Abstract
Background
Oct4, a key stemness transcription factor, is overexpressed in lung cancer. Here, we reveal a novel transcription regulation of long non-coding RNAs (lncRNAs) by Oct4. LncRNAs have emerged as important players in cancer progression.
Methods
Oct4 chromatin-immunoprecipitation (ChIP)-sequencing and several lncRNA databases with literature annotation were integrated to identify Oct4-regulated lncRNAs. Luciferase activity, qRT-PCR and ChIP-PCR assays were conducted to examine transcription regulation of lncRNAs by Oct4. Reconstitution experiments of Oct4 and downstream lncRNAs in cell proliferation, migration and invasion assays were performed to confirm the Oct4-lncRNAs signaling axes in promoting lung cancer cell growth and motility. The expression correlations between Oct4 and lncRNAs were investigated in 124 lung cancer patients using qRT-PCR analysis. The clinical significance of Oct4/lncRNAs signaling axes were further evaluated using multivariate Cox regression and Kaplan-Meier analyses.
Results
We confirmed that seven lncRNAs were upregulated by direct binding of Oct4. Among them, nuclear paraspeckle assembly transcript 1 (NEAT1), metastasis-associated lung adenocarcinoma transcript 1 (MALAT1) and urothelial carcinoma-associated 1 (UCA1) were validated as Oct4 transcriptional targets through promoter or enhancer activation. We showed that lung cancer cells overexpressing NEAT1 or MALAT1 and the Oct4-silenced cells reconstituted with NEAT1 or MALAT1 promoted cell proliferation, migration and invasion. In addition, knockdown of NEAT1 or MALAT1 abolished Oct4-mediated lung cancer cell growth and motility. These cell-based results suggested that Oct4/NEAT1 or Oct4/MALAT1 axis promoted oncogenesis. Clinically, Oct4/NEAT1/MALAT1 co-overexpression was an independent factor for prediction of poor outcome in 124 lung cancer patients.
Conclusions
Our study reveals a novel mechanism by which Oct4 transcriptionally activates NEAT1 via promoter and MALAT1 via enhancer binding to promote cell proliferation and motility, and led to lung tumorigenesis and poor prognosis.
Electronic supplementary material
The online version of this article (doi:10.1186/s12943-017-0674-z) contains supplementary material, which is available to authorized users.
Keywords: Oct4, lncRNA, MALAT1, NEAT1, Transcription regulation, Lung cancer
Background
Oct4, encoded by POU5F1 (POU domain, class 5, transcription factor 1), is a homeodomain transcription factor of the POU family. Oct4, Sox2 and Nanog are well-known pluripotency-associated transcription factors which maintain embryonic stem cells state [1]. Metaplastic transformation, a precancerous condition, has been reported to recapitulate embryonic development. Therefore, the key factors involved in embryonic development may play critical roles in carcinogenesis. Studies have shown that Oct4 is overexpressed in human cancers such as bladder [2], breast [3], cervical cancer [4], oral squamous cell carcinoma [5], hepatocellular carcinoma [6] and lung cancer [7, 8]. In embryonic stem cells, Oct4 has been identified to regulate transcriptions of other transcription factors, chromatin modifiers, long non-coding RNAs (lncRNAs) and microRNAs [9, 10]. For instance, Oct4 regulates lncRNAs expression, such as linc-RoR, which is a key reprogramming factor associated with pluripotency [11]. Oct4 can also interact with Pontin, a chromatin remodeling factor, to regulate the transcription of lncRNAs, including linc1253, a lineage programme repressing lincRNA [12]. However, transcription regulation of lncRNAs by Oct4 in tumorigenesis remains elusive.
LncRNAs is a subset of non-coding RNAs with length ranging from 200 nucleotides to 100,000 nucleotides. According to data obtained using next generation RNA-sequencing, the number of total human lncRNAs is approximately 20,000 transcripts and over 200 lncRNAs are confirmed to be functional [13, 14]. Some lncRNAs are dysregulated in cancers and may serve as potential prognostic markers for specific cancer types [15, 16]. Some lncRNAs have been characterized to possess oncogene-like or tumor suppressor-like function. For instance, Hox Antisense Intergenic RNA (HOTAIR), acts as a bridge between PRC2 chromatin repressive and LSD1/CoREST/REST corepressor complexes to further modulate the metastasis-related gene expressions through changing chromatin states in breast cancer [15, 17]. Another lncRNA, HOXA transcript at the distal tip (HOTTIP), not only promotes pancreatic cancer progression but also confers chemoresistance to gemcitabine, which may be mediated by HOXA13 [18, 19]. Accumulating evidence indicates that lncRNAs play critical roles in cancer biology.
Up to date, most of the studies on lncRNAs focus on the outcome and underlying mechanisms of dysregulated lncRNAs and their potential as prognosis markers. However, little is known about the upstream regulations responsible for aberrant expression of lncRNAs in cancers, especially at the transcriptional level. Our previous study using chromatin-immunoprecipitation sequencing (ChIP-seq) and functional analyses revealed a critical Oct4-driven transcriptional program [8]. Genome-wide analysis of Oct4 targeting of this program suggests a novel role of Oct4-mediated transcriptional regulation of lncRNAs. In the current study, we have shown that Oct4 transcriptionally activated oncogenic lncRNAs expression through promoter- or enhancer-binding regulation. Moreover, Oct4-mediated high expression of lncRNAs such as nuclear paraspeckle assembly transcript 1 (NEAT1) and metastasis-associated lung adenocarcinoma transcript 1 (MALAT1) promoted lung cancer cell proliferation, migration and invasion abilities. Clinical studies further validated the importance of Oct4/NEAT1/MALAT1 signaling axis in lung cancer progression.
Methods
Cell lines and culture conditions
Human lung adenocarcinoma cell line A549 and normal bronchial epithelial cell line BEAS-2B was purchased from American Tissue Culture Company (ATCC). Human lung adenocarcinoma cell line CL1–0 was obtained from Dr. Pan-Chyr Yang (Department of Internal Medicine Medical College, National Taiwan University, Taiwan). All media were supplemented with 10% Fetal Bovine Serum (Gibco, Carlsbad, CA, USA) and 1% penicillin/streptomycin (Gibco). Stable cell line expressing Oct4 or empty vector was established by ectopic transfection of Flag-Oct4 or empty vector plasmid into A549 and CL1–0 cells, and selected with puromycin. Transient transfections of Oct4 in BEAS-2B were carried out with lipofetamine 2000 (Invitrogen, Carslbad, CA, USA).
Transfection of plasmids and RNAi
The plasmids used in the study are listed in Additional file 1: Table S1. The interference RNA (RNAi) for Oct4 was obtained from Invitrogen (# Oct4-HSS143403, Invitrogen). Depletion of NEAT1 or MALAT1 was performed by transfection of smart-pool siRNAs (Dharmacon, Lafayette, CO, USA) at final concentration of 10 nM. Transfections of expression plasmids and RNAi were performed using lipofectamine 2000 (Invitrogen, Carslbad, CA, USA) according to the manufacturer’s protocol.
Chromatin-immunoprecipitation-polymerase chain reaction (ChIP-PCR) assay
Empty vector control and Oct4 stably-overexpressed A549 cells (1 × 107 cells) were cross-linked with 1% formaldehyde for 10 min at 37 °C, followed by preparation of nuclear lysates using Magna ChIP™ protein G Kit (Millipore Co., Billerica, MA, USA). Nuclear lysates were sonicated to shear crosslinked DNA to around 300 ~ 500 bps using Covaris-S2 machine. Chromatin was immunoprecipitated with Oct4 antibody (1:100, # ab-19857, Abcam, Cambridge, UK). Purified chromatin-immunoprecipitated DNA was subjected to PCR analysis using primers for the lncRNA promoter and enhancer regions listed in Additional file 1: Table S2.
RNA extraction and quantitative reverse transcriptase-polymerase chain reaction (qRT-PCR) assays
Four μg of total RNA was reverse transcribed to cDNA using MultiScribe™ reverse transcriptase (Applied Biosystems, Foster City, CA, USA). cDNA was amplified using the Fast SYBR® Green Master Mix (Applied Biosystems). qRT-PCR was used to measure Oct4 mRNA and lncRNA expression using the StepOnePlus™ Real-Time PCR System (Applied Biosystems). The primer sequences and annealing temperature are listed in Additional file 1: Table S3.
Site-directed mutagenesis and luciferase promoter/enhancer activity assays
Mutations (Mut) of Oct4 binding elements within the NEAT1 promoter, MALAT1 enhancer or urothelial carcinoma-associated 1 (UCA1) enhancer were generated by site-directed mutagenesis using wild-type (WT) NEAT1 promoter, MALAT1 enhancer or UCA1 enhancer vectors as templates. The primers used are described in Additional file 1: Table S4.
For luciferase activity assays, cells were seeded the day before transfection. The pGL4-Renilla construct was included as an internal control. After 16 h of co-transfection with empty vector or gene promoter/enhancer vector, and pGL3-Basic or pGL4-Renilla, the dual luciferase reporter assay kit (Promega, Madison, WI, USA) was used to determine gene promoter or enhancer activity. The luminescence was measured with a Turner BioSystems luminometer (Promega). The data are represented as the means of ratio of firefly luciferase to Renilla luciferase activity by triplicate experiments.
RealTime-Glo viability assay
Cell viability was assayed using RealTime-Glo assay (Promega). Briefly, cells were transfected for 24 h and then reseeded at 2 × 103 cells/well in 96-well plates. MT Cell Viability Substrate and NanoLuc Enzyme were diluted and added to each well. The luminescence was measured with a Turner BioSystems luminometer (Promega) at 24, 48 and 72 h.
Transwell migration and invasion assay
The transwell insert with millipore membrane (pore size of 8 μm, Falcon, BD Franklin Lakes, NJ, USA) was used. For transwell migration assay, 2 × 105 A549 cells and 5 × 105 CL1–0 cells were seeded onto the upper chamber with 1 ml serum-free medium. For transwell invasion assay, the transwell inserted membranes were pre-coated with Matri-gel (2.5 mg/ml, Sigma-Aldrich, St. Louis, MO, USA) 1 day before seeding cells. Complete medium with 20% FBS was supplemented into the lower chamber as chemoattractants. The cells were incubated for 16 ~ 24 h and then the cells attached on the reverse side of the membrane were then fixed and stained. Six random views were photographed and quantified under an upright microscope (Nikon E400, Yurakucho, Tokyo, Japan).
Study population
We recruited 124 lung cancer patients from National Cheng Kung University Hospital after obtaining appropriate institutional review board permission and informed consent from the patients. Surgically resected tumor tissue and corresponding normal tissue samples were collected. Total RNA of patient samples were prepared using Trizol reagent (Invitrogen) and reverse transcribed into cDNA as described above. qRT-PCR was conducted to measure the expressions of Oct4, NEAT1 and MALAT1 using the StepOnePlus™ Real-Time PCR System (Applied Biosystems). The expression of the target genes was normalized based on the levels of internal control gene, GAPDH. The primers used for qRT-PCR analyses are described in Additional file 1: Table S3.
Statistical analysis
Pearson χ2 test was used to compare the correlation of Oct4 and lncRNAs expression and clinicopathological parameters in lung cancer patients. Overall survival curves were calculated according to the Kaplan-Meier method, and comparison was performed using the log-rank test. Two-way ANOVA and two-tailed Student’s t-test was used in cell and animal studies. Data represent mean ± SEM. P < 0.05 was considered to be statistically significant.
Results
Unbiased ChIP-seq and ChIP-PCR/qRT-PCR analyses reveal novel lncRNAs controlled by Oct4 transcriptional regulation in lung cancer
We previously performed ChIP-seq in A549 lung cancer cell line stably-overexpressing Oct4 to identify the Oct4 genome-wide DNA binding regions [8]. Notably, genomic loci of some lncRNAs were bound by Oct4. We further identified the potential Oct4-regulated lncRNAs through the following bioinformatic and functional analyses. Firstly, we integrated our in-house ChIP-seq dataset with functional lncRNA database [20] and lncRNA db [21] with literature annotation to search for lncRNAs with oncogenic potential. Secondly, candidate lncRNAs were further selected following the criteria that the Oct4 binding sites located within 100 kb from lncRNAs genomic loci. We then performed ChIP-PCR and qRT-PCR in A549 and CL1–0 cells overexpressing Oct4 to validate the selected lncRNAs for Oct4 protein binding and transcriptional regulation, seven oncogenic lncRNAs were validated (Fig. 1a). A549 and CL1–0 lung cancer cells were confirmed for the Oct4-induced oncogenic effects in vitro and in vivo (Additional file 1: Figure S1). Visualization of Oct4 ChIP-seq targeting revealed significant enrichments of Oct4 binding on three lncRNAs, including the promoter region of NEAT1 and enhancer regions of MALAT1 and UCA1 (Fig. 1b).
Fig. 1.
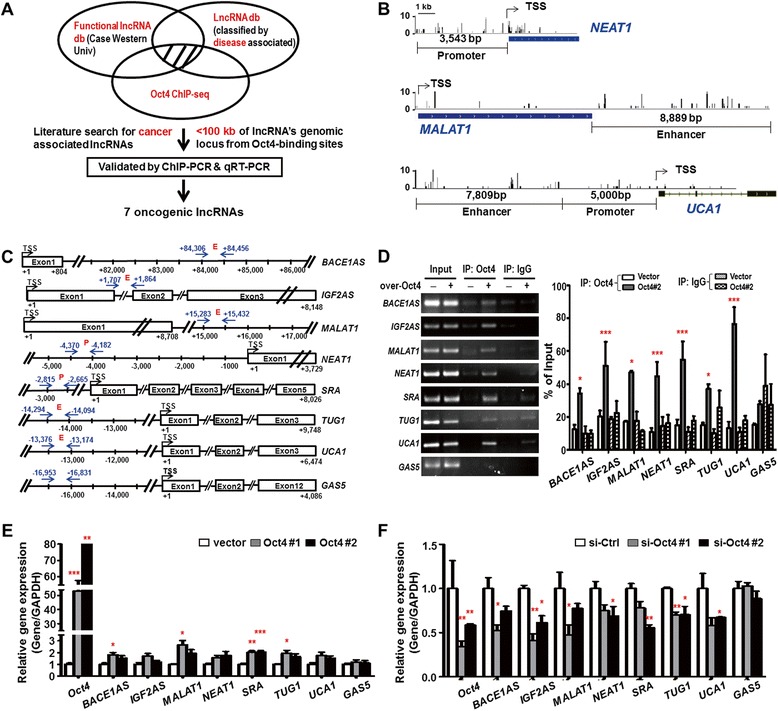
Oncogenic lncRNAs were revealed as Oct4 transcriptional downstream targets in lung cancer using ChIP-seq, ChIP-PCR and qRT-PCR assays. a Flowchart of strategies used to identify Oct4 targeted downstream lncRNAs. b The Oct4 binding peaks at promoter and enhancer regions of three representative lncRNAs NEAT1, MALAT1 and UCA1. c Schematic diagram depicting the ChIP-PCR primers for amplification of the regions including Oct4 binding sites around the eight lncRNAs genomic locus (indicated as blue arrows). TSS: transcription start site as indicated by (+1). Oct4 binding regions were classified as enhancer (E) or promoter (P). d ChIP-PCR analysis of Oct4 occupancy at the binding sites of the eight lncRNAs genomic loci. GAS5 serves as a negative control lncRNA. Results are normalized to input by semi-quantitative analysis. IgG serves as an experimental negative control. Data represent mean ± SEM. P-values were determined by two-way ANOVA. e, f qRT-PCR analysis of eight lncRNAs expressions in A549 cells stably overexpressing Oct4 (Oct4#1, Oct4#2) (e) or Oct4-silenced A549 cells (si-Oct4#1, si-Oct4#2) (f). GAS5 serves as a negative control lncRNA. Target lncRNA expression levels were normalized to GAPDH expression levels. Data represent mean ± SEM. P-values were determined by two-tailed Student t-test. *P < 0.05; **P < 0.01; ***P < 0.001
To validate ChIP-seq data, we performed ChIP-PCR and qRT-PCR to elucidate whether Oct4 transcriptionally regulates the eight selected lncRNAs, including seven oncogenic lncRNAs BACE1AS, IGF2AS, MALAT1, NEAT1, SRA, TUG1 and UCA1, and tumor suppressor lncRNA GAS5. The GAS5 was selected as a negative control lncRNA because it contained no ChIP-seq binding signal of Oct4. We used ALGGEN PROMO and TFSEARCH softwares to identify the putative Oct4 consensus binding elements (5’ATGCAAAT3’) of ChIP-seq regions. Oct4 binding sites within 5 kb upstream of transcription start site (TSS) were defined as promoter region, whereas Oct4 binding regions located at more than 5 kb of TSS or within the gene body were defined as enhancer region according to previous studies [22, 23]. Therefore, we classified the Oct4 binding sites to promoter (P) and enhancer (E) in relation to TSS of the seven lncRNAs (Fig. 1c). The ChIP-PCR results showed that Oct4 indeed targeted the predicted promoter or enhancer regions of the oncogenic lncRNAs but not the negative control lncRNA GAS5 (Fig. 1d).
To further confirm the transcription activation of Oct4 on downstream lncRNAs, qRT-PCR analysis was conducted in both A549 and CL1–0 cells stably expressing Oct4. Results showed that overexpression of Oct4 in A549 increased the expression of the seven oncogenic lncRNAs but not that of the lncRNA GAS5 (Fig. 1e). In contrast, knockdown of Oct4 in A549 cells decreased expression of the oncogenic lncRNAs but not that of GAS5 (Fig. 1f). The results in CL1–0 cells showed a similar trend but to a lesser extent compared to that in A549 cells (Additional file 1: Figure S2). An increased expression of NEAT1 and UCA1 lncRNAs upon Oct4 overexpression was observed in the normal bronchial epithelial cell line BEAS-2B (Additional file 1: Figure S3). These results suggested that Oct4 may transcriptionally upregulate the expression of these oncogenic lncRNAs identified.
Oct4 transcriptionally activates NEAT1 via promoter and activates MALAT1 and UCA1 via enhancer binding
To investigate the mechanism underlying Oct4-mediated lncRNAs upregulation, we selected NEAT1, MALAT1 and UCA1 lncRNAs for further transcriptional analyses. Oct4 was validated to bind at NEAT1 promoter region while to MALAT1 and UCA1 at their enhancer regions (Fig. 1c and d). NEAT1 promoter derived from the Oct4 binding ChIP-seq region was inserted upstream of the luciferase reporter gene (Fig. 2a, left). A549 and CL1–0 cells transiently overexpressing Oct4 significantly induced NEAT1 promoter activity (Fig. 2a, yellow bars). However, Oct4 did not affect the activity of NEAT1 promoter with Oct4 binding element mutated in both lung cancer cells (Fig. 2a, green bars). As for the enhancer construction, reported minimal promoter activity fragment of MALAT1 or UCA1 [24, 25] and the enhancer fragment with Oct4 binding ChIP-seq region of MALAT1 or UCA1 was constructed to the upstream and downstream of luciferase reporter, respectively (Fig. 2b and c, left). A549 and CL1–0 cells transiently overexpressing Oct4 potentiated MALAT1 and UCA1 enhancer activities compared with activities with promoter alone (Fig. 2b and c, bars pink vs. yellow). However, the Oct4-induced enhancer activity was attenuated when the Oct4 binding element within MALAT1 or UCA1 enhancer was mutated in A549 and CL1–0 cells (Fig. 2b and c, bars green vs. pink). Together, we have demonstrated that Oct4 directly regulated NEAT1, MALAT1 and UCA1 lncRNAs transcription through targeting their promoter or enhancer regions.
Fig. 2.
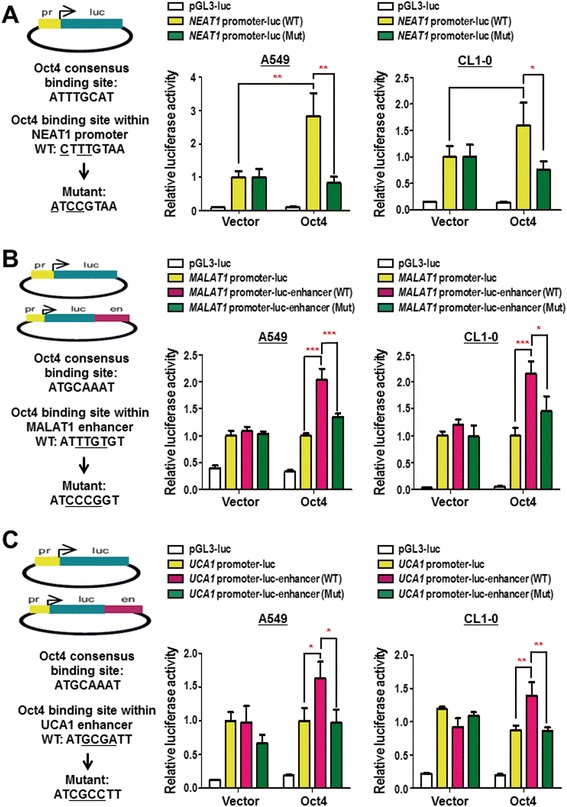
Oct4 promoted oncogenic lncRNAs transcription through activating promoter of NEAT1 and enhancer of MALAT1 and UCA1. a Schematic diagram depicting the construction of promoter activity assay of NEAT1 (upper left). The wild-type (WT) and mutation (Mut) sites at the Oct4 consensus region are shown (lower left). Dual luciferase assay performed in A549 (middle) and CL1–0 (right) cells. b, c Schematic diagram depicting the construction of enhancer activity assay (upper left). The minimal promoters reported were inserted upstream of the luciferase reporter, while the Oct4 binding ChIP-seq regions were inserted downstream as the enhancer plasmids. The WT and Mut sites at the Oct4 consensus region are shown (lower left). Cells were transfected with vector or Oct4 plasmids and luciferase plasmids of NEAT1 (a) MALAT1 (b) or UCA1 (c). Data are mean ± SEM. P-values were determined by two-way ANOVA. *P < 0.05; **P < 0.01; ***P < 0.001
NEAT1 and MALAT1 are downstream effectors of Oct4-induced lung cancer proliferation, migration and invasion
Since oncogenic lncRNAs, NEAT1 and MALAT1, were positively regulated by Oct4 at transcriptional level, we further characterized the oncogenic roles of NEAT1 and MALAT1 lncRNAs in lung cancer cells. Transiently overexpressing NEAT1 or MALAT1 in A549 cells indeed promoted cell proliferation (Fig. 3a and b). In contrast, knockdown of NEAT1 or MALAT1 expression suppressed cell proliferation in A549 cells (Fig. 3c and d). In addition, transwell migration and invasion assays confirmed that NEAT1 or MALAT1 overexpression in A549 cells enhanced cell migration and invasion (Fig. 3e and f), while knockdown of NEAT1 or MALAT1 suppressed cell migration and invasion (Fig. 3g and h). These results indicated that NEAT1 and MALAT1 indeed exerted oncogenic effects in lung cancer cells. The expression levels of NEAT1 and MALAT1 manipulation in functional experiments were confirmed using qRT-PCR analysis and are shown in Additional file 1: Figure S4.
Fig. 3.
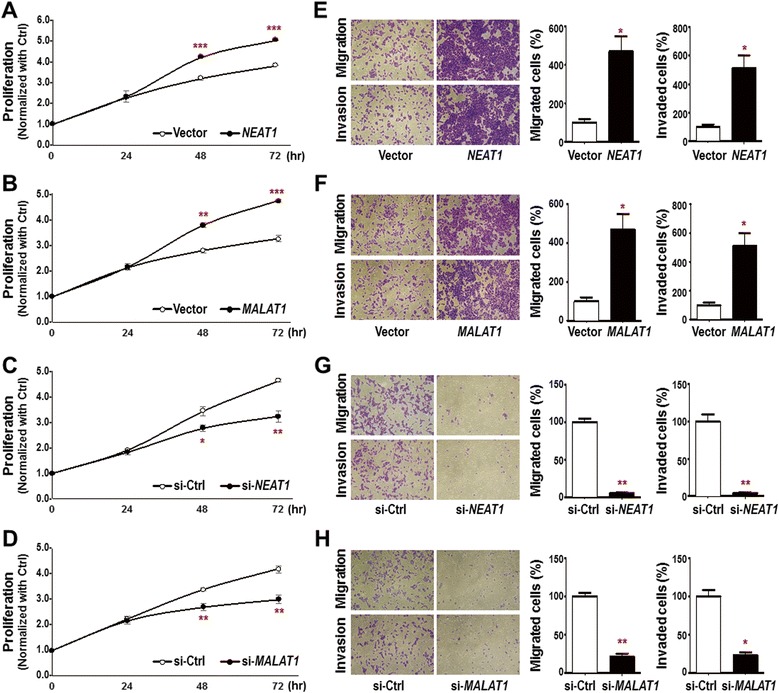
NEAT1 and MALAT1 promoted cell proliferation, migration and invasion in lung cancer cells. a-d Cell proliferation was analyzed by RealTime-Glo viability assay for A549 cells treated with NEAT1 expression vectors (a), MALAT1 expression vectors (b), si-NEAT1 oligo (c) or si-MALAT1 oligo (d) at indicated time points. e-h Cell motility was analyzed by transwell migration and invasion assays for A549 cells treated with NEAT1 expression vectors (e), MALAT1 expression vectors (f), si-NEAT1 oligo (g) or si-MALAT1 oligo (h). Data represent mean ± SEM. P-values were determined by two-tailed Student’s t-test. *P < 0.05; **P < 0.01; ***P < 0.001
To further elucidate whether NEAT1 and MALAT1 contributed to Oct4-mediated oncogenic effects, we conducted reconstitution experiments by transfecting A549 cells with Oct4 expression plasmid alone or together with si-NEAT or si-MALAT1 oligo. On the other hand, A549 cells were transfected with si-Oct4 oligo alone or together with NEAT1 or MALAT1 expression vectors. The expression level of Oct4, NEAT1 and MALAT1 manipulation in reconstitution experiments were confirmed using Western blot and qRT-PCR analyses and results are shown in Additional file 1: Figure S5. The transfected cells were also subjected to proliferation assay, transwell migration and invasion assays. Our results showed that reconstituted expression of NEAT1 or MALAT1 recovered proliferation (Fig. 4a and b), migration and invasion (Fig. 4e and f) abilities, which was downregulated in Oct4-knockdown A549 cells. In contrast, knockdown of NEAT1 or MALAT1 abolished Oct4-promoted A549 cells proliferation (Fig. 4c and d), migration and invasion (Fig. 4g and h) abilities. The results suggested that Oct4/NEAT1 and Oct4/MALAT1 transcriptional axes promote oncogenic effects in lung cancer.
Fig. 4.
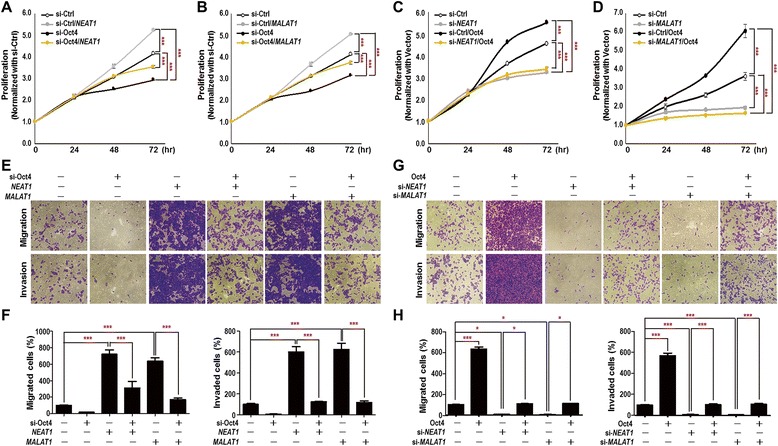
Oct4/NEAT1 or Oct4/MALAT1 signaling axis promoted lung cancer cell proliferation and motility. a-d Cell proliferation assay. Cells were reconstituted with expression vector of NEAT1 (a) or MALAT1 (b) in Oct4-silenced A549 cells, or si-NEAT1 oligo (c) or si-MALAT1 oligo (d) in Oct4-overexpressed A549 cells. Treated cells were then re-seeded and analyzed by RealTime-Glo viability assay at indicated time points. e-h Transwell migration and invasion assays. (e, f). Cells were reconstituted with expression vector of NEAT1 or MALAT1 in Oct4-silenced A549 cells, or (g, h) si-NEAT1 oligo or si-MALAT1 oligo in Oct4-overexpressed A549 cells. Treated cells were re-seeded and analyzed by transwell migration and invasion assays. Results were photographed and quantified. Data represent mean ± SEM. P-values were determined by two-way ANOVA. *P < 0.05; **P < 0.01; ***P < 0.001
Clinical significance of coinciding high expressions of Oct4, NEAT1 and MALAT1 in lung cancer patients
To further validate Oct4/NEAT1 and Oct4/MALAT1 transcriptional axes in lung cancer patients, we examined RNA expression level of Oct4, NEAT1 and MALAT1 using qRT-PCR analysis of samples from 124 lung cancer patients. The overexpression rate for Oct4, NEAT1 and MALAT1 RNA were 85.5%, 90.3% and 88.7%, indicating the oncogenic roles of Oct4, NEAT1 and MALAT1 in lung cancer patients (Table 1). Of note, significant positive correlations were found between Oct4 mRNA and NEAT1 or between Oct4 mRNA and MALAT1 lncRNA expression (P < 0.001 for Oct4 and NEAT1; P < 0.001 for Oct4 and MALAT1) (Table 1).
Table 1.
Alteration of Oct4, NEAT1 and MALAT1 expression levels in relation to clinicopathological parameters in 124 lung cancer patients
| Characteristics | Oct4 | NEAT1 | MALAT1 | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Total | Normal | Overexpression | Normal | Overexpression | Normal | Overexpression | ||||||||||
| n | n | (%) | n | (%) | P a | n | (%) | n | (%) | P a | n | (%) | n | (%) | P a | |
| Overall | 124 | 18 | (14.5) | 106 | (85.5) | 12 | (9.7) | 112 | (90.3) | 14 | (11.3) | 110 | (88.7) | |||
| Tumor stage | ||||||||||||||||
| I + II | 72 | 16 | (22.2) | 56 | (77.8) | 0.005 | 11 | (15.3) | 61 | (84.7) | 0.014 | 12 | (16.7) | 60 | (83.3) | 0.028 |
| III + IV | 51 | 2 | (3.9) | 49 | (96.1) | 1 | (2.0) | 50 | (98.0) | 2 | (3.9) | 49 | (96.1) | |||
| T stage | ||||||||||||||||
| I + II | 104 | 18 | (17.3) | 86 | (82.7) | 0.050 | 11 | (10.6) | 93 | (89.4) | 0.473 | 13 | (12.5) | 91 | (87.5) | 0.361 |
| III + IV | 19 | 0 | (0.0) | 19 | (100.0) | 1 | (5.3) | 18 | (94.7) | 1 | (5.3) | 18 | (94.7) | |||
| N stageb | ||||||||||||||||
| 0 | 71 | 15 | (21.1) | 56 | (78.9) | 0.021 | 11 | (15.5) | 60 | (84.5) | 0.014 | 12 | (16.9) | 59 | (83.1) | 0.029 |
| 1 + 2 | 50 | 3 | (6.0) | 47 | (94.0) | 1 | (2.0) | 49 | (98.0) | 2 | (4.0) | 48 | (96.0) | |||
| M stagec | ||||||||||||||||
| 0 | 116 | 18 | (15.5) | 98 | (84.5) | 0.259 | 12 | (10.3) | 104 | (89.7) | 0.370 | 14 | (12.1) | 102 | (87.9) | 0.329 |
| 1 | 7 | 0 | (0.0) | 7 | (100.0) | 0 | (0.0) | 7 | (100.0) | 0 | (0.0) | 7 | (100.0) | |||
| NEAT1 | ||||||||||||||||
| Normal | 12 | 10 | (83.3) | 2 | (16.7) | <0.001 | − | − | − | − | − | − | − | − | − | |
| Overexpression | 112 | 8 | (7.1) | 104 | (92.9) | − | − | − | − | − | − | − | − | |||
| MALAT1 | ||||||||||||||||
| Normal | 14 | 10 | (71.4) | 4 | (28.6) | <0.001 | 11 | (78.6) | 3 | (21.4) | <0.001 | − | − | − | − | |
| Overexpression | 110 | 8 | (7.3) | 102 | (92.7) | 1 | (0.9) | 109 | (99.1) | − | − | − | − | |||
aThe data were analyzed by Pearson χ2 test. Bold values indicate statistical significance (P < 0.05)
bN Stage: lymph node metastasis
cM Stage: distant metastasis
To determine whether high expression of Oct4, NEAT1 and MALAT1 contributes to poor outcome in lung cancer patients, Kaplan-Meier analysis was performed and data showed that high expression of Oct4 mRNA (P = 0.01), NEAT1 lncRNA (P = 0.006) or MALAT1 lncRNA (P = 0.014) in lung cancer patients was associated with poor overall survival (Fig. 5a-c). Moreover, patients with coinciding high expression of Oct4, NEAT1 and MALAT1 had poorer prognosis compared with other patients (P = 0.002) (Fig. 5d). Univariate Cox regression analysis confirmed that patients with co-overexpressed Oct4, NEAT1 and MALAT1 had poor outcome (P = 0.016, hazard ratio = 2.78, 95% confidence interval = 1.21–6.42; Table 2, left panel). After adjusting for late stage and lymph node metastasis using multivariate Cox regression analysis, co-overexpression of Oct4, NEAT1 and MALAT1 in lung cancer patients showed a relative risk of death of 2.42 (P = 0.039; Table 2, right panel). These clinical studies clearly indicated that Oct4 positively correlates with NEAT1 and MALAT1 expression in lung cancer and that Oct4/NEAT1/MALAT1 co-overexpression is an independent factor for prediction of poor outcome in lung cancer patients.
Fig. 5.
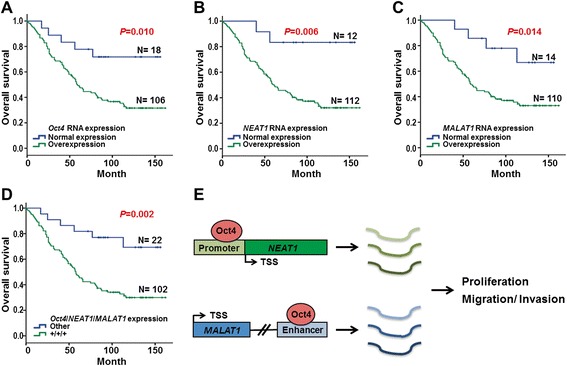
Clinical significances of Oct4/NEAT1/MALAT1 signaling axis in 124 lung cancer patients. qRT-PCR analyses of Oct4, NEAT1 and MALAT1 RNA expression were performed in 124 lung cancer specimens. a-d Overall survival analyzed by Kaplan-Meier method indicated that patients with high RNA expression of Oct4 (a), NEAT1 (b), MALAT1 (c) or coinciding high expression of Oct4, NEAT1 and MALAT1 (d) had poor survival. P-values were determined using log-rank test. e Schematic diagram depicting Oct4 activates NEAT1 transcription via promoter binding and MALAT1 transcription via enhancer binding. NEAT1 and MALAT1 lncRNAs act as downstream effectors of Oct4 to promote lung cancer proliferation, migration and invasion, and thereby lead to lung cancer progression
Table 2.
Cox regression analysis of risk factors for cancer-related death in lung cancer patients
| Characteristics | Univariate analysis | Multivariate analysis | ||
|---|---|---|---|---|
| HRa (95% CIb) | P-valuec | HRa (95% CIb) | P-valuec | |
| Oct4/NEAT1/MALAT1 expression | ||||
| Other | 1.00 | 1.00 | ||
| +/+/+ | 2.78 (1.21–6.42) | 0.016 | 2.42 (1.05–5.62) | 0.039 |
| Age | ||||
| < 65 year-old | 1.00 | - e | ||
| ≥ 65 year-old | 1.03 (0.64–1.67) | 0.896 | - e | - e |
| Gender | ||||
| Female | 1.00 | - e | ||
| Male | 1.50 (0.86–2.61) | 0.150 | - e | - e |
| Smoking habit | ||||
| Non-smoker | 1.00 | - e | ||
| Smoker | 0.96 (0.54–1.72) | 0.891 | - e | - e |
| Tumor typed | ||||
| SCC | 1.00 | - e | ||
| ADC | 1.32 (0.74–2.37) | 0.343 | - e | - e |
| Stage | ||||
| Stage I-II | 1.00 | 1.00 | ||
| Stage III-IV | 1.86 (1.18–2.95) | 0.008 | 1.13 (0.65–1.97) | 0.672 |
| T stage | ||||
| T1–2 | 1.00 | - e | ||
| T3–4 | 1.42 (0.78–2.59) | 0.249 | - e | - e |
| N stage | ||||
| N0 | 1.00 | 1.00 | ||
| ≥ N1 | 2.36 (1.48–3.78) | <0.001 | 1.93 (1.10–3.38) | 0.022 |
| M stage | ||||
| M0 | 1.00 | - e | ||
| ≥ M1 | 1.98 (0.86–4.57) | 0.110 | - e | - e |
a HR hazard ratio
b CI confidence interval
cBold values indicate statistical significance (P < 0.05)
d SCC squamous cell carcinoma, ADC adenocarcinoma
eThe variables without significant HR in the univariate analysis were not included in the multivariate analysis
Discussion
In the present study, we have revealed that Oct4 binds to the genomic loci of lncRNAs through ChIP-seq and bioinformatic analysis (Fig. 1). We then validated that Oct4 bound on the promoter or enhancer regions of lncRNAs (Fig. 1). Dual luciferase activity assay further confirmed that Oct4 potentiated promoter activity of NEAT1 and enhancer activities of MALAT1 and UCA1 lncRNAs (Fig. 2). Moreover, NEAT1 and MALAT1 acted as downstream effectors of Oct4 to promote proliferation, migration and invasion abilities of A549 lung cancer cells (Figs. 3 and 4). Of note, positive correlations between Oct4 mRNA and NEAT1/MALAT1 lncRNAs were evident in lung cancer patient specimens (Fig. 5). Our study provides new evidence that Oct4 transcriptionally regulates lncRNAs expression by targeting their promoter or enhancer regions. NEAT1 and MALAT1 function as Oct4 downstream mediators to promote lung cancer proliferation, migration and invasion (Fig. 5e).
Recently, the roles of NEAT1 in cancer have been uncovered. Studies have reported that NEAT1 is upregulated in prostate cancer, colorectal cancer and lung cancer, and thus associated with poor prognosis in these cancer patients [16, 26–28]. Rubin and associates demonstrated that estrogen receptor transcriptionally activates NEAT1 expression to promote prostate tumorigenesis under the treatment of oestrogen [16]. NEAT1 has also been shown to modulate prostate cancer-specific gene expression through chromatin modifications and thus contributes to cancer progression [16]. However, the upstream mechanisms of NEAT1 overexpression in cancers await to be uncovered. It is until recently that HIF-2α is demonstrated to transactivate NEAT1 transcription under hypoxia, which promotes the formation of paraspeckles, accelerates tumor proliferation and cancer cell survival leading to poor prognosis in breast cancer patient [29]. In our study, we have shown that a well-known stemness transcription factor Oct4 transcriptionally upregulates NEAT1 expression through binding to Oct4 consensus binding element on promoter region (Figs. 1 and 2a), and therefore promoting lung cancer proliferation and motility (Figs. 3 and 4). Notably, NEAT1 is found overexpressed in BRCA1-deficient breast cancer and promotes self-renewal abilities in breast cancer cells through epigenetically suppressing miR-129-5p, which targets to Wnt4 [30]. The last-mentioned study together with our results revealed a potential role of NEAT1 in maintaining stemness properties and suggested that Oct4-mediated NEAT1 upregulation may play critical roles in embryonic or cancer stemness maintenance.
MALAT1 was first identified as a prognosis marker in early-stage metastasizing lung cancer [31]. MALAT1 knockdown in lung cancer cells decreases cell migration abilities [32]. In addition, MALAT1 suppresses expression of anti-metastasis genes such as MIA2 (melanoma inhibitory activity 2) and ROBO1 (roundabout 1), while induces pro-metastasis genes including LPHN2 (latrophilin 2) and ABCA1 (ATP-binding cassette, sub-family A, member 1) to accelerate metastasis [33]. However, MALAT1-promoting lung cancer cell proliferation in different studies are contradictory. For example, MALAT1 has no effect on cell proliferation in vitro and slightly promotes tumor growth in vivo [33]. In contrast, knockdown of MALAT1 in A549 lung cancer cells decreased proliferation [34], which is consistent with our results that MALAT1 plays a role in lung cancer cell proliferation (Fig. 3b and d) and this provides new insight into the role of MALAT1 in various cancer types. MALAT1 has been demonstrated to promote lung, bladder, colorectal, liver, oral and prostate cancer cells proliferation and migration [32, 33, 35–40]. However, the upstream regulatory mechanisms of MALAT1 expression remain unclear, especially at the transcription level. Recently, Sp1 is found to transcriptionally activate MALAT1 expression through targeting the promoter region and the Sp1-MALAT1 axis may play a critical role in cancers [34]. In addition, Wnt signaling pathway acts upstream of MALAT1 transcription, which is mediated by TCF4 binding on MALAT1 promoter in endometrioid endometrial cancer [41]. Importantly, our results provide the first evidence of Oct4-mediated MALAT1 upregulation through enhancer regions.
Conclusions
In conclusion, our genome-wide ChIP-seq analysis reveals a novel role of Oct4 transcription regulation on lncRNAs in lung cancer. We demonstrate for the first time that Oct4 promotes transcription of NEAT1, MALAT1 and UCA1 through targeting their promoter or enhancer regions. Moreover, NEAT1 and MALAT1 function as Oct4 downstream mediators to promote lung cancer proliferation, migration and invasion. Clinical studies confirm that patients with Oct4/NEAT1/MALAT1 high expression had poor outcome. Collectively, our study provides a novel insight into Oc4-lncRNAs as critical axes in lung tumorigenesis.
Acknowledgements
We are grateful to Professors Ying Jin, Tetsuro Hirose and Kannanganattu V. Prasanth for generous help with the Oct4, NEAT1 and MALAT1 expression vectors. We thank the Human Biobank, Research Center of Clinical Medicine, National Cheng Kung University Hospital for providing the lung cancer specimens.
Funding
This work was supported by the Taiwan Ministry of Science and Technology Grant (MOST103-2320-B-006-045-MY3), the Taiwan Ministry of Health and Welfare Grant (MOHW103-TD-B-111-06) and the Aim for the Top University Project Grant (D103-35A02).
Availability of data and materials
All data used during the current study available from the corresponding author on reasonable request.
Authors’ contributions
JJ, YAT, YHL and CCL performed the experiments. JJ, YAT, YHL and CCL did the data analysis in this study. WWL provided clinical samples. JJ, YHL and YCW wrote the paper. All authors read and approved the manuscript. YCW obtained funding.
Competing interests
The authors declare that they have no competing interests.
Consent for publication
Not applicable.
Ethics approval and consent to participate
This study was approved by the Institutional Review Board of National Cheng Kung University Hospital (B-ER-102-451), and patient consent was obtained prior to the initiation of the study.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Abbreviations
- ChIP
Chromatin-immunoprecipitation
- lncRNA
Long non-coding RNA
- MALAT1
Metastasis-associated lung adenocarcinoma transcript 1
- NEAT1
Nuclear paraspeckle assembly transcript 1
- POU5F1
POU domain, class 5, transcription factor 1
- UCA1
Urothelial carcinoma-associated 1
Additional file
Supplementary methods. Anchorage-independent growth assay, Tumor-sphere formation assay, Tumor formation assay, Western blot analysis. Table S1. The plasmids and their characteristics used in the current study. Table S2. The ChIP-PCR primers used in the current study. Table S3. The cDNA primers used in the current study. Table S4. The construction primers of promoters and enhancers used in the current study. Figure S1. Oct4 promoted lung cancer tumorigenesis in vitro and in vivo. A Anchorage-independent assays in empty vector stably-transfected cell line (vector) and two biological replicates of Oct4 stably-overexpressed A549 and CL1-0 cell lines (Oct4#1, Oct4#2). Results were photographed (left) and quantified (right). B Transwell migration and invasion assay analysis of stably-transfected cell lines in A549 and CL1-0 cells. Results were photographed (left) and quantified (right). P-values were determined by two-tailed Student’s t-test. *P < 0.05; **P < 0.01; ***P < 0.001. C In vitro tumor sphere formation assay of A549 lung cancer cells stably expressing Oct4#1 or vector photographed (top) and quantified (middle). In vivo tumor formation assay using limited cell number (100, 1000, and 5000 cells) of vector and Oct4#1 cells. Tumor incidence of mice was analyzed at 8 weeks after implantation. D The immunoblots (upper) and qRT-PCR (lower) confirmed Oct4 expression in A549 and CL1–0 stable clones. Figure S2. Expression of lncRNAs in CL1–0 lung cancer cells manipulated for Oct4. A, B qRT-PCR analysis of eight lncRNAs expression in CL1-0 cells stably overexpressing Oct4 (Oct4#1, Oct4#2) (A) or Oct4-silenced CL1-0 cells (si-Oct4#1, si-Oct4#2) (B). Target lncRNA expression levels were normalized to GAPDH expression levels. Data represent mean ± SEM. P-values were determined by two-tailed Student’s t-test. *P < 0.05; **P < 0.01; ***P < 0.001. Figure S3. Oct4 positively regulated NEAT1 and UCA1 lncRNAs transcription in normal bronchial epithelial BEAS-2B cells. qRT-PCR analysis of selected lncRNAs expressions in BEAS-2B cells overexpressing Oct4. Target lncRNA expression levels were normalized to GAPDH expression levels. Data represent mean ± SEM. P-values were determined by two-tailed Student’s t-test. *P < 0.05; **P < 0.01; ***P < 0.001. Figure S4. RNA expression level of the manipulated NEAT1 and MALAT1 in A549 lung cancer cells. A549 cells transfected with NEAT1 expression vector (A) or MALAT1 expression vector (B), si-NEAT1 oligo (si-NEAT1) (C) or si-MALAT1 oligo (si-MALAT1) (D) were harvested and subjected to qRT-PCR assays for NEAT1 and MALAT1 RNA expression. Data are mean ± SEM. P-values were determined by two-tailed Student’s t-test. *P < 0.05; ***P < 0.001. Figure S5. RNA and protein expression level of the manipulated Oct4, NEAT1 and MALAT1 in A549 lung cancer cells. A549 cells were transfected with expression vectors of NEAT1 (A) or MALAT1 (B) alone or together with si-Oct4 oligo (si-Oct4). A549 cells were transfected with si-NEAT1 oligo (si-NEAT1) (C) or si-MALAT1 oligo (si-MALAT1) (D) alone or together with Oct4 expression vector. Cell lysates were subjected to qRT-PCR assays for Oct4, NEAT1 and MALAT1 RNA expression or Western blot analysis for Oct4 protein expression (inset). GAPDH serves as an internal control. Data are mean ± SEM. P-values were determined by two-way ANOVA. **P < 0.01; ***P < 0.001. (PDF 911 kb)
Footnotes
Electronic supplementary material
The online version of this article (doi:10.1186/s12943-017-0674-z) contains supplementary material, which is available to authorized users.
Contributor Information
Jayu Jen, Email: jayujen1021@gmail.com.
Yen-An Tang, Email: alan.leimow@msa.hinet.net.
Ying-Hung Lu, Email: b091082008@hotmail.com.
Che-Chung Lin, Email: jackie8407@gmail.com.
Wu-Wei Lai, Email: wwlai@mail.ncku.edu.tw.
Yi-Ching Wang, Phone: +886-6-2353535, Email: ycw5798@mail.ncku.edu.tw.
References
- 1.Wang Z, Oron E, Nelson B, Razis S, Ivanova N. Distinct lineage specification roles for NANOG, OCT4, and SOX2 in human embryonic stem cells. Cell Stem Cell. 2012;10:440–454. doi: 10.1016/j.stem.2012.02.016. [DOI] [PubMed] [Google Scholar]
- 2.Chang CC, Shieh GS, Wu P, Lin CC, Shiau AL, Wu CL. Oct-3/4 expression reflects tumor progression and regulates motility of bladder cancer cells. Cancer Res. 2008;68:6281–6291. doi: 10.1158/0008-5472.CAN-08-0094. [DOI] [PubMed] [Google Scholar]
- 3.Ben-Porath I, Thomson MW, Carey VJ, Ge R, Bell GW, Regev A, Weinberg RA. An embryonic stem cell-like gene expression signature in poorly differentiated aggressive human tumors. Nat Genet. 2008;40:499–507. doi: 10.1038/ng.127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Kim BW, Cho H, Choi CH, Ylaya K, Chung JY, Kim JH, Hewitt SM. Clinical significance of OCT4 and SOX2 protein expression in cervical cancer. BMC Cancer. 2015;15:1015. doi: 10.1186/s12885-015-2015-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Chiou SH, Yu CC, Huang CY, Lin SC, Liu CJ, Tsai TH, Chou SH, Chien CS, Ku HH, Lo JF. Positive correlations of Oct-4 and Nanog in oral cancer stem-like cells and high-grade oral squamous cell carcinoma. Clin Cancer Res. 2008;14:4085–4095. doi: 10.1158/1078-0432.CCR-07-4404. [DOI] [PubMed] [Google Scholar]
- 6.Chai S, Ng KY, Tong M, Lau EY, Lee TK, Chan KW, Yuan YF, Cheung TT, Cheung ST, Wang XQ, et al. Octamer 4/microRNA-1246 signaling axis drives Wnt/β-catenin activation in liver cancer stem cells. Hepatology. 2016;64:2062–2076. doi: 10.1002/hep.28821. [DOI] [PubMed] [Google Scholar]
- 7.Karoubi G, Cortes-Dericks L, Gugger M, Galetta D, Spaggiari L, Schmid RA. Atypical expression and distribution of embryonic stem cell marker, OCT4, in human lung adenocarcinoma. J Surg Oncol. 2010;102:689–698. doi: 10.1002/jso.21665. [DOI] [PubMed] [Google Scholar]
- 8.Tang YA, Chen CH, Sun HS, Cheng CP, Tseng VS, Hsu HS, Su WC, Lai WW, Wang YC. Global Oct4 target gene analysis reveals novel downstream PTEN and TNC genes required for drug-resistance and metastasis in lung cancer. Nucleic Acids Res. 2015;43:1593–1608. doi: 10.1093/nar/gkv024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Ng HH, Surani MA. The transcriptional and signalling networks of pluripotency. Nat Cell Biol. 2011;3:490–496. doi: 10.1038/ncb0511-490. [DOI] [PubMed] [Google Scholar]
- 10.Wu T, Pinto HB, Kamikawa YF, Donohoe ME. The BET family member BRD4 interacts with OCT4 and regulates pluripotency gene expression. Stem Cell Rep. 2015;4:390–403. doi: 10.1016/j.stemcr.2015.01.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Loewer S, Cabili MN, Guttman M, Loh YH, Thomas K, Park IH, Garber M, Curran M, Onder T, Agarwal S, et al. Large intergenic non-coding RNA-RoR modulates reprogramming of human induced pluripotent stem cells. Nat Genet. 2010;42:1113–1117. doi: 10.1038/ng.710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Boo K, Bhin J, Jeon Y, Kim J, Shin HJ, Park JE, Kim K, Kim CR, Jang H, Kim IH, et al. Pontin functions as an essential coactivator for Oct4-dependent lincRNA expression in mouse embryonic stem cells. Nat Commun. 2015;6:6810. doi: 10.1038/ncomms7810. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Moran VA, Perera RJ, Khalil AM. Emerging functional and mechanistic paradigms of mammalian long non-coding RNAs. Nucleic Acids Res. 2012;40:6391–6400. doi: 10.1093/nar/gks296. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.St Laurent G, Wahlestedt C, Kapranov P. The landscape of long noncoding RNA classification. Trends Genet. 2015;31:239–251. doi: 10.1016/j.tig.2015.03.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Gupta RA, Shah N, Wang KC, Kim J, Horlings HM, Wong DJ, Tsai MC, Hung T, Argani P, Rinn JL, et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature. 2010;464:1071–1076. doi: 10.1038/nature08975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Chakravarty D, Sboner A, Nair SS, Giannopoulou E, Li R, Hennig S, Mosquera JM, Pauwels J, Park K, Kossai M, et al. The oestrogen receptor alpha-regulated lncRNA NEAT1 is a critical modulator of prostate cancer. Nat Commun. 2014;5:5383. doi: 10.1038/ncomms6383. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Tsai MC, Manor O, Wan Y, Mosammaparast N, Wang JK, Lan F, Shi Y, Segal E, Chang HY. Long noncoding RNA as modular scaffold of histone modification complexes. Science. 2010;329:689–693. doi: 10.1126/science.1192002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Li Z, Zhao X, Zhou Y, Liu Y, Zhou Q, Ye H, Wang Y, Zeng J, Song Y, Gao W, et al. The long non-coding RNA HOTTIP promotes progression and gemcitabine resistance by regulating HOXA13 in pancreatic cancer. J Transl Med. 2015;13:84. doi: 10.1186/s12967-015-0442-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Wang KC, Yang YW, Liu B, Sanyal A, Corces-Zimmerman R, Chen Y, Lajoie BR, Protacio A, Flynn RA, Gupta RA, et al. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature. 2011;472:120–124. doi: 10.1038/nature09819. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Niazi F, Valadkhan S. Computational analysis of functional long noncoding RNAs reveals lack of peptide-coding capacity and parallels with 3′ UTRs. RNA. 2012;18:825–843. doi: 10.1261/rna.029520.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Quek XC, Thomson DW, Maag JL, Bartonicek N, Signal B, Clark MB, Gloss BS, Dinger ME. lncRNAdb v2.0: expanding the reference database for functional long noncoding RNAs. Nucleic Acids Res. 2015;43:D168–D173. doi: 10.1093/nar/gku988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Alheim K, Corness J, Samuelsson MK, Bladh LG, Murata T, Nilsson T, Okret S. Identification of a functional glucocorticoid response element in the promoter of the cyclin-dependent kinase inhibitor p57Kip2. J Mol Endocrinol. 2003;30:359–368. doi: 10.1677/jme.0.0300359. [DOI] [PubMed] [Google Scholar]
- 23.Cruickshanks HA, McBryan T, Nelson DM, Vanderkraats ND, Shah PP, van Tuyn J, Singh Rai T, Brock C, Donahue G, Dunican DS, et al. Senescent cells harbour features of the cancer epigenome. Nat Cell Biol. 2013;15:1495–1506. doi: 10.1038/ncb2879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Wang X, Li M, Wang Z, Han S, Tang X, Ge Y, Zhou L, Zhou C, Yuan Q, Yang M. Silencing of long noncoding RNA MALAT1 by miR-101 and miR-217 inhibits proliferation, migration and invasion of esophageal squamous cell carcinoma cells. J Biol Chem. 2014;290:3925–3935. doi: 10.1074/jbc.M114.596866. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Wu W, Zhang S, Li X, Xue M, Cao S, Chen W. Ets-2 regulates cell apoptosis via the Akt pathway, through the regulation of urothelial cancer associated 1, a long non-coding RNA, in bladder cancer cells. PLoS One. 2013;8:e73920. doi: 10.1371/journal.pone.0073920. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Li Y, Li Y, Chen W, He F, Tan Z, Zheng J, Wang W, Zhao Q, Li J. NEAT expression is associated with tumor recurrence and unfavorable prognosis in colorectal cancer. Oncotarget. 2015;6:27641–27650. doi: 10.18632/oncotarget.4737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Pan LJ, Zhong TF, Tang RX, Li P, Dang YW, Huang SN, Chen G. Upregulation and clinicopathological significance of long non coding NEAT1 RNA in NSCLC tissues. Asian Pac J Cancer Prev. 2015;16:2851–2855. doi: 10.7314/APJCP.2015.16.7.2851. [DOI] [PubMed] [Google Scholar]
- 28.Sun C, Li S, Zhang F, Xi Y, Wang L, Bi Y, et al. Long non-coding RNA NEAT1 promotes non-small cell lung cancer progression through regulation of miR-377-3p-E2F3 pathway. Oncotarget. 2016; doi:10.18632/oncotarget.10108. [DOI] [PMC free article] [PubMed]
- 29.Choudhry H, Albukhari A, Morotti M, Haider S, Moralli D, Smythies J, Schödel J, Green CM, Camps C, Buffa F, et al. Tumor hypoxia induces nuclear paraspeckle formation through HIF-2α dependent transcriptional activation of NEAT1 leading to cancer cell survival. Oncogene. 2015;34:4482–4490. doi: 10.1038/onc.2014.378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Lo PK, Zhang Y, Wolfson B, Gernapudi R, Yao Y, Duru N, et al. Dysregulation of the BRCA1/long non-coding RNA NEAT1 signaling axis contributes to breast tumorigenesis. Oncotarget. 2016; doi:10.18632/oncotarget.11364. [DOI] [PMC free article] [PubMed]
- 31.Ji P, Diederichs S, Wang W, Böing S, Metzger R, Schneider PM, Tidow N, Brandt B, Buerger H, Bulk E, et al. MALAT-1, a novel noncoding RNA, and thymosin beta4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene. 2003;22:8031–8041. doi: 10.1038/sj.onc.1206928. [DOI] [PubMed] [Google Scholar]
- 32.Schmidt LH, Spieker T, Koschmieder S, Schäffers S, Humberg J, Jungen D, Bulk E, Hascher A, Wittmer D, Marra A, et al. The long noncoding MALAT-1 RNA indicates a poor prognosis in non-small cell lung cancer and induces migration and tumor growth. J Thorac Oncol. 2011;6:1984–1992. doi: 10.1097/JTO.0b013e3182307eac. [DOI] [PubMed] [Google Scholar]
- 33.Gutschner T, Hämmerle M, Eissmann M, Hsu J, Kim Y, Hung G, Revenko A, Arun G, Stentrup M, Gross M, et al. The noncoding RNA MALAT1 is a critical regulator of the metastasis phenotype of lung cancer cells. Cancer Res. 2013;73:1180–1189. doi: 10.1158/0008-5472.CAN-12-2850. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Li S, Wang Q, Qiang Q, Shan H, Shi M, Chen B, Zhao S, Yuan L. Sp1 mediated transcriptional regulation of MALAT1 plays a critical role in tumor. J Cancer Res Clin Oncol. 2015;141:1909–1920. doi: 10.1007/s00432-015-1951-0. [DOI] [PubMed] [Google Scholar]
- 35.Tano K, Mizuno R, Okada T, Rakwal R, Shibato J, Masuo Y, Ijiri K, Akimitsu N. MALAT-1 enhances cell motility of lung adenocarcinoma cells by influencing the expression of motilityrelated genes. FEBS Lett. 2010;584:4575–4580. doi: 10.1016/j.febslet.2010.10.008. [DOI] [PubMed] [Google Scholar]
- 36.Fan Y, Shen B, Tan M, Mu X, Qin Y, Zhang F, Liu Y. TGF-β-induced upregulation of malat1 promotes bladder cancer metastasis by associating with suz12. Clin Cancer Res. 2014;20:1531–1541. doi: 10.1158/1078-0432.CCR-13-1455. [DOI] [PubMed] [Google Scholar]
- 37.Xu C, Yang M, Tian J, Wang X, Li Z. MALAT-1: a long non-coding RNA and its important 3′ end functional motif in colorectal cancer metastasis. Int J Oncol. 2011;39:169–175. doi: 10.3892/ijo.2011.1007. [DOI] [PubMed] [Google Scholar]
- 38.Lai MC, Yang Z, Zhou L, Zhu QQ, Xie HY, Zhang F, Wu LM, Chen LM, Zheng SS. Long non-coding RNA MALAT-1 overexpression predicts tumor recurrence of hepatocellular carcinoma after liver transplantation. Med Oncol. 2012;29:1810–1816. doi: 10.1007/s12032-011-0004-z. [DOI] [PubMed] [Google Scholar]
- 39.Zhou X, Liu S, Cai G, Kong L, Zhang T, Ren Y, Wu Y, Mei M, Zhang L, Wang X. Long non coding RNA MALAT1 promotes tumor growth and metastasis by inducing epithelial-Mesenchymal transition in oral squamous cell carcinoma. Sci Rep. 2015;5:15972. doi: 10.1038/srep15972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Wang F, Ren S, Chen R, Lu J, Shi X, Zhu Y, Zhang W, Jing T, Zhang C, Shen J, et al. Development and prospective multicenter evaluation of the long noncoding RNA MALAT-1 as a diagnostic urinary biomarker for prostate cancer. Oncotarget. 2014;5:11091–11102. doi: 10.18632/oncotarget.2691. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Zhao Y, Yang Y, Trovik J, Sun K, Zhou L, Jiang P, Lau TS, Hoivik EA, Salvesen HB, Sun H, et al. A novel wnt regulatory axis in endometrioid endometrial cancer. Cancer Res. 2014;74:5103–5117. doi: 10.1158/0008-5472.CAN-14-0427. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
All data used during the current study available from the corresponding author on reasonable request.


