Abstract
Two so far undetected tRNA genes were found close to delta (δ) sequences at the sup4 locus on chromosome X in the genome of Saccharomyces cerevisiae. The two genes were identified from their abundant transcription products in frog oocytes. Hybridisation experiments allowed the mapping of the transcripts in cloned DNA and DNA sequence analysis revealed the presence of one AGGtRNAArg and one GACtRNAAsp gene. tRNAAsp genes with sequences similar or identical to GACtRNAAsp exist in 14-16 copies per haploid yeast genome, whereas only one copy was detected for AGGtRNAArg. In vivo labelling of total yeast tRNA with 32P followed by hybridisation revealed that the unique AGGtRNAArg gene is transcribed in S. cerevisiae. δ sequences are present 120 bp upstream from the first coding nucleotide in the case of AGGtRNAArg, 80 bp in the case of GACtRNAAsp and 405 bp in the case of the known UACtRNATyr (sup4) gene. δ sequences, as part of Ty elements or alone, were also found by other investigators at similar distances upstream of the mRNA start in mutant alleles of protein-coding yeast genes. Although protein-coding genes are transcribed by RNA polymerase II and tRNA genes by RNA polymerase III, the 5' non-coding region of both types of genes could conceivably have a peculiar DNA or chromatin structure used as preferred landing sites by transposable elements.
Keywords: δ sequences, frog oocyte injection, transposition, tRNA genes, yeast
Full text
PDF



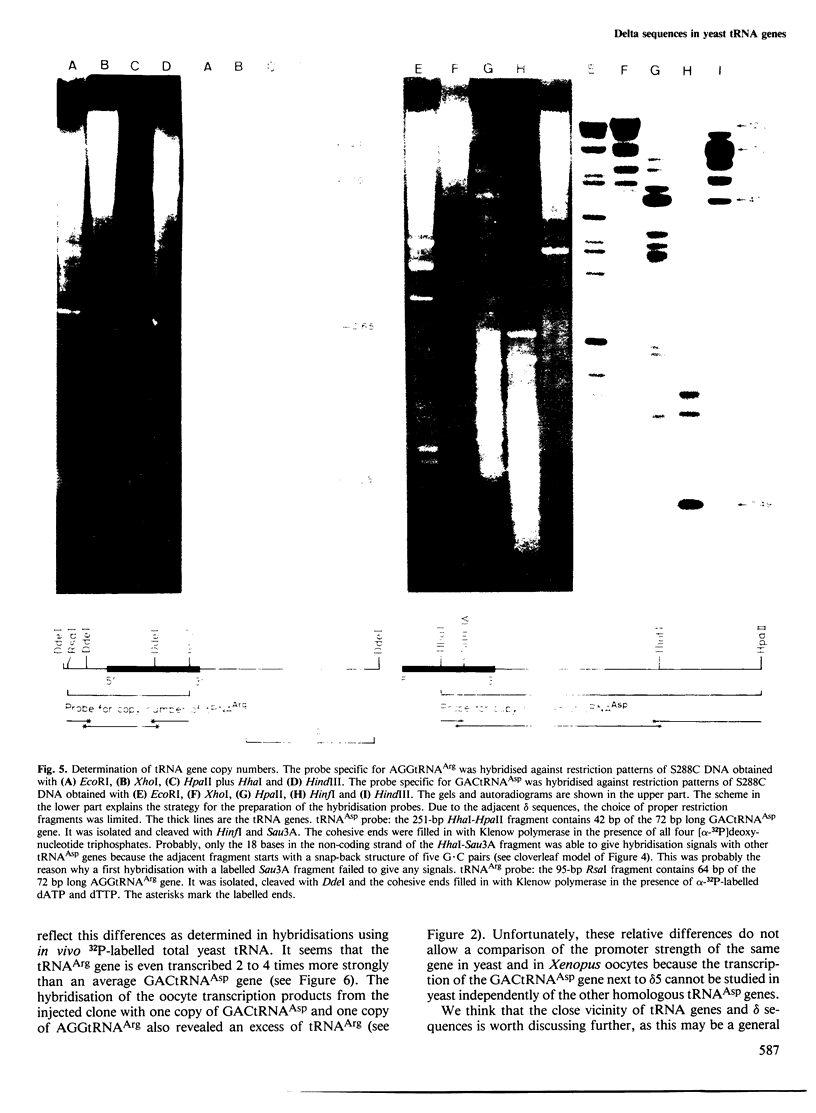
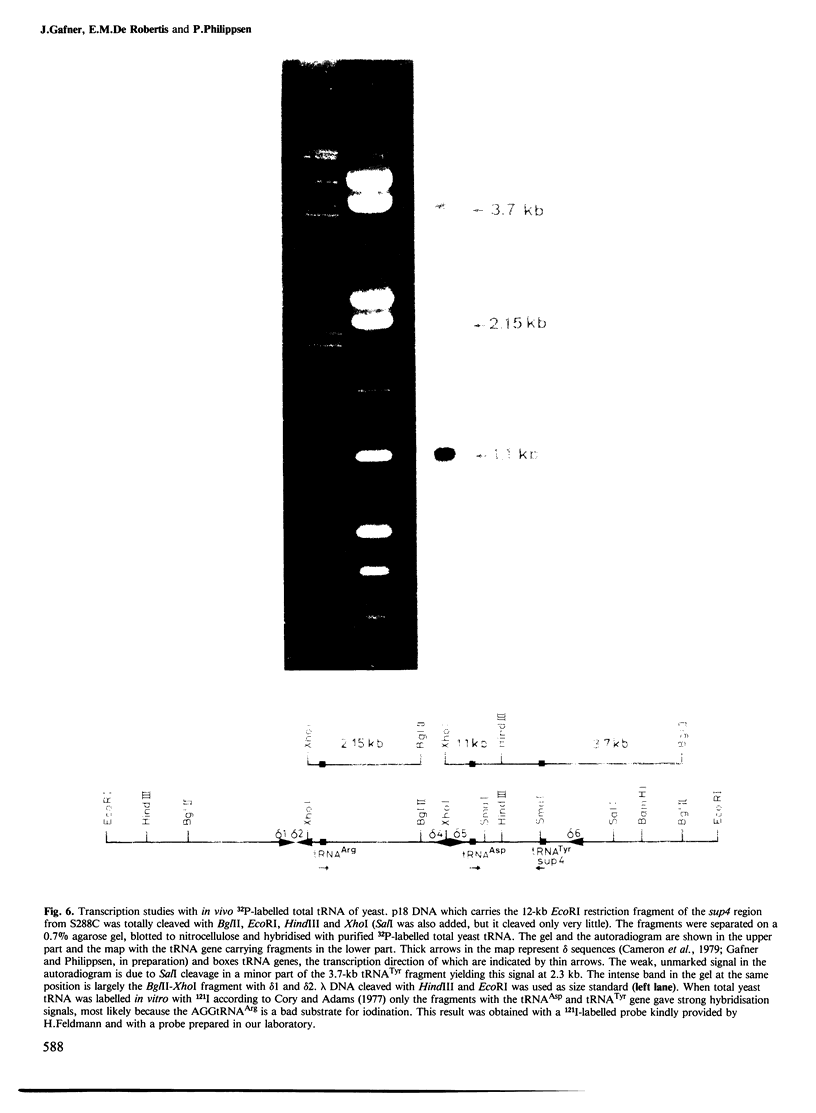



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Andreadis A., Hsu Y. P., Kohlhaw G. B., Schimmel P. Nucleotide sequence of yeast LEU2 shows 5'-noncoding region has sequences cognate to leucine. Cell. 1982 Dec;31(2 Pt 1):319–325. doi: 10.1016/0092-8674(82)90125-8. [DOI] [PubMed] [Google Scholar]
- Beggs J. D. Transformation of yeast by a replicating hybrid plasmid. Nature. 1978 Sep 14;275(5676):104–109. doi: 10.1038/275104a0. [DOI] [PubMed] [Google Scholar]
- Bennetzen J. L., Hall B. D. Codon selection in yeast. J Biol Chem. 1982 Mar 25;257(6):3026–3031. [PubMed] [Google Scholar]
- Brown R. D., Brown D. D. The nucleotide sequence adjoining the 3' end of the genes coding for oocyte-type 5 S ribosomal RNA in Xenopus. J Mol Biol. 1976 Mar 25;102(1):1–14. doi: 10.1016/0022-2836(76)90070-x. [DOI] [PubMed] [Google Scholar]
- Cameron J. R., Loh E. Y., Davis R. W. Evidence for transposition of dispersed repetitive DNA families in yeast. Cell. 1979 Apr;16(4):739–751. doi: 10.1016/0092-8674(79)90090-4. [DOI] [PubMed] [Google Scholar]
- Chaleff D. T., Fink G. R. Genetic events associated with an insertion mutation in yeast. Cell. 1980 Aug;21(1):227–237. doi: 10.1016/0092-8674(80)90130-0. [DOI] [PubMed] [Google Scholar]
- Cory S., Adams J. M. A very large repeating unit of mouse DNA containing the 18S, 28S and 5.8S rRNA genes. Cell. 1977 Aug;11(4):795–805. doi: 10.1016/0092-8674(77)90292-6. [DOI] [PubMed] [Google Scholar]
- De Robertis E. M., Olson M. V. Transcription and processing of cloned yeast tyrosine tRNA genes microinjected into frog oocytes. Nature. 1979 Mar 8;278(5700):137–143. doi: 10.1038/278137a0. [DOI] [PubMed] [Google Scholar]
- DeFranco D., Schmidt O., Söll D. Two control regions for eukaryotic tRNA gene transcription. Proc Natl Acad Sci U S A. 1980 Jun;77(6):3365–3368. doi: 10.1073/pnas.77.6.3365. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eibel H., Gafner J., Stotz A., Philippsen P. Characterization of the yeast mobile element Ty1. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 2):609–617. doi: 10.1101/sqb.1981.045.01.079. [DOI] [PubMed] [Google Scholar]
- Eigel A., Feldmann H. Ty1 and delta elements occur adjacent to several tRNA genes in yeast. EMBO J. 1982;1(10):1245–1250. doi: 10.1002/j.1460-2075.1982.tb00020.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elder R. T., St John T. P., Stinchcomb D. T., Davis R. W., Scherer S., Davis R. W. Studies on the transposable element Ty1 of yeast. I. RNA homologous to Ty1. II. Recombination and expression of Ty1 and adjacent sequences. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 2):581–591. doi: 10.1101/sqb.1981.045.01.075. [DOI] [PubMed] [Google Scholar]
- Errede B., Cardillo T. S., Sherman F., Dubois E., Deschamps J., Wiame J. M. Mating signals control expression of mutations resulting from insertion of a transposable repetitive element adjacent to diverse yeast genes. Cell. 1980 Nov;22(2 Pt 2):427–436. doi: 10.1016/0092-8674(80)90353-0. [DOI] [PubMed] [Google Scholar]
- Errede B., Cardillo T. S., Wever G., Sherman F., Stiles J. I., Friedman L. R., Sherman F. Studies on transposable elements in yeast. I. ROAM mutations causing increased expression of yeast genes: their activation by signals directed toward conjugation functions and their formation by insertion of Ty1 repetitive elements. II. deletions, duplications, and transpositions of the COR segment that encompasses the structural gene of yeast iso-1-cytochrome c. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 2):593–607. [PubMed] [Google Scholar]
- Farabaugh P. J., Fink G. R. Insertion of the eukaryotic transposable element Ty1 creates a 5-base pair duplication. Nature. 1980 Jul 24;286(5771):352–356. doi: 10.1038/286352a0. [DOI] [PubMed] [Google Scholar]
- Feldman H. Arangement of transfer-RNA -genes in yeast. Nucleic Acids Res. 1976 Sep;3(9):2379–2386. doi: 10.1093/nar/3.9.2379. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gafner J., Philippsen P. The yeast transposon Ty1 generates duplications of target DNA on insertion. Nature. 1980 Jul 24;286(5771):414–418. doi: 10.1038/286414a0. [DOI] [PubMed] [Google Scholar]
- Gangloff J., Keith G., Ebel J. P., Dirheimer G. The primary structure of aspartate transfer ribonucleic acid from brewer's yeast. II. Partial digestions with pancreatic ribonuclease and T 1 ribonuclease and derivation of complete sequence. Biochim Biophys Acta. 1972 Jan 31;259(2):210–222. [PubMed] [Google Scholar]
- Goodman H. M., Olson M. V., Hall B. D. Nucleotide sequence of a mutant eukaryotic gene: the yeast tyrosine-inserting ochre suppressor SUP4-o. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5453–5457. doi: 10.1073/pnas.74.12.5453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hawthorne D. C., Leupold U. Suppressors in yeast. Curr Top Microbiol Immunol. 1974;64(0):1–47. doi: 10.1007/978-3-642-65848-8_1. [DOI] [PubMed] [Google Scholar]
- Liebman S., Shalit P., Picologlou S. Ty elements are involved in the formation of deletions in DEL1 strains of Saccharomyces cerevisiae. Cell. 1981 Nov;26(3 Pt 1):401–409. doi: 10.1016/0092-8674(81)90209-9. [DOI] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 1980;65(1):499–560. doi: 10.1016/s0076-6879(80)65059-9. [DOI] [PubMed] [Google Scholar]
- Maxam A. M., Tizard R., Skryabin K. G., Gilbert W. Promotor region for yeast 5S ribosomal RNA. Nature. 1977 Jun 16;267(5612):643–645. doi: 10.1038/267643a0. [DOI] [PubMed] [Google Scholar]
- Montgomery D. L., Boss J. M., McAndrew S. J., Marr L., Walthall D. A., Zitomer R. S. The molecular characterization of three transcriptional mutations in the yeast iso-2-cytochrome c gene. J Biol Chem. 1982 Jul 10;257(13):7756–7761. [PubMed] [Google Scholar]
- Nishikura K., Kurjan J., Hall B. D., De Robertis E. M. Genetic analysis of the processing of a spliced tRNA. EMBO J. 1982;1(2):263–268. doi: 10.1002/j.1460-2075.1982.tb01157.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olson M. V., Hall B. D., Cameron J. R., Davis R. W. Cloning of the yeast tyrosine transfer RNA genes in bacteriophage lambda. J Mol Biol. 1979 Jan 25;127(3):285–295. doi: 10.1016/0022-2836(79)90330-9. [DOI] [PubMed] [Google Scholar]
- Olson M. V., Montgomery D. L., Hopper A. K., Page G. S., Horodyski F., Hall B. D. Molecular characterisation of the tyrosine tRNA genes of yeast. Nature. 1977 Jun 16;267(5612):639–641. doi: 10.1038/267639a0. [DOI] [PubMed] [Google Scholar]
- Philippsen P., Kramer R. A., Davis R. W. Cloning of the yeast ribosomal DNA repeat unit in SstI and HindIII lambda vectors using genetic and physical size selections. J Mol Biol. 1978 Aug 15;123(3):371–386. doi: 10.1016/0022-2836(78)90085-2. [DOI] [PubMed] [Google Scholar]
- Piper P. W. A correlation between a recessive lethal amber suppressor mutation in Saccharomyces cerevisiae and an anticodon change in a minor serine transfer RNA. J Mol Biol. 1978 Jun 25;122(2):217–235. doi: 10.1016/0022-2836(78)90037-2. [DOI] [PubMed] [Google Scholar]
- Roeder G. S., Farabaugh P. J., Chaleff D. T., Fink G. R. The origins of gene instability in yeast. Science. 1980 Sep 19;209(4463):1375–1380. doi: 10.1126/science.6251544. [DOI] [PubMed] [Google Scholar]
- Roeder G. S., Fink G. R. DNA rearrangements associated with a transposable element in yeast. Cell. 1980 Aug;21(1):239–249. doi: 10.1016/0092-8674(80)90131-2. [DOI] [PubMed] [Google Scholar]
- Rothstein R. Deletions of a tyrosine tRNA gene in S. cerevisiae. Cell. 1979 May;17(1):185–190. doi: 10.1016/0092-8674(79)90306-4. [DOI] [PubMed] [Google Scholar]
- Sandmeyer S. B., Olson M. V. Insertion of a repetitive element at the same position in the 5'-flanking regions of two dissimilar yeast tRNA genes. Proc Natl Acad Sci U S A. 1982 Dec;79(24):7674–7678. doi: 10.1073/pnas.79.24.7674. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schweizer E., MacKechnie C., Halvorson H. O. The redundancy of ribosomal and transfer RNA genes in Saccharomyces cerevisiae. J Mol Biol. 1969 Mar 14;40(2):261–277. doi: 10.1016/0022-2836(69)90474-4. [DOI] [PubMed] [Google Scholar]
- Smith H. O. Recovery of DNA from gels. Methods Enzymol. 1980;65(1):371–380. doi: 10.1016/s0076-6879(80)65048-4. [DOI] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Sprinzl M., Grueter F., Spelzhaus A., Gauss D. H. Compilation of tRNA sequences. Nucleic Acids Res. 1980 Jan 11;8(1):r1–r22. [PMC free article] [PubMed] [Google Scholar]
- Sutcliffe J. G. Complete nucleotide sequence of the Escherichia coli plasmid pBR322. Cold Spring Harb Symp Quant Biol. 1979;43(Pt 1):77–90. doi: 10.1101/sqb.1979.043.01.013. [DOI] [PubMed] [Google Scholar]
- Temin H. M. Origin of retroviruses from cellular moveable genetic elements. Cell. 1980 Oct;21(3):599–600. doi: 10.1016/0092-8674(80)90420-1. [DOI] [PubMed] [Google Scholar]
- Tschumper G., Carbon J. Delta sequences and double symmetry in a yeast chromosomal replicator region. J Mol Biol. 1982 Apr 5;156(2):293–307. doi: 10.1016/0022-2836(82)90330-8. [DOI] [PubMed] [Google Scholar]
- Valenzuela P., Bell G. I., Masiarz F. R., DeGennaro L. J., Rutter W. J. Nucleotide sequence of the yeast 5S ribosomal RNA gene and adjacent putative control regions. Nature. 1977 Jun 16;267(5612):641–643. doi: 10.1038/267641a0. [DOI] [PubMed] [Google Scholar]
- Williamson V. M., Young E. T., Ciriacy M. Transposable elements associated with constitutive expression of yeast alcohol dehydrogenase II. Cell. 1981 Feb;23(2):605–614. doi: 10.1016/0092-8674(81)90156-2. [DOI] [PubMed] [Google Scholar]
- del Rey F. J., Donahue T. F., Fink G. R. sigma, a repetitive element found adjacent to tRNA genes of yeast. Proc Natl Acad Sci U S A. 1982 Jul;79(13):4138–4142. doi: 10.1073/pnas.79.13.4138. [DOI] [PMC free article] [PubMed] [Google Scholar]