Abstract
Background
Liver metastasis is common in patients with colorectal cancer (CRC), and is correlated with poor outcome. MicroRNAs (miRNAs) are small non-coding RNAs involved in cancer development and progression, but their role in CRC liver metastasis has not been extensively investigated.
Results
Thirteen miRNAs were deregulated in pCRCs compared to their matched liver metastases. Seventeen miRNAs were chosen for validation, which confirmed significantly reduced expression of miR-99b-5p, miR-377 and miR-200c and increased expression of miR-196b-5p in the tissue of liver metastasis. Furthermore, miR-200c and miR-196b-5p were positively correlated with shorter overall survival in pCRC patients with liver metastasis.
Materials and Methods
Firstly, affymetrix microarrays involving 1036 miRNAs were performed in two pairs of primary CRCs (pCRCs) and their matched liver metastases. Secondly, validation of the results was carried out on an independent cohort of 48 pairs of pCRCs and matched liver metastases using quantitative real-time polymerase chain reaction assay.
Conclusions
We discovered a pCRC liver metastasis-specific miRNA panel including miR-377, miR-99b-5p, miR-200c and miR-196b-5p through intensive validation. These miRNAs may function as prognostic factors in patients with metastatic CRC.
Keywords: colorectal carcinoma, microRNA, liver metastasis
INTRODUCTION
Colorectal cancer (CRC) ranks as the third most prevalent cancer and the third leading cause of cancer mortality worldwide [1]. Liver is the most common site of metastasis. Even after resection of liver metastases, more than half the patients will not be cured due to relapse of liver metastases and emergence of other distant metastases [2, 3]. Therefore, it is critical to develop metastasis-specific molecular biomarkers for patients’ outcome prediction and treatment options.
MicroRNAs (miRNAs) are a class of endogenous, small (17–25 nucleotides), noncoding, single-stranded, evolutionarily conserved RNA molecules that regulate target gene expression by interfering with their transcription or by inhibiting translation. MiRNAs are known to be involved in the process of tumor cell growth, invasion and metastasis of CRC by regulating the expression of down-stream target genes [4–7]. Several studies reported that certain miRNAs were associated with CRC liver metastasis [4, 8–10]. For example, miR-181a and miR-200c promote liver metastases by inducing epithelial-mesenchymal transition in CRC [9, 10]. MiR-192 inhibits liver metastases by down-regulating the expression B-cell lymphoma 2, vascular endothelial growth factor A, and zinc finger E-box-binding homeobox 2 [4].
In our previous study, we found that miR-99b-5p was significantly more expressed in primary tumor than in matched liver metastasis. In vitro, miR-99b-5p was demonstrated to inhibit expression of mechanistic target of rapamycin (mTOR) by directly targeting its 3′ UTR. Moreover, miR-99b-5p was associated with a survival benefit in CRC patients with liver metastasis, suggesting that miR-99b-5p may serve as a predictive factor for metastasis and prognosis of CRC patients [8].
Herein, we report our metastasis-specific miRNA biomarker discovery approach which outlines the extensive miRNA difference between the primary tumor and paired liver metastasis specimens and validates newfound miRNA biomarkers in other independent cohorts of specimens. We also evaluated the correlation of the differential miRNAs expression profile with clinicopathological features and patient prognosis.
RESULTS
Differential miRNAs expression profile in primary tumros and matched liver metastasis tissues
We firstly performed miRNA microarray analysis in two pairs of matched pCRC and liver metastasis frozen specimens. The basic clinicopathological characteristics of these two patients were well matched. We identified 13 microRNAs whose expression levels differed between pCRC and liver metastasis tissues with a fold-change of > 1.5. Among these miRNAs, seven (miR-513a-5p, miR-181a-5p, miR-182-5p, miR-613, miR-152, miR-644a and miR-550a-5p) were up-regulated and six (miR-192-5p, miR-506-3p, miR-99b-5p, miR-29a-3p, miR-27b-3p and miR-934) were down-regulated in liver metastasis tissues compared with pCRC tissues.
For the validation step, qRT-PCR assay was performed in 48 isolated pairs of pCRC and liver metastasis tissues; the clinicopathological characteristics of these 48 patients are shown in Table 1. To extend the study, we added another four microRNAs (miR-196b-5p, miR-377, miR-200c and miR-10a-5p) with fold change between 1.3 and 1.5. As a result, four miRNAs (miR-99b-5p, miR-377, miR-200c and miR-196b-5p) were found to have significant differential expression between pCRC and liver metastasis tissues. MiR-99b-5p, miR-377 and miR-200c were down-regulated, while miR-196b-5p was up-regulated in liver metastasis tissues compared to pCRC (see Table 2, Figure 1 Supplementary Figure 1).
Table 1. The clinicopathological characteristics of 48 colorectal cancer patients with liver metastases.
| Characteristics | No. | Percent |
|---|---|---|
| Gender | ||
| Male | 29 | 60.4 |
| Female | 19 | 39.6 |
| Age, years | ||
| Median (SD) | 52 (28–74) | |
| < 60 | 34 | 70.8 |
| ≥ 60 | 14 | 29.2 |
| Primary tumor site | ||
| Rectum | 17 | 35.4 |
| Colon | 31 | 64.6 |
| pT | ||
| 2+3 | 9 | 18.7 |
| 4 | 39 | 81.3 |
| pN | ||
| 0 | 12 | 25.0 |
| 1 | 21 | 43.8 |
| 2 | 15 | 31.2 |
| Vascular invasion | ||
| Negative | 27 | 62.8 |
| Positive | 16 | 37.2 |
| NK | 5 | |
| Perineural invasion | ||
| Negative | 27 | 62.8 |
| Positive | 16 | 37.2 |
| NK | 5 | |
| Differentiation | ||
| Poor/poor-moderate | 12 | 25.0 |
| moderate/moderate-high | 36 | 75.0 |
| Time of occurrence of liver metastasis | ||
| Synchronous | 43 | 89.6 |
| Metachronous | 5 | 10.4 |
| Combined with other metastases | ||
| No | 41 | 85.4 |
| Yes | 7 | 14.6 |
Abbreviations: NK, not known; NR, not reached; SD, standard deviation.
Table 2. Expression level of miRNAs in the primary tumor and matched liver metastases of 48 colorectal cancer patients with liver metastases.
| miRNA | Tumor status | Median | SEM | Foldchange (primary/ metastasis) | p value |
|---|---|---|---|---|---|
| miR-513a-5p | Primary | 28.9670 | 7.87524 | 0.54 | 0.113 |
| Metastasis | 53.6298 | 15.04555 | |||
| miR-181a-5p | Primary | 2.2210 | 1.11954 | 0.51 | 0.188 |
| Metastasis | 4.3494 | 0.99962 | |||
| miR-182-5p | Primary | 19.8616 | 4.40714 | 1.48 | 0.253 |
| Metastasis | 13.3865 | 4.96276 | |||
| miR-613 | Primary | 64.2181 | 41.80958 | 1.20 | 0.877 |
| Metastasis | 53.7233 | 52.13662 | |||
| miR-152 | Primary | 31.6605 | 18.05150 | 9.32 | 0.114 |
| Metastasis | 3.3972 | 0.93292 | |||
| miR-644a | Primary | 18.0385 | 9.14903 | 0.26 | 0.313 |
| Metastasis | 69.2121 | 54.83062 | |||
| miR--550a-5p | Primary | 179.4224 | 87.74360 | 0.51 | 0.291 |
| Metastasis | 354.2439 | 219.88422 | |||
| miR-196b-5p | Primary | 8.8372 | 4.05013 | 0.18 | 0.046* |
| Metastasis | 49.5967 | 20.74880 | |||
| miR-10a-5p | Primary | 5.4569 | 2.19123 | 0.16 | 0.062 |
| Metastasis | 33.9821 | 16.66163 | |||
| miR-192-5p | Primary | 4304.9628 | 1690.66343 | 4.77 | 0.051 |
| Metastasis | 903.1332 | 275.60286 | |||
| miR-506-3p | Primary | 11.2938 | 5.02323 | 1.07 | 0.908 |
| Metastasis | 10.5453 | 6.00231 | |||
| miR-99b-5p | Primary | 8.8244 | 2.75573 | 6.77 | 0.007* |
| Metastasis | 1.3043 | 0.29901 | |||
| miR-29a-3p- | Primary | 1.3543 | 0.90819 | 4.44 | 0.169 |
| Metastasis | 0.3049 | 0.18774 | |||
| miR-27b-3p | Primary | 2.6432 | 0.61204 | 0.07 | 0.170 |
| Metastasis | 39.7780 | 26.64356 | |||
| miR-377 | Primary | 92.8067 | 40.17213 | 433.47 | 0.026* |
| Metastasis | 0.2141 | 0.13345 | |||
| miR-200c | Primary | 13.4106 | 3.51479 | 3.77 | 0.009* |
| Metastasis | 3.5561 | 0.95874 | |||
| miR-934 | Primary | 15.6664 | 11.61170 | 1.78 | 0.515 |
| Metastasis | 8.8113 | 2.47346 |
Abbreviations: miRNA, microRNA; SEM, standard error of the mean. *p ≤ 0.05.
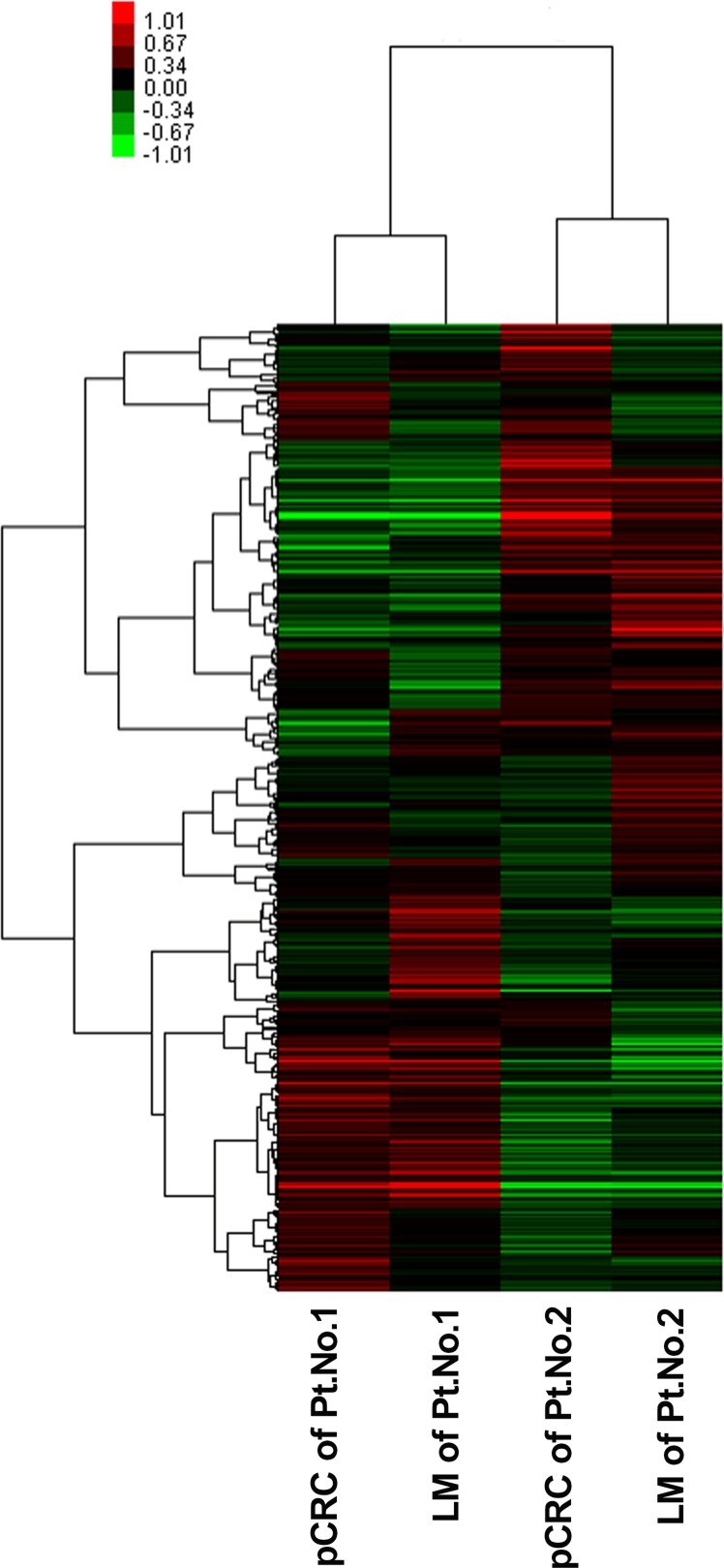
Figure 1. Unsupervised hierarchical cluster analysis of the 528 differentially expressed miRNAs.
Glue Grant Human Transcriptome Array (Affymetrix, Santa Clara, CA, USA) screened the differential expression of miRNA between primary tumor and liver metastases in two pairs of fresh tissue. Green and red represent the down-regulated and up-regulated miRNAs, respectively. LM, liver metastasis; miRNA, microRNA; pCRC, primary CRC; Pt, patient.
MiR-377 and miR-99b-5p expression levels serve as invasive biomarkers in pCRC patients with liver metastasis
To explore the clinical significance of these liver metastasis-specific miRNAs, we further validated the correlation between microRNAs and clinicopathological characteristics. Three up-regulated miRNAs (miR-99b-5p, miR-377 and miR-200c) and one down-regulated miRNA (miR-196b-5p) in pCRC compared to liver metastasis tissues were analyzed. As shown in Table 3, increased miR-377 expression level was associated with advanced lymph node stage (p = 0.043). Likewise, decreased miR-99b-5p expression was positively correlated with perineural invasion in pCRC (p = 0.043).
Table 3. Relationship between the expression level of miRNAs in the primary tumor and clinicopathologic parameters in 48 colorectal cancer patients with liver metastases.
| No. | Percent | miR-377 | miR-200c | miR-99b-5p | miR-196b-5p | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Characteristics | Mean | SEM | p value | Mean | SEM | p value | Mean | SEM | p value | Mean | SEM | p value | ||
| Gender | ||||||||||||||
| Male | 29 | 60.4 | 133.69 | 64.193 | 0.138 | 9.99 | 2.805 | 0.233 | 9.64 | 3.998 | 0.719 | 12.22 | 6.537 | 0.307 |
| Female | 19 | 39.6 | 30.41 | 22.445 | 18.64 | 7.765 | 7.58 | 3.466 | 3.68 | 2.070 | ||||
| Age, years | ||||||||||||||
| Median (SD) | 52 (28–74) | |||||||||||||
| < 60 | 34 | 70.8 | 41.79 | 19.491 | 0.194 | 13.36 | 4.735 | 0.983 | 8.28 | 3.613 | 0.762 | 3.07 | 1.098 | 0.16 |
| ≥ 60 | 14 | 29.2 | 216.69 | 126.472 | 13.53 | 3.864 | 10.15 | 3.668 | 22.83 | 13.215 | ||||
| Primary tumor site | ||||||||||||||
| Rectum | 17 | 35.4 | 41.92 | 25.288 | 0.354 | 19.59 | 9.081 | 0.319 | 15.34 | 6.992 | 0.178 | 4.85 | 2.225 | 0.472 |
| Colon | 31 | 64.6 | 120.71 | 60.452 | 10.02 | 2.183 | 5.25 | 1.709 | 11.02 | 6.156 | ||||
| pT | ||||||||||||||
| 2+3 | 9 | 18.7 | 64.61 | 32.559 | 0.74 | 9.70 | 2.394 | 0.617 | 3.86 | 1.710 | 0.392 | 19.13 | 15.913 | 0.457 |
| 4 | 39 | 81.3 | 99.31 | 48.982 | 14.27 | 4.292 | 9.97 | 3.352 | 6.46 | 3.446 | ||||
| pN | ||||||||||||||
| Negative | 12 | 25.0 | 9.23 | 3.215 | 0.043* | 15.88 | 5.763 | 0.69 | 12.06 | 6.051 | 0.503 | 13.89 | 12.014 | 0.477 |
| Positive | 36 | 75.0 | 120.67 | 52.912 | 12.59 | 4.308 | 7.74 | 3.103 | 7.15 | 3.732 | ||||
| Vascular invasion | ||||||||||||||
| Negative | 27 | 62.8 | 85.06 | 62.303 | 0.673 | 12.37 | 3.394 | 0.525 | 12.91 | 4.688 | 0.088 | 4.17 | 1.549 | 0.216 |
| Positive | 16 | 37.2 | 124.78 | 59.972 | 17.56 | 8.888 | 4.17 | 1.638 | 19.34 | 11.657 | ||||
| NK | 5 | |||||||||||||
| Perineural invasion | ||||||||||||||
| Negative | 27 | 62.8 | 35.52 | 18.413 | 0.151 | 13.30 | 3.371 | 0.742 | 13.45 | 4.710 | 0.043* | 9.61 | 5.442 | 0.953 |
| Positive | 16 | 37.2 | 208.38 | 113.022 | 15.99 | 8.959 | 3.25 | 0.961 | 10.17 | 8.117 | ||||
| NK | 5 | |||||||||||||
| Differentiation | ||||||||||||||
| Poor/poor-moderate | 12 | 25.0 | 124.21 | 79.322 | 0.657 | 7.67 | 2.146 | 0.352 | 3.61 | 1.083 | 0.074 | 12.81 | 10.703 | 0.577 |
| Moderate/moderate-high | 36 | 75.0 | 82.34 | 47.093 | 15.32 | 4.606 | 10.56 | 3.623 | 7.51 | 4.143 | ||||
| Time of occurrence of liver metastasis | ||||||||||||||
| Synchronous | 43 | 89.6 | 99.78 | 44.722 | 0.616 | 10.84 | 2.265 | 0.432 | 9.08 | 3.048 | 0.786 | 9.77 | 4.505 | 0.507 |
| Metachronous | 5 | 10.4 | 32.86 | 20.699 | 35.48 | 28.185 | 6.60 | 4.063 | 0.85 | 0.300 | ||||
| Combined with other metastases | ||||||||||||||
| No | 41 | 85.4 | 88.66 | 43.422 | 0.806 | 11.12 | 2.401 | 0.466 | 9.65 | 3.205 | 0.475 | 7.08 | 3.650 | 0.543 |
| Yes | 7 | 14.6 | 117.11 | 113.770 | 26.84 | 20.083 | 4.00 | 1.473 | 19.15 | 18.443 | ||||
Abbreviations: NK, not known; SD, standard deviation; SEM, standard error of the mean.
MiR-200c and miR-196-5p expression levels serve as prognostic biomarkers in pCRC patients with liver metastasis
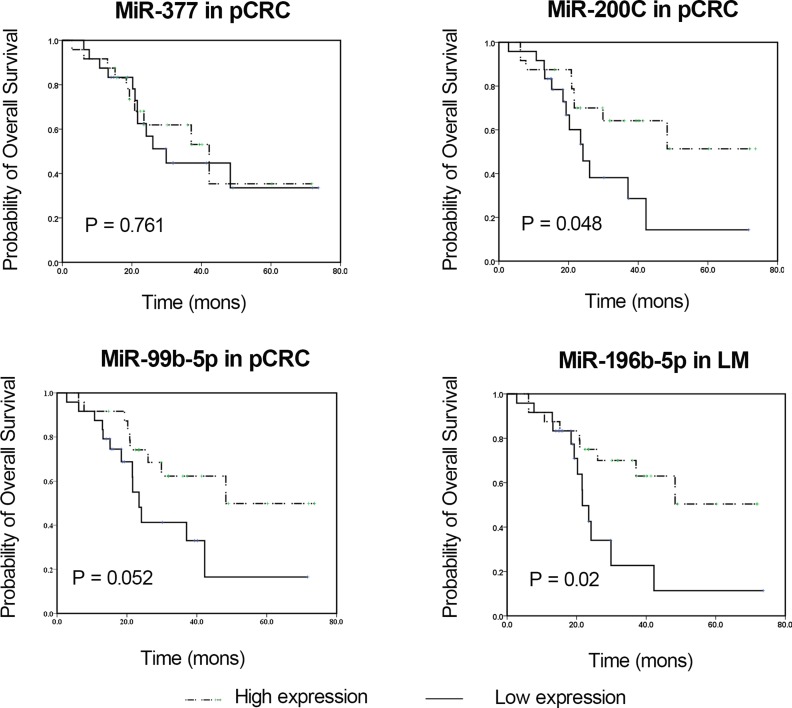
We tested the association between miRNA expression and overall survival by performing survival analysis. As shown in Figure 2 and Table 4, patients with elevated expression levels of miR-200c in pCRC and miR-196b-5p in liver metastases had significantly longer survival by Kaplan-Meier analysis (p = 0.048 and p = 0.02, respectively). Notably, higher expression of miR-99b-5p in pCRC also showed a better trend of survival, although statistical significance was not reached (p = 0.052).
Figure 2. Kaplan-Meier survival curves of CRC patients subdivided by miRNAs levels in pCRC and LM tissues from CRC patients.
Overall survival analysis based on miR-377, miR-200c, miR-99b-5p and miR-196b-5p in pCRC or LM tissues are shown. The p value was determined by log-rank test. LM, liver metastasis; miRNA, microRNA; pCRC, primary CRC.
Table 4. Relationship between the expression level of miRNAs and overall survival in 48 colorectal cancer patients with liver metastases.
| 95% CI | 95% CI | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Primary tumor | Median OS | SEM | Lower bound | Upper bound | Liver metastases | Median OS | SEM | Lower bound | Upper bound | ||
| miR-377 | Low | 29.867 | 5.509 | 19.069 | 40.664 | miR-377 | Low | 24.133 | 13.966 | 0 | 51.506 |
| High | 42.233 | 11.776 | 19.152 | 65.314 | High | 37.067 | NR | NR | NR | ||
| p value | 0.761 | p value | 0.417 | ||||||||
| miR-200c | Low | 24.133 | 3.290 | 17.684 | 30.582 | miR-200c | Low | 26.033 | 5.702 | 14.858 | 37.209 |
| High | NR | NR | NR | NR | High | 48.3 | 12.092 | 24.599 | 72.001 | ||
| p value | 0.048* | p value | 0.266 | ||||||||
| miR-99b-5p | Low | 23.467 | 2.212 | 19.131 | 27.802 | miR-99b-5p | Low | 23.467 | 1.594 | 20.343 | 26.59 |
| High | 48.300 | NR | NR | NR | High | 48.3 | 12.096 | 24.592 | 72.008 | ||
| p value | 0.052 | p value | 0.098 | ||||||||
| miR-196b-5p | Low | 23.467 | 1.71 | 20.115 | 26.819 | miR-196b-5p | Low | 21.633 | 1.686 | 18.329 | 24.937 |
| High | 42.233 | NR | NR | NR | High | NR | NR | NR | NR | ||
| p value | 0.116 | p value | 0.02* | ||||||||
Abbreviations: CI: confidence interval; miRNA, microRNA; NR: not reached; OS, overall survival; SEM, standard error of the mean. *p ≤ 0.05.
DISCUSSION
Patients with liver metastasis in CRC were with a poor prognosis and early recurrence. Previous researches in the fields of molecular and genetic studies have contributed to our understanding of CRC liver metastasis in the cellular level [11, 12]. However, heterogeneity of research methods as well as tumor tissue acquisition limited the diagnostic and prognostic role of specific genes in metastases.
Recent findings have highlighted the role of miRNAs in the tumorigenesis of CRC cells acting as oncogenes or tumor suppressor genes [13, 14]. Similar studies have been performed to gain better insight into genomic changes involved in pCRC initiation and progression [15]. However, there are few studies of miRNA expression accounting for distant metastasis of pCRC. In this study, we aim to identify a differential microRNA expression profile in pCRC and their matched liver metastasis.
In this study, expression profiling of 1036 miRNAs based on an RNA array platform in clinical samples of pCRC tissues with matched liver metastasis tissues was carried out to identify advanced CRC signatures and to use miRNA expression profiles to classifying CRC with liver metastasis. The confirmation of the array data was done by using qRT-PCR. The results, by microarray analysis in two paired colorectal cancers, showed 17 miRNAs were dysregulated in liver metastasis tissues compared with pCRC tissues. These differentially expressed miRNAs were further validated by qRT-PCR in 48 isolated pairs of pCRC and liver metastasis tissues, and four miRNAs (miR-99b-5p, miR-377, miR-200c and miR-196b-5p) were found to have significant differential expression between pCRC tissues and liver metastasis tissues. While examining the potential clinical impact of these four liver metastasis-specific miRNAs, we noticed that up-regulation of miR-200c in pCRC tissues and miR-196b-5p in liver metastasis tissues were associated with significantly longer survival by Kaplan-Meier analysis (p = 0.048 and p = 0.02, respectively).
We previously demonstrated that miR-99b-5p is a tumor suppressing miRNA in metastatic CRC, and that its suppressive effects are mediated chiefly by suppressing mTOR expression [8]. MiR-196b was reported to be highly expressed in CRC tissues and was correlated with rapid progression and poor prognosis [16, 17]. MiR-196b was also a predictive factor in palliative chemotherapy and neoadjuvant radiochemotherapy [18, 19]. Similar to our findings, high expression of miR-200c was observed in liver metastasis [10] and its expression in serum was reported to be a prognostic factor in patients with CRC [20]. It has been found that miR-200c could mediate epithelial-mesenchymal transition in liver metastases by regulating the TGF-TGFegu signal pathway [21, 22]. MiR-377 was a novel biomarker for poor prognosis in gastrointestinal cancer, which induced tumorigenesis by targeting multiple tumor-suppressor genes [23].
Our study is distinct from previous studies in several aspects. First, we directly compared pCRC and matching liver metastasis tissues from the same patients, which avoid heterogeneity due to distinct miRNA from different patients. Second, to improve the predictive value of the miRNA markers, we developed a miRNA-based classifier panel rather than just suggesting each miRNA as a predictor.
In conclusion, four miRNAs (miR-99b-5p, miR-377, miR-200c and miR-196b-5p) were found to be differently expressed between primary tumors of patients with CRC and their matched liver metastases. Moreover, we nominate the candidate microRNAs, miR-200 and miR-196b-5p as potential prognostic markers. Our study suggested that these miRNAs could hopefully serve as metastasis-specific miRNA biomarkers for the management of patients with CRC.
MATERIALS AND METHODS
Patient samples
The fresh-frozen/formalin-fixed, paraffin-embedded primary CRC and matched corresponding liver metastatic tissues were collected from 2004 to 2012 in the Department of Colorectal Cancer, Fudan University Shanghai Cancer Center. The tissue bank of the hospital stored the specimens. The study was approved by the Ethics Committee of Fudan University Shanghai Cancer Center and written informed consent was obtained from all patients.
RNA extraction and miRNA array hybridization
Total RNAs from two CRC patients with liver metastasis were analyzed by microarray. Total RNA was isolated using Trizol (Invitrogen, Carlsbad, CA, USA). RNA was labeled and hybridized to Glue Grant Human Transcriptome Array (Affymetrix, Santa Clara, CA, USA) [24–26] according to manufacturer's instructions. Command Console® Software version 3.1 was used for microarray analysis. A total of 1036 miRNAs were covered using miRBase version 18.0.
Quantitative real-time polymerase chain reaction
miR-513a-5p, miR-181a-5p, miR-182-5p, miR-613, miR-152, miR-644a, miR-550a-5p, miR-192-5p, miR-506-3p, miR-99b-5p, miR-29a-3p, miR-27b-3p, miR-934, miR-196b-5p, miR-200c, miR-377, miR-10a-5p and RNU6B (internal control) specific cDNA syntheses were carried out using the miScript Reverse Transcription Kit (Qiagen, Hilden, Germany) according to the manufacturer's protocols. The expression level of miRNAs above was measured using TaqMan miRNA assays (Applied Biosystems, Carlsbad, CA, USA). U6 was used as an endogenous control and the fold change was calculated using the 2−ΔΔCt method.
Statistical analysis
The data were analyzed with SPSS version 18.0. The student t-test and one factor analysis of variance were used for analysis as appropriate. Kaplan-Meier method and log-rank test were used to estimate the survival. P < 0.05 was considered statistically significant.
SUPPLEMENTARY MATERIALS FIGURES
Acknowledgments
We thank Dr. Menghong Sun (Chief of the Tissue Bank of Fudan University Shanghai Cancer Center) for tumor tissues acquisition and Mary Smith (Ph.D) for language editing.
Abbreviations
- CRC
colorectal cancer
- miRNA
microRNA
- mTOR
mechanistic target of rapamycin
- pCRC
primary colorectal cancer
- qRT-PCR
quantitative real-time polymerase chain reaction
Footnotes
CONFLICTS OF INTEREST
The authors declare that they have no competing interests.
REFERENCES
- 1.Siegel R, Ma J, Zou Z, Jemal A. Cancer statistics, 2014. CA Cancer J Clin. 2014;64:9–29. doi: 10.3322/caac.21208. [DOI] [PubMed] [Google Scholar]
- 2.Adam R, Haller DG, Poston G, Raoul JL, Spano JP, Tabernero J, Van Cutsem E. Toward optimized front-line therapeutic strategies in patients with metastatic colorectal cancer--an expert review from the International Congress on Anti-Cancer Treatment (ICACT) 2009. Ann Oncol. 2010;21:1579–84. doi: 10.1093/annonc/mdq043. [DOI] [PubMed] [Google Scholar]
- 3.Adam R, De Gramont A, Figueras J, Guthrie A, Kokudo N, Kunstlinger F, Loyer E, Poston G, Rougier P, Rubbia-Brandt L, Sobrero A, Tabernero J, Teh C, et al. The oncosurgery approach to managing liver metastases from colorectal cancer: a multidisciplinary international consensus. Oncologist. 2012;17:1225–39. doi: 10.1634/theoncologist.2012-0121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Geng L, Chaudhuri A, Talmon G, Wisecarver JL, Are C, Brattain M, Wang J. MicroRNA-192 suppresses liver metastasis of colon cancer. Oncogene. 2014;33:5332–40. doi: 10.1038/onc.2013.478. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Ma Y, Zhang P, Wang F, Zhang H, Yang Y, Shi C, Xia Y, Peng J, Liu W, Yang Z, Qin H. Elevated oncofoetal miR-17-5p expression regulates colorectal cancer progression by repressing its target gene P130. Nat Commun. 2012;3:1291. doi: 10.1038/ncomms2276. [DOI] [PubMed] [Google Scholar]
- 6.Yuan K, Xie K, Fox J, Zeng H, Gao H, Huang C, Wu M. Decreased levels of miR-224 and the passenger strand of miR-221 increase MBD2, suppressing maspin and promoting colorectal tumor growth and metastasis in mice. Gastroenterology. 2013;145:853–64. doi: 10.1053/j.gastro.2013.06.008. e9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Zhang L, Pickard K, Jenei V, Bullock MD, Bruce A, Mitter R, Kelly G, Paraskeva C, Strefford J, Primrose J, Thomas GJ, Packham G, Mirnezami AH. miR-153 supports colorectal cancer progression via pleiotropic effects that enhance invasion and chemotherapeutic resistance. Cancer Res. 2013;73:6435–47. doi: 10.1158/0008-5472.CAN-12-3308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Li W, Chang J, Wang S, Liu X, Peng J, Huang D, Sun M, Chen Z, Zhang W, Guo W, Li J. miRNA-99b-5p suppresses liver metastasis of colorectal cancer by down-regulating mTOR. Oncotarget. 2015;6:24448–62. doi: 10.18632/oncotarget.4423. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Ji D, Chen Z, Li M, Zhan T, Yao Y, Zhang Z, Xi J, Yan L, Gu J. MicroRNA-181a promotes tumor growth and liver metastasis in colorectal cancer by targeting the tumor suppressor WIF-1. Mol Cancer. 2014;13:86. doi: 10.1186/1476-4598-13-86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Hur K, Toiyama Y, Takahashi M, Balaguer F, Nagasaka T, Koike J, Hemmi H, Koi M, Boland CR, Goel A. MicroRNA-200c modulates epithelial-to-mesenchymal transition (EMT) in human colorectal cancer metastasis. Gut. 2013;62:1315–26. doi: 10.1136/gutjnl-2011-301846. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Shanghai Cancer Report 2007. Shanghai: SCDC; 2007. Shanghai Municipal Center for Disease Control and Prevention. [Google Scholar]
- 12.Simmonds PC, Primrose JN, Colquitt JL, Garden OJ, Poston GJ, Rees M. Surgical resection of hepatic metastases from colorectal cancer: a systematic review of published studies. Br J Cancer. 2006;94:982–99. doi: 10.1038/sj.bjc.6603033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Asangani IA, Rasheed SA, Nikolova DA, Leupold JH, Colburn NH, Post S, Allgayer H. MicroRNA-21 (miR-21) post-transcriptionally downregulates tumor suppressor Pdcd4 and stimulates invasion, intravasation and metastasis in colorectal cancer. Oncogene. 2008;27:2128–36. doi: 10.1038/sj.onc.1210856. [DOI] [PubMed] [Google Scholar]
- 14.Valeri N, Croce CM, Fabbri M. Pathogenetic and clinical relevance of microRNAs in colorectal cancer. Cancer Genomics Proteomics. 2009;6:195–204. doi. [PubMed] [Google Scholar]
- 15.Kanaan Z, Rai SN, Eichenberger MR, Barnes C, Dworkin AM, Weller C, Cohen E, Roberts H, Keskey B, Petras RE, Crawford NP, Galandiuk S. Differential microRNA expression tracks neoplastic progression in inflammatory bowel disease-associated colorectal cancer. Hum Mutat. 2012;33:551–60. doi: 10.1002/humu.22021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Wang YX, Zhang XY, Zhang BF, Yang CQ, Chen XM, Gao HJ. Initial study of microRNA expression profiles of colonic cancer without lymph node metastasis. J Dig Dis. 2010;11:50–4. doi: 10.1111/j.1751-2980.2009.00413.x. [DOI] [PubMed] [Google Scholar]
- 17.Ge J, Chen Z, Li R, Lu T, Xiao G. Upregulation of microRNA-196a and microRNA-196b cooperatively correlate with aggressive progression and unfavorable prognosis in patients with colorectal cancer. Cancer Cell Int. 2014;14:128. doi: 10.1186/s12935-014-0128-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Boisen MK, Dehlendorff C, Linnemann D, Nielsen BS, Larsen JS, Osterlind K, Nielsen SE, Tarpgaard LS, Qvortrup C, Pfeiffer P, Hollander NH, Keldsen N, Hansen TF, et al. Tissue microRNAs as predictors of outcome in patients with metastatic colorectal cancer treated with first line Capecitabine and Oxaliplatin with or without Bevacizumab. PLoS One. 2014;9:e109430. doi: 10.1371/journal.pone.0109430. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Svoboda M, Sana J, Fabian P, Kocakova I, Gombosova J, Nekvindova J, Radova L, Vyzula R, Slaby O. MicroRNA expression profile associated with response to neoadjuvant chemoradiotherapy in locally advanced rectal cancer patients. Radiat Oncol. 2012;7:195. doi: 10.1186/1748-717X-7-195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Toiyama Y, Hur K, Tanaka K, Inoue Y, Kusunoki M, Boland CR, Goel A. Serum miR-200c is a novel prognostic and metastasis-predictive biomarker in patients with colorectal cancer. Ann Surg. 2014;259:735–43. doi: 10.1097/SLA.0b013e3182a6909d. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Shen A, Lin W, Chen Y, Liu L, Chen H, Zhuang Q, Lin J, Sferra TJ, Peng J. Pien Tze Huang inhibits metastasis of human colorectal carcinoma cells via modulation of TGF-beta1/ZEB/miR-200 signaling network. Int J Oncol. 2015;46:685–90. doi: 10.3892/ijo.2014.2772. [DOI] [PubMed] [Google Scholar]
- 22.Tian Y, Pan Q, Shang Y, Zhu R, Ye J, Liu Y, Zhong X, Li S, He Y, Chen L, Zhao J, Chen W, Peng Z, et al. MicroRNA-200 (miR-200) cluster regulation by achaete scute-like 2 (Ascl2): impact on the epithelial-mesenchymal transition in colon cancer cells. J Biol Chem. 2014;289:36101–15. doi: 10.1074/jbc.M114.598383. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Wen X, Wu JQ, Peng W, Feng JF, Tang JH. MicroRNA-377 predicts poor clinical outcome of gastric cancer and induces tumorigenesis by targeting multiple tumor-suppressor genes. Oncol Rep. 2015;34:203–10. doi: 10.3892/or.2015.3981. [DOI] [PubMed] [Google Scholar]
- 24.Xu W, Seok J, Mindrinos MN, Schweitzer AC, Jiang H, Wilhelmy J, Clark TA, Kapur K, Xing Y, Faham M, Storey JD, Moldawer LL, Maier RV, et al. Human transcriptome array for high-throughput clinical studies. Proc Natl Acad Sci USA. 2011;108:3707–12. doi: 10.1073/pnas.1019753108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Qiu JJ, Ye LC, Ding JX, Feng WW, Jin HY, Zhang Y, Li Q, Hua KQ. Expression and clinical significance of estrogen-regulated long non-coding RNAs in estrogen receptor alpha-positive ovarian cancer progression. Oncol Rep. 2014;31:1613–22. doi: 10.3892/or.2014.3000. [DOI] [PubMed] [Google Scholar]
- 26.Sun NX, Ye C, Zhao Q, Zhang Q, Xu C, Wang SB, Jin ZJ, Sun SH, Wang F, Li W. Long noncoding RNA-EBIC promotes tumor cell invasion by binding to EZH2 and repressing E-cadherin in cervical cancer. PLoS One. 2014;9:e100340. doi: 10.1371/journal.pone.0100340. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.