards.
Disclosure Summary: The authors have nothing to disclose.
Funding Statement
This work was supported by a research project grant from the National Health and Medical Research Council (NHMRC) of Australia. G.E.O.M. is a University of Queensland, Vice Chancellor's Senior Research Fellow of the NHMRC, and Natalie Eriksson and Joel Goode are recipients of an Australian Postgraduate Awards.
Footnotes
- ABRA
- actin-binding Rho-activating protein
- CLAMS
- Comprehensive Lab Animal Monitoring System
- DXA
- dual-energy X-ray absorptiometry
- MHC
- myosin heavy chain
- MRI
- magnetic resonance imaging
- Nor-1
- neuron-derived orphan receptor 1
- NR
- nuclear receptor
- Nur77
- neuron-derived clone 77
- Nurr1
- nuclear receptor related 1
- PAS
- periodic acid-Schiff
- PCG-1
- peroxisome proliferator-activated receptor γ coactivator-1
- qPCR
- quantitative PCR
- RER
- respiratory exchange ratio
- STARS
- striated muscle activator of Rho signaling
- TCA
- tricarboxylic acid
- Tg
- transgenic
- TLDA
- TaqMan low-density array
- WT
- wild-type.
References
- 1. Owen OE, Reichard GA Jr, Boden G, Patel MS, Trapp VE. Interrelationships among key tissues in the utilization of metabolic substrate. In Katzen HE, Mahler RJ, eds. Advances in Modern Nutrition. New York, NY: John Wiley & Sons; 1978:517–550. [Google Scholar]
- 2. Zurlo F, Larson K, Bogardus C, Ravussin E. Skeletal muscle metabolism is a major determinant of resting energy expenditure. J Clin Invest. 1990;86:1423–1427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Pearen MA, Eriksson NA, Fitzsimmons RL, et al. The nuclear receptor, Nor-1, markedly increases type II oxidative muscle fibers and resistance to fatigue. Mol Endocrinol. 2012;26:372–384. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Chao LC, Wroblewski K, Zhang Z, et al. Insulin resistance and altered systemic glucose metabolism in mice lacking Nur77. Diabetes. 2009;58:2788–2796. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Chao LC, Wroblewski K, Ilkayeva OR, et al. Skeletal muscle Nur77 expression enhances oxidative metabolism and substrate utilization. J Lipid Res. 2012;53:2610–2619. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Kanzleiter T, Preston E, Wilks D, et al. Overexpression of the orphan receptor Nur77 alters glucose metabolism in rat muscle cells and rat muscle in vivo. Diabetologia. 2010;53:1174–1183. [DOI] [PubMed] [Google Scholar]
- 7. Pearen MA, Myers SA, Raichur S, Ryall JG, Lynch GS, Muscat GE. The orphan nuclear receptor, NOR-1, a target of β-adrenergic signaling, regulates gene expression that controls oxidative metabolism in skeletal muscle. Endocrinology. 2008;149:2853–2865. [DOI] [PubMed] [Google Scholar]
- 8. Maxwell MA, Cleasby ME, Harding A, Stark A, Cooney GJ, Muscat GE. Nur77 regulates lipolysis in skeletal muscle cells. Evidence for cross-talk between the β-adrenergic and an orphan nuclear hormone receptor pathway. J Biol Chem. 2005;280:12573–12584. [DOI] [PubMed] [Google Scholar]
- 9. Chao LC, Zhang Z, Pei L, Saito T, Tontonoz P, Pilch PF. Nur77 coordinately regulates expression of genes linked to glucose metabolism in skeletal muscle. Mol Endocrinol. 2007;21:2152–2163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Kanzleiter T, Schneider T, Walter I, et al. Evidence for Nr4a1 as a cold-induced effector of brown fat thermogenesis. Physiol Genomics. 2005;24:37–44. [DOI] [PubMed] [Google Scholar]
- 11. Kumar N, Liu D, Wang H, Robidoux J, Collins S. Orphan nuclear receptor NOR-1 enhances 3′,5′-cyclic adenosine 5′-monophosphate-dependent uncoupling protein-1 gene transcription. Mol Endocrinol. 2008;22:1057–1064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Roche E, Buteau J, Aniento I, Reig JA, Soria B, Prentki M. Palmitate and oleate induce the immediate-early response genes c-fos and nur-77 in the pancreatic β-cell line INS-1. Diabetes. 1999;48:2007–2014. [DOI] [PubMed] [Google Scholar]
- 13. Susini S, Roche E, Prentki M, Schlegel W. Glucose and glucoincretin peptides synergize to induce c-fos, c-jun, junB, zif-268, and nur-77 gene expression in pancreatic β(INS-1) cells. FASEB J. 1998;12:1173–1182. [PubMed] [Google Scholar]
- 14. Wu X, Wang J, Cui X, et al. The effect of insulin on expression of genes and biochemical pathways in human skeletal muscle. Endocrine. 2007;31:5–17. [DOI] [PubMed] [Google Scholar]
- 15. Rius J, Martínez-Gonzalez J, Crespo J, Badimon L. Involvement of neuron-derived orphan receptor-1 (NOR-1) in LDL-induced mitogenic stimulus in vascular smooth muscle cells: role of CREB. Arterioscler Thromb Vasc Biol. 2004;24:697–702. [DOI] [PubMed] [Google Scholar]
- 16. Wang SC, Myers SA, Eriksson NA, Fitzsimmons RL, Muscat GE. Nr4a1 siRNA expression attenuates α-MSH regulated gene expression in 3T3–L1 adipocytes. Mol Endocrinol. 2011;25:291–306. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Fu Y, Luo L, Luo N, Zhu X, Garvey WT. NR4A orphan nuclear receptors modulate insulin action and the glucose transport system: potential role in insulin resistance. J Biol Chem. 2007;282:31525–31533. [DOI] [PubMed] [Google Scholar]
- 18. Wansa KD, Harris JM, Yan G, Ordentlich P, Muscat GE. The AF-1 domain of the orphan nuclear receptor NOR-1 mediates trans-activation, coactivator recruitment, and activation by the purine anti-metabolite 6-mercaptopurine. J Biol Chem. 2003;278:24776–24790. [DOI] [PubMed] [Google Scholar]
- 19. Kagaya S, Ohkura N, Tsukada T, et al. Prostaglandin A2 acts as a transactivator for NOR1 (NR4A3) within the nuclear receptor superfamily. Biol Pharm Bull. 2005;28:1603–1607. [DOI] [PubMed] [Google Scholar]
- 20. Veum VL, Dankel SN, Gjerde J, et al. The nuclear receptors NUR77, NURR1 and NOR1 in obesity and during fat loss. Int J Obes (Lond). 2012;36:1195–1202. [DOI] [PubMed] [Google Scholar]
- 21. Wang YX, Zhang CL, Yu RT, et al. Regulation of muscle fiber type and running endurance by PPARδ. PLoS Biol. 2004;2:e294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Hakimi P, Yang J, Casadesus G, et al. Overexpression of the cytosolic form of phosphoenolpyruvate carboxykinase (GTP) in skeletal muscle repatterns energy metabolism in the mouse. J Biol Chem. 2007;282:32844–32855. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Wenz T, Rossi SG, Rotundo RL, Spiegelman BM, Moraes CT. Increased muscle PGC-1α expression protects from sarcopenia and metabolic disease during aging. Proc Natl Acad Sci USA. 2009;106:20405–20410. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 24. Yang N, MacArthur DG, Gulbin JP, et al. ACTN3 genotype is associated with human elite athletic performance. Am J Hum Genet. 2003;73:627–631. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Eynon N, Duarte JA, Oliveira J, et al. ACTN3 R577X polymorphism and Israeli top-level athletes. Int J Sports Med. 2009;30:695–698. [DOI] [PubMed] [Google Scholar]
- 26. Quinlan KG, Seto JT, Turner N, et al. α-Actinin-3 deficiency results in reduced glycogen phosphorylase activity and altered calcium handling in skeletal muscle. Hum Mol Genet. 2010;19:1335–1346. [DOI] [PubMed] [Google Scholar]
- 27. MacArthur DG, Seto JT, Raftery JM, et al. Loss of ACTN3 gene function alters mouse muscle metabolism and shows evidence of positive selection in humans. Nat Genet. 2007;39:1261–1265. [DOI] [PubMed] [Google Scholar]
- 28. MacArthur DG, Seto JT, Chan S, et al. An Actn3 knockout mouse provides mechanistic insights into the association between α-actinin-3 deficiency and human athletic performance. Hum Mol Genet. 2008;17:1076–1086. [DOI] [PubMed] [Google Scholar]
- 29. Vissing K, Rahbek SK, Lamon S, et al. Effect of resistance exercise contraction mode and protein supplementation on members of the STARS signalling pathway. J Physiol. 2013;591:3749–3763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Lamon S, Wallace MA, Léger B, Russell AP. Regulation of STARS and its downstream targets suggest a novel pathway involved in human skeletal muscle hypertrophy and atrophy. J Physiol. 2009;587:1795–1803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Wallace MA, Hock MB, Hazen BC, Kralli A, Snow RJ, Russell AP. Striated muscle activator of Rho signalling (STARS) is a PGC-1α/oestrogen-related receptor-α target gene and is upregulated in human skeletal muscle after endurance exercise. J Physiol. 2011;589:2027–2039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Wallace MA, Russell AP. Striated muscle activator of Rho signaling is required for myotube survival but does not influence basal protein synthesis or degradation. Am J Physiol Cell Physiol. 2013;305:C414–C426. [DOI] [PubMed] [Google Scholar]
- 33. Frey N, Barrientos T, Shelton JM, et al. Mice lacking calsarcin-1 are sensitized to calcineurin signaling and show accelerated cardiomyopathy in response to pathological biomechanical stress. Nat Med. 2004;10:1336–1343. [DOI] [PubMed] [Google Scholar]
- 34. Frey N, Frank D, Lippl S, et al. Calsarcin-2 deficiency increases exercise capacity in mice through calcineurin/NFAT activation. J Clin Invest. 2008;118(11):3598–3608. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Berman Y, North KN. A gene for speed: the emerging role of α-actinin-3 in muscle metabolism. Physiology (Bethesda). 2010;25:250–259. [DOI] [PubMed] [Google Scholar]
- 36. Lee S, Yang WK, Song JH, et al. Anti-obesity effects of 3-hydroxychromone derivative, a novel small-molecule inhibitor of glycogen synthase kinase-3. Biochem Pharmacol. 2013;85:965–976. [DOI] [PubMed] [Google Scholar]
- 37. Ryan AS, Ortmeyer HK, Sorkin JD. Exercise with calorie restriction improves insulin sensitivity and glycogen synthase activity in obese postmenopausal women with impaired glucose tolerance. Am J Physiol Endocrinol Metab. 2012;302:E145–E152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Printz RL, Koch S, Potter LR, et al. Hexokinase II mRNA and gene structure, regulation by insulin, and evolution. J Biol Chem. 1993;268:5209–5219. [PubMed] [Google Scholar]
- 39. Rose AJ, Richter EA. Skeletal muscle glucose uptake during exercise: how is it regulated? Physiology (Bethesda). 2005;20:260–270. [DOI] [PubMed] [Google Scholar]
- 40. Fueger PT. Glucose phosphorylation as a barrier to muscle glucose uptake. Clin Exp Pharmacol Physiol. 2005;32:314–318. [DOI] [PubMed] [Google Scholar]
- 41. Wasserman DH, Kang L, Ayala JE, Fueger PT, Lee-Young RS. The physiological regulation of glucose flux into muscle in vivo. J Exp Biol. 2011;214:254–262. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Fueger PT, Shearer J, Krueger TM, et al. Hexokinase II protein content is a determinant of exercise endurance capacity in the mouse. J Physiol. 2005;566:533–541. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Irimia JM, Rovira J, Nielsen JN, Guerrero M, Wojtaszewski JF, Cusso R. Hexokinase 2, glycogen synthase and phosphorylase play a key role in muscle glycogen supercompensation. PloS One. 2012;7:e42453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Sahlin K, Katz A, Henriksson J. Redox state and lactate accumulation in human skeletal muscle during dynamic exercise. Biochem J. 1987;245:551–556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Ruas JL, White JP, Rao RR, et al. A PGC-1α isoform induced by resistance training regulates skeletal muscle hypertrophy. Cell. 2012;151:1319–1331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Adamovich Y, Shlomai A, Tsvetkov P, et al. The protein level of PGC-1α, a key metabolic regulator, is controlled by NADH-NQO1. Mol Cell Biol. 2013;33:2603–2613. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Lerin C, Rodgers JT, Kalume DE, Kim SH, Pandey A, Puigserver P. GCN5 acetyltransferase complex controls glucose metabolism through transcriptional repression of PGC-1α. Cell Metab. 2006;3:429–438. [DOI] [PubMed] [Google Scholar]
- 48. Nemoto S, Fergusson MM, Finkel T. SIRT1 functionally interacts with the metabolic regulator and transcriptional coactivator PGC-1α. J Biol Chem. 2005;280:16456–16460. [DOI] [PubMed] [Google Scholar]
- 49. Viollet B, Foretz M, Guigas B, et al. Activation of AMP-activated protein kinase in the liver: a new strategy for the management of metabolic hepatic disorders. J Physiol. 2006;574:41–53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Blaak EE, Van Baak MA, Kemerink GJ, Pakbiers MT, Heidendal GA, Saris WH. β-Adrenergic stimulation of energy expenditure and forearm skeletal muscle metabolism in lean and obese men. Am J Physiol. 1994;267:E306–E315. [DOI] [PubMed] [Google Scholar]
- 51. Hagström-Toft E, Enoksson S, Moberg E, Bolinder J, Arner P. β-Adrenergic regulation of lipolysis and blood flow in human skeletal muscle in vivo. Am J Physiol. 1998;275:E909–E916. [DOI] [PubMed] [Google Scholar]
- 52. Blaak EE. Basic disturbances in skeletal muscle fatty acid metabolism in obesity and type 2 diabetes mellitus. Proc Nutr Soc. 2004;63:323–330. [DOI] [PubMed] [Google Scholar]
- 53. Fagher B, Liedholm H, Monti M, Moritz U. Thermogenesis in human skeletal muscle as measured by direct microcalorimetry and muscle contractile performance during β-adrenoceptor blockade. Clin Sci (Lond). 1986;70:435–441. [DOI] [PubMed] [Google Scholar]
- 54. Pearen MA, Ryall JG, Maxwell MA, Ohkura N, Lynch GS, Muscat GE. The orphan nuclear receptor, NOR-1, is a target of β-adrenergic signaling in skeletal muscle. Endocrinology. 2006;147:5217–5227. [DOI] [PubMed] [Google Scholar]
- 55. Myers SA, Eriksson N, Burow R, Wang SC, Muscat GE. β-Adrenergic signaling regulates NR4A nuclear receptor and metabolic gene expression in multiple tissues. Mol Cell Endocrinol. 2009;309:101–108. [DOI] [PubMed] [Google Scholar]
- 56. Zhu X, Walton RG, Tian L, et al. Prostaglandin A2 enhances cellular insulin sensitivity via a mechanism that involves the orphan nuclear receptor NR4A3. Horm Metab Res. 2013;45:213–220. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Rangwala SM, Wang X, Calvo JA, et al. Estrogen-related receptor γ is a key regulator of muscle mitochondrial activity and oxidative capacity. J Biol Chem. 2010;285:22619–22629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Seto JT, Lek M, Quinlan KG, et al. Deficiency of α-actinin-3 is associated with increased susceptibility to contraction-induced damage and skeletal muscle remodeling. Hum Mol Genet. 2011;20:2914–2927. [DOI] [PubMed] [Google Scholar]
- 59. Krause MP, Moradi J, Coleman SK, et al. A novel GFP reporter mouse reveals Mustn1 expression in adult regenerating skeletal muscle, activated satellite cells and differentiating myoblasts. Acta Physiol (Oxf). 2013;208:180–190. [DOI] [PubMed] [Google Scholar]
- 60. Kostek MC, Chen YW, Cuthbertson DJ, et al. Gene expression responses over 24 h to lengthening and shortening contractions in human muscle: major changes in CSRP3, MUSTN1, SIX1, and FBXO32. Physiol Genomics. 2007;31:42–52. [DOI] [PubMed] [Google Scholar]
- 61. Knoll R, Buyandelger B, Lab M. The sarcomeric Z-disc and Z-discopathies. J Biomed Biotechnol. 2011;2011:569628. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62. Frey N, Richardson JA, Olson EN. Calsarcins, a novel family of sarcomeric calcineurin-binding proteins. Proc Natl Acad Sci USA. 2000;97:14632–14637. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63. Phielix E, Meex R, Ouwens DM, et al. High oxidative capacity due to chronic exercise training attenuates lipid-induced insulin resistance. Diabetes. 2012;61:2472–2478. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64. Shulman GI, Rothman DL, Jue T, Stein P, DeFronzo RA, Shulman RG. Quantitation of muscle glycogen synthesis in normal subjects and subjects with non-insulin-dependent diabetes by 13C nuclear magnetic resonance spectroscopy. N Engl J Med. 1990;322:223–228. [DOI] [PubMed] [Google Scholar]
- 65. Højlund K, Staehr P, Hansen BF, et al. Increased phosphorylation of skeletal muscle glycogen synthase at NH2-terminal sites during physiological hyperinsulinemia in type 2 diabetes. Diabetes. 2003;52:1393–1402. [DOI] [PubMed] [Google Scholar]
- 66. Schantz PG, Sjöberg B, Svedenhag J. Malate-aspartate and α-glycerophosphate shuttle enzyme levels in human skeletal muscle: methodological considerations and effect of endurance training. Acta Physiol Scand. 1986;128:397–407. [DOI] [PubMed] [Google Scholar]
- 67. Millay DP, Olson EN. Making muscle or mitochondria by selective splicing of PGC-1α. Cell Metab. 2013;17:3–4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68. Stephenson EJ, Stepto NK, Koch LG, Britton SL, Hawley JA. Divergent skeletal muscle respiratory capacities in rats artificially selected for high and low running ability: a role for Nor1? J Appl Physiol. 2012;113:1403–1412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69. Catoire M, Mensink M, Boekschoten MV, et al. Pronounced effects of acute endurance exercise on gene expression in resting and exercising human skeletal muscle. PloS One. 2012;7:e51066. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70. Abramoff MD, Magalhaes PJ, Ram SJ. Image processing with ImageJ. Biophoton Int. 2004;11:36–42. [Google Scholar]
- 71. Pearen MA, Ryall JG, Lynch GS, Muscat GE. Expression profiling of skeletal muscle following acute and chronic β2-adrenergic stimulation: implications for hypertrophy, metabolism and circadian rhythm. BMC Genomics. 2009;10:448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72. Crowther LM, Wang SC, Eriksson NA, Myers SA, Murray LA, Muscat GE. Chicken ovalbumin upstream promoter-transcription factor II regulates nuclear receptor, myogenic, and metabolic gene expression in skeletal muscle cells. Physiol Genomics. 2011;43:213–227. [DOI] [PubMed] [Google Scholar]
- 73. Smyth GK. Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol. 2004;3:Article3. [DOI] [PubMed] [Google Scholar]
- 74. Goni R, Garcia P, Foissac ST. The qPCR data statistical analysis. Integromics White Paper. 2009; Integromics SL:1–9. [Google Scholar]
- 75. Raichur S, Fitzsimmons RL, Myers SA, et al. Identification and validation of the pathways and functions regulated by the orphan nuclear receptor, ROR alpha1, in skeletal muscle. Nucleic Acids Res. 2010;38:4296–4312. [DOI] [PMC free article] [PubMed] [Google Scholar]