Abstract
Improving our understanding of biological motors, both to fully comprehend their activities in vital processes, and to exploit their impressive abilities for use in bionanotechnology, is highly desirable. One means of understanding these systems is through the production of synthetic molecular motors. We demonstrate the use of orthogonal coiled-coil dimers (including both parallel and antiparallel coiled coils) as a hub for linking other components of a previously described synthetic molecular motor, the Tumbleweed. We use circular dichroism, analytical ultracentrifugation, dynamic light scattering, and disulfide rearrangement studies to demonstrate the ability of this six-peptide set to form the structure designed for the Tumbleweed motor. The successful formation of a suitable hub structure is both a test of the transferability of design rules for protein folding as well as an important step in the production of a synthetic protein-based molecular motor.
Protein-based molecular machines are a vital component of living cells and carry out diverse functions including the transportation of numerous cargoes, muscle contraction, and cell division.1,2 Biological motors are highly complex, regulated machines, making them challenging to study but also a motivating target for use in bionanotechnology.3,4 Within the field of synthetic biology, the bottom-up approach5,6 aims to start with simplified, well-characterized, components and modify or extend their functionality, in this case to produce novel nanoscale motors. A key design challenge in this area is the level of reproducibility and transferability of components from one context to another, and determining the ability of components to be combined without unexpected interactions occurring is important.
Entirely synthetic, biologically inspired motors have been designed with varying levels of ability and mimicry, including small molecule machines such as switches, shuttles, and even a robotic arm.7,8 Using more biological starting materials, there has also been significant progress in the field of autonomous DNA motors9,10 and hybrid DNA–protein motors.11 Neither of these areas, however, utilize the same set of initial building blocks as nature: amino acids. The field of synthetic protein motors is far less developed due to the higher complexity of amino acid interactions. There are, however, several examples of combining peptides with small molecules to create synthetic machines12−14 and structures such as a cyclic peptide-based nanopore,15 but there are, to our knowledge, no peptide or protein-based synthetic motors that avoid the use of existing protein motor domains.
This paper describes progress in the realization of a series of previously published motor designs, including the Tumbleweed and SKIP motors.16,17 These motors are designed to undergo directed motion through rectified diffusion using ligand-controlled binding of their “feet” to a DNA track. A peptide “hub” is proposed16 to self-assemble, connect, and organize E. coli protein based “feet” with the ability to bind orthogonal DNA sequences (in the presence of orthogonal ligands) to form the motor molecule (see Figure 1). Construction of a suitable DNA track18 and microfluidic device19,20 have since been demonstrated, along with further simulation studies.21,22
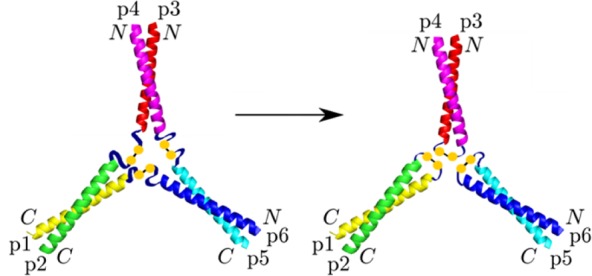
Figure 1.
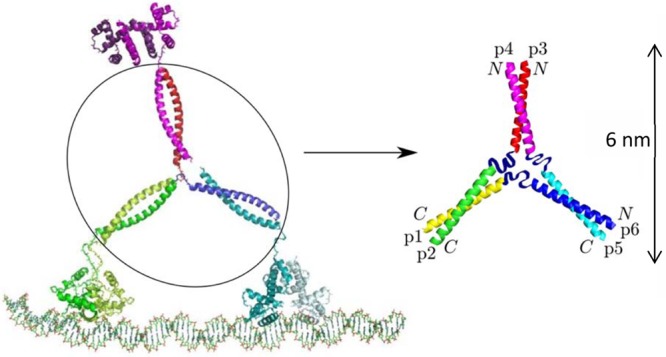
“Y” configuration of the Tumbleweed Hub. The peptide hub termini are indicated, showing the parallel coiled coils p1,p2 and p3,p4, and the antiparallel coiled coil p5,p6. To form a hub, coiled-coil pairs (p1,p2), (p3,p4), and (p5,p6) need to preferentially interact, while p1 and p6, p2 and p3, and p4 and p5 are covalently linked, to ensure that all of the coiled coils are connected to form one structure. The nonhelical central ribbons represent the residues at one end of every peptide that are designed not to contribute to a coiled-coil region. Figure adapted with permission from ref (22). Copyright 2011 American Physical Society.
The proposed hub is composed of three peptide sequences, each of which contains two coiled-coil domains,23 with each domain programmed to assemble with a specified partner to produce the final configuration (see Figure 1). Coiled-coil domains have been selected for this application, primarily due to their ability to undergo programmed assembly; however, the rigidity of the folded domain is also desirable in reducing the probability of backward stepping and hence improving the efficiency of the final motor. The use of coiled-coils in this way is not straightforward as individual coiled-coil sequences have often been found to interact with many other potential partners rather than interacting only with their biologically relevant partner.24,25 It has also been documented in other areas of synthetic biology that once the complexity of synthetic systems increases, unintended cross-talk between components can occur.26,27
To design the high fidelity sequences required in this application we must therefore draw on the wealth of information now available on the folding and interactions of coiled coils.23,28,29 The coiled-coil nanostructure field has advanced on many fronts in recent years including the use of biological coiled-coil domains in novel combinations,30,31 the design of self-assembling peptides from first-principles,32−34 and the use of computational design tools.35−37 From these works we anticipate the need to use a heptad repeat of hydrophobic residues with polar insertions used to specify the orientation of helix interactions. Given that our peptides will each contain two coiled-coil domains, we must consider the formation of intramolecular hairpins to be a primary route to misfolding and use the combination of charged amino acids and polar insertions to the hydrophobic core to specifically design against this eventuality in each peptide. In addition to the coiled-coil oligomerization information, knowledge on how to link domains successfully must be extracted from recent studies38,39 and applied in order to provide sufficient flexibility between domains to allow folding.
In this work we have designed, synthesized, and characterized a linked system of three coiled coils, suitable for the formation of a hub for the Tumbleweed motor. The design in this work is presented as a proof of principle that the combined knowledge within the field can be successfully applied to a range of new applications requiring the ordered colocalization of multiple functional components, including the design of hubs for protein-based synthetic molecular motors.
Results and Discussion
Using the design principles laid out previously38,40 six peptides were designed to form three dimeric coiled coils: (p1,p2), (p3,p4), and (p5,p6). Polar residues are included in various hydrophobic core positions to prevent staggered assembly and electrostatic interactions are optimized to produce the desired dimeric pairings using an algorithm reported previously.40 All contain a tyrosine residue for UV absorption concentration measurements, a single cysteine residue for disulfide bond formation, and a linker of glycine and asparagine intended to form a flexible region outside the coiled-coil domains on one terminus.
The sequences of the six peptides labeled in Figure 1 are shown in Table 1. These were synthesized using solid phase peptide synthesis and purified with high performance liquid chromatography. Several facts must be established in order to confirm the success of the design principles and demonstrate that when all six peptides are mixed, the three required dimeric coiled-coil structures are produced. First we must demonstrate that the designed peptide pairs interact with each other. Second we must show that the structures formed by these pairs are dimeric in nature. Finally we must find that the three designed pairs are produced preferentially over other possible oligomers.
Table 1.
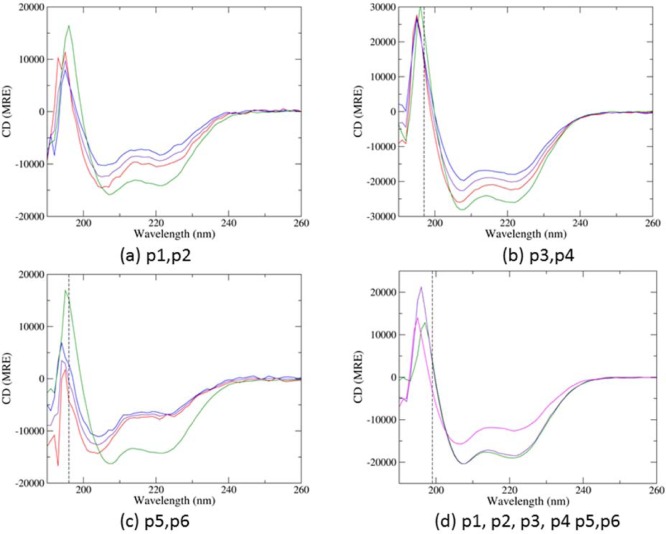
The secondary structures of the peptides and two-peptide combinations were studied with circular dichroism (CD). The individual peptides displayed varying levels of helicity (see Figure S1a). All 15 two-peptide mixtures were made and measured. The CD spectra of the three specified dimers, alongside the spectra of the component peptides, are shown in Figure 2 panels a, b, and c. All three pairs indicate the presence of heterooligomeric interactions, as the measured spectra of the mixtures show greater helicity than those predicted from the sum of the individual component peptide spectra. The data from the other 12 possible combinations of two hub peptides also demonstrate various levels of heterooligomeric interactions to be present. These data show that the specified pairs can be formed. To determine whether the desired pairs are formed preferentially to other interactions, all six peptides were mixed together. The CD spectrum of this mixture is shown in Figure 2d. This has been compared to predicted spectra for six noninteracting species, and to a sample comprising only the three designed pairs. As expected, the combined spectrum is highly similar to that predicted under the assumption that no further interactions are produced when the designed pairs are mixed.
Figure 2.
CD spectra of the three mixtures of the designed pairs of peptides: (a) p1 and p2, (b) p3 and p4, and (c) p5 and p6 (shown in green). Each panel also shows the individual peptide spectra (red and blue, in numerical order), and the theoretical signal for a mixture of the two without interaction (a combination of the two single peptide signals, shown in purple). (d) CD spectrum of all six hub peptides mixed together (green), compared with the theoretical spectrum predicted from the individual spectra (shown in magenta), and from the designed pair spectra (shown in purple). Each sample had a concentration of 20 μM per peptide, in PBS buffer (pH 7.4), measured at 20 °C. Dotted lines represent the wavelength cutoff below which data cannot be reliably obtained, as dictated by the CD instrument.
Analytical ultracentrifugation (AUC) measurements of the designed coiled-coil pairs (Table 2) gave molecular weights consistent with dimer formation, and inconsistent with monomers or oligomers of a higher order than dimers (see Supporting Information, Tables S1 and S2 for further details).
Table 2.
| sample | expected molecular weight/Da | measured molecular weight/Da |
|---|---|---|
| p1,p2 | 7041 | 7000 ± 1000 |
| p3,p4 | 7397 | 7100 ± 1000 |
| p5,p6 | 5816 | 5900 ± 1000 |
Likewise, dynamic light scattering (DLS) measurements of the designed coiled-coil pairs each exhibited species with 3–5 nm hydrodynamic diameters, consistent with the hypothesis of the formation of discrete coiled-coil oligomers, and inconsistent with the formation of fibers (Figure S2).
To demonstrate the fidelity of the interactions and to confirm the utility of the peptides in organizing the motor hub, pairs of the peptides were covalently linked using a disulfide bond between terminal cysteine residues, and the disulfide-bonded peptides, p6–p1, p3–p2, and p4–p5 were produced and characterized.
The CD spectra of the disulfide-bonded peptides reveal little about the system as all of the collected spectra show high levels of helicity (see SI Figure S1b). This is expected for a system where the peptides have been designed to form coiled-coils and have been colocalized by the disulfide tethering causing conditions of high local concentration and hence nonspecific coiled-coil formation.
AUC results are more informative about the system’s interactions (see Table 3). The single disulfide-bonded peptides have the expected masses. (The higher than expected value found for p2–p3 nonetheless corresponds well to a single species fit, and the mass found is far closer to the mass of a single disulfide-bonded peptide than to a pair.) To study the specificity of coiled-coil interactions, the disulfide-bonded peptides were mixed pairwise and allowed to associate. The mixtures of pairs of disulfide-bonded peptides exhibit single species behavior, indicative of the desired specificity of interaction. Furthermore, this species in each case has a mass consistent with the interaction of two disulfide-bonded peptides. The mixture of all three disulfide-bonded peptides gives a mass consistent with the expected mass of 20 248 Da, again supportive of our design criteria.
Table 3.
| sample | expected heterodimeric molecular weight/Da | measured molecular weight/Da |
|---|---|---|
| p6–p1 | 6470 | 7300 ± 1000 |
| p2–p3 | 7096 | 9700 ± 1000 |
| p4–p5 | 6682 | 7600 ± 1000 |
| p6–p1, p2–p3 | 13566 | 16000 ± 1000 |
| p2–p3, p4–p5 | 13778 | 14000 ± 1000 |
| p4–p5, p6–p1 | 13151 | 16000 ± 1000 |
| p6–p1, p2–p3, p4–p5 | 20248 | 19000 ± 1000 |
DLS data of the individual disulfide-bonded peptides gave hydrodynamic diameters of 3–4 nm, while DLS data of pairs of the disulfide-bonded peptides, and of all three disulfide-bonded peptides mixed, showed a shift to higher values, around 5 nm (see Figure S3). The change in size between the single and pairs of disulfide-bonded peptides indicates that there is interaction between the disulfide-bonded peptides. The fact that the size of the three-component mixture is similar to that of the pairs of disulfide-bonded peptides is supportive that a structure such as that shown in Figure 1 is formed, and not a lengthy fiber.
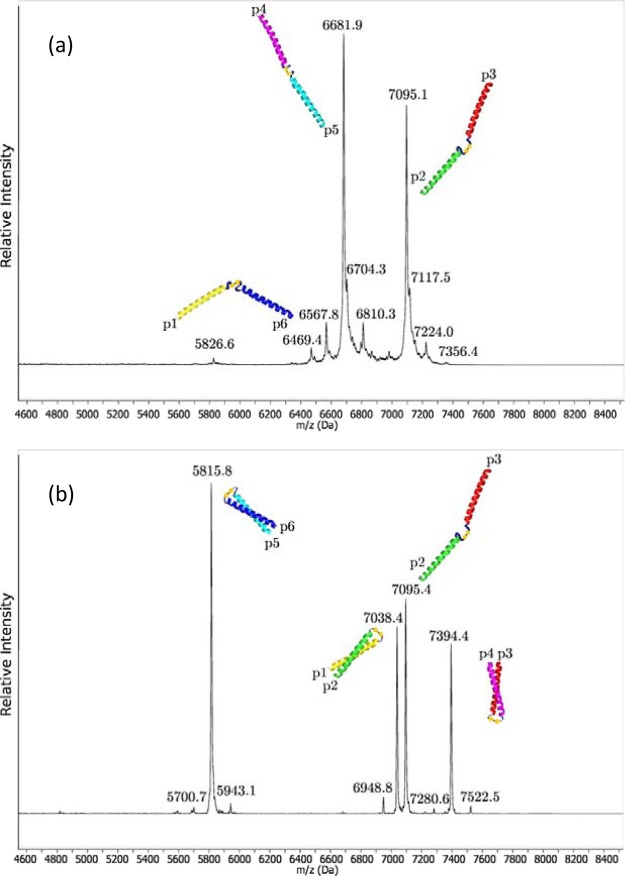
To further verify that the interactions occurring in the final structure are due to the three specified peptide pairs, we carried out disulfide exchange experiments on the linked peptides. We measured the system of three disulfide-bonded peptides, and then changed the system’s redox potential, using reduced glutathione, to allow the disulfide bonds to rearrange. If there are preferred interactions, the disulfide bonds formed when rearrangement is possible should reflect these preferences (see Figure 3). We measured the species in each of these samples using MALDI-ToF mass spectrometry, and the results are shown in Figure 4. Prior to rearrangement, the species present in the system are the disulfide-bonded peptides produced in the laboratory (see Figure 4a). Once the disulfide bonds are given the opportunity to rearrange (see Figure 4b), the species present change. Disulfide-bonded versions of all three of the designed pairs (p1–p2, p3–p4, p6–p5) produce the dominant peaks, along with a fourth peak attributed to the starting peptide 2–3 (that is likely due to a failure to produce a precise 1:1:1 ratio of the starting species). Regardless of the presence of this peak, it is clear that the three designed pairs interact when given the opportunity, while none of the other possible combinations not present in the initial system form during rearrangement (Supporting Information Table S3 for details of peak assignments).
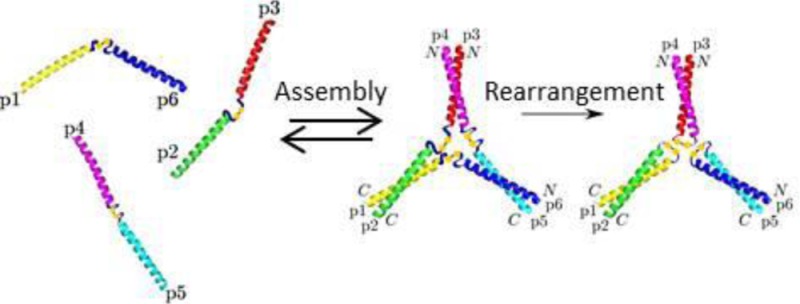
Figure 3.
If the designed dimer peptides preferentially interact, when allowed to rearrange, the disulfide bonds should switch to being between the desired pairs, with the species present in the system moving from p6–p1, p3–p2, and p4–p5 to p1–p2, p3–p4, and p5–p6.
Figure 4.
MALDI spectra of glutathione experiments. The initial spectrum has prominent peaks due to the p4–p5 (MW 6682 Da) and p3–p2 (MW 7095 Da) species, and smaller peaks due to deletions and the p6–p1 peptide (MW 6470 Da). After rearrangement, the p4–p5 and p6–p1 species are no longer seen, while the p6–p5 (MW 5814 Da), p1–p2 (MW 7039 Da) and p3–p4 (MW 7395 Da) species have appeared. All of the trace peaks correlate with impurities in the initial samples and do not correlate with possible rearrangements.
This combined information from CD, DLS, AUC, and disulfide rearrangement experiments indicates that the hub construction should indeed be possible from mixing covalently linked species of peptides p1 and p6, p2 and p3, and p4 and p5, driven by coiled-coil interactions. As such, this has been a successful test of the hypothesis that the knowledge within the peptide design field is now sufficient for the design of self-assembling hubs for diverse applications, including the colocalization of other motor components.
Future extensions of this work could investigate whether it is possible to combine this synthetic biology approach to structure design with active means of actuating the structure. Using the observed ability of peptides41−43 to switch conformation in different conditions could provide a feasible means of adapting this system to undergo conformational changes (and hence changes in movement) in the presence of different stimuli. This will take us one step closer to realizing designed protein motion.
Conclusions
We have demonstrated that a set of six coiled-coil domains can be designed to self-assemble preferentially into three coiled-coil heterodimers. We have shown that by pairing the domains covalently, a hub structure can be formed. This hub can then be used to assemble multiple functional units into the desired 3D configuration. We propose using this technology to produce designed protein motor hubs; however, the methodology exploited here may have wider applications in the design and production of self-assembling junctions in other areas of bionanotechnology including the construction of synthetic cages or other nanoscale devices.
Materials and Methods
Amino acids and PyBOP were purchased from AGTC Bioproducts Ltd. DMF was purchased from Rathburn Chemicals and AGTC Bioproducts Ltd. Rink amide MBHA resin was purchased from Novabiochem. Other synthesis materials, including Aldrithiol-2, were purchased from Sigma-Aldrich. HPLC grade acetonitrile was purchased from Fisher Scientific.
Peptides were synthesized using SPPS using a CEM Liberty 1 peptide synthesizer under standard conditions. HPLC purification was carried out using linear water to acetonitrile gradients, with a PerkinElmer 785A detector and Series 200 LC pump, and a C18 column. MALDI-ToF experiments were carried out using a Bruker Daltonics Autoflex II ToF/ToF machine, and α-cyano-4-hydroxycinnamic acid matrix.
To form each disulfide-bonded peptide, one of the contributing peptides (dissolved in water) was reacted with a 10-fold excess of Aldrithiol-2 (in acetonitrile), and the products were separated by HPLC. The peptide product was then reacted with the other contributing peptide, and through displacement formed a disulfide-bonded peptide. The waste products of this reaction were then removed using HPLC.
Concentrations were determined using UV–vis absorption spectroscopy, and the tyrosine absorption centered around 276 nm: εTyr = 1280 cm–1 mol–1. With the exception of UV–vis spectra for the disulfide exchange experiments, which used a Thermo Scientific Nanodrop 1000, spectra were taken using a Unicam UV2 spectrometer.
CD spectroscopy was carried out using a JASCO J-1500 spectropolarimeter with mini-circulation bath and Peltier stage, using a 1 mm quartz cuvette. Samples were made at a concentration of 20 μM per peptide, in PBS buffer.
AUC sedimentation equilibrium data was collected at 20 °C in PBS buffer using a Beckman Optima XL-I analytical ultracentrifuge with an An-60 Ti rotor. The partial specific volume for each of the peptides/peptide mixtures, and the solvent density, were calculated using Sednterp.
DLS measurements were taken on a Malvern Zetasizer μV using Sarstedt disposable transparent cuvettes. Samples were made at a concentration of 100 μM per peptide, in PBS buffer.
TFA in water solutions (pH 2) were found to quench the disulfide exchange reaction, and hence this was used to both prevent rearrangement of the initial species prior to MALDI analysis, and to quench the system after rearrangement was encouraged in a second sample. Each of the disulfide-bonded peptides, in water solution, was added to both a solution of TFA in water (4 μL TFA in 15 mL water, pH 2) and to a PBS buffer solution containing reduced glutathione. After 10 min, TFA in water was added to the glutathione-containing solution to quench the reaction. These two solutions were then studied using MALDI-ToF mass spectrometry.
Acknowledgments
The authors thank EPSRC for funding L.S.R.S. through a Doctoral Studentship (EP/P505488/1) and Doctoral Prize (EP/M507854/1). A.R.T. and D.N.W. are supported by the ERC (340764), and D.N.W. is a Royal Society Wolfson Research Merit Award holder (WM140008). H.L. is supported by a Swedish Research Council Grant 2015-0612. Data supporting this article is available at http://dx.doi.org/10.15128/r2qr46r080t.
Supporting Information Available
The Supporting Information is available free of charge on the ACS Publications website at DOI: 10.1021/acssynbio.7b00037.
Additional circular dichroism data; dynamic light scattering data; AUC fitting information; MALDI data (PDF)
The authors declare no competing financial interest.
Supplementary Material
References
- Vale R. D. (2003) The Molecular Motor Toolbox for Intracellular Transport. Cell 112, 467–480. 10.1016/S0092-8674(03)00111-9. [DOI] [PubMed] [Google Scholar]
- Alberts B., Johnson A., Lewis J., Raff M., Roberts K., and Walter P. (2002) Molecular Biology of the Cell, 4 ed., Garland Science, New York. [Google Scholar]
- (2014) Hitching a ride with motor proteins. Nat. Nanotechnol. 9, 1–1. 10.1038/nnano.2013.308. [DOI] [PubMed] [Google Scholar]
- van den Heuvel M. G. L.; Dekker C. (2007) Motor proteins at work for nanotechnology. Science 317, 333–336. 10.1126/science.1139570. [DOI] [PubMed] [Google Scholar]
- Bromley E. H. C.; Channon K.; Moutevelis E.; Woolfson D. N. (2008) Peptide and protein building blocks for synthetic biology: From programming biomolecules to self-organized biomolecular systems. ACS Chem. Biol. 3, 38–50. 10.1021/cb700249v. [DOI] [PubMed] [Google Scholar]
- Channon K.; Bromley E. H. C.; Woolfson D. N. (2008) Synthetic biology through biomolecular design and engineering. Curr. Opin. Struct. Biol. 18, 491–498. 10.1016/j.sbi.2008.06.006. [DOI] [PubMed] [Google Scholar]
- Erbas-Cakmak S.; Leigh D. A.; McTernan C. T.; Nussbaumer A. L. (2015) Artificial Molecular Machines. Chem. Rev. 115, 10081–10206. 10.1021/acs.chemrev.5b00146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kassem S.; Lee A. T. L.; Leigh D. A.; Markevicius A.; Sola J. (2016) Pick-up, transport and release of a molecular cargo using a small-molecule robotic arm. Nat. Chem. 8, 138–143. 10.1038/nchem.2410. [DOI] [PubMed] [Google Scholar]
- Chen Y. J.; Groves B.; Muscat R. A.; Seelig G. (2015) DNA nanotechnology from the test tube to the cell. Nat. Nanotechnol. 10, 748–760. 10.1038/nnano.2015.195. [DOI] [PubMed] [Google Scholar]
- Pan J.; Li F. R.; Cha T. G.; Chen H. R.; Choi J. H. (2015) Recent progress on DNA based walkers. Curr. Opin. Biotechnol. 34, 56–64. 10.1016/j.copbio.2014.11.017. [DOI] [PubMed] [Google Scholar]
- Lund K.; Manzo A. J.; Dabby N.; Michelotti N.; Johnson-Buck A.; Nangreave J.; Taylor S.; Pei R. J.; Stojanovic M. N.; Walter N. G.; et al. (2010) Molecular robots guided by prescriptive landscapes. Nature 465, 206–210. 10.1038/nature09012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ikezoe Y.; Fang J.; Wasik T. L.; Shi M. L.; Uemura T.; Kitagawa S.; Matsui H. (2015) Peptide-Metal Organic Framework Swimmers that Direct the Motion toward Chemical Targets. Nano Lett. 15, 4019–4023. 10.1021/acs.nanolett.5b00969. [DOI] [PubMed] [Google Scholar]
- Ikezoe Y.; Washino G.; Uemura T.; Kitagawa S.; Matsui H. (2012) Autonomous motors of a metal-organic framework powered by reorganization of self-assembled peptides at interfaces. Nat. Mater. 11, 1081–1085. 10.1038/nmat3461. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nagatsugi F.; Takahashi Y.; Kobayashi M.; Kuwahara S.; Kusano S.; Chikuni T.; Hagihara S.; Harada N. (2013) Synthesis of peptide-conjugated light-driven molecular motors and evaluation of their DNA-binding properties. Mol. BioSyst. 9, 969–973. 10.1039/c2mb25520k. [DOI] [PubMed] [Google Scholar]
- Watson M. A.; Cockroft S. L. (2016) Man-made molecular machines: membrane bound. Chem. Soc. Rev. 45, 6118. 10.1039/C5CS00874C. [DOI] [PubMed] [Google Scholar]
- Bromley E. H. C.; Kuwada N. J.; Zuckermann M. J.; Donadini R.; Samii L.; Blab G. A.; Gemmen G. J.; Lopez B. J.; Curmi P. M. G.; Forde N. R.; et al. (2009) The Tumbleweed: towards a synthetic protein motor. HFSP J. 3, 204–212. 10.2976/1.3111282. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zuckermann M. J.; Angstmann C. N.; Schmitt R.; Blab G. A.; Bromley E. H. C.; Forde N. R.; Linke H.; Curmi P. M. G. (2015) Motor properties from persistence: a linear molecular walker lacking spatial and temporal asymmetry. New J. Phys. 17, 13. 10.1088/1367-2630/17/5/055017. [DOI] [Google Scholar]
- Kovacic S.; Samii L.; Woolfson D. N.; Curmi P. M. G.; Linke H.; Forde N. R.; Blab G. A. (2012) Design and Construction of a One- Dimensional DNA Track for an Artificial Molecular Motor. J. Nanomater. 2012, 10. 10.1155/2012/109238. [DOI] [Google Scholar]
- Niman C. S.; Beech J. P.; Tegenfeldt J. O.; Curmi P. M. G.; Woolfson D. N.; Forde N. R.; Linke H. (2013) Controlled microfluidic switching in arbitrary time-sequences with low drag. Lab Chip 13, 2389–2396. 10.1039/c3lc50194a. [DOI] [PubMed] [Google Scholar]
- Niman C. S.; Zuckermann M. J.; Balaz M.; Tegenfeldt J. O.; Curmi P. M. G.; Forde N. R.; Linke H. (2014) Fluidic switching in nanochannels for the control of Inchworm: a synthetic biomolecular motor with a power stroke. Nanoscale 6, 15008–15019. 10.1039/C4NR04701J. [DOI] [PubMed] [Google Scholar]
- Kuwada N. J.; Blab G. A.; Linke H. (2010) A classical Master equation approach to modeling an artificial protein motor. Chem. Phys. 375, 479–485. 10.1016/j.chemphys.2010.05.009. [DOI] [Google Scholar]
- Kuwada N. J.; Zuckermann M. J.; Bromley E. H. C.; Sessions R. B.; Curmi P. M. G.; Forde N. R.; Woolfson D. N.; Linke H. (2011) Tuning the performance of an artificial protein motor. Phys. Rev. E 84, 9. 10.1103/PhysRevE.84.031922. [DOI] [PubMed] [Google Scholar]
- Apostolovic B.; Danial M.; Klok H. A. (2010) Coiled coils: attractive protein folding motifs for the fabrication of self-assembled, responsive and bioactive materials. Chem. Soc. Rev. 39, 3541–3575. 10.1039/b914339b. [DOI] [PubMed] [Google Scholar]
- Newman J. R. S.; Keating A. E. (2003) Comprehensive identification of human bZIP interactions with coiled-coil arrays. Science 300, 2097–2101. 10.1126/science.1084648. [DOI] [PubMed] [Google Scholar]
- Kaplan J. B.; Reinke A. W.; Keating A. E. (2014) Increasing the affinity of selective bZIP-binding peptides through surface residue redesign. Protein Sci. 23, 940–953. 10.1002/pro.2477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vilanova C.; Tanner K.; Dorado-Morales P.; Villaescusa P.; Chugani D.; Frias A.; Segredo E.; Molero X.; Fritschi M.; Morales L.; Ramon D.; Pena C.; Pereto J.; Porcar M. (2015) Standards not that standard. J. Biol. Eng. 9, 4. 10.1186/s13036-015-0017-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kwok R. (2010) FIVE HARD TRUTHS FOR SYNTHETIC BIOLOGY. Nature 463, 288–290. 10.1038/463288a. [DOI] [PubMed] [Google Scholar]
- Mason J. M.; Arndt K. M. (2004) Coiled coil domains: Stability, specificity, and biological implications. ChemBioChem 5, 170–176. 10.1002/cbic.200300781. [DOI] [PubMed] [Google Scholar]
- Woolfson D. N. (2005) The design of coiled-coil structures and assemblies. Adv. Protein Chem. 70, 79. 10.1016/S0065-3233(05)70004-8. [DOI] [PubMed] [Google Scholar]
- Gradisar H.; Bozic S.; Doles T.; Vengust D.; Hafner-Bratkovic I.; Mertelj A.; Webb B.; Sali A.; Klavzar S.; Jerala R. (2013) Design of a single-chain polypeptide tetrahedron assembled from coiled-coil segments. Nat. Chem. Biol. 9, 362. 10.1038/nchembio.1248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raman S.; Machaidze G.; Lustig A.; Aebi U.; Burkhard P. (2006) Structure-based design of peptides that self-assemble into regular polyhedral nanoparticles. Nanomedicine 2, 95–102. 10.1016/j.nano.2006.04.007. [DOI] [PubMed] [Google Scholar]
- Burgess N. C.; Sharp T. H.; Thomas F.; Wood C. W.; Thomson A. R.; Zaccai N. R.; Brady R. L.; Serpell L. C.; Woolfson D. N. (2015) Modular Design of Self-Assembling Peptide-Based Nanotubes. J. Am. Chem. Soc. 137, 10554–10562. 10.1021/jacs.5b03973. [DOI] [PubMed] [Google Scholar]
- Fletcher J. M.; Boyle A. L.; Bruning M.; Bartlett G. J.; Vincent T. L.; Zaccai N. R.; Armstrong C. T.; Bromley E. H. C.; Booth P. J.; Brady R. L.; et al. (2012) A Basis Set of de Novo Coiled-Coil Peptide Oligomers for Rational Protein Design and Synthetic Biology. ACS Synth. Biol. 1, 240–250. 10.1021/sb300028q. [DOI] [PubMed] [Google Scholar]
- Fletcher J. M.; Harniman R. L.; Barnes F. R. H.; Boyle A. L.; Collins A.; Mantell J.; Sharp T. H.; Antognozzi M.; Booth P. J.; Linden N.; et al. (2013) Self-Assembling Cages from Coiled-Coil Peptide Modules. Science 340, 595–599. 10.1126/science.1233936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grigoryan G.; Keating A. E. (2006) Structure-based prediction of bZIP partnering specificity. J. Mol. Biol. 355, 1125–1142. 10.1016/j.jmb.2005.11.036. [DOI] [PubMed] [Google Scholar]
- Ramisch S.; Lizatovic R.; Andre I. (2015) Exploring alternate states and oligomerization preferences of coiled-coils by de novo structure modeling. Proteins: Struct., Funct., Genet. 83, 235–247. 10.1002/prot.24729. [DOI] [PubMed] [Google Scholar]
- Thomson A. R.; Wood C. W.; Burton A. J.; Bartlett G. J.; Sessions R. B.; Brady R. L.; Woolfson D. N. (2014) Computational design of water-soluble alpha-helical barrels. Science 346, 485–488. 10.1126/science.1257452. [DOI] [PubMed] [Google Scholar]
- Boyle A. L.; Bromley E. H. C.; Bartlett G. J.; Sessions R. B.; Sharp T. H.; Williams C. L.; Curmi P. M. G.; Forde N. R.; Linke H.; Woolfson D. N. (2012) Squaring the Circle in Peptide Assembly: From Fibers to Discrete Nanostructures by de Novo Design. J. Am. Chem. Soc. 134, 15457–15467. 10.1021/ja3053943. [DOI] [PubMed] [Google Scholar]
- Chen X. Y.; Zaro J. L.; Shen W. C. (2013) Fusion protein linkers: Property, design and functionality. Adv. Drug Delivery Rev. 65, 1357–1369. 10.1016/j.addr.2012.09.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bromley E. H. C.; Sessions R. B.; Thomson A. R.; Woolfson D. N. (2009) Designed alpha-Helical Tectons for Constructing Multicomponent Synthetic Biological Systems. J. Am. Chem. Soc. 131, 928–930. 10.1021/ja804231a. [DOI] [PubMed] [Google Scholar]
- Cerpa R.; Cohen F. E.; Kuntz I. D. (1996) Conformational switching in designed peptides: The helix/sheet transition. Folding Des. 1, 91–101. 10.1016/S1359-0278(96)00018-1. [DOI] [PubMed] [Google Scholar]
- Webber M. J.; Newcomb C. J.; Bitton R.; Stupp S. I. (2011) Switching of self-assembly in a peptide nanostructure with a specific enzyme. Soft Matter 7, 9665–9672. 10.1039/c1sm05610g. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zimenkov Y.; Dublin S. N.; Ni R.; Tu R. S.; Breedveld V.; Apkarian R. P.; Conticello V. P. (2006) Rational design of a reversible pH-responsive switch for peptide self-assembly. J. Am. Chem. Soc. 128, 6770–6771. 10.1021/ja0605974. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.