Abstract
The hidden Markov model (HMM) has been a workhorse of single-molecule data analysis and is now commonly used as a stand-alone tool in time series analysis or in conjunction with other analysis methods such as tracking. Here, we provide a conceptual introduction to an important generalization of the HMM, which is poised to have a deep impact across the field of biophysics: the infinite HMM (iHMM). As a modeling tool, iHMMs can analyze sequential data without a priori setting a specific number of states as required for the traditional (finite) HMM. Although the current literature on the iHMM is primarily intended for audiences in statistics, the idea is powerful and the iHMM’s breadth in applicability outside machine learning and data science warrants a careful exposition. Here, we explain the key ideas underlying the iHMM, with a special emphasis on implementation, and provide a description of a code we are making freely available. In a companion article, we provide an important extension of the iHMM to accommodate complications such as drift.
Main Text
Hidden Markov models (HMMs) (1, 2) provide a method for analyzing sequential time series data and have been a workhorse across fields including biology (3, 4), physics (5, 6, 7), and engineering (8, 9, 10).
The power of HMMs has been heavily exploited in biophysics (11, 12) in the interpretation of single-molecule experiments such as fluorescence resonance energy transfer (FRET) (13), force spectroscopy (14), atomic force microscopy (15), and ion-channel patch clamp (16). The data from these disparate techniques can be analyzed using HMMs because biomolecules or collections of biomolecules are seen as visiting discrete states and the signal emitted from each state is corrupted by noise (see Fig. 1).
Figure 1.
A synthetic time trace illustrating measurements of a hypothetical biomolecule that undergoes conformational transitions. (Left) The state space consists of conformations depicted discretely as . (Middle) Time series of noisy observations, , produced by the biomolecule (blue) and the corresponding noiseless trace (red). Over the time course of the measurements, the biomolecule attains only conformations –, though additional conformations might be visited at subsequent times. For the sake of concreteness only, we label these states in order of appearance from 1 through 5. (Right) Binning the collected observations reveals “emission distributions,” , associated with each conformation. These distributions are highlighted with red lines. The centers (mean values) of the emission distributions are used to obtain the noiseless trace in the middle panel. The illustration on the left is created using data from (47) (PDB: 2N4G). To see this figure in color, go online.
Although HMMs have been hugely successful, the method has encountered a fundamental limitation, which is that the number of different states visited over the course of an experiment must be specified in advance (13). This limitation is too restrictive for complex biomolecules, where state numbers may often be difficult to assess a priori. It also presents a problem when the number of states appears to vary from trace to trace, as would be expected if some states are only rarely visited. As a work-around, problem- and user-dependent pre- or post-processing steps have therefore been suggested in the biophysical literature to winnow down the number of candidate models or to eliminate problematic traces altogether. Popular choices include model-selection tools such as information criteria (e.g., Bayesian and Akaike information criteria (11, 17)) or maximum evidence methods (e.g., (12)). However, thanks to recent mathematics—Bayesian nonparametrics, first introduced in 1973 (18)—an elegant solution is available that largely circumvents these model-selection steps.
In this perspectives article, we provide a description of the concepts and implementation of an important, and to our knowledge new, computational tool that exploits Bayesian nonparametrics: the infinite HMM (iHMM), which was heuristically described in (19) but only fully realized in 2012 (20).
It has recently been suggested that Bayesian nonparametrics are poised to have a deep impact in biophysics (21), and in this field, they have just begun to be exploited (22, 23, 24). However, there is, to our knowledge, no single resource yet available that describes the iHMM, its concepts, or its implementation, as would be required to bring the power of Bayesian nonparametrics to bear on biophysics and accelerate its inevitable widespread adoption. Indeed, although the iHMM tackles a conceptually simple problem, it relies on mathematics, whose literature is inaccessible outside a rarified community of statisticians and computer scientists.
It is for this reason that we have organized our perspective article as follows: Formulation of the iHMM describes the structure of the HMM in its finite (traditional) and infinite (nonparametric) realizations; Inference on the iHMM presents a computational algorithm to perform inference with the iHMM that we make freely available; Results shows results from sample time traces; and Perspectives discusses the potential for further applications to biophysics.
Methods
Formulation of the iHMM
Here, we introduce the HMM and its generalization, the iHMM. To facilitate the presentation, we initially describe the structure of the space within which the biomolecule evolves and subsequently formulate the dynamics. This discussion applies to both HMMs and iHMMs.
In the HMM framework, a system of interest is assumed to alternate successively between different states labeled , where k takes integer values from 1 to some L. We use L to denote the total number of states available to the system, no matter whether all these states are visited or some remain unvisited during the time course of the measurements. For instance, for an experiment on a single protein, the protein is the “system” and the “states” are conformations, such as open/closed conformations of an ion channel (16) or low/high-efficiency states in FRET experiments (25). For an illustration see Fig. 1 (left).
In the standard HMM, we assume that the system’s transitions are governed by Markovian dynamics (1). This means that the system jumps from a state to a state , for example, from one FRET value to another, in a stochastic manner that depends exclusively on and not any other state visited in the past. For this reason, all transitions out of state are fully described by a probability vector , where is the probability of departing from state and arriving at .
A note on our notation is appropriate here. Throughout this survey we adopt the tilde to denote vectors with components over , the system’s state space , such as . Shortly, we will adopt bars to denote vectors with components over time.
Once the system reaches a state , observations are emitted stochastically according to a probability distribution unique to (see Fig. 1, right). We call this the “emission distribution,” . It is often practical to model the emission distributions by a general family and use state-specific parameters, , to distinguish its members. For example, to model Gaussian emissions x, as in Fig. 1, we may choose
| (1) |
where stands for the mean, , and standard deviation (width), , of the observations produced by state . Therefore, for example, we define , , etc., where , are the state FRET efficiencies and , , …) are the corresponding spreads attributed to noise.
Next, we let denote the state of the system at the nth time step of the experiment and its corresponding observation. Thus, we label with n the states of the system as it evolves through time, forming a sequence , with each being equal to some state chosen from . For completeness, we can also assume that, before the first measurement, the system is at a default state, which we denote as . Again, we mention that might contain states that do not appear in this sequence, i.e., states that remain unvisited throughout the experimental time course.
As is common in the statistical literature—and especially since we will borrow from this notation to introduce the iHMM—we express the HMM compactly using the scheme
| (2) |
| (3) |
and depict it schematically in Fig. 2. The ∼ symbol used above indicates that the random variables on the left side of the scheme are “distributed according to”, and therefore sampled from, the probability distributions shown on the right side (26). For example, Eq. 2 denotes a sampling from a categorical probability distribution that is supported on , i.e., equals a state that is taken from with probability .
Figure 2.
Graphical representation of the HMM. In the HMM, a biomolecule of interest transitions between unobserved states according to the probability vectors and generates observations according to the probability distributions that depend on the parameter . Here, following convention, the values are shaded to denote that these quantities are observed, whereas the values are hidden. Arrows denote the dependences among the model variables and red lines denote the model parameters. To see this figure in color, go online.
As can be seen from Eqs. 2 and 3, the HMM models the experimental output in a doubly stochastic manner: 1) the state of the system evolves stochastically within , as expressed by Eq. 2; and 2) the observations from each are also emitted stochastically, as expressed by Eq. 3. In particular, for single-molecule experiments, these two characteristics allow the HMM to elegantly capture 1) the seemingly random biomolecular state switching; and 2) the noise corrupting the measurements (13, 27).
From the experimental measurements, we gain access only to the observations , which take the form of a time series, , over some regularly spaced time intervals . In the most general case, the goals of the HMM are to estimate 1) the underlying state sequence, , which is unobserved during the measurements; and 2) the model parameters, which include of the emission distributions and the state transition probabilities, , associated with the states in . Of course, in practice, these goals are feasible only for those states that are visited during the experimental time course.
Normally, this is the extent of the HMM. That is, one fixes L, i.e., the size of the state space , writes down the likelihood of observing the sequence of observations, , and maximizes this likelihood to estimate the quantities of interest. Extensive literature—that has made its way into standard textbooks, for example (26)—explains each of these steps in detail.
However, in preparation for the iHMM, we take a Bayesian route to accomplish the goals of the HMM (21). This is a computational overkill for some applications but otherwise essential to its nonparametric generalization.
Following the Bayesian paradigm (21, 28), we assign prior probabilities to model parameters including the emission parameters, , and transition probability vectors, . For instance, we may assume a prior given by a common distribution for all states. However, assigning a prior is more subtle, as any choice must ensure that the predicted transitions stay within the system’s state space, . It is specifically the formulation of this prior that fundamentally distinguishes the finite from the infinite variants of the HMM (20), both of which we describe next.
In the finite variant of the HMM (1, 2), before setting the prior on , we must fix L, the total number of states in , or, in the language of single-molecule biophysics, conformations available to the biomolecule. Once L is fixed, the symmetric Dirichlet distribution is a common choice:
| (4) |
Here, denotes the Dirichlet distribution supported in with concentration parameter α (for an explicit definition of the Dirichlet distribution, see the Supporting Material). Basically, this prior asserts that lacking any information on the system’s kinetics, the model is likely to place on average equal probability on any possible transition . In this prior, the value of α can be used to influence the transitions departing each state. For example, with , the prior tends to favor uniform transitions, whereas with , the prior tends to favor sparse transitions. In other words, the resulting sequence, , tends to contain several different states for , whereas it tends to contain fewer for .
The Dirichlet prior in Eq. 4 offers two key advantages: 1) it provides a noninformative prior, since no specific transitions between the L states are preferentially selected; and 2) it combines well with (i.e., is conjugate to) the categorical distribution of Eq. 2, which greatly simplifies model parameter estimation.
Although such priors help lessen the computational burden, pre-setting L ignores the data, and thus, the arbitrary choice of L may cause under- or over-fitting, which, in turn, has far-reaching consequences in estimating state kinetics. Pre-setting L also does not allow the model’s complexity to grow in response to newly available data (e.g., a rare state visited later or in another time series). Resolving the latter issue in a principled fashion can also help avoid cherry-picking data sets that behave more closely to one’s expected or preferred value of L. It is to resolve such issues that the iHMM was developed in the first place.
Now, in the infinite HMM, the key difference is that is assumed to be infinite in size (19). To avoid any confusion, we point out that this assumption is different from forcing the system to visit an infinite number of states, as it might appear at first. Regardless of the size of , the system visits only K different states, where K cannot exceed, for example, the number of steps, N, in the collected time series. In practice, of course, we expect K to be considerably smaller than N.
As we will see, with an infinite state space, the iHMM can recruit as many states as necessary and, in doing so, avoids underfitting by growing the state space as new states are visited (through the effect of the prior), but also avoids overfitting by placing more weight on states already visited (through the effect of the likelihood).
The generalization of the Dirichlet distribution appropriate for the infinite sized is the Dirichlet process (DP),
| (5) |
where is the “base distribution” of the DP, which determines how are distributed on average. The notion of a base is only required for the infinite DP and has no direct analog in the finite Dirichlet distribution. By contrast, α is similar to the concentration of the finite HMM, since it controls the sparsity of the transition probability vectors, . For instance, large values of α make the model more likely to choose values that are similar for all states, whereas low values of α make the model more likely to choose values that differ considerably.
The DP is at the basis of much of nonparametric Bayesian inference. It was only in 2012 that a hierarchical DP (HDP) was invoked to construct the iHMM (20). The basic idea is to select a random—albeit discrete—base distribution with members that 1) are infinite in number, 2) attain nonnegative values, and 3) sum to 1. In the HDP, it is common to sample such a base from a Griffiths-Engen-McCloskey distribution realized through a stick-breaking construction (20, 29, 30),
| (6) |
which ensures all these properties. Intuitively, the stick-breaking above is obtained as follows. We start with a stick of length 1. We break the stick at some location and use the broken fraction for the probability of visiting ; thus, . Subsequently, we break the remaining stick at and use that new fraction for the probability of visiting ; thus, , and so on. In this stick-breaking, the locations are sampled from a β-distribution, , so γ determines the relative location of the break point. Thus, γ can be used to set how rapidly decreases toward zero and ultimately controls the number of states in with an appreciable likelihood of being visited at least once during the experiment.
In summary, the iHMM—together with its priors illustrated in Fig. 3—takes the form
| (7) |
| (8) |
| (9) |
| (10) |
| (11) |
where, for notational simplicity, we use and to gather the transition probability vectors and emission parameters associated with all states in . Here, can be seen as a bi-infinite matrix whose elements are the probabilities for each possible transition , and it is directly analogous to the transition matrix of the finite HMM.
Figure 3.
Graphical representation of the iHMM. The hidden Markov model that formulates the observations to be analyzed (black lines) is shown together with its priors (red lines). For completeness, we also show the concentration parameters α and γ and the prior probability distribution on the emission parameters, H, that fully characterize the iHMM. The key difference from the HMM shown in Fig. 2 is that now the model parameters and are treated as random variables similar to the hidden states, , and observations, . For details, see the main text. To see this figure in color, go online.
Unlike the finite HMM, where we need to be specific about the size of the state space, the iHMM gives us greater flexibility through the hyperparameters α and γ. For instance, larger concentrations are more appropriate for biomolecules with roughly similar transition rates and several conformations, whereas low concentrations are more appropriate for biomolecules with dissimilar transition rates and few conformations. In general, it is possible to sum over these concentrations if we are truly ignorant of the properties of the biomolecule at hand (20, 31).
As we will see shortly, this powerful formalism allows us to infer state numbers robustly, and even if the number of states is apparent—something that is rarely the case for more complex problems (32)—it avoids the critical shortfall of cherry-picking data sets to be analyzed that appear to have similar state numbers.
Inference on the iHMM
Before describing the analysis that iHMMs may offer, we show how to infer quantities such as the hidden state sequence and the parameters from the experimental traces. A working implementation of the algorithm described in this section, equipped with a user interface, can be found in the Supporting Materials. (This code can also be found on the authors’ web site, as well as on GitHub.)
To be clear, on account of the infinite size of , common methods used in finite-state HMMs, such as expectation maximization (1), are inapplicable. Instead, we focus our discussion on a specialized method, the beam sampler (33). Other methods are also described elsewhere (20, 31).
The beam sampler is a special instance of the Gibbs sampler (34), and although it is beyond the scope of this survey, a brief description about its usage might be beneficial here. Similar to all samplers of this family, we use the beam sampler to generate (pseudo) random sequences of the model variables we are interested in inferring. Specifically, in our case these variables consist of the hidden-state sequence and the model parameters . When generating the sequences, we use (7), (8), (9), (10), (11) and the data . As a result, the generated sequences have the same statistical properties as if they consisted of samples from the posterior probability distribution, . Thus, we may use them to compute averages, confidence intervals, and maximum a posteriori estimates or simply to produce histograms that resemble , as we show in the next section.
Overall, Gibbs and related samplers provide a more general approach than, for example, expectation maximization, which provides only the maximum of . Instead, with Gibbs sampling, we fully characterize over its whole domain. For an introduction to the general methodology underlying Gibbs sampling, we refer the interested reader to (35), and from now on, we focus on the iHMM.
Suppose we have a time series of experimental observations . Here, we describe the steps involved in generating samples (superscript r) of our model variables . Given a computed sample , the naive approach for computing the next sample would be to update each of the involved variables conditional upon the other variables and the data (35). For example, we could generate by sampling from . Then, we could generate by sampling from , and so on. Based on the theory of Markov-chain Monte Carlo sampling (see, for example, (34)), this approach is guaranteed to produce samples from the full posterior , as we wish. However, the infinite size of makes this approach impractical, since in each iteration we need to sample infinite-sized , which is computationally infeasible.
It is fundamentally because of this difficulty that we need the beam sampler, which introduces a set of auxiliary variables (slicers) that in each iteration effectively truncate to a finite portion (33, 36). In particular, for each time step n in the data set, consider an auxiliary variable, , and let gather all of them. Since consists of auxiliary variables only, we have
| (12) |
where the sum on the righthand side is considered over all possible values may take. Now, we can use the equality in Eq. 12 to sample from instead of from . The benefit of doing so is that when updating , we also have to sample conditional upon . Thus, by properly choosing the auxiliaries , we make these updates consider only a finite selection of the states . For instance, by having uniformly distributed over the interval , it is sufficient to consider only those states in that have a probability of being visited at least once that is >. Therefore, we can generate the desired samples while maintaining a finite that we expand and compress dynamically according to .
In summary, given , , , and the beam sampler generates a new set of samples through the following steps:
-
1.
Generate by sampling from
-
2.
Expand according to
-
3.
Generate by sampling from
-
4.
Compress according to
-
5.
Generate by sampling from
-
6.
Generate by sampling from
-
7.
Generate by sampling from
where, for simplicity, we have dropped additional dependencies in the conditional probability distributions above.
In principle, , , and are infinite dimensional vectors. However, as mentioned above, the sampler requires the computation of only those components that correspond to the states allows to be visited. Therefore, in step 4 of the above algorithm we can safely discard those components that are absent from the state sequence . By doing so, in each time step, we only need to store vectors of finite dimension, , , and . Subsequently, in step 2 of the following iteration, we can supplement , , and with states as necessary. The discarded states, since they are unvisited, do not produce any of the observations in . Thus, they do not provide any information that could be lost. Similarly, the supplemented states are also not associated with any of the observations in . Thus, when we generate them, we simply have to draw samples from their priors. Consequently, the supplemented states neither require nor introduce extra information, as might first appear to be the case. In summary, expanding and compressing in steps 2 and 4 leaves our inference unaffected.
The Supporting Material provides a pseudocode implementation of the algorithm with further details regarding each of the sampler’s steps for the interested reader.
Results
In this section, we demonstrate the use of the iHMM in the analysis of time-series data. To benchmark the method, we use synthetic time series where the “ground truth” is known. In a companion research article that draws heavily from the formulation described in this article, we show the performance of the method on actual time traces that exhibit further complications, such as drift (37). For such time traces, we need an additional important generalization to the iHMM: the iHMM must be coupled to a continuous process. For now, our synthetic traces are meant to illustrate realistic data with noise, but from which drift is otherwise absent or minimal.
Below, we present the results of the analysis on two types of time trace that are specifically problematic for the HMM: 1) data set 1, consisting of time traces where the visited states are in high number and their emissions show significant overlap; and 2) data set 2, consisting of time traces where some states are rarely visited and the number of states appears to change as more data become available.
Details about the generation of these data sets, as well as more in-depth analyses than those discussed here, can be found in the Supporting Material.
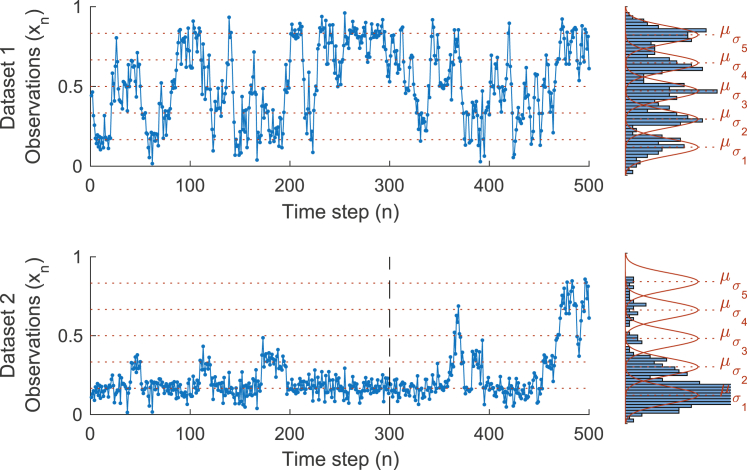
The data sets we analyze are shown in Fig. 4. Specifically, we used these time series to estimate the posterior distributions over 1) the number, K, of different conformations visited by the biomolecule, and 2) the parameters describing the emission distributions, such as the mean value for each conformation , for those states. To accomplish this, we generate samples from using the beam sampler described earlier.
Figure 4.
Synthetic data sets resembling a hypothetical biomolecule undergoing transitions between discrete states that we analyzed with the iHMM. (Left) Time series of noisy observations. During the measuring period, the biomolecule attains five conformations, . The number of conformations are a priori unknown and the iHMM seeks to determine the probability over the number of states, as well as their properties, given the data available. In data set 1, the biomolecule transitions often through every state. By contrast, in data set 2, transitions to some states are rare. As a result, all states in data set 1 are almost equally visited throughout the experiment time course, whereas in data set 2, higher states are visited, by chance, only toward the end of the trace. (Right) The corresponding emission distributions, , as obtained by simply binning the observations (blue) and plotting the exact ones used for the simulations (red). For both data sets, the emission distributions show significant overlap. In all panels, dotted lines indicate the exact mean values, , of the emission distributions. To see this figure in color, go online.
Fig. 5 shows how the number of different states visited in each sampled state sequence for data set 1, which we denote with , changes through the sampler’s iterations. Again, we mention that K counts only those states that are visited within the given trace. Initially, equals the number of states in the state space as specified in the initialization of our method. After ∼50 iterations, drops to 5, which is the correct number of states attained by our hypothetical biomolecule.
Figure 5.
After some iterations, the sampler used in the iHMM to analyze data set 1 of Fig. 4 eventually converges to the correct number of states. The number of visited states, (top), and the means of the emission distributions, (bottom), change throughout the sampler’s iterations. Unlike the HMM, which uses a finite and fixed state space, the iHMM learns the number of available states and grows/shrinks the state space as required by the data.
Subsequently, stays at 5 and occasionally adds extra states that are rapidly eliminated. This behavior is characteristic of our algorithm (34). In the initial iterations, known as “burn-in,” the sampler randomly explores possible choices for the model variables that eventually evolve closer to the ground truth. Once they approach the ground truth, the sampler begins generating them based on the corresponding posterior distribution, . So, after burn-in, values with high are sampled more often than those with low .
The same is true of the other variables we try to estimate. For example, in Fig. 5, we also show how the sampled means of the emission distributions, , change through the iterations. As with , during burn-in, the sampler explores different choices that eventually converge to the ground truth. Afterward, the sampler occasionally explores new emissions, for example, around iterations 150–250, that quickly disappear.
To obtain more quantitative results, we may drop the burn-in samples from the generated sequences and use the remainder of the samples to produce histograms. For example, in Fig. 6, we show histograms of and . These histograms approximate the corresponding posterior probability distributions and thus provide a full characterization of the estimated variables. In these cases, the breadth of the histograms arises because the data we analyze are limited and noisy, so the estimates are associated with some uncertainty that is reflected in the broadening of the posterior distributions.
Figure 6.
We may use samples from the iHMM posterior probability to infer the size of the state space and the location of each state. In particular, we illustrate histograms for (top) and (bottom) using data set 1 of Fig. 4. In both panels, dashed lines indicate the exact (ground-truth) values used to produce the data in Fig. 4. To see this figure in color, go online.
As can be seen, data set 1—although it contains a large number of states that are not well separated—can be successfully analyzed by means of the iHMM. However, data set 2 poses another difficulty. Specifically, the biomolecule visits some states only rarely. In experiments, this would give rise to traces with an unequal number of states. One may naively suggest that longer traces should be collected, but due to experimental limitations (e.g., early photobleaching in FRET experiments), such long traces might not always be available. Instead, a large number of shorter traces, each containing an unknown portion of the complete state space, may need to be analyzed separately and the results combined (38). For this reason, it is crucial to treat all traces on an equal footing without a priori assuming a different number of states individually for each one, as the HMM would require.
To illustrate how the iHMM handles such cases, we use the iHMM to analyze the emission data of Fig. 4 twice, first using only the initial 300 time steps, where only two of the five states are visited, and subsequently using the complete trace, which contains all five states.
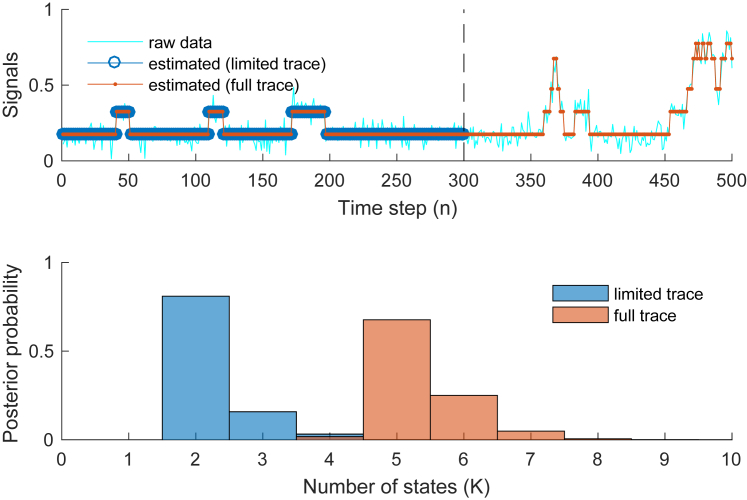
Fig. 7 shows the results of the analyses. Specifically, we show the estimated noiseless traces (Fig. 7, upper) and the estimated numbers of states (Fig. 7, lower). As can be seen, the iHMM successfully estimates the correct number in each trace. Further, as can be seen from the de-noised traces, the portion of the state space, i.e., states and , that is common is estimated accurately from both traces.
Figure 7.
We may use the iHMM to estimate portions of the complete state space such as those contained in different segments of data set 2 provided in Fig. 4. (Upper) Estimated noiseless traces for two cases: 1) using a limited segment of the full trace; and 2) using the full trace. Although only the latter case allows an estimate of all five states, both cases provide similar estimates over those states that they mutually visit. (Lower) Corresponding estimates of the number of states contained in each trace. To see this figure in color, go online.
Perspectives
In this survey, we presented the iHMM (19, 20), a relatively recent development in statistics that has already found diverse applicability in, among other fields, genomics (39), genetics (40), finance (41, 42), tracking (43, 44), machine learning (45, 46), and biophysics (22, 23, 24).
As a modeling tool, the iHMM inherits the characteristics of its predecessor—the finite HMM—but offers greater flexibility in the modeling and analysis of the experimental data. An important advantage is that it does not require the underlying system to have a limited state space as would its predecessor. This feature is of particular importance to biophysics, where it is often the case that little information is available to fix the number of biomolecular states a priori.
Despite the difficulty in prespecifying the complexity of the state space, the finite HMM is a routine choice for the analysis of single-molecule time series (11, 12, 13). Nevertheless, as the experimental techniques improve and data from complex biomolecules are becoming available, the shortcomings of the finite HMM have become more apparent. To this end, methods to circumvent the limitations of HMM have been developed independently of the progress made on the same problems that have exploited Bayesian nonparametrics.
Although it has previously been suggested that single-molecule analysis may benefit from Bayesian nonparametrics (22), to date only limited applications of these methods have been explored. This is in part because models like the iHMM and its implementation are scattered over various sources, but also because of the important language barrier between fields where Bayesian nonparametrics have changed the course of data analysis and the physical sciences. We believe that it is in part for this reason that methods continue to be developed to address consequences of the finiteness of the state space of the HMM in the physical sciences.
The iHMM, described here, presents great advantages in the analysis of single-molecule observations. For one thing, it elegantly addresses those challenges that are presented by the HMM, namely, the problem of having to select a number of states when the emission distributions are unclear, but also to address the problem—currently often addressed in a user-dependent fashion—of how to proceed when different traces present a seemingly different number of states. The key idea is that, just as the HMM performs de-noising while parametrizing transition kinetics, the iHMM takes this logic one step further by additionally learning the number of states in a self-consistent manner.
Although greatly advantageous, the iHMM itself presents an important challenge for experimental data with drift, a common feature of biophysical time traces (13, 25). Indeed, without its explicit consideration, drift would be interpreted as the population of artifact states by a method designed to recruit extra states to accommodate the data. This is a key challenge that has so far not been considered anywhere. Without careful consideration, drift would render the power of the iHMM into a key disadvantage. Although our main emphasis here has been to feature the power of the iHMM to biophysics, our companion research article (37) now tackles the problem of extending iHMMs to deal with drift.
Author Contributions
I.S. and S.P. performed research; I.S. contributed analytic tools and analyzed data; I.S. and S.P. wrote the article.
Editor: Brian Salzberg.
Footnotes
Supporting Materials and Methods and three figures are available at http://www.biophysj.org/biophysj/supplemental/S0006-3495(17)30443-5.
Supporting Material
References
- 1.Rabiner L., Juang B. An introduction to hidden Markov models. IEEE ASSP Mag. 1986;3:4–16. [Google Scholar]
- 2.Eddy S.R. What is a hidden Markov model? Nat. Biotechnol. 2004;22:1315–1316. doi: 10.1038/nbt1004-1315. [DOI] [PubMed] [Google Scholar]
- 3.Yoon B.J. Hidden Markov models and their applications in biological sequence analysis. Curr. Genomics. 2009;10:402–415. doi: 10.2174/138920209789177575. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Krogh A., Brown M., Haussler D. Hidden Markov models in computational biology. Applications to protein modeling. J. Mol. Biol. 1994;235:1501–1531. doi: 10.1006/jmbi.1994.1104. [DOI] [PubMed] [Google Scholar]
- 5.Streit R.L., Barrett R.F. Frequency line tracking using hidden Markov models. IEEE Trans. Acoust. Speech Signal Process. 1990;38:586–598. [Google Scholar]
- 6.Pikrakis A., Theodoridis S., Kamarotos D. Classification of musical patterns using variable duration hidden Markov models. IEEE Trans. Audio Speech Lang. Process. 2006;14:1795–1807. [Google Scholar]
- 7.Dasgupta N., Runkle P., Carin L. Dual hidden Markov model for characterizing wavelet coefficients from multi-aspect scattering data. Signal Process. 2001;81:1303–1316. [Google Scholar]
- 8.Fine S., Singer Y., Tishby N. The hierarchical hidden Markov model: analysis and applications. Mach. Learn. 1998;32:41–62. [Google Scholar]
- 9.Juang B.H., Rabiner L.R. Hidden Markov models for speech recognition. Technometrics. 1991;33:251–272. [Google Scholar]
- 10.Marroquin J.L., Santana E.A., Botello S. Hidden Markov measure field models for image segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 2003;25:1380–1387. [Google Scholar]
- 11.McKinney S.A., Joo C., Ha T. Analysis of single-molecule FRET trajectories using hidden Markov modeling. Biophys. J. 2006;91:1941–1951. doi: 10.1529/biophysj.106.082487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Bronson J.E., Fei J., Wiggins C.H. Learning rates and states from biophysical time series: a Bayesian approach to model selection and single-molecule FRET data. Biophys. J. 2009;97:3196–3205. doi: 10.1016/j.bpj.2009.09.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Blanco M., Walter N.G. Analysis of complex single-molecule FRET time trajectories. Methods Enzymol. 2010;472:153–178. doi: 10.1016/S0076-6879(10)72011-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Long X., Parks J.W., Stone M.D. Mechanical unfolding of human telomere G-quadruplex DNA probed by integrated fluorescence and magnetic tweezers spectroscopy. Nucleic Acids Res. 2013;41:2746–2755. doi: 10.1093/nar/gks1341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kruithof M., van Noort J. Hidden Markov analysis of nucleosome unwrapping under force. Biophys. J. 2009;96:3708–3715. doi: 10.1016/j.bpj.2009.01.048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Venkataramanan L., Sigworth F.J. Applying hidden Markov models to the analysis of single ion channel activity. Biophys. J. 2002;82:1930–1942. doi: 10.1016/S0006-3495(02)75542-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Munro J.B., Altman R.B., Blanchard S.C. Identification of two distinct hybrid state intermediates on the ribosome. Mol. Cell. 2007;25:505–517. doi: 10.1016/j.molcel.2007.01.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Ferguson T.S. A Bayesian analysis of some nonparametric problems. Ann. Stat. 1973;1:209–230. [Google Scholar]
- 19.Beal M.J., Ghahramani Z., Rasmussen C.E. The infinite hidden Markov model. In: Jordan M.L., editor. Advances in Neural Information Processing Systems. MIT Press; Cambridge, MA: 2001. pp. 577–584. [Google Scholar]
- 20.Teh Y.W., Jordan M.I., Blei D.M. Hierarchical Dirichlet processes. J. Am. Stat. Assoc. 2012 [Google Scholar]
- 21.Hines K.E. A primer on Bayesian inference for biophysical systems. Biophys. J. 2015;108:2103–2113. doi: 10.1016/j.bpj.2015.03.042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Hines K.E., Bankston J.R., Aldrich R.W. Analyzing single-molecule time series via nonparametric Bayesian inference. Biophys. J. 2015;108:540–556. doi: 10.1016/j.bpj.2014.12.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Palla, K., D. A. Knowles, and Z. Ghahramani. 2014. A reversible infinite HMM using normalised random measures. Proceedings of the 31st International Conference on Machine Learning. 5:4090–4107.
- 24.Calderon C.P., Bloom K. Inferring latent states and refining force estimates via hierarchical dirichlet process modeling in single particle tracking experiments. PLoS One. 2015;10:e0137633. doi: 10.1371/journal.pone.0137633. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Roy R., Hohng S., Ha T. A practical guide to single-molecule FRET. Nat. Methods. 2008;5:507–516. doi: 10.1038/nmeth.1208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Bishop C. Springer-Verlag; New York: 2006. Pattern Recognition and Machine Learning. [Google Scholar]
- 27.Nir E., Michalet X., Weiss S. Shot-noise limited single-molecule FRET histograms: comparison between theory and experiments. J. Phys. Chem. B. 2006;110:22103–22124. doi: 10.1021/jp063483n. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Yoon J.W., Bruckbauer A., Klenerman D. Bayesian inference for improved single molecule fluorescence tracking. Biophys. J. 2008;94:4932–4947. doi: 10.1529/biophysj.107.116285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Sethuraman J. A constructive definition of Dirichlet priors. Stat. Sin. 1994;4:639–650. [Google Scholar]
- 30.Jim Pitman Poisson-Dirichlet and gem invariant distributions for split-and-merge transformations of an interval partition. Combin. Probab. Comput. 2002;11:501–514. [Google Scholar]
- 31.Fox E.B., Sudderth E.B., Willsky A.S. A sticky HDP-HMM with application to speaker diarization. Ann. Appl. Stat. 2011;5:1020–1056. [Google Scholar]
- 32.Pirchi M., Ziv G., Haran G. Single-molecule fluorescence spectroscopy maps the folding landscape of a large protein. Nat. Commun. 2011;2:493. doi: 10.1038/ncomms1504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Van Gael, J., Y. Saatci, …, Z. Ghahramani. 2008. Beam sampling for the infinite hidden Markov model. In Proceedings of the 25th International Conference on Machine Learning. (ACM), pp. 1088–1095.
- 34.Robert Ch., Casella G. Springer Science & Business Media; Berlin, Germany: 2013. Monte Carlo Statistical Methods. [Google Scholar]
- 35.Robert C., Casella G. Springer Science & Business Media; Berlin, Germany: 2009. Introducing Monte Carlo Methods with R. [Google Scholar]
- 36.Walker S.G. Sampling the Dirichlet mixture model with slices. Commun. Stat. Simul. Comput. 2007;36:45–54. [Google Scholar]
- 37.Sgouralis I., Pressé S. ICON: an adaptation of infinite HMMs for time traces with drift. Biophys. J. 2017;112:2117–2126. doi: 10.1016/j.bpj.2017.04.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Blanco M.R., Martin J.S., Walter N.G. Single molecule cluster analysis dissects splicing pathway conformational dynamics. Nat. Methods. 2015;12:1077–1084. doi: 10.1038/nmeth.3602. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Yau C., Papaspiliopoulos O., Holmes C. Bayesian non-parametric hidden Markov models with applications in genomics. J. R. Stat. Soc. Series B Stat. Methodol. 2011;73:37–57. doi: 10.1111/j.1467-9868.2010.00756.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Xing E.P., Sohn K.A. Hidden Markov Dirichlet process: modeling genetic inference in open ancestral space. Bayesian Anal. 2007;2:501–527. [Google Scholar]
- 41.Dufays A. Infinite-state Markov-switching for dynamic volatility. J. Financ. Econom. 2015;14:418–460. [Google Scholar]
- 42.Shi S., Song Y. Identifying speculative bubbles using an infinite hidden Markov model. J. Financ. Econom. 2014;14:159–184. [Google Scholar]
- 43.Kuettel, D., M. D. Breitenstein, …, V. Ferrari. 2010. What’s going on? discovering spatio-temporal dependencies in dynamic scenes. In 2010 IEEE Conference on Computer Vision and Pattern Recognition (CVPR). (IEEE), pp. 1951–1958.
- 44.Fox, E. B., E. B. Sudderth, and A. S. Willsky. 2007. Hierarchical Dirichlet processes for tracking maneuvering targets. In 2007 10th International Conference on Information Fusion. (IEEE), pp. 1–8.
- 45.Teh Y.W., Jordan M.I. Hierarchical Bayesian nonparametric models with applications. In: Hjort N.L., Holmes C., Müller P., Walker S.G., editors. Bayesian Nonparametrics. Cambridge University Press; 2010. pp. 158–203. [Google Scholar]
- 46.Hu W., Tian G., Maybank S. An improved hierarchical Dirichlet process-hidden Markov model and its application to trajectory modeling and retrieval. Int. J. Comput. Vis. 2013;105:246–268. [Google Scholar]
- 47.Jiang L.-L., Zhao J., Hu H.-Y. Two mutations G335D and Q343R within the amyloidogenic core region of TDP-43 influence its aggregation and inclusion formation. Sci. Rep. 2016;6:23928. doi: 10.1038/srep23928. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.