Abstract
The histopathological evaluation of morphological features in breast tumours provides prognostic information to guide therapy. Adjunct molecular analyses provide further diagnostic, prognostic and predictive information. However, there is limited knowledge of the molecular basis of morphological phenotypes in invasive breast cancer. This study integrated genomic, transcriptomic and protein data to provide a comprehensive molecular profiling of morphological features in breast cancer. Fifteen pathologists assessed 850 invasive breast cancer cases from The Cancer Genome Atlas (TCGA). Morphological features were significantly associated with genomic alteration, DNA methylation subtype, PAM50 and microRNA subtypes, proliferation scores, gene expression and/or reverse-phase protein assay subtype. Marked nuclear pleomorphism, necrosis, inflammation and a high mitotic count were associated with the basal-like subtype, and had a similar molecular basis. Omics-based signatures were constructed to predict morphological features. The association of morphology transcriptome signatures with overall survival in oestrogen receptor (ER)-positive and ER-negative breast cancer was first assessed by use of the Molecular Taxonomy of Breast Cancer International Consortium (METABRIC) dataset; signatures that remained prognostic in the METABRIC multivariate analysis were further evaluated in five additional datasets. The transcriptomic signature of poorly differentiated epithelial tubules was prognostic in ER-positive breast cancer. No signature was prognostic in ER-negative breast cancer. This study provided new insights into the molecular basis of breast cancer morphological phenotypes. The integration of morphological with molecular data has the potential to refine breast cancer classification, predict response to therapy, enhance our understanding of breast cancer biology, and improve clinical management. This work is publicly accessible at www.dx.ai/tcga_breast.
Keywords: PAM50, TCGA, bioinformatics, genomics, mRNA, epithelial tubule formation, histological grade
Introduction
Histopathological analysis of breast tumours plays a central role in the diagnosis of breast cancer. The assessment of histological type [e.g. invasive ductal carcinoma (IDC) or invasive lobular carcinoma (ILC)] and histological grade (a summary score of epithelial tubule formation, mitotic count, and nuclear pleomorphism) are reported to guide clinical management [1–4]. The microscopic assessment of tumour-infiltrating lymphocytes can predict improved response to chemotherapy and prognosis in erb-b2 receptor tyrosine kinase (HER2)-positive breast cancer [5–8]. Beyond these features, breast tumours show an array of other morphological features such as necrosis, the clinical significance of which is not well characterized.
Breast cancer is a heterogeneous disease at both the morphological and molecular levels. The PAM50 molecular ‘intrinsic’ subtypes, luminal A, luminal B, HER2-enriched, basal-like, and normal-like, have distinct biological properties, epidemiological risk factors, responses to therapy, and prognoses, and are associated with specific morphological features [9–13]. The normal-like subtype is highly variable and is not reproducibly defined [14]. Morphological and molecular data complement the characterization of breast cancer phenotypes. For example, basal-like tumours show high histological grade, necrosis, tumour-infiltrating lymphocytes, and fibrotic foci, and are generally IDCs [15–19], whereas HER2-enriched tumours show high histological grade, and may contain apocrine features and ductal carcinoma in situ (DCIS) [20,21].
Few studies have analysed the molecular characteristics of morphological features. These studies were limited by sample sizes (n = 57–212), and investigated one to three features with one or two types of molecular data [22–25]. The Genomic Grade Index (GGI; i.e. MapQuant Dx) is a transcriptomic signature constructed by integrating histological grade with gene expression, and is associated with oestrogen receptor (ER)-positive breast cancer prognosis [22]. GGI, like most first-generation prognostic signatures, is largely a measure of cellular proliferation [14,26,27]. The molecular bases of histological grade components, nuclear pleomorphism, epithelial tubule formation, and mitotic count, as well as other breast tumour morphological features, remain unknown.
This study aimed to comprehensively elucidate the molecular basis of breast cancer morphological phenotypes by integrating genomic, transcriptomic and proteomic data with morphological features, and to determine whether morphology transcriptomic signatures were prognostic in ER-positive or ER-negative breast cancer. To achieve this, a team of 15 international breast cancer pathology experts provided detailed histopathological annotation for 850 invasive breast cancer cases in The Cancer Genome Atlas (TCGA). After we had integrated the consensus assessments of 11 morphological features with TCGA’s molecular profiles, we identified genomic, transcriptomic and proteomic data associated with morphological features. Next, omics-based signatures representative of morphological features were constructed, and the prognostic value of each signature with overall survival was assessed by use of the Molecular Taxonomy of Breast Cancer International Consortium (METABRIC) database [28]. Signature(s) that remained prognostic in the METABRIC multivariate analysis were further evaluated in five additional datasets.
Materials and methods
Images and molecular data
TCGA data generation and processing were performed as previously described; samples were obtained from patients with appropriate consent from institutional review boards [29]. TCGA invasive breast cancer (n = 850) images were assessed via http://cancer.digitalslidearchive.net/[30]. Molecular profiles were retrieved (http://cancergenome.nih.gov/): RNAseq gene expression (Illumina HiSeq RNASeqV2 Level 3.1.9.0); DNA methylation subtypes 1, 2, 3, 4 and 5 (Illumina Infinium DNA chips); microRNA subtypes 1, 2, 3, 4, 5, 6 and 7 (Illumina sequencing); and reverse-phase protein assay (RPPA) subtypes basal, HER2-enriched, luminal A, luminal A/B, ReacI, ReacII and ‘X’ (MD Anderson RPPA Core Facility). PAM50 classification and PAM50 proliferation score were computed [9,31,32].
Genomic alterations implicated in breast cancer (43 somatic mutations, 45 amplifications, 62 deletions, and six multiple alterations, e.g. mutation and amplification) identified with Mutation Significance version 2 [33] and Genomic Identification of Significant Targets in Cancer [34] were retrieved from cBioPortal for Cancer Genomics [31]. Of a total of 156 genomic alterations, 127 genomic alterations were assessed in this study after exclusion of multiple alterations (n = 6) and rare genomic alterations that occurred in fewer than five cases (n = 23). These were excluded to allow separate analyses of mutation, amplification, and deletion, and reduce spurious findings.
Histopathology morphological assessment
Cases were randomly assigned to the pathologists, and images were graded by use of an electronic scoring sheet adapted from the College of American Pathologists’ protocol for invasive breast examination [35] (supplementary material, Figure S1A). For routine clinical features such as histological type, histological grade (nuclear pleomorphism, mitotic count, and epithelial tubule formation), lobular carcinoma in situ (LCIS), and DCIS, the pathologists used criteria applied in clinical practice. For features not commonly assessed in clinical practice, including stromal inflammation, necrosis, the proportion of cancerous epithelium in the invasive portion by area (i.e. epithelial area), lymphovascular invasion, stromal central fibrotic foci, and apocrine features, the pathologists carried out conference calls to discuss the grading criteria, and circulated images for scoring. Images with high consensus diagnoses were circulated as examples for grading. Supplementary material, Figure S1B shows an annotated scoring sheet with additional pathological scoring criteria and details.
To define the final histological type, information from pathology reports and the pathology review committee were integrated [31]. Pathology assessments were converted to integer scores. For the proportion of cancerous epithelium, in cases with discordance, discordant scores were resolved by taking the minimum value. For other morphological features, if the most frequent feature value in the dataset was the maximum of the possible feature values, discordant scores were resolved by taking the minimum value; otherwise, the maximum value was used. This was done to obtain an even distribution of the scores in the final dataset. Table 1 shows the morphological features, and their grading categories and frequencies.
Table 1.
Twelve morphological features graded by the international breast cancer pathology expert committee using clinical categories for 850 TCGA invasive breast cancer cases; detailed annotation for each TCGA case, consensus annotation, example images of each morphological feature and other details can be found at: www.dx.ai/tcga_breast
| Morphological feature | Clinical grading categories | All cases (n =850), n (%) | IDC (n =523), n (%) | ILC (n =117), n (%) | Special histological types (n =117), n (%) |
|---|---|---|---|---|---|
| Histological type | IDC | 523 (61.5) | |||
| ILC | 117 (13.8) | ||||
| Mixed (IDC/ILC) | 93 (10.9) | ||||
| Special types (others) | 117 (13.8) | ||||
| Histological grade | |||||
| Epithelial tubule formation | >75% (well differentiated) | 91 (11.0) | 61 (11.7) | 2 (1.8) | 23 (22.5) |
| 10–75% (moderately differentiated) | 161 (19.4) | 134 (25.6) | 5 (4.5) | 16 (15.7) | |
| <10% (poorly differentiated) | 576 (69.6) | 328 (62.7) | 104 (93.7) | 63 (61.8) | |
| Nuclear pleomorphism | Small regular nuclei | 67 (8.1) | 17 (3.3) | 33 (29.7) | 11 (10.8) |
| Moderate increase in size | 372 (44.9) | 200 (38.2) | 65 (58.6) | 57 (55.9) | |
| Moderate to marked variation in size | 389 (47.0) | 306 (58.5) | 13 (11.7) | 34 (33.3) | |
| Mitotic count | 0–5 per 10 HPFs (low) | 383 (46.7) | 172 (33.3) | 100 (90.1) | 54 (53.5) |
| 6–10 per 10 HPFs (medium) | 194 (23.6) | 146 (28.2) | 9 (8.1) | 18 (17.8) | |
| >10 per 10 HPFs (high) | 244 (29.7) | 199 (38.5) | 2 (1.8) | 29 (28.7) | |
| In situ cancer | |||||
| DCIS | Present | 376 (45.5) | 298 (57.0) | 5 (4.5) | 48 (47.1) |
| Absent | 450 (54.5) | 225 (43.0) | 105 (95.5) | 54 (52.9) | |
| LCIS | Present | 60 (7.3) | 8 (1.6) | 43 (39.1) | 2 (1.9) |
| Absent | 767 (92.7) | 506 (98.4) | 67 (60.9) | 101 (98.1) | |
| Other features | |||||
| Stromal inflammation | Present | 262 (31.8) | 207 (39.9) | 11 (9.9) | 22 (21.4) |
| Absent | 562 (68.2) | 312 (60.1) | 100 (90.1) | 81 (78.6) | |
| Necrosis | Present | 264 (32.0) | 217 (41.6) | 3 (2.7) | 30 (29.7) |
| Absent | 562 (68.0) | 305 (58.4) | 108 (97.3) | 71 (70.3) | |
| Proportion of cancerous epithelium in invasive portion by area (excluding areas of necrosis) | <25% (low) | 78 (9.5) | 29 (5.6) | 20 (18.2) | 13 (12.7) |
| 25–75% (moderate) | 506 (61.5) | 325 (62.5) | 76 (69.1) | 54 (52.9) | |
| >75% (high) | 239 (29.0) | 166 (31.9) | 14 (12.7) | 35 (34.3) | |
| Apocrine features | Absent | 669 (81.0) | 413 (79.0) | 98 (88.3) | 83 (83) |
| 1–5% (minimum) | 24 (2.9) | 16 (3.1) | 2 (1.8) | 3 (3.0) | |
| 6–50% (moderate) | 57 (6.9) | 36 (6.9) | 6 (5.4) | 6 (6.0) | |
| >50% (marked) | 76 (9.2) | 58 (11.1) | 5 (4.5) | 8 (8.0) | |
| Lymphovascular invasion | Present | 204 (24.8) | 142 (27.5) | 15 (13.6) | 24 (23.3) |
| Absent | 617 (75.2) | 375 (72.5) | 95 (86.4) | 79 (76.7) | |
| Stromal central fibrotic focus | Multiple fibrotic foci | 263 (32.1) | 184 (35.6) | 22 (19.8) | 21 (21.2) |
| Absent | 556 (67.9) | 333 (64.4) | 89 (80.2) | 78 (78.8) | |
DCIS, ductal carcinoma in situ; HPF, high-power field; IDC, invasive ductal carcinoma; ILC, invasive lobular carcinoma; LCIS, lobular carcinoma in situ; TCGA, The Cancer Genome Atlas.
% Cancerous Epithelium, Proportion of Cancerous Epithelium in Invasive Portion by Area (excluding Areas of Necrosis)
Inter-rater reliability
Inter-rater reliability was assessed for each morphological feature where cases were graded (using the categories shown in Table 1) by at least two pathologists. Inter-rater reliability was calculated according to Krippendorff’s alpha [36] (irr, version 0.84; R, version 3.2.1) with bootstrapping (100 iterations), and average percentage agreement.
Subsequent exclusion of histological type and re-stratification of morphological features into binary groups
The molecular characterization of histological types was reported separately, and we demonstrated that histological type represents a morphological continuum with a significant proportion of cases with morphological features of both ductal and lobular cancers [31]. Thus, we decided to include the full range of histological types in this article to enable the robust identification of molecular profiles and signatures associated with the remaining 11 morphological phenotypes across all types of invasive breast cancer. To reduce the complexity and to increase statistical power, morphological features were re-stratified into binary categories to determine the association of each morphological feature with a type of molecular data (Table 2). For example, we investigated the association of TP53 mutation in tumours with marked nuclear pleomorphism as compared with tumours with small/moderate nuclear pleomorphism, or genes differentially expressed in tumours with DCIS as compared with tumours without DCIS. All tests of statistical significance were two-sided. Statistical significance was achieved when the p-value was <0.05 or the false discovery rate (FDR) was <0.05.
Table 2.
The re-stratification of the 11 morphological features into binary grading levels for integration with molecular data
| Morphological features | Binary categories |
|---|---|
| Histological grade | |
| Epithelial tubule formation | >10% (well/moderately differentiated) |
| <10% (poorly differentiated) | |
| Nuclear pleomorphism | Small regular nuclei/moderate increase in size |
| Moderate to marked variation in size | |
| Mitotic count | 0–5 per 10 HPFs (low) |
| >6 per 10 HPFs (medium/high) | |
| In situ cancer | |
| DCIS | Present or absent |
| LCIS | Present or absent |
| Other features | |
| Stromal inflammation | Present or absent |
| Necrosis | Present or absent |
| % Cancerous epithelium | <75% (low/moderate) |
| >75% (high) | |
| Apocrine features | Absent/1–50% (minimum/moderate) |
| >50% (marked) | |
| Lymphovascular invasion | Present or absent |
| Stromal central fibrotic focus | Present or absent |
DCIS, ductal carcinoma in situ; HPF, high-power field; LCIS, lobular carcinoma in situ.
Determining the association of morphological features with molecular profiles
Genomic data
The univariate associations of genomic alteration and DNA methylation subtype with morphological features was determined with a chi-square test with Bonferroni adjustment, and Fisher’s exact test with Benjamini–Hochberg multiple testing corrections, respectively.
Transcriptomic data
The associations of PAM50 and microRNA subtypes, PAM50 proliferation score, differential gene expression and gene sets/pathways with morphological features were determined with a chi-square test with Bonferroni’s adjustment, Wilcoxon’s test, limmavoom with the Benjamini–Hochberg correction (version 3.22.1) [37], and piano (version 1.6.2) [38], respectively.
Differential gene expression (n = 15 398) was determined in all cases (i.e. overall, n = 826) and within each PAM50 subtype, except in the normal-like subtype (excluded because of small sample size, n = 24). Gene set enrichment analysis was performed with the C2 Molecular Signatures Database, which includes gene sets from Reactome, BioCarta, and KEGG (version 4.0, n = 4646; www.broadinstitute.org/gsea/msigdb/). Gene sets that were distinctly upregulated or downregulated were reported.
Proteomic data
The association of RPPA subtype with morphological features was determined with a chi-square test with Bonferroni adjustment.
Constructing molecular signatures of morphological features
Elastic-net regularized generalized linear models (glm-net, version 2.0-2) [39] was used to construct molecular signatures of morphological features according to genomic alteration, transcriptomic or both types of data (genomic and transcriptomic). Model performances were assessed according to the cross-validated area under the receiver operator characteristic curve (ROC AUC). To determine which type of molecular data best predicted morphological features, the ROC AUC of models built with genomic alterations, transcriptomic or both types of data were compared by use of a paired t-test.
Molecular signatures for histological grade were also constructed. A histological grade ‘feature’ was created by summing the original scores of epithelial tubule formation, nuclear pleomorphism, and mitotic count. The summed scores (ranging from 3 to 9) were then stratified into low/medium histological grade (summed scores of 3–7) or high histological grade (summed scores of 8 or 9).
Transcriptomic signatures predicted morphological features with the highest ROC AUCs. Thus, morphology transcriptomic signatures were subjected to bootstrapping (1000 iterations) to obtain 95% confidence intervals for gene coefficient estimates. Gene coefficients with 95% confidence intervals crossing zero were dropped from the signature. Gene set enrichment of each transcriptomic signature was performed with Gene Ontology Biological Processes summarized version (DAVID 6.7 [40]). Statistical significance was achieved when the FDR was <0.05.
Survival analyses performed with the METABRIC breast cancer dataset
To determine whether the morphological features’ transcriptomic signatures were prognostic for overall survival, these signatures were compared with established proliferation-based prognostic signatures (GGI [22], OncotypeDx [41], and MammaPrint [42]) and PAM50 subtype determined from the METABRIC (n = 1992) dataset [28].
PAM50 subtypes for METABRIC were retrieved from Prat et al. [43]. Research-based classifications of GGI, OncotypeDx and MammaPrint for each woman were computed with genefu (version 3.1) [44]. Each morphological feature’s signature score was calculated by subtracting the average expression of genes with negative coefficients from the average expression of genes with positive coefficients.
A Cox proportional hazards model was used to assess the univariate associations of clinicopathological variables [age at cancer diagnosis, tumour size (in centimetres), node-positive (spread to regional lymph nodes; yes/no) and clinical grade (1, 2 or 3)], PAM50, GGI, OncotypeDx, MammaPrint and morphology transcriptomic signatures with overall survival [45]. To ensure that the association of our morphology transcriptomic signatures with overall survival was not attributable to chance, the Significance Analysis of Prognostic Signatures (SAPS) algorithm was used to compare the prognostic utility of each morphology transcriptomic signature with ‘random’ transcriptomic signatures of similar size (saps, version 2.0.0) [46]. Hence, a morphology transcriptomic signature was only considered to be significant when the Cox model (Wald test) p-value was <0.05 and an absolute adjusted SAPS score of >1.3 was obtained.
Clinicopathological variables, PAM50, GGI, OncotypeDx and MammaPrint were considered to be significantly associated with survival when p-values were <0.05. Significant variables and/or signatures were subsequently evaluated in a multivariate model, adjusted by treatment (chemotherapy, hormone therapy, combined chemotherapy and hormone therapy, or untreated). Analyses were performed separately in ER-positive and ER-negative breast cancers.
Meta-analysis of significant transcriptomic signatures
The transcriptomic signature of poorly differentiated epithelial tubules remained significantly prognostic in the METABRIC multivariate analysis among ER-positive women. This signature was further evaluated in a meta-analysis consisting of five ER-positive breast cancer gene expression datasets: CAL [47], PNC [48], NKI [42,49], TRANSBIG [50], and GSE25066 [51]. Each gene expression dataset was pre-processed with Weighted Gene Co-Expression Network (version 1.47) [52], and annotated with lumi (version 2.20.2) [53]. These datasets were chosen because they had overall survival data or distant-relapse-free survival, clinical grade information, treatment information (for CAL, NKI, and GSE25066), and at least 10 000 annotated genes. The meta-analysis adjusted for clinical grade and treatment.
Website resource
Data are available at www.dx.ai/tcga_breast. Detailed methodologies are given in Supplementary materials and methods.
Results
Pathology morphological dataset and assessment of inter-rater reliability
From November 2011 to March 2014, 15 pathologists completed 1524 online scoring sheets: 11 cases were reviewed >10 times, 15 cases were reviewed five to nine times, 357 cases were reviewed two to four times, and 467 cases were reviewed once. The annotations and frequencies of morphological assessments are shown in Table 1. The prevalence rates of IDC, ILC and special histological types were similar to those in previous reports (IDC, 50–80%; ILC, 5–15%; special histological types, 1–15%). The proportion of mixed IDC/ILC cases (10.9%) in this study appear to be slightly higher than the 3–7% reported by a limited number of studies [54–59]. Supplementary material, Table S1A–D shows the frequencies stratified by PAM50 subtype for all cases, within IDCs, within ILCs or within special histological types. Raw annotation data are shown in supplementary material, Table S2. Inter-rater reliability was calculated for 383 cases that were reviewed at least twice. There was moderate agreement among pathologists, with percentage agreements ranging from 78% (mitotic count) to 98% (LCIS) (Table 3).
Table 3.
Inter-rater reliability on cases graded by at least two pathologists and Krippendorff’s alpha with bootstrap resampling and percentage agreement; each morphological feature’s grading categories are shown in Table 1
| Morphological feature | Cases (n) | Krippendorff’s alpha | Krippendorff’s alpha with bootstrap resampling (95% confidence interval) | Agreement (%) |
|---|---|---|---|---|
| Histological type | 358 | 0.471 | 0.472 (0.402–0.532) | 85.6 |
| Histological grade | ||||
| Epithelial tubule formation | 316 | 0.544 | 0.547 (0.463–0.621) | 87.4 |
| Nuclear pleomorphism | 318 | 0.522 | 0.520 (0.457–0.590) | 80.8 |
| Mitotic count | 311 | 0.488 | 0.493 (0.421–0.576) | 77.7 |
| In situ cancer | ||||
| Ductal carcinoma in situ | 317 | 0.526 | 0.521 (0.451–0.592) | 89.0 |
| Lobular carcinoma in situ | 317 | 0.298 | 0.303 (0.088–0.507) | 97.5 |
| Other features | ||||
| Stromal inflammation | 315 | 0.544 | 0.534 (0.442–0.593) | 89.8 |
| Necrosis | 317 | 0.591 | 0.581 (0.474–0.669) | 90.6 |
| Proportion of cancerous epithelium in invasive portion by area (excluding areas of necrosis) | 312 | 0.472 | 0.467 (0.387–0.538) | 79.2 |
| Apocrine features | 314 | 0.164 | 0.189 (0.076–0.318) | 90.3 |
| Lymphovascular invasion | 312 | 0.423 | 0.413 (0.327–0.515) | 90.1 |
| Stromal central fibrotic focus | 311 | 0.256 | 0.262 (0.155–0.367) | 82.7 |
Morphological features are associated with molecular data
Table 4 summarizes the associations of each morphological feature with various molecular data (details are shown in supplementary material, Table S3 and Figure S2). Differential gene expression was performed in all cases and within each PAM50 subtype, except in the normal-like subtype (supplementary material, Table S4A). Owing to small sample sizes, differential gene expression associated with the presence of LCIS was performed for all cases and within luminal A cases.
Table 4.
An overview of molecular data significantly associated with morphological features
| Stromal inflammation, necrosis, nuclear pleomorphism, and mitotic count |
The presence of necrosis and inflammation, a medium/high mitotic count and marked nuclear pleomorphism was associated with the following:
|
| Epithelial tubule formation |
Poorly differentiated epithelial tubules were associated with:
|
| LCIS |
The presence of LCIS was associated with:
|
| DCIS |
| Tumours with DCIS were enriched for proliferation gene sets Upregulated genes in DCIS are linked to the breast microenvironment, especially myoepithelial cells |
| Apocrine features |
The presence of marked apocrine features was associated with:
|
| Lymphovascular invasion |
The lymphovascular invasion feature remains mainly morphological but was also associated with:
|
| Stromal central fibrotic foci |
| The presence of fibrotic foci remains mainly morphological Fibrotic focus was not associated with any gene, but showed downregulated inflammation gene sets |
| Proportion of cancerous epithelium in invasive portion by area (excluding areas of necrosis) |
A high proportion of cancerous epithelium was associated with:
|
chr, chromosome; DCIS, ductal carcinoma in situ; IL, interleukin; LCIS, lobular carcinoma in situ.
Inflammation, necrosis, nuclear pleomorphism, and mitotic count
Inflammation, necrosis, marked nuclear pleomorphism and medium/high mitotic counts co-occurred in the tumours and shared many genomic alterations (FDR of <0.05; Figure 1, Table 4). In the TCGA publication, DNA methylation subtypes 3 and 5 are enriched for luminal B and basal-like, respectively, whereas TCGA microRNA subtypes 4 and 5 are associated with basal-like [29]. Therefore, the presence of inflammation and necrosis, marked nuclear pleomorphism and medium/high mitotic counts are distinctly associated with the highly proliferative basal-like subtype (Table 4).
Figure 1.
Heatmap and unsupervised hierarchical clustering of the 38 significant genomic alterations and 11 morphological features based on the degree and direction of the associations. Inflammation and necrosis, marked nuclear pleomorphism and medium/high mitotic counts are clustered together, as they share many common genomic alterations.
These four morphological features had 15 common upregulated genes involved in cellular proliferation (MYBL2, CHEK1, CENPA, MELK, MEMO1, NASP, RCC2, LTV1, and C1orf135) [60,61], MYC activation (CDCA7) [62–65], and DNA and RNA metabolism (PIF1, RBM17, AMD1, and RPIA). The function of C17orf96 is unknown. These four morphological features also had 13 common downregulated genes involved in membrane signalling (CBLN4, ELFN1, LTBP3, LRP10, TENC1, and TPCN1), including GTPase activity (RAPGEF3 and TBC1D13), transcription (CAMTA2, CRY2, and LOC653501), the cytoskeleton (KIF13B), and lysosome positioning (C10orf32). The plethora of differentially expressed genes associated with necrosis, marked nuclear pleomorphism and medium/high mitotic counts were enriched for proliferation gene sets, whereas the presence of stromal inflammation was enriched for inflammation gene signatures (FDR of <0.05; Figure 2; a detailed heatmap is shown in supplementary material, Figure S3). Collectively, molecular data suggest that tumorigenesis involving these four proliferative basal-like morphological features may be driven by MYB-regulated and MYC-regulated pathways, and, potentially in conjunction with TP53 pathways, in invasive breast cancer [60,66,67].
Figure 2.
Heatmap summarizing the FDRs of 485 significant pathways and unsupervised hierarchical clustering of morphological features. Features are clustered into two groups characterized mainly by proliferation and inflammation. Detailed pathways are presented in supplementary material, Figure S3. The proliferation cluster had increased cell proliferation and metabolism, and decreased inflammation and membrane receptor signalling. The inflammation cluster comprised largely immune-related signatures.
Epithelial tubule formation
Other studies reported the association of TP53, 8q24.21 (MYC), 19q12 (CCNE1), 20p13.2 (ZNF217) and 9p21.3 (MTAP) with histologic grade [24,25]. However, when focussing on the individual components of histological grade, poorly differentiated epithelial tubules shared only a few molecular traits with medium/high mitotic counts and marked nuclear pleomorphism: TP53 mutation, high PAM50 proliferative score, basal-like subtype classified according to methylation, and microRNA data. The molecular traits of poorly differentiated epithelial tubules were common with those of LCIS (i.e. CDH1 mutation, PAM50 luminal A subtype, and inflammation gene sets), although there was no correlation between the two morphological features (supplementary material, Table S3B). P2RY11, which encodes a G-protein-coupled receptor activated by extracellular adenosine and uridine [68], was the top differentially expressed gene (2.3-fold increase) in tumours with poorly differentiated epithelial tubules as compared with well-differentiated/moderately differentiated epithelial tubules (supplementary material, Table S4B). The role of P2RY11 in breast tissue remains unknown, and could be evaluated as a potential pharmacological target.
LCIS
LCISs are precursor lesions for ILCs, defined by the hallmark CDH1 loss-of-function mutation, and are almost exclusive to luminal A tumours [31,69]. Regardless of histological type, the presence of LCIS was also associated with DNA methylation subtype 1, downregulation of proliferation gene sets, and enrichment for cytokines/immune-signalling pathways (despite not being associated with the presence of morphological inflammation). The expression of HMGCS2, a breast apocrine carcinoma marker involved in the anabolic ketogenesis pathway, was increased by 8.6-fold in tumours with LCIS (supplementary material, Table S4B) [70]. Other top-ranking upregulated genes were those involved in inflammation (GP2 and C7) [71], alcohol metabolism (ADH1B), fat metabolism (ADIPOQ) [72,73], transcription (TFAP2B), transmembrane proteins (TMEM132C and SLC7A4), and genes with unknown function (TFF1 and TUSC5) [74,75]. At the same time, the mRNA expression levels of extracellular matrix proteins (MMP1, CDH1, EPYC, COL11A1, HAPLN1, and IBSP) were significantly lower. Thus, APIPOQ and HMGCS2 overexpression suggest that the manifestation of LCIS may reflect abnormal hormone and fatty acid levels in the breast tissues, impaired fatty acid oxidation, and mitochondrial dysfunction [76,77]. Mitochondrial dysfunction can lead to inflammation, tumorigenesis, dysregulation of cell–cell adhesion, discohesive morphology, and invasion [76,78–81]. These characteristics of mitochondrial dysregulation are supported by our differential gene expression analyses. It would also be interesting to investigate the association of lifestyle factors such as obesity or alcohol consumption with LCIS or histological type [72,82,83].
DCIS
The co-existence of DCIS with prominent features (i.e. strong molecular profiles) such as marked nuclear pleomorphism and poorly differentiated epithelial tubules may have masked our ability to decipher the molecular basis of DCIS (supplementary material, Table S3B). DCIS was associated with 40 differentially expressed genes, and was enriched for proliferation and cell–cell junction pathways (supplementary material, Figure S3). The top-ranking upregulated genes in breast cancers associated with DCIS were those encoding epithelial proteins (CALML3, ANXA8L1, and ANXA8) [84,85], extracellular matrix proteins (KRT14, KRT6B, KRT17, and MMP10) [86], desmosomes (DSG3 and DSC3) that connect adjacent myoepithelial cells [87], and proteins involved in myoepithelial cell differentiation (ACTA1) [88] and CCL21-related chemotaxis resulting in epithelial–mesenchymal transition and metastasis [89–91]. These results support reports that the progression of DCIS to invasive breast cancer is influenced by changes in microenvironmental factors, especially in myoepithelial cells [87,92]. Proliferating cancerous ductal cells exert pressure against the myoepithelial cells and basement membrane. When the myoepithelial cells cannot sustain the pressure and rate of basement membrane turnover, they lose their cell–cell adhesion capabilities and allow the cancerous cells to invade into the surrounding tissues [87]. The 11 downregulated genes in breast cancers associated with DCIS are newly associated with breast cancer [cytoskeleton-related (HOOK2 and ARHGEF18), mitochondrial iron–sulphur cluster assembly pathway (C1orf69), gene regulation (MAFG and WDR37), GTPase activity (TBC1D13 and RAB43), lipid synthesis (CLN8), and neuronal components (PRX, LOC100130093 and OPA3); supplementary material, Table S4B]. Their involvement in DCIS and/or invasive breast cancer warrants further elucidation.
Apocrine features
This is the first study to characterize the molecular basis of apocrine features (Table 4). Upregulated genes and enriched pathways associated with marked apocrine features include increased lipid and membrane transport (ABCC2, ABCA12, ABCC11, HAPLN1, FIBCD1, and FAM155B), lipid and/or cell metabolism (DHRS2, HGD, IYD, and HHIPL2), apoptotic, diabetes and cholesterol pathways. Downregulated genes and pathways with marked apocrine features were those encoding gastropeptides (NPYIR and PI16) and serine peptidase (KLK11), and those involved in alcohol/drug metabolism (ADH1B and CYP4F22), secretion (AQP5), the extracellular matrix (HAPLN1), and cytokine signalling (C7 and DARC). These genes have been investigated as markers of proliferation or metastasis [93–99], breast cancer risk [100], prognosis [101,102], and response to therapy [103–106]. Our work suggests that drug resistance may occur in tumours with marked apocrine features with overexpression of ATP-binding cassette transporter mRNA, and new drugs targeting aquaporin water channels may not work in these tumours [103,107].
Lymphovascular invasion and fibrotic foci
Neither genomic alteration nor PAM50 subtype was associated with the presence of lymphovascular invasion [25], fibrotic foci, or a high proportion of cancerous epithelium. Interleukin-12-related and integrin-related neutrophil pathways and extracellular matrix organization gene sets were downregulated in the presence of lymphovascular invasion. The presence of fibrotic foci was linked to upregulated integrin and extracellular matrix organization gene sets, and down-regulated inflammation gene sets. The lack of distinct molecular profiles for lymphovascular invasion and fibrotic foci may be attributable to their low frequencies, and suggests that these features remain largely morphological.
Transcriptomic signatures of morphological features
Genomic alterations, gene expression or both data types were used to construct signatures of morphological features. The ROC AUCs of multivariate models built with transcriptomic and combined data outperformed models constructed with genomic alterations (p ≤ 0.001; supplementary material, Table S5A). There was no difference in the ROC AUCs between transcriptomic and combined data, indicating that the addition of genomic alteration data did not enhance the performance of transcriptomic signatures in predicting morphological features (p = 0.139; supplementary material, Table S5B). Thus, only transcriptomic signatures were subjected to bootstrapping and further explored. The transcriptomic signatures of morphological features ranged from one gene (LRRC32) for the proportion of cancerous epithelium to 110 genes for poorly differentiated epithelial tubules (supplementary material, Table S5C, D).
The stromal inflammation signature is enriched for the suppression of T-cell activation, driven by its strongest (positive) coefficient, CTLA4. The increase in the level of CTLA4 in breast cancer prevents the anti-tumour T-cell response [108]. Its monoclonal antibody, anti-CTLA4, when used in synergy with other therapeutic agents (e.g. trastuzumab), blocks immune checkpoints, and induces anti-tumour immunity, resulting in tumour regression in preclinical (HER2) breast cancer models [109–112]. However, the blocking of immune checkpoints with antibodies against programmed cell death protein 1 and its ligand is more effective than using anti-CTLA4 [109,113,114]. Future work could evaluate whether this inflammation signature can identify women who may benefit from anti-CTLA4 therapy, as well as investigating how CTLA4 contributes to tumour immunity [115].
Signatures for medium/high mitotic count, marked nuclear pleomorphism and high histological grade were enriched for cell proliferation, further confirming that these features are proliferation-related (supplementary material, Table S5E). No enrichment was obtained for other signatures.
The epithelial tubule formation transcriptomic signature was prognostic in ER-positive breast cancer
In METABRIC ER-positive women (n = 1494), age at cancer diagnosis, tumour size, node-positive status and the transcriptomic signature for poorly differentiated epithelial tubules remained prognostic in the multivariate model (p < 0.05; Table 5). In ER-negative women (n = 434), no feature was prognostic in the multivariate model.
Table 5.
The variables significantly associated with overall survival in METABRIC oestrogen receptor (ER)-positive and ER-negative women in the univariate and subsequent multivariate analyses
| Type of variable | Univariate analysis
|
Multivariate analysis
|
|||||||
|---|---|---|---|---|---|---|---|---|---|
| Hazard ratio (eb) | 95% CI for hazard ratio
|
p-value | Hazard ratio (eb) | 95% CI for hazard ratio
|
p-value | ||||
| Lower | Upper | Lower | Upper | ||||||
| ER-positive breast cancer | |||||||||
| Age at initial pathological diagnosis | Clinicopathological | 1.048 | 1.040 | 1.056 | <1.00E-16 | 1.046 | 1.037 | 1.055 | <1.00E-16 |
| Tumour size | Clinicopathological | 1.217 | 1.172 | 1.264 | <1.00E-16 | 1.148 | 1.098 | 1.199 | 1.03E-09 |
| Node-positive | Clinicopathological | 1.916 | 1.642 | 2.235 | 1.11E-16 | 1.534 | 1.257 | 1.872 | 2.50E-05 |
| Clinical grade (METABRIC) | Clinicopathological | 1.340 | 1.182 | 1.519 | 4.85E-06 | – | – | – | – |
| Genomic Grade Index | Established signature | 1.639 | 1.406 | 1.911 | 2.64E-10 | – | – | – | – |
| OncotypeDx | Established signature | 1.619 | 1.309 | 2.002 | 8.86E-06 | – | – | – | – |
| MammaPrint | Established signature | 1.421 | 1.191 | 1.695 | 9.67E-05 | – | – | – | – |
| Marked nuclear pleomorphism | Transcriptomic signature | 1.334 | 1.231 | 1.446 | 2.41E-12 | – | – | – | – |
| High histological grade | Transcriptomic signature | 1.311 | 1.211 | 1.420 | 2.08E-11 | – | – | – | – |
| Medium/high mitotic count | Transcriptomic signature | 1.339 | 1.229 | 1.460 | 2.93E-11 | – | – | – | – |
| Poorly differentiated epithelial tubules | Transcriptomic signature | 1.907 | 1.554 | 2.338 | 5.83E-10 | 1.308 | 1.005 | 1.703 | 4.57E-02 |
| Necrosis | Transcriptomic signature | 1.348 | 1.173 | 1.548 | 2.54E-05 | – | – | – | – |
| HER2-enriched | PAM50 | 1.423 | 1.141 | 1.776 | 1.75E-03 | – | – | – | – |
| Luminal A | PAM50 | 0.787 | 0.670 | 0.925 | 3.60E-03 | – | – | – | – |
| Basal-like | PAM50 | 1.511 | 1.138 | 2.006 | 4.36E-03 | – | – | – | – |
| Luminal B | PAM50 | 1.252 | 1.064 | 1.473 | 6.72E-03 | – | – | – | – |
| Normal-like | PAM50 | 0.638 | 0.503 | 0.811 | 2.32E-04 | – | – | – | – |
| ER-negative breast cancer | |||||||||
| Node-positive | Clinicopathological | 1.628 | 1.229 | 2.156 | 6.69E-04 | – | – | – | – |
| Tumour size | Clinicopathological | 1.084 | 1.023 | 1.149 | 6.03E-03 | – | – | – | – |
CI, confidence interval; METABRIC, Molecular Taxonomy of Breast Cancer International Consortium.
Clinicopathological variables were obtained from METABRIC. Transcriptomic signatures for morphological features were developed in this study. Research-based classifications of Genomic Grade Index, OncotypeDx and MammaPrint (i.e. established signatures) were computed with genefu.
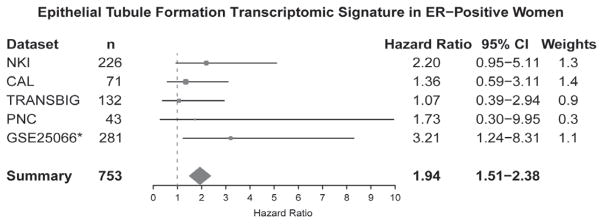
The transcriptomic signature for poorly differentiated epithelial tubules in ER-positive women was further evaluated in a meta-analysis across five publicly available gene expression datasets (Figure). The summary hazard ratio was 1.94 (95% confidence interval 1.51–2.38).
The epithelial tubule formation transcriptomic signature is distinct and least correlated with proliferation
The transcriptomic signature for poorly differentiated epithelial tubules was distinct from the signatures for medium/high mitotic count, marked nuclear pleomorphism, and high histological grade (Figure 4). To determine whether the transcriptomic signature for poorly differentiated epithelial tubules was the least correlated with proliferation, the PAM50 proliferation score for each woman in the METABRIC dataset was calculated and correlated with the transcriptomic signature scores of nuclear pleomorphism, mitotic count, and epithelial tubule formation. PAM50 proliferation scores were more highly correlated with medium/high mitotic count [Spearman’s rho = 0.878 (ER-positive) and Spearman’s rho = 0.919 (ER-negative)] and marked nuclear pleomorphism (rho = 0.852 and rho = 0.904) than with poorly differentiated epithelial tubules (rho = 0.351 and rho = 0.616) in ER-positive and ER-negative invasive breast cancers (p < 0.001).
Figure 4.
(A) There were 17 overlapping genes between the transcriptomic signatures for medium/high mitotic count and marked nuclear pleomorphism, whereas genes predictive of poorly differentiated epithelial tubules were distinct. (B) Most of the genes in the transcriptomic signature for high histological grade were common to the signatures for medium/high mitotic count and marked nuclear pleomorphism, but were distinct from of poorly differentiated epithelial tubules.
Discussion
Little is known about the molecular characteristics of various morphological features in invasive breast cancer. We comprehensively unravelled the molecular portraits of breast cancer histopathological phenotypes by bridging histopathological annotations with the molecular profiles in the TCGA database. This article represents the largest cross-section of cases and pathologists to examine breast cancer histopathological phenotypes to date. Our data support the central role of proliferation driving histological grade. Inflammation, necrosis, medium/high mitotic count and marked nuclear pleomorphism frequently co-exist in breast tumours, are associated with basal-like subtypes, and have similar molecular bases. LCIS has a distinct molecular profile that may be linked to mitochondrial dysfunction, whereas genes that are differentially expressed in DCIS are intimately associated with myoepithelial cells. Lymphovascular invasion and fibrotic foci are mainly morphological, with few significant molecular traits.
Some morphological features harbour molecular traits that may confer drug resistance or serve as pharmacological targets. Our signatures can act as surrogate representation of morphological features, enabling future studies to link the signatures to response to therapy, with the long-term aim of improving clinical management. Personalized or refined breast cancer classification can be achieved by combining the observation of morphological features with molecular and immunohistochemistry data. Collectively, this study provides new insights into the molecular basis of breast cancer morphological phenotypes, and could potentially facilitate the future development of diagnostic and prognostic tools for breast cancer.
Most databases, including METABRIC, usually provide information on histological grade but not its components such as epithelial tubule formation. We were unable to directly determine whether the pathological measure of epithelial tubules is independently prognostic, or whether our transcriptomic signature of epithelial tubule formation adds prognostic information or is superior to pathological assessment. However, if high histological grade can function as a surrogate for poorly differentiated epithelial tubules, our multivariate analyses show that the epithelial tubule formation signature is more prognostic than clinical histological grade, and indirectly demonstrate that our signature adds prognostic information for ER-positive breast cancer. Nevertheless, prognostic signatures for ER-positive breast cancer are well established [22,41,42], and more research is needed to discover clinically useful prognostic signatures for ER-negative breast cancer.
At the molecular level, epithelial tubule formation is least similar to mitotic count and nuclear pleomorphism, and shares traits with LCIS and inflammation. The transcriptomic signature for poorly differentiated epithelial tubules is distinct from high histological grade, but not significantly enriched for any gene sets. The signature’s genes are involved in proliferation, mitochondrial metabolism, membrane signalling, cellular adhesion, oxidative stress, extracellular matrix organization, and inflammation. These gene functions are a mix of selective molecular traits associated with medium/high mitotic count, marked nuclear pleomorphism, LCIS, and inflammation. We speculate that our transcriptomic signature for poorly differentiated epithelial tubules is unique and prognostically superior, because it contains genes that represent a wide range of tumour biology.
The failure to detect any association of DCIS, fibrotic foci or apocrine features with PAM50 subtypes may be attributable to our study utilizing PAM50 classification by molecular data instead of immunohistochemistry, using different grading criteria, and investigating these features within tumours of invasive breast cancer [116–119]. Our transcriptomic signature for fibrotic foci was not prognostic, despite previous studies reporting that IDCs or luminal B tumours with fibrotic foci have a poorer prognosis [120,121]. The relevance of fibrotic foci as a prognostic factor requires further investigation, which should take into consideration its size, breast cancer histological type, and PAM50 classification.
Fourteen TCGA cases were inadequate for scoring, owing to poor image quality or insufficient invasive cancer being present. Despite adherence to clinical definitions or agreed consensus scoring criteria, our histopathological analyses may be influenced by the variation in histology quality and the use of images instead of slides for scoring. For example, the high-power field used to count mitotic bodies at the highest magnification (×40) on a web browser is infuenced by computer monitor size (hence, the high-power field for each pathologist varies) and the difficulty in distinguishing between mitotic figures from pynknotic nuclei, owing to the lack of a Z-axis. The pathologists used their best judgement in counting cells in mitosis. However, the mitotic count (as it was scored) was highly concordant with the PAM50 proliferation score and enrichment for proliferation gene sets in this study, signifying that both mitotic count and gene expression were adequately tracking proliferation. Another limitation of this study is that we focused exclusively on a set of known morphological features that could be scored manually by experienced breast pathologists. It is likely that there are additional morphological patterns (e.g. various types of stromal reaction pattern) beyond those included in our study that are biologically important and will provide additional insights into the molecular underpinnings of breast cancer pathology.
In conclusion, breast tumour pathological phenotypes are driven by distinct underlying sets of molecular alterations. The integration of morphological with molecular data has great potential to refine breast cancer classification, predict response to therapy, enhance our understanding of breast cancer biology, and improve clinical management.
Supplementary Material
Images of the electronic scoring sheets: (A) used by the pathologists to annotate images; (B) annotated scoring sheet with pathological scoring criteria and details.
Boxplots displaying PAM50 proliferation scores for each morphological feature.
Expanded heatmap of Figure 2, with detailed gene sets.
Tables summarising the distribution of morphological features within each PAM50 subtype for (A) all cases; (B) invasive ductal cases; (C) invasive lobular cases; and (D) special histological types.
Detailed data for morphological features, PAM50, DNA methylation, miRNA and RPPA subtypes, PAM50 proliferation scores and genomic alterations for 850 TCGA cases.
(A) The association of molecular subtypes PAM50, DNA methylation, RPPA, and microRNA with morphological features. (B) Association of morphological features with each other.
(A) The frequency of each morphological feature in binary groups and the number of differentially expressed genes, stratified by PAM50 subtype, excluding Normal-like. (B) Top 10 up- or down-regulated differentially-expressed genes (limma FDR<0.05) for each morphological feature, ranked by Log2 fold change (Log2FC).
(A) Multivariate ROCAUC performances of genomic alterations, gene expression and combined data. (B) Paired t-test of ROCAUC of genomic alterations, gene expression and combined data models. (C) Summary of transcriptomic signatures and their enrichments. (D) Transcriptomic signatures of morphological features. (E) Gene set enrichments for transcriptomic signatures of morphological feature.
Figure 3.
The prognostic significance of the transcriptomic signature for poorly differentiated epithelial tubules in ER-positive women was further validated in a meta-analysis across five cohorts. *The endpoint for GSE25066 is distant relapse-free survival; the endpoints for all other datasets are overall survival. The summary hazard ratio estimate is a weighted average. Weights are the reciprocal of the estimated variance (square of standard error for the analysis). CI, confidence interval.
Acknowledgments
The data used in this study were in whole or in part based on the data generated by the TCGA Research Network: http://cancergenome.nih.gov/. Funding for this project was provided by the Klarman Family Foundation (AHB), the National Cancer Institute of the National Institutes of Health (SPORE grant P50CA168504; AHB), and the National Library of Medicine of the National Institutes of Health Career Development Award (Number K22LM011931; AHB).
Footnotes
Conflicts of interest statement CMP is an equity stock holder and Board of Director Member of BioClassifier. CMP is also listed an inventor on patent applications on the Breast PAM50 assay. AHB is an equity stock holder and Board of Director Member of PathAI. The remaining authors declare no competing financial interests.
Author contributions statement
The authors contributed in the following way: AHB, CMP: conception and design of the study; AHB, SCL, GMKT, REF, KHA, LCC, YC, KCJ, NBJ, SJS, KS, GK, DAE, PHT, KK, FMW, JSRF: provided pathology annotations and clinical input; YJH, AHB, BHK, DMAG, GMC, GC, KAH, CMP: retrieved and acquired other relevant data; YJH, AHB: analysed and interpreted the data; JCJ, SRF, RP, DMAG, BHK, GMC, DAG: provided website and/or database support; YJH, AHB, CMP, SCL, GMKT, DMAG, BHK, KCJ, MS, JSRF, PHT, SJS, DAE, REF, KAH, KK: involved in the writing, reviewing and revision of the manuscript.
References
- 1.Elston CW, Elston CW, Ellis IO, et al. Pathological prognostic factors in breast cancer. I. The value of histological grade in breast cancer: experience from a large study with long-term follow-up. Histopathology. 1991;19:403–410. doi: 10.1111/j.1365-2559.1991.tb00229.x. [DOI] [PubMed] [Google Scholar]
- 2.Galea MH, Blamey RW, Elston CE, et al. The Nottingham prognostic index in primary breast cancer. Breast Cancer Res Treat. 1992;22:207–219. doi: 10.1007/BF01840834. [DOI] [PubMed] [Google Scholar]
- 3.Loi S, Sirtaine N, Piette F, et al. Prognostic and predictive value of tumor-infiltrating lymphocytes in a phase III randomized adjuvant breast cancer trial in node-positive breast cancer comparing the addition of docetaxel to doxorubicin with doxorubicin-based chemotherapy: BIG 02-98. J Clin Oncol. 2013;31:860–867. doi: 10.1200/JCO.2011.41.0902. [DOI] [PubMed] [Google Scholar]
- 4.Rakha EA, Reis-Filho JS, Baehner F, et al. Breast cancer prognostic classification in the molecular era: the role of histological grade. Breast Cancer Res. 2010;12:207. doi: 10.1186/bcr2607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Salgado R, Denkert C, Campbell C, et al. Tumor-infiltrating lymphocytes and associations with pathological complete response and event-free survival in HER2-positive early-stage breast cancer treated with lapatinib and trastuzumab: a secondary analysis of the NeoALTTO Trial. JAMA Oncol. 2015;1:448–454. doi: 10.1001/jamaoncol.2015.0830. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Adams S, Gray RJ, Demaria S, et al. Prognostic value of tumor-infiltrating lymphocytes in triple-negative breast cancers from two phase III randomized adjuvant breast cancer trials: ECOG 2197 and ECOG 1199. J Clin Oncol. 2014;32:2959–2966. doi: 10.1200/JCO.2013.55.0491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Gentles AJ, Newman AM, Liu CL, et al. The prognostic landscape of genes and infiltrating immune cells across human cancers. Nat Med. 2015;21:938–945. doi: 10.1038/nm.3909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Denkert C, Loibl S, Noske A, et al. Tumor-associated lymphocytes as an independent predictor of response to neoadjuvant chemotherapy in breast cancer. J Clin Oncol. 2010;28:105–113. doi: 10.1200/JCO.2009.23.7370. [DOI] [PubMed] [Google Scholar]
- 9.Parker JS, Mullins M, Cheung MCU, et al. Supervised risk predictor of breast cancer based on intrinsic subtypes. J Clin Oncol. 2009;27:1160–1167. doi: 10.1200/JCO.2008.18.1370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Sorlie T, Perou CM, Tibshirani R, et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci USA. 2001;98:10869–10874. doi: 10.1073/pnas.191367098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Millikan RC, Newman B, Tse CK, et al. Epidemiology of basal-like breast cancer. Breast Cancer Res Treat. 2008;109:123–139. doi: 10.1007/s10549-007-9632-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Chia SK, Bramwell VH, Tu D, et al. A 50-gene intrinsic subtype classifier for prognosis and prediction of benefit from adjuvant tamoxifen. Clin Cancer Res. 2012;18:4465–4472. doi: 10.1158/1078-0432.CCR-12-0286. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Perou CM, Sørlie T, Eisen MB, et al. Molecular portraits of human breast tumours. Nature. 2000;406:747–752. doi: 10.1038/35021093. [DOI] [PubMed] [Google Scholar]
- 14.Wirapati P, Sotiriou C, Kunkel S, et al. Meta-analysis of gene expression profiles in breast cancer: toward a unified understanding of breast cancer subtyping and prognosis signatures. Breast Cancer Res. 2008;10:R65. doi: 10.1186/bcr2124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Livasy CA, Karaca G, Nanda R, et al. Phenotypic evaluation of the basal-like subtype of invasive breast carcinoma. Mod Pathol. 2006;19:264–271. doi: 10.1038/modpathol.3800528. [DOI] [PubMed] [Google Scholar]
- 16.Fulford LG, Easton DF, Reis-Filho JS, et al. Specific morphological features predictive for the basal phenotype in grade 3 invasive ductal carcinoma of breast. Histopathology. 2006;49:22–34. doi: 10.1111/j.1365-2559.2006.02453.x. [DOI] [PubMed] [Google Scholar]
- 17.Carey LA, Perou CM, Livasy CA, et al. Race, breast cancer subtypes, and survival in the Carolina Breast Cancer Study. JAMA. 2006;295:2492–2502. doi: 10.1001/jama.295.21.2492. [DOI] [PubMed] [Google Scholar]
- 18.Dabbs DJ, Chivukula M, Carter G, et al. Basal phenotype of ductal carcinoma in situ: recognition and immunohistologic profile. Mod Pathol. 2006;19:1506–1511. doi: 10.1038/modpathol.3800678. [DOI] [PubMed] [Google Scholar]
- 19.Reis-Filho JS, Milanezi F, Steele D, et al. Metaplastic breast carcinomas are basal-like tumours. Histopathology. 2006;49:10–21. doi: 10.1111/j.1365-2559.2006.02467.x. [DOI] [PubMed] [Google Scholar]
- 20.Tamimi RM, Baer HJ, Marotti J, et al. Comparison of molecular phenotypes of ductal carcinoma in situ and invasive breast cancer. Breast Cancer Res. 2008;10:R67. doi: 10.1186/bcr2128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Allison KH. Molecular pathology of breast cancer: what a pathologist needs to know. Am J Clin Pathol. 2012;138:770–780. doi: 10.1309/AJCPIV9IQ1MRQMOO. [DOI] [PubMed] [Google Scholar]
- 22.Sotiriou C, Wirapati P, Loi S, et al. Gene expression profiling in breast cancer: understanding the molecular basis of histologic grade to improve prognosis. J Natl Cancer Inst. 2006;98:262–272. doi: 10.1093/jnci/djj052. [DOI] [PubMed] [Google Scholar]
- 23.Farshid G, Balleine RL, Cummings M, et al. Morphology of breast cancer as a means of triage of patients for BRCA1 genetic testing. Am J Surg Pathol. 2006;30:1357–1366. doi: 10.1097/01.pas.0000213273.22844.1a. [DOI] [PubMed] [Google Scholar]
- 24.Fidalgo F, Rodrigues TC, Pinilla M, et al. Lymphovascular invasion and histologic grade are associated with specific genomic profiles in invasive carcinomas of the breast. Tumor Biol. 2015;36:1835–1848. doi: 10.1007/s13277-014-2786-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Langerod A, Zhao H, Borgan Ø, et al. TP53 mutation status and gene expression profiles are powerful prognostic markers of breast cancer. Breast Cancer Res. 2007;9:R30. doi: 10.1186/bcr1675. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Desmedt C, Haibe-Kains B, Wirapati P, et al. Biological processes associated with breast cancer clinical outcome depend on the molecular subtypes. Clin Cancer Res. 2008;14:5158–5165. doi: 10.1158/1078-0432.CCR-07-4756. [DOI] [PubMed] [Google Scholar]
- 27.Venet D, Dumont JE, Detours V. Most random gene expression signatures are significantly associated with breast cancer outcome. PLoS Comput Biol. 2011;7:e1002240. doi: 10.1371/journal.pcbi.1002240. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Curtis C, Shah SP, Chin S-F, et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature. 2012;486:346–352. doi: 10.1038/nature10983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.The Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature. 2012;490:61–70. doi: 10.1038/nature11412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Gutman DA, Cobb J, Somanna D, et al. Cancer Digital Slide Archive: an informatics resource to support integrated in silico analysis of TCGA pathology data. J Am Med Inf Assoc. 2013;20:1091–1098. doi: 10.1136/amiajnl-2012-001469. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Ciriello G, Gatza ML, Beck AH, et al. Comprehensive molecular portraits of invasive lobular breast cancer. Cell. 2015;163:506–519. doi: 10.1016/j.cell.2015.09.033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Nielsen TO, Parker JS, Leung S, et al. A comparison of PAM50 intrinsic subtyping with immunohistochemistry and clinical prognostic factors in tamoxifen-treated estrogen receptor-positive breast cancer. Clin Cancer Res. 2010;16:5222–5232. doi: 10.1158/1078-0432.CCR-10-1282. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Lawrence MS, Stojanov P, Polak P, et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature. 2013;499:214–218. doi: 10.1038/nature12213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Mermel CH, Schumacher SE, Hill B, et al. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 2011;12:R41. doi: 10.1186/gb-2011-12-4-r41. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Lester SC, Bose S, Chen Y-Y, et al. Protocol for the examination of specimens from patients with invasive carcinoma of the breast. Arch Pathol Lab Med. 2009;133:1515–1538. doi: 10.5858/133.10.1515. [DOI] [PubMed] [Google Scholar]
- 36.Krippendorff K. Content analysis: An introduction to its methodology. 3. Thousand Oaks, CA: Sage; 2013. pp. 221–250. [Google Scholar]
- 37.Ritchie ME, Phipson B, Wu D, et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43:e47. doi: 10.1093/nar/gkv007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Väremo L, Nielsen J, Nookaew I. Enriching the gene set analysis of genome-wide data by incorporating directionality of gene expression and combining statistical hypotheses and methods. Nucleic Acids Res. 2013;41:4378–4391. doi: 10.1093/nar/gkt111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Friedman J, Hastie T, Tibshirani R. Regularization Paths for Generalized Linear Models via Coordinate Descent. J Stat Softw. 2010;33:1–22. [PMC free article] [PubMed] [Google Scholar]
- 40.Dennis G, Jr, Sherman BT, Hosack DA, et al. DAVID: Database for Annotation, Visualization, and Integrated Discovery. Genome Biol. 2003;4:P3. [PubMed] [Google Scholar]
- 41.Paik S, Shak S, Tang G, et al. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N Engl J Med. 2004;351:2817–2826. doi: 10.1056/NEJMoa041588. [DOI] [PubMed] [Google Scholar]
- 42.van ’t Veer LJ, Dai H, van de Vijver MJ, et al. Gene expression profiling predicts clinical outcome of breast cancer. Nature. 2002;415:50–536. doi: 10.1038/415530a. [DOI] [PubMed] [Google Scholar]
- 43.Prat A, Carey LA, Adamo B, et al. Molecular features and survival outcomes of the intrinsic subtypes within HER2-positive breast cancer. J Natl Cancer Inst. 2014;106:dju152. doi: 10.1093/jnci/dju152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Gendoo DMA, Ratanasirigulchai N, Schröder MS, et al. Genefu: an R/Bioconductor package for computation of gene expression-based signatures in breast cancer. Bioinformatics. 2016;32:1097–1099. doi: 10.1093/bioinformatics/btv693. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Haibe-Kains B, Desmedt C, Loi S, et al. A three-gene model to robustly identify breast cancer molecular subtypes. J Natl Cancer Inst. 2012;104:311–325. doi: 10.1093/jnci/djr545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Beck AH, Knoblauch NW, Hefti MM, et al. Significance analysis of prognostic signatures. PLoS Comput Biol. 2013;9:e1002875. doi: 10.1371/journal.pcbi.1002875. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Chin K, DeVries S, Fridlyand J, et al. Genomic and transcriptional aberrations linked to breast cancer pathophysiologies. Cancer Cell. 2006;10:529–541. doi: 10.1016/j.ccr.2006.10.009. [DOI] [PubMed] [Google Scholar]
- 48.Dedeurwaerder S, Desmedt C, Calonne E, et al. DNA methylation profiling reveals a predominant immune component in breast cancers. EMBO Mol Med. 2011;3:726–741. doi: 10.1002/emmm.201100801. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.van de Vijver MJ, He YD, van ’t Veer LJ, et al. A Gene-expression signature as a predictor of survival in breast cancer. N Engl J Med. 2002;347:1999–2009. doi: 10.1056/NEJMoa021967. [DOI] [PubMed] [Google Scholar]
- 50.Desmedt C, Piette F, Loi S, et al. Strong time dependence of the 76-gene prognostic signature for node-negative breast cancer patients in the TRANSBIG multicenter independent validation series. Clin Cancer Res. 2007;13:3207–3214. doi: 10.1158/1078-0432.CCR-06-2765. [DOI] [PubMed] [Google Scholar]
- 51.Hatzis C, Pusztai L, Valero V, et al. A genomic predictor of response and survival following taxane-anthracycline chemotherapy for invasive breast cancer. JAMA. 2011;305:1873–1881. doi: 10.1001/jama.2011.593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Langfelder P, Horvath S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics. 2008;9:559. doi: 10.1186/1471-2105-9-559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Du P, Kibbe WA, Lin SM. lumi: a pipeline for processing Illumina microarray. Bioinformatics. 2008;24:1547–1548. doi: 10.1093/bioinformatics/btn224. [DOI] [PubMed] [Google Scholar]
- 54.Weigelt B, Geyer FC, Reis-Filho JS. Histological types of breast cancer: how special are they? Mol Oncol. 2010;4:192–208. doi: 10.1016/j.molonc.2010.04.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Albrektsen G, Heuch I, Thoresen SØ, et al. Histological type and grade of breast cancer tumors by parity, age at birth, and time since birth: a register-based study in Norway. BMC Cancer. 2010;10:226. doi: 10.1186/1471-2407-10-226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Li CI, Uribe DJ, Daling JR. Clinical characteristics of different histologic types of breast cancer. Br J Cancer. 2005;93:1046–1052. doi: 10.1038/sj.bjc.6602787. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Phipps AI, Li CI. Breast cancer biology and clinical characteristics. In: Li CI, editor. Breast Cancer Epidemiology. Springer; New York: 2010. pp. 21–46. [Google Scholar]
- 58.Dillon D, Guidi A, Schnitt S. Pathology of invasive breast cancer. In: Harris J, Lippman M, Morrow M, et al., editors. Diseases of the Breast. 5. Lippincott Williams & Wilkins Health; Philadelphia: 2014. pp. 381–410. [Google Scholar]
- 59.Arps DP, Healy P, Zhao L, et al. Invasive ductal carcinoma with lobular features: a comparison study to invasive ductal and invasive lobular carcinomas of the breast. Breast Cancer Res Treat. 2013;138:719–726. doi: 10.1007/s10549-013-2493-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Thorner A, Hoadley K, Parker JS, et al. In vitro and in vivo analysis of B-Myb in basal-like breast cancer. Oncogene. 2009;28:742–751. doi: 10.1038/onc.2008.430. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Ma CX, Cai S, Li S, et al. Targeting Chk1 in p53-deficient triple-negative breast cancer is therapeutically beneficial in human-in-mouse tumor models. J Clin Invest. 2012;122:1541–1552. doi: 10.1172/JCI58765. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Osthus RC, Karim B, Prescott JE, et al. The Myc target gene JPO1/CDCA7 is frequently overexpressed in human tumors and has limited transforming activity in vivo. Cancer Res. 2005;65:5620–5627. doi: 10.1158/0008-5472.CAN-05-0536. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Goto Y, Hayashi R, Muramatsu T, et al. JPO1/CDCA7, a novel transcription factor E2F1-induced protein, possesses intrinsic transcriptional regulator activity. Biochim Biophys Acta Gene Struct Expr. 2006;1759:60–68. doi: 10.1016/j.bbaexp.2006.02.004. [DOI] [PubMed] [Google Scholar]
- 64.Prescott JE, Osthus RC, Lee LA, et al. A novel c-Myc-responsive gene, JP01, participates in neoplastic transformation. J Biol Chem. 2001;276:48276–48284. doi: 10.1074/jbc.M107357200. [DOI] [PubMed] [Google Scholar]
- 65.Gill RM, Gabor TV, Couzens AL, et al. The MYC-associated protein CDCA7 is phosphorylated by AKT to regulate MYC-dependent apoptosis and transformation. Mol Cell Biol. 2013;33:498–513. doi: 10.1128/MCB.00276-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Xu J, Chen Y, Olopade OI. MYC and breast cancer. Genes Cancer. 2010;1:629–640. doi: 10.1177/1947601910378691. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Ulz P, Heitzer E, Speicher MR. Co-occurrence of MYC amplification and TP53 mutations in human cancer. Nat Genet. 2016;48:104–106. doi: 10.1038/ng.3468. [DOI] [PubMed] [Google Scholar]
- 68.Moore DJ, Chambers JK, Wahlin JP, et al. Expression pattern of human P2Y receptor subtypes: a quantitative reverse transcription-polymerase chain reaction study. Biochim Biophys Acta. 2001;1521:107–119. doi: 10.1016/s0167-4781(01)00291-3. [DOI] [PubMed] [Google Scholar]
- 69.Vos CB, Cleton-Jansen AM, Berx G, et al. E-cadherin inactivation in lobular carcinoma in situ of the breast: an early event in tumorigenesis. Br J Cancer. 1997;76:1131–1133. doi: 10.1038/bjc.1997.523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Gromov P, Espinoza JA, Talman ML, et al. FABP7 and HMGCS2 are novel protein markers for apocrine differentiation categorizing apocrine carcinoma of the breast. PLoS One. 2014;9:e112024. doi: 10.1371/journal.pone.0112024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Clive KS, Tyler JA, Clifton GT, et al. The GP2 peptide: a HER2/neu-based breast cancer vaccine. J Surg Oncol. 2012;105:452–458. doi: 10.1002/jso.21723. [DOI] [PubMed] [Google Scholar]
- 72.Karaduman M, Bilici A, Ozet A, et al. Tissue levels of adiponectin in breast cancer patients. Med Oncol. 2007;24:361–366. doi: 10.1007/s12032-007-0021-0. [DOI] [PubMed] [Google Scholar]
- 73.Libby EF, Liu J, Li Y, et al. Globular adiponectin enhances invasion in human breast cancer cells. Oncol Lett. 2016;11:633–641. doi: 10.3892/ol.2015.3965. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Gillesby B, Zacharewski T. pS2 (TFF1) levels in human breast cancer tumor samples: correlation with clinical and histological prognostic markers. Breast Cancer Res Treat. 1999;2:253–265. doi: 10.1023/a:1006215310169. [DOI] [PubMed] [Google Scholar]
- 75.Bubnov V, Moskalev E, Petrovskiy Y, et al. Hypermethylation of TUSC5 genes in breast cancer tissue. Exp Oncol. 2012;34:370–372. [PubMed] [Google Scholar]
- 76.Camarero N, Mascaró C, Mayordomo C, et al. Ketogenic HMGCS2 is a c-Myc target gene expressed in differentiated cells of human colonic epithelium and down-regulated in colon cancer. Mol Cancer Res. 2006;4:645–653. doi: 10.1158/1541-7786.MCR-05-0267. [DOI] [PubMed] [Google Scholar]
- 77.Le May C, Pineau T, Bigot K, et al. Reduced hepatic fatty acid oxidation in fasting PPAR3 null mice is due to impaired mitochondrial hydroxymethylglutaryl-CoA synthase gene expression. FEBS Lett. 2000;475:163–166. doi: 10.1016/s0014-5793(00)01648-3. [DOI] [PubMed] [Google Scholar]
- 78.Carracedo A, Cantley LC, Pandolfi PP. Cancer metabolism: fatty acid oxidation in the limelight. Nat Rev Cancer. 2013;13:227–232. doi: 10.1038/nrc3483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Yadava N, Schneider SS, Jerry DJ, et al. Impaired mitochondrial metabolism and mammary carcinogenesis. J Mammary Gland Biol Neoplasia. 2013;18:75–87. doi: 10.1007/s10911-012-9271-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Kamp DW, Shacter E, Weitzman SA. Chronic inflammation and cancer: the role of the mitochondria. Oncology. 2011;25:400–410. 413. [PubMed] [Google Scholar]
- 81.Jeong YJ, Bong JG, Park SH, et al. Expression of leptin, leptin receptor, adiponectin, and adiponectin receptor in ductal carcinoma in situ and invasive breast cancer. J Breast Cancer. 2011;14:96–103. doi: 10.4048/jbc.2011.14.2.96. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Shield KD, Soerjomataram I, Rehm J. Alcohol use and breast cancer: a critical review. Alcohol Clin Exp Res. 2016;40:1166–1181. doi: 10.1111/acer.13071. [DOI] [PubMed] [Google Scholar]
- 83.Kaklamani VG, Hoffmann TJ, Thornton TA, et al. Adiponectin pathway polymorphisms and risk of breast cancer in African Americans and Hispanics in the Women’s Health Initiative. Breast Cancer Res Treat. 2013;139:461–468. doi: 10.1007/s10549-013-2546-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Stein T. Annexin A8 is up-regulated during mouse mammary gland involution and predicts poor survival in breast cancer. Clin Cancer Res. 2005;11:6872–6879. doi: 10.1158/1078-0432.CCR-05-0547. [DOI] [PubMed] [Google Scholar]
- 85.Rogers MS, Foleyt MA, Crotty TB, et al. Loss of immunoreactivity for human calmodulin-like protein is an early event in breast cancer development. Neoplasia. 1999;1:220–225. doi: 10.1038/sj.neo.7900029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Köhrmann A, Kammerer U, Kapp M, et al. Expression of matrix metalloproteinases (MMPs) in primary human breast cancer and breast cancer cell lines: new findings and review of the literature. BMC Cancer. 2009;9:188. doi: 10.1186/1471-2407-9-188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Adriance MC, Inman JL, Petersen OW, et al. Myoepithelial cells: good fences make good neighbors. Breast Cancer Res. 2005;7:190–197. doi: 10.1186/bcr1286. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Tamiolakis D, Papadopoulos N, Cheva A, et al. Immunohistochemical expression of alpha-smooth muscle actin in infiltrating ductal carcinoma of the breast with productive fibrosis. Eur J Gynaecol Oncol. 2002;23:469–471. [PubMed] [Google Scholar]
- 89.Li F, Zou Z, Suo N, et al. CCL21/CCR7 axis activating chemotaxis accompanied with epithelial–mesenchymal transition in human breast carcinoma. Med Oncol. 2014;31:180. doi: 10.1007/s12032-014-0180-8. [DOI] [PubMed] [Google Scholar]
- 90.Weitzenfeld P, Kossover O, Körner C, et al. Chemokine axes in breast cancer: factors of the tumor microenvironment reshape the CCR7-driven metastatic spread of luminal-A breast tumors. J Leukoc Biol. 2016;99:1009–1025. doi: 10.1189/jlb.3MA0815-373R. [DOI] [PubMed] [Google Scholar]
- 91.Pang M, Georgoudaki A, Lambut L, et al. TGF-Beta1-induced EMT promotes targeted migration of breast cancer cells through the lymphatic system by the activation of CCR7/CCL21-mediated chemotaxis. Oncogene. 2016;35:748–760. doi: 10.1038/onc.2015.133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Schnitt SJ. The transition from ductal carcinoma in situ to invasive breast cancer: the other side of the coin. Breast Cancer Res. 2009;11:101. doi: 10.1186/bcr2228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Crawford NPS, Walker RC, Lukes L, et al. The diasporin pathway:a tumor progression-related transcriptional network that predicts breastcancer survival. Clin Exp Metastasis. 2008;25:357–369. doi: 10.1007/s10585-008-9146-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Shi Z, Zhang T, Luo L, et al. Aquaporins in human breast cancer: identification and involvement in carcinogenesis of breast cancer. J Surg Oncol. 2012;106:267–272. doi: 10.1002/jso.22155. [DOI] [PubMed] [Google Scholar]
- 95.Jung HJ, Park J-Y, Jeon H-S, et al. Aquaporin-5: a marker protein for proliferation and migration of human breast cancer cells. PLoS One. 2011;6:e28492. doi: 10.1371/journal.pone.0028492. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Su ML, Chang TM, Chiang CH, et al. Inhibition of chemokine (C-C Motif) receptor 7 sialylation suppresses CCL19-stimulated proliferation, invasion and anti-anoikis. PLoS One. 2014;9:e98823. doi: 10.1371/journal.pone.0098823. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Liu L, Xu Q, Cheng L, et al. NPY1R is a novel peripheral blood marker predictive of metastasis and prognosis in breast cancer patients. Oncol Lett. 2015;9:891–896. doi: 10.3892/ol.2014.2721. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Ehrenfeld P, Manso L, Pavicic MF, et al. Bioregulation of kallikrein-related peptidases 6, 10 and 11 by the Kinin b1 receptor in breast cancer cells. Anticancer Res. 2014;34:6925–6938. [PubMed] [Google Scholar]
- 99.Yau C, Esserman L, Moore DH, et al. A multigene predictor of metastatic outcome in early stage hormone receptor-negative and triple-negative breast cancer. Breast Cancer Res. 2010;12:R85. doi: 10.1186/bcr2753. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Wang J, Scholtens D, Holko M, et al. Lipid metabolism genes in contralateral unaffected breast and estrogen receptor status of breast cancer. Cancer Prev Res. 2013;6:321–330. doi: 10.1158/1940-6207.CAPR-12-0304. [DOI] [PubMed] [Google Scholar]
- 101.Lee SJ, Chae YS, Kim JG, et al. AQP5 expression predicts survival in patients with early breast cancer. Ann Surg Oncol. 2013;21:375–383. doi: 10.1245/s10434-013-3317-7. [DOI] [PubMed] [Google Scholar]
- 102.Yamada A, Ishikawa T, Ota I, et al. High expression of ATP-binding cassette transporter ABCC11 in breast tumors is associated with aggressive subtypes and low disease-free survival. Breast Cancer Res Treat. 2013;137:773–782. doi: 10.1007/s10549-012-2398-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Park S, Shimizu C, Shimoyama T, et al. Gene expression profiling of ATP-binding cassette (ABC) transporters as a predictor of the morphological response to neoadjuvant chemotherapy in breast cancer patients. Breast Cancer Res Treat. 2006;99:9–17. doi: 10.1007/s10549-006-9175-2. [DOI] [PubMed] [Google Scholar]
- 104.Hlavác V, Brynychová V, Václavíková R, et al. The expression profile of ATP-binding cassette transporter genes in breast carcinoma. Pharmacogenomics. 2013;14:515–529. doi: 10.2217/pgs.13.26. [DOI] [PubMed] [Google Scholar]
- 105.Sensorn I, Sirachainan E, Chamnanphon M, et al. Association of CYP3A4/5, ABCB1 and ABCC2 polymorphisms and clinical outcomes of Thai breast cancer patients treated with tamoxifen. Pharmgenomics Pers Med. 2013;6:93–98. doi: 10.2147/PGPM.S44006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Litviakov NV, Cherdyntseva NV, Tsyganov MM, et al. Deletions of multidrug resistance gene loci in breast cancer leads to the down-regulation of its expression and predict tumor response to neoadjuvant chemotherapy. Oncotarget. 2016;5:7829–7841. doi: 10.18632/oncotarget.6953. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Mobasheri A, Barrett-Jolley R. Aquaporin water channels in the mammary gland: from physiology to pathophysiology and neoplasia. J Mammary Gland Biol Neoplasia. 2014;19:91–102. doi: 10.1007/s10911-013-9312-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Mao H, Zhang L, Yang Y, et al. New insights of CTLA-4 into its biological function in breast cancer. Curr Cancer Drug Targets. 2010;10:728–736. doi: 10.2174/156800910793605811. [DOI] [PubMed] [Google Scholar]
- 109.Loi S. Tumor infiltrating lymphocytes (TILs) indicate trastuzumab benefit in early-stage HER2-positive breast cancer (HER2+ BC) Cancer Res. 2013;73:S1–S5. [Google Scholar]
- 110.Jure-Kunkel M, Masters G, Girit E, et al. Synergy between chemotherapeutic agents and CTLA-4 blockade in preclinical tumor models. Cancer Immunol Immunother. 2013;62:1533–1545. doi: 10.1007/s00262-013-1451-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Wang Q, Li SH, Wang H, et al. Concomitant targeting of tumor cells and induction of T-cell response synergizes to effectively inhibit trastuzumab-resistant breast cancer. Cancer Res. 2012;72:4417–4428. doi: 10.1158/0008-5472.CAN-12-1339-T. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Demaria S, Kawashima N, Yang AM, et al. Immune-mediated inhibition of metastases after treatment with local radiation and CTLA-4 blockade in a mouse model of breast cancer. Clin Cancer Res. 2005;11:728–734. [PubMed] [Google Scholar]
- 113.Mittendorf EA, Philips AV, Meric-Bernstam F, et al. PD-L1 expression in triple-negative breast cancer. Cancer Immunol Res. 2014;2:361–370. doi: 10.1158/2326-6066.CIR-13-0127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Stagg J, Loi S, Divisekera U, et al. Anti-ErbB-2 mAb therapy requires type I and II interferons and synergizes with anti-PD-1 or anti-CD137 mAb therapy. Proc Natl Acad Sci USA. 2011;108:7142–7147. doi: 10.1073/pnas.1016569108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Janakiram M, Abadi YM, Sparano JA, et al. T cell coinhibition and immunotherapy in human breast cancer. Discov Med. 2012;14:229–236. [PMC free article] [PubMed] [Google Scholar]
- 116.Hwang ES, McLennan JL, Moore DH, et al. Ductal carcinoma in situ in BRCA mutation carriers. J Clin Oncol. 2007;25:642–647. doi: 10.1200/JCO.2005.04.0345. [DOI] [PubMed] [Google Scholar]
- 117.Hannemann J, Velds A, Halfwerk JBG, et al. Classification of ductal carcinoma in situ by gene expression profiling. Breast Cancer Res. 2006;8:R61. doi: 10.1186/bcr1613. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Done SJ, Eskandarian S, Bull S, et al. p53 Missense mutations in microdissected high-grade ductal carcinoma in situ of the breast. J Natl Cancer Inst. 2001;93:700–7004. doi: 10.1093/jnci/93.9.700. [DOI] [PubMed] [Google Scholar]
- 119.Carraro DM, Elias EV, Andrade VP. Ductal carcinoma in situ of the breast: morphological and molecular features implicated in progression. Biosci Rep. 2014;34:19–28. doi: 10.1042/BSR20130077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Mujtaba SS, Ni Y-B, Tsang JYS, et al. Fibrotic focus in breast carcinomas: relationship with prognostic parameters and biomarkers. Ann Surg Oncol. 2013;20:2842–2849. doi: 10.1245/s10434-013-2955-0. [DOI] [PubMed] [Google Scholar]
- 121.Hasebe T, Sasaki S, Imoto S, et al. Prognostic significance of fibrotic focus in invasive ductal carcinoma of the breast: a prospective observational study. Mod Pathol. 2002;15:502–516. doi: 10.1038/modpathol.3880555. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Images of the electronic scoring sheets: (A) used by the pathologists to annotate images; (B) annotated scoring sheet with pathological scoring criteria and details.
Boxplots displaying PAM50 proliferation scores for each morphological feature.
Expanded heatmap of Figure 2, with detailed gene sets.
Tables summarising the distribution of morphological features within each PAM50 subtype for (A) all cases; (B) invasive ductal cases; (C) invasive lobular cases; and (D) special histological types.
Detailed data for morphological features, PAM50, DNA methylation, miRNA and RPPA subtypes, PAM50 proliferation scores and genomic alterations for 850 TCGA cases.
(A) The association of molecular subtypes PAM50, DNA methylation, RPPA, and microRNA with morphological features. (B) Association of morphological features with each other.
(A) The frequency of each morphological feature in binary groups and the number of differentially expressed genes, stratified by PAM50 subtype, excluding Normal-like. (B) Top 10 up- or down-regulated differentially-expressed genes (limma FDR<0.05) for each morphological feature, ranked by Log2 fold change (Log2FC).
(A) Multivariate ROCAUC performances of genomic alterations, gene expression and combined data. (B) Paired t-test of ROCAUC of genomic alterations, gene expression and combined data models. (C) Summary of transcriptomic signatures and their enrichments. (D) Transcriptomic signatures of morphological features. (E) Gene set enrichments for transcriptomic signatures of morphological feature.