ABSTRACT
The phospholipid (PL) composition of bacterial membranes varies as a function of growth rate and in response to changes in the environment. While growth adaptation can be explained by biochemical feedback in the PL synthesis pathway, recent transcriptome studies have revealed that the expression of PL synthesis genes can also be tuned in response to various stresses. We previously showed that the BasRS two-component pathway controls the expression of the diacylglycerol kinase gene, dgkA, in Escherichia coli (A. Wahl, L. My, R. Dumoulin, J. N. Sturgis, and E. Bouveret, Mol Microbiol, 80:1260–1275, 2011, https://doi.org/10.1111/j.1365-2958.2011.07641.x). In this study, we set up a strategy to identify the mutation responsible for the upregulation of pssA observed in the historical pssR1 mutant and supposedly corresponding to a transcriptional repressor (C. P. Sparrow and J. Raetz, J Biol Chem, 258:9963–9967, 1983). pssA encodes phosphatidylserine synthase, the first step of phosphatidylethanolamine synthesis. We showed that this mutation corresponded to a single nucleotide change in the anti-Shine-Dalgarno sequence of the 16S rRNA encoded by the rrnC operon. We further demonstrated that this mutation enhanced the translation of pssA. Though this effect appeared to be restricted to PssA among phospholipid synthesis enzymes, it was not specific, as evidenced by a global effect on the production of unrelated proteins.
IMPORTANCE Bacteria adjust the phospholipid composition of their membranes to the changing environment. In addition to enzymatic regulation, stress response regulators control specific steps of the phospholipid synthesis pathway. We wanted to identify a potential regulator controlling the expression of the phosphatidylserine synthase gene. We showed that it was not the previously suggested hdfR gene and instead that a mutation in the anti-Shine-Dalgarno sequence of 16S RNA was responsible for an increase in pssA translation. This example underlines the fact that gene expression can be modulated by means other than specific regulatory processes.
KEYWORDS: Escherichia coli, hdfR, maoP, phosphatidylserine synthase, phospholipid synthesis, pssA, pssR, ribosomal mutations, rrnC, yifE
INTRODUCTION
Phospholipids, which are the building blocks of bacterial membranes, are synthesized by a series of enzymes localized in the inner membrane. The biochemistry of the phospholipid synthesis pathway has been well deciphered, especially in the Escherichia coli model bacterium (1). Phospholipid composition is very tightly controlled and is modulated by various parameters, such as growth conditions, growth rate, and stresses. Most of this control is thought to occur at the enzymatic activity level (1). However, transcriptomic data have recently uncovered the possibility that several signaling pathways control phospholipid synthesis through a regulation of enzyme expression levels in E. coli. Indeed, the expression of all the genes for phospholipid synthesis seems to correlate with the growth rate through ppGpp and stringent response control (2, 3). Furthermore, different stress response pathways control specific steps of phospholipid synthesis. For example, the σS alternative sigma factor activates the expression of cfa, encoding cyclopropane synthase (4, 5). Membranes are indeed enriched in cyclopropanated phospholipids in stationary phase. Another example is the activation of the expression of plsB by σE during envelope stress (6, 7).
Interestingly, some time ago, Raetz and collaborators performed extensive mutagenesis screens to isolate genes involved in phospholipid synthesis (8). Among them, they were able to isolate potential regulators of specific steps of the pathway. Notably, one mutant showed increased expression of dgkA (8, 9). We were able to trace this to a point mutation in the BasS sensor protein and showed further that dgkA is part of the BasRS regulon (7). Following this first identification, we then decided to focus on a second regulator that was reported by Sparrow and Raetz to control the expression of pssA (10). In that study, a mutant exhibiting a specific increase in PssA expression was isolated. The mutation was clearly functioning in trans, and the authors hypothesized that there was a mutation in a putative PssR repressor of pssA expression. The position of the mutation was precisely mapped by three-point analysis of random general transduction at 84 min on the chromosome, just upstream of the ilv locus (10). Yet, the pssR gene was never cloned or identified afterwards. Following genome sequencing and annotation, because of this initial physical mapping of pssR, an open reading frame (ORF) was annotated as pssR. This first annotation was later the object of corrections, and the yifA and pssR neighbor ORFs were fused into one single hdfR ORF (11, 12). However, no effect of the hdfR mutant on phospholipids was observed (11).
The goal of our study was to identify pssR and elucidate the regulation of pssA expression. Instead of finding a specific transcriptional regulator, we show that a single mutation in the anti-Shine-Dalgarno (anti-SD) sequence of 16S rRNA encoded by the rrnC operon is responsible for the large amount of PssA enzyme observed in the original mutant strain. This effect, while very strong on PssA, is not specific, since the expression of several unrelated genes is also affected.
RESULTS
Increase of pssA expression in the AC5 pssR1 mutant strain.
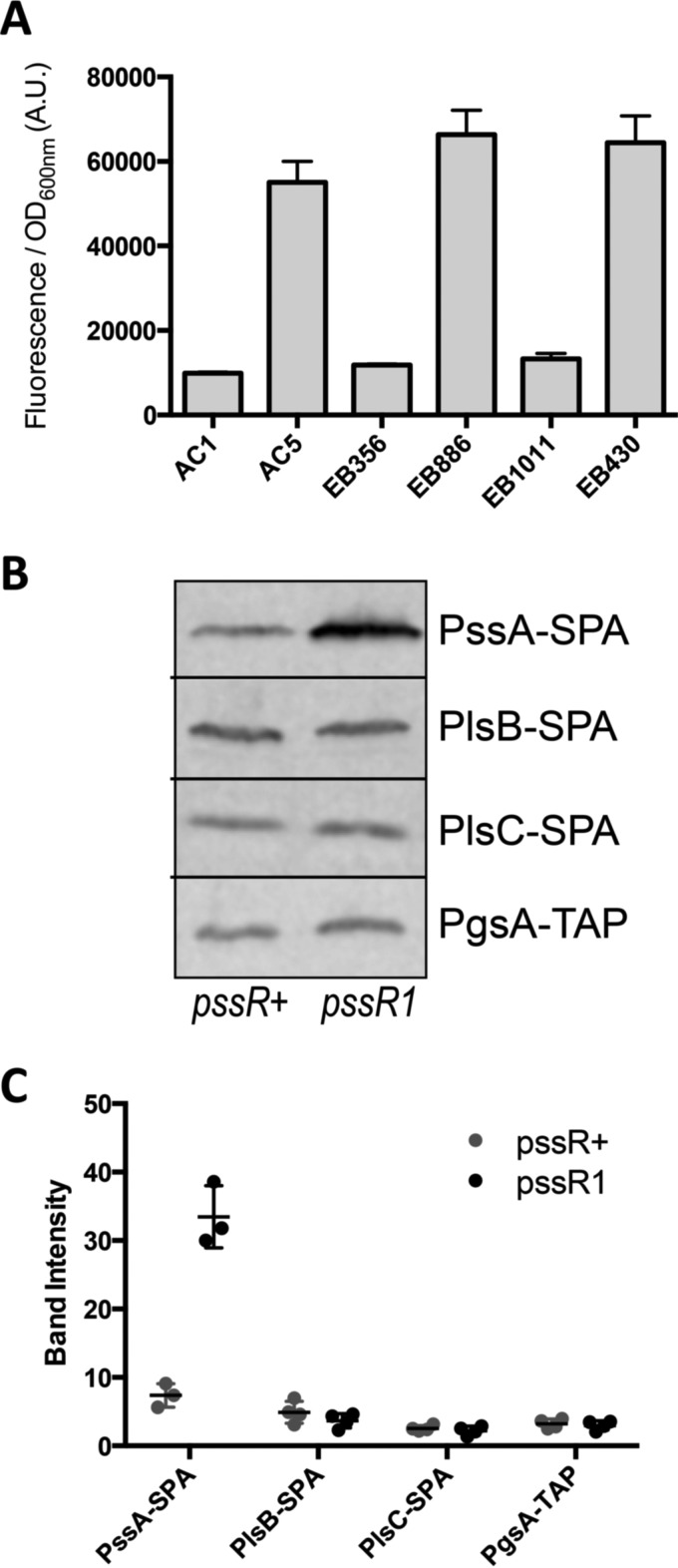
Using the AC1 (pssR+) and AC5 (pssR1) strains provided by Raetz's laboratory (10), we first wanted to reproduce the upregulation of pssA reported in the original paper. To this end, we used a translational fusion of the entire promoter and ORF of the pssA gene with green fluorescent protein (GFP) in a plasmid. We observed a 6-fold overexpression of pssA in the AC5 strain compared with that in AC1 (Fig. 1A). Then, we also wanted to verify that the same effect was obtained for the amounts of the PssA protein expressed from its natural locus on the chromosome. We constructed recombinant strains in which the endogenous PssA was tagged at its C terminus with the sequential peptide affinity (SPA) tag while still being expressed from its natural promoter. The amount of PssA-SPA protein as revealed by Western blotting was increased 4-fold in the AC5 (pssR1) genetic background compared with that in the parental AC1 background (Fig. 1B and C).
FIG 1.
Upregulation of pssA in the pssR1 mutant. (A) Strains AC1 (pssR+), AC5 (pssR1), AC1 ΔhdfR::Chlr (EB356), AC5 ΔhdfR::Chlr (EB886), MG ΔhdfR pssR+ (EB1011), and MG ΔhdfR pssR1 (EB430) (Table 2) were transformed with the pssA-GFP translational fusion and the corresponding control plasmids (pEB1853 and pEB1900), and expression of pssA-GFP was measured as described in Materials and Methods. Data are the means from four replicates and the error bars show the standard deviations. (B) AC1 and AC5 strains producing tagged phospholipid enzymes, EB160 and EB161 (PssA-SPA), EB1052 and EB1053 (PlsB-SPA), EB1050 and EB1051 (PlsC-SPA), and EB1054 and EB1055 (PgsA-TAP) (Table 2), were grown in LB at 37°C and stopped in exponential growth phase. Whole-cell extracts were separated by 10% SDS-PAGE and the SPA-tagged or TAP-tagged proteins were analyzed by Western blotting using anti-Flag or PAP antibodies, respectively. (C) The experiment shown in panel B was repeated with three or four replicates for each strain, and the Western blots were quantified on a Li-Cor imager. The error bars show the standard deviations.
To test if the pssR1 mutation was indeed linked to the ilv genetic locus identified in the original study and to assess if the upregulation effect was independent of the specific genetic background of the strains, we transduced the pssR1 mutation from the AC5 strain to the wild-type MG1655 reference strain. For this, we first introduced a chloramphenicol resistance cassette to replace the hdfR ORF through direct recombination, both in the AC1 and AC5 strains. The hdfR gene is located close to the ilv locus, at approximately 10,000 bp. The cassette insertion did not interfere with the upregulation of pssA in the AC5 genetic background (Fig. 1A, compare EB356/EB886 with AC1/AC5). The ΔhdfR::cat marker was then transduced from the two strains into MG1655. pssA expression was found to be upregulated in the EB430 strain obtained from the AC5 transduction compared with that in the EB1011 strain obtained from the AC1 transduction (Fig. 1A). This confirmed that pssR1 was genetically linked to hdfR but suggested that pssR1 was distinct from hdfR (see below). Furthermore, this showed that the effect of the mutation on pssA expression was independent of the particular genetic background of the strain used.
Sparrow and Raetz originally suggested that the observed regulation was specific to pssA because the enzymatic activities of 6 other phospholipid (PL) synthesis enzymes were not modified (the activities tested at that time corresponded to PlsB, CdsA, PgsA, phosphatidylglycerophosphate [PGP], DgkA, and Psd enzymes) (10). To test directly that the production of PL synthesis enzymes other than PssA was not affected, we followed the same strategy used to monitor PssA protein amounts, building recombinant strains expressing PlsB-SPA, PlsC-SPA, or PgsA-TAP chimeric proteins (13). In contrast to that of PssA, and in accordance with the observations of Sparrow and Raetz, the amounts of these three proteins were not modified by the presence of the pssR1 mutation (Fig. 1B and C).
Identification of the mutated gene causing pssA overexpression.
To identify the location of the pssR1 mutation, we performed genome resequencing of AC1 and AC5 strains. Relative to the MG1655 reference genome, approximately 200 point mutations were found throughout the AC1 and AC5 genomes, yet all these mutations were identical in AC1 and AC5. By contrast, 19 mutations were found only in AC5, all restricted to a region between positions 3923000 and 3996000 of the MG1655 reference genome (Table 1). This region is compatible with the window of approximately 100 kb corresponding to the P1 transduction used to isolate the pssR locus (10). This also highlights the high mutagenesis rate that was used in the original paper. Our whole-genome sequencing results thus appeared perfectly consistent with the conclusions obtained from the initial genetic experiments.
TABLE 1.
Genome sequence differences between AC1 and AC5 strains identified by genome resequencing
| Positiona | Nucleotide in: |
Amino acid changeb | Geneb | |
|---|---|---|---|---|
| AC1 | AC5 | |||
| 3923644 | C | T | G13S | rsmG |
| 3926991 | G | A | L5 | asnC |
| 3934295 | C | T | T173I | rbsA |
| 3935773 | C | T | G162 | rbsC |
| 3943345 | C | T | NA | rrsC |
| 3945437 | C | T | NA | rrlC |
| 3945582 | C | T | NA | rrlC |
| 3946315 | C | T | NA | rrlC |
| 3947259 | C | T | G237S | hdfR |
| 3979126 | C | T | R176C | wzyE |
| yifK | ||||
| 3982939 | C | T | NA | Intergenic |
| 3985435 | C | T | M191I | aslA |
| 3988482 | C | T | S195 | hemX |
| 3990406 | C | T | A121T | hemC |
| 3993390 | C | T | F746 | cyaA |
| 3993903 | A | G | C53Rc | cyaY |
| 3994950 | C | T | F63 | dapF |
| 4035734 | T | G | NA | rrsA |
| 4038695 | T | C | NA | rrlA |
Positions of the mutations on the E. coli K-12 MG1655 reference genome as given in the Ecocyc database (21).
Missense mutations and the corresponding genes are highlighted in bold. The yifK gene used for mapping pssR1 is also indicated. NA, not applicable.
In the MG1655 reference strain, residue 53 is arginine.
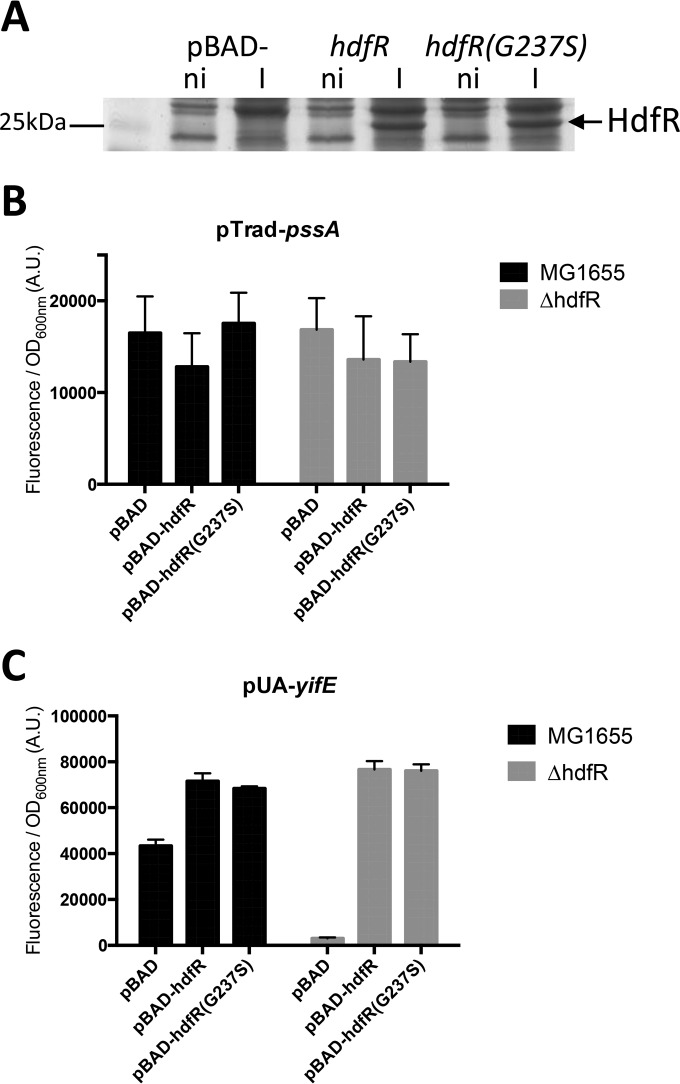
The next step was then to identify among the 19 mutations the one(s) responsible for pssA upregulation. The position of the G237S mutation found in the hdfR ORF in AC5 (Table 1) was in very good agreement with the genetic mapping of pssR (i.e., 84 min, near the ilv genes). At some point, hdfR was even annotated as potential pssR (12). This, besides the fact that it encodes a transcriptional regulator of the LysR family, made HdfR a very good candidate for the PssR repressor that we were looking for. However, when we measured the expression of pssA in a ΔhdfR strain compared with that in a wild-type strain, no effect was observed (Fig. 2B, compare the pBAD bars). This was expected, as the deletion of hdfR did not suppress the difference of pssA expression in AC1 and AC5 strains (Fig. 1A). To test further if the G237S mutation in HdfR had a specific effect, we cloned the wild-type and mutated versions of hdfR in the pBAD24 expression vector. Wild-type and mutated HdfR proteins were produced in similar amounts upon induction with arabinose (Fig. 2A). We did not observe any significant effect on pssA expression with either wild-type HdfR or HdfR(G237S) mutant overproduction (Fig. 2B). To test if the G237S mutation in HdfR had any effect on HdfR function, we first tried to use a transcriptional fusion of gltB, a gene whose expression was reported to be regulated by HdfR (14), but we did not observe any signal for this fusion under any of the conditions tested (data not shown). Transcriptional regulators of the LysR family often regulate a divergently transcribed neighboring gene (15). Therefore, we tested the expression of yifE, localized next to hdfR in an opposite orientation. YifE has been recently renamed MaoP, as it was shown to be involved in macrodomain organization of the Ori region (16). Interestingly, both hdfR and yifE have been identified in a genetic screen for genes promoting the stress-induced mutagenesis response in E. coli (17). We observed that HdfR strongly activates yifE expression (Fig. 2C). Indeed, yifE expression was abolished in a ΔhdfR mutant, while it was upregulated when HdfR was overproduced (Fig. 2C). However, it was clear that the G237S mutation of HdfR had no effect on yifE expression (Fig. 2C). In summary, the G237S mutation in HdfR does not seem to influence HdfR function. Furthermore, overproduction or deletion of hdfR had no effect on pssA expression, demonstrating that hdfR is not pssR.
FIG 2.
hdfR is not pssR. (A) Wild-type MG1655 was transformed by the pBAD24, pBAD-hdfR, or pBAD-hdfR(G237S) plasmid. Cells were grown in LB at 37°C and expression was induced with 0.5% arabinose for 2 h. Whole-cell extracts were separated by 10% SDS-PAGE, and proteins were stained with Coomassie blue. Strains MG1655 and EB814 (ΔhdfR) were transformed simultaneously with the fusion pTrad-pssA (pEB1611) (B) or pUA-yifE (C) and the pBAD-hdfR or pBAD-hdfR(G237S) plasmid (Table 3). Cultures were performed in LB supplemented with ampicillin, kanamycin, and 0.5% arabinose, and expression was monitored as described in Materials and Methods. Data are the means from 4 replicates and the error bars show the standard deviations.
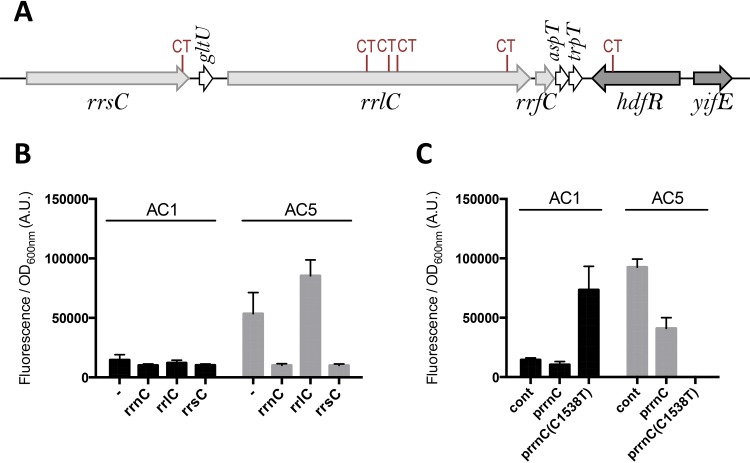
Using a similar approach to the one detailed above for HdfR (overproduction of wild-type or mutant versions of the proteins, or deletion of the genes), we also ruled out any link between rsmG(G13S) or rbsA(T173I) mutation (Table 1) and pssR (data not shown). Finally, we used an MG1655 strain containing a ΔyifK::Kanr resistance cassette localized between wzyE and aslA at approximately 30 kb from hdfR (Table 1) to transduce back into AC5 wild-type regions of the chromosome together with the ΔyifK::Kanr marker. The transductants were then screened by measuring the expression of pssA and by sequencing the genes originally mutated in AC5. Using this strategy, we were able to obtain a strain (EB1068) with all the genes from hdfR to rrlA that were mutated in AC5 (Table 1) restored to a wild-type sequence. This strain still displayed an overproduction of PssA (data not shown). Therefore, we finally investigated the possibility that the only mutations left for study, the ones in the rrnC ribosomal operon localized just upstream of hdfR on the chromosome (Table 1; Fig. 3A), are responsible for the overproduction of PssA. We deleted the entire rrnC operon or only the rrsC or rrlC genes by direct recombination in the AC1 and AC5 strains. When rrlC (encoding 23S RNA) was deleted alone, leaving the mutation in rrsC, the overproduction of PssA was still observed in the AC5 background compared with expression in the AC1 background (Fig. 3B). However, when the entire operon or only the rrsC gene was deleted, overproduction of PssA was no longer observed in the AC5 background (Fig. 3B). This left the single mutation C1538T at the extremity of rrsC as the only one that could be responsible for PssA overproduction. This mutation is located at the 3′ extremity of 16S RNA, in the anti-SD sequence (Fig. 3A).
FIG 3.
The rrsC(C1538T) mutation is responsible for the increase in pssA expression in AC5. (A) rrnC operon. The mutations present in AC5 but not in AC1 are indicated in red. (B) AC1 and AC5 strains with portions of the rrnC operon deleted (strains EB350, EB349, EB1062, EB352, EB351, and EB1063) (Table 2) were transformed with the pssA-GFP translational fusion (plasmid pEB1853), and expression was monitored as described in Materials and Methods. Data are the means from 4 replicates and the error bars show the standard deviations. (C) Strains AC1 and AC5 were transformed with the pssA-GFP reporter fusion (plasmid pEB1853) and the indicated plasmids [cont, pEB0354; prrnC, pEB1858; prrnC(C1538T), pEB1866]. Expression was monitored as described in Materials and Methods. Data are the means from 4 replicates and the error bars show the standard deviations.
To finally prove that the mutation C1538T in rrsC was by itself responsible for the upregulation of pssA, we transformed AC1 and AC5 strains with plasmids bearing the entire rrnC operon, with or without the C1538T mutation in rrsC. The wild-type rrnC plasmid strongly reduced the upregulation of pssA in AC5 (Fig. 3C), whereas the mutated rrnC plasmid provoked the upregulation of pssA in AC1 (Fig. 3C). It was impossible to obtain clones from the transformation of AC5 with the mutated rrnC plasmid, suggesting that this mutation C1538T in the anti-SD sequence had deleterious effects, as already reported (18).
The C1538T mutation in AC5 increases pssA translation level.
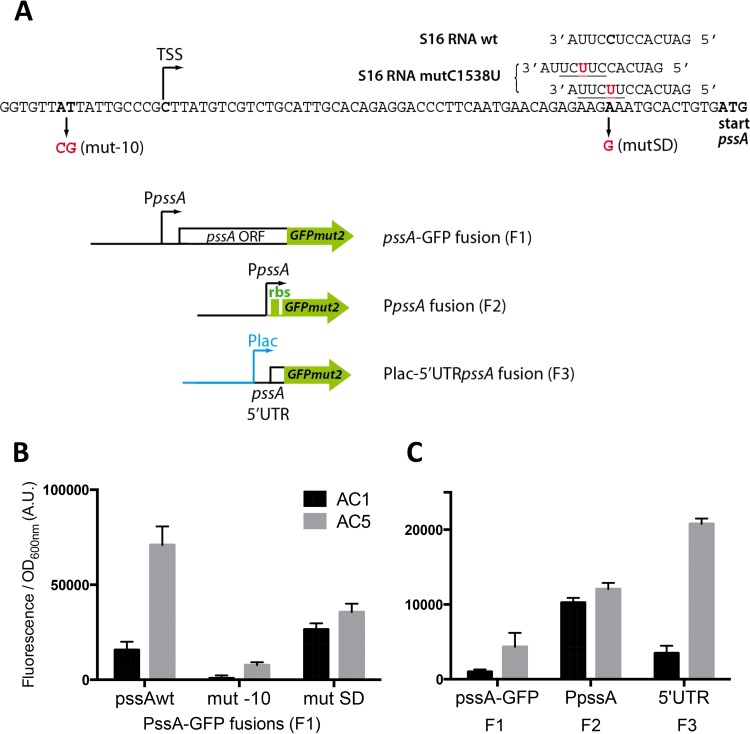
Given the nature of the mutation responsible for the overexpression of pssA, we suspected that the regulation happened at the initiation of translation. To test this, we dissected the promoter and the 5′ untranslated region (5′ UTR) of pssA to produce different transcriptional or translational fusions with GFP (Fig. 4A). The transcriptional start site of pssA has been mapped 59 nucleotides before its ATG start codon in a global transcriptome sequencing (RNA-seq) study (19). To verify this position, we first mutated the corresponding −10 promoter sequence in the pssA-GFP translational fusion (F1) (Fig. 4A). The mutation abolished the expression of pssA (Fig. 4B), confirming the +1 position, which we then used as the limit for constructing strict transcriptional and translational fusions (F2 and F3, respectively) (Fig. 4A). The expression levels of GFP under the control of the promoter of pssA (PpssA) and an artificial mRNA leader sequence were not very different in AC1 and AC5 strains (Fig. 4C, F2 bars). On the contrary, the expression of GFP under the control of the Plac promoter and the 5′ UTR of pssA was increased 6-fold in AC5 compared with that in AC1 (Fig. 4C, F3 bars), which is very similar to the 4- to 5-fold increase observed with the global pssA-GFP fusion (F1) (Fig. 4B and C). This demonstrated that the effect of the pssR1 mutant on pssA expression was posttranscriptional. We reasoned that the C1538U mutation in the anti-SD 16S RNA sequence might enable better recognition of the SD sequence of pssA. Indeed, the SD sequence of pssA is not optimal, but the C1538U mutation improves the base pairing at 2 putative positions on pssA: AGAAG and AAGAA (Fig. 4A). We tried to restore a better base pairing with the wild-type 16S RNA by introducing a mutation at the second position in the pssA SD sequence (Fig. 4A). The mutation abolished overexpression of pssA in AC5 (Fig. 4B). However, though designed as a compensatory mutation, it did not trigger in AC1 expression as strong as that of wild-type pssA in AC5. Another possibility is that this mutation simply increases by 1 nucleotide the base pairing of the upstream position.
FIG 4.
The rrsC(C1538T) mutation affects pssA translation initiation. (A) The 5′ UTR sequence of pssA and wild-type (wt) or mutated (mutC1538U) anti-SD sequences of 16S RNA are shown. Mutations are indicated in red. TSS, transcription start site. The different constructions of GFP fusions with portions of the pssA gene are shown below. (B and C) Strains AC1 and AC5 were transformed with the indicated reporter fusions (plasmids pEB1853, pEB1864, and pEB1878 for panel B and plasmids pEB1611, pEB1856, and pEB1895 for panel C). Expression was monitored as described in Materials and Methods. Data are the means from 4 replicates and the error bars show the standard deviations.
Together, these results demonstrated that the initiation of pssA translation is increased in AC5, presumably due to enhanced base pairing of the SD sequence of pssA with the C1538U mutated anti-SD sequence of 16S RNA.
Differential proteome analysis of the AC1 and AC5 strains.
Since we showed that the pssR1 genotype could be reduced to the C1538T mutation in rrsC, it was difficult to imagine that the effect was restricted to pssA expression. Indeed, the C1538T mutation mapped to the extremity of 16S RNA in the anti-SD sequence and was thus expected to affect the translation of a larger number of genes. It has to be noted that this mutation was studied before, shown to be deleterious, and also described to affect the expression of several genes (18). We wanted to obtain a global view of the genes whose expression was impacted by the mutation. To this end, we performed a quantitative proteomic analysis of the AC1 and AC5 strains to identify differentially expressed proteins. This permitted us to identify and quantify 1,318 proteins (see Table S1 in the supplemental material). A stringent statistical analysis revealed 85 proteins whose expression levels were modified at least 2-fold between the compared strains (highlighted in green and blue in Table S1). Thus, 43 proteins were found to be more expressed in the AC5 strain, whereas 42 were found to be enriched in the AC1 strain. Among the differentially expressed proteins, PssA was found to be overexpressed more than 6 times [log2(fold change) = 2.75] in AC5 compared with that in AC1, which technically validated the experiment.
To validate these data, we choose in this table several proteins (highlighted in bold in Table S1) for which a strain was available in a collection of E. coli SPA-tagged strains (13). The holC, aspA, smpB, ybbA, and ygeV SPA-tagged constructions were transduced in the AC1 and AC5 strains, and the amounts of the proteins were quantified by Western blotting using a monoclonal antibody directed against the SPA tag. We confirmed the increase of HolC, YgeV, YbbA, and AspA protein expression in the AC5 strain compared with that in the AC1 strain while, in contrast to the observation of the proteomics analysis, SmpB-SPA levels were found to be decreased in the AC5 strain (see Fig. S1).
Therefore, while PssA was apparently the only PL synthesis enzyme upregulated in the pssR1 mutant (10; our results), the mutation in the anti-SD sequence of 16S RNA encoded by the rrnC operon affected the translation of several other genes.
DISCUSSION
Global effect of the C1538T mutation in 16S RNA on gene expression and physiology.
The C1538T mutation has been reported to be strongly toxic, as the mutated rRNA operon could not be expressed from a plasmid even in a wild-type strain (18). By contrast, we were able to transform the wild-type AC1 with the mutated plasmid (using a plasmid similar to the one used in reference 18) (Fig. 3C). We did observe a slight growth phenotype for the AC5 strain in which the mutated rrnC operon was expressed from its natural chromosomal locus. Additionally, it was indeed impossible to transform further the AC5 strain using a plasmid bearing the same mutated rRNA operon, demonstrating the deleterious effect of the mutation. In the same study, a series of proteins whose production was affected by the mutation were identified by comparative proteomic two-dimensional (2D) gel analysis (18). From the 8 proteins reported to be affected by the mutation, we found only methionyl-tRNA synthetase (MetG) to be significantly enriched in our proteomic analyses of the AC1 and AC5 strains (see Table S1 in the supplemental material). This discrepancy might be explained by the different levels of expression of the mutated rrnC operon (from the inducible plasmid in reference 18 and from the chromosomal locus in our study) but also by the low accuracy of the method used previously. We are confident in our proteomics analysis, given that PssA was identified and that 4 proteins randomly chosen in the set were confirmed to be similarly affected in a different experiment. We obtained a contradictory result only for the SmpB protein. However, given the role of SmpB in trans translation (20), a possible explanation is that the SPA tag perturbed the function of the SmpB protein, which in return would affect broadly the translation of many proteins, including itself.
In reference 18, the authors hypothesized that the amount of protein produced reflected the change in complementarity between the mutated rRNA and the SD sequence in the mRNA. When we looked at the proteins we identified by proteomics analysis, apart from PssA (see below), it was difficult to pinpoint specific features of the SD sequences for the overproduced or reduced proteins. In some cases, it was possible to correlate upregulation or downregulation with a better or worse base pairing with the mutated 16S RNA, as described in reference 18. However, the expression of many genes might be indirectly affected by the increased or decreased translation of others. Furthermore, because we performed the analysis in AC1 and AC5 strains, it is possible that the additional mutations present in AC5 (Table 1) also affect the expression of some genes.
PssA translation is affected by the C1538T mutation in 16S RNA.
In this study, we showed that a mutation in the anti-SD sequence of the 16S RNA encoded by the rrnC operon results in a strong increase of PssA translation. We demonstrated that this mutation corresponds to the putative pssR regulator described in reference 10. These results completely rule out the hdfR gene as a candidate for pssR and indicate that HdfR is not involved in lipid metabolism regulation. HdfR has been reported to be a negative regulator of the flagellar master regulator flhDC and activates the gltBD operon (11, 14). It might be involved in a complex regulatory network involving H-NS and other regulators involved in acid stress resistance (14). However, its binding site has not been identified and we could not observe regulation of the gltBD promoter. By contrast, it seems that the adjacent yifE gene might be a genuine HdfR target gene, given its strong regulation and its classic genetic organization relative to hdfR. It was shown recently that YifE protein (renamed MaoP) is involved in the chromosome organization of the Ori macrodomain by binding to the specific maoS sequence localized in the hdfR-yifE intergenic region (16). It will be very interesting to understand the role of HdfR regulation and its connection with the MaoP/maoS site-specific control of the Ori macrodomain.
The fact that mutation C1538T in the 16S RNA rrsC coding gene corresponds to the pssR1 allele meant that we did not find a specific regulator of pssA expression. While no phospholipid synthesis gene other than pssA seemed to be affected by the mutation, its nature excludes a specific effect. Indeed, we showed by proteomics analyses that the expression of several genes was affected. However, we did not systematically verify that it was a direct effect on their translation. We suggest that the effect of the C1538T mutation depends on the nature of the SD sequence of a given gene. The strong effect on pssA translation reflects the suboptimal nature of the SD sequence of pssA, which might be better recognized by the mutated anti-SD sequence of 16S RNA. This weak initiation of translation might be important in the control of pssA expression and highlights the fact that gene expression can be modulated by means other than specific regulatory processes.
MATERIALS AND METHODS
Strain constructions.
For all strain and plasmid constructions, the Ecocyc website (21) was used for sequence retrieval. The construction of the various strains is described succinctly in Table 2. Deletions of hdfR, rrnC, rrlC, and rrsC were performed by direct recombination of PCR fragments obtained using pKD3 or pKD4 plasmids as the templates, following the procedure described by Datsenko and Wanner (22). Deletion mutant alleles obtained by recombination, from the Keio collection (23), or from tagged strains from the DY330 strain collection (13) were transduced from one background to another by generalized transduction with the phage P1 (24).
TABLE 2.
E. coli K-12 strains
| Lab codea | Name | Description | Reference |
|---|---|---|---|
| EB944 | MG1655 | Wild-type parental E. coli K-12 strain | |
| EB801 | BW25113 ΔhdfR::Kanr | 23 | |
| EB466 | BW25113 ΔyifK::Kanr | 23 | |
| EB814 | MG1655 ΔhdfR° | Transduction of ΔhdfR::Kanr from EB801 to MG1655, cassette removed using pCP20 plasmid | This work |
| EB815 | AC1 | pssR+ | 10 |
| EB816 | AC5 | pssR1 | 10 |
| EB356 | AC1 ΔhfdR::cat | Product of PCR with ebm1094/1081 on pKD3 of the cat cassette recombined in AC1 | This work |
| EB886 | AC5 ΔhdfR::cat | Product of PCR with ebm1094/1081 on pKD3 of the cat cassette recombined in AC5 | This work |
| EB1011 | MG ΔhdfR::cat pssR+ | Transduction of ΔhdfR::cat from EB356 to MG1655 | This work |
| EB430 | MG ΔhdfR::cat pssR1 | Transduction of ΔhdfR::cat from EB886 to MG1655 | This work |
| EB1068 | AC5 ΔyifK::Kanr (JB70) | pssR1, wild-type hdfR; transduction of ΔyifK::Kanr from EB466 to EB886 | This work |
| NA | DY330 series | Collection of strains with SPA or TAP tags on the chromosome | 13 |
| EB160 | AC1/PssA-SPA | Transduction of pssA-SPA-Kanr from the SPA collection to AC1 | This work |
| EB161 | AC5/PssA-SPA | Transduction of pssA-SPA-Kanr from the SPA collection to AC5 | This work |
| EB1052 | AC1/PlsB-SPA | Transduction of plsB-SPA-Kanr from the SPA collection to AC1 | This work |
| EB1053 | AC5/PlsB-SPA | Transduction of plsB-SPA-Kanr from the SPA collection to AC5 | This work |
| EB1050 | AC1/PlsC-SPA | Transduction of plsC-SPA-Kanr from the SPA collection to AC1 | This work |
| EB1051 | AC5/PlsC-SPA | Transduction of plsC-SPA-Kanr from the SPA collection to AC5 | This work |
| EB1054 | AC1/PgsA-TAP | Transduction of pgsA-TAP-Kanr from the SPA collection to AC1 | This work |
| EB1055 | AC5/PgsA-TAP | Transduction of pgsA-TAP-Kanr from the SPA collection to AC5 | This work |
| EB350 | AC1 ΔrrnC | Product of PCR with ebm1624/1620 on pKD4 recombined in AC1 | This work |
| EB352 | AC5 ΔrrnC | Product of PCR with ebm1624/1620 on pKD4 recombined in AC5 | This work |
| EB349 | AC1 ΔrrlC | Product of PCR with ebm1619/1620 on pKD4 recombined in AC1 | This work |
| EB351 | AC5 ΔrrlC | Product of PCR with ebm1619/1620 on pKD4 recombined in AC5 | This work |
| EB1062 | AC1 ΔrrsC | Product of PCR with ebm1624/1640 on pKD4 recombined in AC1 | This work |
| EB1063 | AC5 ΔrrsC | Product of PCR with ebm1624/1640 on pKD4 recombined in AC5 | This work |
NA, not applicable.
Plasmid constructions.
Plasmid constructions are described succinctly in Table 3. The hdfR ORF was amplified by PCR using AC1 and AC5 genomic DNA as the templates and cloned in the pBAD24 vector to obtain plasmids pEB1408 and pEB1501, respectively. Plasmid pcSacB-rrnC, which contains the entire rrnC operon, was PCR mutagenized to obtain pcSacB-rrnC(C1538T).
TABLE 3.
Plasmids
| Lab code | Namea | Descriptionb | Reference |
|---|---|---|---|
| pEB0268 | pKD3 | Ampr, FRT-Chlr-FRT cassette | 22 |
| pEB0269 | pKD4 | Ampr, FRT-Kanr-FRT cassette | 22 |
| pEB0267 | pKD46 | Ts, Ampr, lambda Red genes | 22 |
| pEB0266 | pCP20 | Ts, Ampr Chlr, FLP recombinase gene | 31 |
| pEB0227 | pBAD24 | Ampr, PBAD promoter | 32 |
| pEB1408 | pBAD-hdfR | PCR with ebm870/871 oligonucleotides on AC1 genomic DNA (EcoRI/XhoI) in pBAD24 (EcoRI/SalI) | This work |
| pEB1501 | pBAD-hdfR(G237S) | PCR with ebm870/871 oligonucleotides on AC5 genomic DNA (EcoRI/XhoI) in pBAD24 (EcoRI/SalI) | This work |
| pEB0898 | pUA66 | Kanr, sc101 ori, GFPmut2 | 25 |
| pUA-UPyifE | Region −196 to +99 relative to the start codon of yifE | 25 | |
| pEB1427 | pTrad-GFPmut2 | Kanr, sc101 ori, Δrbs, PCR mutagenesis on pUA66 with ebm927/928 oligonucleotides | This work |
| pEB1611 | pTrad-pssA (F1) | Kanr, sc101 ori, PCR with 427/1192 oligonucleotides in pEB1427 (XhoI/BamHI) | This work |
| pEB1895 | pUA-Plac 5′ UTR pssA (F3) | Kanr, sc101 ori, PCR with ebm1656/1657 oligonucleotides in pEB1427 | This work |
| pEB1856 | pUA-PpssA (F2) | Kanr, sc101 ori, PCR with ebm1205/1405 oligonucleotides in pUA66 (XhoI/BamHI) | This work |
| pEB1323 | pZEgfp | Ampr, pBR322 ori, GFPmut3aav | 26 |
| pEB1900 | pZE-Trad-GFPmut2 | Ampr, pBR322 ori, Δrbs, pEB1427 (XhoI/XbaI) insert in pEB1323 (XhoI/XbaI) | This work |
| pEB1853 | pPssA-GFPmut2 (F1) | Ampr, pBR322 ori, PCR with 427/1192 oligonucleotides in pEB1900 (XhoI/BamHI) | This work |
| pEB1864 | pPssA_mut10 | Ampr, PCR mutagenesis on pEB1853 with ebm1631/1632 oligonucleotides | This work |
| pEB1878 | pPssA_mutSD | Ampr, PCR mutagenesis on pEB1853 with ebm1654/1655 oligonucleotides | This work |
| pEB1858 | pcSacB-rrnC | Kanr | S. Quan, unpublished data |
| pEB1866 | pcSacB-rrnC rrsC(C1538T) | PCR mutagenesis on pEB1858 with ebm1641/1642 oligonucleotides | This work |
Trad signifies translational fusion.
Ts, thermosensitive replication; ori, origin of replication; Δrbs, deletion of ribosome binding site on the vector; Ampr, ampicillin resistance; Kanr, kanamycin resistance; Chlr, chloramphenicol resistance; FRT, flippase recognition target.
Transcriptional or translational fusions with GFP were constructed in the pUA66 vector backbone (25). When available, transcriptional fusions were retrieved from the Zaslaver collection (25), or else the promoter regions were PCR amplified and cloned between XhoI and BamHI restriction sites. The pssA-GFPmut2 translational fusions were also transferred in the pZE plasmid (26).
Measure of expression using transcriptional and translational fusions with GFP.
E. coli strains were transformed by the indicated plasmids derived from pUA66 (25) or pZE-GFPmut2 (pEB1900), and the selection plates were incubated at 37°C for 16 h. Six hundred microliters of LB medium supplemented with the required antibiotics was inoculated (4 to 6 replicates for each assay) and grown for 16 h at 30°C in 96-well polypropylene plates with 2.2-ml wells under aeration and agitation. Fluorescent intensity measurements were performed in a Tecan infinite M200 plate reader. One hundred fifty microliters from each well was transferred into a black Greiner 96-well plate for reading the optical density at 600 nm (OD600) and fluorescence (excitation, 485 nm; emission, 530 nm). The expression levels were calculated by dividing the intensity of fluorescence by the OD600. After mean values were calculated, the values from the control vector were subtracted. The results are given in arbitrary units because the intensity of fluorescence is acquired with an optimal and variable gain; hence, the absolute values cannot be compared between different types of experimental and growth conditions.
AC1 and AC5 strain genome resequencing.
Genomic DNA of AC1 and AC5 strains was prepared using the Wizard genomic DNA purification kit from Promega. Library construction, Illumina sequencing, and data analysis were performed by GATC Biotech.
Proteomics analysis of AC1 and AC5 strains.
AC1 and AC5 strains were grown in duplicates in 100 ml LB at 37°C until an OD600 of 2 was reached. Protein preparation and mass spectrometry-based proteomic analyses were realized as described in reference 27. Briefly, extracted proteins were stacked in the top of an SDS-PAGE gel (NuPAGE 4 to 12%; Invitrogen) before in-gel digestion was performed using trypsin (sequencing grade; Promega). Resulting peptides were analyzed in duplicate by online nanoscale liquid chromatography tandem mass spectrometry (nanoLC-MS/MS) (UltiMate 3000 and LTQ-Orbitrap Velos Pro; Thermo Scientific) using a 120-min gradient. Peptides and proteins were identified and quantified using MaxQuant (version 1.5.7.4 [28]) and Uniprot database (February 2017 version, Escherichia coli K-12 taxonomy). For statistical analysis, we used ProStaR (29). Proteins identified in the reverse and contaminant databases and proteins exhibiting less than 4 intensity values in one condition were discarded from the list. After log2 transformation, intensity values were normalized by median centering before missing value imputation (replacing missing values by the 1 percentile value of each column); statistical testing was conducted using a limma t test. Differentially expressed proteins were sorted out using a log2(fold change) cutoff of 1 and a false discovery rate threshold on P values of 1% using the Benjamini-Hochberg method.
SDS-PAGE and Western blotting.
SDS-PAGE, electrotransfer onto nitrocellulose membranes, and Western blot analyses were performed as previously described (30). Monoclonal anti-Flag M2, used for SPA tag detection, and PAP antibody, used for TAP tag detection, were purchased from Sigma. The relative amounts of fatty acid (FA) synthesis enzymes fused to the SPA tag or TAP tag were quantified by 10% SDS-PAGE and Western blotting. The amounts produced were then quantified using Alexa Fluor 680 anti-mouse IgG fluorescent secondary antibodies (Invitrogen) on an Odyssey Fc imager from LI-COR Biosciences.
Supplementary Material
ACKNOWLEDGMENTS
Our thoughts go to the late C. O. Raetz who first gave us the idea to follow the study on pssR. We thank the members of his laboratory, Hak Suk Chung and Jinshi Zhao, who were able to retrieve the historical AC1 and AC5 strains and send them to us. We thank Mickael Canestrari and Racym Bouharaoua for their help in the lab and James Sturgis for his constant support and help with the manuscript. We are indebted to P. Moreau from the LCB laboratory for managing the Keio collection. We thank Ciaran Condon for the gift of the rrnC plasmid and Rick Gourse for important suggestions. We also thank Sandrine Ollagnier for her help. Y.C. and L.B. are grateful for the support of the discovery platform and informatics group at EDyP.
This work was funded by the CNRS and Aix-Marseille University. Proteomic experiments were partly supported by the ProFi grant (ANR-10-INBS-08-01).
Footnotes
Supplemental material for this article may be found at https://doi.org/10.1128/JB.00202-17.
REFERENCES
- 1.Parsons JB, Rock CO. 2013. Bacterial lipids: metabolism and membrane homeostasis. Prog Lipid Res 52:249–276. doi: 10.1016/j.plipres.2013.02.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Durfee T, Hansen AM, Zhi H, Blattner FR, Jin DJ. 2008. Transcription profiling of the stringent response in Escherichia coli. J Bacteriol 190:1084–1096. doi: 10.1128/JB.01092-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Traxler MF, Summers SM, Nguyen HT, Zacharia VM, Hightower GA, Smith JT, Conway T. 2008. The global, ppGpp-mediated stringent response to amino acid starvation in Escherichia coli. Mol Microbiol 68:1128–1148. doi: 10.1111/j.1365-2958.2008.06229.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Wang AY, Cronan JEJ. 1994. The growth phase-dependent synthesis of cyclopropane fatty acids in Escherichia coli is the result of an RpoS(KatF)-dependent promoter plus enzyme instability. Mol Microbiol 11:1009–1017. doi: 10.1111/j.1365-2958.1994.tb00379.x. [DOI] [PubMed] [Google Scholar]
- 5.Eichel J, Chang YY, Riesenberg D, Cronan JEJ. 1999. Effect of ppGpp on Escherichia coli cyclopropane fatty acid synthesis is mediated through the RpoS sigma factor (sigmaS). J Bacteriol 181:572–576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Rhodius VA, Suh WC, Nonaka G, West J, Gross CA. 2006. Conserved and variable functions of the sigmaE stress response in related genomes. PLoS Biol 4:e2. doi: 10.1371/journal.pbio.0040002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Wahl A, My L, Dumoulin R, Sturgis JN, Bouveret E. 2011. Antagonistic regulation of dgkA and plsB genes of phospholipid synthesis by multiple stress responses in Escherichia coli. Mol Microbiol 80:1260–1275. doi: 10.1111/j.1365-2958.2011.07641.x. [DOI] [PubMed] [Google Scholar]
- 8.Raetz CR, Kantor GD, Nishijima M, Jones ML. 1981. Isolation of Escherichia coli mutants with elevated levels of membrane enzymes. A trans-acting mutation controlling diglyceride kinase. J Biol Chem 256:2109–2112. [PubMed] [Google Scholar]
- 9.Walsh JP, Loomis CR, Bell RM. 1986. Regulation of diacylglycerol kinase biosynthesis in Escherichia coli. A trans-acting dgkR mutation increases transcription of the structural gene. J Biol Chem 261:11021–11027. [PubMed] [Google Scholar]
- 10.Sparrow CP, Raetz CR. 1983. A trans-acting regulatory mutation that causes overproduction of phosphatidylserine synthase in Escherichia coli. J Biol Chem 258:9963–9967. [PubMed] [Google Scholar]
- 11.Ko M, Park C. 2000. H-NS-Dependent regulation of flagellar synthesis is mediated by a LysR family protein. J Bacteriol 182:4670–4672. doi: 10.1128/JB.182.16.4670-4672.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Riley M, Abe T, Arnaud MB, Berlyn MK, Blattner FR, Chaudhuri RR, Glasner JD, Horiuchi T, Keseler IM, Kosuge T, Mori H, Perna NT, Plunkett Gr Rudd KE, Serres MH, Thomas GH, Thomson NR, Wishart D, Wanner BL. 2006. Escherichia coli K-12: a cooperatively developed annotation snapshot–2005. Nucleic Acids Res 34:1–9. doi: 10.1093/nar/gkj405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Butland G, Peregrin-Alvarez JM, Li J, Yang W, Yang X, Canadien V, Starostine A, Richards D, Beattie B, Krogan N, Davey M, Parkinson J, Greenblatt J, Emili A. 2005. Interaction network containing conserved and essential protein complexes in Escherichia coli. Nature 433:531–537. doi: 10.1038/nature03239. [DOI] [PubMed] [Google Scholar]
- 14.Krin E, Danchin A, Soutourina O. 2010. Decrypting the H-NS-dependent regulatory cascade of acid stress resistance in Escherichia coli. BMC Microbiol 10:273. doi: 10.1186/1471-2180-10-273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Maddocks SE, Oyston PC. 2008. Structure and function of the LysR-type transcriptional regulator (LTTR) family proteins. Microbiology 154:3609–3623. doi: 10.1099/mic.0.2008/022772-0. [DOI] [PubMed] [Google Scholar]
- 16.Valens M, Thiel A, Boccard F. 2016. The MaoP/maoS site-specific system organizes the Ori region of the E. coli chromosome into a macrodomain. Plos Genet 12:e1006309. doi: 10.1371/journal.pgen.1006309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Al Mamun AA, Lombardo MJ, Shee C, Lisewski AM, Gonzalez C, Lin D, Nehring RB, Saint-Ruf C, Gibson JL, Frisch RL, Lichtarge O, Hastings PJ, Rosenberg SM. 2012. Identity and function of a large gene network underlying mutagenic repair of DNA breaks. Science 338:1344–1348. doi: 10.1126/science.1226683. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Jacob WF, Santer M, Dahlberg AE. 1987. A single base change in the Shine-Dalgarno region of 16S rRNA of Escherichia coli affects translation of many proteins. Proc Natl Acad Sci U S A 84:4757–4761. doi: 10.1073/pnas.84.14.4757. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Thomason MK, Bischler T, Eisenbart SK, Forstner KU, Zhang A, Herbig A, Nieselt K, Sharma CM, Storz G. 2015. Global transcriptional start site mapping using differential RNA sequencing reveals novel antisense RNAs in Escherichia coli. J Bacteriol 197:18–28. doi: 10.1128/JB.02096-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Karzai AW, Susskind MM, Sauer RT. 1999. SmpB, a unique RNA-binding protein essential for the peptide-tagging activity of SsrA (tmRNA). EMBO J 18:3793–3799. doi: 10.1093/emboj/18.13.3793. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Keseler IM, Mackie A, Peralta-Gil M, Santos-Zavaleta A, Gama-Castro S, Bonavides-Martinez C, Fulcher C, Huerta AM, Kothari A, Krummenacker M, Latendresse M, Muniz-Rascado L, Ong Q, Paley S, Schroder I, Shearer AG, Subhraveti P, Travers M, Weerasinghe D, Weiss V, Collado-Vides J, Gunsalus RP, Paulsen I, Karp PD. 2013. EcoCyc: fusing model organism databases with systems biology. Nucleic Acids Res 41:D605-12. doi: 10.1093/nar/gks1027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Datsenko KA, Wanner BL. 2000. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci U S A 97:6640–6645. doi: 10.1073/pnas.120163297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Baba T, Ara T, Hasegawa M, Takai Y, Okumura Y, Baba M, Datsenko KA, Tomita M, Wanner BL, Mori H. 2006. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol Syst Biol 2:2006.0008. doi: 10.1038/msb4100050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Miller JH. 1992. A short course in bacterial genetics: a laboratory manual and handbook for Escherichia coli and related bacteria, vol 2 Cold Spring Harbor Laboratory Press, Plainview, NY. [Google Scholar]
- 25.Zaslaver A, Bren A, Ronen M, Itzkovitz S, Kikoin I, Shavit S, Liebermeister W, Surette MG, Alon U. 2006. A comprehensive library of fluorescent transcriptional reporters for Escherichia coli. Nat Methods 3:623–628. doi: 10.1038/nmeth895. [DOI] [PubMed] [Google Scholar]
- 26.de Jong H, Ranquet C, Ropers D, Pinel C, Geiselmann J. 2010. Experimental and computational validation of models of fluorescent and luminescent reporter genes in bacteria. BMC Syst Biol 4:55. doi: 10.1186/1752-0509-4-55. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Milbradt J, Kraut A, Hutterer C, Sonntag E, Schmeiser C, Ferro M, Wagner S, Lenac T, Claus C, Pinkert S, Hamilton ST, Rawlinson WD, Sticht H, Couté Y, Marschall M. 2014. Proteomic analysis of the multimeric nuclear egress complex of human cytomegalovirus. Mol Cell Proteomics 8:2132–2146. doi: 10.1074/mcp.M113.035782. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Cox J, Mann M. 2008. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat Biotechnol 12:1367–1372. doi: 10.1038/nbt.1511. [DOI] [PubMed] [Google Scholar]
- 29.Wieczorek S, Combes F, Lazar C, Giai Gianetto Q, Gatto L, Dorffer A, Hesse AM, Couté Y, Ferro M, Bruley C, Burger T. 2017. DAPAR & ProStaR: software to perform statistical analyses in quantitative discovery proteomics. Bioinformatics 33:135–136. doi: 10.1093/bioinformatics/btw580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Gully D, Moinier D, Loiseau L, Bouveret E. 2003. New partners of acyl carrier protein detected in Escherichia coli by tandem affinity purification. FEBS Lett 548:90–96. doi: 10.1016/S0014-5793(03)00746-4. [DOI] [PubMed] [Google Scholar]
- 31.Cherepanov PP, Wackernagel W. 1995. Gene disruption in Escherichia coli: TcR and KmR cassettes with the option of Flp-catalyzed excision of the antibiotic-resistance determinant. Gene 158:9–14. doi: 10.1016/0378-1119(95)00193-A. [DOI] [PubMed] [Google Scholar]
- 32.Guzman LM, Belin D, Carson MJ, Beckwith J. 1995. Tight regulation, modulation, and high-level expression by vectors containing the arabinose PBAD promoter. J Bacteriol 177:4121–4130. doi: 10.1128/jb.177.14.4121-4130.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.