Abstract
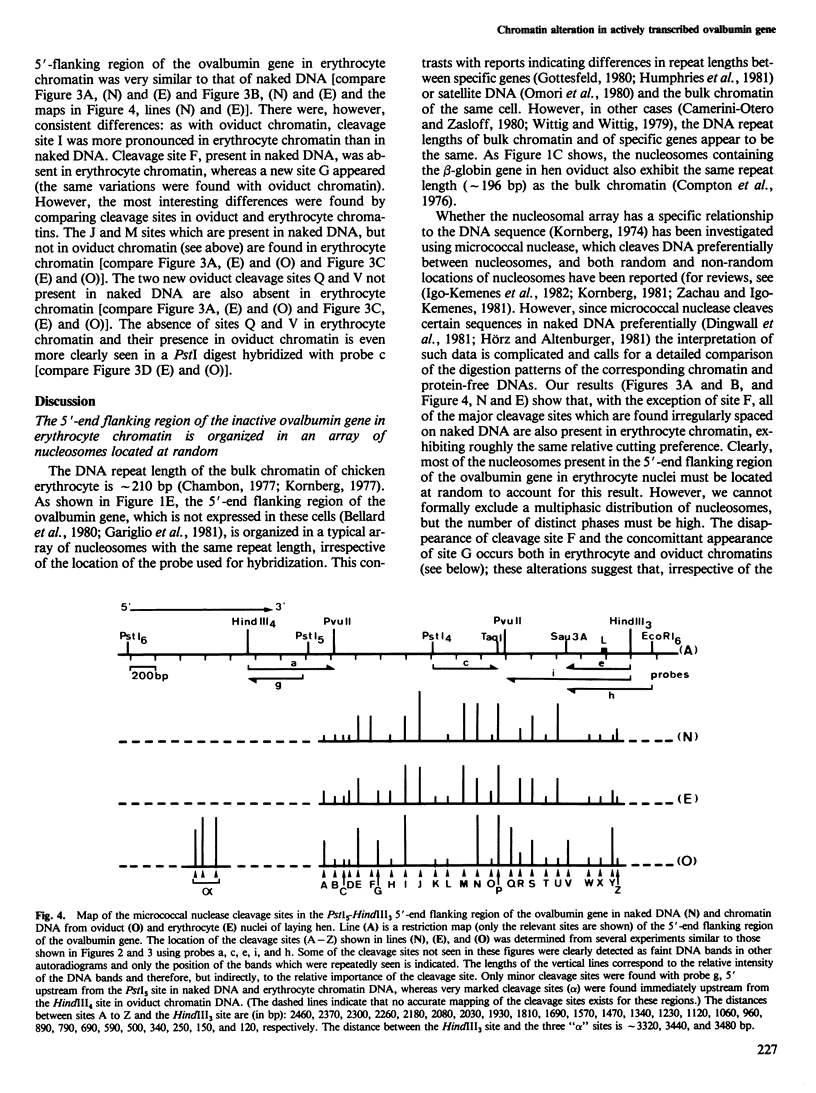
We examined the chromatin organizations of approximately 3 kb of DNA in the 5'-end flanking region of the ovalbumin gene in chicken erythrocyte and oviduct cell nuclei. With specific DNA probes and an indirect end-labeling technique, we analysed the pattern of the DNA fragments obtained after micrococcal nuclease digestion and generated comparative maps of the nuclease cuts. This region of the chicken genome displays a "typical" chromatin arrangement in erythrocyte nuclei, with nucleosomes apparently positioned at random. In contrast, in oviduct nuclei, the same region has an "altered" chromatin structure, and lacks a typical nucleosomal array. The existence of specifically positioned proteins and of alterations in the DNA secondary structure in this region of the oviduct chromatin is suggested by comparison of the nuclease cleavage maps which reveals specific changes: disappearance of nuclease cuts present in "naked" and erythrocyte chromatin DNAs, and appearance of new cuts absent from these DNAs.
Full text
PDF
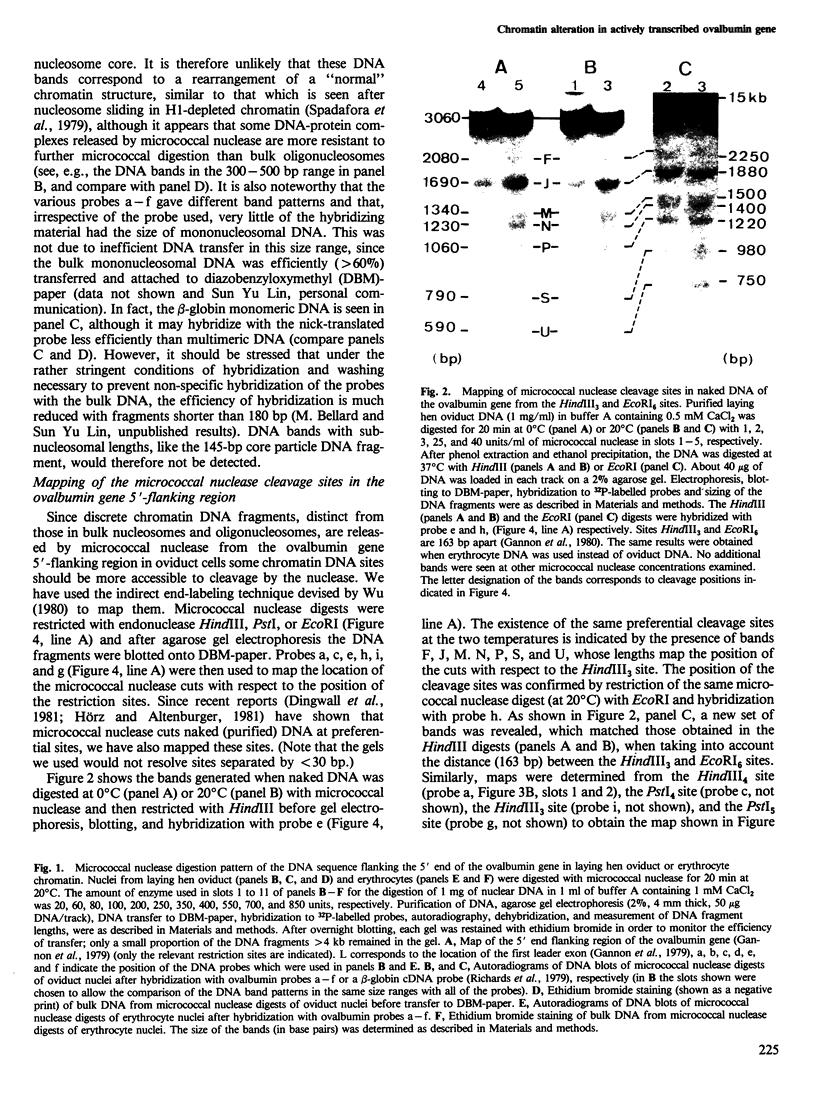




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bellard M., Gannon F., Chambon P. Nucleosome structure III: the structure and transcriptional activity of the chromatin containing the ovalbumin and globin genes in chick oviduct nuclei. Cold Spring Harb Symp Quant Biol. 1978;42(Pt 2):779–791. doi: 10.1101/sqb.1978.042.01.078. [DOI] [PubMed] [Google Scholar]
- Bellard M., Kuo M. T., Dretzen G., Chambon P. Differential nuclease sensitivity of the ovalbumin and beta-globin chromatin regions in erythrocytes and oviduct cells of laying hen. Nucleic Acids Res. 1980 Jun 25;8(12):2737–2750. doi: 10.1093/nar/8.12.2737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bloom K. S., Anderson J. N. Conformation of ovalbumin and globin genes in chromatin during differential gene expression. J Biol Chem. 1979 Oct 25;254(20):10532–10539. [PubMed] [Google Scholar]
- Breathnach R., Chambon P. Organization and expression of eucaryotic split genes coding for proteins. Annu Rev Biochem. 1981;50:349–383. doi: 10.1146/annurev.bi.50.070181.002025. [DOI] [PubMed] [Google Scholar]
- Camerini-Otero R. D., Zasloff M. A. Nucleosomal packaging of the thymidine kinase gene of herpes simplex virus transferred into mouse cells: an actively expressed single-copy gene. Proc Natl Acad Sci U S A. 1980 Sep;77(9):5079–5083. doi: 10.1073/pnas.77.9.5079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chambon P. Summary: the molecular biology of the eukaryotic genome is coming of age. Cold Spring Harb Symp Quant Biol. 1978;42(Pt 2):1209–1234. doi: 10.1101/sqb.1978.042.01.122. [DOI] [PubMed] [Google Scholar]
- Cremisi C. The appearance of DNase I hypersensitive sites at the 5' end of the late SV40 genes is correlated with the transcriptional switch. Nucleic Acids Res. 1981 Nov 25;9(22):5949–5964. doi: 10.1093/nar/9.22.5949. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dingwall C., Lomonossoff G. P., Laskey R. A. High sequence specificity of micrococcal nuclease. Nucleic Acids Res. 1981 Jun 25;9(12):2659–2673. doi: 10.1093/nar/9.12.2659. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dretzen G., Bellard M., Sassone-Corsi P., Chambon P. A reliable method for the recovery of DNA fragments from agarose and acrylamide gels. Anal Biochem. 1981 Apr;112(2):295–298. doi: 10.1016/0003-2697(81)90296-7. [DOI] [PubMed] [Google Scholar]
- Felsenfeld G. Chromatin. Nature. 1978 Jan 12;271(5641):115–122. doi: 10.1038/271115a0. [DOI] [PubMed] [Google Scholar]
- Gannon F., Jeltsch J. M., Perrin F. A detailed comparison of the 5'-end of the ovalbumin gene cloned from chicken oviduct and erythrocyte DNA. Nucleic Acids Res. 1980 Oct 10;8(19):4405–4421. doi: 10.1093/nar/8.19.4405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gannon F., O'Hare K., Perrin F., LePennec J. P., Benoist C., Cochet M., Breathnach R., Royal A., Garapin A., Cami B. Organisation and sequences at the 5' end of a cloned complete ovalbumin gene. Nature. 1979 Mar 29;278(5703):428–434. doi: 10.1038/278428a0. [DOI] [PubMed] [Google Scholar]
- Garel A., Axel R. Selective digestion of transcriptionally active ovalbumin genes from oviduct nuclei. Proc Natl Acad Sci U S A. 1976 Nov;73(11):3966–3970. doi: 10.1073/pnas.73.11.3966. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gariglio P., Bellard M., Chambon P. Clustering of RNA polymerase B molecules in the 5' moiety of the adult beta-globin gene of hen erythrocytes. Nucleic Acids Res. 1981 Jun 11;9(11):2589–2598. doi: 10.1093/nar/9.11.2589. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gottesfeld J. M. Organization of 5S genes in chromatin of Xenopus laevis. Nucleic Acids Res. 1980 Feb 25;8(4):905–922. [PMC free article] [PubMed] [Google Scholar]
- Groudine M., Eisenman R., Weintraub H. Chromatin structure of endogenous retroviral genes and activation by an inhibitor of DNA methylation. Nature. 1981 Jul 23;292(5821):311–317. doi: 10.1038/292311a0. [DOI] [PubMed] [Google Scholar]
- Herbomel P., Saragosti S., Blangy D., Yaniv M. Fine structure of the origin-proximal DNAase I-hypersensitive region in wild-type and EC mutant polyoma. Cell. 1981 Sep;25(3):651–658. doi: 10.1016/0092-8674(81)90172-0. [DOI] [PubMed] [Google Scholar]
- Hewish D. R., Burgoyne L. A. Chromatin sub-structure. The digestion of chromatin DNA at regularly spaced sites by a nuclear deoxyribonuclease. Biochem Biophys Res Commun. 1973 May 15;52(2):504–510. doi: 10.1016/0006-291x(73)90740-7. [DOI] [PubMed] [Google Scholar]
- Humphries S. E., Young D., Carroll D. Chromatin structure of the 5S ribonucleic acid genes of Xenopus laevis. Biochemistry. 1979 Jul 24;18(15):3223–3231. doi: 10.1021/bi00582a006. [DOI] [PubMed] [Google Scholar]
- Hörz W., Altenburger W. Sequence specific cleavage of DNA by micrococcal nuclease. Nucleic Acids Res. 1981 Jun 25;9(12):2643–2658. doi: 10.1093/nar/9.12.2643. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jackson D. A., McCready S. J., Cook P. R. RNA is synthesized at the nuclear cage. Nature. 1981 Aug 6;292(5823):552–555. doi: 10.1038/292552a0. [DOI] [PubMed] [Google Scholar]
- Jakobovits E. B., Bratosin S., Aloni Y. A nucleosome-free region in SV40 minichromosomes. Nature. 1980 May 22;285(5762):263–265. doi: 10.1038/285263a0. [DOI] [PubMed] [Google Scholar]
- Keene M. A., Corces V., Lowenhaupt K., Elgin S. C. DNase I hypersensitive sites in Drosophila chromatin occur at the 5' ends of regions of transcription. Proc Natl Acad Sci U S A. 1981 Jan;78(1):143–146. doi: 10.1073/pnas.78.1.143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keene M. A., Elgin S. C. Micrococcal nuclease as a probe of DNA sequence organization and chromatin structure. Cell. 1981 Nov;27(1 Pt 2):57–64. doi: 10.1016/0092-8674(81)90360-3. [DOI] [PubMed] [Google Scholar]
- Kornberg R. D. Chromatin structure: a repeating unit of histones and DNA. Science. 1974 May 24;184(4139):868–871. doi: 10.1126/science.184.4139.868. [DOI] [PubMed] [Google Scholar]
- Kornberg R. D. Structure of chromatin. Annu Rev Biochem. 1977;46:931–954. doi: 10.1146/annurev.bi.46.070177.004435. [DOI] [PubMed] [Google Scholar]
- Kornberg R. The location of nucleosomes in chromatin: specific or statistical. Nature. 1981 Aug 13;292(5824):579–580. doi: 10.1038/292579a0. [DOI] [PubMed] [Google Scholar]
- Kuo M. T., Mandel J. L., Chambon P. DNA methylation: correlation with DNase I sensitivity of chicken ovalbumin and conalbumin chromatin. Nucleic Acids Res. 1979 Dec 20;7(8):2105–2113. doi: 10.1093/nar/7.8.2105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lamb M. M., Daneholt B. Characterization of active transcription units in Balbiani rings of Chironomus tentans. Cell. 1979 Aug;17(4):835–848. doi: 10.1016/0092-8674(79)90324-6. [DOI] [PubMed] [Google Scholar]
- Lawson G. M., Tsai M. J., O'Malley B. W. Deoxyribonuclease I sensitivity of the nontranscribed sequences flanking the 5' and 3' ends of the ovomucoid gene and the ovalbumin and its related X and Y genes in hen oviduct nuclei. Biochemistry. 1980 Sep 16;19(19):4403–4441. doi: 10.1021/bi00560a004. [DOI] [PubMed] [Google Scholar]
- LeMeur M., Glanville N., Mandel J. L., Gerlinger P., Palmiter R., Chambon P. The ovalbumin gene family: hormonal control of X and Y gene transcription and mRNA accumulation. Cell. 1981 Feb;23(2):561–571. doi: 10.1016/0092-8674(81)90152-5. [DOI] [PubMed] [Google Scholar]
- Levy A., Noll M. Chromatin fine structure of active and repressed genes. Nature. 1981 Jan 15;289(5794):198–203. doi: 10.1038/289198a0. [DOI] [PubMed] [Google Scholar]
- Lohr D. E. Detailed analysis of the nucleosomal organization of transcribed DNA in yeast chromatin. Biochemistry. 1981 Oct 13;20(21):5966–5972. doi: 10.1021/bi00524a007. [DOI] [PubMed] [Google Scholar]
- Mathis D., Oudet P., Chambon P. Structure of transcribing chromatin. Prog Nucleic Acid Res Mol Biol. 1980;24:1–55. doi: 10.1016/s0079-6603(08)60670-4. [DOI] [PubMed] [Google Scholar]
- McGhee J. D., Wood W. I., Dolan M., Engel J. D., Felsenfeld G. A 200 base pair region at the 5' end of the chicken adult beta-globin gene is accessible to nuclease digestion. Cell. 1981 Nov;27(1 Pt 2):45–55. doi: 10.1016/0092-8674(81)90359-7. [DOI] [PubMed] [Google Scholar]
- McKnight G. S., Palmiter R. D. Transcriptional regulation of the ovalbumin and conalbumin genes by steroid hormones in chick oviduct. J Biol Chem. 1979 Sep 25;254(18):9050–9058. [PubMed] [Google Scholar]
- Omori A., Igo-Kemenes T., Zachau H. G. Different repeat lengths in rat satellite I DNA containing chromatin and bulk chromatin. Nucleic Acids Res. 1980 Nov 25;8(22):5363–5375. doi: 10.1093/nar/8.22.5363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Palmiter R. D. Quantitation of parameters that determine the rate of ovalbumin synthesis. Cell. 1975 Mar;4(3):189–189. doi: 10.1016/0092-8674(75)90167-1. [DOI] [PubMed] [Google Scholar]
- Richards R. I., Shine J., Ullrich A., Wells J. R., Goodman H. M. Molecular cloning and sequence analysis of adult chicken betal globin cDNA. Nucleic Acids Res. 1979 Nov 10;7(5):1137–1146. doi: 10.1093/nar/7.5.1137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Samal B., Worcel A., Louis C., Schedl P. Chromatin structure of the histone genes of D. melanogaster. Cell. 1981 Feb;23(2):401–409. doi: 10.1016/0092-8674(81)90135-5. [DOI] [PubMed] [Google Scholar]
- Saragosti S., Moyne G., Yaniv M. Absence of nucleosomes in a fraction of SV40 chromatin between the origin of replication and the region coding for the late leader RNA. Cell. 1980 May;20(1):65–73. doi: 10.1016/0092-8674(80)90235-4. [DOI] [PubMed] [Google Scholar]
- Scott W. A., Wigmore D. J. Sites in simian virus 40 chromatin which are preferentially cleaved by endonucleases. Cell. 1978 Dec;15(4):1511–1518. doi: 10.1016/0092-8674(78)90073-9. [DOI] [PubMed] [Google Scholar]
- Senear A. W., Palmiter R. D. Multiple structural features are responsible for the nuclease sensitivity of the active ovalbumin gene. J Biol Chem. 1981 Feb 10;256(3):1191–1198. [PubMed] [Google Scholar]
- Spadafora C., Oudet P., Chambon P. Rearrangement of chromatin structure induced by increasing ionic strength and temperature. Eur J Biochem. 1979 Oct;100(1):225–235. doi: 10.1111/j.1432-1033.1979.tb02053.x. [DOI] [PubMed] [Google Scholar]
- Stalder J., Larsen A., Engel J. D., Dolan M., Groudine M., Weintraub H. Tissue-specific DNA cleavages in the globin chromatin domain introduced by DNAase I. Cell. 1980 Jun;20(2):451–460. doi: 10.1016/0092-8674(80)90631-5. [DOI] [PubMed] [Google Scholar]
- Sundin O., Varshavsky A. Staphylococcal nuclease makes a single non-random cut in the simian virus 40 viral minichromosome. J Mol Biol. 1979 Aug 15;132(3):535–546. doi: 10.1016/0022-2836(79)90274-2. [DOI] [PubMed] [Google Scholar]
- Varshavsky A. J., Sundin O. H., Bohn M. J. SV40 viral minichromosome: preferential exposure of the origin of replication as probed by restriction endonucleases. Nucleic Acids Res. 1978 Oct;5(10):3469–3477. doi: 10.1093/nar/5.10.3469. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Varshavsky A. J., Sundin O., Bohn M. A stretch of "late" SV40 viral DNA about 400 bp long which includes the origin of replication is specifically exposed in SV40 minichromosomes. Cell. 1979 Feb;16(2):453–466. doi: 10.1016/0092-8674(79)90021-7. [DOI] [PubMed] [Google Scholar]
- Weintraub H., Larsen A., Groudine M. Alpha-Globin-gene switching during the development of chicken embryos: expression and chromosome structure. Cell. 1981 May;24(2):333–344. doi: 10.1016/0092-8674(81)90323-8. [DOI] [PubMed] [Google Scholar]
- Wilhelm X., Champagne M. Dissociation de la nucléoprotéine d'érythrocytes de poulets par les sels. Eur J Biochem. 1969 Aug;10(1):102–109. [PubMed] [Google Scholar]
- Wittig B., Wittig S. A phase relationship associates tRNA structural gene sequences with nucleosome cores. Cell. 1979 Dec;18(4):1173–1183. doi: 10.1016/0092-8674(79)90230-7. [DOI] [PubMed] [Google Scholar]
- Wong Y. C., O'Connell P., Rosbash M., Elgin S. C. DNase I hypersensitive sites of the chromatin for Drosophila melanogaster ribosomal protein 49 gene. Nucleic Acids Res. 1981 Dec 21;9(24):6749–6762. doi: 10.1093/nar/9.24.6749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu C., Gilbert W. Tissue-specific exposure of chromatin structure at the 5' terminus of the rat preproinsulin II gene. Proc Natl Acad Sci U S A. 1981 Mar;78(3):1577–1580. doi: 10.1073/pnas.78.3.1577. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu C. The 5' ends of Drosophila heat shock genes in chromatin are hypersensitive to DNase I. Nature. 1980 Aug 28;286(5776):854–860. doi: 10.1038/286854a0. [DOI] [PubMed] [Google Scholar]
- Wu C., Wong Y. C., Elgin S. C. The chromatin structure of specific genes: II. Disruption of chromatin structure during gene activity. Cell. 1979 Apr;16(4):807–814. doi: 10.1016/0092-8674(79)90096-5. [DOI] [PubMed] [Google Scholar]
- Zachau H. G., Igo-Kemenes T. Face to phase with nucleosomes. Cell. 1981 Jun;24(3):597–598. doi: 10.1016/0092-8674(81)90084-2. [DOI] [PubMed] [Google Scholar]