Abstract
Depression is one of the most prevalent and debilitating psychiatric disorders worldwide. Recently, we showed that both relatively short and relatively long cytosine–adenine–guanine (CAG) repeats in the huntingtin gene (HTT) are associated with an increased risk of lifetime depression. However, to what extent the variations in CAG repeat length in the other eight polyglutamine disease-associated genes (PDAGs) are associated with depression is still unknown. We determined the CAG repeat sizes of ATXN1, ATXN2, ATXN3, CACNA1A, ATXN7, TBP, ATN1 and AR in two well-characterized Dutch cohorts—the Netherlands Study of Depression and Anxiety and the Netherlands Study of Depression in Older Persons—including 2165 depressed and 1058 non-depressed individuals—aged 18–93 years. The association between PDAG CAG repeat size and the risk for depression was assessed via binary logistic regression. We found that the odds ratio (OR) for lifetime depression was significantly higher for individuals with >10, compared with subjects with ≤10, CAG repeats in both ATXN7 alleles (OR=1.90, confidence interval (CI) 1.26–2.85). For TBP we found a similar association: A CAG repeat length exceeding the median in both alleles was associated with an increased risk for lifetime depression (OR=1.33, CI 1.00–1.76). In conclusion, we observed that carriers of either ATXN7 or TBP alleles with relatively large CAG repeat sizes in both alleles had a substantially increased risk of lifetime depression. Our findings provide critical evidence for the notion that repeat polymorphisms can act as complex genetic modifiers of depression.
Introduction
Depression is one of the most common psychiatric disorders with an estimated lifetime prevalence of 14.6% in high-income and 11.1% in low-income countries.1 Worldwide, depression is responsible for 74.5 million disability-adjusted life years2 and in 2010 the total economic burden of depression in the United States alone was estimated at $210.5 billion,3 indicating that depression is a major strain on society. In order to create new and more effective therapeutic and preventive strategies, unravelling the pathogenesis of depression is imperative and elucidation of its genetic determinants is a critical step in this process.
The heritability of major depressive disorder (MDD) has been estimated to be between 30 and 50%.4 Nevertheless, genome-wide association studies (GWASs) have shown limited success in identifying its genetic basis. In a mega-analysis with more than 18 000 subjects from European ancestry, no single-nucleotide polymorphism (SNP) reached genome-wide significance.4 However, recently, a meta-analysis of results from three studies, together containing 180 866 individuals, found two lead SNPs associated with depression.5 In addition, another study combining three cohorts in a joint analysis with a total of 478 240 subjects found 17 independent SNPs significantly associated with a diagnosis of MDD,6 suggesting that GWASs can successfully identify genetic associations with highly polygenic phenotypes. However, the identified effect sizes were small. Moreover, to gain sufficient statistical power the sample sizes required were extremely large and, as a consequence, unavoidably heterogeneous. Therefore, although GWASs have contributed greatly to genetic mapping of complex human traits, their success has been limited by the fact that, aside from SNPs, these studies are unsuitable to assess the contribution of other genetic polymorphisms, especially DNA repeat sequences.7
Tandem repeats constitute ~3% of the human genome, a higher percentage than the entirety of the protein-coding sequences,8 substantially contributing to genetic variation.9, 10 Many rare hereditary disorders are caused by expansions of simple DNA repeat sequences.11 However, the association of DNA repeat sequences with more common diseases is largely unknown. Polyglutamine diseases are the most prevalent disorders caused by an expanded DNA repeat sequence.11, 12 These diseases are caused by an expansion of a trinucleotide repeat (cytosine–adenine–guanine (CAG)) in the translated region of various genes. As CAG encodes the amino acid glutamine, the expansion in the trinucleotide repeat sequence results in an elongated polyglutamine domain in the associated proteins.12 The most common polyglutamine disease is Huntington disease, a severe neurodegenerative disorder characterized by both motor and neuropsychiatric impairment. Huntington disease is caused by a CAG repeat expansion in exon 1 of the huntingtin (HTT) gene.13, 14 Recently, we demonstrated that both relatively short and relatively large CAG repeats in the longer HTT allele are associated with lifetime depression, suggesting that repeat polymorphisms could also act as complex genetic modifiers of depression and account for part of its ‘missing heritability’.15
Apart from Huntington disease, eight other polyglutamine disorders, all of which are neurodegenerative disorders, are frequently associated with considerable neuropsychiatric impairment (Table 1).16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35 These rare disorders result from relatively large repeat expansions in polyglutamine disease-associated genes (PDAGs). However, to what extent more common repeat length variations in the normal range of these genes could act as genetic modifiers of depression in the general population is still unknown. Hence, here we aimed to assess the contribution of CAG repeat length variations in these other PDAGs to depression susceptibility using data from two well-defined Dutch cohorts: The Netherlands Study of Depression and Anxiety (NESDA) and the Netherlands Study of Depression in Old Persons (NESDO).
Table 1. Summary genotyped polyglutamine disease-associated genes.
| Gene | Disease | Protein |
CAG repeat ranges |
Allele | Mean | Median | N | Range | No. of intermediatea | No. of pathologicala | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Normal | Pathological | ||||||||||
| ATXN1 | SCA1 | Ataxin-1 | 6–39 | 41–83 | Short | 29.22 | 29.00 | 3228 | 18–35 | 0 | 0 |
| Long | 30.76 | 30.00 | 3228 | 26–39 | 0 | 0 | |||||
| ATXN2 | SCA2 | Ataxin-2 | 14–32 | 33–500 | Short | 21.91 | 22.00 | 3236 | 14–24 | 0 | 0 |
| Long | 22.35 | 22.00 | 3236 | 22–36 | 0 | 2 (33, 36) | |||||
| ATXN3 | SCA3 | Ataxin-3 | 12–44 | 52–87 | Short | 18.96 | 20.00 | 3235 | 13–33 | 0 | 0 |
| Long | 24.27 | 23.00 | 3235 | 13–49 | 1 (49) | 0 | |||||
| CACNA1A | SCA6 | CACNA1A | 4–18 | 20–33 | Short | 10.52 | 11.00 | 3220 | 4–14 | 0 | 0 |
| Long | 12.45 | 13.00 | 3220 | 4–22 | 0 | 1 (22) | |||||
| ATXN7 | SCA7 | Ataxin-7 | 3–19 | 37–460 | Short | 10.04 | 10.00 | 2979 | 7–15 | 0 | 0 |
| Long | 10.85 | 10.00 | 2979 | 7–19 | 0 | 0 | |||||
| TBP | SCA17 | TBP | 25–43 | 45–66 | Short | 36.31 | 37.00 | 3238 | 27–39 | 0 | 0 |
| Long | 37.89 | 38.00 | 3238 | 32–48 | 1 (44) | 1 (48) | |||||
| ATN1 | DRPLA | Atrophin-1 | 3–38 | 48–93 | Short | 12.36 | 14.00 | 3228 | 5–22 | 0 | 0 |
| Long | 15.47 | 15.00 | 3228 | 8–30 | 0 | 0 | |||||
| AR | SMBA | Androgen receptor | 6–36 | 38–72 | Short | 20.80 | 21.00 | 3228 | 8–36 | 0 | 0 |
| Long | 22.90 | 23.00 | 3228 | 8–36 | 0 | 0 | |||||
Abbreviations: CACNA1A, calcium channel, voltage-dependent P/Q type, α 1A subunit; CAG, cytosine–adenine–guanine; DRPLA, dentatorubropallidoluysian atrophy; SBMA, spinal bulbar muscular atrophy; SCA, spinocerebellar ataxia; TBP, thymine-adenine-thymine-adenine (TATA) box-binding protein.
Materials and methods
We genotyped eight PDAGs (ATXN1, ATXN2, ATXN3, CACNA1A, ATXN7, TBP, ATN1 and AR) in all participants with sufficient amounts of DNA available from blood samples of two well-characterized Dutch cohorts: the NESDA (cohort 1) and the NESDO (cohort 2) cohorts (Table 1 and Supplementary Figures 1–17).
Cohort 1
The NESDA is a cohort study among 2981 participants aged 18–65 years.36 The participants were recruited from the general population, general practices and mental health-care institutes. The sample included 1973 subjects with a lifetime diagnosis of depression and/or dysthymia (including 1925 patients with MDD), 635 subjects with a lifetime anxiety disorder without lifetime depression and 373 healthy controls.36 Diagnoses were made in accordance with the Diagnostic and Statistical Manuel of Mental Disorders Fourth Edition criteria using the WHO Composite International Diagnostic Interview.37
Cohort 2
The NESDO is a cohort study among 510 participants aged 60–93 years.38 The participants were recruited from both general practices and mental health-care institutes. The sample included 378 depressed individuals (360 with MDD) and 132 healthy controls. The same methods for diagnosing depression were used as in the NESDA.38
Genotyping
A polymerase chain reaction (PCR) was performed in a TProfessional thermocycler (Biometra, Westburg, Leusden, the Netherlands) with labelled primers flanking the CAG stretch of the PDAGs (Biolegio, Nijmegen, the Netherlands; Supplementary Table 1). The PCR was performed using 10 ng of genomic DNA, 1 × OneTaq mastermix (New England Biolabs, Ipswich, MA, USA, OneTaq Hot start with GC Buffer mastermix), 1 μl of primer Mix A or B (Supplementary Table 1) and Aqua B. Braun water to a final volume of 10 μl. The PCR was run with 27 cycles of 30 s, denaturation at 94 °C, 1 min of annealing at 60 °C and 2 min elongation at 68 °C, preceded by 5 min of initial denaturation at 94 °C. Final elongation was performed at 69 °C for 5 min. Every PCR included a negative control without genomic DNA and a reference sample of CEPH 1347-02 genomic DNA. The PCR products were run on an ABI 3730 automatic DNA sequencer (Applied Biosystems, Foster City, CA, USA) and analysed using the GeneMarker software version 2.4.0. (SoftGenetics, State College, PA, USA). For every analysis, we included three controls with known CAG repeat lengths for each PDAG to assure that every run was performed reliably. All assessments were made with cases and controls randomized on plates and blinding with respect to disease status information.
Statistical analysis
Binary logistic regression was used to assess whether CAG repeat sizes in the two alleles of each PDAG were associated with the risk of lifetime depression. For each PDAG, in an initial model, the presence of lifetime depression (that is, MDD and/or dysthymia) was set as the dependent variable and the CAG repeat lengths of both alleles were used as the independent variables. To assess interaction effects between the two alleles or nonlinear effects, a product term of the two alleles and a quadratic term for each allele were added to the model. In case the product term or the quadratic terms were not significant, these predictors were removed from the model and the analysis was repeated. Subsequently, we adjusted the results for the effects of sex, age and education level (coded as ‘basic’, ‘intermediate’ and ‘high’36, 38) in order to assess whether the effect of CAG repeat size variations was independent of these well-established risk factors for depression. The Nagelkerke R2 was used to assess the proportion of variability explained by the predictors in the model. To account for potential effects of heteroscedasticity and influential points, all statistical significance tests were based on robust estimators of s.e.'s. Moreover, to visualise our results as well as to assure that the results were not unduly affected by violated model assumptions, we also applied a non-parametric method: we calculated the odds ratios (ORs) for a lifetime diagnosis of depression per CAG repeat group. The CAG repeat groups were defined based on the median CAG repeat size of each allele. A division as such resulted in three adequately sized groups: I, both alleles⩽their median; II, the relatively shorter allele⩽the median and the relatively longer allele>the median; III, both alleles>their median. Subsequently, the ORs were compared with the Fisher’s exact test. In order to account for multiple testing, we applied a false discovery rate correction as described by Benjamini and Hochberg, assuming eight independent tests with a two-sided α of 0.05.39 All data are displayed as means and 95% confidence intervals (CIs), unless otherwise specified. All analyses were performed in SPSS version 23.0 (IBM SPSS Statistics for Windows, IBM, Armonk, NY, USA).
Results
Risk of lifetime depression increases with higher ATXN7 CAG repeat sizes
In total, we were able to genotype a total of 2979 participants for ATXN7 (Table 1). This included 1998 depressed subjects and 981 non-depressed subjects. For 512 individuals, too little DNA material was available to determine the CAG repeat length in ATXN7 (Supplementary Table 2). These lacking samples were missing completely at random. The CAG repeat lengths ranged from 7 to 19 repeats (Table 1).
We found a significant association between the risk of lifetime depression and the CAG repeat length of ATXN7 in the shorter allele (β=0.202, P=0.006). Adjusting for the effects of gender, age and education hardly changed these results (β=0.184, P=0.013; Supplementary Table 3). After adjusting for multiple testing, this association remained significant. Inclusion of the CAG repeat length of the ATXN7 allele in the model increased the proportion of explained genetic variation from the baseline model including only gender, age and level of education by 0.004 (that is, the R2 increased from 0.024 to 0.028). Considering our control group included subjects with anxiety disorders without comorbid depression (n=348), we also performed a sensitivity analysis by excluding these individuals from the control group. Neither the parameter estimates nor their significance were materially altered by this procedure (β=0.215, P=0.009).
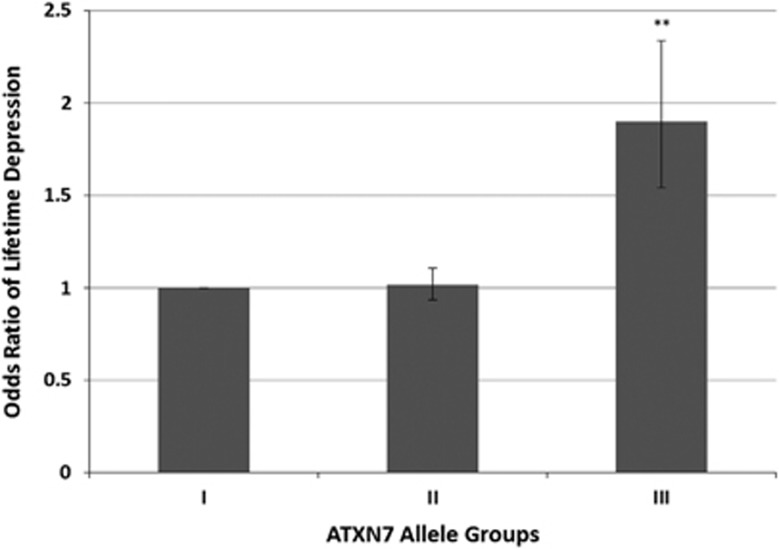
For the non-parametric method, we divided all subjects based on the median CAG repeat size for each ATXN7 allele (median relatively short allele=10, median relatively long allele=10) and compared the ORs of lifetime depression among the groups (Table 2). The group containing the largest number of individuals was used as the reference category. Comparing the other groups to the reference category demonstrated the significant effect of the shorter allele on the risk of lifetime depression (Figure 1). When the relatively short allele and thus also the relatively long allele both contained a CAG repeat length larger than 10 repeats, the odds for lifetime depression almost doubled compared with the reference category in which both ATXN7 alleles had a CAG repeat number equal to or smaller than the median.
Table 2. The distribution of participants in three ATXN7 categories.
| ATXN7 category | Allele | Mean | Range | Lifetime depressiona | No lifetime depressionb | Odds ratio (95% CI) |
|---|---|---|---|---|---|---|
| Ic | Short | 9.94 | 7–10 | 1220 | 619 | 1.00 |
| Long | 10.00 | 7–10 | ||||
| II | Short | 9.97 | 7–10 | 662 | 331 | 1.01 (0.86–1.20) |
| Long | 12.18 | 11–19 | ||||
| III | Short | 11.90 | 11–15 | 116 | 31 | (1.26–2.85)d |
| Long | 12.54 | 11–18 |
Abbreviations: I, both alleles of ATXN7 contain a CAG repeat number ⩽10; II, the relatively longer ATXN7 allele contains a CAG repeat number >10 and the relatively shorter allele contains a CAG repeat number ⩽10; III, both alleles of ATXN7 contain a CAG repeat number >10; CAG, cytosine–adenine–guanine; CI, confidence interval.
Figures represent the number of subjects per ATXN7 category in the group with lifetime depression.
Figures represent the number of subjects per ATXN7 category in the group with no lifetime depression.
This category contains the largest number of subjects and was therefore chosen as the reference category for odds ratio calculation.
P<0.01 by Fisher’s exact test in comparison with the reference category.
Figure 1.
Odds ratio for lifetime depression per ATXN7 category. The odds ratio for lifetime depression increases significantly when both the relatively long ATXN7 allele and the relatively short ATXN7 allele exceed the median CAG repeat number of 10. The odds of having lifetime depression almost doubles. The group having both alleles with a CAG repeat number ⩽10 was the largest and, therefore, was set as the reference category. I, both alleles of ATXN7 contain a CAG repeat number ⩽10; II, the relatively longer ATXN7 allele contains a CAG repeat number >10 and the relatively shorter allele contains a CAG repeat number ⩽10; III, both alleles of ATXN7 contain a CAG repeat number >10. Error bars indicate±s.e. **P<0.01 by the Fisher’s exact test in comparison with the reference category; CAG, cytosine–adenine–guanine.
TBP CAG repeat sizes in both alleles interact to affect the risk of lifetime depression
A total of 3238 individuals were genotyped for TBP (Table 1). This included 2180 people diagnosed with lifetime depression and 1058 non-depressed people. For 253 individuals, too little DNA material was available to determine the CAG repeat length in TBP (Supplementary Table 2). The lacking samples were missing completely at random. For TBP, we found CAG repeat lengths ranging between 27 and 48 repeats (Table 1).
We found the interaction between the number of CAG repeats in the two TBP alleles to be significantly associated with the risk of lifetime depression (TBP short allele: β=−2.270, P=0.004; TBP long allele: β=−2.112, P=0.006; TBP interaction term: β=0.060, P=0.005). When adjusting for the effects of gender, age and education, the results hardly changed (TBP short allele: β=−2.302, P=0.004; TBP long allele: β=−2.150, P=0.005; TBP interaction term: β=0.061, P=0.005; Supplementary Table 3). Furthermore, the results remained significant after performing the sensitivity analysis and applying the false discovery rate correction.39 The proportion of explained genetic variation increased by 0.005 from the baseline model including only gender, age and level of education (that is, the R2 increased from 0.023 to 0.028).
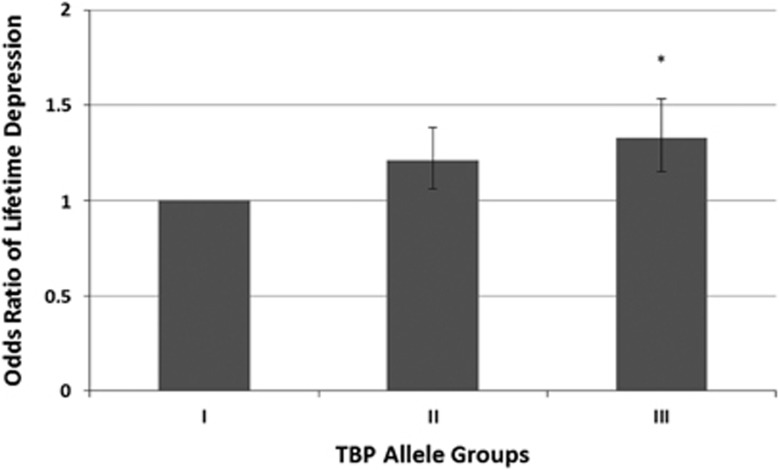
For the non-parametric method, we divided all subjects in three groups based on the median CAG repeat size for each TBP allele (median relatively short TBP allele=37, median relatively long TBP allele=38) and compared the ORs of lifetime depression among the groups (Table 3). The largest group was set as the reference category. The comparison of the groups demonstrated the significant effect of the interaction between the relatively short TBP allele and the relatively long TBP allele on the risk of depression. The risk of lifetime depression was significantly higher when both TBP alleles had a CAG repeat length exceeding their median compared with the reference category, in which both alleles had a CAG repeat number equal to or lower than their median (Figure 2).
Table 3. The distribution of participants in three TBP categories.
| TBP category | Allele | Mean | Range | Lifetime depressiona | No lifetime depressionb | Odds ratio (95% CI) |
|---|---|---|---|---|---|---|
| Ic | Short | 35.78 | 27–37 | 1462 | 716 | 1.00 |
| Long | 37.47 | 32–38 | ||||
| II | Short | 36.04 | 30–37 | 230 | 93 | 1.21 (0.94–1.57) |
| Long | 39.30 | 39–48 | ||||
| III | Short | 38.09 | 38–39 | 201 | 74 | 1.33 (1.00–1.76)d |
| Long | 39.28 | 39–44 |
I, both TBP alleles contain a CAG repeat number⩽their median; II, the relatively longer TBP allele contains a CAG repeat number>the median and the relatively shorter TBP allele contains a CAG repeat number⩽the median; III, both TBP alleles contain a CAG repeat number>their median (median short allele=37, median long allele=38); CAG, cytosine–adenine–guanine; CI, confidence interval.
Figures represent the number of subjects per TBP category in the group with lifetime depression.
Figures represent the number of subjects per TBP category in the group with no lifetime depression.
This category contains the largest number of subjects and was therefore chosen as the reference category for odds ratio calculation.
P<0.05 by Fisher’s exact test in comparison with the reference category.
Figure 2.
Odds ratio for lifetime depression per TBP category. The odds ratio for lifetime depression increases significantly when both the relatively shorter TBP allele and the relatively longer TBP allele exceed their median CAG repeat number (median short allele=37, median long allele=38). The group with both alleles being equal to or smaller than their medians was the largest and, therefore, is defined as the reference category. I, both TBP alleles contain a CAG repeat number ⩽ their median; II, the relatively longer TBP allele contains a CAG repeat number>the median and the relatively shorter TBP allele contains a CAG repeat number ⩽ the median; III, both TBP alleles contain a CAG repeat number > their median. Error bars indicate±one s.e. *P<0.05 by the Fisher’s exact test in comparison with the reference category. CAG, cytosine–adenine–guanine; TBP, thymine-adenine-thymine-adenine (TATA) box-binding protein.
CAG repeat sizes in other PDAGs were not associated with risk of lifetime depression
The number of CAG repeats in ATXN1, ATXN2, ATXN3, CACNA1A, ATN1 and AR were not associated with the risk of lifetime depression. Neither the main effect of the alleles, nor the interactions between the two alleles, or the quadratic terms, were significantly associated with the presence of lifetime depression in the combined cohort. As AR is located on the X-chromosome, we performed one analysis stratifying the data by gender and another analysis using only either the relatively short or the relatively long allele in the model. None of the two approaches demonstrated a significant association between CAG repeat number in AR and depression (Supplementary Table 3).
The effects of HTT, ATXN7 and TBP CAG repeat sizes are independent
In previous research, we demonstrated that both the main term of the longer HTT allele and the quadratic term of the longer HTT allele are also significantly associated with a diagnosis of lifetime depression.15 To assess the contribution of the CAG repeat polymorphisms in all PDAGs to depression risk, we examined the degree to which HTT, ATXN7 and TBP CAG repeat sizes explain depression heritability. To this end, we applied a multivariate model with the CAG repeat sizes in these three PDAGs as predictors. In addition, we adjusted for the effects of gender, age and education. We found that the parameter estimates as well as their associated statistical significances hardly changed, indicating that the effects of HTT, ATXN7 and TBP CAG repeat sizes on depression susceptibility are mutually independent (HTT long allele: β=−0.286, P=0.018; HTT long allele quadratic term β=0.006, P=0.017; ATXN7 short allele: β=0.210, P=0.005; TBP short allele: β=−2.728, P=0.001; TBP long allele, β=−2.566; P=0.001; TBP interaction term: β=0.072, P=0.001). Inclusion of the CAG repeat sizes in HTT, ATXN7 and TBP in the model increased R2 by 0.014 from the baseline model including only gender, age and level of education (from 0.024 to 0.038), indicating that the CAG repeat lengths in these PDAGs can account for an additional 1.4% of the genetic variation on the observed probability scale. We also derived the R2 on the liability scale as described previously.40 Assuming that depression has a lifetime prevalence of ~15% in the Netherlands41 and adjusting for the oversampling of patients with depression in our cohort, the R2 on the liability scale was 0.0191, indicating that CAG repeat size polymorphisms in HTT, ATXN7 and TBP together can account for ~1.9% of depression heritability.
The prevalence of intermediate and pathological PGDA alleles
In total, four genotyped subjects had a CAG repeat number in the pathological range of a PDAG. One individual had a CAG repeat number of 22 in CACNA1A, one subject had a CAG repeat number of 48 in TBP and two people had a CAG repeat number of 33 and 36 in ATXN2 (Table 1). All four were depressed, but at the ages of 53 years for CACNA1A, 59 years for TBP, and 26 and 31 years for ATXN2, none had been diagnosed with the respective diseases (that is, SCA6, SCA17 and SCA2).
One person had a CAG repeat number in the longer TBP allele of 44, belonging to the intermediate range. This individual was also diagnosed with lifetime depression. Furthermore, one depressed individual had an intermediate CAG repeat number in the longer allele of ATXN3 (that is, 49 repeats). For ATXN1, ATXN7, ATN1 and AR all subjects had a CAG repeat number within the normal range (Table 1).
When we included the two depressed individuals with a CAG repeat length in the reduced penetrance range of HTT found previously,15 we observed a trend for a higher proportion of subjects with a CAG repeat in the intermediate or pathological range of a PDAG in the depressed group (P=0.059).
Discussion
To our knowledge, this is the first study assessing the influence of CAG repeat size variations in the normal range of PDAGs—other than HTT—on the risk of lifetime depression. Interestingly, we found an association between the presence of lifetime depression and the CAG repeat length in two PDAGs, that is, ATXN7 and TBP. The main relationship we found was that when the CAG repeat number of both alleles in either ATXN7 or TBP was relatively large, the odds for a diagnosis of lifetime depression markedly increased. Moreover, six genotyped individuals had a CAG repeat number in at least one of their PDAGs that extended into the intermediate or pathological range (two intermediate, four pathological). All of these subjects were diagnosed with lifetime depression. These findings are in support of the hypothesis that repeat polymorphisms may act as complex genetic modifiers of depression and thus could account for part of its ‘missing heritability’.
In our study, we found that the odds of lifetime depression almost doubled in individuals with a CAG repeat length of >10 in both ATXN7 alleles. Similarly, we found that the risk of lifetime depression markedly increased (OR=1.33) in individuals who had a CAG repeat length exceeding the median in both TBP alleles. These effect sizes are considerable, especially when compared with the effect sizes of the two most significant SNPs found in a recent GWAS: OR=0.955 (95% CI 0.943–0.968) and OR=1.051 (95% CI 1.036–1.067).6 Although we cannot fully exclude potential modifying effects of SNPs in linkage disequilibrium (LD) with ATXN7 and TBP, the fact that such SNPs in these PDAGs have not been detected before in GWASs suggests that the influence of other genetic variants in LD with the CAG repeat size in these genes is likely to be minimal.42 Furthermore, we found that CAG repeat size polymorphisms in HTT, ATXN7 and TBP together can account for ~1.9% of depression heritability. Using the same population prevalence of 15% for depression, the meta-analysis in the recent GWAS calculated a heritability of 5.9%.6 The amount of variance explained by these SNPs is larger than the amount explained by the CAG repeat polymorphisms we found. However, the meta-analysis in this GWAS examined SNPs in the entire genome of 326 113 individuals,6 whereas our estimation was based on the effects of CAG repeat variations in only nine PDAGs. This fact suggests that investigating additional repeat polymorphisms in the human genome could lead to the identification of many more novel genetic determinants of depression. The importance of investigating repeat polymorphisms in association with health and disease has been described in previous literature and begins to gain more recognition within the field of genetic research.9, 10, 43, 44, 45, 46
CAG repeat numbers exceeding 36 in ATXN7 are responsible for the severe neurodegenerative disorder spinocerebellar ataxia type 7 (SCA7).34 SCA7 is a progressive autosomal-dominant neurodegenerative disorder primarily characterized by cerebellar ataxia and macular degeneration.47 Although depressive symptoms are a frequent finding in many neurodegenerative disorders, the prevalence of depression has hardly been assessed in SCA7.48, 49, 50, 51 Our findings suggest that depression might be an underappreciated feature of SCA7, which needs further characterisation. A CAG repeat number in TBP larger than 48 causes the severe progressive neurodegenerative disorder SCA17.34 Aside from cerebellar ataxia, dementia and pyramidal symptoms, in 67% of the cases SCA17 is also accompanied by psychiatric signs and symptoms, including depression, behavioural changes as well as psychosis.52 Initial symptoms of SCA17 have been described to include depression.53
The gene ATXN7 encodes the protein ataxin-7. Apart from several important cellular functions,51 ataxin-7 is an integral part of the TATA-binding protein-free TAF-containing complex.54 TATA-binding protein-free TAF-containing complex allows for the initiation of transcription via RNA polymerase II in the absence of the RNA polymerase II transcription factor D.55 Interestingly, the TATA-box-binding protein (TBP) that is encoded by TBP, is the DNA-binding subunit of RNA polymerase II transcription factor D. TBP anchors RNA polymerase II transcription factor D to the TATA-box upstream of the first codon, allowing for the initiation of transcription via RNA polymerase II.56 Thus, TBP as well as ataxin-7 seem to have a part in the initiation of transcription via RNA polymerase II. RNA polymerase II catalyses the transcription of DNA to synthesise precursors of mRNA, snRNA and microRNA.57, 58 Numerous studies indicate an altered mRNA expression of proteins such as the serotonin 5-HT1A receptor,59, 60 brain-derived neurotropic factor61, 62, 63, 64 and corticotrophin-releasing factor65, 66 in subjects suffering from MDD compared with controls. Furthermore, the evidence supporting a role for the dysfunction of microRNA-mediated regulated gene expression in MDD is increasing.67, 68, 69, 70, 71 Therefore, CAG repeat variations in either ATXN7 or TBP could modulate the function of RNA polymerase II, and thereby lead to changes in mRNA and microRNA expression that have previously been associated with MDD.59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71 Hence, we intend to investigate the effect of CAG repeat length variations within the normal range in ATXN7 and TBP on the expression and function of proteins thought to be associated with depression in a cellular model.
The presence of one subject with a CAG repeat size in the pathological range of CACNA1A, one subject with a CAG repeat size in the pathological range of TBP and two subjects with a pathological CAG repeat size in ATXN2 within our two cohorts is remarkable. For these individuals, depression is apparently the first symptom with which SCA6, SCA17 or SCA2 manifests itself. The prevalence of an autosomal-dominant cerebral ataxia among Europeans is estimated to be between 1 and 3 per 100 000, of which 2% are diagnosed with SCA6, <1% with SCA17 and 10% with SCA2,72 resulting in a prevalence of 0.02–0.06 per 100 000, <0.01–0.03 per 10 000 and 0.10–0.30 per 100 000 for SCA6, SCA17 and SCA2, respectively. These numbers are in stark contrast to the substantially higher prevalence estimates in patients with depression in our study, 90 per 100 000 for SCA6, 50 per 100 000 for SCA17 and 50 per 100 000 for SCA2. Previously, we also found two individuals with incompletely penetrant HTT alleles (that is, those containing 36–39 CAG repeats) in this same population, both of whom suffered from depression.15 The difference in proportion of individuals with intermediate or pathological CAG repeat lengths in PDAGs indeed tended to be higher in depressed compared with control subjects, although likely due to the relative rarity of these expanded alleles the results did not reach statistical significance. Our findings are in line with another study that estimated the prevalence of incompletely penetrant HTT alleles in MDD patients to be ~3 in 1000, whereas such alleles were absent in the control group.73 Together, these findings suggest that depression could be the first manifestation of polyglutamine diseases and that, conversely, polyglutamine diseases might be underdiagnosed in patients with depression.
In this study, we analysed a homogenous population by using samples from two Dutch cohorts for our analysis, thereby minimising the impact of population stratification. However, the use of a uniform group of individuals could also be seen as a limitation of our study, as it might have consequences for the generalisability of our findings. However, our results appear robust and consistent, although confirmation of our findings in other populations, as well as the elucidation of their pathophysiological basis, is warranted.
In conclusion, we observed a significant association between the risk of lifetime depression and CAG repeat size in ATXN7 and TBP. A relatively large CAG repeat number in both alleles of either ATXN7 or TBP substantially increased depression risk. Our findings add more critical evidence to the notion that repeat polymorphisms could act as complex genetic modifiers of depression and, therefore, could partially account for its ‘missing heritability’. In addition, our findings indicate that the role of DNA repeat polymorphisms as potential genetic modifiers of other psychiatric disorders also needs further scrutiny.
Acknowledgments
This study was supported by a VENI-grant (#91615080) from the Netherlands Organization of Scientific Research and a Marie Sklodowska-Curie Individual Fellowship grant from the European Union (Horizon 2020, #701130; NAA). The infrastructure for the NESDA study (www.nesda.nl) has been funded through the Geestkracht programme of the Netherlands Organisation for Health Research and Development (Zon-Mw, grant number 10-000-1002) and participating universities (VU University Medical Center, Leiden University Medical Centre, University Medical Centre Groningen). The infrastructure for the NESDO study (http://nesdo.amstad.nl) is funded through the Fonds NutsOhra (project 0701-065), Stichting tot Steun VCVGZ, NARSAD The Brain and Behaviour Research Fund (grant ID 41080), and the participating universities and mental health-care organisations (VU University Medical Centre, Leiden University Medical Center, University Medical Centre Groningen, UMC St Radboud, and GGZ inGeest, GG Net, GGZ Nijmegen and Parnassia).
Footnotes
Supplementary Information accompanies the paper on the Translational Psychiatry website (http://www.nature.com/tp).
The authors declare no conflict of interest.
Supplementary Material
References
- Bromet E, Andrade LH, Hwang I, Sampson NA, Alonso J, de Girolamo G et al. Cross-national epidemiology of DSM-IV major depressive episode. BMC Med 2011; 9: 90. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Whiteford HA, Degenhardt L, Rehm J, Baxter AJ, Ferrari AJ, Erskine HE et al. Global burden of disease attributable to mental and substance use disorders: findings from the Global Burden of Disease Study 2010. Lancet 2013; 382: 1575–1586. [DOI] [PubMed] [Google Scholar]
- Greenberg PE, Fournier AA, Sisitsky T, Pike CT, Kessler RC. The economic burden of adults with major depressive disorder in the United States (2005 and 2010). J Clin Psychiatry 2015; 76: 155–162. [DOI] [PubMed] [Google Scholar]
- Major Depressive Disorder Working Group of the Psychiatric GC, Ripke S, Wray NR, Lewis CM, Hamilton SP, Weissman MM et al. A mega-analysis of genome-wide association studies for major depressive disorder. Mol Psychiatry 2013; 18: 497–511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Okbay A, Baselmans BM, De Neve JE, Turley P, Nivard MG, Fontana MA et al. Genetic variants associated with subjective well-being, depressive symptoms, and neuroticism identified through genome-wide analyses. Nat Genet 2016; 48: 624–633. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hyde CL, Nagle MW, Tian C, Chen X, Paciga SA, Wendland JR et al. Identification of 15 genetic loci associated with risk of major depression in individuals of European descent. Nat Genet 2016; 48: 1031–1036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Manolio TA. Genomewide association studies and assessment of the risk of disease. N Engl J Med 2010; 363: 166–176. [DOI] [PubMed] [Google Scholar]
- Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J et al. Initial sequencing and analysis of the human genome. Nature 2001; 409: 860–921. [DOI] [PubMed] [Google Scholar]
- Hannan AJ. Tandem repeat polymorphisms: modulators of disease susceptibility and candidates for 'missing heritability'. Trends Genet 2010; 26: 59–65. [DOI] [PubMed] [Google Scholar]
- Hannan AJ. TRPing up the genome: tandem repeat polymorphisms as dynamic sources of genetic variability in health and disease. Discov Med 2010; 10: 314–321. [PubMed] [Google Scholar]
- Mirkin SM. Expandable DNA repeats and human disease. Nature 2007; 447: 932–940. [DOI] [PubMed] [Google Scholar]
- Shao J, Diamond MI. Polyglutamine diseases: emerging concepts in pathogenesis and therapy. Hum Mol Genet 2007; 16 Spec No. 2: R115–R123. [DOI] [PubMed] [Google Scholar]
- Bates G, Harper P, Jones L. Huntington's Disease, 3rd edn. Oxford University Press: : New York, NY, USA, 2002. [Google Scholar]
- THsDCR G. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington's disease chromosomes. The Huntington's Disease Collaborative Research Group. Cell 1993; 72: 971–983. [DOI] [PubMed] [Google Scholar]
- Gardiner SL, van Belzen MJ, Boogaard MW, van Roon-Mom WMC, Rozing MP, Albert M et al Huntingtin gene repeat size variations affect risk of lifetime depression (submitted for publication). [DOI] [PMC free article] [PubMed]
- Orr HT, Chung MY, Banfi S, Kwiatkowski TJ Jr., Servadio A, Beaudet AL et al. Expansion of an unstable trinucleotide CAG repeat in spinocerebellar ataxia type 1. Nat Genet 1993; 4: 221–226. [DOI] [PubMed] [Google Scholar]
- Chung MY, Ranum LP, Duvick LA, Servadio A, Zoghbi HY, Orr HT. Evidence for a mechanism predisposing to intergenerational CAG repeat instability in spinocerebellar ataxia type I. Nat Genet 1993; 5: 254–258. [DOI] [PubMed] [Google Scholar]
- Matilla T, Volpini V, Genis D, Rosell J, Corral J, Davalos A et al. Presymptomatic analysis of spinocerebellar ataxia type 1 (SCA1) via the expansion of the SCA1 CAG-repeat in a large pedigree displaying anticipation and parental male bias. Hum Mol Genet 1993; 2: 2123–2128. [DOI] [PubMed] [Google Scholar]
- Pulst SM, Nechiporuk A, Nechiporuk T, Gispert S, Chen XN, Lopes-Cendes I et al. Moderate expansion of a normally biallelic trinucleotide repeat in spinocerebellar ataxia type 2. Nat Genet 1996; 14: 269–276. [DOI] [PubMed] [Google Scholar]
- Sanpei K, Takano H, Igarashi S, Sato T, Oyake M, Sasaki H et al. Identification of the spinocerebellar ataxia type 2 gene using a direct identification of repeat expansion and cloning technique, DIRECT. Nat Genet 1996; 14: 277–284. [DOI] [PubMed] [Google Scholar]
- Giunti P, Sweeney MG, Harding AE. Detection of the Machado-Joseph disease/spinocerebellar ataxia three trinucleotide repeat expansion in families with autosomal dominant motor disorders, including the Drew family of Walworth. Brain 1995; 118(Pt 5): 1077–1085. [DOI] [PubMed] [Google Scholar]
- Maciel P, Gaspar C, DeStefano AL, Silveira I, Coutinho P, Radvany J et al. Correlation between CAG repeat length and clinical features in Machado-Joseph disease. Am J Hum Genet 1995; 57: 54–61. [PMC free article] [PubMed] [Google Scholar]
- Maruyama H, Nakamura S, Matsuyama Z, Sakai T, Doyu M, Sobue G et al. Molecular features of the CAG repeats and clinical manifestation of Machado-Joseph disease. Hum Mol Genet 1995; 4: 807–812. [DOI] [PubMed] [Google Scholar]
- Zhuchenko O, Bailey J, Bonnen P, Ashizawa T, Stockton DW, Amos C et al. Autosomal dominant cerebellar ataxia (SCA6) associated with small polyglutamine expansions in the alpha 1A-voltage-dependent calcium channel. Nat Genet 1997; 15: 62–69. [DOI] [PubMed] [Google Scholar]
- Matsuyama Z, Kawakami H, Maruyama H, Izumi Y, Komure O, Udaka F et al. Molecular features of the CAG repeats of spinocerebellar ataxia 6 (SCA6). Hum Mol Genet 1997; 6: 1283–1287. [DOI] [PubMed] [Google Scholar]
- David G, Abbas N, Stevanin G, Durr A, Yvert G, Cancel G et al. Cloning of the SCA7 gene reveals a highly unstable CAG repeat expansion. Nat Genet 1997; 17: 65–70. [DOI] [PubMed] [Google Scholar]
- Benton CS, de Silva R, Rutledge SL, Bohlega S, Ashizawa T, Zoghbi HY. Molecular and clinical studies in SCA-7 define a broad clinical spectrum and the infantile phenotype. Neurology 1998; 51: 1081–1086. [DOI] [PubMed] [Google Scholar]
- Del-Favero J, Krols L, Michalik A, Theuns J, Lofgren A, Goossens D et al. Molecular genetic analysis of autosomal dominant cerebellar ataxia with retinal degeneration (ADCA type II) caused by CAG triplet repeat expansion. Hum Mol Genet 1998; 7: 177–186. [DOI] [PubMed] [Google Scholar]
- Johansson J, Forsgren L, Sandgren O, Brice A, Holmgren G, Holmberg M. Expanded CAG repeats in Swedish spinocerebellar ataxia type 7 (SCA7) patients: effect of CAG repeat length on the clinical manifestation. Hum Mol Genet 1998; 7: 171–176. [DOI] [PubMed] [Google Scholar]
- Nakamura K, Jeong SY, Uchihara T, Anno M, Nagashima K, Nagashima T et al. SCA17, a novel autosomal dominant cerebellar ataxia caused by an expanded polyglutamine in TATA-binding protein. Hum Mol Genet 2001; 10: 1441–1448. [DOI] [PubMed] [Google Scholar]
- Koide R, Kobayashi S, Shimohata T, Ikeuchi T, Maruyama M, Saito M et al. A neurological disease caused by an expanded CAG trinucleotide repeat in the TATA-binding protein gene: a new polyglutamine disease? Hum Mol Genet 1999; 8: 2047–2053. [DOI] [PubMed] [Google Scholar]
- Nagafuchi S, Yanagisawa H, Sato K, Shirayama T, Ohsaki E, Bundo M et al. Dentatorubral and pallidoluysian atrophy expansion of an unstable CAG trinucleotide on chromosome 12p. Nat Genet 1994; 6: 14–18. [DOI] [PubMed] [Google Scholar]
- La Spada AR, Wilson EM, Lubahn DB, Harding AE, Fischbeck KH. Androgen receptor gene mutations in X-linked spinal and bulbar muscular atrophy. Nature 1991; 352: 77–79. [DOI] [PubMed] [Google Scholar]
- Bettencourt C, Hensman-Moss D, Flower M, Wiethoff S, Brice A, Goizet C et al. DNA repair pathways underlie a common genetic mechanism modulating onset in polyglutamine diseases. Ann Neurol 2016; 79: 983–990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fan HC, Ho LI, Chi CS, Chen SJ, Peng GS, Chan TM et al. Polyglutamine (PolyQ) diseases: genetics to treatments. Cell Transplant 2014; 23: 441–458. [DOI] [PubMed] [Google Scholar]
- Penninx BW, Beekman AT, Smit JH, Zitman FG, Nolen WA, Spinhoven P et al. The Netherlands Study of Depression and Anxiety (NESDA): rationale, objectives and methods. Int J Methods Psychiatr Res 2008; 17: 121–140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wittchen HU, Robins LN, Cottler LB, Sartorius N, Burke JD, Regier D. Cross-cultural feasibility, reliability and sources of variance of the Composite International Diagnostic Interview (CIDI). The Multicentre WHO/ADAMHA Field Trials. Br J Psychiatry 1991; 159: 645–653, 658. [DOI] [PubMed] [Google Scholar]
- Comijs HC, van Marwijk HW, van der Mast RC, Naarding P, Oude Voshaar RC, Beekman AT et al. The Netherlands study of depression in older persons (NESDO); a prospective cohort study. BMC Res Notes 2011; 4: 524. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benjamini Y, Hochberg Y. Controling the false discovery rate: a practical and powerful approach to multiple testing. J R StatSoc Ser B 1995; 57: 289–300. [Google Scholar]
- Lee SH, Goddard ME, Wray NR, Visscher PM. A better coefficient of determination for genetic profile analysis. Genet Epidemiol 2012; 36: 214–224. [DOI] [PubMed] [Google Scholar]
- Andrade L, Caraveo-Anduaga JJ, Berglund P, Bijl RV, De Graaf R, Vollebergh W et al. The epidemiology of major depressive episodes: results from the International Consortium of Psychiatric Epidemiology (ICPE) Surveys. Int J Method Psychiatr Res 2003; 12: 3–21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Okbay A, Baselmans BM, De Neve JE, Turley P, Nivard MG. Genetic variants associated with subjective well-being, depressive symptoms, and neuroticism identified through genome-wide analyses. Nat Genet 2016; 48: 624–633. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kashi Y, King D, Soller M. Simple sequence repeats as a source of quantitative genetic variation. Trends Genet 1997; 13: 74–78. [DOI] [PubMed] [Google Scholar]
- Kashi Y, King DG. Simple sequence repeats as advantageous mutators in evolution. Trends Genet 2006; 22: 253–259. [DOI] [PubMed] [Google Scholar]
- Sawaya S, Bagshaw A, Buschiazzo E, Kumar P, Chowdhury S, Black MA et al. Microsatellite tandem repeats are abundant in human promoters and are associated with regulatory elements. PLoS ONE 2013; 8: e54710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sawaya SM, Lennon D, Buschiazzo E, Gemmell N, Minin VN. Measuring microsatellite conservation in mammalian evolution with a phylogenetic birth-death model. Genome Biol Evol 2012; 4: 636–647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lebre AS, Brice A. Spinocerebellar ataxia 7 (SCA7). Cytogenet Genome Res 2003; 100: 154–163. [DOI] [PubMed] [Google Scholar]
- Pla P, Orvoen S, Saudou F, David DJ, Humbert S. Mood disorders in Huntington's disease: from behavior to cellular and molecular mechanisms. Front Behav Neurosci 2014; 8: 135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chaudhuri KR, Schapira AH. Non-motor symptoms of Parkinson's disease: dopaminergic pathophysiology and treatment. Lancet Neurol 2009; 8: 464–474. [DOI] [PubMed] [Google Scholar]
- Schmitz-Hubsch T, Coudert M, Tezenas du Montcel S, Giunti P, Labrum R, Durr A et al. Depression comorbidity in spinocerebellar ataxia. Move Disorders 2011; 26: 870–876. [DOI] [PubMed] [Google Scholar]
- Rub U, Schols L, Paulson H, Auburger G, Kermer P, Jen JC et al. Clinical features, neurogenetics and neuropathology of the polyglutamine spinocerebellar ataxias type 1, 2, 3, 6 and 7. Progr Neurobiol 2013; 104: 38–66. [DOI] [PubMed] [Google Scholar]
- Stevanin G, Brice A. Spinocerebellar ataxia 17 (SCA17) and Huntington's disease-like 4 (HDL4). Cerebellum 2008; 7: 170–178. [DOI] [PubMed] [Google Scholar]
- Bauer P, Laccone F, Rolfs A, Wullner U, Bosch S, Peters H et al. Trinucleotide repeat expansion in SCA17/TBP in white patients with Huntington's disease-like phenotype. J Med Genet 2004; 41: 230–232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Helmlinger D, Hardy S, Sasorith S, Klein F, Robert F, Weber C et al. Ataxin-7 is a subunit of GCN5 histone acetyltransferase-containing complexes. Hum Mol Genet 2004; 13: 1257–1265. [DOI] [PubMed] [Google Scholar]
- Wieczorek E, Brand M, Jacq X, Tora L. Function of TAF(II)-containing complex without TBP in transcription by RNA polymerase II. Nature 1998; 393: 187–191. [DOI] [PubMed] [Google Scholar]
- Lescure A, Lutz Y, Eberhard D, Jacq X, Krol A, Grummt I et al. The N-terminal domain of the human TATA-binding protein plays a role in transcription from TATA-containing RNA polymerase II and III promoters. EMBO J 1994; 13: 1166–1175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kornberg RD. Eukaryotic transcriptional control. Trends Cell Biol 1999; 9: M46–M49. [PubMed] [Google Scholar]
- Sims RJ 3rd, Mandal SS, Reinberg D. Recent highlights of RNA-polymerase-II-mediated transcription. Curr Opin Cell Biol 2004; 16: 263–271. [DOI] [PubMed] [Google Scholar]
- Lopez-Figueroa AL, Norton CS, Lopez-Figueroa MO, Armellini-Dodel D, Burke S, Akil H et al. Serotonin 5-HT1A, 5-HT1B, and 5-HT2A receptor mRNA expression in subjects with major depression, bipolar disorder, and schizophrenia. Biol Psychiatry 2004; 55: 225–233. [DOI] [PubMed] [Google Scholar]
- Lopez JF, Chalmers DT, Little KY, Watson SJ. A.E. Bennett Research Award. Regulation of serotonin1A, glucocorticoid, and mineralocorticoid receptor in rat and human hippocampus: implications for the neurobiology of depression. Biol Psychiatry 1998; 43: 547–573. [DOI] [PubMed] [Google Scholar]
- Cattaneo A, Gennarelli M, Uher R, Breen G, Farmer A, Aitchison KJ et al. Candidate genes expression profile associated with antidepressants response in the GENDEP study: differentiating between baseline 'predictors' and longitudinal 'targets'. Neuropsychopharmacology 2013; 38: 377–385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- D'Addario C, Dell'Osso B, Galimberti D, Palazzo MC, Benatti B, Di Francesco A et al. Epigenetic modulation of BDNF gene in patients with major depressive disorder. Biol Psychiatry 2013; 73: e6–e7. [DOI] [PubMed] [Google Scholar]
- Pandey GN, Dwivedi Y, Rizavi HS, Ren X, Zhang H, Pavuluri MN. Brain-derived neurotrophic factor gene and protein expression in pediatric and adult depressed subjects. Progr Neuro-psychopharmacol Biol Pychiatry 2010; 34: 645–651. [DOI] [PubMed] [Google Scholar]
- Zhou L, Xiong J, Lim Y, Ruan Y, Huang C, Zhu Y et al. Upregulation of blood proBDNF and its receptors in major depression. J Affect Disord 2013; 150: 776–784. [DOI] [PubMed] [Google Scholar]
- Austin MC, Janosky JE, Murphy HA. Increased corticotropin-releasing hormone immunoreactivity in monoamine-containing pontine nuclei of depressed suicide men. Mol Psychiatry 2003; 8: 324–332. [DOI] [PubMed] [Google Scholar]
- Raadsheer FC, van Heerikhuize JJ, Lucassen PJ, Hoogendijk WJ, Tilders FJ, Swaab DF. Corticotropin-releasing hormone mRNA levels in the paraventricular nucleus of patients with Alzheimer's disease and depression. Am J Psychiatry 1995; 152: 1372–1376. [DOI] [PubMed] [Google Scholar]
- Geaghan M, Cairns MJ. MicroRNA and posttranscriptional dysregulation in psychiatry. Biol Psychiatry 2015; 78: 231–239. [DOI] [PubMed] [Google Scholar]
- Hommers LG, Domschke K, Deckert J. Heterogeneity and individuality: microRNAs in mental disorders. J Neural Trans 2015; 122: 79–97. [DOI] [PubMed] [Google Scholar]
- Hunsberger JG, Austin DR, Chen G, Manji HK. MicroRNAs in mental health: from biological underpinnings to potential therapies. Neuromol Med 2009; 11: 173–182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Issler O, Chen A. Determining the role of microRNAs in psychiatric disorders. Nat Rev Neurosci 2015; 16: 201–212. [DOI] [PubMed] [Google Scholar]
- O'Connor RM, Grenham S, Dinan TG, Cryan JF. microRNAs as novel antidepressant targets: converging effects of ketamine and electroconvulsive shock therapy in the rat hippocampus. Int J Neuropsychopharmacol 2013; 16: 1885–1892. [DOI] [PubMed] [Google Scholar]
- Durr A. Autosomal dominant cerebellar ataxias: polyglutamine expansions and beyond. Lancet Neurol 2010; 9: 885–894. [DOI] [PubMed] [Google Scholar]
- Perlis RH, Smoller JW, Mysore J, Sun M, Gillis T, Purcell S et al. Prevalence of incompletely penetrant Huntington's disease alleles among individuals with major depressive disorder. Am J Psychiatry 2010; 167: 574–579. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.