Abstract
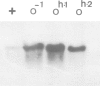
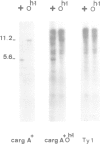
The Saccharomyces cerevisiae gene cargA + or CAR1 , encoding arginase has been cloned by recovering function in transformed yeast cells. It was used to analyse RNA and chromosomal DNA from six strains bearing cis-dominant regulatory mutations leading to constitutive arginase synthesis. The DNA from the four cargA + O- strains in which constitutive arginase synthesis was independent of the mating-type functions showed no detectable differences with the wild- typye . The cargA + O- mutations were, therefore, small alterations, possibly single base substitutions. On the other hand, the cargA + Oh-1 and cargA + Oh-2 mutations, leading to a constitutive and mating-type dependent arginase synthesis, were identified as insertions. Their size and restriction pattern strongly suggested that they were induced by the Ty1 yeast transposable element. This was confirmed by cloning and analysis of the cargA + Oh-1 mutant gene. The concentration of arginase RNA was significantly increased in the mutants, indicating that the regulation of arginase synthesis was exerted, at least in part, at the level of RNA synthesis or stability. In the cargA + Oh-2 strain the Ty1 element was located at a distance of approximately 600 base pairs from the insertion present in the cargA + Oh-1 strain. This result suggests either a surprisingly large arginase regulatory region or an indirect influence of the Ty1 element on gene expression over long distances.
Full text
PDF






Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Alwine J. C., Kemp D. J., Parker B. A., Reiser J., Renart J., Stark G. R., Wahl G. M. Detection of specific RNAs or specific fragments of DNA by fractionation in gels and transfer to diazobenzyloxymethyl paper. Methods Enzymol. 1979;68:220–242. doi: 10.1016/0076-6879(79)68017-5. [DOI] [PubMed] [Google Scholar]
- Beggs J. D. Transformation of yeast by a replicating hybrid plasmid. Nature. 1978 Sep 14;275(5676):104–109. doi: 10.1038/275104a0. [DOI] [PubMed] [Google Scholar]
- Birnboim H. C., Doly J. A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res. 1979 Nov 24;7(6):1513–1523. doi: 10.1093/nar/7.6.1513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cameron J. R., Loh E. Y., Davis R. W. Evidence for transposition of dispersed repetitive DNA families in yeast. Cell. 1979 Apr;16(4):739–751. doi: 10.1016/0092-8674(79)90090-4. [DOI] [PubMed] [Google Scholar]
- Chevallier M. R., Bloch J. C., Lacroute F. Transcriptional and translational expression of a chimeric bacterial-yeast plasmid in yeasts. Gene. 1980 Oct;11(1-2):11–19. doi: 10.1016/0378-1119(80)90082-7. [DOI] [PubMed] [Google Scholar]
- Crabeel M., Charlier D., Cunin R., Glansdorff N. Cloning and endonuclease restriction analysis of argF and of the control region of the argECBH bipolar operon in Escherichia coli. Gene. 1979 Mar;5(3):207–231. doi: 10.1016/0378-1119(79)90079-9. [DOI] [PubMed] [Google Scholar]
- Cryer D. R., Eccleshall R., Marmur J. Isolation of yeast DNA. Methods Cell Biol. 1975;12:39–44. doi: 10.1016/s0091-679x(08)60950-4. [DOI] [PubMed] [Google Scholar]
- Davis R. W., Thomas M., Cameron J., St John T. P., Scherer S., Padgett R. A. Rapid DNA isolations for enzymatic and hybridization analysis. Methods Enzymol. 1980;65(1):404–411. doi: 10.1016/s0076-6879(80)65051-4. [DOI] [PubMed] [Google Scholar]
- Deschamps J., Wiame J. M. Mating-type effect on cis mutations leading to constitutivity of ornithine transaminase in diploid cells of Saccharomyces cerevisiae. Genetics. 1979 Jul;92(3):749–758. doi: 10.1093/genetics/92.3.749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dubois E., Grenson M., Wiame J. M. Release of the "ammonia effect" on three catabolic enzymes by NADP-specific glutamate dehydrogenaseless mutations in Saccharomyces cerevisiae. Biochem Biophys Res Commun. 1973 Feb 20;50(4):967–972. doi: 10.1016/0006-291x(73)91500-3. [DOI] [PubMed] [Google Scholar]
- Dubois E., Grenson M., Wiame J. M. The participation of the anabolic glutamate dehydrogenase in the nitrogen catabolite repression of arginase in Saccharomyces cerevisiae. Eur J Biochem. 1974 Oct 2;48(2):603–616. doi: 10.1111/j.1432-1033.1974.tb03803.x. [DOI] [PubMed] [Google Scholar]
- Dubois E., Hiernaux D., Grennon M., Wiame J. M. Specific induction of catabolism and its relation to repression of biosynthesis in arginine metabolism of Saccharomyces cerevisiae. J Mol Biol. 1978 Jul 15;122(4):383–406. doi: 10.1016/0022-2836(78)90417-5. [DOI] [PubMed] [Google Scholar]
- Dubois E., Vissers S., Grenson M., Wiame J. M. Glutamine and ammonia in nitrogen catabolite repression of Saccharomyces cerevisiae. Biochem Biophys Res Commun. 1977 Mar 21;75(2):233–239. doi: 10.1016/0006-291x(77)91033-6. [DOI] [PubMed] [Google Scholar]
- Errede B., Cardillo T. S., Sherman F., Dubois E., Deschamps J., Wiame J. M. Mating signals control expression of mutations resulting from insertion of a transposable repetitive element adjacent to diverse yeast genes. Cell. 1980 Nov;22(2 Pt 2):427–436. doi: 10.1016/0092-8674(80)90353-0. [DOI] [PubMed] [Google Scholar]
- Gafner J., Philippsen P. The yeast transposon Ty1 generates duplications of target DNA on insertion. Nature. 1980 Jul 24;286(5771):414–418. doi: 10.1038/286414a0. [DOI] [PubMed] [Google Scholar]
- Grenson M., Hou C. Ammonia inhibition of the general amino acid permease and its suppression in NADPH-specific glutamate dehydrogenaseless mutants of saccharomyces cerevisiae. Biochem Biophys Res Commun. 1972 Aug 21;48(4):749–756. doi: 10.1016/0006-291x(72)90670-5. [DOI] [PubMed] [Google Scholar]
- Grunstein M., Hogness D. S. Colony hybridization: a method for the isolation of cloned DNAs that contain a specific gene. Proc Natl Acad Sci U S A. 1975 Oct;72(10):3961–3965. doi: 10.1073/pnas.72.10.3961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hinnen A., Hicks J. B., Fink G. R. Transformation of yeast. Proc Natl Acad Sci U S A. 1978 Apr;75(4):1929–1933. doi: 10.1073/pnas.75.4.1929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- LOWRY O. H., ROSEBROUGH N. J., FARR A. L., RANDALL R. J. Protein measurement with the Folin phenol reagent. J Biol Chem. 1951 Nov;193(1):265–275. [PubMed] [Google Scholar]
- Lemoine Y., Dubois E., Wiame J. M. The regulation of urea amidolyase of Saccharomyces cerevisiae: mating type influence on a constitutivity mutation acting in cis. Mol Gen Genet. 1978 Nov 9;166(3):251–258. [PubMed] [Google Scholar]
- McMaster G. K., Carmichael G. G. Analysis of single- and double-stranded nucleic acids on polyacrylamide and agarose gels by using glyoxal and acridine orange. Proc Natl Acad Sci U S A. 1977 Nov;74(11):4835–4838. doi: 10.1073/pnas.74.11.4835. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Messenguy F., Penninckx M., Wiame J. M. Interaction between arginase and ornithine carbamoyltransferase in Saccharomyces cerevisiae. The regulatory site for ornithine. Eur J Biochem. 1971 Sep 24;22(2):277–286. doi: 10.1111/j.1432-1033.1971.tb01542.x. [DOI] [PubMed] [Google Scholar]
- Messenguy F., Wiame J. -M. The control of ornithinetranscarbamylase activity by arginase in Saccharomyces cerevisiae. FEBS Lett. 1969 Apr;3(1):47–49. doi: 10.1016/0014-5793(69)80093-1. [DOI] [PubMed] [Google Scholar]
- Penninckx M., Simon J. P., Wiame J. M. Interaction between arginase and L-ornithine carbamoyltransferase in Saccharomyces cerevisiae. Purification of S. cerevisiae enzymes and evidence that these enzymes as well as rat-liver arginase are trimers. Eur J Biochem. 1974 Nov 15;49(2):429–442. doi: 10.1111/j.1432-1033.1974.tb03848.x. [DOI] [PubMed] [Google Scholar]
- Petes T. D., Broach J. R., Wensink P. C., Hereford L. M., Fink G. R., Botstein D. Isolation and analysis of recombinant DNA molecules containing yeast DNA. Gene. 1978 Sep;4(1):37–49. doi: 10.1016/0378-1119(78)90013-6. [DOI] [PubMed] [Google Scholar]
- Ramos F., Wiame J. M. Synthesis and activation of asparagine in asparagine auxotrophs of Saccharomyces cerevisiae. Eur J Biochem. 1979 Mar;94(2):409–417. doi: 10.1111/j.1432-1033.1979.tb12908.x. [DOI] [PubMed] [Google Scholar]
- Rigby P. W., Dieckmann M., Rhodes C., Berg P. Labeling deoxyribonucleic acid to high specific activity in vitro by nick translation with DNA polymerase I. J Mol Biol. 1977 Jun 15;113(1):237–251. doi: 10.1016/0022-2836(77)90052-3. [DOI] [PubMed] [Google Scholar]
- Roeder G. S., Farabaugh P. J., Chaleff D. T., Fink G. R. The origins of gene instability in yeast. Science. 1980 Sep 19;209(4463):1375–1380. doi: 10.1126/science.6251544. [DOI] [PubMed] [Google Scholar]
- Rothstein R. J., Sherman F. Dependence on mating type for the overproduction of iso-2-cytochrome c in the yeast mutant CYC7-H2. Genetics. 1980 Apr;94(4):891–898. doi: 10.1093/genetics/94.4.891. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Southern E. Gel electrophoresis of restriction fragments. Methods Enzymol. 1979;68:152–176. doi: 10.1016/0076-6879(79)68011-4. [DOI] [PubMed] [Google Scholar]
- Waldron C., Lacroute F. Effect of growth rate on the amounts of ribosomal and transfer ribonucleic acids in yeast. J Bacteriol. 1975 Jun;122(3):855–865. doi: 10.1128/jb.122.3.855-865.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]