Abstract
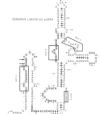
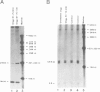
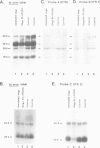
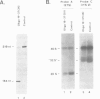
DNA oligonucleotide complementary to sequences in the 5' third of U3 snRNA were injected into Xenopus oocyte nuclei to disrupt endogenous U3 snRNA. The effect of this treatment on rRNA processing was examined. We found that some toads have a single rRNA processing pathway, whereas in other toads, two rRNA processing pathways can coexist in a single oocyte. U3 snRNA disruption in toads with the single rRNA processing pathway caused a reduction in 20S and '32S' pre-rRNA. In addition, in toads with two rRNA processing pathways, an increase in '36S' pre-rRNA of the second pathway is observed. This is the first in vivo demonstration that U3 snRNA plays a role in rRNA processing. Cleavage site #3 is at the boundary of ITS 1 and 5.8S and links all of the affected rRNA intermediates: 20S and '32S' are the products of site #3 cleavage in the first pathway and '36S' is the substrate for cleavage at site #3 in the second pathway. We postulate that U3 snRNP folds pre-rRNA into a conformation dictating correct cleavage at processing site #3.
Full text
PDF


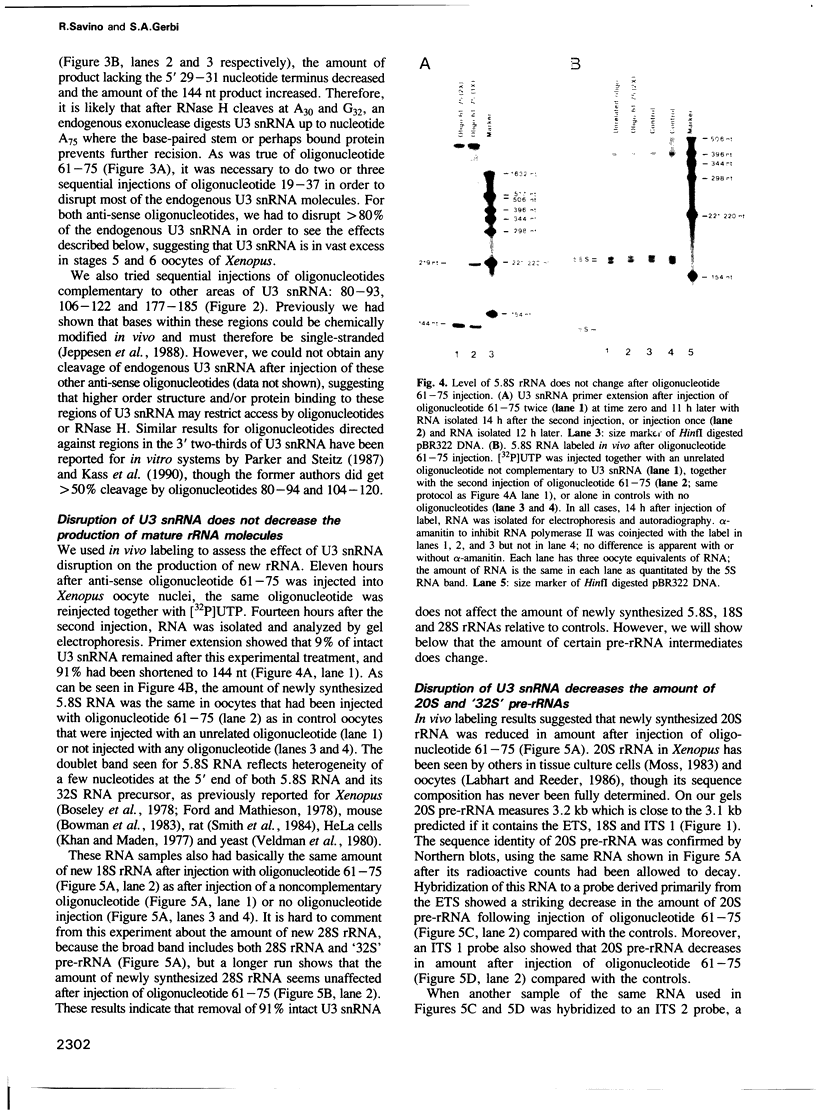






Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Anderson D. M., Smith L. D. Synthesis of heterogeneous nuclear RNA in full-grown oocytes of Xenopus laevis (Daudin). Cell. 1977 Jul;11(3):663–671. doi: 10.1016/0092-8674(77)90083-6. [DOI] [PubMed] [Google Scholar]
- Bachellerie J. P., Michot B., Raynal F. Recognition signals for mouse pre-rRNA processing. A potential role for U3 nucleolar RNA. Mol Biol Rep. 1983 May;9(1-2):79–86. doi: 10.1007/BF00777477. [DOI] [PubMed] [Google Scholar]
- Black D. L., Chabot B., Steitz J. A. U2 as well as U1 small nuclear ribonucleoproteins are involved in premessenger RNA splicing. Cell. 1985 Oct;42(3):737–750. doi: 10.1016/0092-8674(85)90270-3. [DOI] [PubMed] [Google Scholar]
- Boseley P. G., Tuyns A., Birnstiel M. L. Mapping of the Xenopus laevis 5.8S rDNA by restriction and DNA sequencing. Nucleic Acids Res. 1978 Apr;5(4):1121–1137. doi: 10.1093/nar/5.4.1121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bowman L. H., Goldman W. E., Goldberg G. I., Hebert M. B., Schlessinger D. Location of the initial cleavage sites in mouse pre-rRNA. Mol Cell Biol. 1983 Aug;3(8):1501–1510. doi: 10.1128/mcb.3.8.1501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cazenave C., Chevrier M., Nguyen T. T., Hélène C. Rate of degradation of [alpha]- and [beta]-oligodeoxynucleotides in Xenopus oocytes. Implications for anti-messenger strategies. Nucleic Acids Res. 1987 Dec 23;15(24):10507–10521. doi: 10.1093/nar/15.24.10507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Christofori G., Keller W. 3' cleavage and polyadenylation of mRNA precursors in vitro requires a poly(A) polymerase, a cleavage factor, and a snRNP. Cell. 1988 Sep 9;54(6):875–889. doi: 10.1016/s0092-8674(88)91263-9. [DOI] [PubMed] [Google Scholar]
- Dabeva M. D., Dudov K. P., Hadjiolov A. A., Emanuilov I., Todorov B. N. Intranuclear maturation pathways of rat liver ribosomal ribonucleic acids. Biochem J. 1976 Dec 15;160(3):495–503. doi: 10.1042/bj1600495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dahlberg A. E., Dahlberg J. E., Lund E., Tokimatsu H., Rabson A. B., Calvert P. C., Reynolds F., Zahalak M. Processing of the 5' end of Escherichia coli 16S ribosomal RNA. Proc Natl Acad Sci U S A. 1978 Aug;75(8):3598–3602. doi: 10.1073/pnas.75.8.3598. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dawid I. B., Wellauer P. K. A reinvestigation of 5' leads to 3' polarity in 40S ribosomal RNA precursor of Xenopus laevis. Cell. 1976 Jul;8(3):443–448. doi: 10.1016/0092-8674(76)90157-4. [DOI] [PubMed] [Google Scholar]
- Denhardt D. T. A membrane-filter technique for the detection of complementary DNA. Biochem Biophys Res Commun. 1966 Jun 13;23(5):641–646. doi: 10.1016/0006-291x(66)90447-5. [DOI] [PubMed] [Google Scholar]
- Dumont J. N. Oogenesis in Xenopus laevis (Daudin). I. Stages of oocyte development in laboratory maintained animals. J Morphol. 1972 Feb;136(2):153–179. doi: 10.1002/jmor.1051360203. [DOI] [PubMed] [Google Scholar]
- Epstein P., Reddy R., Busch H. Multiple states of U3 RNA in Novikoff hepatoma nucleoli. Biochemistry. 1984 Nov 6;23(23):5421–5425. doi: 10.1021/bi00318a007. [DOI] [PubMed] [Google Scholar]
- Ford P. J., Mathieson T. The nucleotide sequences of 5.8-S ribosomal RNA from Xenopus laevis and Xenopus borealis. Eur J Biochem. 1978 Jun 1;87(1):199–214. doi: 10.1111/j.1432-1033.1978.tb12367.x. [DOI] [PubMed] [Google Scholar]
- Frendewey D., Krämer A., Keller W. Different small nuclear ribonucleoprotein particles are involved in different steps of splicing complex formation. Cold Spring Harb Symp Quant Biol. 1987;52:287–298. doi: 10.1101/sqb.1987.052.01.034. [DOI] [PubMed] [Google Scholar]
- Gegenheimer P., Watson N., Apirion D. Multiple pathways for primary processing of ribosomal RNA in Escherichia coli. J Biol Chem. 1977 May 10;252(9):3064–3073. [PubMed] [Google Scholar]
- Gerbi S. A., Jeppesen C., Stebbins-Boaz B., Ares M., Jr Evolution of eukaryotic rRNA: constraints imposed by RNA interactions. Cold Spring Harb Symp Quant Biol. 1987;52:709–719. doi: 10.1101/sqb.1987.052.01.080. [DOI] [PubMed] [Google Scholar]
- Hamm J., Dathan N. A., Mattaj I. W. Functional analysis of mutant Xenopus U2 snRNAs. Cell. 1989 Oct 6;59(1):159–169. doi: 10.1016/0092-8674(89)90878-7. [DOI] [PubMed] [Google Scholar]
- Hannon G. J., Maroney P. A., Branch A., Benenfield B. J., Robertson H. D., Nilsen T. W. Accurate processing of human pre-rRNA in vitro. Mol Cell Biol. 1989 Oct;9(10):4422–4431. doi: 10.1128/mcb.9.10.4422. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hughes J. M., Konings D. A., Cesareni G. The yeast homologue of U3 snRNA. EMBO J. 1987 Jul;6(7):2145–2155. doi: 10.1002/j.1460-2075.1987.tb02482.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jeppesen C., Stebbins-Boaz B., Gerbi S. A. Nucleotide sequence determination and secondary structure of Xenopus U3 snRNA. Nucleic Acids Res. 1988 Mar 25;16(5):2127–2148. doi: 10.1093/nar/16.5.2127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kass S., Tyc K., Steitz J. A., Sollner-Webb B. The U3 small nucleolar ribonucleoprotein functions in the first step of preribosomal RNA processing. Cell. 1990 Mar 23;60(6):897–908. doi: 10.1016/0092-8674(90)90338-f. [DOI] [PubMed] [Google Scholar]
- Khan M. S., Maden B. E. Nucleotide sequence relationships between vertebrate 5.8 S ribosomal RNAs. Nucleic Acids Res. 1977 Jul;4(7):2495–2505. doi: 10.1093/nar/4.7.2495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- King T. C., Schlessinger D. S1 nuclease mapping analysis of ribosomal RNA processing in wild type and processing deficient Escherichia coli. J Biol Chem. 1983 Oct 10;258(19):12034–12042. [PubMed] [Google Scholar]
- King T. C., Sirdeshmukh R., Schlessinger D. RNase III cleavage is obligate for maturation but not for function of Escherichia coli pre-23S rRNA. Proc Natl Acad Sci U S A. 1984 Jan;81(1):185–188. doi: 10.1073/pnas.81.1.185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- King T. C., Sirdeskmukh R., Schlessinger D. Nucleolytic processing of ribonucleic acid transcripts in procaryotes. Microbiol Rev. 1986 Dec;50(4):428–451. doi: 10.1128/mr.50.4.428-451.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krainer A. R., Maniatis T. Multiple factors including the small nuclear ribonucleoproteins U1 and U2 are necessary for pre-mRNA splicing in vitro. Cell. 1985 Oct;42(3):725–736. doi: 10.1016/0092-8674(85)90269-7. [DOI] [PubMed] [Google Scholar]
- Krämer A., Keller W., Appel B., Lührmann R. The 5' terminus of the RNA moiety of U1 small nuclear ribonucleoprotein particles is required for the splicing of messenger RNA precursors. Cell. 1984 Aug;38(1):299–307. doi: 10.1016/0092-8674(84)90551-8. [DOI] [PubMed] [Google Scholar]
- Kuhn A., Grummt I. 3'-end formation of mouse pre-rRNA involves both transcription termination and a specific processing reaction. Genes Dev. 1989 Feb;3(2):224–231. doi: 10.1101/gad.3.2.224. [DOI] [PubMed] [Google Scholar]
- Labhart P., Reeder R. H. Characterization of three sites of RNA 3' end formation in the Xenopus ribosomal gene spacer. Cell. 1986 May 9;45(3):431–443. doi: 10.1016/0092-8674(86)90329-6. [DOI] [PubMed] [Google Scholar]
- Lamond A. I., Sproat B., Ryder U., Hamm J. Probing the structure and function of U2 snRNP with antisense oligonucleotides made of 2'-OMe RNA. Cell. 1989 Jul 28;58(2):383–390. doi: 10.1016/0092-8674(89)90852-0. [DOI] [PubMed] [Google Scholar]
- Lillie J. W., Green M., Green M. R. An adenovirus E1a protein region required for transformation and transcriptional repression. Cell. 1986 Sep 26;46(7):1043–1051. doi: 10.1016/0092-8674(86)90704-x. [DOI] [PubMed] [Google Scholar]
- Maser R. L., Calvet J. P. U3 small nuclear RNA can be psoralen-cross-linked in vivo to the 5' external transcribed spacer of pre-ribosomal-RNA. Proc Natl Acad Sci U S A. 1989 Sep;86(17):6523–6527. doi: 10.1073/pnas.86.17.6523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moss T. A transcriptional function for the repetitive ribosomal spacer in Xenopus laevis. Nature. 1983 Mar 17;302(5905):223–228. doi: 10.1038/302223a0. [DOI] [PubMed] [Google Scholar]
- Mowry K. L., Steitz J. A. snRNP mediators of 3' end processing: functional fossils? Trends Biochem Sci. 1988 Nov;13(11):447–451. doi: 10.1016/0968-0004(88)90220-4. [DOI] [PubMed] [Google Scholar]
- Pan Z. Q., Prives C. Assembly of functional U1 and U2 human-amphibian hybrid snRNPs in Xenopus laevis oocytes. Science. 1988 Sep 9;241(4871):1328–1331. doi: 10.1126/science.2970672. [DOI] [PubMed] [Google Scholar]
- Parker K. A., Bruzik J. P., Steitz J. A. An in vitro interaction between the human U3 snRNP and 28S rRNA sequences near the alpha-sarcin site. Nucleic Acids Res. 1988 Nov 25;16(22):10493–10509. doi: 10.1093/nar/16.22.10493. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parker K. A., Steitz J. A. Structural analysis of the human U3 ribonucleoprotein particle reveal a conserved sequence available for base pairing with pre-rRNA. Mol Cell Biol. 1987 Aug;7(8):2899–2913. doi: 10.1128/mcb.7.8.2899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Porter G. L., Brennwald P. J., Holm K. A., Wise J. A. The sequence of U3 from Schizosaccharomyces pombe suggests structural divergence of this snRNA between metazoans and unicellular eukaryotes. Nucleic Acids Res. 1988 Nov 11;16(21):10131–10152. doi: 10.1093/nar/16.21.10131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Prestayko A. W., Tonato M., Busch H. Low molecular weight RNA associated with 28 s nucleolar RNA. J Mol Biol. 1970 Feb 14;47(3):505–515. doi: 10.1016/0022-2836(70)90318-9. [DOI] [PubMed] [Google Scholar]
- Purtell M. J., Anthony D. D. Changes in ribosomal RNA processing paths in resting and phytohemagglutinin-stimulated guinea pig lymphocytes. Proc Natl Acad Sci U S A. 1975 Sep;72(9):3315–3319. doi: 10.1073/pnas.72.9.3315. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rebagliati M. R., Melton D. A. Antisense RNA injections in fertilized frog eggs reveal an RNA duplex unwinding activity. Cell. 1987 Feb 27;48(4):599–605. doi: 10.1016/0092-8674(87)90238-8. [DOI] [PubMed] [Google Scholar]
- Shuttleworth J., Colman A. Antisense oligonucleotide-directed cleavage of mRNA in Xenopus oocytes and eggs. EMBO J. 1988 Feb;7(2):427–434. doi: 10.1002/j.1460-2075.1988.tb02830.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith S. D., Banerjee N., Sitz T. O. Gene heterogeneity: a basis for alternative 5.8S rRNA processing. Biochemistry. 1984 Jul 31;23(16):3648–3652. doi: 10.1021/bi00311a011. [DOI] [PubMed] [Google Scholar]
- Stroke I. L., Weiner A. M. The 5' end of U3 snRNA can be crosslinked in vivo to the external transcribed spacer of rat ribosomal RNA precursors. J Mol Biol. 1989 Dec 5;210(3):497–512. doi: 10.1016/0022-2836(89)90126-5. [DOI] [PubMed] [Google Scholar]
- Tague B. W., Gerbi S. A. Processing of the large rRNA precursor: two proposed categories of RNA-RNA interactions in eukaryotes. J Mol Evol. 1984;20(3-4):362–367. doi: 10.1007/BF02104742. [DOI] [PubMed] [Google Scholar]
- Toniolo D., Basilico C. Processing of ribosomal RNA in a temperature sensitive mutant of BHK cells. Biochim Biophys Acta. 1976 Apr 2;425(4):409–418. doi: 10.1016/0005-2787(76)90005-8. [DOI] [PubMed] [Google Scholar]
- Tyc K., Steitz J. A. U3, U8 and U13 comprise a new class of mammalian snRNPs localized in the cell nucleolus. EMBO J. 1989 Oct;8(10):3113–3119. doi: 10.1002/j.1460-2075.1989.tb08463.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vance V. B., Thompson E. A., Bowman L. H. Transfection of mouse ribosomal DNA into rat cells: faithful transcription and processing. Nucleic Acids Res. 1985 Oct 25;13(20):7499–7513. doi: 10.1093/nar/13.20.7499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Veldman G. M., Brand R. C., Klootwijk J., Planta R. Some characteristics of processing sites in ribosomal precursor RNA of yeast. Nucleic Acids Res. 1980 Jul 11;8(13):2907–2920. doi: 10.1093/nar/8.13.2907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weinberg R. A., Penman S. Processing of 45 s nucleolar RNA. J Mol Biol. 1970 Jan 28;47(2):169–178. doi: 10.1016/0022-2836(70)90337-2. [DOI] [PubMed] [Google Scholar]
- Weinberg R. A., Penman S. Small molecular weight monodisperse nuclear RNA. J Mol Biol. 1968 Dec;38(3):289–304. doi: 10.1016/0022-2836(68)90387-2. [DOI] [PubMed] [Google Scholar]
- Wellauer P. K., Dawid I. B. Secondary structure maps of ribosomal RNA and DNA. I. Processing of Xenopus laevis ribosomal RNA and structure of single-stranded ribosomal DNA. J Mol Biol. 1974 Oct 25;89(2):379–395. doi: 10.1016/0022-2836(74)90526-9. [DOI] [PubMed] [Google Scholar]
- Winicov I. Alternate temporal order in ribosomal RNA maturation. J Mol Biol. 1976 Jan 15;100(2):141–155. doi: 10.1016/s0022-2836(76)80145-3. [DOI] [PubMed] [Google Scholar]
- Wise J. A., Weiner A. M. Dictyostelium small nuclear RNA D2 is homologous to rat nucleolar RNA U3 and is encoded by a dispersed multigene family. Cell. 1980 Nov;22(1 Pt 1):109–118. doi: 10.1016/0092-8674(80)90159-2. [DOI] [PubMed] [Google Scholar]
- Zieve G., Penman S. Small RNA species of the HeLa cell: metabolism and subcellular localization. Cell. 1976 May;8(1):19–31. doi: 10.1016/0092-8674(76)90181-1. [DOI] [PubMed] [Google Scholar]
- Zillmann M., Zapp M. L., Berget S. M. Gel electrophoretic isolation of splicing complexes containing U1 small nuclear ribonucleoprotein particles. Mol Cell Biol. 1988 Feb;8(2):814–821. doi: 10.1128/mcb.8.2.814. [DOI] [PMC free article] [PubMed] [Google Scholar]