Abstract
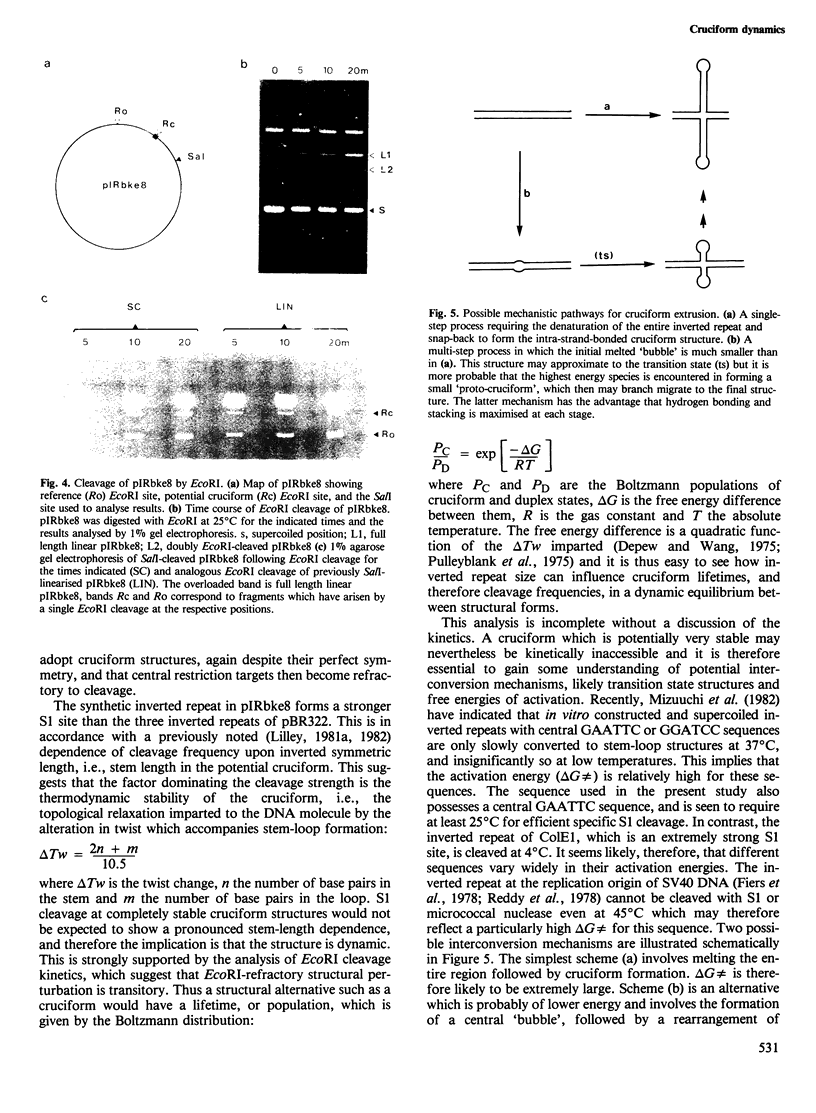
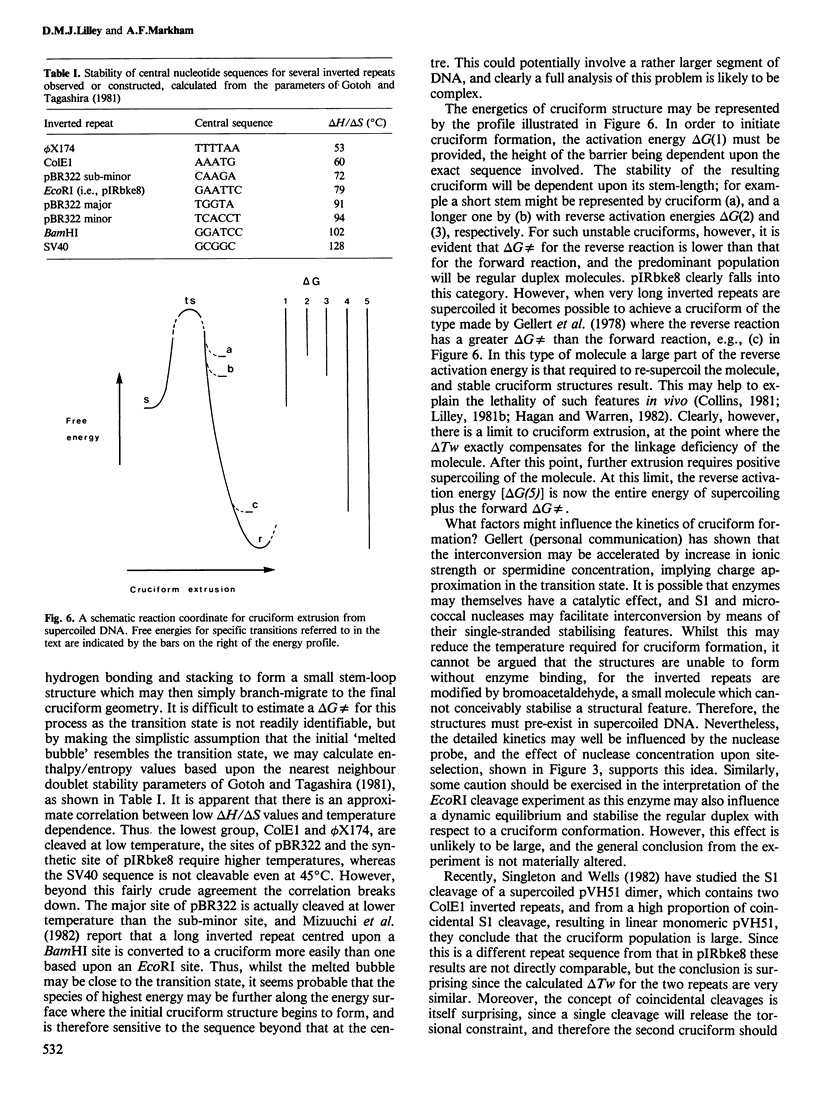
An inverted repeat has been created in a plasmid by ligation of two 13 nucleotide synthetic oligonucleotides into the cloning vector pAT153. The resulting recombinant plasmid, pIRbke8, is hypersensitive to cleavage by the single-strand-specific nuclease S1, and to modification by the single-strand-selective reagent bromoacetaldehyde, when the plasmid is negatively supercoiled. The new inverted repeat is a stronger S1 site than those derived from pBR322, but, in contrast to the ColE1 and phi X174 RF inverted repeats, these repeats share a similar temperature dependence. The kinetics of EcoRI cleavage at the centre of the synthetic inverted repeat have been studied in supercoiled and linear molecules. It is found that in the supercoiled molecule this target is not refractory to EcoRI cleavage to an extent which is greater than the resolution of the experiment. We conclude that in this molecule the cruciform is in a dynamic equilibrium with the regular duplex, in which the cruciform constitutes a relatively small subpopulation of conformational species.
Full text
PDF


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bauer W., Vinograd J. Interaction of closed circular DNA with intercalative dyes. II. The free energy of superhelix formation in SV40 DNA. J Mol Biol. 1970 Feb 14;47(3):419–435. doi: 10.1016/0022-2836(70)90312-8. [DOI] [PubMed] [Google Scholar]
- Benham C. J. Stable cruciform formation at inverted repeat sequences in supercoiled DNA. Biopolymers. 1982 Mar;21(3):679–696. doi: 10.1002/bip.360210314. [DOI] [PubMed] [Google Scholar]
- Betz J. L., Sadler J. R. Variants of a cloned synthetic lactose operator. I. A palindromic dimer lactose operator derived from one stand of the cloned 40-base pair operator. Gene. 1981 Jan-Feb;13(1):1–12. doi: 10.1016/0378-1119(81)90038-x. [DOI] [PubMed] [Google Scholar]
- Birnboim H. C., Doly J. A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res. 1979 Nov 24;7(6):1513–1523. doi: 10.1093/nar/7.6.1513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boyer H. W., Roulland-Dussoix D. A complementation analysis of the restriction and modification of DNA in Escherichia coli. J Mol Biol. 1969 May 14;41(3):459–472. doi: 10.1016/0022-2836(69)90288-5. [DOI] [PubMed] [Google Scholar]
- Cohen S. N., Chang A. C., Hsu L. Nonchromosomal antibiotic resistance in bacteria: genetic transformation of Escherichia coli by R-factor DNA. Proc Natl Acad Sci U S A. 1972 Aug;69(8):2110–2114. doi: 10.1073/pnas.69.8.2110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Collins J. Instability of palindromic DNA in Escherichia coli. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 1):409–416. doi: 10.1101/sqb.1981.045.01.055. [DOI] [PubMed] [Google Scholar]
- Depew D. E., Wang J. C. Conformational fluctuations of DNA helix. Proc Natl Acad Sci U S A. 1975 Nov;72(11):4275–4279. doi: 10.1073/pnas.72.11.4275. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Edge M. D., Green A. R., Heathcliffe G. R., Meacock P. A., Schuch W., Scanlon D. B., Atkinson T. C., Newton C. R., Markham A. F. Total synthesis of a human leukocyte interferon gene. Nature. 1981 Aug 20;292(5825):756–762. doi: 10.1038/292756a0. [DOI] [PubMed] [Google Scholar]
- Fiers W., Contreras R., Haegemann G., Rogiers R., Van de Voorde A., Van Heuverswyn H., Van Herreweghe J., Volckaert G., Ysebaert M. Complete nucleotide sequence of SV40 DNA. Nature. 1978 May 11;273(5658):113–120. doi: 10.1038/273113a0. [DOI] [PubMed] [Google Scholar]
- Fuller F. B. The writhing number of a space curve. Proc Natl Acad Sci U S A. 1971 Apr;68(4):815–819. doi: 10.1073/pnas.68.4.815. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gellert M., Mizuuchi K., O'Dea M. H., Ohmori H., Tomizawa J. DNA gyrase and DNA supercoiling. Cold Spring Harb Symp Quant Biol. 1979;43(Pt 1):35–40. doi: 10.1101/sqb.1979.043.01.007. [DOI] [PubMed] [Google Scholar]
- Hagan C. E., Warren G. J. Lethality of palindromic DNA and its use in selection of recombinant plasmids. Gene. 1982 Jul-Aug;19(1):147–151. doi: 10.1016/0378-1119(82)90199-8. [DOI] [PubMed] [Google Scholar]
- Hsieh T. S., Wang J. C. Thermodynamic properties of superhelical DNAs. Biochemistry. 1975 Feb 11;14(3):527–535. doi: 10.1021/bi00674a011. [DOI] [PubMed] [Google Scholar]
- Lilley D. M. Hairpin-loop formation by inverted repeats in supercoiled DNA is a local and transmissible property. Nucleic Acids Res. 1981 Mar 25;9(6):1271–1289. doi: 10.1093/nar/9.6.1271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lilley D. M. In vivo consequences of plasmid topology. Nature. 1981 Jul 23;292(5821):380–382. doi: 10.1038/292380a0. [DOI] [PubMed] [Google Scholar]
- Lilley D. M. The inverted repeat as a recognizable structural feature in supercoiled DNA molecules. Proc Natl Acad Sci U S A. 1980 Nov;77(11):6468–6472. doi: 10.1073/pnas.77.11.6468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. A new method for sequencing DNA. Proc Natl Acad Sci U S A. 1977 Feb;74(2):560–564. doi: 10.1073/pnas.74.2.560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mizuuchi K., Mizuuchi M., Gellert M. Cruciform structures in palindromic DNA are favored by DNA supercoiling. J Mol Biol. 1982 Apr 5;156(2):229–243. doi: 10.1016/0022-2836(82)90325-4. [DOI] [PubMed] [Google Scholar]
- Mukai F. H., Margolin P. ANALYSIS OF UNLINKED SUPPRESSORS OF AN O degrees MUTATION IN SALMONELLA. Proc Natl Acad Sci U S A. 1963 Jul;50(1):140–148. doi: 10.1073/pnas.50.1.140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Panayotatos N., Wells R. D. Cruciform structures in supercoiled DNA. Nature. 1981 Feb 5;289(5797):466–470. doi: 10.1038/289466a0. [DOI] [PubMed] [Google Scholar]
- Peck L. J., Nordheim A., Rich A., Wang J. C. Flipping of cloned d(pCpG)n.d(pCpG)n DNA sequences from right- to left-handed helical structure by salt, Co(III), or negative supercoiling. Proc Natl Acad Sci U S A. 1982 Aug;79(15):4560–4564. doi: 10.1073/pnas.79.15.4560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Platt J. R. POSSIBLE SEPARATION OF INTERTWINED NUCLEIC ACID CHAINS BY TRANSFER-TWIST. Proc Natl Acad Sci U S A. 1955 Mar 15;41(3):181–183. doi: 10.1073/pnas.41.3.181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pulleyblank D. E., Shure M., Tang D., Vinograd J., Vosberg H. P. Action of nicking-closing enzyme on supercoiled and nonsupercoiled closed circular DNA: formation of a Boltzmann distribution of topological isomers. Proc Natl Acad Sci U S A. 1975 Nov;72(11):4280–4284. doi: 10.1073/pnas.72.11.4280. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reddy V. B., Thimmappaya B., Dhar R., Subramanian K. N., Zain B. S., Pan J., Ghosh P. K., Celma M. L., Weissman S. M. The genome of simian virus 40. Science. 1978 May 5;200(4341):494–502. doi: 10.1126/science.205947. [DOI] [PubMed] [Google Scholar]
- Richardson J. P. Effects of supercoiling on transcription from bacteriophage PM2 deoxyribonucleic acid. Biochemistry. 1974 Jul 16;13(15):3164–3169. doi: 10.1021/bi00712a025. [DOI] [PubMed] [Google Scholar]
- Sanzey B. Modulation of gene expression by drugs affecting deoxyribonucleic acid gyrase. J Bacteriol. 1979 Apr;138(1):40–47. doi: 10.1128/jb.138.1.40-47.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Secrist J. A., 3rd, Barrio J. R., Leonard N. J., Villar-Palasi C., Gilman A. G. Fluorescent modification of adenosine 3',5'-monophosphate: spectroscopic properties and activity in enzyme systems. Science. 1972 Jul 21;177(4045):279–280. doi: 10.1126/science.177.4045.279. [DOI] [PubMed] [Google Scholar]
- Sherratt D. J., Dougan G., Saul M., Sunar B., Twigg A., Warren G. Structure and function of plasmid ColE1 and related plasmids. Contrib Microbiol Immunol. 1979;6:100–110. [PubMed] [Google Scholar]
- Singleton C. K., Klysik J., Stirdivant S. M., Wells R. D. Left-handed Z-DNA is induced by supercoiling in physiological ionic conditions. Nature. 1982 Sep 23;299(5881):312–316. doi: 10.1038/299312a0. [DOI] [PubMed] [Google Scholar]
- Singleton C. K., Wells R. D. Relationship between superhelical density and cruciform formation in plasmid pVH51. J Biol Chem. 1982 Jun 10;257(11):6292–6295. [PubMed] [Google Scholar]
- Sutcliffe J. G. Complete nucleotide sequence of the Escherichia coli plasmid pBR322. Cold Spring Harb Symp Quant Biol. 1979;43(Pt 1):77–90. doi: 10.1101/sqb.1979.043.01.013. [DOI] [PubMed] [Google Scholar]
- Trucksis M., Depew R. E. Identification and localization of a gene that specifies production of Escherichia coli DNA topoisomerase I. Proc Natl Acad Sci U S A. 1981 Apr;78(4):2164–2168. doi: 10.1073/pnas.78.4.2164. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Twigg A. J., Sherratt D. Trans-complementable copy-number mutants of plasmid ColE1. Nature. 1980 Jan 10;283(5743):216–218. doi: 10.1038/283216a0. [DOI] [PubMed] [Google Scholar]
- Vinograd J., Lebowitz J., Radloff R., Watson R., Laipis P. The twisted circular form of polyoma viral DNA. Proc Natl Acad Sci U S A. 1965 May;53(5):1104–1111. doi: 10.1073/pnas.53.5.1104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vinograd J., Lebowitz J., Watson R. Early and late helix-coil transitions in closed circular DNA. The number of superhelical turns in polyoma DNA. J Mol Biol. 1968 Apr 14;33(1):173–197. doi: 10.1016/0022-2836(68)90287-8. [DOI] [PubMed] [Google Scholar]
- Vologodskii A. V., Frank-Kamenetskii M. D. Theoretical study of cruciform states in superhelical DNAs. FEBS Lett. 1982 Jul 5;143(2):257–260. doi: 10.1016/0014-5793(82)80111-7. [DOI] [PubMed] [Google Scholar]
- Wang A. H., Quigley G. J., Kolpak F. J., Crawford J. L., van Boom J. H., van der Marel G., Rich A. Molecular structure of a left-handed double helical DNA fragment at atomic resolution. Nature. 1979 Dec 13;282(5740):680–686. doi: 10.1038/282680a0. [DOI] [PubMed] [Google Scholar]
- Wang J. C. Interactions between twisted DNAs and enzymes: the effects of superhelical turns. J Mol Biol. 1974 Aug 25;87(4):797–816. doi: 10.1016/0022-2836(74)90085-0. [DOI] [PubMed] [Google Scholar]
- Yang H. L., Heller K., Gellert M., Zubay G. Differential sensitivity of gene expression in vitro to inhibitors of DNA gyrase. Proc Natl Acad Sci U S A. 1979 Jul;76(7):3304–3308. doi: 10.1073/pnas.76.7.3304. [DOI] [PMC free article] [PubMed] [Google Scholar]