Abstract
Targeted protein degradation is a promising strategy for drug design and functional assessment. Several small molecule approaches have been developed that localize target proteins to ubiquitin ligases, inducing ubiquitination and subsequent degradation by the 26S proteasome. We discovered that the degradation of a target protein can also be induced by a recognition ligand linked to tert-butyl carbamate (Boc3)-protected arginine (B3A). Here we show that this process requires the proteasome, but does not involve ubiquitination of the target protein. B3A does not perturb the structure of the target protein; instead a B3A-ligand stabilizes its target protein. B3A ligands stimulate activity of purified 20S proteasome, emonstrating that the tag binds directly to the 20S proteasome. Moreover, purified 20S proteasome is sufficient to degrade target proteins in the presence of their respective B3A-linked recognition ligands. These observations suggest a simple model for B3A-mediated degradation wherein the B3A tag localizes target proteins directly to the 20S proteasome. Thus B3A ligands are the first example of a ubiquitin-free strategy for targeted protein degradation.
Graphical abstract

Targeted protein degradation is an emerging strategy in drug design 1, 2. Several drugs, most notably fulvestrant and other selective estrogen receptor downregulators (SERDs), induce degradation of their target protein. SERDs change the conformation of the estrogen receptor, exposing a hydrophobic surface that triggers ubiquitination and subsequent degradation 3. Other examples of this phenomenon include a CaCCinh-A01 calcium-activated chloride channel inhibitor 4, the ErbB2/HER2 ligand CL-1033 5, the androgen receptor antagonist bicalutamide 6 and RNA polymerase inhibitor BMH-21 7. All of these compounds were discovered serendipitously.
Several laboratories have recently developed targeted degradation methods by linking a recognition ligand for a target protein to a ligand for an E3 ubiquitin ligase, thus inducing ubiquitination and degradation of the target 8–11. Crews and colleagues have also devised a hydrophobic tagging strategy by attaching a recognition ligand to adamantyl and other hydrophobic moieties (e.g., HyT13; Figure 1; 12–17). Adamantyl-tagged recognition ligands destabilize their target proteins and recruit Hsp70, inducing ubiquitination and eventual degradation via the proteasome 12, 14. Adamantyl-tagged recognition ligands can also induce the unfolded protein response characterized by phosphorylation of eIF2 and increased expression of XBP1s 18.
Figure 1. Mechanisms of small molecule targeted protein degradation.
A) Several methods, e.g. PROTACS and SNIPERS, localize the target protein to an E3 ubiquitin ligase via a chimeric ligand, causing ubiquitination and subsequent degradation through the 26S proteasome 1. B) Mechanism of hydrophobic tag induced degradation, e.g., HaloTag/HyT 14, 15. Ligand binds to Halo protein. Halo protein unfolds, causing ubiquitination and degradation via the 26S proteasome. C) Possible mechanisms of Boc3Arg mediated degradation. D) Compounds used in this work.
We serendipitously discovered that tert-butyl carbamate (Boc3)-protected arginine (hereafter B3A) acts as a small molecule chemical degron to elicit targeted protein degradation 19. We attached B3A to ethacrynic acid (EA-B3A), a covalent inhibitor of glutathione transferases (GST), and EA-B3A induced the degradation of GST-α and GST-α-EGFP fusion proteins as well as endogenous GST-π in cells and lysates. Similarly, trimethoprim-linked B3A (TMP-B3A) induced the degradation of its target, E. coli DHFR (eDHFR), as well as eDHFR-EGFP fusion proteins, demonstrating that degradation does not require covalent attachment of the tag to the target protein. Importantly, B3A ligands decreased the half-lives of their respective target proteins in the presence of the translation inhibitor cycloheximide, confirming that the B3A tag induced degradation.
At first glance, B3A appears to be another example of hydrophobic tagging described by the Crews laboratory. However, B3A does not appear to induce ubiquitination of the target protein 19, nor does it induce phosphorylation of eIF2 20, indicating that the mechanism of B3A induced degradation is distinct from that of adamantly-tagged recognition ligands. Here we show that B3A ligands activate the 20S proteasome, and the 20S proteasome degrades target proteins in a B3A-dependent manner. Thus we propose that B3A binding localizes the target protein to the 20S proteasome, resulting in degradation.
RESULTS AND DISCUSSION
B3A does not induce ubiquitination
Previously reported methods to induce targeted protein degradation rely on ubiquitination of the target protein 1, 8, 10, 11. However, we failed to observe the high molecular weight bands indicative of ubiquitin conjugation when B3A-induced degradation was blocked by the addition of proteasome inhibitors 19. We performed several additional experiments to more rigorously probe for ubiquitination of the target protein.
HA-ubiquitin (HA-Ub) pulldown experiments are perhaps the most sensitive method to detect ubiquitination 21. Therefore we probed the ubiquitination of endogenous GST-π in HEK293T cells transiently expressing HA-Ub. These cells were treated with either ethacrynic acid (EA) or EA-B3A in the presence of the proteasome inhibitor bortezomib, and lysates were prepared in the presence of the pan-deubiquitinating enzyme inhibitor PR-619. No high molecular weight GST-π was observed, indicating that EA-B3A did not induce ubiquitination (Figure 2a). Similar results were observed in Cos1 cells (Supplementary Figure S1). These experiments strongly suggest that EA-B3A does not induce ubiquitination of GST-π.
Figure 2. EA-B3A does not increase the ubiquitination of target proteins.
A) Endogenous GST-π. HEK293T cells expressing HA-Ub were treated with bortezomib (1 μM) followed by EA or EA-B3A (40 μM) for 2 h. Lysates (5 mg/ml) were prepared in the presence of the pan-DUB inhibitor PR-619 and ubiquitinated proteins were isolated with anti-HA beads. Protein was analyzed by western blot using anti-GST-π; anti-HA (HA-ubiquitin) and anti-tubulin. B) Cos1 cells were transfected with DHFR-HA-GFP-FLAG and treated with either TMP (80 μM) or TMP-B3A (80 μM) for 3 h in the presence of bortezomib (1 μM). Lysates were prepared in 0.3% SDS containing buffer supplemented with irreversible DUB inhibitor WP1130 and then immunoprecipitated with anti-FLAG. Protein analyzed by western blot using anti-HA (DHFR-HA-GFP-FLAG) and anti-K48-linked ubiquitin. C) PYR-41 does not block the B3A-induced degradation of endogenous GST-π. HEK293T cells were preincubated with cycloheximide (CHX) (50 μg/ml) for 30 min, then treated with EA-B3A (40 μM), PYR-41 (40 μM), DMSO (vehicle only), or EA-B3A (40 μM) with PYR-41 (40 μM) and MG132 (10 μM) for 3 h and blotted for K48 ubiquitination. Same condition without MG132 was used when blotting GST-π. D) Plots show mean ± s.d. for experiments as in panel D, (N=3). E) b-AP15 does not block B3A-induced degradation of eDHFR. A Cos1 cell lysate (1 mg/ml) was preincubated with b-AP15 (20 μM) for 20 min before addition of eDHFR (800nM) and TMP-B3A (1.6 μM) were added to the assay.
We also investigated the ubiquitination of an HA and FLAG-tagged fusion protein eDHFR-HA-GFP-FLAG. Cos1 cells transiently expressing eDHFR-HA-GFP-FLAG were treated with bortezomib followed by TMP or TMP-B3A. Cells were lysed in the presence of 0.3% SDS and the irreversible deubiquitinating enzyme (DUB) inhibitor WP1130 22. eDHFR-HA-GFP-FLAG was immunoprecipitated with anti-FLAG antibody and analyzed (Figure 2b). No high molecular weight bands were observed in an anti-HA blot in either the TMP or TMP-B3A treated samples. Furthermore, less K48-linked ubiquitin was immunoprecipitated in the TMP-B3A-treated cells than in TMP treated cells. These data provide further evidence that ubiquitination is not involved in B3A-mediated protein degradation.
B3A-induced degradation does not require UAE1
Ubiquitin-activating enzyme 1 (UAE1) is required for the majority of cellular ubiquitination 23. PYR-41 is a cell permeable small molecule inhibitor of UAE1 24. A significant decrease in K48-linked high molecular weight ubiquitin conjugates was observed when HEK293T cells were treated with the proteasome inhibitor MG132 followed by PYR-41 (Supplementary Figure S2a), although no effect was observed in the absence of MG132. These observations are consistent with reports from other laboratories 25. PYR-41 also inhibits DUBs, which stabilizes ubiquitination and can obscure the effect of UAE1 inhibition, explaining these relatively modest effects 26. Having confirmed that PYR-41 inhibited ubiquitination in HEK293T cells, we then tested the effect of PYR-41 on B3A-induced degradation. Cells were pre-treated with cycloheximide to inhibit protein biosynthesis, then treated with both EA-B3A and PYR-41. No effect on the degradation of GST-π was observed (Figure 2c, d), further suggesting that B3A-induced degradation does not involve ubiquitination.
B3A-induced degradation does not involve the 26S
B3A-induced degradation is blocked by proteasome inhibitors such as bortezomib and MG132 19. However, these inhibitors do not distinguish the core 20S proteasome from the 26S proteasome that is responsible for the degradation of ubiquitinated proteins. The 26S proteasome contains the 19S regulatory complex, which includes the DUBs USP14 and UCH37. Inhibition of these DUBs by the small molecule b-AP15 leads to inhibition of 26S proteasome 27. Therefore we investigated the degradation of eDHFR in Cos1 cell lysates treated with b-AP15 to determine if B3A-induced degradation required the 26S proteasome. Inhibition of UCH37 was confirmed by activity profiling (Supplementary Figure S2b). TMP-B3A induced the degradation of eDHFR similarly in both untreated and b-AP15 treated lysates (Figure 2e). No accumulation of polyubiquitinated eDHFR was observed. These observations strongly suggest that B3A-mediated eDHFR degradation does not involve the ubiquitin-26S proteasome pathway.
B3A does not destabilize the target protein
Hydrophobic tagging is proposed to induce protein degradation by destabilizing the target protein 14. Therefore we investigated whether the B3A tag destabilized the eDHFR. The structure and folding of this small monomeric protein has been extensively characterized by other laboratories 28. The melting temperature for eDHFR was 52.4 ± 0.4 °C as measured by circular dichroism spectroscopy, in good agreement with previous reports (literature Tm = 49–51 °C; 29, 30). The presence of TMP increased the melting point of eDHFR by 14 degrees (Tm = 66.7 ± 1.8 °C; Supplementary Figure S3). Such stabilization is commonly observed when ligands bind to proteins. A similar increase in the value of Tm was observed in the presence of TMP-B3A (Tm = 64.6 ± 1.0 °C). Thus TMP-B3A does not destabilize its target protein.
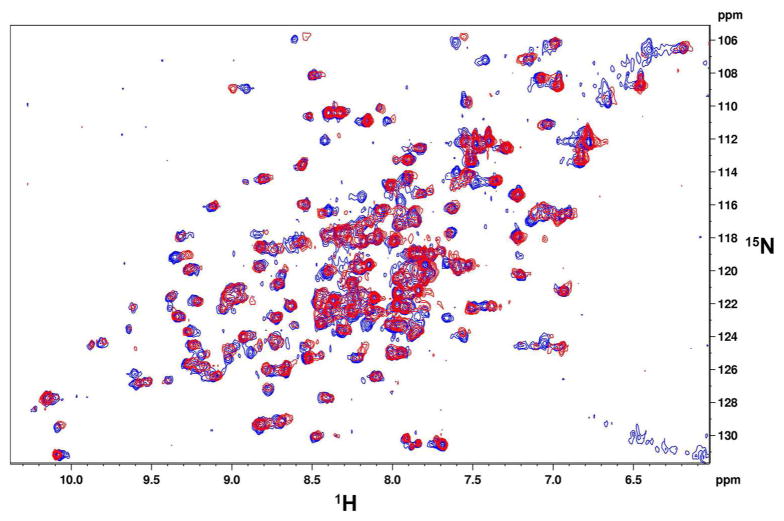
It is possible that the B3A tag might disrupt the structure of eDHFR, perhaps by embedding into the protein. Such a process would likely require a significant rearrangement of the protein structure. Therefore we analyzed the structure of the eDHFR•TMP-B3A complex using NMR spectroscopy. The 1H-15N HSQC NMR spectra of the eDHFR•TMP and eDHFR•TMP-B3A complexes are almost identical, showing that the B3A tag does not embed into the target protein (Figure 3).
Figure 3. TMP-B3A does not destabilize eDHFR.
2D 1H-15N HSQC spectra of eDHFR•TMP (blue) and eDHFR•TMP-B3A (red). NMR samples contained 0.1 mM DHFR and 0.14 mM of ligands in 25 mM Phosphate (pH 7.6), 100 mM KCl, 1 mM DTT and 10% D2O.
SAR of B3A-induced degradation
A series of EA-B3A derivatives were synthesized to explore the structure activity relationship (SAR) of B3A induced protein degradation (Figure 1d). As reported previously, degradation of GST-α1 modified with EA-B3A was observed in HeLa cell lysates within 2 h (Figure 4a–c). No degradation was observed when GST-α1 was modified with EA. Shorter and rigid linkers are expected to limit the interactions between B3A and the target protein, but replacement of the six carbon diamine linker with cyclohexane-1,4-diamine (EA-CY-B3A, 2) or ethylenediamine (EA-C2-B3A, 3) had no effect on degradation (Figure 4a). In contrast, no degradation was observed when B3A was replaced with di-butyl carbamate-protected ornithine (B2O, 4) or butyl carbamate-protected-protected glycine (B1G, 5) (Figure 4b). This observation suggests that all three boc groups are required for B3A-induced degradation. Lastly, NH2-B3A, which consists of B3A and the 6 carbon diamine linker but no recognition ligand, inhibits the EA-B3A induced degradation of GST-α1 in lysates (Figure 4c). Collectively these observations further indicate that the B3A tag interacts with another protein factor distinct from the target protein.
Figure 4. SAR of B3A-mediated degradation.

a) GST-α1 was modified with EA derivatives and added to a HeLa cell lysate. b) SAR of EA-B1G, EA-B2O and EA-B3A. c) NH2-B3A inhibits degradation. All plots show mean ± s.e.m.
B3A ligands activate the 20S proteasome
Although proteasome inhibitors block B3A-induced degradation, the above experiments suggest that the 26S proteasome is not involved. Therefore B3A-induced degradation may instead utilize the 20S proteasome. Hydrophobic peptides are known to activate the 20S proteasome 31, 32, so we hypothesized that B3A might similarly activate the 20S proteasome, thereby docking the target protein to the 20S proteasome. Therefore we investigated the effect of various B3A-containing ligands on the activities of the 20S proteasome (Figure 5). All five B3A ligands activated the trypsin-like subunits by 2–16-fold (Figure 5a and S4a). Activation of the trypsin-like subunits was also observed with EA-B1G and EA-B2O. No activation of the chymotrypsin-like activity was observed under these conditions. The activity of the 20S proteasome varies with different monovalent cations 32, so we also tried different assay buffers, and a modest stimulation (1.4–1.7-fold) of the chymotrypsin-like activity was observed with three B3A ligands, NH2-B3A, Ac-B3A and TMP-B3A when the reaction was performed in sodium chloride (Figure 5b and Supplementary S4b). EA-B3A and Cbz-B3A failed to activate the chymotrypsin-like activity and EA-B1G and EA-B2O inhibited. No activation of the postglutamyl activity was observed (Supplementary Figure S4c). In contrast to the 20S proteasome, none of the activities of the 26S proteasome were increased by the presence of NH2-B3A (Supplementary Figure S4d). In addition, the adamantyl ligand Ad-NH2 failed to stimulate the 20S proteasome (Supplementary Figure S5a, b), which further demonstrates that there are significant mechanistic differences between the adamantyl system and B3A.
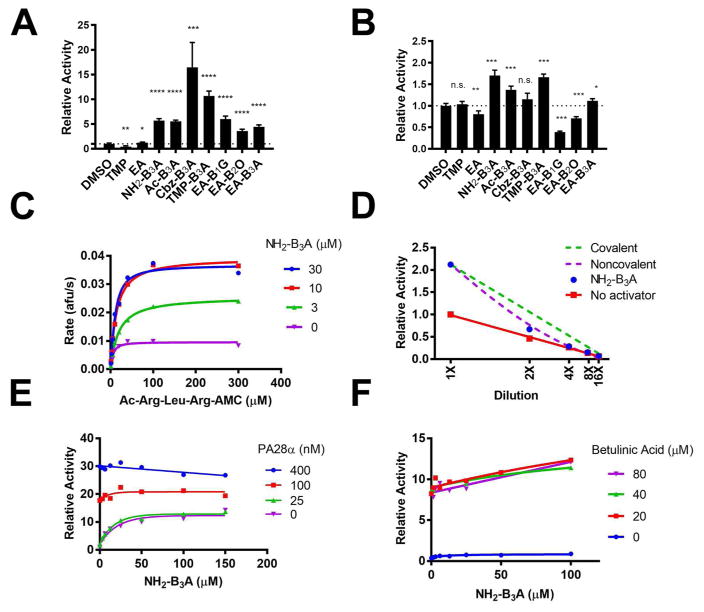
Figure 5. Effect of B3A containing ligands on proteasome activity.
a) Effect of B3A containing ligands on 20S proteasome-catalyzed hydrolysis of Ac-Arg-Leu-Arg-AMC (100 μM). b) Effects of B3A containing ligands on 20S proteasome-catalyzed hydrolysis of Suc-Leu-Leu-Val-Tyr-AMC (100 μM). c) Effect of NH2-B3A on the activity of the 20S proteasome on the trypsin-like active sites. d) B3A activates the 20S proteasome via a noncovalent interaction. 20S proteasome (0.7 nM) was first incubated with Suc-Leu-Leu-Val-Tyr-AMC (25 μM) and NH2-B3A (25 μM) for 45 min to reach an activated steady state. The proteasome solution was diluted and activity was measured. The nonlinear behavior shows that activation is not stable to dilution, as expected if NH2-B3A dissociates from the 20S proteasome. Simulations of a covalent interaction and a hypothetical noncovalent interacton (Kact = 30 μM in the simulation) are also shown. e) NH2-B3A has no effect on PA28α activated 20S proteasome (1 nM), indicating it binds at a different site. Activity was profiled with Ac-Arg-Leu-Arg-AMC (25 μM) at 37 °C. (n.s.: p > 0.05, *: p < 0.05, **: p < 0.01, ***: p < 0.001, ****: p < 0.0001). f) NH2-B3A binds to a different site than betulinic acid.
We characterized the activation of the 20S proteasome by NH2-B3A in more detail (Figure 5d and Supplementary Table S1). The values of kcat/Km increased for both the trypsin-like and chymotrypsin-like activities, albeit with different underlying causes. For the trypsin-like activity, the values of kcat increased with increasing B3A ligand while Km was unchanged. In contrast, the value of Km decreased with increasing B3A ligand while the value of kcat/Km was unchanged for the chymotrypsin-like proteasome activity. For the postglutamyl activity, the value of kcat/Km decreased with increasing NH2-B3A due to an increase in the values of Km. These observations indicate that B3A ligands allosterically modulate all of the proteasome active sites.
B3A interacts noncovalently with 20S
We performed a dilution experiment to determine if proteasome activation involved covalent or noncovalent interactions with B3A. Proteasome was activated by incubation with substrate and NH2-B3A, then diluted into substrate (Figure 5e). The nonlinear behavior shows that activation is not stable to dilution, as expected if B3A dissociates from the 20S proteasome.
B3A has no effect on other 20S activators
The 20S proteasome is also regulated by PA28α, which accelerates peptide degradation in an ATP-independent manner 33, 34. We measured the effects of NH2-B3A on PA28α stimulated 20S proteasome (Figure 5e). PA28α alone stimulated the trypsin-like activity of 20S proteasome by 10 – 15 fold at a concentration of 400 nM. NH2-B3A does not affect the stimulatory effect of PA28α nor further activate the 20S proteasome. This observation suggests that these two activators likely bind to different sites.
Betulinic acid is a well recognized small molecule activator of the 20S proteasome 35. Consistent with previous reports, betulinic acid stimulated the trypsin-like activities of the 20S proteasome by 8-fold (Figure 5f). Addition NH2-B3A to saturating concentrations of betulinic acid elicited further stimulation of the proteasome, indicating that these molecules bind to different sites on the proteasome.
20S is sufficient for B3A-induced degradation
We characterized the degradation of eDHFR and GST-α1 to determine if the 20S proteasome was sufficient to degrade proteins in a B3A-dependent manner. eDHFR was a poor substrate of the 20S proteasome (Figure 6a). TMP protected eDHFR from degradation as expected given that TMP stabilizes eDHFR. Equimolar NH2-B3A did not significantly increase degradation of eDHFR, also as expected since this concentration of B3A ligand is not sufficient to activate the 20S proteasome. In contrast, degradation increased in the presence of equimolar TMP-B3A. No degradation was observed with a mixture of equimolar NH2-B3A and TMP, confirming that B3A must be directly linked to the recognition ligand. Importantly, excess NH2-B3A inhibited TMP-B3A-induced degradation. These observations are consistent with a model that the B3A ligands bind to a site on the proteasome, localizing the target protein for degradation.
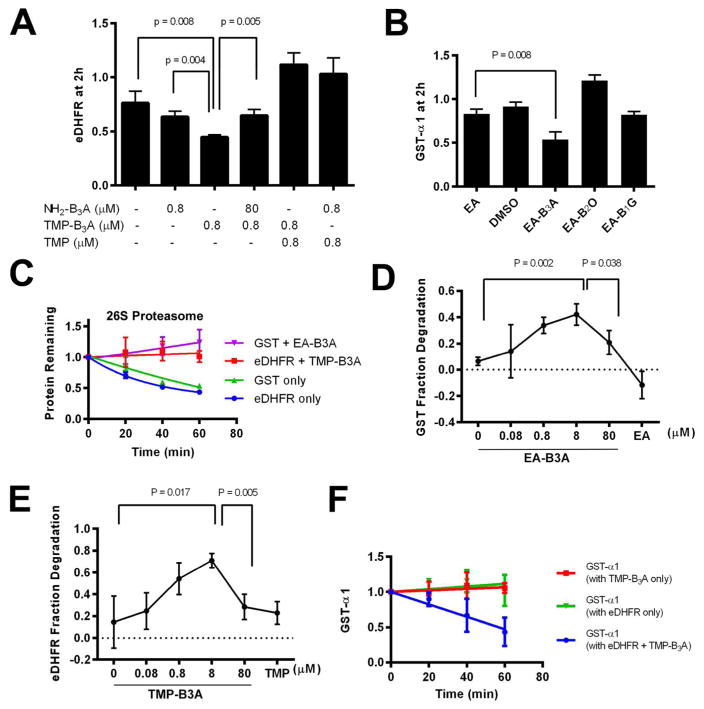
Figure 6. The 20S proteasome is sufficient to induce B3A-dependent degradation.
a) TMP-B3A induces degradation of eDHFR by the 20S proteasome. eDHFR (800 nM) was incubated with purified 20S proteasome (5 nM) at 37 °C for 2 h (N=3). b) EA-B3A induces degradation of GST-α1 by the 20S proteasome. GST-α1 (800 nM) was incubated with purified 20S proteasome (5 nM) at 37 °C for 2 h (N=3). c) Degradation of eDHFR and GST by 26S proteasome in vitro. eDHFR (1.6 μM) or GST-α1 (1.6 μM) in the absence and presence of their respective recognition ligands were incubated with 26S proteasome (1 nM) at 37 °C. d) eDHFR (800 nM) was incubated with TMP-B3A and 20S proteasome (5 nM) at 37 °C for 2 h. e) GST-α1 (800 nM) was incubated with EA-B3A and 20S proteasome (5 nM) at 37 °C for 1 h. f) TMP-B3A induces the degradation of both GST-α1 and eDHFR. A mixture of eDHFR and GST-α1 (both 160 nM) were incubated with 20S proteasome (5 nM) in the absence and presence of TMP-B3A (1.6 μM).
In principle, degradation should release the B3A ligand, allowing it to bind and degrade another target protein. However, substoichiometric TMP-B3A did not accelerate the degradation of eDHFR, indicating that the B3A ligand does not act catalytically under these conditions.
Likewise, little degradation of GST-α1 was observed in the absence of B3A ligands or in the presence of EA (Figure 6b). EA-B3A induced the degradation of GST-α1, but no degradation was observed with EA-B2O or EA-B1G. Thus the purified 20S proteasome displays the same SAR as GST-α1 degradation in lysates. These observations strongly suggest that the 20S proteasome is responsible for B3A-mediated degradation in cells.
In contrast to the above observations, TMP-B3A and EA-B3A protected their respective target proteins from degradation by the 26S proteasome, further indicating that the 20S proteasome is responsible for B3A-mediated degradation (Figure 6c).
Collectively these observations suggest degradation is mediated by a ternary complex between target protein, B3A-linked recognition ligand and 20S proteasome. This model predicts that excess recognition ligand will inhibit degradation by blocking access to the proteasome. A bell-shaped dose dependence is indeed observed for both the TMP-B3A mediated degradation of eDHFR and the EA-B3A mediate degradation of GST-α1 (Figure 6d, e). These observations demonstrate that localization of the target protein to the 20S proteasome, rather than the stimulation of proteasome activity, is the critical feature of B3A-induced degradation.
Selectivity of B3A-induced degradation
We monitored the reaction of mixtures of eDHFR and GST-α1 with the 20S proteasome and their respective recognition ligands in order to test the selectivity of degradation. The addition of EA-B3A caused the selective degradation of its target GST-α1 as expected (Supplementary Figure S6). However, the addition of TMP-B3A caused the degradation of both its target eDHFR and the non-target protein GST-α1 (Figure 6f). No GST-α1 degradation was observed in the absence of eDHFR, indicating that degradation did not result from a promiscuous interaction between TMP-B3A and GST-α1. This observation indicates that the 20S proteasome complex with eDHFR•TMP-B3A can promiscuously degrade other proteins, which suggests that off-target protein degradation may be an issue for some B3A ligands/target protein pairs.
B3A ligands activate the proteasome in living cells
We used the luminogenic proteasome substrate Z-Leu-Arg-Arg-aminoluciferin to assay the ability of B3A ligands to activate the proteasome in living cells 36. A strong luminescent signal was observed when HELA cells were treated with Z-Leu-Arg-Arg-aminoluciferin. This signal was reduced by 40% when cells were treated with the proteasome inhibitor bortezomib, indicating that at least 40% of the activity can be attributed to the proteasome. Addition of Cbz-B3A and TMP-B3A increased luminescence in the absence of bortezomib, but had no effect on luminescence in the presence of bortezomib (Figure 7). These observations indicate that both Cbz-B3A and TMP-B3A activate the proteasome in cells. NH2-B3A did not activate the proteasome in cells, possibly because it was not permeable.
Figure 7. B3A ligands activate the 20S proteasome in cells.
a) HELA cells were treated with the luminogenic proteasome substrate Z-Leu-Arg-Arg-aminoluciferin and the indicated activators (80 μM) in the absence (solid bars) and presence of bortezomib (striped bars, 20 μM). b) Effect of B3A ligands on stability of G76V-Ub-GFP. Cos1 cells expressing G76V-Ub-GFP were treated with 100 μM B3A ligands for 8 h. GFP was quantified by FACS analysis (N=4).
The above assay does not distinguish between the 20S and 26S proteasomes. Therefore we performed the UbG76V-GFP assay to monitor 26S proteasome activity 37. The G76V mutation prevents the cleavage of ubiquitin from GFP, leading to its degradation by the 26S proteasome. Inhibition of the proteasome by bortezomib treatment causes an increase of UbG76V-GFP as expected 38, 39. An approximately 15% decrease in UbG76V-GFP was observed when cells were treated with Cbz-B3A, suggesting that B3A ligands do not activate the 26S proteasome in cells.
B3A ligands localize their targets to 20S
The experiments described above indicate that, unlike other methods, B3A does not induce ubiquitination of the target protein. Instead, our experiments suggest that B3A localizes the target protein to the 20S proteasome (Figure 8). B3A activates purified 20S proteasome, demonstrating that it binds to the 20S proteasome. We have also found that proteasome subunits α7 and β7 were strongly enriched in the pool of proteins bound to B3A-linked resin (Supplementary Table S2) 20, further substantiating an interaction between B3A and the proteasome. While it is possible that other protein factors are involved in B3A-mediated degradation within a cell, purified 20S proteasome is sufficient for the B3A-mediated degradation of a target protein in vitro. Thus B3A ligands provide proof of concept for a new strategy of targeted protein degradation. Intriguingly, this strategy could be applied to other AAA+ proteases such as ClpXP or FtsH 41. These proteases are present in virtually every organism, and so this strategy has the potential to extend beyond the reach of ubiquitination targeting methods.
Figure 8.
Model for B3A-mediated protein degradation.
It’s noteworthy that TMP-B3A induces the degradation of eDHFR even as it stabilizes the protein fold. The 20S proteasome degrades intrinsically unfolded proteins, and can latch onto unfolded segments to degrade folded proteins 42, 43. Both eDHFR and GST-α contain flexible segments that could be vulnerable. We propose that the B3A ligand localizes the target protein to 20S proteasome, greatly increasing the probability that a flexible segment enters the barrel. This model predicts that the efficiency of degradation will depend on the interaction between the recognition ligand and target and the lifetime of the ternary complex. Optimization of these parameters should improve the efficiency of degradation. For example, the efficiency of eDHFR degradation might improve with a lower affinity recognition ligand that does not stabilize the folded structure to the extent observed with TMP.
Potential of B3A-induced degradation in drug discovery
It’s important to recognize that functional assessment and drug discovery have fundamentally different objectives and requirements. Ideally, functional assessment requires rapid and surgical removal of the target protein so that the resulting phenotypes can be confidently assigned. The methods that rely on E3 ubiquitin ligases appear to satisfy both of these criteria 10. Hydrophobic tagging with adamantyl ligands also induces rapid degradation, but can cause a transient unfolded protein response that may complicate the functional assessment of target proteins 18. B3A does not induce the unfolded protein response 20, but can activate the 20S proteasome and induce the degradation of non-target proteins. In addition, some B3A-ligands inhibit protein translation 20. Both phenomena depend on the specific B3A-ligand and target protein, which will complicate the use of B3A induced degradation in functional assignment.
In contrast, the goal of a drug is to elicit a clinically beneficial response. The great advantage of targeted protein degradation for drug discovery is the potential to uncouple pharmacokinetics from pharmacodynamics. The effects of a conventional inhibitor usually depend on its concentration in plasma, and therefore wane as the drug is excreted or metabolized 44. In contrast, the effect of targeted degradation will endure until the cell resynthesizes the target protein, which may be long after the drug has cleared the patient. Selectivity is not as important in this context. In fact many drugs are successful precisely because they interact with many targets 45. In the case of B3A ligands, the ability to inhibit translation may further enhance the effects of target degradation.
To our knowledge, B3A ligands are the first small molecules to induce protein degradation by localization to the proteasome. Though the importance of the 26S proteasome in cellular protein degradation is widely recognized, the substantial contribution of the 20S proteasome is often overlooked 46. The 20S proteasome is typically 2–3-fold more abundant than the 26S proteasome 47–50. The 20S proteasome degrades misfolded and oxidatively damaged proteins, as well as proteins with large regions of disorder in an ubiquitin-independent and ATP-independent manner 51. Localization of a target protein to the 26S proteasome has long been known to be sufficient for degradation 52, but the potential of the 20S proteasome for targeted protein degradation has been largely overlooked. Intriguingly, B3A binding appears to be independent of other activators such as PA28α and betulinic acid, suggesting that the 20S proteasome may harbor several potential binding sites that can be exploited with localization ligands. Thus proteasome docking is a promising new strategy for targeted protein
METHODS
Detailed Methods are available in the Supplementary Material.
Proteasome catalytic activity assays
20S Proteasome purified from human erythrocytes (catalog #E-360), 26S proteasome (catalog #E-365) and PA28α (catalog #E-381) was purified from transfected HEK cells were purchased from BostonBiochem. For peptide AMC substrates using a BioTek plate reader. For protein degradation assays, the target protein (eDHFR or GST) was preincubated with its respective inhibitor (TMP-B3A or EA-B3A) then mixed with proteasome. Aliquots were quenched with 3.5 μL of 5X SDS loading buffer at the desired time intervals and analyzed by SDS-PAGE and immunoblotting. Protein was visualized with the appropriate primary and secondary antibodies using ECL Plus (GE, Bucks, UK) after transfer. Densitometry was carried out using software IMAGE-J V 1.44p.
DUB activity profiling
Cos1 cell pellets were lysed on ice by vortexing with glass beads and crude lysate was centrifuged for 5 min. The lysate (1.5 mg/ml) was incubated with b-AP15 (or 1% DMSO control) for 20 min, then HA-ubiquitin-VME (250 nM) was added and incubated for 25 min. The reaction mixture (20 μl) was removed and quenched in 2X (final concentration) reducing SDS-PAGE loading buffer.
In cell proteasome activity
Proteasome activity in cells was measured using Proteasome-Glo™ Cell-Based Assay (Promega)36 using Z-leucine-arginine-arginine-aminoluciferin. For the ubiquitinG76V-GFP assay, Cos1 cells expressing ubiquitinG76V-GFP were treated with compound for 8 h. Cells were harvested and analyzed on a Beckman FACS-Calibur. All data were analyzed using FlowJo V10, from TreeStar (Ashland, OR).
Supplementary Material
Acknowledgments
We thank A. Gershenson (University of Massachusetts Amherst) for assistance with the protein stability experiments.
FUNDING SOURCES
This work was funded by NIH grants 5 RO1 GM100921 (LH) and 5 T32 GM007122 (RTC). MJCL was supported by a Howard Hughes Medical Institute International Student Research Fellowship.
References
- 1.Toure M, Crews CM. Small-Molecule PROTACS: New Approaches to Protein Degradation. Angew Chem Int Ed Engl. 2016;55:1966–1973. doi: 10.1002/anie.201507978. [DOI] [PubMed] [Google Scholar]
- 2.Buckley DL, Crews CM. Small-molecule control of intracellular protein levels through modulation of the ubiquitin proteasome system. Angew Chem Int Ed Engl. 2014;53:2312–2330. doi: 10.1002/anie.201307761. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Wittmann BM, Sherk A, McDonnell DP. Definition of functionally important mechanistic differences among selective estrogen receptor down-regulators. Cancer Res. 2007;67:9549–9560. doi: 10.1158/0008-5472.CAN-07-1590. [DOI] [PubMed] [Google Scholar]
- 4.Bill A, Hall ML, Borawski J, Hodgson C, Jenkins J, Piechon P, Popa O, Rothwell C, Tranter P, Tria S, Wagner T, Whitehead L, Gaither LA. Small molecule-facilitated degradation of ANO1 protein: a new targeting approach for anticancer therapeutics. J Biol Chem. 2014;289:11029–11041. doi: 10.1074/jbc.M114.549188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Citri A, Alroy I, Lavi S, Rubin C, Xu W, Grammatikakis N, Patterson C, Neckers L, Fry DW, Yarden Y. Drug-induced ubiquitylation and degradation of ErbB receptor tyrosine kinases: implications for cancer therapy. EMBO J. 2002;21:2407–2417. doi: 10.1093/emboj/21.10.2407. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Waller AS, Sharrard RM, Berthon P, Maitland NJ. Androgen receptor localisation and turnover in human prostate epithelium treated with the antiandrogen, casodex. J Mol Endocrinol. 2000;24:339–351. doi: 10.1677/jme.0.0240339. [DOI] [PubMed] [Google Scholar]
- 7.Peltonen K, Colis L, Liu H, Trivedi R, Moubarek MS, Moore HM, Bai B, Rudek MA, Bieberich CJ, Laiho M. A targeting modality for destruction of RNA polymerase I that possesses anticancer activity. Cancer cell. 2014;25:77–90. doi: 10.1016/j.ccr.2013.12.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Itoh Y, Ishikawa M, Naito M, Hashimoto Y. Protein knockdown using methyl bestatin-ligand hybrid molecules: design and synthesis of inducers of ubiquitination-mediated degradation of cellular retinoic acid-binding proteins. J Am Chem Soc. 2010;132:5820–5826. doi: 10.1021/ja100691p. [DOI] [PubMed] [Google Scholar]
- 9.Lu J, Qian Y, Altieri M, Dong H, Wang J, Raina K, Hines J, Winkler JD, Crew AP, Coleman K, Crews CM. Hijacking the E3 Ubiquitin Ligase Cereblon to Efficiently Target BRD4. Chem Biol. 2015;22:755–763. doi: 10.1016/j.chembiol.2015.05.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Winter GE, Buckley DL, Paulk J, Roberts JM, Souza A, Dhe-Paganon S, Bradner JE. DRUG DEVELOPMENT. Phthalimide conjugation as a strategy for in vivo target protein degradation. Science. 2015;348:1376–1381. doi: 10.1126/science.aab1433. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Zengerle M, Chan KH, Ciulli A. Selective Small Molecule Induced Degradation of the BET Bromodomain Protein BRD4. ACS Chem Biol. 2015;10:1770–1777. doi: 10.1021/acschembio.5b00216. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Neklesa TK, Noblin DJ, Kuzin A, Lew S, Seetharaman J, Acton TB, Kornhaber G, Xiao R, Montelione GT, Tong L, Crews CM. A bidirectional system for the dynamic small molecule control of intracellular fusion proteins. ACS Chem Biol. 2013;8:2293–2300. doi: 10.1021/cb400569k. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Neklesa TK, Crews CM. Chemical biology: Greasy tags for protein removal. Nature. 2012;487:308–309. doi: 10.1038/487308a. [DOI] [PubMed] [Google Scholar]
- 14.Tae HS, Sundberg TB, Neklesa TK, Noblin DJ, Gustafson JL, Roth AG, Raina K, Crews CM. Identification of hydrophobic tags for the degradation of stabilized proteins. Chembiochem. 2012;13:538–541. doi: 10.1002/cbic.201100793. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Neklesa TK, Tae HS, Schneekloth AR, Stulberg MJ, Corson TW, Sundberg TB, Raina K, Holley SA, Crews CM. Small-molecule hydrophobic tagging-induced degradation of HaloTag fusion proteins. Nat Chem Biol. 2011;7:538–543. doi: 10.1038/nchembio.597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Xie T, Lim SM, Westover KD, Dodge ME, Ercan D, Ficarro SB, Udayakumar D, Gurbani D, Tae HS, Riddle SM, Sim T, Marto JA, Janne PA, Crews CM, Gray NS. Pharmacological targeting of the pseudokinase Her3. Nat Chem Biol. 2014;10:1006–1012. doi: 10.1038/nchembio.1658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Gustafson JL, Neklesa TK, Cox CS, Roth AG, Buckley DL, Tae HS, Sundberg TB, Stagg DB, Hines J, McDonnell DP, Norris JD, Crews CM. Small-Molecule-Mediated Degradation of the Androgen Receptor through Hydrophobic Tagging. Angew Chem Int Ed Engl. 2015;54:9659–9662. doi: 10.1002/anie.201503720. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Raina K, Noblin DJ, Serebrenik YV, Adams A, Zhao C, Crews CM. Targeted protein destabilization reveals an estrogen-mediated ER stress response. Nat Chem Biol. 2014;10:957–962. doi: 10.1038/nchembio.1638. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Long MJ, Gollapalli DR, Hedstrom L. Inhibitor mediated protein degradation. Chem Biol. 2012;19:629–637. doi: 10.1016/j.chembiol.2012.04.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Coffey RT, Shi Y, Long MJ, Marr MT, 2nd, Hedstrom L. Ubiquilin Mediated Small Molecule Inhibition of Mammalian Target of Rapamycin Complex 1 (mTORC1) Signaling. J Biol Chem. 2016;291:5221–5233. doi: 10.1074/jbc.M115.691584. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Callis J, Ling R. Preparation, characterization, and use of tagged ubiquitins. Methods in enzymology. 2005;399:51–64. doi: 10.1016/S0076-6879(05)99004-6. [DOI] [PubMed] [Google Scholar]
- 22.Kapuria V, Peterson LF, Fang D, Bornmann WG, Talpaz M, Donato NJ. Deubiquitinase inhibition by small-molecule WP1130 triggers aggresome formation and tumor cell apoptosis. Cancer Res. 2010;70:9265–9276. doi: 10.1158/0008-5472.CAN-10-1530. [DOI] [PubMed] [Google Scholar]
- 23.Schulman BA, Harper JW. Ubiquitin-like protein activation by E1 enzymes: the apex for downstream signalling pathways. Nat Rev Mol Cell Biol. 2009;10:319–331. doi: 10.1038/nrm2673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Yang Y, Kitagaki J, Dai RM, Tsai YC, Lorick KL, Ludwig RL, Pierre SA, Jensen JP, Davydov IV, Oberoi P, Li CC, Kenten JH, Beutler JA, Vousden KH, Weissman AM. Inhibitors of ubiquitin-activating enzyme (E1), a new class of potential cancer therapeutics. Cancer Res. 2007;67:9472–9481. doi: 10.1158/0008-5472.CAN-07-0568. [DOI] [PubMed] [Google Scholar]
- 25.Mi L, Gan N, Cheema A, Dakshanamurthy S, Wang X, Yang DC, Chung FL. Cancer preventive isothiocyanates induce selective degradation of cellular alpha- and beta-tubulins by proteasomes. J Biol Chem. 2009;284:17039–17051. doi: 10.1074/jbc.M901789200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Kapuria V, Peterson LF, Showalter HD, Kirchhoff PD, Talpaz M, Donato NJ. Protein cross-linking as a novel mechanism of action of a ubiquitin-activating enzyme inhibitor with anti-tumor activity. Biochem Pharmacol. 2011;82:341–349. doi: 10.1016/j.bcp.2011.05.012. [DOI] [PubMed] [Google Scholar]
- 27.D’Arcy P, Brnjic S, Olofsson MH, Fryknas M, Lindsten K, De Cesare M, Perego P, Sadeghi B, Hassan M, Larsson R, Linder S. Inhibition of proteasome deubiquitinating activity as a new cancer therapy. Nat Med. 2011;17:1636–1640. doi: 10.1038/nm.2536. [DOI] [PubMed] [Google Scholar]
- 28.Schnell JR, Dyson HJ, Wright PE. Structure, dynamics, and catalytic function of dihydrofolate reductase. Annu Rev Biophys Biomol Struct. 2004;33:119–140. doi: 10.1146/annurev.biophys.33.110502.133613. [DOI] [PubMed] [Google Scholar]
- 29.Tey LH, Loveridge EJ, Swanwick RS, Flitsch SL, Allemann RK. Highly site-selective stability increases by glycosylation of dihydrofolate reductase. FEBS J. 2010;277:2171–2179. doi: 10.1111/j.1742-4658.2010.07634.x. [DOI] [PubMed] [Google Scholar]
- 30.Ionescu RM, Smith VF, O’Neill JC, Jr, Matthews CR. Multistate equilibrium unfolding of Escherichia coli dihydrofolate reductase: thermodynamic and spectroscopic description of the native, intermediate, and unfolded ensembles. Biochemistry. 2000;39:9540–9550. doi: 10.1021/bi000511y. [DOI] [PubMed] [Google Scholar]
- 31.Huang L, Chen CH. Proteasome regulators: activators and inhibitors. Curr Med Chem. 2009;16:931–939. doi: 10.2174/092986709787581860. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Kisselev AF, Kaganovich D, Goldberg AL. Binding of hydrophobic peptides to several non-catalytic sites promotes peptide hydrolysis by all active sites of 20 S proteasomes. Evidence for peptide-induced channel opening in the alpha-rings. The Journal of biological chemistry. 2002;277:22260–22270. doi: 10.1074/jbc.M112360200. [DOI] [PubMed] [Google Scholar]
- 33.Chu-Ping M, Slaughter CA, DeMartino GN. Purification and characterization of a protein inhibitor of the 20S proteasome (macropain) Biochim Biophys Acta. 1992;1119:303–311. doi: 10.1016/0167-4838(92)90218-3. [DOI] [PubMed] [Google Scholar]
- 34.Cascio P. PA28alphabeta: the enigmatic magic ring of the proteasome? Biomolecules. 2014;4:566–584. doi: 10.3390/biom4020566. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Huang L, Ho P, Chen CH. Activation and inhibition of the proteasome by betulinic acid and its derivatives. FEBS Lett. 2007;581:4955–4959. doi: 10.1016/j.febslet.2007.09.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Moravec RA, O’Brien MA, Daily WJ, Scurria MA, Bernad L, Riss TL. Cell-based bioluminescent assays for all three proteasome activities in a homogeneous format. Anal Biochem. 2009;387:294–302. doi: 10.1016/j.ab.2009.01.016. [DOI] [PubMed] [Google Scholar]
- 37.Dantuma NP, Lindsten K, Glas R, Jellne M, Masucci MG. Short-lived green fluorescent proteins for quantifying ubiquitin/proteasome-dependent proteolysis in living cells. Nat Biotechnol. 2000;18:538–543. doi: 10.1038/75406. [DOI] [PubMed] [Google Scholar]
- 38.Lawson AP, Long MJ, Coffey RT, Qian Y, Weerapana E, El Oualid F, Hedstrom L. Naturally Occurring Isothiocyanates Exert Anticancer Effects by Inhibiting Deubiquitinating Enzymes. Cancer Res. 2015;75:5130–5142. doi: 10.1158/0008-5472.CAN-15-1544. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Liu G, Rogers J, Murphy CT, Rongo C. EGF signalling activates the ubiquitin proteasome system to modulate C. elegans lifespan. EMBO J. 2011;30:2990–3003. doi: 10.1038/emboj.2011.195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Crews CM, Georg G, Wang S. Inducing Protein Degradation as a Therapeutic Strategy. J Med Chem. 2016 doi: 10.1021/acs.jmedchem.6b00735. [DOI] [PubMed] [Google Scholar]
- 41.Olivares AO, Baker TA, Sauer RT. Mechanistic insights into bacterial AAA+ proteases and protein-remodelling machines. Nat Rev Microbiol. 2016;14:33–44. doi: 10.1038/nrmicro.2015.4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Baugh JM, Viktorova EG, Pilipenko EV. Proteasomes can degrade a significant proportion of cellular proteins independent of ubiquitination. J Mol Biol. 2009;386:814–827. doi: 10.1016/j.jmb.2008.12.081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Pickering AM, Davies KJ. Degradation of damaged proteins: the main function of the 20S proteasome. Prog Mol Biol Transl Sci. 2012;109:227–248. doi: 10.1016/B978-0-12-397863-9.00006-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Tummino PJ, Copeland RA. Residence time of receptor-ligand complexes and its effect on biological function. Biochemistry. 2008;47:5481–5492. doi: 10.1021/bi8002023. [DOI] [PubMed] [Google Scholar]
- 45.Morphy R, Kay C, Rankovic Z. From magic bullets to designed multiple ligands. Drug Discov Today. 2004;9:641–651. doi: 10.1016/S1359-6446(04)03163-0. [DOI] [PubMed] [Google Scholar]
- 46.Ben-Nissan G, Sharon M. Regulating the 20S proteasome ubiquitin-independent degradation pathway. Biomolecules. 2014;4:862–884. doi: 10.3390/biom4030862. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Tanahashi N, Murakami Y, Minami Y, Shimbara N, Hendil KB, Tanaka K. Hybrid proteasomes. Induction by interferon-gamma and contribution to ATP-dependent proteolysis. The Journal of biological chemistry. 2000;275:14336–14345. doi: 10.1074/jbc.275.19.14336. [DOI] [PubMed] [Google Scholar]
- 48.Brooks P, Fuertes G, Murray RZ, Bose S, Knecht E, Rechsteiner MC, Hendil KB, Tanaka K, Dyson J, Rivett J. Subcellular localization of proteasomes and their regulatory complexes in mammalian cells. Biochem J. 2000;346(Pt 1):155–161. [PMC free article] [PubMed] [Google Scholar]
- 49.Babbitt SE, Kiss A, Deffenbaugh AE, Chang YH, Bailly E, Erdjument-Bromage H, Tempst P, Buranda T, Sklar LA, Baumler J, Gogol E, Skowyra D. ATP hydrolysis-dependent disassembly of the 26S proteasome is part of the catalytic cycle. Cell. 2005;121:553–565. doi: 10.1016/j.cell.2005.03.028. [DOI] [PubMed] [Google Scholar]
- 50.Fabre B, Lambour T, Delobel J, Amalric F, Monsarrat B, Burlet-Schiltz O, Bousquet-Dubouch MP. Subcellular distribution and dynamics of active proteasome complexes unraveled by a workflow combining in vivo complex cross-linking and quantitative proteomics. Mol Cell Proteomics. 2013;12:687–699. doi: 10.1074/mcp.M112.023317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Jariel-Encontre I, Bossis G, Piechaczyk M. Ubiquitin-independent degradation of proteins by the proteasome. Biochim Biophys Acta. 2008;1786:153–177. doi: 10.1016/j.bbcan.2008.05.004. [DOI] [PubMed] [Google Scholar]
- 52.Janse DM, Crosas B, Finley D, Church GM. Localization to the proteasome is sufficient for degradation. J Biol Chem. 2004;279:21415–21420. doi: 10.1074/jbc.M402954200. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.