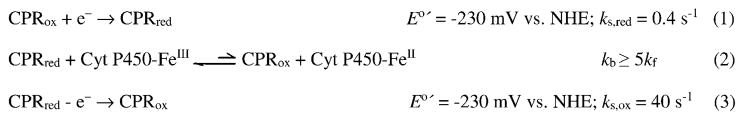Abstract
This article reviews recent progress from our laboratory in electrochemiluminescence (ECL) arrays designed for screening toxicity-related chemistry of chemical and drug candidates. Cytochrome P450s and metabolic bioconjugation enzymes convert lipophilic chemicals in our bodies by oxidation and bioconjugation that can lead to toxic metabolites. DNA can be used as an easily measurable toxicity-related endpoint, targeting DNA oxidation and addcut formation with metabolites. ECL using guanosines in the DNA strands as co-reactants have been used in high throughput arrays utilizing DNA-enzyme films fabricated layer-by-layer. This review describes approaches developed to provide new high throughput ECL arrays to aid in toxicity assessment for drug and chemical product development.
Keywords: Metabolites, Toxicity Chemistry, Electrochemiluminescence, DNA Arrays, DNA Damage, Cytochrome P450s
1 Introduction
Organic chemicals used in agriculture, personal care, home applications industry and elsewhere are potential sources of disease-causing substances [1]. Metabolites of chemicals generated by cytochrome P450 (cyt P450) enzymes [2] and bioconjugation enzymes in the human body can form covalent adducts with nucleobases of DNA and may cause DNA oxidation in major pathways of chemical toxicity [3]. While only a fraction of these metabolites are acutely toxic, a wide range of substances cause metabolite-related damage to DNA, proteins and other biomolecules. DNA oxidation can occur due to attack by reactive oxygen species (ROS) [4,5] including singlet oxygen, superoxide and hydroxyl radicals to form most notably 8-oxo-2-deoxyguanosine on DNA strands [6]. Alternatively, formation of adducts usually involves SN2 coupling reactions of DNA nucleobases with electrophilic metabolites (Scheme 1).
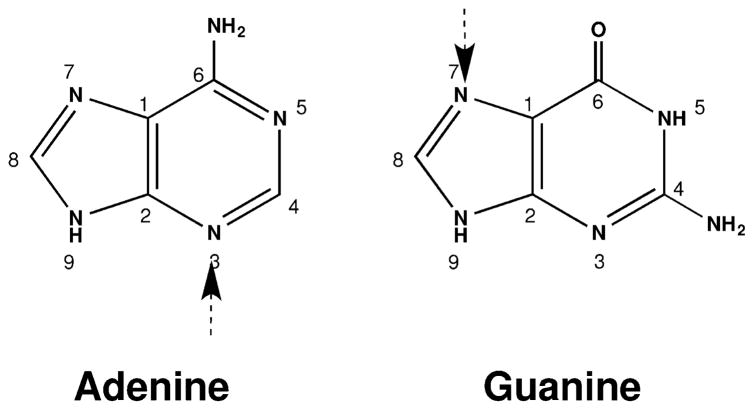
Scheme 1.
Predominant DNA adduct formation sites at adenine and guanine.
We have developed approaches to toxicity screening of chemicals by monitoring the DNA damage, mainly metabolite-related adduct formation and oxidation. This review focuses on clectrochemiluminescence (ECL) arrays to screen for possible toxicity chemistry. These toxicity screening arrays feature metabolite generation in thin films and require sensor film architecture that combines metabolic enzymes, an ECL dye that is usually a polymer, and polyion “glue” to hold the film together [7,8]. In order to develop devices for metabolic toxicity screening, nearly all of our approaches are based on films formed layer-by-layer (LbL) in microwells on the arrays. The general operation of these array devices requires 2 steps. First the test molecules are reacted with metabolic enzymes and, metabolites formed may react to damage DNA in the films [9–12]. Second, DNA damage is monitored by ECL [13–18]. Voltammetry can also be used [19–23], but is more restrictive on array size and is not covered in this review.
When molecules or their metabolites can damage DNA, they are called genotoxic molecules. This term is applied to both parent compounds and their metabolites, even when only the metabolite is the actual damaging agent. Once identified by these arrays, toxicity-related pathways of genotoxic chemicals can be assessed using magnetic bead bioreactors coupled with HPLC-MS/MS [7,8,24].
In section 2 below, we briefly describe sensors based on cyt P450 enzyme and DNA, since this work led up to the eventual development of the present ECL arrays. In section 3, we discuss microfluidic ECL arrays and provide illustrative examples of high throughput metabolic screening for potentially toxic chemistry. In section 4, we describe an approach to measure DNA oxidation using ECL arrays. In the concluding section, we summarize key features, and discuss prospects for the future.
2 Electrochemical Toxicity Biosensors
2.1 Cytochrome P450-containing Biosensors
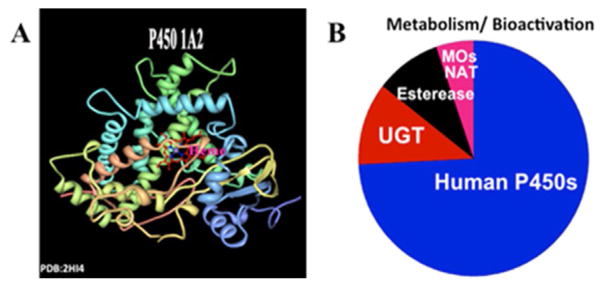
Cyt P450s are a key human iron heme enzyme superfamily involved in 75% or more of oxidative metabolic reactions in the human liver and other organs (Scheme 2) [25,26]. Cyt P450s enzymes are present in nearly all organs of the human body and they are integral to the metabolism of drugs and other xenobiotics [2]. On the catalytic pathway from the parent compound to the metabolic product, cyt P450s can mediate formation of reactive species that react with DNA.
Scheme 2.
Ribbon structure of (A) human cyt P450 1A2 and (B) the role of cyt P450s and selected bioconjugation enzymes in human metabolism.
2.1.1 Catalytic Pathway of Cyt P450
Catalytic reactions and reductive electrochemistry of cyt P450s both center on the iron heme prosthetic group. The natural catalytic cycle of cyt P450 involves a complex pathway that features donation of electrons from NADPH via cytochrome P450 reductase (CPR) resulting in the reduction of cyt P450-FeIII to cyt P450-FeII. Dioxygen then binds to cyt P450-FeII, followed by events including a second electron reduction by NADPH-CPR and protonation to yield a cyt P450-FeIII-hydroperoxo complex. This species becomes protonated and loses water to produce the presumed catalytically active ferryloxy species P450-FeIV=O that transfers oxygen to bound substrate (e.g. RH) to form product (e.g. ROH). Cytochrome b5 serves as a second electron donor for some cyt P450s [27].
2.1.2 Cyt P450 Catalysis on Electrodes
Bioelectronic initiation of catalytic reactions of cyt P450s in films on electrodes can simplify applications by replacing electron donation from the expensive NADPH reductant with electron donation from an electrical circuit. In order to electrochemically activate cyt P450s in toxicity screening applications, we spent considerable effort investigating its fundamental voltammetric properties in thin films. We previously reviewed research on cyt P450 catalysis in films on electrodes [28]. Our early focus was achieving thin LbL film environments to facilitate reversible FeIII/FeII electron transfer (ET) of cyt P450 on an electrode surface [29–31]. However, this reduction in the natural catalytic pathway of cyt P450s is achieved by ET from cyt P450 reductase (CPR) that has previously accepted electrons from NADPH. To mimic the natural catalytic pathway, we need to send electrons first to CPR in a film that contained cyt P450s.
We explored ways to place proteins in film environments that facilitate electron exchange with underlying electrodes for electrode-driven cyt P450 biocatalysis [28] including (1) adsorbed DNA films that bind cyt P450s, (2) cyt P450s in multilayered films of insoluble lipids [29–30]; and (3) LbL films of cyt P450s made with polyion films that can also contain DNA. Gilardi and coworkers developed fusion proteins in which an initial electron acceptor protein is linked to a cyt P450 to activate the cyt P450 FeIII. In these systems, electrons flow from the electrode, to the initial electron acceptor and then to the cyt P450 [32]. Catalytic activity in these fusion proteins can be optimized by controlling the electron transfer rate due to longer lived cyt P450 iron-peroxy species [33]. Our most successful approach for a natural catalysis pathway employed microsomal CPR/cyt P450 films on electrodes with a large excess of cyt P450 over CPR, similar to the situation in vivo. Cyclic voltammetry (CV) of this system revealed that it closely mimics the details of the natural cyt P450 catalytic pathway [34]. We also found that a similar pathway can be accessed in microsomes containing multiple cyt P450s and CPR [35]. We showed by CV that electrons in these systems flow from the electrode to CPR and then to the cyt P450, the same as in the in vivo pathway. Spectroscopic studies of CO binding showed that cyt P450s are in the native form in these films. The model best fitting the data includes an equilibrium redox reaction involving a CPR-cyt P450 complex that facilitates efficient catalytic turnover of the excess cyt P450s. Microsomal CPR/cyt P450 films gave midpoint potentials near −490 mV vs. SCE (−250 mV vs. NHE) (Figure 1A). The measured CV potentials were not influenced by CO, and ks values were near 40 s−1. Midpoint potentials and ks values were consistent with that of CPR alone in similar films. With only cyt P450s in the films (no CPR), midpoint potentials were very different at −340 mV vs. SCE (−100 mV vs. NHE) (Figure 1B), ~40 mV shifts were found with added CO which only happens with iron heme proteins but not CPR, and ks was 2–18 s−1 [34].
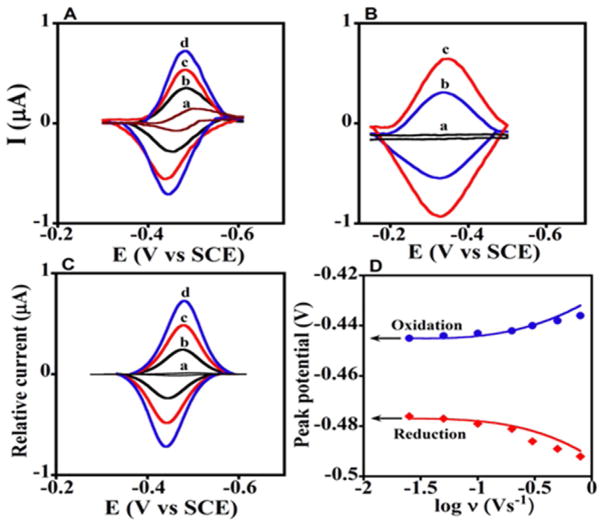
Fig. 1.
CVs of microsomal LbL films cyt P450 1A2 with and without CPR on PG electrodes. (A) Background subtracted CVs of {a} CPR films at 0.3 Vs−1 with no cyt P450, and cyt P450 1A2/CPR films at scan rates {b} 0.1, {c} 0.2, and {d} 0.3 Vs−1. (B) Background subtracted CVs of cyt P450 1A2 film with no CPR {a} polyion film control; 0.1 Vs−1 and {b} Cyt P450 films at 0.1 and {c} 0.2 Vs−1. (C) Digitally simulated CVs corresponding to {a} reversible electron transfer for only CPR film at 0.3 Vs−1, and {b to d} the ErCEo-model using parameters in Scheme 2 for cyt P450 1A2/CPR films at scan rates {b} 0.1, {c} 0.2, and {d} 0.3 Vs−1 showing excellent agreement with experimental CVs in Figure 1A. (D) Influence of scan rate on oxidation (blue circles) and reduction (red diamonds) peak potentials for cyt P450 1A2/CPR films with theoretical peak potentials (lines) simulated using the ErCEo model. Reproduced with permission from ref. [35] American Chemical Society, copyright 2011.
Peak currents were much larger for CPR/cyt P450 films (Figure 1A) than for CPR alone. This is explained by Scheme 3. CPR is reduced by the electrode but cyt P450s are not (eq. 1), CPRred exists in a redox equilibrium with cyt P450-FeIII to yield CPRox as a product (eq. 2) and electrochemical oxidation of CPRred (eq. 3) competes with this equilibrium. We used Scheme 3 to accurately simulate CVs (compare Figures 1A and 1C) and trumpet plots (Figure 1D) for CPR/cyt P450 films after establishing the best fit parameters shown in Scheme 3. Thus, the microsomal CPR/cyt P450 film is a model for the natural catalytic pathway [34].
Scheme 3.
ErCEo simulation model for CVs of cyt P450s+ CPR.
Catalytic oxidation of classic cyt P450 substrate 4-(methylnitrosamino)-1-(3-pyridyl)-1-butanone (NNK) with the CPR/cyt P450 films further confirmed the electrode→CPR→cyt P450 pathway. The rate of formation of the product of this reaction in bulk electrolysis was slightly larger than that achieved in the same system using NADPH to deliver electrons [35].
CV simulations based on Scheme 3 serve to characterize equilibrium between cyt P450 and CPR (eqn. 1), which most likely features a complex within which electron transfer occurs [32]. The best fit was found for kb ≥ 5kf and K = kf/kb ≤ 0.2 lies in favor of CPRred (in absence of O2), this is why CPRox is reduced electrochemically in cyt P450/CPR films before cyt P450. The difference in formal potential (ΔE0) between CPR and cyt P450 is ΔE0=[RT/nF] ln(K), where R is the gas constant, T is absolute temperature, and F is Faraday’s constant. Thus, K<0.2 predicts that cyt P450 formal potential is ~40 mV negative of CPRox in the films at 25 °C, so that CPRox is the more the easily reduced protein in the film. O2 present in catalytic oxidations reacts with P450-FeII and the equilibrium in eq. 2 shifts to the right to help drive catalytic conversion when the substrate is present [34].
3 ECL Arrays
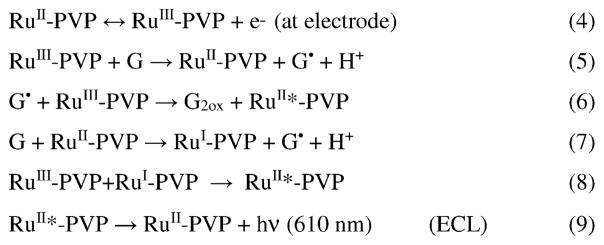
Using ECL to measure DNA damage provides a sensitive and a simpler approach to arrays than other electrochemical detection methods because it does not require individually addressable sensors. Arrays can thus be designed with a single patterned working electrode as we discuss below. A useful ECL scheme for sensors includes a high energy redox process involving Ru(bpy)32+ dye (RuBPY) and co-reactant tri-propylamine (TprA) that is driven electrochemically to emit ECL light [36,37]. A charge-coupled device (CCD) camera can be used to detect ECL light from arrays. We showed in 2003 [14] that guanine bases in intact DNA can serve as ECL co-reactants for metallopolymer (bis-2,2′-bipyridyl) ruthenium polyvinyl-pyridine([Ru(bpy)2(PVP)10]2+ or Ru-PVP) to produce visible light (Scheme 4), thus replacing TprA. This opened the door to ECL arrays for toxicity screening based on DNA damage by metabolites, since structural disorder in the damaged nucleotide provided better access of RuIII sites and faster reaction with guanines. This in turn provides more ECL light for damaged DNA (Scheme 4). Oxidation of RuII-PVP at ~1.2 V vs. Ag/AgCl generates RuIII-PVP that catalytically oxidizes guanines to guanine radicals (G•). RuIII-PVP metallopolymer reacts with G• on the DNA to produce electronically excited RuII*-PVP, which in turn emits visible ECL light at 610 nm. An alternative pathway involves generation of RuI-PVP, that reacts with RuIII-PVP to give RuII*-PVP (eq. 7–9).
Scheme 4.
ECL pathways using DNA as co-reactant, G=guanine.
3.1 Arrays for Genotoxicity Chemistry
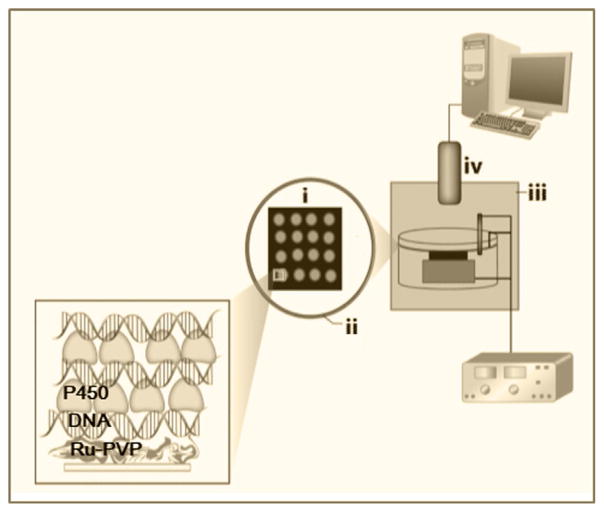
We first developed single sensors [3,14] and manually run ECL arrays [7,13,15,16] aimed at identifying possibly toxic pathways for test chemical and based on thin LbL films of DNA, RuPVP, and metabolic enzymes (Scheme 5). This work progressed to microfluidic approaches that partly automate the reaction chemistry [8,18]. The microsomal cyt P450/CPR films enabled electrochemical bioactivation of the natural cyt P450 catalytic cycle in these arrays [34]. Higher-throughput array formats for ECL-based toxicity sensors were fabricated on a single pyrolytic graphite (PG) wafer serving as an enzyme reactor and analytical electrode with spatially separated spots (Scheme 6). The reactor chip of conductive pyrolytic graphite (PG) has 64 microwells created by heat-transfer from a computer printed pattern onto the PG. These microwells made from computer toner ink have ~10 nm high hydrophobic sides that help to hold 1 μL solution droplets in the microwells during adsorption of LbL film layers (Scheme 6c). Individual spots containing layers of DNA, metabolic enzyme and ECL dye Ru-PVP are deposited into the wells on the chip, which is then connected to a copper “shoe” in a microfluidic housing via silver epoxy, and insulated with acrylic polymer so only the conductive PG face is exposed.
Scheme 5.
Schematic representation of sensor system for ECL detection, showing microwell-patterned PG array electrode the DNA-enzyme-Ru-PVP spots with Ag/AgCl reference (i) and Pt-wire counter electrodes (ii). For ECL readout, the device is placed into a dark box (iii) and connected to a potentiostat that applies the necessary voltage while a charge coupled device camera (CCD) detects the light (iv). A computer is interfaced for data capture and analysis.
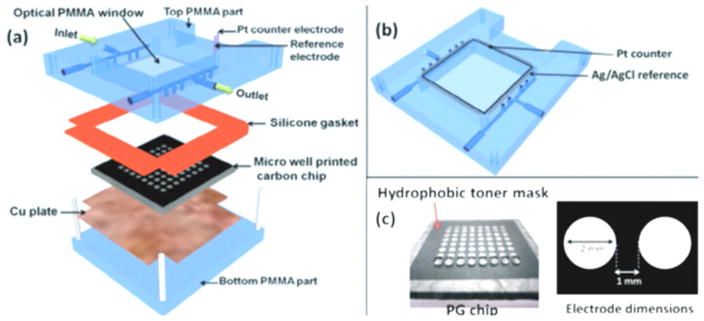
Scheme 6.
ECL microfluidic array assembly into flow cell: (a), underside view of reference and counter electrode wires in the top poly(methylmethacrylate) (PMMA) plate: (b) pyrolytic graphite (PG) chip with printed microwells in (c). The first row is shown containing 1 μL water droplets. The reaction system is operated by connection to a syringe pump. Reproduced with permission from ref. [18] The Royal Society of Chemistry, copyright 2013.
Enzyme/RuPVP/DNA films are grown in the micro-wells by the LbL method using 1 μL drops of each adsorbate solution successively for 20–30 min at 4°C in a humidified chamber, with intermediate washing. Sources of enzymes for the films include human liver microsomes (HLM), microsomal epoxide hydrolase (EH), human cytosol, pooled human liver S9 enzyme fractions (HS9), supersomes containing a specific cyt P450 and CPR, and rat liver microsomes (RLM). Microsomes typically contain a collection of cyt P450s and CPR. Additional bioconjugation enzymes can be added at will. Mass, amount of enzyme, and nominal thickness are estimated using a quartz crystal microbalance (QCM).
This completed system is essentially a microfluidic reactor operated by introducing enzyme cofactors, oxygen, and reactants to the microwell chip in the microfluidic flow cell (Scheme 6) at a constant flow rate using a syringe pump. Microsomal cyt P450s are activated electrochemically in the electrode→CPR→cyt P450 pathway [18]. The metabolic enzyme reactions are allowed to proceed for the desired time, and then the array is washed with buffer. ECL light developed at 1.25 V vs. Ag/AgCl is detected by a CCD camera for 180 s in a dark box.
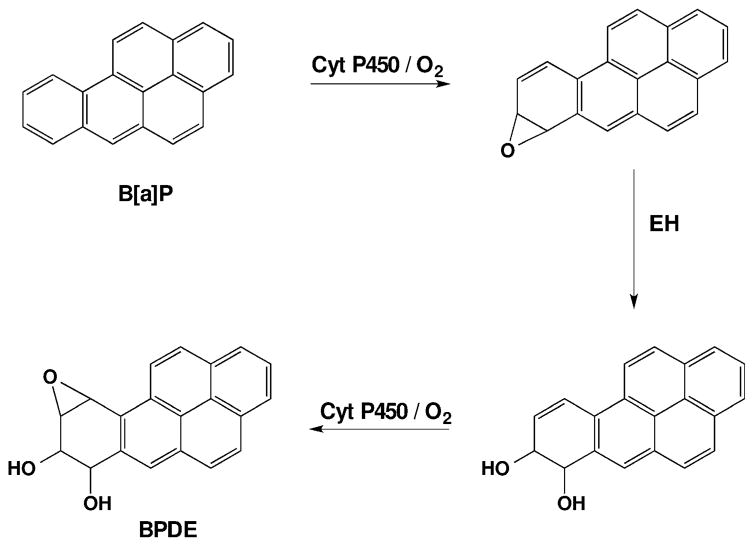
The microfluidic array reactor was first tested using benzo[a]pyrene (B[a]P), which is oxidized by cyt P450s to give B[a]P-7,8-epoxide [18]. Hydrolysis of B[a]P-7,8-ep-oxide catalyzed by the enzyme epoxide hydrolase (EH) yields B[a]P-7,8-dihydrodiol. Subsequent oxidation of this dihydrodiol catalyzed by cyt P450s results in B[a]P-7,8-di-hydrodiol-9,10-epoxide (BPDE) (Scheme 7). BPDE is the major reactant that forms adducts with guanine and adenine bases in DNA. Enzymes used for these experiments included human liver microsomes (HLM), rat liver microsomes (RLM), human microsomal epoxide hydrolase (EH), pooled human liver S9 enzyme fractions (HS9), and supersomes of cyt P450 1B1 and 1A2. Slightly larger relative rates of DNA damage were found when EH was included in the spots with cyt P450 enzymes, suggesting formation of the major damage agent BPDE when both cyt P450 and EH are present, leading to more DNA damage (Figure 2). LC-MS/MS analysis of products of magnetic bead reactors coated with the same enzyme/DNA films as in the arrays resulted in product ion spectra of dG-BPDE and dA-BPDE that confirmed formation of these adducts when cyt P450 and EH were on the beads. The arrays were also used to study inhibition of cyt P450 enzymes by measuring the decrease in DNA damage in the presence of known inhibitors. Results were consistent with reported enzyme activities and inhibition [18].
Scheme 7.
Metabolic reactions of benzo[a]pyrene leading to DNA-reactive metabolite BPDE.
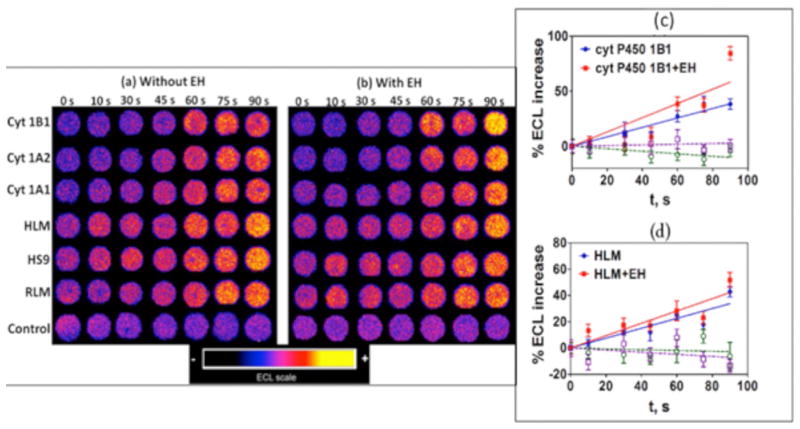
Fig. 2.
ECL array results for oxygenated 25 μM B[a]P in pH 7.4 phosphate buffer with electrochemical activation of cyt P450s at −0.65 V vs. Ag/AgCl (0.14 M KCl): (a and b) Reconstructed ECL array data from RuIIPVP/enzyme/DNA spots for reaction times 0–90 s. Control spots contained cyt P450 1B1 without (a) and with (b) EH subjected to the same reaction conditions as above without activation of cyt P450s. (c and d) Influence of enzyme reaction time on ECL increase for (c) cyt P450 1B1, and (d) human liver microsomes (HML). Controls for cyt P450s (green) and cyt P450s+EH (purple) behaved equivalently without substrate or with substrate but no activation of cyt P450s. Reproduced from ref. [18] with permission of the Royal Society of Chemistry, copyright 2013.
3.2 Multiple Component Organ Specific Enzymes Arrays
Standard toxicity bioassays typically rely on liver enzyme metabolism. However, other tissues can also metabolize the drugs and xenobiotic chemicals and contribute significantly to toxic reactions. The 64-microwell microfluidic chip system described above was utilized for organ specific metabolic enzymes studies of some common pollutants. The only difference from the B[a]P application was that metabolic enzyme sources included organ-specific microsomes (insoluble tissue fractions), cytosols (soluble tissue fractions) and supersomes from human liver, lung, intestine, and kidney. The PG microwell chips were outfitted with enzymes according to Scheme 8 with the aim of identifying relative contributions of metabolic enzymes that are responsible for conversion and DNA damage from a test compound in each human organ. Guided by this information, individual metabolite-nucleobase structures and formation rates were also measured by LC-MS/MS using a 96-well plate biocolloid reactor with magnetic beads to establish molecular pathways for DNA damage. This approach provided results to give a detailed, comprehensive picture of possible genotoxicity chemistry linked to individual human organs. With these methods, known genotoxic agents 4-(methylnitrosamino)-1-(3-pyr-idyl)-1-butanone (NNK), 2-acetylaminofluorene (2-AAF) and styrene revealed organ-based differences in rates of DNA damage from major metabolite-nucleobase adducts. Correlations between cell-free DNA damage in these studies with Comet assays that monitor DNA damage in living cells from the same organs were found [38].
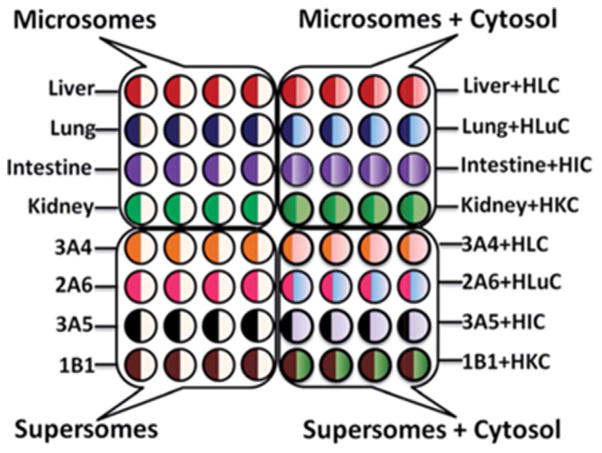
Scheme 8.
Experimental design for 64-nanowell ECL chip with films of organ-derived enzymes and RuIIPVP/DNA for metabolic toxicity assays of test compounds. Symbols: HLC, H=human, L=liver, C=cytosol, Lu=lung, I=intestine, K=kidney, number-letter such as 3A4 indicate supersomes containing the denoted single cyt P450 manifold. Reproduced from ref. [38] with permission of the Royal Society of Chemistry, copyright 2015.
4 ECL Arrays for DNA Oxidation
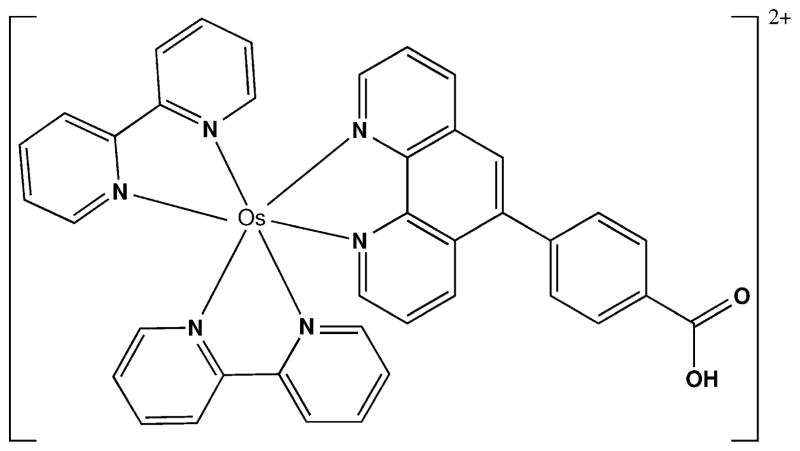
The microwell arrays described above were adapted to measure oxidative DNA damage caused by reactive oxygen species (ROS). Guanosines in DNA are oxidized by ROS to form 8-oxo-7,8-dihydro-2-deoxyguanosine (8-oxodG), a biomarker for oxidative damage, and we found that 8-oxodG serves as a coreactant for certain Os complexes to produce ECL [39]. Detection of 8-oxodG in intact ds-DNA without hydrolysis was done using the osmium complex [Os(bpy)2(phen-benz-COOH)]2+ {bpy= 2,2′-bipyridine; phen-benz-COOH=(4-(1,10-phenanthro-lin-6-yl)benzoic acid)} (Scheme 9) in stable composite films made with Nafion and reduced graphene oxide (OsNG). These OsNG films were assembled in the printed microwells on the PG chip, and used for ECL detection of 8-oxodG in oligonucleotides [40].
Scheme 9.
Structure of [Os(bpy)2(phen-benz-COOH)]2+.
Reduced graphene oxide in the OsNG films helped to enhance the oxidation current and thus the % ECL increase that results from the catalytic oxidation of 8-oxodG in DNA strands in the films. Nafion contributes mechanical and electrochemical stability to the film. Oxidized polyguanylic acid (PolyG) or DNA solutions were drop-cast into the OsNG-containing microwells in the arrays and dried. The array was fitted into the fluidic device (Scheme 6a), and buffer was delivered to the detection chamber to detect ECL at 0.9 V versus Ag/AgCl by using a CCD camera.
No external co-reactant is needed since the analyte 8-oxodG serves this purpose [40]. ECL generation involves selective catalytic oxidation of 8-oxodG by [Os(bpy)2-(phen-benz-COOH)]2+ in a complex redox pathway similar to Scheme 3 (except involving Os centers with 8-oxodG as co-reactant) to yield electronically excited [Os(bpy)2(phen-benz-COOH)]2+* in the films that decays by emitting ECL light at 746 nm. Figure 3a shows typical ECL data for oxidation of poly(deoxyguanosine) (polyG) and DNA oxidized by Fenton’s reagent.
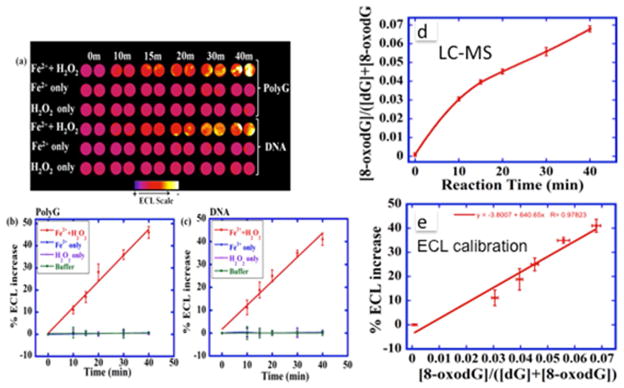
Fig. 3.
ECL-derived data related to nucleotide oxidation. (a) Reconstructed, recolorized ECL image showing ECL for polyG and DNA treated with Fenton′s reagent in pH 7.2 PBS for time in min. (top image). Controls are polynucleotides treated only with buffer, with only FeSO4 or with only H2O2. Graphs show relative %ECL increase for (b) polyG and (c) DNA. (d) UPLC-MS/MS for ECL array calibration showing measured ratio [8-oxodG]/([dG]+[8-oxodG]) vs. reaction time with Fenton′s reagent. (e) Array calibration as % ECL increase vs. ratio [8-oxo dG]/([dG]+[8-oxodG]). Reproduced from ref. [40] with permission American Chemical Society, Copyright 2016.
We constructed a calibration curve (Figure 3e) for the array in the form of % ECL increase vs. ratio of 8-oxodG to total dG, by measuring amounts of 8-oxodG directly by Ultrahigh Pressure Liquid Chromatography (UPLC)-MS/MS. Figure 3d depicts data for relative amounts of 8-oxodG in DNA oxidized by Fenton’s reagent vs. reaction time determined in this way. These data enable % ECL for the same samples analyzed on the array to be plotted vs. relative amount of 8-oxodG (Figure 3e), establishing calibration that is independent of the mode of DNA oxidation. From ECL arrays, 64 samples can be analyzed using only 1 μg DNA per microwell, with detection limit 1500 8-oxodG per 106 nucleobases, which was sufficient to examine DNA damage resulting from metabolite-mediated ROS production (data not shown) [40].
5 Summary
The discussion above illustrates development of micro-fluidic arrays capable of detecting multiple types of DNA damage by ECL. This approach is well suited to the task since ECL is sensitive, easily measured with a camera, and requires only a single patterned working electrode for the array. Films of DNA and metabolic enzymes aid in uncovering genotoxic chemistry that may result in genotoxic reactions of the test compounds and metabolites in vivo. ECL approaches involving films of DNA, enzymes and an ECL metallopolymer dye in microfluidic array reactors provide relative DNA damage rates that correlate with LC-MS measures of DNA adducts and other toxicity metrics and link results to genotoxic chemistry. The use of an Os-complex as ECL dye enables detection of oxidation in intact DNA without hydrolysis. Combination of array results with high throughput LC-MS/MS approaches using magnetic beads coated with similar films can be used to establish detailed toxicity-related chemical pathways. Our future work will be aimed towards combined detection of DNA oxidation and adduction and other types of damage on the same array. However, it is important to realize the complexity of human toxicity prediction, and the methods reviewed here must be combined with a complete strategy involving a reliable cohort of bioassays and computational algorithms for the most accurate predictions [7].
Acknowledgments
This work was supported financially by National Institute of Environmental Health Sciences (NIEHS), NIH, USA. Grant ES03154 (JFR). The authors thank coworkers and collaborators named in joint publications for their many excellent contributions to this research, without which progress would not have been possible.
Biographies

Kiran Bano received her PhD in electrochemistry from the Monash University, Australia under the supervision of Dr. Jie Zhang and Prof. Alan M. Bond in 2014. Her research was based on electrode kinetics studies using large amplitude Fourier transformed ac voltammetry of systems applicable in sensor application. Currently, she is a postdoctoral fellow at the University of Connecticut under the supervision of Prof. James F. Rusling. Her current work is focused on cyt P450-generated metabolic toxicity studies using ECL based biosensors.

James F. Rusling was awarded a Ph. D. from Clarkson University in 1979. He is Professor of Chemistry at University of Connecticut and Professor of Surgery and member of the Neag Cancer Center at University of Connecticut Health Center, as well as adjunct Professor of Chemistry at National Univ. of Ireland. Galway. Current research includes low cost, automated cancer diagnostic arrays measuring biomarker proteins, ECL and mass spectrometric arrays for toxicity screening and fundamental bioelectrochemistry.
References
- 1.Travis CC, Hester ST. Environ Sci Tech. 1991;25:814. [Google Scholar]
- 2.Guengerich FP. Chem Res Toxicology. 2001;14:611. doi: 10.1021/tx0002583. [DOI] [PubMed] [Google Scholar]
- 3.Rusling JF, Hvastkovs EG, Schenkman JB. Drug Metabolism Handbook. John Wiley & Sons, Inc; 2008. [Google Scholar]
- 4.Durackova Z. Physiol Res. 2010;59:459. doi: 10.33549/physiolres.931844. [DOI] [PubMed] [Google Scholar]
- 5.Reuter S, Gupta SC, Chaturvedi MM, Aggarwal BB. Free radical biology & medicine. 2010;49:1603. doi: 10.1016/j.freeradbiomed.2010.09.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.White B, Smyth MR, Stuart JD, Rusling JF. J Am Chem Soc. 2003;125:6604. doi: 10.1021/ja0343252. [DOI] [PubMed] [Google Scholar]
- 7.Hvastkovs EG, Schenkman JB, Rusling JF. Ann Rev Anal Chem. 2012;5:79. doi: 10.1146/annurev.anchem.111808.073659. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Rusling JF, Wasalathanthri DP, Schenkman JB. Soft Matter. 2014;10:8145. doi: 10.1039/c4sm01679c. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Wang B, Jansson I, Schenkman JB, Rusling JF. Anal Chem. 2005;77:1361. doi: 10.1021/ac0485536. [DOI] [PubMed] [Google Scholar]
- 10.Ma H, Hu N, Rusling JF. Langmuir. 2000;16:4969. [Google Scholar]
- 11.Rusling JF, Hvastkovs EG, Schenkman JB. Curr Opin Drug Disc & Devel. 2007;10:67. [PubMed] [Google Scholar]
- 12.Rusling JF. Biosens Bioelect. 2004;20:1022. doi: 10.1016/j.bios.2004.06.033. [DOI] [PubMed] [Google Scholar]
- 13.Hvastkovs EG, So M, Krishnan S, Bajrami B, Tarun M, Jansson I, Schenkman JB, Rusling JF. Anal Chem. 2007;79:1897. doi: 10.1021/ac061975q. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Dennany L, Forster RJ, Rusling JF. J Am Chem Soc. 2003;125:5213. doi: 10.1021/ja0296529. [DOI] [PubMed] [Google Scholar]
- 15.Krishnan S, Hvastkovs EG, Bajrami B, Choudhary D, Schenkman JB, Rusling JF. Anal Chem. 2008;80:5279. doi: 10.1021/ac800763r. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Pan S, Zhao L, Schenkman JB, Rusling JF. Anal Chem. 2011;83:2754. doi: 10.1021/ac200050n. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Krishnan S, Hvastkovs EG, Bajrami B, Schenkman JB, Rusling JF. Mol BioSyst. 2009;5:163. doi: 10.1039/b815910f. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Wasalathanthri DP, Malla S, Bist I, Tang CK, Faria RC, Rusling JF. Lab on chip. 2013;13:4554. doi: 10.1039/c3lc50698c. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Wasalathanthri DP, Faria RC, Malla S, Joshi AA, Schenkman JB, Rusling JF. Analyst. 2013;138:171. doi: 10.1039/c2an35993f. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Krishnan S, Hvastkovs EG, Bajrami B, Jansson I, Schenkman JB, Rusling JF. Chem Commun. 2007;17:1713. doi: 10.1039/b703012f. [DOI] [PubMed] [Google Scholar]
- 21.So M, Hvastkovs EG, Schenkman JB, Rusling JF. Biosens Bioelect. 2007;23:492. doi: 10.1016/j.bios.2007.06.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Wasalathanthri DP, Mani V, Tang CK, Rusling JF. Anal Chem. 2011;83:9499. doi: 10.1021/ac202269t. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Wang B, Rusling JF. Anal Chem. 2003;75:4229. doi: 10.1021/ac034097u. [DOI] [PubMed] [Google Scholar]
- 24.Zhao L, Schenkman JB, Rusling JF. Anal Chem. 2010;82:10172. doi: 10.1021/ac102317a. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Schenkman JB, Greim H, editors. Cytochrome P450. Springer-Verlag; Berlin: 1993. [Google Scholar]
- 26.Shimada T, Yamazaki H, Mimura M, Wakamiya N, Ueng YF, Guengerich FP, Inui Y. Drug metab Dispos. 1996;24:515. [PubMed] [Google Scholar]
- 27.Ortiz de Montellano PR, editor. Cytochrome P450. 3. Kluwer/Plenum; New York: 2005. [Google Scholar]
- 28.Krishnan S, Schenkman JB, Rusling JF. J Phys Chem B. 2011;115:8371. doi: 10.1021/jp201235m. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Lu Z, Lvov Y, Jansson II, Schenkman JB, Rusling JF. J Colloid Interface Sci. 2000;224:162. doi: 10.1006/jcis.1999.6685. [DOI] [PubMed] [Google Scholar]
- 30.Lvov YM, Lu Z, Schenkman JB, Zu X, Rusling JF. J Am Chem Soc. 1998;120:4073. [Google Scholar]
- 31.Yang J, Zhang Z, FRusling J. Electroanal. 2002:1494. [Google Scholar]
- 32.Gilardi G, Meharenna YT, Tsotsou GE, Sadeghi SJ, Fairhead M, Giannini S. Biosens Bioelect. 2002;17:133. doi: 10.1016/s0956-5663(01)00286-x. [DOI] [PubMed] [Google Scholar]
- 33.Fleming BD, Johnson DL, Bond AM, Martin LL. Expert Opin Drug Metab Toxicol. 2006;2:581. doi: 10.1517/17425255.2.4.581. [DOI] [PubMed] [Google Scholar]
- 34.Krishnan S, Wasalathanthri D, Zhao L, Schenkman JB, Rusling JF. J Am Chem Soc. 2011;133:1459. doi: 10.1021/ja108637s. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Wasalathanthri DP, Malla S, Faria RC, Rusling JF. Electroanalysis. 2012;24:2049. doi: 10.1002/elan.201200373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Bard AJ. Electrogenerated Chemiluminescence. Marcel Dekker; New York: 2004. [Google Scholar]
- 37.Forster RJ, Bertoncello P, Keyes TE. Annu Rev Anal Chem. 2009;2:359. doi: 10.1146/annurev-anchem-060908-155305. [DOI] [PubMed] [Google Scholar]
- 38.Wasalathanthri DP, Li D, Song D, Zheng Z, Choudhary D, Jansson I, Lu X, Schenkman JB, Rusling JF. Chem Sci. 2015;6:2457. doi: 10.1039/c4sc03401e. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Dennany L, Forster RJ, White B, Smyth M, Rusling JF. J Am Chem Soc. 2004;126:8835. doi: 10.1021/ja048615+. [DOI] [PubMed] [Google Scholar]
- 40.Bist I, Song B, Mosa IM, Keyes TE, Martin A, Forster RJ, Rusling JF. ACS Sens. 2016;1:272. doi: 10.1021/acssensors.5b00189. [DOI] [PMC free article] [PubMed] [Google Scholar]