Abstract
Microsporidia are ubiquitous intracellular pathogens whose opportunistic nature led to their increased recognition with the rise of the AIDS pandemic. As the RNA world was largely unexplored in this parasitic lineage, we developed a dedicated in silico methodology to carry out exhaustive identification of ncRNAs across the Encephalitozoon and Nosema genera. Thus, the previously missing U1 small nuclear RNA (snRNA) and small nucleolar RNAs (snoRNAs) targeting only the LSU rRNA were highlighted and were further validated using 5' and 3'RACE-PCR experiments. Overall, the 15 ncRNAs that were found shared between Encephalitozoon and Nosema spp. may represent the minimal core set required for parasitic life. Interestingly, the systematic presence of a CCC- or GGG-like motif in 5' of all ncRNA and mRNA gene transcripts regardless of the RNA polymerase involved suggests that the RNA polymerase machineries in microsporidia species could use common factors. Our data provide additional insights in accordance with the simplification processes observed in these reduce genomes and underline the usefulness of sequencing closely related species to help identify highly divergent ncRNAs in these parasites.
Keywords: Microsporidia, ncRNA prediction, ncRNA structure, genome evolution, transcriptional regulation signals
1. Introduction
Microsporidia are unicellular eukaryotic parasites divided into more than 187 genera and 1,500 distinct species of varying medical, veterinary and economic impacts.1 These highly-adapted, obligate intracellular pathogens were recently shown to cluster at the base of the fungal kingdom as a sister-group to chytrid pathogen Rozella allomycis.2 Unlike most of their fungal relatives however, Microsporidia cannot thrive without a host and can only survive in the outside environment as spores. The intracellular lifestyle of Microsporidia is obligate and likely irreversible, as all but one species (i.e. Mitosporidium daphniae)3 feature highly-reduced mitochondria known as mitosomes that are incapable of generating ATP via oxidative phosphorylation and gene-depleted nuclear genomes, reflecting a heavy host-dependence towards a large number of essential cellular components that they are no longer able to produce.4 Of the circa 1,800–2,600 proteins5 encoded in the microsporidian gene repertoire, a total of about 800 proteins form a reduced core set involved in critical cellular processes and pathogenicity, including spore formation, invasion and host-parasite relationships,4,6,7 with the remaining proteome postulated to reflect their adaptation to different niches.8 However, while recent investigations have painted a clearer picture of the microsporidian pan-proteome, we still know very little about the types and distribution of non-coding RNAs (ncRNAs) that are present in Microsporidia.
In eukaryotic cells, three RNA polymerases (RNA Pol I, II and III) are involved in the transcription processes of ncRNAs. RNA Pol I is specialized in the high level synthesis of the large rRNA precursor from a single type promoter.9 RNA Pol III transcribes numerous ncRNAs, including tRNAs, 5S rRNA, and a variety of other types of ncRNA such as U6 snRNA, RNase P RNA and SRP RNA, by recognizing cis-acting elements located within the transcribed region.10 Finally, RNA Pol II, which transcribes protein-coding genes, is also devoted to the production of ncRNA such as other small nuclear RNAs (snRNAs) and small nucleolar RNAs (snoRNAs).11
Non-coding RNAs are diverse and biologically relevant molecules involved in various crucial cellular processes.12–15 For example, snRNAs, snoRNAs and RNase ncRNAs are found in most eukaryotic cells and are involved in the splicing of eukaryotic pre-mRNAs, ribosome-related 2'-O-methylation (C/D snoRNA) or pseudouridylation (H/ACA snoRNA), and miscellaneous maturation processes, respectively. However, current ncRNA investigations are hampered by our limited prediction, detection and identification capabilities, which lag far behind those coding for proteins. Indeed, ncRNAs lack the signatures that make protein-coding gene prediction possible (e.g. codon usage, splicing signals, open reading frame length, sequence bias) and similarity searches are often inefficient due to the lack of primary sequence conservation across distantly related genomes.16,17 The most successful approaches to identify ncRNAs exploit the idea that functionally significant RNA secondary and/or tertiary structures will be conserved in distant species, as function is usually dependent on structure, and structures can be maintained despite high sequence divergence by compensatory mutations. Predicting consensus secondary/tertiary structures can thus be far more useful than alignments of closely related primary sequences when working across large evolutionary distances or with sequences displaying extremely fast rates of evolution, as is the case with most microsporidian genomes.18
Most approaches currently used to identify ncRNAs start from the analysis of transcriptome data, which are then mapped onto genomic data.19 This circumscribes the scope of the search to a smaller subset of transcribed and therefore potentially functional sequences, a clear advantage when the genomes under scrutiny are huge. However, distinguishing between function and transcriptional noise can be difficult, and the tools that are available to process transcriptome data are still in flux. Here, we used instead the reverse approach, from genome prediction to transcriptome validation, to investigate the presence of ncRNAs in the genomes of Microsporidia from the four Encephalitozoon species. This approach is well-suited to Encephalitozoon genomes due to their extreme size reduction (smaller than 3 Mbp), high-levels of gene synteny and coding density, and characterized CCC-/GGG-like transcriptional initiation signals.6,20–22 Using this approach, we identified a total of ten new ncRNAs in the Encephalitozoon spp., including the previously missing U1 snRNA and nine snoRNA-like genes. Their presence and that of eight previously predicted ncRNAs were further confirmed by 5' and 3′ RACE-PCR experiments in E. cuniculi. To strengthen the relevance of our comparative approach and to better evaluate the overall catalogue of ncRNA in microsporidian genomes, we also applied the same method to the available genomes from the genus Nosema. Most of the ncRNA found in the four Encephalitozoon species were also identified in Nosema spp. This suggests that the repertoire of ncRNA may be relatively reduced in the microsporidian genomes and, taken altogether, that these ncRNAs may very well represent the minimal core set required by these parasites for their survival.
2. Material and methods
2.1. Detection and characterization of microsporidian ncRNAs
Conventional approaches used to annotate coding DNA sequences (CDS) consider the first AUG codon in an open reading frame as the transcriptional initiation site (TIS). However, this is often inaccurate and can cause the erroneous inclusion of intergenic regions and/or other small overlapping unpredicted genes in the 5' end of the CDS. To mitigate this and help identify ncRNA that could potentially be found in these overlapping segments, we first revisited the latest annotations from Encephalitozoon cuniculi (accession codes AL391737.2, AL590442.2-AL590451.2), Encephalitozoon intestinalis (accession codes CP001942.1-CP001952.1), Encephalitozoon hellem (accession codes CP002713.1-CP002724.1) and Encephalitozoon romaleae (accession codes CP003518.1-CP003530.1) genomes to better predict TIS and intergenic regions by taking advantage of transcriptional signals and the increased resolution in comparative analysis provided by the growing set of Encephalitozoon genomes.6,21,23 From these curated annotations we then used an “all-versus-all” BLASTN approach (word size: 7 nt, low-complexity filter disabled) to compare the intergenic regions of these four genomes and search for the presence of small but highly conserved stretches of DNA reflecting elevated levels of conservation—and potentially functional importance—in these genomes with the highest rates of sequence evolution in eukaryotes.18 BLASTN searches were also further performed in a directed fashion by specifically comparing intergenic regions from the same locus in the four species, which was made possible due to the extreme levels of synteny found across all Encephalitozoon genomes (1,824 colinear genes in 55 syntenic blocks).20 BLASTN results were manually explored and flanking regions analysed to identify putative transcriptional regulation signals. Multiple sequence alignments were performed with Clustal Omega24 to validate sequence conservation between the four Encephalitozoon genomes.
The presence of ncRNAs in species from the genus Nosema was first investigated by screening the Nosema genomes (N. ceranae (accession codes JPQZ01000001-JPQZ01000536), N. apis (accession codes ANPH01000001-ANPH01001133) and N. bombycis (accession codes ACJZ01000001-ACJZ01003558)) for the Encephalitozoon ncRNA genes identified as described above using BLASTN searches (word size: 7 nts, Match/Mismatch Scores (1, −1), Gap Costs (Existence: 0 Extension: 2) and low-complexity filter disabled). Next, because accurate annotation and TIS prediction in Nosema genomes has been exclusively carried out for N. ceranae,25 only the intergenic regions from this species were extracted and used as queries against the N. apis and N. bombycis genomes for genus-specific BLASTN searches (same parameters), with putative hits investigated for the presence of transcriptional regulation signals as described above. Canonical secondary structures were predicted using locARNA26 and refined manually. In silico functional annotations of identified ncRNAs were conducted using the RFAM 12.0 database.27 ncRNAs lacking predicted functions after this step were annotated according to the expertize available in our group as follows. The C/D box snoRNA genes were annotated based on their conserved C (RUGAUGA) and D (CUGA) sequence motifs located at the 5' and 3′ ends, respectively, and based on a short terminal stem located 2 nt upstream and of the C and D boxes, respectively. Most of the C/D box candidate snoRNA genes also further contained another copy of the C and D motifs, albeit less conserved, located in the central region of the snoRNA sequence. H/ACA box snoRNA genes were annotated based on the presence of two hairpins and of the two H (ANANNA) and ACA (ACA) sequence motifs, located in the intergenic region between the two hairpins and at the 3′ end immediately following the second hairpin, respectively. The U1 snRNA was annotated based on its well-conserved secondary structure and the presence of the 5′ end highly conserved SS (ACUUAC) motif.
To identify snoRNA targets in microsporidian rRNA, rRNA alignments between E. cuniculi, human and S. cerevisiae were performed to highlight homologous targets positions. In-house scripts were used to predict putative modified positions from both E. cuniculi snoRNA and rRNA sequences. rRNA alignments were built using multalin,28 (http://multalin.toulouse.inra.fr) for human, yeast and Encephalitozoon genomes. Positions modified in S. cerevisiae and human rRNA were obtained from the Human snoRNA29 and the S. cerevisiae snoRNA30 database, respectively. rRNA targets of Encephalitozoon box C/D and H/ACA snoRNA were then predicted by searching in rRNAs for putative base pairings to the expected regions in snoRNA (upstream the D or D’ box for C/D box snoRNA and in the pocket of the stem loop for H/ACA box snoRNA). E. cuniculi candidate targets were reported in the alignment. Positions labeled as modified in human and/or yeast and predicted as modified in E. cuniculi were kept as true positive modifications. SnoRNA actors of these modifications were assigned the same identifier than in human/yeast when possible. Identification of consensus sequences of A and B boxes from tRNA sequences have been performed with the MEME tool.31
2.2. Cell culture
Human Foreskin Fibroblast (HFF) host cells (ATCC SCRC-1041) were infected by approximately 109 spores of E. cuniculi GB-M1 (kindly provided by Prof. Elisabeth U. Canning, Imperial College of Science, Technology and Medicine, London, UK) during 2 h in 75 cm2 flasks. Cultures were washed three times with PBS (1×) to remove spores that did not invade host cells and then incubated for two further days as described previously.32 Infected cells were maintained in 5% CO2 at 37 °C in minimum essential medium (MEM) supplemented with 5% fetal calf serum, 2 mM glutamine (Invitrogen, Carlsbad, CA, USA) and 20 µg/ml gentamicin.
2.3. Total RNA isolation and polyuridylation
Total RNA from E. cuniculi-infected HFF cells were extracted using TRIzol (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instructions. Total RNA integrity and purity were verified on the 2200 TapeStation system (Agilent Technologies, Santa Clara, CA, USA). Three micrograms of total RNAs were polyuridylated using a polyU polymerase in 25 µl reactions according to the manufacturer’s instructions (New England Biolabs, Ipswich, MA, USA), purified by phenol/chloroform extraction followed by ethanol precipitation, and resuspended in RNase-free water.
2.4. 5′RACE-PCR experiments and 3′ end amplification
5′ cDNA ends were characterized with the SMARTer RACE Amplification kit (Clontech Laboratories, Inc., Mountain View, CA, USA) according to the manufacturer’s recommendations. The reverse transcription (RT) reaction step was performed with 200 ng of E. cuniculi total polyuridylated RNAs using a universal poly(A)-stem-loop RT primer.33 This first strand reaction products were diluted with 50 µl of tricine-EDTA buffer and used both for 5′ RACE-PCR (0.2 µM of each specific primer, 0.2 mM dNTPs, 2 U of Taq polymerase) according to the manufacturer’s recommendations and 3′ end amplification using specific 3′ primers and the universal reverse primer on an Eppendorf Mastercycler gradient PCR machine with the following cycling parameters: 10 cycles of touch-down PCR (denaturation: 94 °C for 30s; annealing: 55–68 °C for 30 s; extension: 72 °C for 30 s), followed by 30 cycles of regular PCR with annealing at 52 °C. Specific 5′ and 3′ primers were defined using KASpOD software.34
2.5. PCR products sequencing
Amplification products were analysed by electrophoresis on 1.5% agarose gels. Bands of the expected sizes were excised and purified using the Wizard SV Gel and PCR Clean-Up System (Promega, Madison, WI, USA). Purified PCR products were directly sequenced with the specific primers from the RACE amplifications. For weak band signal, PCR products were ligated into the pCR II TOPO vector (TOPO TA Cloning Kit Dual Promoter, Invitrogen) and transformed into chemically competent XL1-Blue Escherichia coli cells following the Inoue method.35 All sequences were determined using the Sanger dideoxynucleotides chemistry by MWG Operon (Ebersberg, Germany) with the SP6 primers.
3. Results
3.1. Identification and validation of non-coding RNA genes in Encephalitozoon and Nosema spp
Using a synteny-driven “all-versus-all” BLASTN approach, a total of 10 new putative ncRNAs were predicted in the genomes of the four closely related Encephalitozoon species E. cuniculi, E. intestinalis, E. romaleae and E. hellem (Supplementary Table S1), raising the total to 18 with the eight ncRNAs (RNase P, MRP, SRP, U3 C/D snoRNA, U2, U4, U5 and U6 snRNAs) previously described in the RFAM 12.0 database27 for at least one of these genomes. Similarity and synteny-based ortholog searches in three AT-rich Nosema genomes (N. ceranae, N. apis and N. bombycis) retrieved only four out of the ten Encephalitozoon newly predicted ncRNAs, suggesting that the primary sequences of ncRNAs are overall poorly conserved throughout Microsporidia. However, using only intergenic regions of N. ceranae to screen N. apis and N. bombycis ones, we were able to identify seven additional ncRNAs. Thus, with the RNase P, U2, U3 and U6 ncRNA genes available in the RFAM 12.0 database, Nosema species would possess 15 ncRNA genes that are orthologous to Encephalitozoon ones (Supplementary Table S1).
All 18 E. cuniculi ncRNAs were then validated by 5′ and 3′ RACE-PCR (Supplementary Fig. S1) and Sanger sequencing of the full-length products. This approach allowed to precisely define their transcriptional initiation and termination sites (Fig. 1). The lengths of ncRNA primary sequences were found to be relatively well conserved between the four Encephalitozoon species and the three Nosema species (Supplementary Fig. S2) but otherwise to be generally shorter when compared with their respective yeast and human orthologs (Supplementary Table S1). For instance, the sequence of the SRP RNA is 60 nt shorter than its S. cerevisiae and H. sapiens homologs predicted in silico.36 However, U4 and U6 snRNAs are longer than their human and yeast orthologs and include unexpected additional stems located near their 3′ end (Supplementary Fig. S3). Despite the overall size reduction and the lack of sequence similarity compared with the genus and other phyla, the Encephalitozoon ncRNAs were predicted to fold into canonical secondary structures, with the exception of an uncommon Alu domain in the Encephalitozoon SRP RNA (Fig. 2 and Supplementary Table S2).
Figure 1.
Encephalitozoon cuniculi ncRNAs validated by 5′ and 3′ RACE-PCR and their potential regulation signals. Black boxes correspond to the potential TATA box signals for transcription initiation by RNA polymerases II or III. The potential transcriptional motifs (CCC/GGG-like signals) are boxed. For brevity, the ncRNA sequences were only represented by the transcription start site, the end of transcription and the corresponding gene name.
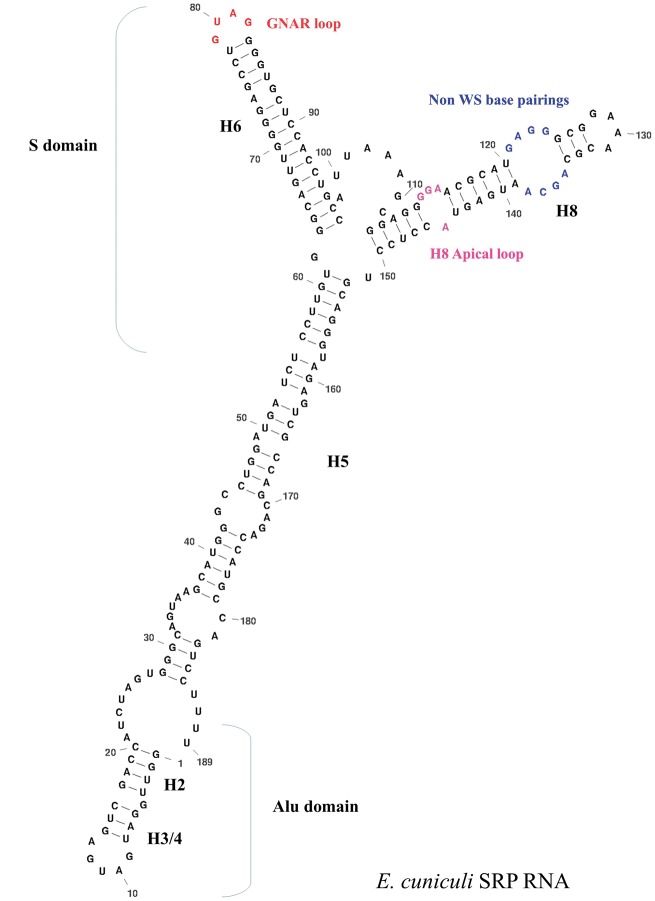
Figure 2.
Proposed secondary structure for the E. cuniculi SRP non-coding RNA. Specific domains of the molecule are given.
Analyses of the upstream regions for all the 18 E. cuniculi ncRNAs revealed the presence of a conserved CCC or GGG motif located in close proximity of the transcriptional start site (TSS) (Fig. 1). Complementary analyses of upstream regions have also highlighted a TATA-like sequence for all the genes with the exception of the E. cuniculi and E. intestinalis U46 C/D snoRNA (Fig. 1 and Supplementary Table S2). For this last gene, in these two species, the TSS upstream region presents a higher cytosine and guanine percentage, such that we could locate only multiple CCC or GGG-like motifs. Searches for DNA signals implicated in 5′ transcript processing for the other Encephalitozoon and Nosema species have shown the same signals for all the predicted ncRNA genes (Supplementary Table S2). In S. cerevisiae and more generally in eukaryotic cells, tRNA, U6 snRNA, RNase P RNA and SRP RNA genes are conventionally under the control of the RNA polymerase III (RNA Pol III), specifically recruited by A or B box sequence motifs.37 While these cis-acting elements were unambiguously identified for all Encephalitozoon and Nosema tRNA genes, none could be found for the other three ncRNA families (Supplementary Table S2).
3.2. Annotation of new ncRNAs
All ten newly predicted ncRNAs in the Encephalitozoon genus were assigned putative functions based on their predicted secondary structures (Supplementary Fig. S3). With the exception of the U1 spliceosomal snRNA, all newly identified ncRNAs fall within the C/D box or H/ACA snoRNA families. The U1 snRNA in Encephalitozoon species has retained the typical cloverleaf-like structure (Fig. 3a) with a four-way junction and a terminal stem-loop.38 However, the sequence of their SL1 and SL2 loops, which are key sites for snRNP specific proteins binding, differ significantly from their S. cerevisiae and human homologs and lack recognizable U1-70k (GAUCRYGARR) and U1A (YUGCAYUY) canonical protein binding sites.39 In agreement with motifs present at intron boundaries in Encephalitozoon, U1 snRNA displays a conserved human-like ACUUACC 5′ splicing site (SS) motif. A binding site for the Sm proteins was identified at the expected position but differs from human and yeast Sm sites at positions 6, 8 and 9 (Fig. 3a and b). Using the above characteristics derived from the Encephalitozoon spp. U1 snRNAs, we were able to successfully confirm the presence of U1 orthologs in the three genomes from the Nosema genus (Fig. 3b). Remarkably, in microsporidia the length of the SL3 variable stem loop is shorter than in human and yeast. Interestingly, the SL2 stemloop sequence seems much more conserved between Encephalitozoon and Nosema species than SL1, whose sequence seems more genus specific.
Figure 3.
Proposed secondary structure for the E. cuniculi U1 snRNA (a) and multiple sequence alignment with U1 snRNA from other microsporidia, human and yeast (b). In the secondary structure (a), the 5′ SS and Sm conserved motifs are shaded orange and blue, respectively. (b) Multiple sequence alignment of U1 snRNA from Encephalitozoon (Ec, Eh, Ei and Er), Nosema (Na, Nc and Nb), Human (Hs) and S. cerevisiae (Sc) species. Nucleotides are coloured according to the stem they belong to. Ec, E. cuniculi; Ei, E. intestinalis; Eh, E. hellem; Er, E. romaleae; Na, N. apis; Nc, N. ceranae; Nb, N. bombycis. [n]: insertion of n nucleotides in Sc.
Three C/D box snoRNAs were identified as homolog of U31/snR67, U38/snR61 and U40-U46/snR63 in human/S. cerevisiae based on the presence of the box C (RUGAUGA) and D (CUGA) conserved sequence elements, of a terminal stem, and of nucleotide targets in microsporidian rRNAs that are complementary to the nucleotides located upstream the D or D' boxes (Fig. 4a). The ncRNA located on E. cuniculi chromosome I (Supplementary Table S1) also displays C and D motifs characteristic of C/D box snoRNAs as well as an expected terminal stem (Supplementary Fig. S3). However, this ncRNA does not display any clear target in microsporidian rRNAs.
Figure 4.
Putative rRNA targets for the C/D and H/ACA snoRNAs encoded by E. cuniculi. Base-pairing interactions between: (a) Putative 2′-O methylated target regions in the 23S LSU rRNA and guide sequences within C/D box snoRNAs and (b) putative Ψ target regions in LSU rRNA and guide sequences within H/ACA box snoRNAs.
Five of the newly-predicted ncRNAs were annotated as box H/ACA snoRNAs (Supplementary Table S1 and Fig. S2). In all cases, putative 23S rRNA pseudouridylation pockets were found located 14 to 15 nt upstream from box H (ANANNA) and/or ACA terminal motifs (Fig. 4b), with hairpin secondary structures derived from multiple sequence alignment consensus compatible with those commonly found in snoRNAs H/ACA box. Based on their expected targets, we were able to assign human and S. cerevisiae orthologs to four (ACA10/snR37, U65/snR34, ACA27/snR42 and U19/snR191) of the five microsporidian H/ACA box snoRNAs. In human and S. cerevisiae, ACA10/snR37 targets two positions, respectively in 28S and 18S rRNA. Remarkably, in all available Encephalitozoon species—supposed more closely related to S. cerevisiae—ACA10/snR37 snoRNA contains only one pseudouridylation pocket which target the 23S rRNA at the expected conserved position. We were unable to propose rRNA or snRNA targets for the H/ACA snoRNA encoded by E. cuniculi chromosome II (Supplementary Table S1), which is conserved in Nosema genus but apparently not in S. cerevisiae and human.
4. Discussion
Predictive searches based on homology are inherently tied to the similarities that exist between targets and queries and, as sequences diverge, the signature elements that could help identify biologically-relevant relationships slowly fade away. Non-coding RNAs, in particular, are rather difficult to predict due to their general lack level of primary sequence homology and to the limited availability of software-based tools capable of investigating their secondary or tridimensional structures. In this study, we used the availability of multiple closely related Encephalitozoon and Nosema genomes as well as the presence of transcriptional signals to improve ncRNA gene prediction in Microsporidia from precisely delineated intergenic regions6,21 (here we note that intronic ncRNAs are highly unlikely in these species because few introns are present in Microsporidia and their introns are very short). Using our approach, we were able to identify ten new ncRNAs that are involved in pre-RNA processing and rRNA modification, thus in effect more than doubling the number of known ncRNAs encoded in these species.
4.1. A complete RNA machinery of the major spliceosome?
Despite their eukaryotic ancestry, microsporidia from the genus Encephalitozoon are intron poor, with less than 50 known spliceosomal introns per genome,6,40 and it is unclear if this scarcity is a consequence of or rather the cause behind the apparent reduced complexity of the spliceosome apparatus in these organisms. The spliceosome of E. cuniculi was predicted based on genome data to be much simpler than that of other eukaryotes and to be composed of only 37 proteins39,41,42 compared to its human (∼200) and yeast (∼100) counterparts, yet it clearly remains functional as indicated by splicing studies.44,45 However, up until now, many ncRNA components considered vital to the function of the eukaryotic spliceosome were missing from microsporidian annotations. In particular, although four U1-associated proteins were identified in the E. cuniculi genome, the U1 snRNA previously escaped bioinformatics detection despite large screenings39,41,43 that predicted the presence of U1 in only one microsporidia from the genus Antonospora. Our study represents the first evidence of the presence of U1 in Encephalitozoon and three Nosema species and confirms, as expected from splicing studies, that microsporidia possess a complete set of snRNA for the major spliceosome. With this small but whole set of snRNAs, its few splicing proteins and the scarce introns that are encoded in its genome, E. cuniculi can be now considered one of the simplest complete systems to investigate eukaryotic splicing mechanisms.
4.2. Towards a minimal set of compact ncRNA genes
Microsporidia from the genus Encephalitozoon are models of extreme compaction with thoroughly streamlined genomes, a streamlining so pervasive that the functions left encoded within are likely essential to the survival of these parasites. As such, the subset of Encephalitozoon ncRNAs (15 out of 18) that is shared with Nosema species may very well represent the minimal set of ncRNA genes that is viable for any spliceosome-dependent eukaryote. The reductionist pressure exerted on the Encephalitozoon genomes resulted in a drastic genes loss during microsporidian evolution, in the shortening of the protein coding sequences and rRNA-coding regions but also in a general reduction in ncRNA size. A careful analysis of the secondary structure of each ncRNA revealed that their cores were reduced to the structural features that are preserved in all living cells, and that further reduction is unlikely without disrupting the underlying biological function(s). For example, the highly derived P3 helix and 3′ domains of the RNase P and MRP RNAs, which are not critical for RNA processing,45 are highly reduced in Encephalitozoon spp. and compatible with the absence of the interacting protein Rpp20 from these organisms.46 Another example of high reduction is the SRP RNA, which lacks a canonical Alu domain in Encephalitozoon species. Encephalitozoon spp. lacks apparently the protein heterodimers that interact with this domain, i.e. SRP9/SRP14 and SRP68/SRP72, and while these heterodimers could potentially be replaced by structurally related proteins, the reduced size of the microsporidian SRP RNA strongly argues against this.
With a total number of snoRNAs that is but a fifth of that encountered in the budding yeast S. cerevisiae,47 the Encephalitozoon and Nosema genomes are pushing our understanding of how far ribosomes can devolve to yet remain functional. Here, all of the conserved microsporidian snoRNAs targeted modifications are realized in key regions of rRNAs, which argues in favor of an essential role for ribosome function.48,49 In the snoRNAs we identified here, even those with reduced structures have retained their characteristic target-binding pattern and sequence motifs, suggesting that very little is left to prune. Remarkably, we did not find any snoRNA involved in the modification of SSU rRNA or snRNA. The only H/ACA box snoRNA ortholog able to modify SSU rRNAs in human and S. cerevisiae lacks the necessary 3′ pocket in Microsporidia, which begs the question: are SSU-modifying snoRNAs present at all in Microsporidia? We cannot exclude the possibility that highly divergent specimens are left to be discovered, but in light of the highly reduced microsporidian ribosomal rRNAs, they may be no longer necessary.
4.3. A simplified RNA polymerase III recruitment?
Non-coding RNAs in eukaryotes are transcribed by one of the three RNA polymerases known as RNA Pol I, II and III, with transcription initiation signals specific to each polymerase. Most genes transcribed by RNA Pol III fall into well-defined groups depending on the location or type of cis-acting elements, which constitute their promoters.10,50,51 In a significant subset of genes under control of the RNA Pol III, two regions known as A and B boxes are involved in the binding of the transcriptional factor TFIIIC.10 As expected, we identified these cis-acting A and B boxes—harboring the canonical consensuses TRGYNNANNNG and GWTCRANNC—for all tRNA genes in E. cuniculi and N. ceranae genomes (Supplementary Table S2). However, we could not locate these signals or any other putative Pol III-related signal in others microsporidian ncRNA genes known to be transcribed in eukaryotes by RNA Pol III (U6 snRNA, RNase P and MRP RNA). Because they play an important role in the formation of the L-shaped tertiary structure of the tRNA molecule, the A and B boxes could be subjected to more selection pressure when associated with tRNA genes, limiting consequently their variation.10
Given these results, two major assumptions can be formulated. The first hypothesis is that although identifiable in tRNA genes, the A and B boxes used for the recruitment of the RNA Pol III in microsporidian ncRNAs are probably too degenerated to be detected by a homology-based method. This is consistent with the fact that these genomes display some of the highest rates of sequence evolution in eukaryotes.18 The high rate of sequence divergence could also explain why the 5S rRNA internal control region (ICR) C-box required for transcription and relatively well conserved between S. cerevisiae and Xenopus borealis rRNA 5S genes,52 could not be characterized in microsporidia despite their phylogenetic proximity with S. cerevisiae. The second hypothesis is based on an in vitro observation showing that in S. cerevisiae, TFIIIB can recruit RNA Pol III without the help of TFIIIC.53 Thus, a TATA box, without the need of any other transcriptional signal, could directly ensure TFIIIB and ultimately RNA Pol III recruitment for multiple rounds of accurately initiated transcription. In this scenario, the A and B boxes would not be required at all, and could be missing entirely from the Encephalitozoon ncRNAs.
Interestingly, the systematic presence of a perfect CCC or GGG motif in 5′ of all ncRNA and mRNA gene transcripts in E. cuniculi,6,23,42 regardless of the RNA polymerase involved (I, II or III), suggests that the RNA polymerase machinery in Encephalitozoon species could perhaps use common factors, in agreement with the apparent overall simplification of transcriptional processes in Microsporidia.
In any case, novel experimental strategies will be required to understand more accurately the transcriptional regulation of coding and non-coding genes in Microsporidia. Notably, a better characterization of their general and specific transcription factors will further help identify which components of the transcriptional machinery are essential in these ultra-compact genome.
Supplementary data
Supplementary data are available at www.dnaresearch.oxfordjournals.org.
Conflict of interest
None declared.
Funding
This work was supported in part by the National Institute of Allergy and Infectious Diseases of the National Institutes of Health under award number R15AI128627 to Jean-François Pombert. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Supplementary Material
References
- 1. Vavra J., Lukes J.. 2013, Microsporidia and ‘the art of living together’. Adv. Parasitol., 82, 253–319. [DOI] [PubMed] [Google Scholar]
- 2. James T.Y., Pelin A., Bonen L., et al. 2013, Shared signatures of parasitism and phylogenomics unite Cryptomycota and microsporidia. Curr. Biol., 23, 1548–53. [DOI] [PubMed] [Google Scholar]
- 3. Haag K.L., James T.Y., Pombert J.F., et al. 2014, Evolution of a morphological novelty occurred before genome compaction in a lineage of extreme parasites. Proc. Natl. Acad. Sci. USA, 111, 15480–5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Corradi N. 2015, Microsporidia: eukaryotic intracellular parasites shaped by gene loss and horizontal gene transfers. Annu. Rev. Microbiol., 69, 167–83. [DOI] [PubMed] [Google Scholar]
- 5. Peyretaillade E., Boucher D., Parisot N., et al. 2015, Exploiting the architecture and the features of the microsporidian genomes to investigate diversity and impact of these parasites on ecosystems. Heredity (Edinb), 114, 441–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Peyretaillade E., Parisot N., Polonais V., et al. 2012, Annotation of microsporidian genomes using transcriptional signals. Nat. Commun., 3, 1137. [DOI] [PubMed] [Google Scholar]
- 7. Corradi N., Slamovits C.H.. 2011, The intriguing nature of microsporidian genomes. Brief. Funct. Genomics, 10, 115–24. [DOI] [PubMed] [Google Scholar]
- 8. Cuomo C.A., Desjardins C.A., Bakowski M.A., et al. 2012, Microsporidian genome analysis reveals evolutionary strategies for obligate intracellular growth. Genome Res., 22, 2478–88. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Hannan K.M., Hannan R.D., Rothblum L. I.. 1998, Transcription by RNA polymerase I. Front. Biosci., 3, d376–98. [DOI] [PubMed] [Google Scholar]
- 10. Orioli A., Pascali C., Pagano A., Teichmann M., Dieci G.. 2012, RNA polymerase III transcription control elements: themes and variations. Gene, 493, 185–94. [DOI] [PubMed] [Google Scholar]
- 11. Arndt K.M., Reines D.. 2015, Termination of transcription of short noncoding RNAs by RNA polymerase II. Annu. Rev. Biochem., 84, 381–404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Amaral P.P., Dinger M.E., Mercer T.R., Mattick J.S.. 2008, The eukaryotic genome as an RNA machine. Science, 319, 1787–9. [DOI] [PubMed] [Google Scholar]
- 13. Huang B., Zhang R.. 2014, Regulatory non-coding RNAs: revolutionizing the RNA world. Mol. Biol. Rep., 41, 3915–23. [DOI] [PubMed] [Google Scholar]
- 14. Huttenhofer A., Schattner P., Polacek N.. 2005, Non-coding RNAs: hope or hype? Trends Genet., 21, 289–97. [DOI] [PubMed] [Google Scholar]
- 15. Matera A.G., Terns R.M., Terns M.P.. 2007, Non-coding RNAs: lessons from the small nuclear and small nucleolar RNAs. Nat. Rev. Mol. Cell. Biol., 8, 209–20. [DOI] [PubMed] [Google Scholar]
- 16. Washietl S., Will S., Hendrix D.A., et al. 2012, Computational analysis of noncoding RNAs. Wiley Interdiscip. Rev. RNA, 3, 759–78. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Sridhar J., Gunasekaran P.. 2013, Computational small RNA prediction in bacteria. Bioinform. Biol. Insights, 7, 83–95. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Thomarat F., Vivares C.P., Gouy M.. 2004, Phylogenetic analysis of the complete genome sequence of Encephalitozoon cuniculi supports the fungal origin of microsporidia and reveals a high frequency of fast-evolving genes. J. Mol. Evol., 59, 780–91. [DOI] [PubMed] [Google Scholar]
- 19. Veneziano D., Nigita G., Ferro A.. 2015, Computational approaches for the analysis of ncRNA through deep sequencing techniques. Front. Bioeng. Biotechnol., 3, 77. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Pombert J.F., Selman M., Burki F., et al. 2012, Gain and loss of multiple functionally related, horizontally transferred genes in the reduced genomes of two microsporidian parasites. Proc. Natl. Acad. Sci. USA, 109, 12638–43. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Belkorchia A., Gasc C., Polonais V., et al. 2015, The prediction and validation of small CDSs expand the gene repertoire of the smallest known eukaryotic genomes. PLoS One, 10, e0139075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Cornman R.S., Chen Y.P., Schatz M.C., et al. 2009, Genomic analyses of the microsporidian Nosema ceranae, an emergent pathogen of honey bees. PLoS Pathog., 5, e1000466. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Peyretaillade E., Goncalves O., Terrat S., et al. 2009, Identification of transcriptional signals in Encephalitozoon cuniculi widespread among Microsporidia phylum: support for accurate structural genome annotation. BMC Genomics, 10, 607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Sievers F., Wilm A., Dineen D., et al. 2011, Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol., 7, 539. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Pelin A., Selman M., Aris-Brosou S., Farinelli L., Corradi N.. 2015, Genome analyses suggest the presence of polyploidy and recent human-driven expansions in eight global populations of the honeybee pathogen Nosema ceranae. Environ. Microbiol., 17, 4443–58. [DOI] [PubMed] [Google Scholar]
- 26. Smith C., Heyne S., Richter A.S., Will S., Backofen R.. 2010, Freiburg RNA Tools: a web server integrating INTARNA, EXPARNA and LOCARNA. Nucleic Acids Res., 38, W373–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Nawrocki E.P., Burge S.W., Bateman A., et al. 2015, Rfam 12.0: updates to the RNA families database. Nucleic Acids Res., 43, D130–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Corpet F. 1988, Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res., 16, 10881–90. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Lestrade L., Weber M. J.. 2006, snoRNA-LBME-db, a comprehensive database of human H/ACA and C/D box snoRNAs. Nucleic Acids Res., 34, D158–62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Piekna-Przybylska D., Decatur W.A., Fournier M.J.. 2007, New bioinformatic tools for analysis of nucleotide modifications in eukaryotic rRNA. RNA, 13, 305–12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Bailey T.L., Boden M., Buske F.A., et al. 2009, MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res., 37, W202–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Peyret P., Katinka M.D., Duprat S., et al. 2001, Sequence and analysis of chromosome I of the amitochondriate intracellular parasite Encephalitozoon cuniculi (Microspora). Genome Res., 11, 198–207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Mei Q., Li X., Meng Y., et al. 2012, A facile and specific assay for quantifying microRNA by an optimized RT-qPCR approach. PLoS One, 7, e46890. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Parisot N., Denonfoux J., Dugat-Bony E., Peyret P., Peyretaillade E.. 2012, KASpOD: a web service for highly specific and explorative oligonucleotide design. Bioinformatics, 28, 3161–2. [DOI] [PubMed] [Google Scholar]
- 35. Sambrook J., Russell D.W.. 2006, The inoue method for preparation and transformation of competent E. coli: “ultra-competent” cells. CSH Protoc., 2006. [DOI] [PubMed] [Google Scholar]
- 36. Rosenblad M.A., Zwieb C., Samuelsson T.. 2004, Identification and comparative analysis of components from the signal recognition particle in protozoa and fungi. BMC Genomics, 5, 5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Willis I. M. 1993, RNA polymerase III. Genes, factors and transcriptional specificity. Eur. J. Biochem., 212, 1–11. [DOI] [PubMed] [Google Scholar]
- 38. Krol A., Westhof E., Bach M., Luhrmann R., Ebel J.P., Carbon P.. 1990, Solution structure of human U1 snRNA. Derivation of a possible three-dimensional model. Nucleic Acids Res., 18, 3803–11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Hudson A.J., Stark M.R., Fast N.M., Russell A.G., Rader S.D.. 2015, Splicing diversity revealed by reduced spliceosomes in C. merolae and other organisms. RNA Biol., 12, 1–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Lee R.C., Gill E.E., Roy S.W., Fast N.M.. 2010, Constrained intron structures in a microsporidian. Mol. Biol. Evol., 27, 1979–82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Davila Lopez M., Rosenblad M.A., Samuelsson T.. 2008, Computational screen for spliceosomal RNA genes aids in defining the phylogenetic distribution of major and minor spliceosomal components. Nucleic Acids Res., 36, 3001–10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Katinka M.D., Duprat S., Cornillot E., et al. 2001, Genome sequence and gene compaction of the eukaryote parasite Encephalitozoon cuniculi. Nature, 414, 450–3. [DOI] [PubMed] [Google Scholar]
- 43. Grisdale C.J., Bowers L.C., Didier E.S., Fast N.M.. 2013, Transcriptome analysis of the parasite Encephalitozoon cuniculi: an in-depth examination of pre-mRNA splicing in a reduced eukaryote. BMC Genomics, 14, 207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Gill E.E., Lee R.C., Corradi N., et al. 2010, Splicing and transcription differ between spore and intracellular life stages in the parasitic microsporidia. Mol. Biol. Evol., 27, 1579–84. [DOI] [PubMed] [Google Scholar]
- 45. Li X., Zaman S., Langdon Y., Zengel J.M., Lindahl L.. 2004, Identification of a functional core in the RNA component of RNase MRP of budding yeasts. Nucleic Acids Res., 32, 3703–11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Davila Lopez M., Rosenblad M.A., Samuelsson T.. 2009, Conserved and variable domains of RNase MRP RNA. RNA Biol., 6, 208–20. [DOI] [PubMed] [Google Scholar]
- 47. Cruz J. A., Westhof E.. 2011, Identification and annotation of noncoding RNAs in Saccharomycotina. C R Biol., 334, 671–8. [DOI] [PubMed] [Google Scholar]
- 48. Bachellerie J. P., Cavaille J., Huttenhofer A.. 2002, The expanding snoRNA world. Biochimie, 84, 775–90. [DOI] [PubMed] [Google Scholar]
- 49. Yu Y.T., Meier U.T.. 2014, RNA-guided isomerization of uridine to pseudouridine–pseudouridylation. RNA Biol., 11, 1483–94. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Schramm L., Hernandez N.. 2002, Recruitment of RNA polymerase III to its target promoters. Genes Dev., 16, 2593–620. [DOI] [PubMed] [Google Scholar]
- 51. Acker J., Conesa C., Lefebvre O.. 2013, Yeast RNA polymerase III transcription factors and effectors. Biochim. Biophys. Acta, 1829, 283–95. [DOI] [PubMed] [Google Scholar]
- 52. Braun B.R., Riggs D.L., Kassavetis G.A., Geiduschek E.P.. 1989, Multiple states of protein-DNA interaction in the assembly of transcription complexes on Saccharomyces cerevisiae 5S ribosomal RNA genes. Proc. Natl. Acad. Sci. USA, 86, 2530–4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Kassavetis G.A., Braun B.R., Nguyen L.H., Geiduschek E.P.. 1990, S. cerevisiae TFIIIB is the transcription initiation factor proper of RNA polymerase III, while TFIIIA and TFIIIC are assembly factors. Cell, 60, 235–45. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.