Summary
Recent reports have documented the differentiation of human pluripotent stem cells toward the skeletal myogenic lineage using transgene- and cell purification-free approaches. Although these protocols generate myocytes, they have not demonstrated scalability, safety, and in vivo engraftment, which are key aspects for their future clinical application. Here we recapitulate one prominent protocol, and show that it gives rise to a heterogeneous cell population containing myocytes and other cell types. Upon transplantation, the majority of human donor cells could not contribute to myofiber formation. As a proof-of-principle, we incorporated the inducible PAX7 lentiviral system into this protocol, which then enabled scalable expansion of a homogeneous population of skeletal myogenic progenitors capable of forming myofibers in vivo. Our findings demonstrate the methods for scalable expansion of PAX7+ myogenic progenitors and their purification are critical for practical application to cell replacement treatment of muscle degenerative diseases.
Keywords: pluripotent stem cells, skeletal muscle, morphogens, purification, PAX7+ myogenic progenitors, MHC+ myocytes, transgene, engraftment, stem cell therapy
Highlights
-
•
Tested CDM muscle differentiation protocol generates a heterogeneous population
-
•
The population does not contribute to myofiber formation in vivo
-
•
Regulated PAX7 expression allows expansion of homogeneous myogenic progenitors
-
•
CDM iPAX7 myogenic progenitors are endowed with in vivo regenerative potential
In this report, Perlingeiro and colleagues test a chemically defined monolayer (CDM) differentiation protocol and find that it generates a heterogeneous myogenic cell population that does not contribute to myofiber formation in vivo. They demonstrate that conditional expression of PAX7 enables the expansion of a homogeneous population of myogenic progenitors that have in vivo regenerative potential.
Introduction
The ability to self-renew and to differentiate into all somatic cell types makes pluripotent stem (PS) cells an attractive cell source for therapeutic applications. Since the isolation of human embryonic stem (hES) cells, subsequently boosted by the technology of reprogramming somatic cells into induced PS (iPS) cells, several protocols have been reported for differentiating human PS (hPS) cells toward a specific lineage (Tabar and Studer, 2014), with dozens of these applied to in vitro disease modeling, and many fewer to in vivo regeneration (Darabi et al., 2012, Hargus et al., 2010, Kriks et al., 2011, Kroon et al., 2008, Laflamme et al., 2007, Sebastiano et al., 2014, Wang et al., 2012). The development of tissue-specific stem cells with proven in vivo therapeutic potential from hPS cells is evidently a far more challenging milestone than making specific differentiated cell types. However, proof-of-concept for in vivo regeneration with demonstrated functional improvement has been achieved for the skeletal muscle lineage, even though this tissue has been extremely difficult to generate from hPS cells because of the complex morphogenetic events that result in somitogenesis and key somite patterning cues, e.g., notochord and neural tube, do not occur in vitro. We have shown that conditional expression of PAX3 or PAX7 in early unpatterned mesoderm overcomes this roadblock in embryoid body (EB) culture, allowing for the generation of hPS cell-derived myogenic progenitor cells that expand scalably, engraft extensively, seed the stem cell compartment, and, most importantly, promote improved muscle contractility of transplanted dystrophic muscles (Darabi et al., 2012). Other reports have made use of MYOD to drive myogenic differentiation from hPS cells (Albini et al., 2013, Goudenege et al., 2012, Tedesco et al., 2012, Young et al., 2016). One common feature of these approaches is lentiviral delivery of the transgene. In the last few years, several reports have documented the use of transgene-free monolayer cultures for the differentiation of hPS cells toward the skeletal myogenic lineage through activation of WNT signaling and modulation of other signaling pathways (Barberi et al., 2007, Borchin et al., 2013, Chal et al., 2015, Choi et al., 2016, Shelton et al., 2014, Tan et al., 2013, Xi et al., 2017, Young et al., 2016). These protocols allow for the derivation of myocytes expressing myosin heavy chain (MHC); however, their translational applicability is undetermined due to several important unknowns associated with one or more of the following important aspects, including (1) purity of myogenic cell cultures, (2) scalability of a myogenic progenitor cell population, and (3) in vivo regenerative potential. To date, these aspects have not been comprehensively addressed. In the present study we recapitulate a more prominent one of these recently published protocols (Chal et al., 2015).
Results
Heterogeneity of Monolayer-Differentiated Cultures
To validate the feasibility of a transgene- and purification-free approach to generate myogenic progenitors from hPS cells for therapeutic application, here we recapitulated the chemically defined monolayer (CDM) protocol recently published by Chal et al. (2015), which is characterized by the combination of WNT activation and BMP inhibition. We began these studies using H9 ES cells as this hPS cell line was utilized in the reported protocol and, as described (Chal et al., 2015), H9 cultures were maintained without passaging for 50 days (Figure 1A). Within the first 7–10 days from the start of the protocol, we observed extensive overgrowth. Nevertheless, remaining viable cells differentiated into multinucleated myotubes. Importantly, as cultures progressed, careful observation revealed morphological heterogeneity (Figure 1B, upper panel). The anticipated multinucleated myotubes were interspersed with structures resembling neuronal aggregates displaying neurite-like connections, with no significant morphological differences observed between day 25 and day 50 cultures. This heterogeneity was confirmed by immunostaining (Figure 1B) and gene expression (Figure S1A) for MHC and βIII-tubulin (TUBB3), which identify myocytes (Schiaffino and Reggiani, 1996) and neurons (Dennis et al., 2002), respectively. We also observed cells that were negative for both MHC and TUBB3 (Figure 1B), implying the presence of other unidentified cell types. Gene expression for pluripotency markers revealed high levels of SOX2 and KLF4, but not of OCT4 (Figure S1B). Since SOX2 and KLF4 are markers of neural progenitors during early neurogenesis (Cimadamore et al., 2013, Qin and Zhang, 2012), their expression likely reflects the presence of contaminating neural cells in these cultures. Similar heterogeneity was observed among five additional hPS cell lines (four iPS cell lines and the H1 ES cell line), which showed highly variable degree of MHC+ myocyte differentiation (Figures 1B, S1C, and S2).
Figure 1.
In Vitro and In Vivo Skeletal Myogenic Differentiation Potential of Transgene-free hPS Cell-Derived Myogenic Cells Generated Using the Monolayer Method
(A) Schematic diagram of differentiating hPS cells in monolayer using only small molecules without passaging (i = CHIR99021 and LDN; ii = CHIR99021, LDN, and FGF2; iii = LDN, FGF2, HGF, and IGF1; iv = IGF1; v = HGF and IGF1).
(B) Representative bright field image and immunofluorescence analysis for MHC and TUBB3 of CDM-H9 cells (after 50 days) and other CDM-hPS cells (after 30 days). MHC in red; TUBB3 in green; DAPI (nuclei) in blue. Scale bars, 200 μm (n = 4 biological replicates).
(C) Representative immunofluorescence analysis for PAX7 and MHC of CDM-H9 cells after 30 days (top panel: 20 neighbor images under 10× magnification were joined together using “tiles imaging” mode). PAX7 in yellow; MHC in red; DAPI (nuclei) in blue. Scale bars, 200 μm (n = 4 biological replicates).
(D) Western blot analysis of CDM-H9 cells at different time points. Mouse satellite (mSat) cells and iPAX7+CDM-H9 myogenic progenitors (sorted for PAX7+ and expanded for 4 days in the presence of Dox) were used as positive controls for PAX7 expression. Non-induced iPAX7+CDM-H9 cells (−Dox) served as negative control. Actin (ACT) was used as a housekeeping protein. Approximately 100,000 cells were used for each protein sample. Lane 1, day 20 of CDM-H9; lane 2, day 30 of CDM-H9; lane 3, day 40 of CDM-H9; lane 4, iPAX7+CDM-H9 without Dox; lane 5, mSat; lane 6, iPAX7+CDM-H9 with Dox (n = 2 biological replicates).
(E) Representative immunohistochemistry analysis for LMNA-C and DYS of transplanted CDM-H9 cells at day 30 which showed PAX7+ sub-population within the culture (left panel). Number of cells positive for LMNA-C and DYS was quantified for each biological replicate of each muscle section (right panel). LMNA-C in green; DYS in red; DAPI (nuclei) in blue. Scale bars, 200 μm (n = 4 biological replicates).
CDM-Derived Cultures Lack Muscle Engraftment Potential
Next we investigated the in vivo regenerative potential of CDM-H9 myogenic cells by injecting day 25 cultures into cardiotoxin-injured muscles of NOD scid gamma (NSG) mice. Immunostaining for human LAMIN-AC (LMNA-C) revealed the presence of human donor cells in transplanted muscles (Figure S1D). However, we failed to detect donor-derived myofibers as no signal was found for human SPECTRIN (SPEC) and DYSTROPHIN (DYS) (Figures S1D and S1E), suggesting that injected cells survived the intramuscular transplantation but failed to contribute to muscle regeneration.
As reported (Chal et al., 2015, Chal et al., 2016), we were able to detect a putative PAX7+ sub-population, along with MHC+ cells at day 30 CDM cultures by immunofluorescence staining (Figure 1C). However, western blot analysis showed no signal for PAX7 expression in these CDM cultures, contrasting to satellite cells and PAX7-induced hPS cell-derived myogenic progenitors (Figure 1D). This could be due to the limited number of PAX7+ cells within these CDM-differentiated cultures. Nevertheless, next we transplanted day 30 myogenic CDM-H9 cultures, which coincided with PAX7 detection by immunostaining (Figure 1C). As before (Figure S1D), human donor-derived cells were detected, but minimal contribution to muscle regeneration was observed (Figure 1E). Thus, the high level of heterogeneity, limited number of PAX7-expressing cells, and, importantly, minimal in vivo regenerative potential, raises questions about the suitability of this transgene-free CDM approach for clinical applications.
CDM Protocol Incorporating Expansion
Despite the overgrowth, most of the protocols to date involving serum-free CDM approaches for both skeletal (Barberi et al., 2007, Borchin et al., 2013, Chal et al., 2015, Shelton et al., 2014) and cardiac (Lian et al., 2012, Mummery et al., 2012) muscle differentiation do not involve passaging. It is plausible that the maintenance of cells at high density, together with the presence of morphogens, is a requirement for triggering both skeletal and cardiac myogenesis in CDM conditions. Major caveats for these culture conditions are the heterogeneity of cell preparations, lack of scalability, and cell death. Recently, Chal et al. (2016) reported a follow-up updated protocol, which describes that their CDM-derived cultures can be passaged if fetal bovine serum (FBS) is added to culture medium (EXP). In the presence of these less-defined culture conditions (i.e., animal serum supplemented), we found that EXP-H9 cells were expandable (Figure 2A). Upon multiple passaging, fewer cell aggregates were observed but, nevertheless, TUBB3+ neurons were still detected in both expansion (Figures 2B and 2C) and terminal differentiation (DIFF) (Figures 3A and 3B) phases of H9 cells, albeit at lower frequency. In the expansion phase, a sub-population of MYOD- and MYOG-expressing cells was detected for differentiating EXP-H9 cells, but the frequency of PAX7+ nuclei was low (Figures 2B and 2C). Under terminal differentiation conditions, which consisted of horse serum (HS)-supplemented medium (Chal et al., 2016), these DIFF-H9 cultures showed generation of MHC+ myocytes (Figures 3A and 3B).
Figure 2.
Characterization of Transgene-free hPS Cell-Derived Myogenic Cells in Expansion Stage
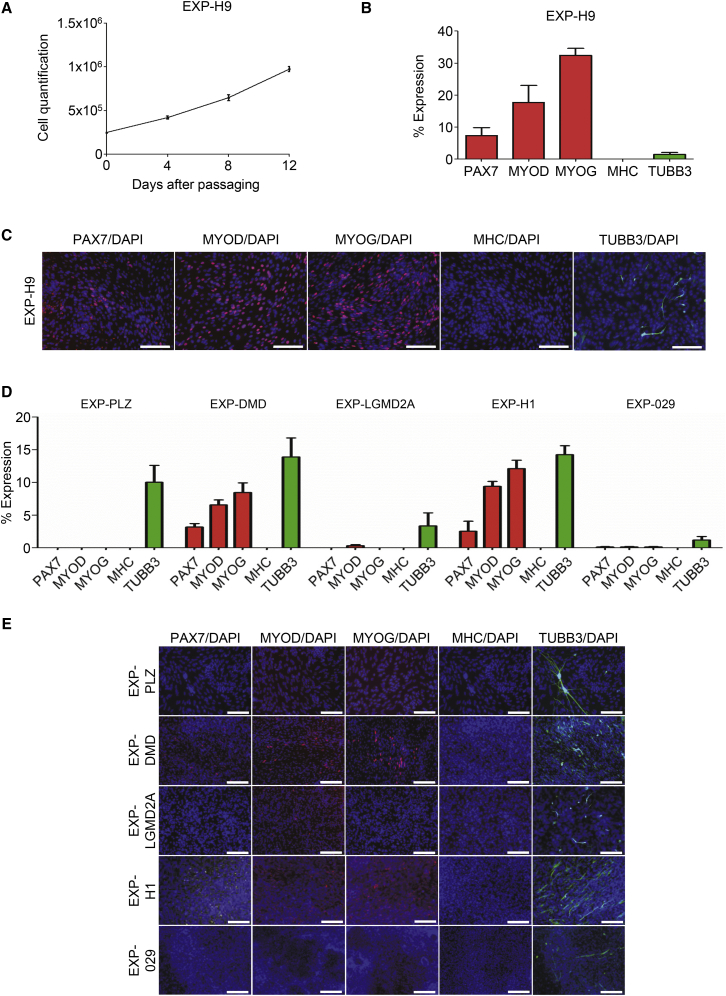
(A) Growth curve analysis of EXP-H9 cells that were CDM-derived for 30 days, followed by expansion in serum-containing conditions for 12 days (values represent mean ± SEMs; n = 3 biological replicates).
(B and C) Quantification (B) and representative immunofluorescence analyses for PAX7, MYOD, MYOG, MHC, and TUBB3 of EXP-H9 cells (C). PAX7, MYOD, MYOG, and MHC in red; TUBB3 in green; DAPI (nuclei) in blue. Scale bars, 200 μm (values represent mean ± SEMs; n = 4 biological replicates).
(D and E) Quantification (D) and representative immunofluorescence analyses for PAX7, MYOD, MYOG, MHC, and TUBB3 expression of EXP-PLZ, -DMD, -LGMD2A, -H1, and -029 cells that were CDM-derived for 30 days, followed by expansion in serum-containing medium for 12 days (E). Cells were stained as described above in (C). Experiments involving EXP-H1 and -029 cells were performed by an independent laboratory. Scale bars, 200 μm (values represent mean ± SEMs; n = 3 biological replicates).
Figure 3.
Characterization of Transgene-free hPS Cell-Derived Myogenic Cells in Terminal Differentiation Stage
(A and B) Quantification (A) and representative immunofluorescence analyses for MHC and TUBB3 of DIFF-H9 cells that were CDM-derived for 30 days, expanded in serum-containing medium for 12 days, and terminally differentiated for one week in HS-containing culture medium (B). MHC in red; TUBB3 in green; DAPI in blue. Scale bars, 200 μm (values represent mean ± SEM; n = 4 biological replicates).
(C and D) Quantification (C) and representative immunofluorescence analyses for MHC and TUBB3 of DIFF-PLZ, -DMD, -LGMD2A, -H1, and -029 cells (DIFF-H1 and -029 cells by an independent lab group) that were cultured (D). The cells were stained as described in (B). Scale bars, 200 μm (values represent mean ± SEM; n = 3 biological replicates).
(E and F) Immunohistochemistry analysis for LMNA-C and DYS in mice that had been transplanted with EXP-H9 cells that were CDM differentiated for 30 days and then expanded in serum-containing medium for 12 days (E). LMNA-C in green; DYS in red; DAPI (nuclei) in blue. Scale bars, 200 μm. The respective quantifications were recorded from muscle sections (F) (values represent mean ± SEM; n = 4 biological replicates).
To further confirm these results, we tested this protocol in several other hPS cell lines and observed a similar pattern, although there was high variability in terms of myogenic potential, as demonstrated by PAX7, MYOD, and MYOG expression, and the presence of neuronal TUBB3+ cells (Figures 2D and 2E). Accordingly, some of these hPS cell lines showed poor differentiation into MHC+ cells (Figures 3C and 3D), confirming our initial results.
Next we examined the in vivo regenerative potential of this mixed population of myoblasts and putative satellite cells. For this, we used EXP-H9 cell preparations with the most promising in vitro results (Figures 2C and 3B). Six weeks post injection, very few donor-derived cells were detected, indicating low survivability of transplanted cells (Figures 3E and 3F). Overall, the new approach, no longer xenogen-free, allows cell scalability but still raises concerns for its application for cell therapy as it still generates a heterogeneous population of both myocytes and neurons that do not efficiently contribute to myofiber formation in vivo.
PAX7 Enables Expansion and Engraftment from Monolayer-Derived Cells
Myogenic progenitors and adult muscle stem cells express PAX7, a paired box transcription factor essential for the commitment and maintenance of these cells (Gros et al., 2005, Lagha et al., 2008, Seale et al., 2000). Previous studies demonstrated that transplantable myogenic progenitors produced from both murine (Darabi et al., 2011) and hPS cells can be expanded by maintaining expression of PAX7 (Darabi et al., 2012, Demestre et al., 2015). Therefore, we reasoned that one of the requirements for the generation of myogenic progenitors is the robust expression of this important muscle regulator. To test whether PAX7 expression could expand hPS cell-derived myogenic progenitors produced under monolayer culture, we implemented the conditional doxycycline (Dox)-inducible PAX7 (iPAX7) lentiviral system to the CDM protocol using the H9 hES cell line (iPAX7+CDM; Figure 4A). After 4 days of PAX7 induction, PAX7+ myogenic progenitor cells were purified based on the expression of co-translated red fluorescent protein (RFP) by fluorescence-activated cell sorting (FACS; approximately >95% sorting efficiency; Figure S3A), and expanded using the same serum-free culture medium. Only PAX7-induced cultures (+Dox) displayed significant proliferative capacity, whereas control non-induced PAX7 (–Dox) counterparts did not expand under these serum-free conditions (Figure 4B). This highlights the importance of PAX7 induction for the scalability of myogenic progenitors in serum-free conditions. As expected, proliferating Dox-induced myogenic progenitors stained positive for PAX7 (Figure 4C, lower panel). Upon Dox withdrawal, cells underwent robust terminal differentiation, as shown by the homogeneous presence of MHC+ multinucleated myotubes and the absence of TUBB3 expression (Figure 4D, middle panel). In the absence of PAX7 induction, cultures were negative for MHC and only a few TUBB3+ cells were detected (Figure 4D, upper panel). Similar differentiation was observed with the iPAX7-PLZ hiPS cell line with Dox treatment (Figure 4D, lower panel). Next we investigated whether PAX7-induced myogenic progenitors generated using the CDM protocol would contribute to in vivo regeneration, as previously reported using the EB method (Darabi et al., 2012). As distinct from the results obtained with CDM-H9 cells (Figures 1E and S1D), PAX7-induced CDM-derived cells gave rise to muscle fibers expressing DYS, which co-localized with human LMNA-C (Figures 4E and 4F).
Figure 4.
Skeletal Myogenic Differentiation and Engraftment Ability of Myogenic Progenitors Derived from hPS Cells Using the iPAX7 System under Serum-free Conditions
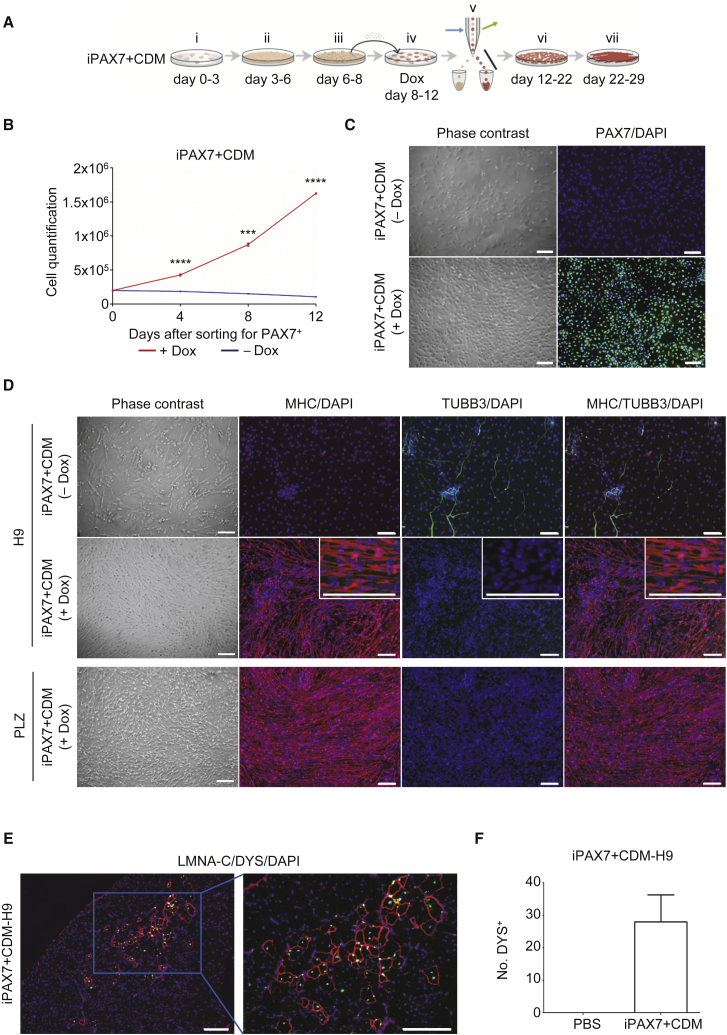
(A) Schematic diagram of differentiating iPAX7-hPS cells using the PAX7 conditional expression system in CDM protocol (i–iii = same culture conditions as shown in Figure 1A; iv = passaging with Dox and FGF2; v = sorting for PAX7+ cells; vi = propagation of PAX7+ cells with Dox and FGF2; vii = HGF and IGF1).
(B) Growth curve analysis of iPAX7+CDM-H9 cells that had been cultured with or without Dox. Cells were counted after day 12 of differentiation. Number of cells was recorded every 4 days. (values represent mean ± SEM; n = 3 biological replicates; ∗∗∗p ≤ 0.001, ∗∗∗∗p ≤ 0.0001).
(C) Representative bright field image and immunofluorescence staining for PAX7 in iPAX7+CDM-H9 cells that had been cultured with or without Dox. iPAX7+CDM-H9 cells cultured with Dox were sorted for PAX7+ after 4 days of Dox induction (A), and expanded for 4 days, whereas non-induced cells (−Dox) were maintained in the same culture dish. PAX7 in green; DAPI (nuclei) in blue. Scale bars, 200 μm (n = 3 biological replicates).
(D) Representative bright field image and immunofluorescence analysis for MHC and TUBB3 of iPAX7+CDM-hPS cells differentiated until day 29 with or without Dox induction, and then subjected to terminal differentiation. MHC in red; TUBB3 in green; DAPI (nuclei) in blue. Scale bars, 200 μm (n = 3 biological replicates).
(E and F) Immunofluorescence analysis for LMNA-C and DYS in mice that had been transplanted with day 22 PAX7-induced H9 myogenic progenitors (E). LMNA-C in green; DYS in red; DAPI (nuclei) in blue. Scale bars, 200 μm. The respective quantifications were recorded from muscle sections (F) (values represent mean ± SEM; n = 4 biological replicates).
Discussion
One of the major obstacles for the application of cell replacement therapy for muscular dystrophy patients is the generation of large quantities of PAX7+ stem cells/myogenic progenitors, which, upon transplantation, contribute to efficient myofiber and stem cell engraftment. Here we demonstrate that myogenic cells generated from multiple hPS cell lines using one prominent CDM protocol (Chal et al., 2015, Chal et al., 2016) are incompatible for muscle cell therapy due to the generation of a non-scalable heterogeneous population of myocytes and other cell types. Although incorporation of animal serum allowed cell expansion in the update protocol (Chal et al., 2016), in addition to losing the value of a xenogen-free system, it still produced a heterogeneous population of cells, which lacked the ability to survive and form myofibers within the host. In addition, derivation of PAX7+ cells, which would represent presumptive satellite-like cells, displayed inconsistent results and suggested that myogenic commitment observed under chemical-defined conditions may be achieved through activation of MYOD expression rather than a PAX7+ muscle stem cell population (Abujarour et al., 2014, Albini et al., 2013, Salani et al., 2012, Tanaka et al., 2013).
Importantly, we show that exogenous PAX7 expression in these CDM cultures enables efficient myogenic progenitor propagation in vitro and in vivo regenerative potential. In addition, due to the presence of a reporter system (PAX7-ires-RFP), PAX7 expression allows purification of myogenic progenitors, thus eliminating other unwanted lineage-derived cells (particularly neurons) which are observed in cultures from the CDM protocol and likely account for the large clusters of DYS− cells observed upon their intramuscular transplantation into NSG mice. This aspect could be ameliorated with the inclusion of a purification step. In this regard, Barberi and colleagues have improved their original CDM protocol (Barberi et al., 2007) by developing a stringent FACS purification strategy based on the negative selection of human natural killer-1 (HNK-1/CD57), which excludes neural and neural crest cells, and the expression of the muscle-specific nicotinic acetylcholine receptor (ACHR), the chemokine receptor CXCR4, and the hepatocyte growth factor (HGF) receptor (also known as C-MET) (Borchin et al., 2013). These authors concluded that the HNK-1−ACHR+C-MET+ fraction, positive or negative for CXCR4, allowed for the isolation of PAX3+/PAX7+ progenitors. Another recent study has applied negative selection of HNK-1/CD57 combined with expression of NCAM to isolate myoblast cells from CDM cultures (Choi et al., 2016). Because both reports were solely focused on in vitro studies, it remains unknown whether such purification strategies would result in better outcome in terms of in vivo regeneration for myogenic progenitors obtained from CDM cultures.
Although our conclusions cannot be extended to other transgene-free CDM methods, our findings suggest that PAX7-transgene expression is important for generating a scalable population of myogenic progenitors with robust in vitro myogenic differentiation potential and capable of contributing to muscle regeneration in vivo. A limitation here is that it remains to be investigated whether myogenic progenitors obtained from iPAX7+CDM cultures are as functional as the ones previously reported using the EB system (Darabi et al., 2011, Darabi et al., 2012). We have previously shown that a fraction of transplanted cells remains mononuclear, and displays key features of skeletal muscle stem cells, including satellite cell localization in combination with co-expression of PAX7 and human LMNA-C, response to re-injury, and contribution to muscle regeneration in secondary transplantation assays, and long-term engraftment. Future studies will investigate whether the monolayer method reported in this study in conjunction with iPAX7 delivery yields engrafted human satellite-like cells.
Experimental Procedures
hPS Cell Cultures and Initial Differentiation
Undifferentiated H9 (ES cell), H1 (ES cell), PLZ (wild-type iPS cell), 029 (wild-type iPS cell), 1705 (Duchenne muscular dystrophy; DMD iPS cell), RK2 (limb-girdle MD type 2A; LGMD2A iPS cell), iPAX7-H9 (ES cell), and iPAX7-PLZ (iPS cell) hPS cells were cultured on Matrigel-coated dishes (BD Biosciences) in mTeSR1 medium (STEMCELL Technologies). Cells were passaged as aggregates. For generation of PAX7+ myogenic progenitors from hPS cells using the Dox-inducible PAX7 system in the CDM protocol, cells were dissociated at day 8 of iPAX7 + CDM differentiation by trypsinization and were seeded onto fresh Matrigel-coated 6-well dishes (300,000 cells/well) with K-CDM medium (DMEM/F12 medium [Life Technologies] supplemented with 15% knockout serum replacement [Life Technologies], 1% non-essential amino acids [Gibco], 1% penicillin/streptomycin [Gibco], 0.1 mM 2-mercaptoethanol [Gibco]) plus 10 μM ROCK inhibitor (Y-27632, Sigma), 10 ng/mL fibroblast growth factor 2 (FGF-2) (R&D Systems), and with 1 ng/mL of Dox (Sigma). After a day, the cells were washed once with PBS and changed to fresh K-CDM medium plus 10 ng/mL FGF-2 and with 1 ng/mL Dox. After 4 days of Dox induction, PAX7+ cells were isolated by FACS. PAX7+ cells were seeded onto a Matrigel-coated culture dish and cultured for an additional ∼10 days in the K-CDM medium supplemented with 1 ng/mL Dox and 10 ng/mL FGF-2. The medium was changed every 3 days and the cells were passaged by trypsinization every 3–4 days. After a total of ∼14 days of Dox induction, the cells were allowed to be confluent (>95%) in a Matrigel-coated 24-well dish. The cells were washed with PBS once and changed to K-CDM medium supplemented with 10 ng/mL hepatocyte growth factor (HGF) (R&D Systems) and 2 ng/mL insulin growth factor 1 (IGF1) (R&D Systems). The cells were cultured for 7–10 days with change of medium every second day.
Serum-free Monolayer Myogenic Differentiation of hPS Cells
The procedure was performed as described previously (Chal et al., 2015, Chal et al., 2016), and a detailed description is provided in Supplemental Experimental Procedures.
Expansion of hPS Cell-Derived Myogenic Progenitors from the Serum-free Monolayer Myogenic Differentiation
As described previously (Chal et al., 2016), after 30 days of serum-free monolayer myogenic differentiation, cells were harvested with trypsin, neutralized with FBS -containing medium (Gibco), and filtered through a 70 μm cell strainer. The cells were seeded onto fresh Matrigel-coated 12-well dishes (250,000 cells/well) with skeletal muscle growth medium-2 (SkGM; Lonza) supplemented with 10 μM Y-27632. After 24 hr, the cells were washed once with PBS and replaced with fresh SkGM medium. After expanding for 12 days, the cells were terminally differentiated by switching to low-glucose DMEM medium (Life Technologies) supplemented with 2% HS (Gibco) and culturing for 1 week.
Transplantation Studies
All animal experiments were carried out according to protocols approved by the University of Minnesota Institutional Animal Care and Use Committee. Cell preparations were injected directly into the tibialis anterior muscles of 6–7-week-old NSG male mice (Jackson Laboratory) that had been pre-injured with cardiotoxin (Sigma), at 500,000 cells/10 μL of PBS, as described previously (Darabi et al., 2012). After 24 hr, cells that had been pre-treated with 10 μM Y-27632 for 24 hr (CDM-H9 [day 25 and 30 culture], EXP-H9 [12 days of expansion in animal serum condition], and iPAX7+CDM-H9 [day 22]) were harvested using trypsin or cell-dissociation buffer (Gibco). Engraftment was assessed 6 weeks later.
qRT-PCR
A detailed description is provided in Supplemental Experimental Procedures.
Immunofluorescence Staining
A detailed description is provided in Supplemental Experimental Procedures.
Western Blot
A detailed description is provided in Supplemental Experimental Procedures.
Statistics
Differences between samples were assessed by using the Student's two-tailed t test for independent samples.
Author Contributions
J.K. designed and conducted experiments, analyzed the data, and wrote the manuscript. A.M. designed and conducted experiments, interpreted the data, and wrote the manuscript. S.C., V.K.P.O., J.W., and R.D. performed experiments. M.K. interpreted the data. R.C.R.P. supervised the study, analyzed the data, and wrote the manuscript.
Acknowledgments
This project was supported by the National Institute of Arthritis and Musculoskeletal and Skin Diseases of the NIH under Award Number R01 AR055299 and the Parent Project Muscular Dystrophy (PPMD grant no. 00031645). We thank the generous support from MyDirectives. The monoclonal antibody to MHC was obtained from the Developmental Studies Hybridoma Bank developed under the auspices of the NICHD and maintained by the University of Iowa. We thank Cynthia Dekay for assistance in graphic design.
Published: May 18, 2017
Footnotes
Supplemental Information includes Supplemental Experimental Procedures and three figures and can be found with this article online at http://dx.doi.org/10.1016/j.stemcr.2017.04.022.
Supplemental Information
References
- Abujarour R., Bennett M., Valamehr B., Lee T.T., Robinson M., Robbins D., Le T., Lai K., Flynn P. Myogenic differentiation of muscular dystrophy-specific induced pluripotent stem cells for use in drug discovery. Stem Cells Transl. Med. 2014;3:149–160. doi: 10.5966/sctm.2013-0095. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Albini S., Coutinho P., Malecova B., Giordani L., Savchenko A., Forcales S., Puri P.L. Epigenetic reprogramming of human ES cells into skeletal muscle cells and generation of contractile myospheres. Cell Rep. 2013;3:661–670. doi: 10.1016/j.celrep.2013.02.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barberi T., Bradbury M., Dincer Z., Panagiotakos G., Socci N.D., Studer L. Derivation of engraftable skeletal myoblasts from human embryonic stem cells. Nat. Med. 2007;13:642–648. doi: 10.1038/nm1533. [DOI] [PubMed] [Google Scholar]
- Borchin B., Chen J., Barberi T. Derivation and FACS-mediated purification of PAX3+/PAX7+ skeletal muscle precursors from human pluripotent stem cells. Stem Cell Reports. 2013;1:620–631. doi: 10.1016/j.stemcr.2013.10.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chal J., Oginuma M., Al Tanoury Z., Gobert B., Sumara O., Hick A., Bousson F., Zidouni Y., Mursch C., Moncuquet P. Differentiation of pluripotent stem cells to muscle fiber to model Duchenne muscular dystrophy. Nat. Biotech. 2015;33:962–969. doi: 10.1038/nbt.3297. [DOI] [PubMed] [Google Scholar]
- Chal J., Al Tanoury Z., Hestin M., Gobert B., Aivio S., Hick A., Cherrier T., Nesmith A.P., Parker K.K., Pourquie O. Generation of human muscle fibers and satellite-like cells from human pluripotent stem cells in vitro. Nat. Protoc. 2016;11:1833–1850. doi: 10.1038/nprot.2016.110. [DOI] [PubMed] [Google Scholar]
- Choi I.Y., Lim H.T., Estrellas K., Mula J., Cohen T.V., Zhang Y., Donnelly C.J., Richard J.P., Kim Y.J., Kim H. Concordant but varied phenotypes among Duchenne muscular dystrophy patient-specific myoblasts derived using a human iPSC-based model. Cell Rep. 2016;15:2301–2312. doi: 10.1016/j.celrep.2016.05.016. [DOI] [PubMed] [Google Scholar]
- Cimadamore F., Amador-Arjona A., Chen C., Huang C.T., Terskikh A.V. SOX2–LIN28/let-7 pathway regulates proliferation and neurogenesis in neural precursors. Proc. Natl. Acad. Sci. USA. 2013;110:E3017–E3026. doi: 10.1073/pnas.1220176110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Darabi R., Pan W., Bosnakovski D., Baik J., Kyba M., Perlingeiro R.R. Functional myogenic engraftment from mouse iPS cells. Stem Cell Rev. 2011;7:948–957. doi: 10.1007/s12015-011-9258-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Darabi R., Arpke R.W., Irion S., Dimos John T., Grskovic M., Kyba M., Perlingeiro R.R. Human ES- and iPS-derived myogenic progenitors restore DYSTROPHIN and improve contractility upon transplantation in dystrophic mice. Cell Stem Cell. 2012;10:610–619. doi: 10.1016/j.stem.2012.02.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Demestre M., Orth M., Föhr K.J., Achberger K., Ludolph A.C., Liebau S., Boeckers T.M. Formation and characterisation of neuromuscular junctions between hiPSC derived motoneurons and myotubes. Stem Cell Res. 2015;15:328–336. doi: 10.1016/j.scr.2015.07.005. [DOI] [PubMed] [Google Scholar]
- Dennis K., Uittenbogaard M., Chiaramello A., Moody S.A. Cloning and characterization of the 5'-flanking region of the rat neuron-specific class III beta-tubulin gene. Gene. 2002;294:269–277. doi: 10.1016/s0378-1119(02)00801-6. [DOI] [PubMed] [Google Scholar]
- Goudenege S., Lebel C., Huot N.B., Dufour C., Fujii I., Gekas J., Rousseau J., Tremblay J.P. Myoblasts derived from normal hESCs and dystrophic hiPSCs efficiently fuse with existing muscle fibers following transplantation. Mol. Ther. 2012;20:2153–2167. doi: 10.1038/mt.2012.188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gros J., Manceau M., Thome V., Marcelle C. A common somitic origin for embryonic muscle progenitors and satellite cells. Nature. 2005;435:954–958. doi: 10.1038/nature03572. [DOI] [PubMed] [Google Scholar]
- Hargus G., Cooper O., Deleidi M., Levy A., Lee K., Marlow E., Yow A., Soldner F., Hockemeyer D., Hallett P.J. Differentiated Parkinson patient-derived induced pluripotent stem cells grow in the adult rodent brain and reduce motor asymmetry in Parkinsonian rats. Proc. Natl. Acad. Sci. USA. 2010;107:15921–15926. doi: 10.1073/pnas.1010209107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kriks S., Shim J.-W., Piao J., Ganat Y.M., Wakeman D.R., Xie Z., Carrillo-Reid L., Auyeung G., Antonacci C., Buch A. Dopamine neurons derived from human ES cells efficiently engraft in animal models of Parkinson's disease. Nature. 2011;480:547–551. doi: 10.1038/nature10648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kroon E., Martinson L.A., Kadoya K., Bang A.G., Kelly O.G., Eliazer S., Young H., Richardson M., Smart N.G., Cunningham J. Pancreatic endoderm derived from human embryonic stem cells generates glucose-responsive insulin-secreting cells in vivo. Nat. Biotech. 2008;26:443–452. doi: 10.1038/nbt1393. [DOI] [PubMed] [Google Scholar]
- Laflamme M.A., Chen K.Y., Naumova A.V., Muskheli V., Fugate J.A., Dupras S.K., Reinecke H., Xu C., Hassanipour M., Police S. Cardiomyocytes derived from human embryonic stem cells in pro-survival factors enhance function of infarcted rat hearts. Nat. Biotech. 2007;25:1015–1024. doi: 10.1038/nbt1327. [DOI] [PubMed] [Google Scholar]
- Lagha M., Sato T., Bajard L., Daubas P., Esner M., Montarras D., Relaix F., Buckingham M. Regulation of skeletal muscle stem cell behavior by Pax3 and Pax7. Cold Spring Harb. Symp. Quant. Biol. 2008;73:307–315. doi: 10.1101/sqb.2008.73.006. [DOI] [PubMed] [Google Scholar]
- Lian X., Hsiao C., Wilson G., Zhu K., Hazeltine L.B., Azarin S.M., Raval K.K., Zhang J., Kamp T.J., Palecek S.P. Robust cardiomyocyte differentiation from human pluripotent stem cells via temporal modulation of canonical Wnt signaling. Proc. Natl. Acad. Sci. USA. 2012;109:E1848–E1857. doi: 10.1073/pnas.1200250109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mummery C.L., Zhang J., Ng E.S., Elliott D.A., Elefanty A.G., Kamp T.J. Differentiation of human ES and iPS cells to cardiomyocytes: a methods overview. Circ. Res. 2012;111:344–358. doi: 10.1161/CIRCRESAHA.110.227512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qin S., Zhang C.L. Role of Krüppel-like factor 4 in neurogenesis and radial neuronal migration in the developing cerebral cortex. Mol. Cell. Biol. 2012;32:4297–4305. doi: 10.1128/MCB.00838-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Salani S., Donadoni C., Rizzo F., Bresolin N., Comi G.P., Corti S. Generation of skeletal muscle cells from embryonic and induced pluripotent stem cells as an in vitro model and for therapy of muscular dystrophies. J. Cell. Mol. Med. 2012;16:1353–1364. doi: 10.1111/j.1582-4934.2011.01498.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schiaffino S., Reggiani C. Molecular diversity of myofibrillar proteins: gene regulation and functional significance. Physiol. Rev. 1996;76:371–423. doi: 10.1152/physrev.1996.76.2.371. [DOI] [PubMed] [Google Scholar]
- Seale P., Sabourin L.A., Girgis-Gabardo A., Mansouri A., Gruss P., Rudnicki M.A. Pax7 is required for the specification of myogenic satellite cells. Cell. 2000;102:777–786. doi: 10.1016/s0092-8674(00)00066-0. [DOI] [PubMed] [Google Scholar]
- Sebastiano V., Zhen H.H., Haddad B., Bashkirova E., Melo S.P., Wang P., Leung T.L., Siprashvili Z., Tichy A., Li J. Human COL7A1-corrected induced pluripotent stem cells for the treatment of recessive dystrophic epidermolysis bullosa. Sci. Transl. Med. 2014;6:264ra163. doi: 10.1126/scitranslmed.3009540. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shelton M., Metz J., Liu J., Carpenedo R.L., Demers S.-P., Stanford W.L., Skerjanc I.S. Derivation and expansion of PAX7-positive muscle progenitors from human and mouse embryonic stem cells. Stem Cell Reports. 2014;3:516–529. doi: 10.1016/j.stemcr.2014.07.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tabar V., Studer L. Pluripotent stem cells in regenerative medicine: challenges and recent progress. Nat. Rev. Genet. 2014;15:82–92. doi: 10.1038/nrg3563. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tan J.Y., Sriram G., Rufaihah A.J., Neoh K.G., Cao T. Efficient derivation of lateral plate and paraxial mesoderm subtypes from human embryonic stem cells through GSKi-mediated differentiation. Stem Cells Dev. 2013;22:1893–1906. doi: 10.1089/scd.2012.0590. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tanaka A., Woltjen K., Miyake K., Hotta A., Ikeya M., Yamamoto T., Nishino T., Shoji E., Sehara-Fujisawa A., Manabe Y. Efficient and reproducible myogenic differentiation from human iPS cells: prospects for modeling miyoshi myopathy in vitro. PLoS One. 2013;8:e61540. doi: 10.1371/journal.pone.0061540. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tedesco F.S., Gerli M.F.M., Perani L., Benedetti S., Ungaro F., Cassano M., Antonini S., Tagliafico E., Artusi V., Longa E. Transplantation of genetically corrected human iPSC-derived progenitors in mice with limb-girdle muscular dystrophy. Sci. Transl. Med. 2012;4:140ra89. doi: 10.1126/scitranslmed.3003541. [DOI] [PubMed] [Google Scholar]
- Wang Y., Zheng C.G., Jiang Y., Zhang J., Chen J., Yao C., Zhao Q., Liu S., Chen K., Du J. Genetic correction of [beta]-thalassemia patient-specific iPS cells and its use in improving hemoglobin production in irradiated SCID mice. Cell Res. 2012;22:637–648. doi: 10.1038/cr.2012.23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xi H., Fujiwara W., Gonzalez K., Jan M., Liebscher S., Van Handel B., Schenke-Layland K., Pyle A.D. In vivo human somitogenesis guides somite development from hPSCs. Cell Rep. 2017;18:1573–1585. doi: 10.1016/j.celrep.2017.01.040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Young C.S., Hicks M.R., Ermolova N.V., Nakano H., Jan M., Younesi S., Karumbayaram S., Kumagai-Cresse C., Wang D., Zack J.A. A single CRISPR-Cas9 deletion strategy that targets the majority of DMD patients restores dystrophin function in hiPSC-derived muscle cells. Cell Stem Cell. 2016;18:533–540. doi: 10.1016/j.stem.2016.01.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.