Abstract
Aim:
The aim of this work was to detect antibiotic resistance in Gram-negative bacteria isolated from subclinical mastitis in cattle in West Bengal.
Materials and Methods:
The milk samples were collected from the cattle suffering with subclinical mastitis in West Bengal. The milk samples were inoculated into the nutrient broth and incubated at 37°C. On the next day, the growth was transferred into nutrient agar and MacConkey agar. All the pure cultures obtained from nutrient agar slant were subjected to Gram-staining and standard biochemical tests. All the bacterial isolates were tested in vitro for their sensitivity to different antibiotics commonly used in veterinary practices. All Gram-negative isolates including positive control were subjected to polymerase chain reaction (PCR) for detection of blaCTX-M, blaTEM, blaSHV, blaVIM, tetA, tetB, tetC, and tetM genes considered for extended-spectrum β-lactamase (ESBL), metallo-β-lactamase, and tetracycline resistance.
Results:
In total, 50 Gram-negative organisms (Escherichia coli, Proteus, Pseudomonas, Klebsiella, and Enterobacter) were isolated from milk samples of subclinical mastitis infected cattle. Among these Gram-negative isolates, 48% (24/50) were found either ESBL producing or tetracycline resistant. Out of total 50 Gram-negative isolates, blaCTX-M was detected in 18 (36%) isolates, and 6 (12%) harbored blaTEM genes in PCR. None of the isolates carried blaSHV genes. Further, in this study, 5 (10%) isolates harbored tet(A) gene, and 8 (16%) isolates carried tet(B) gene. No tet(C) gene was detected from the isolates.
Conclusion:
This study showed emerging trend of antibiotic-resistant Gram-negative bacteria associated with subclinical mastitis in cattle in West Bengal, India.
Keywords: antibiotic resistance, cattle, Escherichia coli, India, sub-clinical mastitis
Introduction
Subclinical mastitis is the most fatal infection confronted by the dairy industry today with significant economic losses worldwide including India [1]. It also poses a major public health risk due to transmission possibility of zoonotic bacteria or their toxin along with antibiotic resistance genes [2]. Most of the subclinical mastitic animals do not produce characteristic symptoms of mastitis and are persistent shedder of zoonotic bacteria without adequate awareness of farmers regarding the transmission possibility [3].
Subclinical mastitis is considered to have a multifactorial etiology including several groups of microorganisms such as bacteria, virus, fungi, yeast, and algae [4]. Among the Gram-negative bacterial etiological agents, the major group includes coliform bacteria (Escherichia coli, Enterobacter, Klebsiella), Pseudomonas and Serratia [5].
Early detection of subclinical mastitis and dry cow therapy with proper antibiotics, use of post-milking teat disinfectants and effective pre-milking hygiene becomes relevant to minimize economic losses in dairy farms and to prevent zoonotic transmission [6]. However, the etiological and commensal bacteria present in animals and exposed to the antimicrobial pressure, develop survival strategies through evolutionary adaptations [7]. Gram-negative bacteria specially Enterobacteriaceae organisms mostly produce β-lactamase enzymes to prevent the action of β-lactam antibiotics. There are more than 1000 β-lactamase enzymes that can be classified under four main classes, i.e., A-D [8]. The most clinically important Class A enzymes, found in Enterobacteriaceae, are known as extended-spectrum β-lactamases (ESBLs). It can confer resistance to a variety of β-lactam antibiotics, including penicillins, 2nd, 3rd and 4th-generation cephalosporins and monobactams (e.g., aztreonam), but usually not the carbapenems or the cephamycins (e.g., cefoxitin). There are three classical ESBLs, i.e., TEM (except TEM-1), SHV (except SHV-1 and 2), and CTX-M [9]. Among them, CTX-M is observed as the most prevalent type worldwide [10]. Other than ESBL genes, possession of tetracycline resistance gene is also common in Gram-negative bacteria [11].
In India, subclinical mastitis in cattle and buffaloes caused by Gram-positive bacteria such as Staphylococcus spp. are reported from different states such as Karnataka, Punjab, and others [3,12]. However, etiological correlation of Gram-negative bacteria with subclinical mastitis in Indian cattle is less explored. This study was conducted to know the occurrence of Gram-negative bacteria in subclinical mastitis in cattle associated with antibiotic resistance potential.
Materials and Methods
Ethical approval
The study was approved by Instituional Animal Ethics Committee, WBUAFS.
Sampling
The udders of the suspected animals were examined for fibrosis, inflammatory swellings, visible injury, tick infestation, atrophy of the tissue, and swelling of supramammary lymph nodes. The size and consistency of mammary quarters were inspected for the presence of any abnormalities, such as disproportional symmetry, swelling, firmness, and blindness. Information relating to the previous health history of the mammary quarters and causes of blindness was obtained from the owners of the farm. The mastitic milk was collected aseptically as described earlier [13]. The collected milk samples were transported to the laboratory maintaining the cold chain.
Isolation of Gram-negative bacteria
Initially, the milk samples were inoculated into the nutrient broth (HiMedia, India) and incubated at 37°C. On the next day, the growth was transferred into nutrient agar and MacConkey agar (HiMedia, India). The convex glistening single colonies with greenish discolouration in nutrient agar and the pink and pale colored colonies in MacConkey agar were isolated into nutrient agar slant as a pure culture. All the pure cultures obtained from nutrient agar slant were subjected to Gram-staining and standard biochemical tests as described earlier [14].
Antibiotic sensitivity test
All the bacterial isolates were tested in vitro for their sensitivity to different antibiotics commonly used in veterinary practices. The antibiotic disks oxytetracycline, amikacin, gentamicin, amoxicillin and clavulanic acid, amoxicillin sulbactam, ceftriaxone and sulbactam, ceftriaxone and tazobactam, enrofloxacin, ceftriaxone and cefotaxime (HiMedia) were selected for the study. The interpretation was done in accordance to performance standards for antimicrobial disks susceptibility tests, Clinical Laboratory Standard Institute [15].
Polymerase chain reaction (PCR)-based detection of ESBL and tetracycline resistance genes
For PCR-based detection of major ESBL genes (blaTEM, blaSHV, blaCTX-M) and tetracycline resistance (tet(A), tet(B) and tet(C)) genes from all the bacterial isolates, DNA was extracted as per the method described by Bonnet et al. [16]. All Gram-negative isolates including positive control were subjected to PCR for detection of blaCTX-M, blaTEM, blaSHV, blaVIM, tetA, tetB, tetC, and tetM genes considered for ESBL, metallo-β-lactamase, and tetracycline resistance. The PCR was performed in a thermocycler (BioRad, USA) with the primers (Imperial life sciences, India) and the cycle conditions as described earlier [16].
Results
In this study, 50 Gram-negative organisms (E. coli, Proteus, Pseudomonas, Klebsiella, and Enterobacter) were isolated from milk samples of subclinical mastitic cattle. Among these Gram-negative isolates, 48% (24/50) isolates were detected phenotypically as either ESBL producing or tetracycline resistant in antibiotic sensitivity test.
Out of total 50 Gram-negative isolates, blaCTX-M was detected in 18 (36%) isolates, and 6 (12%) harbored blaTEM genes in PCR (Figures-1 and 2). None of the isolates carried blaSHV genes. Further, in this study, 5 (10%) isolates harbored tet(A) gene and 8 (16%) isolates carried tet(B) gene (Figures-3 and 4). No tet(C) gene was detected in the isolates.
Figure-1.
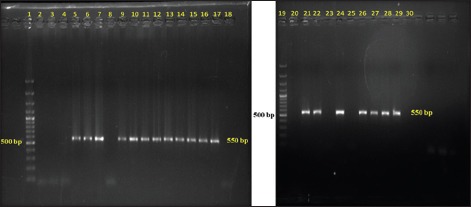
Gel electrophoresis image of polymerase chain reaction amplified products of blaCTX-M gene of Gram-negative isolates; Lane 1 - 1500 bp DNA ladder; Lane 2-14, 16-19 - Isolated Escherichia coli samples; Lane 15 - Positive control for blaCTX-M gene; Lane 20 - Negative control.
Figure-2.
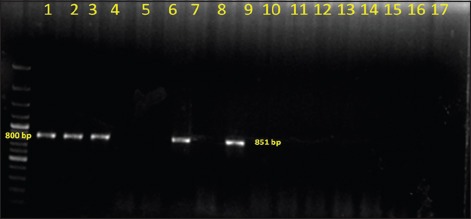
Gel electrophoresis image of polymerase chain reaction amplified products of blaTEM gene of Gram-negative isolates; Lane 1 - 1500 bp DNA ladder; Lane 2-8, 10-16 - Isolated Escherichia coli samples; Lane 15 - Positive control for blaTEM gene; Lane 20 - Negative control.
Figure-3.
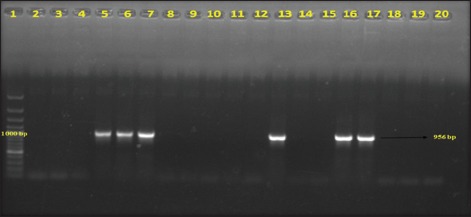
Gel electrophoresis image of polymerase chain reaction amplified products of tetA gene of Gram-negative isolates; Lane 1 - 1500 bp DNA ladder; Lane 2-16, 18-19 - Isolated Escherichia coli samples; Lane 17 - Positive control for tetA gene; Lane 20 - Negative control.
Figure-4.
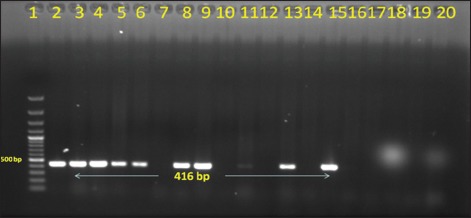
Gel electrophoresis image of polymerase chain reaction amplified products of tetB gene of Gram-negative isolates. Lane 1 - 1500 bp DNA ladder; Lane 2-14, 16-19 - Isolated Escherichia coli samples; Lane 15 - Positive control for tetB gene; Lane 20 - Negative control.
Discussion
Subclinical mastitis is the most fatal infection confronted by dairy industry today with significant economic losses worldwide including India [1]. Moreover, with the extensive use of β-lactam antibiotics, cattle and other ruminants represent a considerable source for the transmission of antibiotic resistance genes (e.g., ESBL) or antibiotic resistant strains to the human intestinal bacterial flora [17]. This study was conducted to know the occurrence of Gram-negative bacteria in subclinical mastitis in cattle associated with antibiotic resistance potential.
Among these Gram-negative isolates, 48% (24/50) were either ESBL producing or tetracycline resistant in this study. Earlier reports from different countries revealed the lower prevalence of ESBL-producing Enterobacteriaceae isolates (0.4-13%) associated with bovine mastitis [18-20]. In India, few reports are available regarding detection of ESBL/New Delhi metallo-β-lactamase-producing E. coli in milk samples collected from clinical or subclinical mastitic cattle [17,21]. The prevalence study of ESBL-producing Gram-negative bacteria with substantial numbers of mastitic milk samples is not apparently available in India to compare the present finding. However, the study indicates about alarming rise in the occurrence of ESBL-producing Gram-negative isolates in subclinical mastitic cattle.
Out of total 50 Gram-negative isolates, blaCTX-M was detected in 18 (36%) isolates, and 6 (12%) harbored blaTEM genes in PCR. None of the isolates carried blaSHV genes. At present, CTX-M is the major ESBL enzyme produced by different clonal complexes of Enterobacteriaceae which mostly replaced the SHV and TEM-type ESBLs during the last decade [22]. It was also observed that CTX-M and TEM were the most prevalent bla-encoded enzyme in human clinical isolates worldwide [23-25]. As well as, CTX-M ESBL producing Klebsiella pneumonae was also isolated from the cases of bovine mastitis [26].
Further, in this study, 5 (10%) isolates harbored tet(A) gene and 8 (16%) isolates carried tet(B) gene. No tet(C) gene was detected from the isolates. Earlier studies indicated the presence of tet genes in Gram-negative bacteria isolated from bovine mastitis and other infections [27,28].
Conclusion
The present study showed emerging trend of antibiotic resistant gram negative bacteria associated with sub-clinical mastitis in cattle in West Bengal, India.
Authors’ Contributions
AD: Conducted study. CG, UB, PSJ, AC and IS: Planned the study. AD and IS: Drafted and revised the manuscript. All authors read and approved the final manuscript.
Acknowlegments
The authors provide sincere gratitude to honorable Vice Chancellor, West Bengal University of Animal and Fishery Sciences, Kolkata for necessary infrastructures.
Competing Interests
The authors declare that they have no competing interests.
References
- 1.Joshi G. Status of mastitis as an emerging disease in improved and periurban dairy farms in India. Ann. N. Y. Acad. Sci. 2006;10:1373–3177. doi: 10.1196/annals.1373.007. [DOI] [PubMed] [Google Scholar]
- 2.Ibrahim H.M.M, Ahmed A.M, El-seedy Y.Y, El-Khodery S.A. Distribution of multidrug-resistant gram negative bacteria causing clinical mastitis in dairy cows. Glob Vet. 2015;15:268–277. [Google Scholar]
- 3.Prabhu K.N, Isloor S, Hegde R, Rathnamma D, Veeregowda B.M, Narasimha Murthy H.N, Shome R, Suryanarayana V.V.S. Development of polymerase chain reaction for detection of predominant streptococcal isolates causing sub clinical bovine mastitis. Indian J. Biotechnol. 2013;12:208–212. [Google Scholar]
- 4.Mallikarjunaswamy M.C, Krishnamurthy G.V. Antibiogram of bacterial pathogens isolated from bovine sub-clinical mastitis cases. Indian Vet J. 1997;74:885–886. [Google Scholar]
- 5.Samanta I. Veterinary Bacteriology. 1st ed. New Delhi, India: New India Publishing Agency; 2013. [Google Scholar]
- 6.Idriss S.E, Foltys V, Tančin V, Kirchnerová K, Tančinová1 D, Zaujec K. Mastitis pathogens and their resistance against antimicrobial agents in dairy cows in Nitra, Slovakia. Slovak J. Anim. Sci. 2014;47:33–38. [Google Scholar]
- 7.Allen H.K, Stanton T.B. Altered egos: Antibiotic effects on food animal micro biomes. Ann Rev. Microbiol. 2014;68:297–315. doi: 10.1146/annurev-micro-091213-113052. [DOI] [PubMed] [Google Scholar]
- 8.Bush K, Jacoby G.A. Updated functional classification of beta-lactamases. Antimicrob. Agents Chemother. 2010;54:969–976. doi: 10.1128/AAC.01009-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.EFSA Panel on Biological Hazards. Scientific opinion on the public health risks of bacterial strains producing extended-spectrum β-lactamases and/or AmpC β-lactamases in food and food-producing animals. EFSA J. 2011;9:2322. [Google Scholar]
- 10.Carattoli A. Plasmids and the spread of resistance. Int. J. Med. Microbiol. 2013;303:298–304. doi: 10.1016/j.ijmm.2013.02.001. [DOI] [PubMed] [Google Scholar]
- 11.Wellington E.M.H, Boxall A.B.A, Cross P, Feil E.J, Gaze W.H, Hawkey P.M, Johnson-Rollings A.S, Jones D.L, Lee N.M, Otten W, Thomas C.M, Williams A.P. The role of the natural environment in the emergence of antibiotic resistance in Gram-negative bacteria. Lancet Infect. Dis. 2013;13:155–165. doi: 10.1016/S1473-3099(12)70317-1. [DOI] [PubMed] [Google Scholar]
- 12.Kaur M, Ramneek Bansal B.K, Mukhopadhyay C.S, Arora J.S. Status of sub-clinical mastitis and associated risk factors in Indian water buffalo in Doaba region of Punjab, India. Indian J. Dairy Sci. 2015;68:483–487. [Google Scholar]
- 13.Schalm O.W, Carroll E.J, Jain N.C. Bovine Mastitis. Philadelphia: Lea and Febiger; 1971. [Google Scholar]
- 14.Quinn P.J, Carter M.E, Markey B.K, Carter G.R. Clinical Veterinary Microbiology. UK: Mosby; 1994. [Google Scholar]
- 15.CLSI. Performance standards for antimicrobial disk and dilution susceptibility tests for bacteria isolated from animals: Approved standard. 3rd ed. Wayne, PA: M31-A3. CLSI; 2010. pp. 13–23. [Google Scholar]
- 16.Bonnet R, Recule C, Baraduc R, Chanal C, Sirot I, De Champs C, Sirot J. Effect of D240G substitution in a novel ESBL CTX-M-27. J. Antimicrob. Chemother. 2003;52:29–35. doi: 10.1093/jac/dkg256. [DOI] [PubMed] [Google Scholar]
- 17.Bandyopadhyay S, Samanta I, Bhattacharyya D, Nanda P.K, Kar D, Chowdhury J, Dandapat P, Das A.K, Batul N, Mondal B, Dutta T.K, Das G, Das B.C, Naskar S, Bandyopadhyay U.K, Das S.C, Bandyopadhyay S. Co-infection of methicillin-resistant Staphylococcus epidermidis methicillin-resistant Staphylococcus aureus and extended spectrum β-lactamase producing Escherichia coli in bovine mastitis - Three cases reported from India. Vet. Q. 2015;35:56–61. doi: 10.1080/01652176.2014.984365. [DOI] [PubMed] [Google Scholar]
- 18.Hu G.Z, Kuang X.H, Yuan L, Mo J, Pan Y.S, Fu X.L, Xu K.L, Li L.F. Detection of antibiotic susceptibility of ESBL producer Enterobacteriaceae against 21 agents. Zhongguo Ren Shou Gong Huan Bing Za Zhi. 2006;22:884–887. [Google Scholar]
- 19.Dahmen S, Métayer V, Gay E, Madec J.Y, Haenni M. Characterization of extended-spectrum beta-lactamase (ESBL)-carrying plasmids and clones of Enterobacteriaceae causing cattle mastitis in France. Vet. Microbiol. 2013;162:793–799. doi: 10.1016/j.vetmic.2012.10.015. [DOI] [PubMed] [Google Scholar]
- 20.Freitag C, Michael G.B, Kadlec K, Hassel M, Schwarz S. Detection of plasmid-borne extended-spectrum β-lactamase (ESBL) genes in Escherichia coli isolates from bovine mastitis. Vet. Microbiol. 2016 doi: 10.1016/j.vetmic.2016.08.010. (In Press) [DOI] [PubMed] [Google Scholar]
- 21.Ghatak S, Singha A, Sen A, Guha C, Ahuja A, Bhattacharjee U, Das S, Pradhan N.R, Puro K, Jana C, Dey T.K, Prashantkumar K.L, Das A, Shakuntala I, Biswas U, Jana P.S. Detection of New Delhi metallo-beta-lactamase and extended-spectrum beta-lactamase genes in Escherichia coli isolated from mastitic milk samples. Transbound. Emer. Dis. 2013;60:385–389. doi: 10.1111/tbed.12119. [DOI] [PubMed] [Google Scholar]
- 22.Cantón R, Coque T.M. The CTX-M β-lactamase pandemic. Curr. Opin. Microbiol. 2006;9:466–475. doi: 10.1016/j.mib.2006.08.011. [DOI] [PubMed] [Google Scholar]
- 23.Carattoli A, Garcia-Fernandez A, Varesi P, Fortini D, Gerardi S, Penni A, Mancini C, Giordano A. Molecular epidemiology of Escherichia coli producing extended-spectrum β-lactamases isolated in Rome, Italy. J. Clin. Microbiol. 2008;46:103–108. doi: 10.1128/JCM.01542-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Kiratisin P, Apisarnthanarak A, Laesripa C, Saifon P. Molecular characterization and epidemiology of extended-spectrum β- lactamase-producing Escherichia coli and Klebsiella pneumonia isolates causing health care-associated infection in Thailand, where the CTX-M family is endemic. Antimicrob. Agents Chemother. 2008;52:2818–2824. doi: 10.1128/AAC.00171-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Smet A, Martel A, Persoons D, Dewulf J, Heyndrickx M, Catry B, Herman L, Haesebrouck F, Butaye P. Diversity of extended-spectrum β-lactamases and Class C β-lactamases among cloacal Escherichia coli isolates in Belgian broiler farms. Antimicrob. Agents Chemother. 2008;52:1238–1243. doi: 10.1128/AAC.01285-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Locatelli C, Licia S, Giuliano P, Paolo M. CTX-M1 ESBL-producing Klebsiella pneumoniae sub sp pneumoniae isolated from cases of bovine mastitis. J. Clin. Microbiol. 2010;48:3822. doi: 10.1128/JCM.00941-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Ahmed A.M, Shimamoto T. Molecular characterization of antimicrobial resistance in Gram negative bacteria isolated from bovine mastitis in Egypt. Microbiol. Immunol. 2011;55:318–327. doi: 10.1111/j.1348-0421.2011.00323.x. [DOI] [PubMed] [Google Scholar]
- 28.Card R, Zhang J, Das P, Cook C, Woodford N, Anjum M.F. Evaluation of an expanded microarray for detecting antibiotic resistance genes in a broad range of gram-negative bacterial pathogens. Antimicrob. Agents Chemother. 2013;57:458–465. doi: 10.1128/AAC.01223-12. [DOI] [PMC free article] [PubMed] [Google Scholar]


