Abstract
We previously cloned a DNA fragment from Saccharomyces cerevisiae that suppressed the alkylation sensitivity of Escherichia coli glycosylase deficient mutants and we showed that it apparently contained a gene for 3-methyl-adenine DNA glycosylase (MAG). Here we establish the identity of the MAG gene by sequence analysis and describe its in vivo function and expression in yeast cells. The MAG DNA glycosylase specifically protects yeast cells against the killing effects of alkylating agents. It does not protect cells against mutation; indeed, it appears to generate mutations which presumably result from those apurinic sites produced by the glycosylase that escape further repair. The MAG gene, which we mapped to chromosome V, is not allelic with any of the RAD genes and appears to be allelic to the unmapped MMS-5 gene. From its sequence the MAG glycosylase is predicted to contain 296 amino acids and have a molecular weight of 34,293 daltons. A 137 amino acid stretch of the MAG glycosylase displays 27.0% identity and 63.5% similarity with the E. coli AlkA glycosylase. Transcription of the MAG gene, like that of the E. coli alkA gene, is greatly increased when yeast cells are exposed to relatively non-toxic levels of alkylating agents.
Full text
PDF


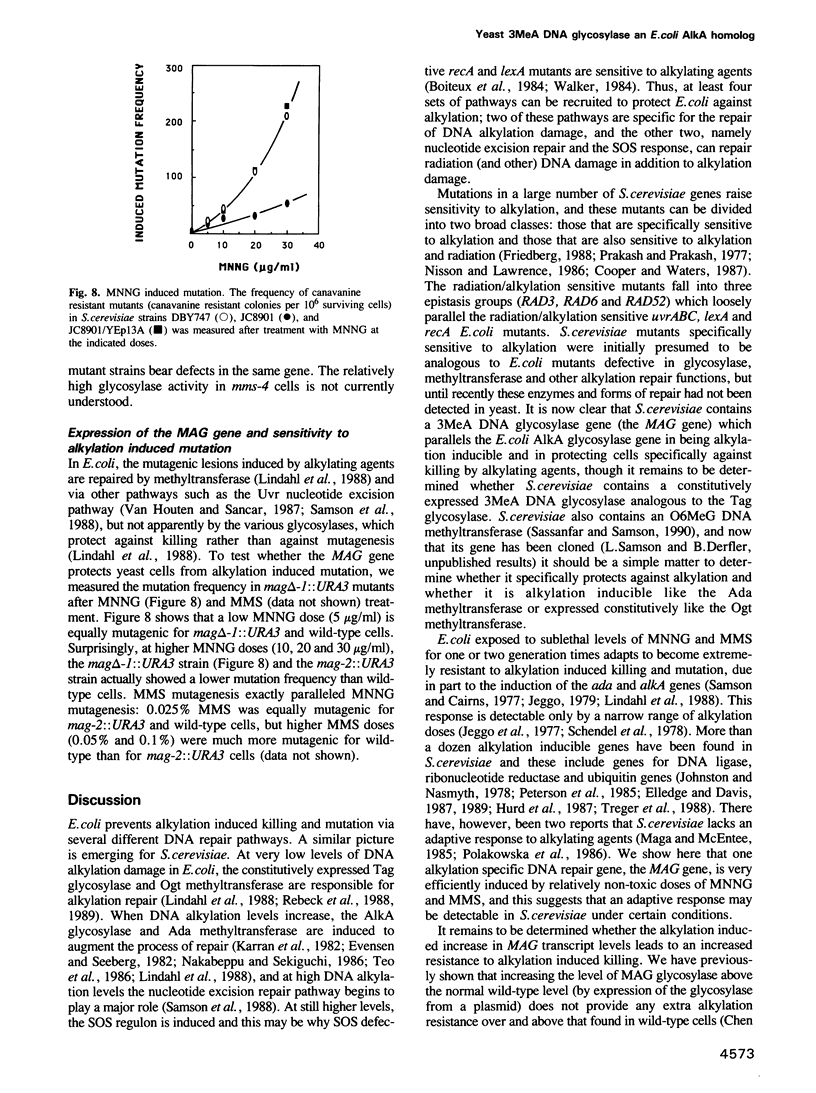


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Alani E., Cao L., Kleckner N. A method for gene disruption that allows repeated use of URA3 selection in the construction of multiply disrupted yeast strains. Genetics. 1987 Aug;116(4):541–545. doi: 10.1534/genetics.112.541.test. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Banga S. S., Boyd J. B., Valerie K., Harris P. V., Kurz E. M., de Riel J. K. denV gene of bacteriophage T4 restores DNA excision repair to mei-9 and mus201 mutants of Drosophila melanogaster. Proc Natl Acad Sci U S A. 1989 May;86(9):3227–3231. doi: 10.1073/pnas.86.9.3227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bennetzen J. L., Hall B. D. Codon selection in yeast. J Biol Chem. 1982 Mar 25;257(6):3026–3031. [PubMed] [Google Scholar]
- Bernstein H., Bernstein C. Bacteriophage T4 genetic homologies with bacteria and eucaryotes. J Bacteriol. 1989 May;171(5):2265–2270. doi: 10.1128/jb.171.5.2265-2270.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boiteux S., Huisman O., Laval J. 3-Methyladenine residues in DNA induce the SOS function sfiA in Escherichia coli. EMBO J. 1984 Nov;3(11):2569–2573. doi: 10.1002/j.1460-2075.1984.tb02175.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen J., Derfler B., Maskati A., Samson L. Cloning a eukaryotic DNA glycosylase repair gene by the suppression of a DNA repair defect in Escherichia coli. Proc Natl Acad Sci U S A. 1989 Oct;86(20):7961–7965. doi: 10.1073/pnas.86.20.7961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cigan A. M., Donahue T. F. Sequence and structural features associated with translational initiator regions in yeast--a review. Gene. 1987;59(1):1–18. doi: 10.1016/0378-1119(87)90261-7. [DOI] [PubMed] [Google Scholar]
- Clarke N. D., Kvaal M., Seeberg E. Cloning of Escherichia coli genes encoding 3-methyladenine DNA glycosylases I and II. Mol Gen Genet. 1984;197(3):368–372. doi: 10.1007/BF00329931. [DOI] [PubMed] [Google Scholar]
- Cole G. M., Schild D., Lovett S. T., Mortimer R. K. Regulation of RAD54- and RAD52-lacZ gene fusions in Saccharomyces cerevisiae in response to DNA damage. Mol Cell Biol. 1987 Mar;7(3):1078–1084. doi: 10.1128/mcb.7.3.1078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cooper A. J., Waters R. A complex pattern of sensitivity to simple monofunctional alkylating agents exists amongst the rad mutants of Saccharomyces cerevisiae. Mol Gen Genet. 1987 Aug;209(1):142–148. doi: 10.1007/BF00329849. [DOI] [PubMed] [Google Scholar]
- Doolittle R. F., Johnson M. S., Husain I., Van Houten B., Thomas D. C., Sancar A. Domainal evolution of a prokaryotic DNA repair protein and its relationship to active-transport proteins. Nature. 1986 Oct 2;323(6087):451–453. doi: 10.1038/323451a0. [DOI] [PubMed] [Google Scholar]
- Downes C. S. DNA repair. Views of unity and diversity. Nature. 1988 Mar 17;332(6161):208–209. doi: 10.1038/332208a0. [DOI] [PubMed] [Google Scholar]
- Elledge S. J., Davis R. W. DNA damage induction of ribonucleotide reductase. Mol Cell Biol. 1989 Nov;9(11):4932–4940. doi: 10.1128/mcb.9.11.4932. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elledge S. J., Davis R. W. Identification and isolation of the gene encoding the small subunit of ribonucleotide reductase from Saccharomyces cerevisiae: DNA damage-inducible gene required for mitotic viability. Mol Cell Biol. 1987 Aug;7(8):2783–2793. doi: 10.1128/mcb.7.8.2783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Evensen G., Seeberg E. Adaptation to alkylation resistance involves the induction of a DNA glycosylase. Nature. 1982 Apr 22;296(5859):773–775. doi: 10.1038/296773a0. [DOI] [PubMed] [Google Scholar]
- Fitzgerald M., Shenk T. The sequence 5'-AAUAAA-3'forms parts of the recognition site for polyadenylation of late SV40 mRNAs. Cell. 1981 Apr;24(1):251–260. doi: 10.1016/0092-8674(81)90521-3. [DOI] [PubMed] [Google Scholar]
- Friedberg E. C. Deoxyribonucleic acid repair in the yeast Saccharomyces cerevisiae. Microbiol Rev. 1988 Mar;52(1):70–102. doi: 10.1128/mr.52.1.70-102.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gallagher P. E., Brent T. P. Partial purification and characterization of 3-methyladenine-DNA glycosylase from human placenta. Biochemistry. 1982 Dec 7;21(25):6404–6409. doi: 10.1021/bi00268a013. [DOI] [PubMed] [Google Scholar]
- Guarente L. Regulatory proteins in yeast. Annu Rev Genet. 1987;21:425–452. doi: 10.1146/annurev.ge.21.120187.002233. [DOI] [PubMed] [Google Scholar]
- Henikoff S. Unidirectional digestion with exonuclease III creates targeted breakpoints for DNA sequencing. Gene. 1984 Jun;28(3):351–359. doi: 10.1016/0378-1119(84)90153-7. [DOI] [PubMed] [Google Scholar]
- Hurd H. K., Roberts C. W., Roberts J. W. Identification of the gene for the yeast ribonucleotide reductase small subunit and its inducibility by methyl methanesulfonate. Mol Cell Biol. 1987 Oct;7(10):3673–3677. doi: 10.1128/mcb.7.10.3673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jeggo P., Defais T. M., Samson L., Schendel P. An adaptive response of E. coli to low levels of alkylating agent: comparison with previously characterised DNA repair pathways. Mol Gen Genet. 1977 Nov 29;157(1):1–9. doi: 10.1007/BF00268680. [DOI] [PubMed] [Google Scholar]
- Jeggo P. Isolation and characterization of Escherichia coli K-12 mutants unable to induce the adaptive response to simple alkylating agents. J Bacteriol. 1979 Sep;139(3):783–791. doi: 10.1128/jb.139.3.783-791.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnston L. H., Nasmyth K. A. Saccharomyces cerevisiae cell cycle mutant cdc9 is defective in DNA ligase. Nature. 1978 Aug 31;274(5674):891–893. doi: 10.1038/274891a0. [DOI] [PubMed] [Google Scholar]
- Johnston L. H., White J. H., Johnson A. L., Lucchini G., Plevani P. The yeast DNA polymerase I transcript is regulated in both the mitotic cell cycle and in meiosis and is also induced after DNA damage. Nucleic Acids Res. 1987 Jul 10;15(13):5017–5030. doi: 10.1093/nar/15.13.5017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karran P., Hjelmgren T., Lindahl T. Induction of a DNA glycosylase for N-methylated purines is part of the adaptive response to alkylating agents. Nature. 1982 Apr 22;296(5859):770–773. doi: 10.1038/296770a0. [DOI] [PubMed] [Google Scholar]
- Karran P., Lindahl T., Ofsteng I., Evensen G. B., Seeberg E. Escherichia coli mutants deficient in 3-methyladenine-DNA glycosylase. J Mol Biol. 1980 Jun 15;140(1):101–127. doi: 10.1016/0022-2836(80)90358-7. [DOI] [PubMed] [Google Scholar]
- Kelley M. R., Venugopal S., Harless J., Deutsch W. A. Antibody to a human DNA repair protein allows for cloning of a Drosophila cDNA that encodes an apurinic endonuclease. Mol Cell Biol. 1989 Mar;9(3):965–973. doi: 10.1128/mcb.9.3.965. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laval F. Repair of methylated bases in mammalian cells during adaptive response to alkylating agents. Biochimie. 1985 Mar-Apr;67(3-4):361–364. doi: 10.1016/s0300-9084(85)80081-x. [DOI] [PubMed] [Google Scholar]
- Lindahl T., Sedgwick B., Sekiguchi M., Nakabeppu Y. Regulation and expression of the adaptive response to alkylating agents. Annu Rev Biochem. 1988;57:133–157. doi: 10.1146/annurev.bi.57.070188.001025. [DOI] [PubMed] [Google Scholar]
- Lipman D. J., Pearson W. R. Rapid and sensitive protein similarity searches. Science. 1985 Mar 22;227(4693):1435–1441. doi: 10.1126/science.2983426. [DOI] [PubMed] [Google Scholar]
- Loeb L. A. Apurinic sites as mutagenic intermediates. Cell. 1985 Mar;40(3):483–484. doi: 10.1016/0092-8674(85)90191-6. [DOI] [PubMed] [Google Scholar]
- Madura K., Prakash S. Nucleotide sequence, transcript mapping, and regulation of the RAD2 gene of Saccharomyces cerevisiae. J Bacteriol. 1986 Jun;166(3):914–923. doi: 10.1128/jb.166.3.914-923.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maga J. A., McEntee K. Response of S. cerevisiae to N-methyl-N'-nitro-N-nitrosoguanidine: mutagenesis, survival and DDR gene expression. Mol Gen Genet. 1985;200(2):313–321. doi: 10.1007/BF00425442. [DOI] [PubMed] [Google Scholar]
- Male R., Haukanes B. I., Helland D. E., Kleppe K. Substrate specificity of 3-methyladenine-DNA glycosylase from calf thymus. Eur J Biochem. 1987 May 15;165(1):13–19. doi: 10.1111/j.1432-1033.1987.tb11188.x. [DOI] [PubMed] [Google Scholar]
- Male R., Helland D. E., Kleppe K. Purification and characterization of 3-methyladenine-DNA glycosylase from calf thymus. J Biol Chem. 1985 Feb 10;260(3):1623–1629. [PubMed] [Google Scholar]
- McClanahan T., McEntee K. Specific transcripts are elevated in Saccharomyces cerevisiae in response to DNA damage. Mol Cell Biol. 1984 Nov;4(11):2356–2363. doi: 10.1128/mcb.4.11.2356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakabeppu Y., Kondo H., Sekiguchi M. Cloning and characterization of the alkA gene of Escherichia coli that encodes 3-methyladenine DNA glycosylase II. J Biol Chem. 1984 Nov 25;259(22):13723–13729. [PubMed] [Google Scholar]
- Nakabeppu Y., Miyata T., Kondo H., Iwanaga S., Sekiguchi M. Structure and expression of the alkA gene of Escherichia coli involved in adaptive response to alkylating agents. J Biol Chem. 1984 Nov 25;259(22):13730–13736. [PubMed] [Google Scholar]
- Nakabeppu Y., Sekiguchi M. Regulatory mechanisms for induction of synthesis of repair enzymes in response to alkylating agents: ada protein acts as a transcriptional regulator. Proc Natl Acad Sci U S A. 1986 Sep;83(17):6297–6301. doi: 10.1073/pnas.83.17.6297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakatsuru Y., Nemoto N., Nakagawa K., Masahito P., Ishikawa T. O6-methylguanine DNA methyltransferase activity in liver from various fish species. Carcinogenesis. 1987 Aug;8(8):1123–1127. doi: 10.1093/carcin/8.8.1123. [DOI] [PubMed] [Google Scholar]
- Nisson P. E., Lawrence C. W. The isolation and characterization of an alkylating-agent-sensitive yeast mutant, ngs1. Mutat Res. 1986 May;165(3):129–137. doi: 10.1016/0167-8817(86)90047-7. [DOI] [PubMed] [Google Scholar]
- Percival K. J., Klein M. B., Burgers P. M. Molecular cloning and primary structure of the uracil-DNA-glycosylase gene from Saccharomyces cerevisiae. J Biol Chem. 1989 Feb 15;264(5):2593–2598. [PubMed] [Google Scholar]
- Peterson T. A., Prakash L., Prakash S., Osley M. A., Reed S. I. Regulation of CDC9, the Saccharomyces cerevisiae gene that encodes DNA ligase. Mol Cell Biol. 1985 Jan;5(1):226–235. doi: 10.1128/mcb.5.1.226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Polakowska R., Perozzi G., Prakash L. Alkylation mutagenesis in Saccharomyces cerevisiae: lack of evidence for an adaptive response. Curr Genet. 1986;10(9):647–655. doi: 10.1007/BF00410912. [DOI] [PubMed] [Google Scholar]
- Popoff S. C., Spira A. I., Johnson A. W., Demple B. Yeast structural gene (APN1) for the major apurinic endonuclease: homology to Escherichia coli endonuclease IV. Proc Natl Acad Sci U S A. 1990 Jun;87(11):4193–4197. doi: 10.1073/pnas.87.11.4193. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Prakash L., Prakash S. Isolation and characterization of MMS-sensitive mutants of Saccharomyces cerevisiae. Genetics. 1977 May;86(1):33–55. doi: 10.1093/genetics/86.1.33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rebeck G. W., Coons S., Carroll P., Samson L. A second DNA methyltransferase repair enzyme in Escherichia coli. Proc Natl Acad Sci U S A. 1988 May;85(9):3039–3043. doi: 10.1073/pnas.85.9.3039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rebeck G. W., Smith C. M., Goad D. L., Samson L. Characterization of the major DNA repair methyltransferase activity in unadapted Escherichia coli and identification of a similar activity in Salmonella typhimurium. J Bacteriol. 1989 Sep;171(9):4563–4568. doi: 10.1128/jb.171.9.4563-4568.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robinson G. W., Nicolet C. M., Kalainov D., Friedberg E. C. A yeast excision-repair gene is inducible by DNA damaging agents. Proc Natl Acad Sci U S A. 1986 Mar;83(6):1842–1846. doi: 10.1073/pnas.83.6.1842. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ruby S. W., Szostak J. W., Murray A. W. Cloning regulated yeast genes from a pool of lacZ fusions. Methods Enzymol. 1983;101:253–269. doi: 10.1016/0076-6879(83)01019-8. [DOI] [PubMed] [Google Scholar]
- Ruby S. W., Szostak J. W. Specific Saccharomyces cerevisiae genes are expressed in response to DNA-damaging agents. Mol Cell Biol. 1985 Jan;5(1):75–84. doi: 10.1128/mcb.5.1.75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sakumi K., Nakabeppu Y., Yamamoto Y., Kawabata S., Iwanaga S., Sekiguchi M. Purification and structure of 3-methyladenine-DNA glycosylase I of Escherichia coli. J Biol Chem. 1986 Nov 25;261(33):15761–15766. [PubMed] [Google Scholar]
- Samson L., Cairns J. A new pathway for DNA repair in Escherichia coli. Nature. 1977 May 19;267(5608):281–283. doi: 10.1038/267281a0. [DOI] [PubMed] [Google Scholar]
- Samson L., Derfler B., Waldstein E. A. Suppression of human DNA alkylation-repair defects by Escherichia coli DNA-repair genes. Proc Natl Acad Sci U S A. 1986 Aug;83(15):5607–5610. doi: 10.1073/pnas.83.15.5607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Samson L., Thomale J., Rajewsky M. F. Alternative pathways for the in vivo repair of O6-alkylguanine and O4-alkylthymine in Escherichia coli: the adaptive response and nucleotide excision repair. EMBO J. 1988 Jul;7(7):2261–2267. doi: 10.1002/j.1460-2075.1988.tb03066.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sassanfar M., Samson L. Identification and preliminary characterization of an O6-methylguanine DNA repair methyltransferase in the yeast Saccharomyces cerevisiae. J Biol Chem. 1990 Jan 5;265(1):20–25. [PubMed] [Google Scholar]
- Schendel P. F., Defais M., Jeggo P., Samson L., Cairns J. Pathways of mutagenesis and repair in Escherichia coli exposed to low levels of simple alkylating agents. J Bacteriol. 1978 Aug;135(2):466–475. doi: 10.1128/jb.135.2.466-475.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shine J., Dalgarno L. The 3'-terminal sequence of Escherichia coli 16S ribosomal RNA: complementarity to nonsense triplets and ribosome binding sites. Proc Natl Acad Sci U S A. 1974 Apr;71(4):1342–1346. doi: 10.1073/pnas.71.4.1342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steinum A. L., Seeberg E. Nucleotide sequence of the tag gene from Escherichia coli. Nucleic Acids Res. 1986 May 12;14(9):3763–3772. doi: 10.1093/nar/14.9.3763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Teo I., Sedgwick B., Kilpatrick M. W., McCarthy T. V., Lindahl T. The intracellular signal for induction of resistance to alkylating agents in E. coli. Cell. 1986 Apr 25;45(2):315–324. doi: 10.1016/0092-8674(86)90396-x. [DOI] [PubMed] [Google Scholar]
- Treger J. M., Heichman K. A., McEntee K. Expression of the yeast UB14 gene increases in response to DNA-damaging agents and in meiosis. Mol Cell Biol. 1988 Mar;8(3):1132–1136. doi: 10.1128/mcb.8.3.1132. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Valerie K., de Riel J. K., Henderson E. E. Genetic complementation of UV-induced DNA repair in Chinese hamster ovary cells by the denV gene of phage T4. Proc Natl Acad Sci U S A. 1985 Nov;82(22):7656–7660. doi: 10.1073/pnas.82.22.7656. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Houten B., Sancar A. Repair of N-methyl-N'-nitro-N-nitrosoguanidine-induced DNA damage by ABC excinuclease. J Bacteriol. 1987 Feb;169(2):540–545. doi: 10.1128/jb.169.2.540-545.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walker G. C. Mutagenesis and inducible responses to deoxyribonucleic acid damage in Escherichia coli. Microbiol Rev. 1984 Mar;48(1):60–93. doi: 10.1128/mr.48.1.60-93.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wallace S. S. AP endonucleases and DNA glycosylases that recognize oxidative DNA damage. Environ Mol Mutagen. 1988;12(4):431–477. doi: 10.1002/em.2860120411. [DOI] [PubMed] [Google Scholar]
- van Duin M., de Wit J., Odijk H., Westerveld A., Yasui A., Koken M. H., Hoeijmakers J. H., Bootsma D. Molecular characterization of the human excision repair gene ERCC-1: cDNA cloning and amino acid homology with the yeast DNA repair gene RAD10. Cell. 1986 Mar 28;44(6):913–923. doi: 10.1016/0092-8674(86)90014-0. [DOI] [PubMed] [Google Scholar]
- van Duin M., van Den Tol J., Hoeijmakers J. H., Bootsma D., Rupp I. P., Reynolds P., Prakash L., Prakash S. Conserved pattern of antisense overlapping transcription in the homologous human ERCC-1 and yeast RAD10 DNA repair gene regions. Mol Cell Biol. 1989 Apr;9(4):1794–1798. doi: 10.1128/mcb.9.4.1794. [DOI] [PMC free article] [PubMed] [Google Scholar]