Abstract
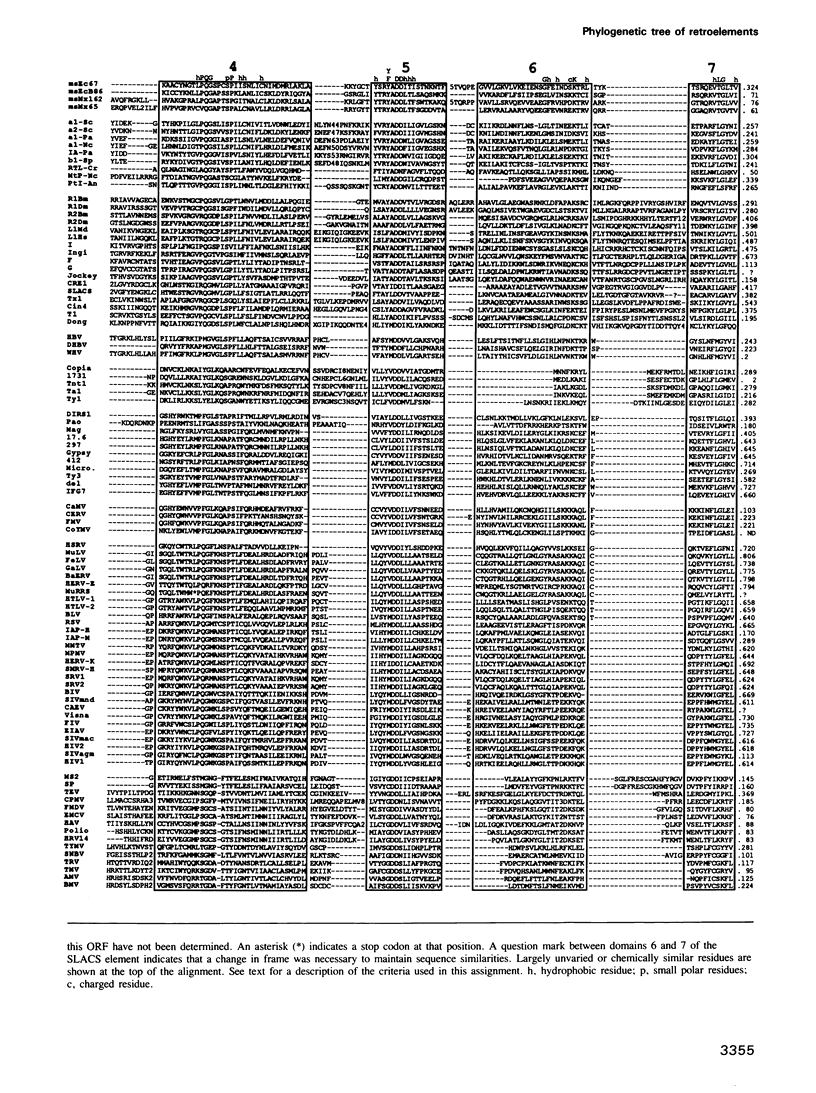
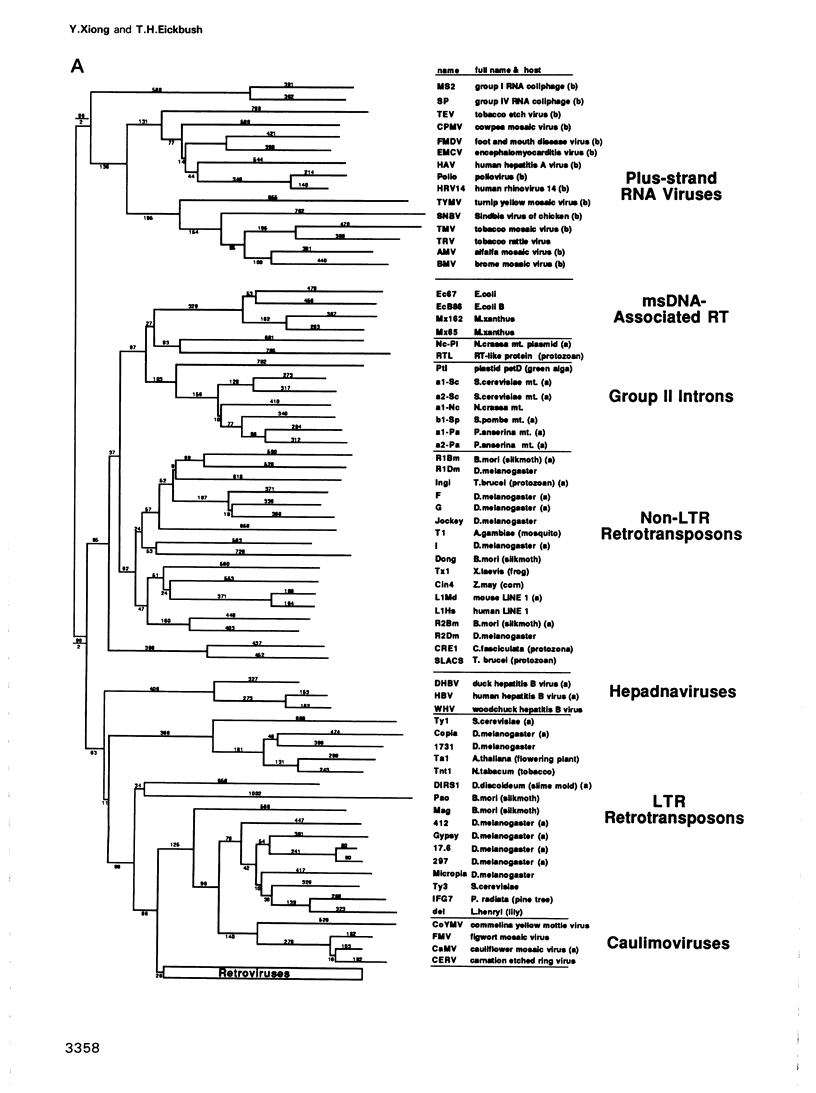
To study the evolutionary relationship of reverse transcriptase (RT) containing genetic elements, a phylogenetic tree of 82 retroelements from animals, plants, protozoans and bacteria was constructed. The tree was based on seven amino acid domains totalling 178 residues identified in all RTs. We have also identified these seven domains in the RNA-directed RNA polymerases from various plus-strand RNA viruses. The sequence similarity of these RNA polymerases to RT suggests that these two enzymes evolved from a common ancestor, and thus RNA polymerase can be used as an outgroup to root the RT tree. A comparison of the genetic organization of the various RT containing elements and their position on the tree allows several inferences concerning the origin and evolution of these elements. The most probable ancestor of current retroelements was a retrotransposable element with both gag-like and pol-like genes. On one major branch of the tree, organelle and bacterial sequences (e.g. group II introns and bacterial msDNA) appear to have captured the RT sequences from retrotransposons which lack long terminal repeats (LTRs). On the other major branch, acquisition of LTRs gave rise to two distinct groups of LTR retrotransposons and three groups of viruses: retroviruses, hepadnaviruses and caulimoviruses.
Full text
PDF







Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Aksoy S., Williams S., Chang S., Richards F. F. SLACS retrotransposon from Trypanosoma brucei gambiense is similar to mammalian LINEs. Nucleic Acids Res. 1990 Feb 25;18(4):785–792. doi: 10.1093/nar/18.4.785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Argos P. A sequence motif in many polymerases. Nucleic Acids Res. 1988 Nov 11;16(21):9909–9916. doi: 10.1093/nar/16.21.9909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baltimore D. RNA-dependent DNA polymerase in virions of RNA tumour viruses. Nature. 1970 Jun 27;226(5252):1209–1211. doi: 10.1038/2261209a0. [DOI] [PubMed] [Google Scholar]
- Besansky N. J. A retrotransposable element from the mosquito Anopheles gambiae . Mol Cell Biol. 1990 Mar;10(3):863–871. doi: 10.1128/mcb.10.3.863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boeke J. D., Corces V. G. Transcription and reverse transcription of retrotransposons. Annu Rev Microbiol. 1989;43:403–434. doi: 10.1146/annurev.mi.43.100189.002155. [DOI] [PubMed] [Google Scholar]
- Boer P. H., Gray M. W. Genes encoding a subunit of respiratory NADH dehydrogenase (ND1) and a reverse transcriptase-like protein (RTL) are linked to ribosomal RNA gene pieces in Chlamydomonas reinhardtii mitochondrial DNA. EMBO J. 1988 Nov;7(11):3501–3508. doi: 10.1002/j.1460-2075.1988.tb03226.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cappello J., Handelsman K., Lodish H. F. Sequence of Dictyostelium DIRS-1: an apparent retrotransposon with inverted terminal repeats and an internal circle junction sequence. Cell. 1985 Nov;43(1):105–115. doi: 10.1016/0092-8674(85)90016-9. [DOI] [PubMed] [Google Scholar]
- Chakrabarti L., Guyader M., Alizon M., Daniel M. D., Desrosiers R. C., Tiollais P., Sonigo P. Sequence of simian immunodeficiency virus from macaque and its relationship to other human and simian retroviruses. Nature. 1987 Aug 6;328(6130):543–547. doi: 10.1038/328543a0. [DOI] [PubMed] [Google Scholar]
- Clare J., Farabaugh P. Nucleotide sequence of a yeast Ty element: evidence for an unusual mechanism of gene expression. Proc Natl Acad Sci U S A. 1985 May;82(9):2829–2833. doi: 10.1073/pnas.82.9.2829. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coletta M., Amiconi G., Bellelli A., Bertollini A., Carsky J., Castagnola M., Condò S., Brunori M. Alteration of T-state binding properties of naturally glycated hemoglobin, HbA1c. J Mol Biol. 1988 Sep 5;203(1):233–239. doi: 10.1016/0022-2836(88)90104-0. [DOI] [PubMed] [Google Scholar]
- Covey S. N. Amino acid sequence homology in gag region of reverse transcribing elements and the coat protein gene of cauliflower mosaic virus. Nucleic Acids Res. 1986 Jan 24;14(2):623–633. doi: 10.1093/nar/14.2.623. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Delassus S., Sonigo P., Wain-Hobson S. Genetic organization of gibbon ape leukemia virus. Virology. 1989 Nov;173(1):205–213. doi: 10.1016/0042-6822(89)90236-5. [DOI] [PubMed] [Google Scholar]
- Donahue P. R., Hoover E. A., Beltz G. A., Riedel N., Hirsch V. M., Overbaugh J., Mullins J. I. Strong sequence conservation among horizontally transmissible, minimally pathogenic feline leukemia viruses. J Virol. 1988 Mar;62(3):722–731. doi: 10.1128/jvi.62.3.722-731.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Doolittle R. F., Feng D. F., Johnson M. S., McClure M. A. Origins and evolutionary relationships of retroviruses. Q Rev Biol. 1989 Mar;64(1):1–30. doi: 10.1086/416128. [DOI] [PubMed] [Google Scholar]
- Fawcett D. H., Lister C. K., Kellett E., Finnegan D. J. Transposable elements controlling I-R hybrid dysgenesis in D. melanogaster are similar to mammalian LINEs. Cell. 1986 Dec 26;47(6):1007–1015. doi: 10.1016/0092-8674(86)90815-9. [DOI] [PubMed] [Google Scholar]
- Feng D. F., Doolittle R. F. Progressive sequence alignment as a prerequisite to correct phylogenetic trees. J Mol Evol. 1987;25(4):351–360. doi: 10.1007/BF02603120. [DOI] [PubMed] [Google Scholar]
- Field D. J., Sommerfield A., Saville B. J., Collins R. A. A group II intron in the Neurospora mitochondrial coI gene: nucleotide sequence and implications for splicing and molecular evolution. Nucleic Acids Res. 1989 Nov 25;17(22):9087–9099. doi: 10.1093/nar/17.22.9087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Finnegan D. J. Retroviruses and transposable elements--which came first? Nature. 1983 Mar 10;302(5904):105–106. doi: 10.1038/302105a0. [DOI] [PubMed] [Google Scholar]
- Finnegan D. J. Transposable elements in eukaryotes. Int Rev Cytol. 1985;93:281–326. doi: 10.1016/s0074-7696(08)61376-5. [DOI] [PubMed] [Google Scholar]
- Fourcade-Peronnet F., d'Auriol L., Becker J., Galibert F., Best-Belpomme M. Primary structure and functional organization of Drosophila 1731 retrotransposon. Nucleic Acids Res. 1988 Jul 11;16(13):6113–6125. doi: 10.1093/nar/16.13.6113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fukasawa M., Miura T., Hasegawa A., Morikawa S., Tsujimoto H., Miki K., Kitamura T., Hayami M. Sequence of simian immunodeficiency virus from African green monkey, a new member of the HIV/SIV group. Nature. 1988 Jun 2;333(6172):457–461. doi: 10.1038/333457a0. [DOI] [PubMed] [Google Scholar]
- Gabriel A., Yen T. J., Schwartz D. C., Smith C. L., Boeke J. D., Sollner-Webb B., Cleveland D. W. A rapidly rearranging retrotransposon within the miniexon gene locus of Crithidia fasciculata. Mol Cell Biol. 1990 Feb;10(2):615–624. doi: 10.1128/mcb.10.2.615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garrett J. E., Knutzon D. S., Carroll D. Composite transposable elements in the Xenopus laevis genome. Mol Cell Biol. 1989 Jul;9(7):3018–3027. doi: 10.1128/mcb.9.7.3018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garvey K. J., Oberste M. S., Elser J. E., Braun M. J., Gonda M. A. Nucleotide sequence and genome organization of biologically active proviruses of the bovine immunodeficiency-like virus. Virology. 1990 Apr;175(2):391–409. doi: 10.1016/0042-6822(90)90424-p. [DOI] [PubMed] [Google Scholar]
- Girones R., Cote P. J., Hornbuckle W. E., Tennant B. C., Gerin J. L., Purcell R. H., Miller R. H. Complete nucleotide sequence of a molecular clone of woodchuck hepatitis virus that is infectious in the natural host. Proc Natl Acad Sci U S A. 1989 Mar;86(6):1846–1849. doi: 10.1073/pnas.86.6.1846. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grandbastien M. A., Spielmann A., Caboche M. Tnt1, a mobile retroviral-like transposable element of tobacco isolated by plant cell genetics. Nature. 1989 Jan 26;337(6205):376–380. doi: 10.1038/337376a0. [DOI] [PubMed] [Google Scholar]
- Habili N., Symons R. H. Evolutionary relationship between luteoviruses and other RNA plant viruses based on sequence motifs in their putative RNA polymerases and nucleic acid helicases. Nucleic Acids Res. 1989 Dec 11;17(23):9543–9555. doi: 10.1093/nar/17.23.9543. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hamilton W. D., Boccara M., Robinson D. J., Baulcombe D. C. The complete nucleotide sequence of tobacco rattle virus RNA-1. J Gen Virol. 1987 Oct;68(Pt 10):2563–2575. doi: 10.1099/0022-1317-68-10-2563. [DOI] [PubMed] [Google Scholar]
- Hansen L. J., Chalker D. L., Sandmeyer S. B. Ty3, a yeast retrotransposon associated with tRNA genes, has homology to animal retroviruses. Mol Cell Biol. 1988 Dec;8(12):5245–5256. doi: 10.1128/mcb.8.12.5245. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hattori M., Kuhara S., Takenaka O., Sakaki Y. L1 family of repetitive DNA sequences in primates may be derived from a sequence encoding a reverse transcriptase-related protein. Nature. 1986 Jun 5;321(6070):625–628. doi: 10.1038/321625a0. [DOI] [PubMed] [Google Scholar]
- Hull R., Sadler J., Longstaff M. The sequence of carnation etched ring virus DNA: comparison with cauliflower mosaic virus and retroviruses. EMBO J. 1986 Dec 1;5(12):3083–3090. doi: 10.1002/j.1460-2075.1986.tb04614.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Inouye S., Herzer P. J., Inouye M. Two independent retrons with highly diverse reverse transcriptases in Myxococcus xanthus. Proc Natl Acad Sci U S A. 1990 Feb;87(3):942–945. doi: 10.1073/pnas.87.3.942. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Inouye S., Hsu M. Y., Eagle S., Inouye M. Reverse transcriptase associated with the biosynthesis of the branched RNA-linked msDNA in Myxococcus xanthus. Cell. 1989 Feb 24;56(4):709–717. doi: 10.1016/0092-8674(89)90593-x. [DOI] [PubMed] [Google Scholar]
- Jakubczak J. L., Xiong Y., Eickbush T. H. Type I (R1) and type II (R2) ribosomal DNA insertions of Drosophila melanogaster are retrotransposable elements closely related to those of Bombyx mori. J Mol Biol. 1990 Mar 5;212(1):37–52. doi: 10.1016/0022-2836(90)90303-4. [DOI] [PubMed] [Google Scholar]
- Kamer G., Argos P. Primary structural comparison of RNA-dependent polymerases from plant, animal and bacterial viruses. Nucleic Acids Res. 1984 Sep 25;12(18):7269–7282. doi: 10.1093/nar/12.18.7269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kück U. The intron of a plastid gene from a green alga contains an open reading frame for a reverse transcriptase-like enzyme. Mol Gen Genet. 1989 Aug;218(2):257–265. doi: 10.1007/BF00331276. [DOI] [PubMed] [Google Scholar]
- Lambowitz A. M. Infectious introns. Cell. 1989 Feb 10;56(3):323–326. doi: 10.1016/0092-8674(89)90232-8. [DOI] [PubMed] [Google Scholar]
- Lampson B. C., Sun J., Hsu M. Y., Vallejo-Ramirez J., Inouye S., Inouye M. Reverse transcriptase in a clinical strain of Escherichia coli: production of branched RNA-linked msDNA. Science. 1989 Feb 24;243(4894 Pt 1):1033–1038. doi: 10.1126/science.2466332. [DOI] [PubMed] [Google Scholar]
- Lazcano A., Fastag J., Gariglio P., Ramírez C., Oró J. On the early evolution of RNA polymerase. J Mol Evol. 1988;27(4):365–376. doi: 10.1007/BF02101199. [DOI] [PubMed] [Google Scholar]
- Lim D., Maas W. K. Reverse transcriptase-dependent synthesis of a covalently linked, branched DNA-RNA compound in E. coli B. Cell. 1989 Mar 10;56(5):891–904. doi: 10.1016/0092-8674(89)90693-4. [DOI] [PubMed] [Google Scholar]
- Loeb D. D., Padgett R. W., Hardies S. C., Shehee W. R., Comer M. B., Edgell M. H., Hutchison C. A., 3rd The sequence of a large L1Md element reveals a tandemly repeated 5' end and several features found in retrotransposons. Mol Cell Biol. 1986 Jan;6(1):168–182. doi: 10.1128/mcb.6.1.168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Michaille J. J., Mathavan S., Gaillard J., Garel A. The complete sequence of mag, a new retrotransposon in Bombyx mori. Nucleic Acids Res. 1990 Feb 11;18(3):674–674. doi: 10.1093/nar/18.3.674. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Michel F., Lang B. F. Mitochondrial class II introns encode proteins related to the reverse transcriptases of retroviruses. Nature. 1985 Aug 15;316(6029):641–643. doi: 10.1038/316641a0. [DOI] [PubMed] [Google Scholar]
- Mietz J. A., Grossman Z., Lueders K. K., Kuff E. L. Nucleotide sequence of a complete mouse intracisternal A-particle genome: relationship to known aspects of particle assembly and function. J Virol. 1987 Oct;61(10):3020–3029. doi: 10.1128/jvi.61.10.3020-3029.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mount S. M., Rubin G. M. Complete nucleotide sequence of the Drosophila transposable element copia: homology between copia and retroviral proteins. Mol Cell Biol. 1985 Jul;5(7):1630–1638. doi: 10.1128/mcb.5.7.1630. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nargang F. E., Bell J. B., Stohl L. L., Lambowitz A. M. The DNA sequence and genetic organization of a Neurospora mitochondrial plasmid suggest a relationship to introns and mobile elements. Cell. 1984 Sep;38(2):441–453. doi: 10.1016/0092-8674(84)90499-9. [DOI] [PubMed] [Google Scholar]
- Oda T., Ikeda S., Watanabe S., Hatsushika M., Akiyama K., Mitsunobu F. Molecular cloning, complete nucleotide sequence, and gene structure of the provirus genome of a retrovirus produced in a human lymphoblastoid cell line. Virology. 1988 Dec;167(2):468–476. [PubMed] [Google Scholar]
- Olmsted R. A., Hirsch V. M., Purcell R. H., Johnson P. R. Nucleotide sequence analysis of feline immunodeficiency virus: genome organization and relationship to other lentiviruses. Proc Natl Acad Sci U S A. 1989 Oct;86(20):8088–8092. doi: 10.1073/pnas.86.20.8088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ono M., Yasunaga T., Miyata T., Ushikubo H. Nucleotide sequence of human endogenous retrovirus genome related to the mouse mammary tumor virus genome. J Virol. 1986 Nov;60(2):589–598. doi: 10.1128/jvi.60.2.589-598.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poch O., Sauvaget I., Delarue M., Tordo N. Identification of four conserved motifs among the RNA-dependent polymerase encoding elements. EMBO J. 1989 Dec 1;8(12):3867–3874. doi: 10.1002/j.1460-2075.1989.tb08565.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Priimägi A. F., Mizrokhi L. J., Ilyin Y. V. The Drosophila mobile element jockey belongs to LINEs and contains coding sequences homologous to some retroviral proteins. Gene. 1988 Oct 30;70(2):253–262. doi: 10.1016/0378-1119(88)90197-7. [DOI] [PubMed] [Google Scholar]
- Richins R. D., Scholthof H. B., Shepherd R. J. Sequence of figwort mosaic virus DNA (caulimovirus group). Nucleic Acids Res. 1987 Oct 26;15(20):8451–8466. doi: 10.1093/nar/15.20.8451. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rogers J. H. The origin and evolution of retroposons. Int Rev Cytol. 1985;93:187–279. doi: 10.1016/s0074-7696(08)61375-3. [DOI] [PubMed] [Google Scholar]
- Saigo K., Kugimiya W., Matsuo Y., Inouye S., Yoshioka K., Yuki S. Identification of the coding sequence for a reverse transcriptase-like enzyme in a transposable genetic element in Drosophila melanogaster. Nature. 1984 Dec 13;312(5995):659–661. doi: 10.1038/312659a0. [DOI] [PubMed] [Google Scholar]
- Saitou N., Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987 Jul;4(4):406–425. doi: 10.1093/oxfordjournals.molbev.a040454. [DOI] [PubMed] [Google Scholar]
- Schmidt M., Wirth T., Kröger B., Horak I. Structure and genomic organization of a new family of murine retrovirus-related DNA sequences (MuRRS). Nucleic Acids Res. 1985 May 24;13(10):3461–3470. doi: 10.1093/nar/13.10.3461. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schwarz-Sommer Z., Leclercq L., Göbel E., Saedler H. Cin4, an insert altering the structure of the A1 gene in Zea mays, exhibits properties of nonviral retrotransposons. EMBO J. 1987 Dec 20;6(13):3873–3880. doi: 10.1002/j.1460-2075.1987.tb02727.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smyth D. R., Kalitsis P., Joseph J. L., Sentry J. W. Plant retrotransposon from Lilium henryi is related to Ty3 of yeast and the gypsy group of Drosophila. Proc Natl Acad Sci U S A. 1989 Jul;86(13):5015–5019. doi: 10.1073/pnas.86.13.5015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sonigo P., Barker C., Hunter E., Wain-Hobson S. Nucleotide sequence of Mason-Pfizer monkey virus: an immunosuppressive D-type retrovirus. Cell. 1986 May 9;45(3):375–385. doi: 10.1016/0092-8674(86)90323-5. [DOI] [PubMed] [Google Scholar]
- Temin H. M., Mizutani S. RNA-dependent DNA polymerase in virions of Rous sarcoma virus. Nature. 1970 Jun 27;226(5252):1211–1213. doi: 10.1038/2261211a0. [DOI] [PubMed] [Google Scholar]
- Temin H. M. Origin of retroviruses from cellular moveable genetic elements. Cell. 1980 Oct;21(3):599–600. doi: 10.1016/0092-8674(80)90420-1. [DOI] [PubMed] [Google Scholar]
- Temin H. M. Reverse transcriptases. Retrons in bacteria. Nature. 1989 May 25;339(6222):254–255. doi: 10.1038/339254a0. [DOI] [PubMed] [Google Scholar]
- Thayer R. M., Power M. D., Bryant M. L., Gardner M. B., Barr P. J., Luciw P. A. Sequence relationships of type D retroviruses which cause simian acquired immunodeficiency syndrome. Virology. 1987 Apr;157(2):317–329. doi: 10.1016/0042-6822(87)90274-1. [DOI] [PubMed] [Google Scholar]
- Toh H., Hayashida H., Miyata T. Sequence homology between retroviral reverse transcriptase and putative polymerases of hepatitis B virus and cauliflower mosaic virus. 1983 Oct 27-Nov 2Nature. 305(5937):827–829. doi: 10.1038/305827a0. [DOI] [PubMed] [Google Scholar]
- Toh H., Kikuno R., Hayashida H., Miyata T., Kugimiya W., Inouye S., Yuki S., Saigo K. Close structural resemblance between putative polymerase of a Drosophila transposable genetic element 17.6 and pol gene product of Moloney murine leukaemia virus. EMBO J. 1985 May;4(5):1267–1272. doi: 10.1002/j.1460-2075.1985.tb03771.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsujimoto H., Hasegawa A., Maki N., Fukasawa M., Miura T., Speidel S., Cooper R. W., Moriyama E. N., Gojobori T., Hayami M. Sequence of a novel simian immunodeficiency virus from a wild-caught African mandrill. Nature. 1989 Oct 12;341(6242):539–541. doi: 10.1038/341539a0. [DOI] [PubMed] [Google Scholar]
- Voytas D. F., Ausubel F. M. A copia-like transposable element family in Arabidopsis thaliana. Nature. 1988 Nov 17;336(6196):242–244. doi: 10.1038/336242a0. [DOI] [PubMed] [Google Scholar]
- Webster T. A., Patarca R., Lathrop R. H., Smith T. F. Potential structural motifs for reverse transcriptases. Mol Biol Evol. 1989 May;6(3):317–320. doi: 10.1093/oxfordjournals.molbev.a040543. [DOI] [PubMed] [Google Scholar]
- Weiner A. M., Deininger P. L., Efstratiadis A. Nonviral retroposons: genes, pseudogenes, and transposable elements generated by the reverse flow of genetic information. Annu Rev Biochem. 1986;55:631–661. doi: 10.1146/annurev.bi.55.070186.003215. [DOI] [PubMed] [Google Scholar]
- Xiong Y., Eickbush T. H. Similarity of reverse transcriptase-like sequences of viruses, transposable elements, and mitochondrial introns. Mol Biol Evol. 1988 Nov;5(6):675–690. doi: 10.1093/oxfordjournals.molbev.a040521. [DOI] [PubMed] [Google Scholar]
- Xiong Y., Eickbush T. H. The site-specific ribosomal DNA insertion element R1Bm belongs to a class of non-long-terminal-repeat retrotransposons. Mol Cell Biol. 1988 Jan;8(1):114–123. doi: 10.1128/mcb.8.1.114. [DOI] [PMC free article] [PubMed] [Google Scholar]