Abstract
DNA sequences of cloned histone coding sequences and spacers of sea urchin species that diverged long ago in evolution were compared. The highly repeated H4 and H3 genes active during early embryogenesis had evolved (in their silent sites) at a rate (0.5-0.6% base changes/Myr) similar to single-copy protein-coding genes and nearly as fast as spacer DNA (0.7% base changes/Myr) and unique DNA. Thus, evolution in the major histone genes conforms to a universal evolutionary clock based on the rate of base sequence change. By contrast, the H4 and H3 coding sequences and a non-transcribed spacer of the DNA clone h19 of Psammechinus miliaris show an exceptionally low rate of sequence evolution only 1/100 to 1/200 that predicted from the clock hypothesis. According to the classical model of gene inheritance, the h19 DNA sequences in the Psammechinus genome require unusual conservation mechanisms by selection at the level of the gene and spacer sequences. An alternative explanation could be recent horizontal gene transfer of a histone gene cluster from the very distantly related Strongylocentrotus dröbachiensis to the P. miliaris genome.
Keywords: codon selection, spacer DNA, mutational drift, horizontal gene transfer
Full text
PDF

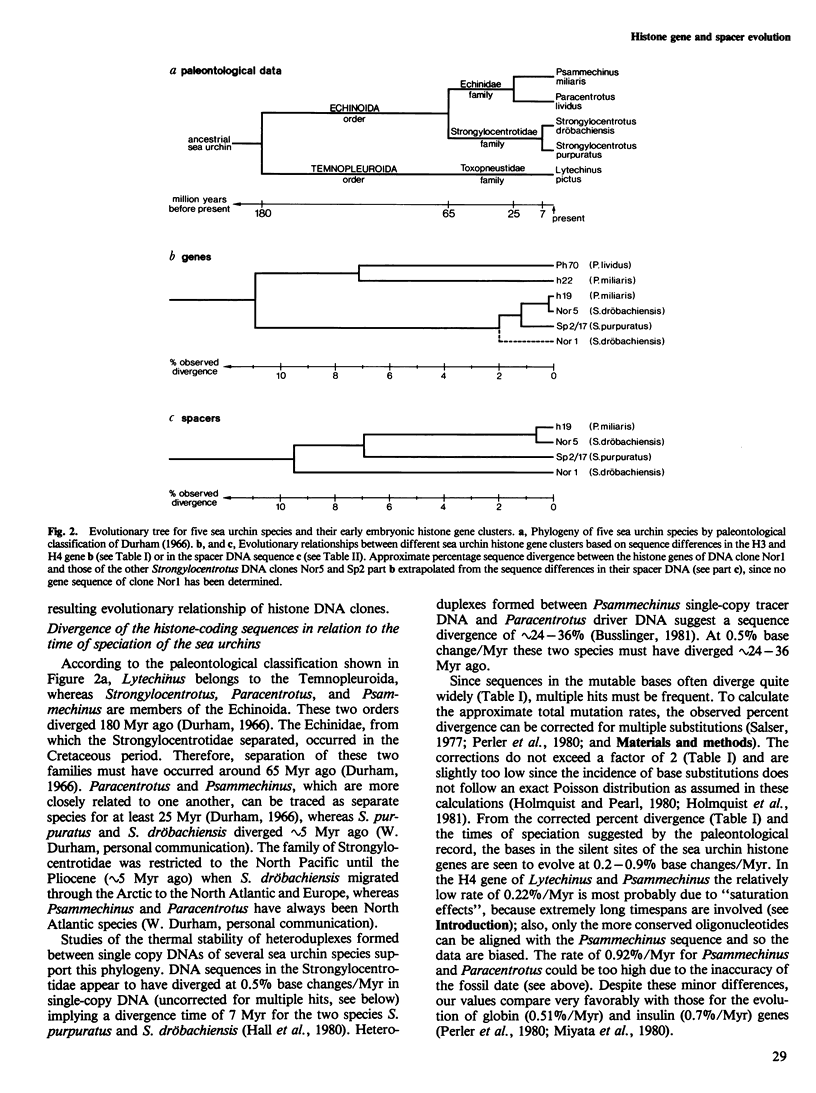
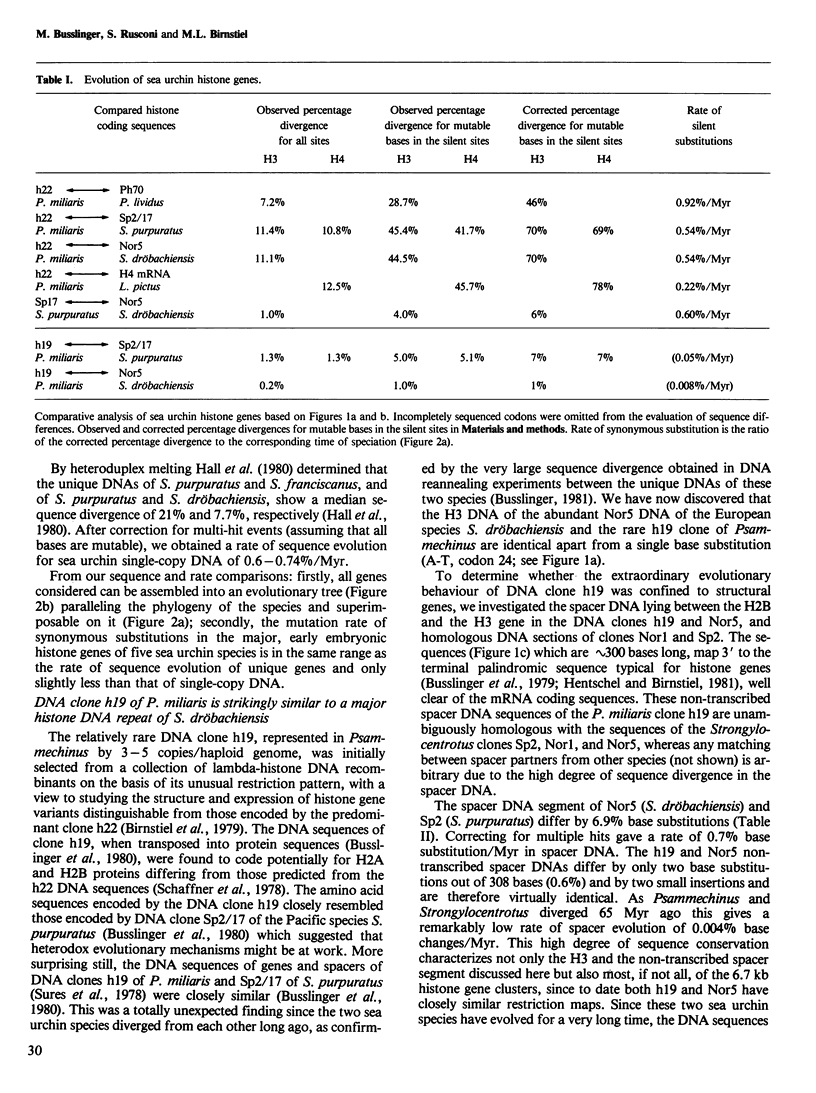
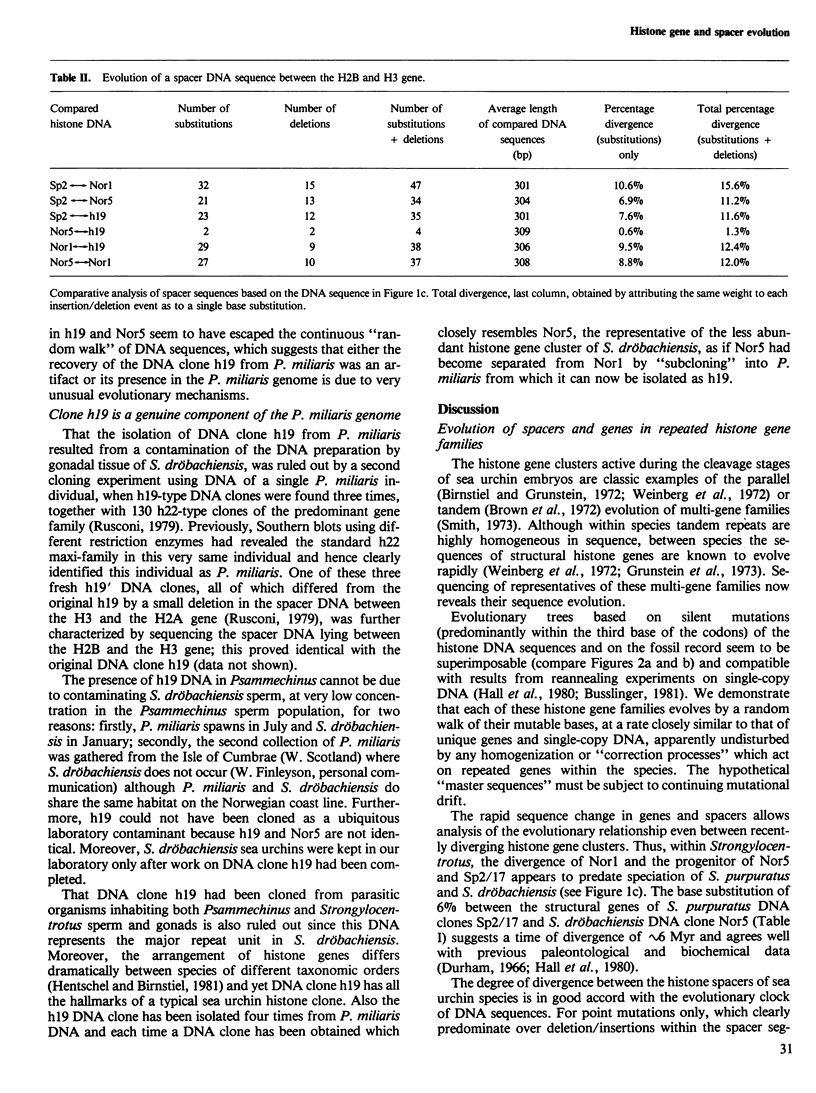


Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Benveniste R. E., Todaro G. J. Evolution of C-type viral genes: inheritance of exogenously acquired viral genes. Nature. 1974 Dec 6;252(5483):456–459. doi: 10.1038/252456a0. [DOI] [PubMed] [Google Scholar]
- Benveniste R. E., Todaro G. J. Evolution of type C viral genes: evidence for an Asian origin of man. Nature. 1976 May 13;261(5556):101–108. doi: 10.1038/261101a0. [DOI] [PubMed] [Google Scholar]
- Bishop J. M. Enemies within: the genesis of retrovirus oncogenes. Cell. 1981 Jan;23(1):5–6. doi: 10.1016/0092-8674(81)90263-4. [DOI] [PubMed] [Google Scholar]
- Boseley P., Moss T., Mächler M., Portmann R., Birnstiel M. Sequence organization of the spacer DNA in a ribosomal gene unit of Xenopus laevis. Cell. 1979 May;17(1):19–31. doi: 10.1016/0092-8674(79)90291-5. [DOI] [PubMed] [Google Scholar]
- Bossi L., Ruth J. R. The influence of codon context on genetic code translation. Nature. 1980 Jul 10;286(5769):123–127. doi: 10.1038/286123a0. [DOI] [PubMed] [Google Scholar]
- Brown D. D., Wensink P. C., Jordan E. A comparison of the ribosomal DNA's of Xenopus laevis and Xenopus mulleri: the evolution of tandem genes. J Mol Biol. 1972 Jan 14;63(1):57–73. doi: 10.1016/0022-2836(72)90521-9. [DOI] [PubMed] [Google Scholar]
- Busslinger M., Portmann R., Birnsteil M. L. A regulatory sequence near the 3' end of sea urchin histone genes. Nucleic Acids Res. 1979 Jul 11;6(9):2997–3008. doi: 10.1093/nar/6.9.2997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Busslinger M., Portmann R., Irminger J. C., Birnstiel M. L. Ubiquitous and gene-specific regulatory 5' sequences in a sea urchin histone DNA clone coding for histone protein variants. Nucleic Acids Res. 1980 Mar 11;8(5):957–977. doi: 10.1093/nar/8.5.957. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Calos M. P., Miller J. H. Transposable elements. Cell. 1980 Jul;20(3):579–595. doi: 10.1016/0092-8674(80)90305-0. [DOI] [PubMed] [Google Scholar]
- Childs G., Maxson R., Cohn R. H., Kedes L. Orphons: dispersed genetic elements derived from tandem repetitive genes of eucaryotes. Cell. 1981 Mar;23(3):651–663. doi: 10.1016/0092-8674(81)90428-1. [DOI] [PubMed] [Google Scholar]
- Clarkson S. G., Smith H. O., Schaffner W., Gross K. W., Birnstiel M. L. Integration of eukaryotic genes for 5S RNA and histone proteins into a phage lambda receptor. Nucleic Acids Res. 1976 Oct;3(10):2617–2632. doi: 10.1093/nar/3.10.2617. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Crick F. General model for the chromosomes of higher organisms. Nature. 1971 Nov 5;234(5323):25–27. doi: 10.1038/234025a0. [DOI] [PubMed] [Google Scholar]
- Efstratiadis A., Kafatos F. C., Maniatis T. The primary structure of rabbit beta-globin mRNA as determined from cloned DNA. Cell. 1977 Apr;10(4):571–585. doi: 10.1016/0092-8674(77)90090-3. [DOI] [PubMed] [Google Scholar]
- Fitch W. M. Estimating the total number of nucleotide substitutions since the common ancestor of a pair of homologous genes: comparison of several methods and three beta hemoglobin messenger RNA's. J Mol Evol. 1980 Dec;16(3-4):153–209. doi: 10.1007/BF01804976. [DOI] [PubMed] [Google Scholar]
- Grunstein M., Diamond K. E., Knoppel E., Grunstein J. E. Comparison of the early histone H4 gene sequence of Strongylocentrotus purpuratus with maternal, early, and late histone H4 mRNA sequences. Biochemistry. 1981 Mar 3;20(5):1216–1223. doi: 10.1021/bi00508a025. [DOI] [PubMed] [Google Scholar]
- Grunstein M., Schedl P. Isolation and sequence analysis of sea urchin (Lytechinus pictus) histone H4 messenger RNA. J Mol Biol. 1976 Jun 25;104(2):323–349. doi: 10.1016/0022-2836(76)90275-8. [DOI] [PubMed] [Google Scholar]
- Grunstein M., Schedl P., Kedes L. Sequence analysis and evolution of sea urchin (Lytechinus pictus and Strongylocentrotus purpuratus) histone H4 messenger RNAs. J Mol Biol. 1976 Jun 25;104(2):351–369. doi: 10.1016/0022-2836(76)90276-x. [DOI] [PubMed] [Google Scholar]
- Hentschel C. C., Birnstiel M. L. The organization and expression of histone gene families. Cell. 1981 Aug;25(2):301–313. doi: 10.1016/0092-8674(81)90048-9. [DOI] [PubMed] [Google Scholar]
- Holmquist R., Pearl D. Theoretical foundations for quantitative paleogenetics. Part III: The molecular divergence of nucleic acids and proteins for the case of genetic events of unequal probability. J Mol Evol. 1980 Dec;16(3-4):211–267. doi: 10.1007/BF01804977. [DOI] [PubMed] [Google Scholar]
- Iida S., Meyer J., Arber W. Genesis and natural history of IS-mediated transposons. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 1):27–43. doi: 10.1101/sqb.1981.045.01.006. [DOI] [PubMed] [Google Scholar]
- Kedes L. H., Chang A. C., Houseman D., Cohen S. N. Isolation of histone genes from unfractionated sea urchin DNA by subculture cloning in E. coli. Nature. 1975 Jun 12;255(5509):533–538. doi: 10.1038/255533a0. [DOI] [PubMed] [Google Scholar]
- Kedes L. H. Histone genes and histone messengers. Annu Rev Biochem. 1979;48:837–870. doi: 10.1146/annurev.bi.48.070179.004201. [DOI] [PubMed] [Google Scholar]
- King J. L., Jukes T. H. Non-Darwinian evolution. Science. 1969 May 16;164(3881):788–798. doi: 10.1126/science.164.3881.788. [DOI] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. A new method for sequencing DNA. Proc Natl Acad Sci U S A. 1977 Feb;74(2):560–564. doi: 10.1073/pnas.74.2.560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miyata T., Hayashida H. Extraordinarily high evolutionary rate of pseudogenes: evidence for the presence of selective pressure against changes between synonymous codons. Proc Natl Acad Sci U S A. 1981 Sep;78(9):5739–5743. doi: 10.1073/pnas.78.9.5739. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miyata T., Yasunaga T., Nishida T. Nucleotide sequence divergence and functional constraint in mRNA evolution. Proc Natl Acad Sci U S A. 1980 Dec;77(12):7328–7332. doi: 10.1073/pnas.77.12.7328. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perler F., Efstratiadis A., Lomedico P., Gilbert W., Kolodner R., Dodgson J. The evolution of genes: the chicken preproinsulin gene. Cell. 1980 Jun;20(2):555–566. doi: 10.1016/0092-8674(80)90641-8. [DOI] [PubMed] [Google Scholar]
- Russell G. J., Walker P. M., Elton R. A., Subak-Sharpe J. H. Doublet frequency analysis of fractionated vertebrate nuclear DNA. J Mol Biol. 1976 Nov;108(1):1–23. doi: 10.1016/s0022-2836(76)80090-3. [DOI] [PubMed] [Google Scholar]
- Salser W. Globin mRNA sequences: analysis of base pairing and evolutionary implications. Cold Spring Harb Symp Quant Biol. 1978;42(Pt 2):985–1002. doi: 10.1101/sqb.1978.042.01.099. [DOI] [PubMed] [Google Scholar]
- Schaffner W., Kunz G., Daetwyler H., Telford J., Smith H. O., Birnstiel M. L. Genes and spacers of cloned sea urchin histone DNA analyzed by sequencing. Cell. 1978 Jul;14(3):655–671. doi: 10.1016/0092-8674(78)90249-0. [DOI] [PubMed] [Google Scholar]
- Shaw R. F., Bloom R. W., Bowman J. E. Hemoglobin and the genetic code. Evolution of protection against somatic mutation. J Mol Evol. 1977 May 13;9(3):225–230. doi: 10.1007/BF01796111. [DOI] [PubMed] [Google Scholar]
- Smith G. P. Unequal crossover and the evolution of multigene families. Cold Spring Harb Symp Quant Biol. 1974;38:507–513. doi: 10.1101/sqb.1974.038.01.055. [DOI] [PubMed] [Google Scholar]
- Spinelli G., Gianguzza F., Casano C., Acierno P., Burckhardt J. Evidences of two different sets of histone genes active during embryogenesis of the sea urchin Paracentrotus lividus. Nucleic Acids Res. 1979 Feb;6(2):545–560. doi: 10.1093/nar/6.2.545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sures I., Lowry J., Kedes L. H. The DNA sequence of sea urchin (S. purpuratus) H2A, H2B and H3 histone coding and spacer regions. Cell. 1978 Nov;15(3):1033–1044. doi: 10.1016/0092-8674(78)90287-8. [DOI] [PubMed] [Google Scholar]
- Walker P. M. How different are the DNAs from related animals? Nature. 1968 Jul 20;219(5151):228–232. doi: 10.1038/219228a0. [DOI] [PubMed] [Google Scholar]
- Weinberg E. S., Birnstiel M. L., Purdom I. F., Williamson R. Genes coding for polysomal 9S RNA of sea urchins: conservation and divergence. Nature. 1972 Nov 24;240(5378):225–228. doi: 10.1038/240225a0. [DOI] [PubMed] [Google Scholar]
- Wilson A. C., Carlson S. S., White T. J. Biochemical evolution. Annu Rev Biochem. 1977;46:573–639. doi: 10.1146/annurev.bi.46.070177.003041. [DOI] [PubMed] [Google Scholar]
- van Ooyen A., van den Berg J., Mantei N., Weissmann C. Comparison of total sequence of a cloned rabbit beta-globin gene and its flanking regions with a homologous mouse sequence. Science. 1979 Oct 19;206(4416):337–344. doi: 10.1126/science.482942. [DOI] [PubMed] [Google Scholar]



