Abstract
The prokaryote Myxococcus xanthus is a model for cell interactions important in multicellular behavior. We used the transposon TnphoA to specifically identify genes for cell-surface factors involved in cell interactions. From a library of 10,700 insertions of TnphoA, we isolated 36 that produced alkaline phosphatase activity. Three TnphoA insertions tagged cell motility genes, called cgl, which control the adventurous movement of cells. The products of the tagged cgl genes could function in trans upon other cells and were localized primarily in the cell envelope and extracellular space, consistent with TnphoA tagging genes for extracellular factors controlling motility.
Full text
PDF
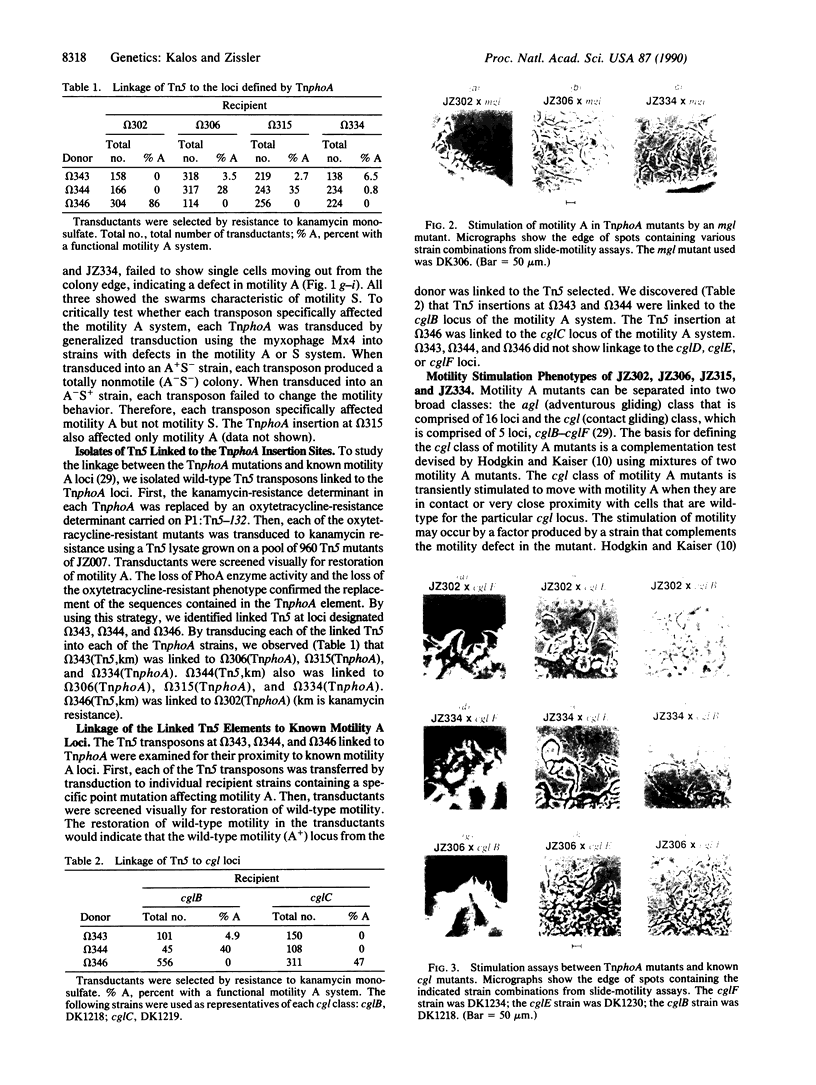
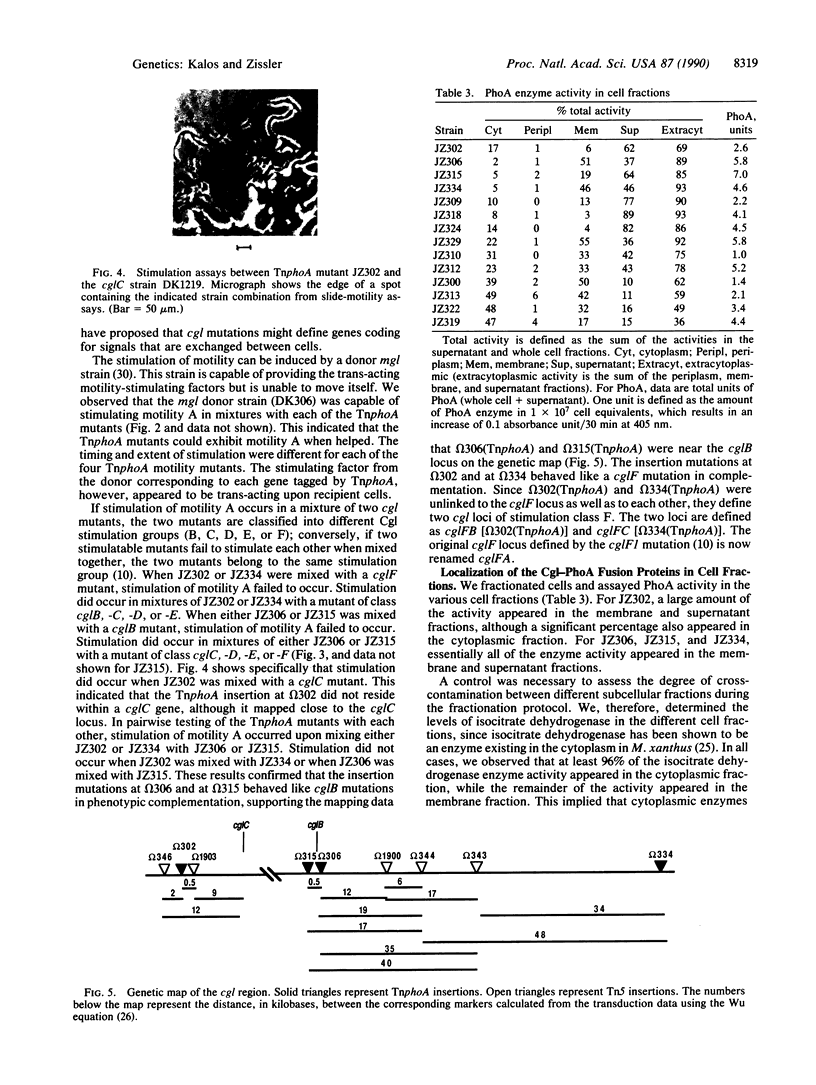




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Avery L., Kaiser D. In situ transposon replacement and isolation of a spontaneous tandem genetic duplication. Mol Gen Genet. 1983;191(1):99–109. doi: 10.1007/BF00330896. [DOI] [PubMed] [Google Scholar]
- Boquet P. L., Manoil C., Beckwith J. Use of TnphoA to detect genes for exported proteins in Escherichia coli: identification of the plasmid-encoded gene for a periplasmic acid phosphatase. J Bacteriol. 1987 Apr;169(4):1663–1669. doi: 10.1128/jb.169.4.1663-1669.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boyd D., Manoil C., Beckwith J. Determinants of membrane protein topology. Proc Natl Acad Sci U S A. 1987 Dec;84(23):8525–8529. doi: 10.1073/pnas.84.23.8525. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Breton A. M., Jaoua S., Guespin-Michel J. Transfer of plasmid RP4 to Myxococcus xanthus and evidence for its integration into the chromosome. J Bacteriol. 1985 Feb;161(2):523–528. doi: 10.1128/jb.161.2.523-528.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burchard R. P. Gliding motility of prokaryotes: ultrastructure, physiology, and genetics. Annu Rev Microbiol. 1981;35:497–529. doi: 10.1146/annurev.mi.35.100181.002433. [DOI] [PubMed] [Google Scholar]
- DWORKIN M. Nutritional requirements for vegetative growth of Myxococcus xanthus. J Bacteriol. 1962 Aug;84:250–257. doi: 10.1128/jb.84.2.250-257.1962. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Geisselsoder J., Campos J. M., Zusman D. R. Physical characterization of bacteriophage MX4, a generalized transducing phage for Myxococcus xanthus. J Mol Biol. 1978 Feb 25;119(2):179–189. doi: 10.1016/0022-2836(78)90432-1. [DOI] [PubMed] [Google Scholar]
- Gutierrez C., Barondess J., Manoil C., Beckwith J. The use of transposon TnphoA to detect genes for cell envelope proteins subject to a common regulatory stimulus. Analysis of osmotically regulated genes in Escherichia coli. J Mol Biol. 1987 May 20;195(2):289–297. doi: 10.1016/0022-2836(87)90650-4. [DOI] [PubMed] [Google Scholar]
- Hodgkin J., Kaiser D. Cell-to-cell stimulation of movement in nonmotile mutants of Myxococcus. Proc Natl Acad Sci U S A. 1977 Jul;74(7):2938–2942. doi: 10.1073/pnas.74.7.2938. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoffman C. S., Wright A. Fusions of secreted proteins to alkaline phosphatase: an approach for studying protein secretion. Proc Natl Acad Sci U S A. 1985 Aug;82(15):5107–5111. doi: 10.1073/pnas.82.15.5107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Manoil C., Beckwith J. A genetic approach to analyzing membrane protein topology. Science. 1986 Sep 26;233(4771):1403–1408. doi: 10.1126/science.3529391. [DOI] [PubMed] [Google Scholar]
- Manoil C., Beckwith J. TnphoA: a transposon probe for protein export signals. Proc Natl Acad Sci U S A. 1985 Dec;82(23):8129–8133. doi: 10.1073/pnas.82.23.8129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin S., Sodergren E., Masuda T., Kaiser D. Systematic isolation of transducing phages for Myxococcus xanthus. Virology. 1978 Jul 1;88(1):44–53. doi: 10.1016/0042-6822(78)90108-3. [DOI] [PubMed] [Google Scholar]
- Michaelis S., Inouye H., Oliver D., Beckwith J. Mutations that alter the signal sequence of alkaline phosphatase in Escherichia coli. J Bacteriol. 1983 Apr;154(1):366–374. doi: 10.1128/jb.154.1.366-374.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nelson D. R., Cumsky M. G., Zusman D. R. Localization of myxobacterial hemagglutinin in the periplasmic space and on the cell surface of Myxococcus xanthus during developmental aggregation. J Biol Chem. 1981 Dec 10;256(23):12589–12595. [PubMed] [Google Scholar]
- Orndorff P., Stellwag E., Starich T., Dworkin M., Zissler J. Genetic and physical characterization of lysogeny by bacteriophage MX8 in Myxococcus xanthus. J Bacteriol. 1983 May;154(2):772–779. doi: 10.1128/jb.154.2.772-779.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rigby P. W., Dieckmann M., Rhodes C., Berg P. Labeling deoxyribonucleic acid to high specific activity in vitro by nick translation with DNA polymerase I. J Mol Biol. 1977 Jun 15;113(1):237–251. doi: 10.1016/0022-2836(77)90052-3. [DOI] [PubMed] [Google Scholar]
- Shimkets L. J. Control of morphogenesis in myxobacteria. Crit Rev Microbiol. 1987;14(3):195–227. doi: 10.3109/10408418709104439. [DOI] [PubMed] [Google Scholar]
- Sodergren E., Cheng Y., Avery L., Kaiser D. Recombination in the Vicinity of Insertions of Transposon Tn 5 in MYXOCOCCUS XANTHUS. Genetics. 1983 Oct;105(2):281–291. doi: 10.1093/genetics/105.2.281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sodergren E., Kaiser D. Insertions of Tn5 near genes that govern stimulatable cell motility in Myxococcus. J Mol Biol. 1983 Jun 25;167(2):295–310. doi: 10.1016/s0022-2836(83)80337-4. [DOI] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Starich T., Zissler J. Movement of multiple DNA units between Myxococcus xanthus cells. J Bacteriol. 1989 May;171(5):2323–2336. doi: 10.1128/jb.171.5.2323-2336.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taylor R. K., Manoil C., Mekalanos J. J. Broad-host-range vectors for delivery of TnphoA: use in genetic analysis of secreted virulence determinants of Vibrio cholerae. J Bacteriol. 1989 Apr;171(4):1870–1878. doi: 10.1128/jb.171.4.1870-1878.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taylor R. K., Miller V. L., Furlong D. B., Mekalanos J. J. Use of phoA gene fusions to identify a pilus colonization factor coordinately regulated with cholera toxin. Proc Natl Acad Sci U S A. 1987 May;84(9):2833–2837. doi: 10.1073/pnas.84.9.2833. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Watson B. F., Dworkin M. Comparative intermediary metabolism of vegetative cells and microcysts of Myxococcus xanthus. J Bacteriol. 1968 Nov;96(5):1465–1473. doi: 10.1128/jb.96.5.1465-1473.1968. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu T. T. A model for three-point analysis of random general transduction. Genetics. 1966 Aug;54(2):405–410. doi: 10.1093/genetics/54.2.405. [DOI] [PMC free article] [PubMed] [Google Scholar]