Abstract
The vast majority of eukaryotes (fungi, plants, animals, slime mold, and euglena) synthesize Asn-linked glycans (Alg) by means of a lipid-linked precursor dolichol-PP-GlcNAc2Man9Glc3. Knowledge of this pathway is important because defects in the glycosyltransferases (Alg1-Alg12 and others not yet identified), which make dolichol-PP-glycans, lead to numerous congenital disorders of glycosylation. Here we used bioinformatic and experimental methods to characterize Alg glycosyltransferases and dolichol-PP-glycans of diverse protists, including many human pathogens, with the following major conclusions. First, it is demonstrated that common ancestry is a useful method of predicting the Alg glycosyltransferase inventory of each eukaryote. Second, in the vast majority of cases, this inventory accurately predicts the dolichol-PP-glycans observed. Third, Alg glycosyltransferases are missing in sets from each organism (e.g., all of the glycosyltransferases that add glucose and mannose are absent from Giardia and Plasmodium). Fourth, dolichol-PP-GlcNAc2Man5 (present in Entamoeba and Trichomonas) and dolichol-PP- and N-linked GlcNAc2 (present in Giardia) have not been identified previously in wild-type organisms. Finally, the present diversity of protist and fungal dolichol-PP-linked glycans appears to result from secondary loss of glycosyltransferases from a common ancestor that contained the complete set of Alg glycosyltransferases.
Keywords: evolution, N-glycans, protist
The majority of eukaryotes studied to date (fungi, plants, animals, slime mold, and euglena) synthesize Asn-linked glycans (Alg) by means of a lipid-linked precursor dolichol-PP-GlcNAc2Man9Glc3 (Fig. 1A) (1-3). Each of the 14 sugars is added to the lipid-linked precursor by means of a specific glycosyltransferase (Alg1-Alg12 and others as yet specified), which were numbered according to the order of their discovery rather than by the sequence of enzymatic steps (4-6). Defects in these Alg glycosyltransferases lead to numerous congenital disorders of glycosylation, which can cause dysmorphic features and mental retardation (7). We and others used the budding yeast Saccharomyces cerevisiae mutants to characterize many but not all of the Alg glycosyltransferases, which are present on the cytosolic aspect of the ER. These glycosyltransferases include Alg7, which adds phospho-GlcNAc to dolichol phosphate, and Alg1, Alg2, and Alg11, which add the first, second, and fifth Man residues, respectively (Fig. 1A). Dolichol-PP-GlcNAc2Man5 is flipped into the lumen of the ER by a flippase (Rft1) (8). Within the ER lumen, mannosyltransferases Alg3, Alg9, and Alg12 make dolichol-PP-GlcNAc2Man9 by using dolichol-P-Man (made by Dpm1) as the sugar donor (Fig. 1A) (1, 9). Also within the ER lumen, glucosyltransferases Alg6, Alg8, and Alg10 make dolichol-PP-GlcNAc2Man9Glc3 by using dolichol-P-Glc (made by Alg5) as the sugar donor (Fig. 1A).
Fig. 1.
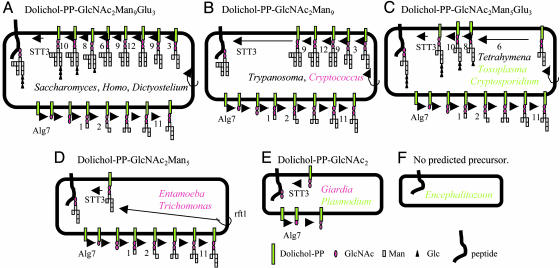
The inventory of Alg glycosyltransferases and predicted dolichol-linked glycans vary dramatically among protists and fungi. Predicted Alg glycosyltransferases and dolichol-linked glycans of Saccharomyces cerevisiae, Homo sapiens, and Dictyostelium discoideum (A), Trypanosoma cruzi, Trypanosoma brucei, Leishmania major, and Cryptococcus neoformans (B), Tetrahymena thermophilia, Toxoplasma gondii, and Cryptosporidium parvum (C), Entamoeba histolytica and Trichomonas vaginalis (D), Plasmodium falciparum and Giardia lamblia (E), and Encephalitozoon cuniculi (F) (see also Table 1). With the exceptions of Saccharomyces and Homo, sets of Alg glycosyltransferases were identified here. Names of organisms, whose dolichol-PP-linked glycans were previously identified (e.g., Saccharomyces), are indicated in black. Names of organisms, whose dolichol-PP-linked glycans were identified here (e.g., Cryptococcus), are indicated in red. Names of organisms, whose dolichol-PP-linked glycans have not yet been identified (e.g., Cryptosporidium), are indicated in green.
An oligosaccharyltransferase (OST), which contains a catalytic peptide, STT3, transfers the dolichol-PP-linked oligosaccharide to “sequon” Asn residues (N-X-T/S) on nascent peptides (Fig. 1A) (10-12). A UDP-Glc:glycoprotein glucosyltransferase adds glucose to N-glycans of improperly folded proteins, which are retained in the ER by conserved glucose-binding lectins (calnexin/calreticulin) (13). Although the Alg glycosyltransferases in the lumen of ER appear to be eukaryote-specific, archaea and Campylobacter sp. glycosylate the sequon Asn and/or contain glycosyltransferases with domains like those of Alg1, Alg2, Alg7, and STT3 (1, 14-16).
Protists, unicellular eukaryotes, suggest three notable exceptions to the N-linked glycosylation path described in yeast and animals (17). First, the kinetoplastid Trypanosoma cruzi (cause of Chagas myocarditis), fails to glucosylate the dolichol-PP-linked precursor and so makes dolichol-PP-GlcNAc2Man9 (18). Another kinetoplastid Leishmania mexicana (cause of skin ulcers) lacks the mannosylating activities of Alg9 and Alg12 and makes dolichol-PP-GlcNAc2Man6 (Alg9 adds both the 7th and 9th Man residues) (18). Second, Tetrahymena pyriformis, which is a free-living ciliate, lacks all of the mannosylating activity in the ER lumen and makes dolichol-PP-GlcNAc2Man5Glc3 (19). Third, it has been difficult to identify N-glycans from either Giardia lamblia (cause of diarrhea) or Plasmodium falicparum (cause of severe malaria) (20-22).
Because these observations of unique protist glycans were made before identification of multiple Alg glycosyltransferases and/or whole-genome sequencing of these protists, numerous important questions remain concerning the diversity of N-glycan precursors among eukaryotes. First, are the same Alg glycosyltransferases conserved across all eukaryotes, and are these Alg glycosyltransferases specific to eukaryotes or are they also present in prokaryotes? Second, are kinetoplastids [Trypanosoma cruzi and Trypanosoma brucei (cause of African sleeping sickness)] missing the genes encoding the glucosylating enzymes or are the genes present but silent, and similarly, is Tetrahymena missing the set of genes encoding mannosylating enzymes in the lumen of the ER? Third, what Alg glycosyltransferases are missing from Giardia and Plasmodium? Fourth, what is the diversity of predicted Alg glycosyltransferases of other protists (e.g., Entamoeba histolytica, Trichomonas vaginalis, Toxoplasma gondii, and Cryptosporidium parvum, which cause dysentery, vaginitis, birth defects, and diarrhea, respectively), and what are the Alg glycosyltransferases of Encephalitozoon cuniculi (an opportunistic fungus with a dramatically reduced genome) and Cryptococcus neoformans (an opportunistic fungus that is distantly related to Saccharomyces)? Fifth, do the predicted Alg glycosyltransferases correlate with the dolichol-PP-linked glycans of each protist or fungus? Sixth, does the pattern distribution of Alg glycosyltransferases across protists, fungi, and metazoa suggest whether these glycosyltransferases have been added to or lost from eukaryotes during their evolution (23, 24)?
Materials and Methods
Use of Bioinformatics to Identify Alg Glycosyltransferases from Diverse Protists and Fungi. Twelve Saccharomyces Alg glycosyltransferases, DPM1, and STT3 were used to search predicted proteins of eukaryotes, which have been sequenced in their entirety or near entirety (Plasmodium falciparum, Encephalitozoon cuniculi, Cryptosporidium parvum, Giardia lamblia, Homo sapiens, and Arabidopsis thaliana) in the nr protein database of the National Center for Biotechnology Information by using psi-blast (25-30). blastp or tblastn searches of Entamoeba histolytica, Trichomonas vaginalis, Trypanosoma brucei, Trypanosoma cruzi, and Tetrhymena thermophilia were performed at web sites managed by The Institute for Genomic Research (www.tigr.org). Predicted proteins of the slime mold Dictyostelium discoideum and kinetoplastid Leishmania major were performed on the Sanger Institute genedb web site (www.genedb.org), and the apicomplexan Toxoplasma gondii-predicted proteins were searched at toxodb (http://ToxoDB.org). Transmembrane helices were predicted by using the Phobius combined transmembrane topology and signal peptide predictor (31).
Alignments of protein sequences were made by using clustalw (http://www.ebi.ac.uk/clustalw), and manual adjustments and trimming of the alignments were performed with jalview (32). Phylogenetic trees were constructed from the positional variation with maximum likelihood by using quartet puzzling (33, 34).
Identification of Dolichol-PP-Linked Precursors and N-Linked Glycans. Genome project strains of Entamoeba histolytica, Trichomonas vaginalis, Giardia lamblia and Cryptococcus neoformans were grown axenically and labeled with 200 μCi (1 Ci = 37 GBq) [2-3H]Man, [6-3H]GlcN, or [3H]Glc in a Glc-free medium for 10 min in a final volume of 250 μl (6). Dolichol-PP-linked glycans were extracted with chloroform/methanol/water, dried, and hydrolyzed in 0.1 M HCl for 45 min at 90°C. Glycans were neutralized and resuspended in 0.1 M acetic acid and 1% butanol for separation on a 1-m Biogel P-4 superfine column (BioRad). Standards were GlcNAc2Man5 from a Saccharomyces alg3Δ mutant incubated with 14C-Man, GlcNAc2Man9 from a Saccharomyces alg6Δ mutant, and unlabeled GlcNAc and GlcNAc2.
For identification of N-glycans, Entamoeba, Trichomonas, and Giardia were labeled with mannose and GlcN for 2 h in medium containing 0.1% glucose before washing and lyophilization. The dry-cell pellet was delipidated with chloroform/methanol/water, glycosylphosphatidylinositol precursors were removed with water-saturated butanol, and glycogen and other free glycans were removed with 50% methanol (35). The clean pellet was finely resuspended by using a manual homogenizer in 500 μl Tris·HCl (0.1M, pH 8) and incubated with 50 milliunits of peptide-N-glycosidase F (PNGaseF) at 37°C for 16 h. Negative controls omitted the PNGaseF, and the peptide:N-glycanase supernatant was chromatographed on a P-4 column. Radioactivity was measured by scintillation counting, and peaks were isolated for treatment with glycosidases. The putative GlcNAc2Man5 from Entamoeba and Trichomonas and putative GlcNAc2Man7 from Cryptococcus were treated with α-1,2-mannosidase, whereas the putative GlcNAc2 of Giardia was cleaved with chitiobiase.
In Vitro Synthesis of Glycopeptides by Using Intact Protist Membranes as a Source of Dolichol-PP-Glycans. Total cellular membranes were prepared from cultures of Trichomonas, Entamoeba, and Cryptococcus. The membranes were incubated for 2-90 min at 37°C with the membrane permeable tripeptide acceptor, 5 μM Nα-Ac-Asn-[125I]-Tyr-Thr-NH2 (NYT), in the presence of deoxynojiromycin to ensure that the glycopeptide products were not degraded by glucosidases I and II (36). Glycopeptide products were collected by binding to immobilized Con A and separated on HPLC by using standards from Saccharomyces that included Man5GlcNAc2-NYT, Man9GlcNAc2-NYT, Glc3Man5GlcNAc2-NYT, and Glc3Man9GlcNAc2-NYT (37).
Results and Discussion
A Common Ancestor of Eukaryotes and Archaea May Have Contained STT3 and Alg7, but the Remaining Alg Glycosyltransferases Appear to Be Eukaryote-Specific. Similarities between eukaryotic cytosolic Alg glycosyltransferases (Alg1, Alg2, and Alg7) and STT3 and their prokaryotic counterparts suggest their common origin (1, 24), but phylogenetic methods have not been used to test this idea. STT3 is present in all eukaryotes examined except Encephalitozoon (Fig. 1, Table 1, and see below) and is present in multiple copies in some protists [e.g., two in Entamoeba, three in Trichomonas and Trypanosoma brucei, and four in Leishmania (data not shown)]. Homologues of STT3 are also present in the bacterium Campylobacter jejuni and both divisions of archaea, euryarchaeota and crenarchaeota (16). The hydrophobicity plots of the eukaryotic and prokaryotic STT3 closely resemble each other, each containing 10 to 15 predicted transmembrane helices (data not shown). In addition, eukaryotic STT3 show a 25-45% positional identity with each other over a ≈700-amino acid (90%) overlap and a 19-23% positional identity with prokaryotic STT3 over a ≈600-amino acid (80%) overlap. Phylogenetic analyses of STT3 show distinct eukaryotic and archael clades, although it was not possible to determine whether eukaryotic STT3 are more similar to homologues of euryarchaeota or crenarchaeota (data not shown). The Campylobacter STT3 gene appears to have been laterally transferred from archaea. These sequence comparisons, as well as the demonstration of OST activity of Campylobacter STT3 to an N-X-T/S sequon (16), are consistent with the idea that a common ancestor to eukaryotes and archaea contained STT3.
Table 1. Predicted Alg glycosyltransferases of representative eukaryotes.
| Cytosol GlcNAc
|
Cytosol Man
|
ER lumen Man
|
ER lumen Glc
|
OST
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Genus/species abbreviation | Alg7 | Alg1 | Alg2 | Alg11 | Rft1 | Alg3 | Alg9 | Alg12 | Dpm1 | Alg5 | Alg6 | Alg8 | Alg10 | STT3* |
| Sc/Hs/Dd | yes | yes | yes | yes | yes | yes | yes | yes | yes | yes | yes | yes | yes | 1/2/1 |
| Tb/Tc/Lm/Cn | yes | yes | yes | yes | yes | yes | yes | y/y/n/y | yes | no | no | no | no | 3/1/4/1 |
| Eh/Tv | yes | yes | yes | yes | yes | no | no | no | y/n | n/y | no | no | no | 2/3 |
| Tt/Cp/Tg | yes | yes | yes | y/y/n | y/y/n | no | no | no | yes | yes | yes | y/n/y | y/n/y | 1/1/1 |
| Pf/Gl | yes | no | no | no | no | no | no | no | yes | no | no | no | no | 1/1 |
| Ec | no | no | no | no | no | no | no | no | yes | no | no | no | no | 0 |
Sc, Saccharomyces cerevisiae; Hs, Homo sapiens; Dd, Dictyostelium discoideum; Tb, Trypanosoma brucei; Tc, Trypanosoma cruzi; Lm, Leishmania mexicana; Cn, Cryptococcus neoformans; Eh, Entamoeba histolytica; Tv, Trichomonas vaginalis; Tt, Tetrahymena thermophilia; Tg, Toxoplasma gondii; Cp, Cryptosp parvum; Pf, Plasmodium falciparum; Gl, Giardia lamblia; Ec, Encephalitozoon cuniculi.
No. of STT3 subunits in each organism
Alg7, which is a UDP-GlcNAc:dolichol-phosphate GlcNAc-1-phosphate transferase, is the first enzyme in the synthesis of dolichol-PP-linked glycans and is present in all eukaryotes examined except Encephalitozoon (Fig. 1, Table 1, and see below). Proteins similar to Alg7 are predicted from whole-genome sequences of some but not all archaea and bacteria. The hydrophobicity plots of the eukaryotic and prokaryotic Alg7 closely resemble each other, each containing 8 to 12 predicted transmembrane helices (data not shown). Eukaryotic Alg7 show a 28-40% positional identity with each other over an ≈310-amino acid (80%) overlap and show an ≈19-23% positional identity with prokaryotic sequences over a 240-amino acid (60%) overlap. Phylogenetic analyses of Alg7 show distinct eukaryotic, archaeal, and bacterial clades (Fig. 2A). Although eukaryote Alg7 are much more similar to archaeal than bacterial homologues, it was again not possible to determine whether eukaryotic Alg7 was more similar to those of euryarchaeota or crenarchaeota. These results and the presence of dolichol-PP-linked glycans in archaea (14) are consistent with the idea that a common ancestor to eukaryotes and archaea contained Alg7.
Fig. 2.
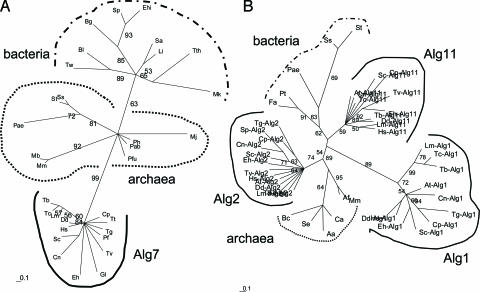
Common ancestry is a useful method of predicting the Alg glycotransferase inventory of each eukaryote. Phylogenetic reconstructions by using the maximum likelihood method of representative eukaryotic and prokaryotic Alg7 (A) and Alg1, Alg2, and Alg11 (B). Branch lengths are proportionate to differences between sequences, and numbers at nodes indicate bootstrap values for 100 replicates. Eukaryotes include Arabidopsis thaliana (At), Cryptococcus neoformans (Cn), Cryptosporidium parvum (Cp), Dictyostelium discoideum (Dd), Entamoeba histolytica (Eh), Giardia lamblia (Gl), Homo sapiens (Hs), Leishmania major (Lm), Plasmodium falciparum (Pf), Saccharomyces cerevisiae (Sc), Schizosaccharomyces pombe (Sp), Tetrahymena thermophilia (Tt), Toxoplasma gondii (Tg), Trichomonas vaginalis (Tv), Trypanosoma brucei (Tb), and Trypanosoma cruzi (Tc). Archaea include Euryarchaeota Archaeoglobus fulgidus (Af), Ferroplasma acidarmanus (Fa), Methanococcoides burtonii (Mb), Methanococcus jannaschii (Mj), Methanopyrus kandleri (Mk), Methanosarcina mazei (Mm), Picrophilus torridus (Pt), Pyrococcus abyssi (Pab), Pyrococcus furiosus (Pfu), Pyrococcus horikoshii (Ph), and Crenarchaeota Pyrobaculum aerophilum (Pae), Sulfolobus solfataricus (Ss), and Sulfolobus tokodaii (St). Bacteria include Actinobacillus actinomycetemcomitans (Aa), Bifidobacterium longum (Bl), Borrelia garinii (Bg), Burkholderia cepacia (Bc), Clostridium acetobutylicum (Ca), Enterococcus hirae (Ehi), Listeria innocua (Li), Staphylococcus aureus (Sa) Streptococcus pneumoniae (Spn), Synechococcus elongates (Se), Thermus thermophilus (Tth), and Tropheryma whipplei (Tw). Not all organisms are present in each tree.
In the case of Alg1, Alg2, and Alg11, which add the 1st-, 2nd-, and 5th-mannose residues to the dolichol-PP-linked precursor, respectively, <50% of each eukaryotic protein is alignable with homologues of prokaryotes, and Alg1, Alg2, and Alg11 each have transmembrane domains that are absent from prokaryotic glycosyltransferases (data not shown). Phylogenetic analyses show eukaryotic Alg1, Alg2, and Alg11 form distinct clades, which are well supported by bootstrap values (Fig. 2B). The relationship of eukaryotic Alg1, Alg2 and Alg11 to archaeal and bacterial glycosyltransferases, however, is unresolved, suggesting that cytosolic mannosylating enzymes are eukaryote-specific and that their precise origins are not clear. These results then do not support the recent hypothesis that the set of cytosolic Alg glycosyltransferases were present in an archaeal ancestor of eukaryotes (24).
In the case of Alg5 and Dpm1 that make dolichol-P-Glc and dolichol-P-Man, respectively (9), phylogenetic analyses show Alg5 and Dpm1 form distinct clades, which are well supported by bootstrap values (data not shown). The Dpm1 clade is itself divided into two clades. Dpm1 clade A contains enzymes with a C-terminal transmembrane helix (TMH) (Saccharomyces, Entamoeba, Trypanosoma, and Leishmania). Dpm1 clade B contains enzymes with no TMH (plants, animals, and fungi) or an N-terminal TMH (Plasmodium). Numerous organisms lacking TMH in their Dpm1 have Dpm2 homologues, which contain two predicted TMH and associate with Dpm1 (data not shown). Conversely, Saccharomyces, Plasmodium, and Entamoeba, which contain TMH in their Dpm1, are missing Dpm2 homologues. Remarkably Trichomonas contains no Dpm1 but has multiple copies of Alg5.
Phylogenetic methods also show that Alg glycosyltransferases in the lumen of the ER, which are unique to eukaryotes but are often similar to each other (e.g., Alg6 and Alg8 or Alg9 and Alg12), may be clearly identified from diverse eukaryotes (data not shown) (15). Alg3 and Alg10, which are present only in eukaryotes, are not similar to other Alg glycosyltransferases. In organisms that contain them, luminal Alg glycosyltransferases are single-copy. These results suggest the possibility (tested in the next four sections) that Alg glycosyltransferase repertoire may be used to correctly predict the dolichol-PP-linked glycans made by each organism.
Sets of Alg Glycosyltransferases Correlate Precisely with Known Dolichol-PP-Linked Glycans. Alg glycosyltransferases were examined first from organisms from which dolichol-PP-linked precursors have been characterized. The slime mold Dictyostelium discoideum, which makes dolichol-PP-GlcNAc2Man9Glc3, contains all 12 Alg glycosyltransferases that have been molecularly characterized (Fig. 1A and Table 1) (1, 3). In contrast, Trypanosoma cruzi, which makes dolichol-PP-GlcNAc2Man9, and Trypanosoma brucei are missing the set of genes encoding glucosylating enzymes in the ER lumen (Fig. 1B) (18). Leishmania major, which causes visceral leishmaniasis, is also missing the Alg12 gene and likely makes dolichol-PP-GlcNAc2Man7. The ciliate Tetrahymena thermophilia, which makes dolichol-PP-GlcNAc2Man5Glc3 as described for Tetrahymena pyriformis (ref. 19; unpublished data), is lacking the set of genes encoding mannosylating enzymes in the ER lumen (Fig. 1C). Plasmodium falciparum, from which it has been difficult to identify N-glycans, is missing all of the Alg glycosyltransferases except Alg7 and STT3 (Fig. 1E) (21, 22). The absence of the other Alg glycosyltransferases makes it likely that putative mannosylated N-glycans of Plasmodium were contaminants from host cells (38). These results suggest that the absence of glycosyltransferase activity (18, 19) is caused by the absence of the gene encoding the enzyme rather than an inactive gene so that sets of predicted Alg glycosyltransferases correlate precisely with experimentally determined dolichol-PP-linked glycans.
Whole-Genome Sequences of Select Protists and Fungi Predict Additional Diversity in Alg Glycosyltransferases and Subsequently Dolichol-PP-Linked Glycans. Like kinetoplastids, the fungus Cryptococcus neoformans is lacking the set of glucosylating enzymes in the ER lumen and is predicted to make dolichol-PP-GlcNAc2Man9 (Fig. 1B, Table 1, and see below) (18). Like Tetrahymena to which it is similar, Toxoplasma gondii and Cryptosporidium parvum are missing the set of luminal mannosylating enzymes (Fig. 1C) (19). Cryptosporidium is also missing Alg8 and Alg10 and likely makes dolichol-PP-GlcNAc2Man5Glc (Table 1). Entamoeba histolytica and Trichomonas vaginalis are missing sets of luminal Alg glycosyltransferases that add Man and Glc to lipid-linked precursors and likely make dolichol-PP-GlcNAc2Man5 (Fig. 1D and see below).
Like Plasmodium, Giardia lamblia is missing all Alg glycosyltransferases except Alg7 and STT3 (Fig. 1E, Table 1, and see below). The presence of Rft1 in all eukaryotes except Giardia, Plasmodium, and Encephalitozoon (see below) supports the idea that Rft1 flips dolichol-PP-GlcNAc2Man5 into the lumen of the ER (8). Why Toxoplasma is also missing Rft1 is not clear.
Encephalitozoon cuniculi, whose genome has been sequenced in its entirety, lacks all Alg glycosyltransferases and is missing STT3 (Fig. 1F and Table 1) (28). Consistent with the absence of N-glycans, Encephalitozoon, like Plasmodium and Giardia, is missing UDP-Glc:glycoprotein glucosyltransferase, glucosidases I and II, calreticulin/calnexin, ERGIC-53, and α-1,2-mannosidases, which operate on N-linked glycans in the ER and Golgi apparatus (unpublished data) (13, 24).
Entamoeba and Trichomonas Make the Predicted Dolichol-PP-GlcNAc2Man5, Whereas Cryptococcus Makes Dolichol-PP-GlcNAc2-Man7-9. The major dolichol-PP-linked glycan of Entamoeba histolytica and Trichomonas vaginalis contains GlcNAc2Man5, which was predicted from the Alg glycosyltransferases of these protists (Fig. 3 A and B). Each peak digests with α-1,2-mannosidase to GlcNAc2Man3 and mannose (data not shown). To show that the labeled dolichol-PP-linked precursor is the same as that transferred to nascent peptides by the OST, membranes of Entamoeba and Trichomonas were incubated with an iodinated tripeptide NYT. As expected, the products of Entamoeba and Trichomonas in vitro comigrate with the GlcNAc2Man5 standard (Fig. 3 D and E). Finally, when Entamoeba and Trichomonas are briefly labeled with 3H-Man in vivo and N-glycans are released with PNGaseF, a major product is GlcNAc2Man5 (data not shown).
Fig. 3.
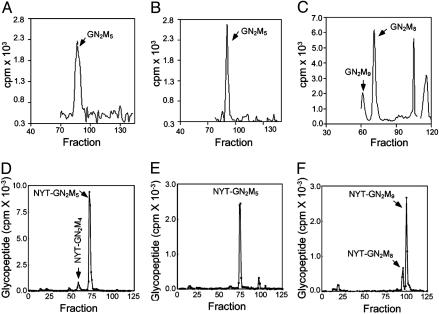
Predicted Trichomonas, Entamoeba, and Cryptococcus dolichol-PP-glycans were identified in vivo and in vitro. Dolichol-PP-linked precursors from Trichomonas vaginalis (A), Entamoeba histolytica (B), and Cryptococcus neoformans (C), as well as in vitro OST assays by using membranes from Trichomonas (D), Entamoeba (E), and Cryptococcus (F). In A-C, glycans were labeled in vivo with [3H]Man and separated on a P-4 column, whereas in D-F, glycans were transferred to a radio-iodinated tripeptide NYT in vitro, captured with Con A, and separated by HPLC.
Although we have not isolated each Alg enzyme individually and have not shown its glycosyltransferase activity in vitro, correct predictions of previously uncharacterized dolichol-PP-linked glycans of Entamoeba and Trichomonas strongly suggest each Alg glycosyltransferase is functioning as expected. The presence of UDP-Glc:glycoprotein glucosyltransferase, glucosidase II, calreticulin/calnexin, and ERGIC-53 in Entamoeba and Trichomonas (unpublished data) suggests these enzymes and/or lectins function with N-glycans built on GlcNAc2Man5 rather than the usual GlcNAc2Man9 (13, 24).
When the fungus Cryptococcus neoformans is labeled with 3H-Man, the major peak on P-4 runs with GlcNAc2Man7-8, whereas a minor peak runs with GlcNAc2Man9, which was predicted from their complement of Alg genes (Fig. 3C). However, GlcNAc2Man9 is the most abundant glycan transferred to the iodinated peptide in vitro (Fig. 3F).
Giardia Dolichol-PP- and N-Linked Glycans Contain GlcNAc1-2. No Giardia lamblia dolichol-PP- or N-linked glycans are labeled with 3H-Man, consistent with the absence of cytosolic Alg glycosyltransferases that add Man to dolichol-PP-linked precursors (Fig. 1E and Table 1). In contrast, when Giardia is labeled with 3H-GlcN, dolichol-PP- and N-linked glycans include GlcNAc and GlcNAc2 (diacetylchitobiose) (solid lines in Figs. 4A and 4B). As expected, Giardia dolichol-PP- and N-linked GlcNAc2 are cleaved with chitobiase to GlcNAc (dotted lines in Fig. 4 A and B). These results, which suggest there is no further modification of N-glycans in the Golgi apparatus of Giardia, are consistent with (i) binding of wheat germ agglutinin to the surface of Giardia (39), and (ii) the prediction that ≈90 secreted proteins of Giardia have ≥10 predicted sites for N-linked glycosylation (unpublished data) (10). These results suggest that the Alg enzyme that makes dolichol-PP-GlcNAc2, which has not yet been molecularly characterized, is present in Giardia (1).
Fig. 4.
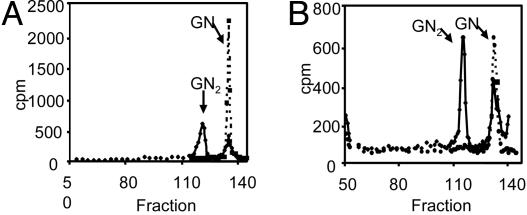
Giardia dolichol- and N-linked glycans are composed of GlcNAc and GlcNAc2. Dolichol-PP-linked glycans (A) and N-linked glycans (B) of Giardia lamblia, each labeled in vivo with [3H]GlcN and separated on a P-4 column. Solid lines indicate labeled products, whereas dotted lines indicate products of digestion of the excised GlcNAc2 peak with chitobiase. Giardia dolichol- and N-linked glycans are composed of GlcNAc and GlcNAc2.
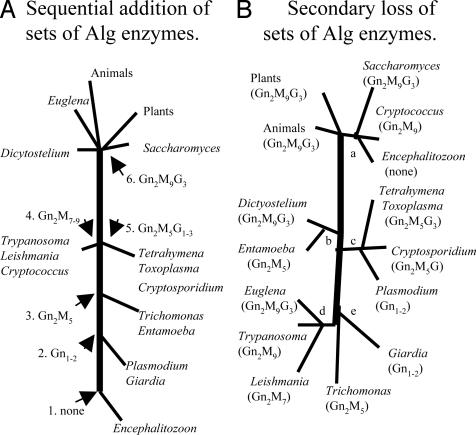
Origins of Eukaryotic Alg Glycosyltransferase Diversity. The present diversity of fungal and protist precursors for N-glycosylation may have resulted from development of an increasingly complex series of Alg glycosyltransferases with eukaryotic evolution (Fig. 5A) or from secondary loss of sets of Alg glycosyltransferases (Fig. 5B) (23, 24). Fig. 5A is drawn so that organisms with the least complex N-glycans are at the bottom, whereas those with the most complex glycans are at the top. Each step in Fig. 5A indicates the addition of a particular set of sugars to the N-glycan precursor. For example, an ancestral eukaryote like Encephalitozoon lacked all dolichol-PP-linked precursors (step 1) until STT3 and Alg7 were obtained to make organisms like Giardia and Plasmodium (step 2). Subsequently cytosolic mannosylating enzymes were added to make ancestors resembling Entamoeba and Trichomonas (step 3), followed by the addition of luminal mannosylating enzymes as in Trypanosoma and Cryptococcus (step 4) (18) or of luminal glucosylating enzymes as in Tetrahymena and Cryptosporidium (step 5) (19). The final result was organisms with an entire set of Alg glycosyltransferases and a complete 14-sugar dolichol-PP-linked precursor (step 6) as in Saccharomyces, Euglena, Dictyostelium, animals, and plants (1).
Fig. 5.
The present diversity of protist and fungal dolichol-PP-lined glycans appears to result from secondary loss glycosyltransferases from a common ancestor that contained the complete set of Alg glycosyltransferases. Models suggesting sequential addition (A) or secondary loss (B) of Alg glycosyltransferases during eukaryotic evolution are shown. Nodes, which are labeled numerically in A and alphabetically in B, are explained in the text.
The difficulties with the Fig. 5A model are (i) Alg7 and STT3 appear to have been present in an ancestor to both eukaryotes and prokaryotes and should be present in Encephalitozoon (16), (ii) it is impossible to determine whether luminal mannosylation (step 4) came before or after luminal glucosylation (step 5), and (iii) the model is in disagreement with rRNA and protein phylogenies, which do not place Encephalitozoon at the base of the phylogenetic tree and do not pair Giardia with Plasmodium, Entamoeba with Trichomonas, Trypanosoma with Cryptococcus, or Saccharomyces with Euglena (40-42).
In the Fig. 5B model, which groups organisms according to rRNA and protein phylogenies, the distribution of Alg glycosyltransferases is best rationalized by secondary loss (23). At node a (fungi), there is loss of luminal glucosylating enzymes in Cryptococcus and all Alg glycosyltransferases in Encephalitozoon, whereas at node b (amebozoa), all luminal Alg glycosyltransferases are lost from Entamoeba. At node c (ciliates and apicomplexa), some luminal glucosylating enzymes are lost from Cryptosporidium, whereas all luminal glucosylating enzymes and all cytosolic mannosylating enzymes are lost from Plasmodium. At node d (kinetoplastids and Euglena), there is loss of luminal glucosylating enzymes from Trypanosoma and an additional loss of one mannosylating enzyme from Leishmania major. If one assumes that Trypanosoma, Giardia, and Trichomonas all branched at the same time from the base of the tree (node e), there are additional secondary losses from Giardia and Trichomonas. Finally, if one assumes the “big bang” hypothesis for eukaryotic origins (43), the common ancestor must have had all of the Alg glycosyltransferases, and all of the differences among extant eukaryotes are due to secondary loss.
A hybrid of the Fig. 5 models, which we cannot rule out because of poor resolution at the base of the eukaryotic phylogenetic tree, suggests that a common eukaryotic ancestor contained Alg7 and STT3. In this hybrid model, Giardia branched off before acquisition of mannosylating and glucosylating enzymes. Still, secondary losses of Alg glycosyltransferases at nodes a-d remain a major factor in the diversity of N-glycan precursors among protists and fungi. Similarly, secondary loss explains the absence of most mitochondrial function in microaerophilic protists such as Giardia, Entamoeba, and Trichomonas (44, 45).
Significance. The bioinformatic approach here allowed us to quickly and accurately predict the dolichol-PP-glycans made by each protist and fungus, which we confirmed for free-living organisms (Giardia, Entamoeba, Trichomonas, and Cryptococcus) and have yet to confirm for intracellular pathogens (Plasmodium, Toxoplasma, Cryptosporidium, and Encephalitozoon). Although it has previously been assumed that the diversity of eukaryotic N-glycans results primarily from differential modification of a common GlcNAc2Man9Glc3 precursor in the ER and Golgi apparatus (1, 24), these results suggest that there are major differences in the dolichol-PP-glycans transferred to the nascent peptide. Secondary losses of Alg glycosyltransferases and mitochondrial function (44, 45) suggest all extant eukaryotes may derive from a relatively complex last common ancestor, and that simple, deeply branching eukaryotes with a primary absence of important biochemical pathways may no longer exist (23). It remains to be determined how the diversity of dolichol-PP-glycans effects OST function, protein-folding in the ER, and modification of glycans in the Golgi apparatus, as well as the antigenicity of glycoproteins on surfaces of these important human pathogens.
Acknowledgments
We thank Ann-Marie Surette and Charles Specht for help with protist and fungal cultures; Kosuke Hashimoto for help with collection and alignment of Alg sequences; Prashanth Vishwanath for advice on phylogenetic methods; investigators at The Institute for Genomic Research and The Sanger Institute for release of preliminary sequence data for numerous protists and fungi; and Temple Smith and Armando Parodi for their comments on this manuscript. This work was supported in part by National Institutes of Health Grants AI44070 and AI48082 (to J.S.), GM43768 (to R.G.), and GM31318 (to P.W.R.).
Abbreviations: Alg, Asn-linked glycan; OST; oligosaccharyltransferase; TMH, transmembrane helix.
References
- 1.Burda, P. & Aebi, M. (1999) Biochim. Biophys. Acta. 1426, 239-257. [DOI] [PubMed] [Google Scholar]
- 2.de la Canal, L. & Parodi, A. J. (1985) Comp. Biochem. Physiol. B Biochem. Mol. Biol. 81, 803-805. [Google Scholar]
- 3.Ivatt, R. L., Das, O. P., Henderson, E. J. & Robbins, P. W. (1984) Cell 38, 561-567. [DOI] [PubMed] [Google Scholar]
- 4.Huffaker, T. & Robbins, P. W. (1983) Proc. Natl. Acad. Sci. USA 80, 7466-7470. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Reiss, G., te Heesen, S., Zimmerman, J., Robbins, P. W. & Aebi, M. (1996) Glycobiology 6, 493-498. [DOI] [PubMed] [Google Scholar]
- 6.Cipollo, J. F., Trimble, R. B., Chi, J. H., Yan, Q. & Dean, N. (2001) J. Biol. Chem. 276, 21828-21840. [DOI] [PubMed] [Google Scholar]
- 7.Aebi, M. & Hennet, T. (2001) Trends Cell Biol. 11, 136-141. [DOI] [PubMed] [Google Scholar]
- 8.Helenius, J., Ng, D. T., Marolda, C. L., Walter, P., Valvano, M. A. & Aebi, M. (2002) Nature 415, 447-450. [DOI] [PubMed] [Google Scholar]
- 9.Orlean, P., Albright, C. & Robbins, P. W. (1988) J. Biol. Chem. 263, 17499-17507. [PubMed] [Google Scholar]
- 10.Kornfeld, R. & Kornfeld, S. (1985) Annu. Rev. Biochem. 54, 631-664. [DOI] [PubMed] [Google Scholar]
- 11.Silberstein, S. & Gilmore, R. (1996) FASEB J. 10, 849-858. [PubMed] [Google Scholar]
- 12.Yan, Q. & Lennarz, W. J. (2002) J. Biol. Chem. 277, 47692-47700. [DOI] [PubMed] [Google Scholar]
- 13.Parodi, A. J. (2000) Biochem. J. 348, 1-13. [PMC free article] [PubMed] [Google Scholar]
- 14.Lechner, J. & Wieland, F. (1989) Annu. Rev. Biochem. 58, 173-194. [DOI] [PubMed] [Google Scholar]
- 15.Oriol, R., Martinez-Duncker, I., Chantret, I., Mollicone, R. & Codogno, P. (2002) Mol. Biol. Evol. 19, 1451-1463. [DOI] [PubMed] [Google Scholar]
- 16.Wacker, M., Linton, D., Hitchen, P. G., Nita-Lazar, M., Haslam, S. M., North, S. J., Panico, M., Morris, H. R., Dell, A., Wren, B. W. & Aebi, M. (2002) Science 298, 1790-1793. [DOI] [PubMed] [Google Scholar]
- 17.Guha-Niyogi, A., Sullivan, D. R. & Turco, S. J. (2001) Glycobiology 11, 45R-59R. [DOI] [PubMed] [Google Scholar]
- 18.Parodi, A. J. (1993) Glycobiology 3, 193-199. [DOI] [PubMed] [Google Scholar]
- 19.Yagodnik, C., de la Canal, L. & Parodi, A. J. (1987) Biochemistry 26, 5937-5943. [Google Scholar]
- 20.Adam, R. D. (2001) Clin. Microbiol. Rev. 14, 447-475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Berhe, S., Gerold, P., Kedees, M. H., Holder, A. A. & Schwarz, R. T. (2000) Exp. Parasitol. 94, 194-197. [DOI] [PubMed] [Google Scholar]
- 22.Gowda, D. C., Gupta, P. & Davidson, E. A. (1997) J. Biol. Chem. 272, 6428-6439. [DOI] [PubMed] [Google Scholar]
- 23.Dacks, J. B. & Doolittle, W. F. (2001) Cell 107, 419-425. [DOI] [PubMed] [Google Scholar]
- 24.Helenius, A. & Aebi, M. (2004) Annu. Rev. Biochem. 73, 1019-1049. [DOI] [PubMed] [Google Scholar]
- 25.Abrahamsen, M. S., Templeton, T. J., Enomoto, S., Abrahante, J. E., Zhu, G., Lancto, C. A., Deng, M., Liu, C., Widmer, G., Tzipori, S., et al. (2004) Science 304, 441-445. [DOI] [PubMed] [Google Scholar]
- 26.Altschul, S. F., Madden, T. L., Schaffer, A. A., Zhang, J., Zhang, Z., Miller, W. & Lipman, D. J. (1997) Nucleic Acids Res. 25, 3389-3402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Gardner, M. J., Hall, N., Fung, E., White, O., Berriman, M., Hyman, R. W., Carlton, J. M., Pain, A., Nelson, K. E., Bowman, S. et al. (2002) Nature 419, 498-511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Katinka, M. D., Duprat, S., Cornillot, E., Metenier, G., Thomarat, F., Prensier, G., Barbe, V., Peyretaillade, E., Brottier, P., Wincker, P., et al. (2001) Nature 414, 450-453. [DOI] [PubMed] [Google Scholar]
- 29.McArthur, A. G., Morrison, H. G., Nixon, J. E. J., Passamaneck, N. Q. E., Kim, U., Hinkle, G., Crocker, M. K., Holder, M. E., Farr, R., Reich, C. I., et al. (2000) FEMS Microbiol. Lett. 189, 271-273. [DOI] [PubMed] [Google Scholar]
- 30.Mewes, H. W., Albermann, K., Bahr, M., Frishman, D., Gleissner, A., Hani, J., Heumann, K., Kleine, K., Maierl, A., Oliver, S. G., et al. (1997) Nature 387, 7-65. [DOI] [PubMed] [Google Scholar]
- 31.Kall, L., Krogh, A. & Sonnhammer, E. L. (2004) J. Mol. Biol. 338, 1027-1036. [DOI] [PubMed] [Google Scholar]
- 32.Thompson, J. D., Higgins, D. G. & Gibson, T. J. (1994) Nucleic Acids Res. 22, 4673-4680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Jones, D. T., Taylor, W. R. & Thornton, J. M. (1992) Comput. Appl. Biosci. 8, 275-282. [DOI] [PubMed] [Google Scholar]
- 34.Strimmer, K. & Von Haeseler, A. (1997) Proc. Natl. Acad. Sci. USA 94, 6815-6819. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.McConville, M. J., Thomas-Oates, J. E., Ferguson, M. A. J. & Homans, S. W. J. Biol. Chem. 265, 19611-19623. [PubMed]
- 36.Kelleher, D. J., Kreibich, G. & Gilmore, R. (1992) Cell 69, 55-65. [DOI] [PubMed] [Google Scholar]
- 37.Kelleher, D. J., Karaoglu, D. & Gilmore, R. (2001) Glycobiology 11, 321-333. [DOI] [PubMed] [Google Scholar]
- 38.Kimura, E. A., Couto, A. S., Peres, V. J., Casal, O. L. & Katzin, A. M. (1996) J. Biol. Chem. 271, 14452-14461. [DOI] [PubMed] [Google Scholar]
- 39.Ortega-Barria, E., Ward, H. D., Evans, J. E. & Pereira, M. E. (1990) Mol. Biochem. Parasitol. 43, 151-165. [DOI] [PubMed] [Google Scholar]
- 40.Baldouf, S. L. (2003) Science 300, 1703-1706. [DOI] [PubMed] [Google Scholar]
- 41.Bapteste, E., Brinkmann, H., Lee, J. A., Moore, D. V., Sensen, C. W., Gordon, P., Durufle, L., Gaasterland, T., Lopez, P., Müller, M. & Philippe, H. (2002) Proc. Natl. Acad. Sci. USA 99, 1414-1419. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Sogin, M. L. & Silberman, J. D. (1998) Int. J. Parasitol. 28, 11-20. [DOI] [PubMed] [Google Scholar]
- 43.Philippe, H., Germot, A. & Moreira, D. (2000) Curr. Opin. Genet. Dev. 10, 596-601. [DOI] [PubMed] [Google Scholar]
- 44.Embley, T. M., van der Giezen, M., Horner, D. S., Dyal, P. L. & Foster, P. (2003) Philos. Trans. R. Soc. Lond. B. 358, 191-201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Mai, Z., Ghosh, S., Frisardi, M., Rosenthal, B., Rogers, R. & Samuelson, J. (1999) Mol. Cell. Biol. 19, 2198-2205. [DOI] [PMC free article] [PubMed] [Google Scholar]