Abstract
Dps is a multifunctional homododecameric protein that oxidizes Fe2+ ions accumulating them in the form of Fe2O3 within its protein cavity, interacts with DNA tightly condensing bacterial nucleoid upon starvation and performs some other functions. During the last two decades from discovery of this protein, its ferroxidase activity became rather well studied, but the mechanism of Dps interaction with DNA still remains enigmatic. The crucial role of lysine residues in the unstructured N-terminal tails led to the conventional point of view that Dps binds DNA without sequence or structural specificity. However, deletion of dps changed the profile of proteins in starved cells, SELEX screen revealed genomic regions preferentially bound in vitro and certain affinity of Dps for artificial branched molecules was detected by atomic force microscopy. Here we report a non-random distribution of Dps binding sites across the bacterial chromosome in exponentially growing cells and show their enrichment with inverted repeats prone to form secondary structures. We found that the Dps-bound regions overlap with sites occupied by other nucleoid proteins, and contain overrepresented motifs typical for their consensus sequences. Of the two types of genomic domains with extensive protein occupancy, which can be highly expressed or transcriptionally silent only those that are enriched with RNA polymerase molecules were preferentially occupied by Dps. In the dps-null mutant we, therefore, observed a differentially altered expression of several targeted genes and found suppressed transcription from the dps promoter. In most cases this can be explained by the relieved interference with Dps for nucleoid proteins exploiting sequence-specific modes of DNA binding. Thus, protecting bacterial cells from different stresses during exponential growth, Dps can modulate transcriptional integrity of the bacterial chromosome hampering RNA biosynthesis from some genes via competition with RNA polymerase or, vice versa, competing with inhibitors to activate transcription.
Introduction
Dps was initially described as an abundant protein of starved cells (The DNA-binding Protein from Starved cells) [1] and later became a prototype for the Dps family of bacterial ferritins. Playing a key role in genome condensation [2], it is actively produced during stationary phase reaching 85,000–180,000 molecules per cell [3, 4], while exponentially growing cells of Escherichia coli (E. coli) possess only 6,000–8,500 monomers of Dps. Proteins from the Dps family are composed of 12 identical or similar subunits, and form spherical particles with an internal cavity of ~4.5 nm. In E. coli this protein is a homododecamer with 2–3 tetrahedral symmetry [5].
Dps is a multifunctional protein protecting bacterial cells from oxidative stress, UV- and γ-radiation, as well as metal ion toxicity [6–11]. Proteins of this family also play a role in biofilm formation [12], and were found among outer membrane proteins and fimbria [13]. The protective function is mostly conferred by DNA-binding and ferroxidase activities of Dps, which are topologically separated, but function jointly to preserve DNA integrity and cellular viability [14]. Operating as a ferritin, Dps oxidizes supertoxic but essential ferrous ions and accumulates them within its cavity in the form of iron oxides [15, 16]. Ferroxidase activity of Dps is relatively well studied, and it is not very different from other ferritins. Although Dps oxidizes Fe(II) predominantly using hydrogen peroxide, rather than oxygen, and 12 catalytic centers are formed by surfaces of the 2 adjacent subunits, rather than being located within the 4-helix bundle of each subunit [17].
An ability to bind and condense genomic DNA is a specific feature of proteins from the Dps-family [18]. Both inner [5] and outer [5, 19] surfaces of the E. coli Dps are charged negatively, thus it is not clear how this protein was evolutionary selected for interaction with negatively charged DNA. Based on the X-ray structure [5] and the work of Ceci at al. [18], it is thought that the N-terminal tails, lacking a DNA-binding module, but containing positively charged amino acid residues, are responsible for interaction with DNA, for Dps self-aggregation, and for binding/aggregation-mediated DNA condensation. Deletion of the first 8 amino acids including Lys5 and Lys8, or 18 amino acids with Lys10 and Arg18 decreased both self-aggregation and DNA condensation [18]. Nonetheless, the mutant Dps molecules still bound DNA as ‘beads on a string’, whereas Dps of Listeria innocua and two Dps-like proteins of Bacillus anthracis possessing short N-termini were not able to do this [17, 20, 21]. On the other hand, Dps from Helicobacter pylori (NAP), which does not have positively charged N-termini at all, binds DNA exploiting its positively charged protein surface [22]. The situation with the DNA-binding modules becomes even more ambiguous, if to take into account that the N-terminal tails of the two Dps proteins from Lactococcus lactis form alpha-helices potentially capable of binding to a large groove of DNA, but do not show any dependence on the presence of Lys residues in their structure [23].
Mycobacterium smegmatis genome also encodes two proteins of the Dps family [24, 25]. Of them, Dps1 is produced predominantly at stationary phase, whereas Dps2 is synthesized constitutively [26, 27]. Both these proteins participate in phase-dependent DNA packaging into toroid-like structures, but only toroids formed by Dps2 are further converted into more stable coral reef structures [28]. The mode of DNA packaging may, therefore, be different even for highly homologous proteins. In the E. coli cells highly packed Dps—DNA co-crystals were also registered at the late stationary phase. Their formation during steady growth was observed both with [29] and without [30–32] mild overproduction of Dps. However, in exponentially growing E. coli cells, no sign of crystallization has been detected, even in conditions of the plasmid-born Dps overproduction [30], or active production of endogenous Dps in response to oxidative stress [33]. Intracellular concentration of Dps is, therefore, not the only factor that is required to switch on protective heterochromatization of genomic DNA in E. coli. At the same time, the Dps orthologs from Staphylococcus aureus can transform nuleoid into a condensed state in both exponential and stationary phases [34]. Such a diversity of features and the lack of clear correlation between structural organization and functional properties make it challenging to understand the mechanism of Dps-DNA interaction.
The presence of functionally important [18] positively charged modules, suggests a simple electrostatic interaction between Dps and DNA without any sequence or structural specificity [1, 2, 18, 22, 30]. If so, deletion of dps would not have a large effect on the pattern of gene expression, and this was likely the case for the proteome of exponentially growing cell cultures [1]. However, the spectrum of proteins produced in bacteria during starvation showed dramatic and differential alterations [1]. These changes were explained by the lack of Dps-mediated global structural impact on the bacterial chromosome or by regulatory effects from transcription factors, occupying the sites occasionally bound by Dps and released in its absence. For all that, genomic SELEX revealed sequences that were preferably bound by Dps in vitro (data deposited in DataBase TEC https://shigen.nig.ac.jp/ecoli/tec/top/about [35]), and we detected an ability of Dps to choose one of the two competing DNA fragments for complex formation [19]. Dps, therefore, may have some sequence or structural selectivity. Some preference of Dps for the ends of linear DNA fragments and even higher affinity for the branching point of artificial 3-way junction molecules was detected by atomic force microscopy [19]. Based on these data, it was assumed that Dps tends to bind those sites in the genomic DNA, which have locally increased “concentration” of DNA providing a platform for the maximal number of N-termini. Given that N-termini are grouped in triades around ferritin-like vertices of the dodecamer, the 3-way junction anchoring the Dps particles at the branching point seemed to be an optimal target for interaction. However, bent DNA, Holliday junctions, or any other structural features, increasing the availability of DNA for N-termini may be attractive as well. Here, we checked if Dps has any structural or sequence preferences at the genome-wide scale.
ChIP-seq experiments performed for exponentially grown cells revealed 451 sites with affinity to Dps and about 1200 genomic regions remained unbound. The newly revealed targets significantly overlap with the Dps binding sites found by SELEX [35]. In comparison to the control unbound sequences, they appeared to be enriched with inverted repeats and apparently overlap with binding sites of several other proteins of the bacterial nucleoid. Biological significance of a combined or permutable interaction of several structural proteins within one and the same genomic region is discussed. We checked the possibility of Dps participation in transcription regulation, and differential changes in intracellular amount of five mRNAs, but not all, were detected in response to the dps gene deletion. In all cases apparently regulatory effects can be explained by relieved competition with Dps for nucleoid proteins recognizing specific sequence motifs in their targets. Thus, it became clear that Dps can “specifically” modulate transcription of at least those genes that are regulated by transcription factors binding their DNA-targets with a lower affinity than Dps.
Materials and methods
Bacterial strains
All experiments, except protein production described in the next paragraph, were done using E. coli K-12 MG1655 cells. The dps-null mutant was first made in E. coli BW25113 as described in [36] using primers dps_del_1 and dps_del_2 (S1 Table), and then the mutation was transferred to E. coli K12 MG1655 by P1 transduction as described in [37]. After removal of the kanamycin cassette with pCP20, the whole-genome sequencing (Illumina, MiSeq) was used to check the presence of required deletion and to exclude the dps transposition in some other place. Bacterial cells were grown aerobically in M9 medium supplemented with 0.2% glucose and 5% LB at 37°C under constant shaking (~120 rpm) in a water bath and harvested at OD ~0.6.
Anti-Dps antibodies
The Dps protein was purified to homogeneity as described in [19]. In brief, the E. coli dps gene was amplified with primers dps_cl_1 and dps_cl_2 (S1 Table) and cloned into the plasmid pGEMΔXba [38] using the Xba I site. No tag was added to the recombinant gene. The nucleotide sequence of the insert was checked by direct sequencing. The gene was expressed in E. coli BL21(DE3) cells grown in LB Medium in the presence of ampicillin (100 μg/mL). Its transcription was induced by 0.02 mM IPTG at OD600 0.4–0.6, and biosynthesis of the protein was allowed for 12 h. Dps was purified using ion exchange chromatography on DEAE-Sephadex A-25 (GE Healthcare, Sweden) and gel-filtration on Sephadex G-200 (Pharmacia, Sweden). The protein was dialyzed against the buffer containing 50 mM Tris-HCl (pH 8.0), 50 mM NaCl, 0.1 mM EDTA and concentrated on Amicon Millipore 3kDa (Merck, USA) spin columns. The final purity of the native protein was higher than 95%. Two New Zealand rabbits were immunized subcutaneously with 100 μg of Dps emulsified in incomplete Freund’s adjuvant. Three booster injections were given using 75 μg of Dps emulsified in the same adjuvant with an interval of two weeks between injections. Animal health was daily monitored to ensure their healthy state at the end of experiment. Blood, collected from the rabbits 14 days after the last immunization, was used to prepare antisera. Antibodies were purified from the antisera by ion exchange chromatography on DEAE-Toyopearl as described in [39]. Final concentration of anti-Dps IgG determined with spectrophotometer ND-1000 (NanoDrop Technologies Inc., USA) was 3mg/ml. Antibodies were precipitated by 60% ammonium sulfate and stored at 4°C until use. All works with animals were carried out in accordance with the “Guidelines for Biomedical Research Using Animals” accepted in the Institute of Cell Biophysics 30.12.2011 and have been approved by the Animal Care and Use Committee of the Institute of Cell Biophysics of Russian Academy of Sciences. At the end of the experiment, the rabbits were euthanized with CO2 inhalation according to the GLP standards for euthanasia using carbon dioxide.
ChIP-seq experiments
ChIP-seq experiments were performed in duplicate using similar growth condition and protocols for chromatin immunoprecipitation. The first experiment was done in the Centre for Genomic Regulation (CRG, Barcelona, Spain), while the second one in the Immanuel Kant Baltic Federal University (Kaliningrad, Russia). In both cases, the chromatin was cross-linked, isolated and precipitated basically as suggested in [40] with certain modifications. Bacterial cells were grown aerobically at 37°C until OD600~0.6, then formaldehyde was added to a final concentration of 1% and incubation was allowed for 20 minutes. Cross-linking was stopped with glycine (final concentration of 450 mM). After 5 minutes of incubation, the cells were pelleted by centrifugation at 14,000 rpm for 15 minutes (+4°C), washed twice with 5 ml of PBS and resuspended in 1.3 ml of ice-cold immunoprecipitation buffer prepared from 50 ml of buffer containing 100 mM NaCl, 50 mM Tris-HCl (pH 8.1), 5 mM EDTA, 0.2% NaN3, 0.5% SDS, and 25 ml of buffer containing 100 mM Tris-HCl (pH 8.6); 100 mM NaCl; 5 mM EDTA; 0.2% NaN3, 5% Triton-X-100. Then phenylmethylsulfonyl fluoride (final concentration of 1 mM) or 20 μl of Protease Inhibitor Cocktail (PIC, Sigma) for the first and the second experiment, respectively, were added followed by incubation for 30 min at +4°C.
To shear the chromatin, in the first experiment samples were subjected to 18 cycles of sonication on Bioruptor (Diagenode, USA). In the second experiment, 40 cycles of sonication on 150 Soniprep Plus (MSE, UK) were applied. Cell debris was removed by centrifugation (15 min at 14,000 rpm and 4°C). Supernatant was checked for shearing quality and used for further procedures. The size of the obtained DNA fragments was in the range of 150–300 bp and 300–500 bp for the first and the second experiment, respectively.
For immunoprecipitation, 800 μl of collected chromatin (~1000 μg of protein) were incubated on a rotating wheel over night at 4°C with 10 μg of either rabbit anti-Dps antibodies (experimental sample) or rabbit pre-immune IgG (negative control). Next day 30 μl of Ultra Link Protein A/G beads (Thermo Scientific, USA) were added to the samples, and incubation was allowed for a further 2 hours at 4°C on a rotating wheel. The beads were washed 3 times with 1 ml of the low-salt buffer (50 mM HEPES pH 7.5, 140 mM NaCl, 1% Triton X-100, 1xPIC) and then with 1 ml of the high-salt buffer (500 mM NaCl). Immunoprecipitated DNA-protein complexes were removed from the beads by 3 h shaking at 65°C and 1000 rpm in 110 μl of freshly prepared elution buffer, containing 100 mM NaHCO3 and 1% SDS. After centrifugation at 3000 rpm for 5 minutes, 100 μl of the supernatant was transferred to a new tube, and the DNA was purified with a PCR Purification Kit (Qiagen, Germany). The DNA concentration was measured on Qubit 2.0 using Qubit dsDNA HS Assay kit. A total of 8–10 identical samples were combined and concentrated to a volume of 30–50 μl containing 5–10 ng of the DNA. Before library preparation, the samples were qPCR-checked for the enrichment with the target DNA. Assuming that Dps can bind the regulatory region of its own gene, two fragments from the dps regulatory region amplified with primer pairs dps_F1 –dps_R1 and dps_F2 –dps_R2 (S1 Table) were used for validation.
Library preparation
ChIP-seq libraries were prepared from 5–10 ng of the DNA samples with the NebNext Ultra DNA Library Prep Kit for Illumina (New England Biolabs, MA, USA) following the manufacturer’s instructions. For the final amplification of the library 15 PCR cycles were used. Size distribution and concentration of the amplicons was checked on the Bioanalyzer 2100 (Agilent, USA). In the first experiment, the maximum was at about 300 bp, and ChIP libraries were sequenced using 50 nt single-end read protocol on the Illumina HiSeq system (Illumina 2000, USA) of the Genomics Facility in the Centre for Genomic Regulation (Barcelona). In the second experiment, the maximum was at about 450 bp, and samples were sequenced using standard paired-end 2*150 nt protocol on the MiSeq system (Illumina, USA) in the Immanuel Kant Baltic Federal University (Kaliningrad).
Chip-seq data evaluation
Raw data were quality checked with the FastQC software (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/). The output files obtained from Illumina HiSeq contained 32,860,548 and 47,147,298 of 50 bp single-end reads in the control and immunoprecipitated samples, respectively. The output files from Illumina MiSeq contained, respectively, 10,028,552 and 11,570,890 reads, varying in the range from 35 to 150 nucleotides. All four sets were quality filtered on the Galaxy server (Filter by Quality tool, Q≥20) providing 99% probability of correct sequencing for all nucleotides in each read [41]. As a result, our sets contained 31,656,551 and 45,396,252 sequences in the control and immunoprecipitated samples of the first experiment, and, respectively, 7,493,528 and 8,214,737 reads for the second experiment. The data discussed in this publication have been deposited in NCBI's Gene Expression Omnibus [42] and are accessible through GEO Series accession number GSE102091 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE102091).
Two different approaches were used to align sequence reads to the genome of E. coli K-12 MG1655 (U00096.3). First, quality controlled reads were mapped using the CLC Genomics Workbench version 7.5.1 (CLC GW, Bio-Qiagen, Aarhus, Denmark) with either default settings for the length and similarity fractions (0.5 and 0.8, respectively) or with the most stringent criteria (1.0 and 1.0, respectively). Reads with sequences aligned to multiple genomic regions were ignored. In both cases, peak calling provided as much as several thousand of the Dps-bound areas with p-value less than e-3.
To apply the second approach variable in length reads from Illumina MiSeq were first trimmed from both sides to obtain a set of standard 50 nt sequences taken from the middle of longer reads. Reads shorter than 50 nt were discarded. Then, all four sets from both experiments were aligned to the genome using the Matcher program [43] (available at: http://www.mathcell.ru/DnaRnaTools/Matcher.zip). This software maps only 5’-ends if the reads correspond to the top strand of genomic DNA, or only 3’-ends, if they are aligned to the bottom strand. As such, the signals from fully complementary reads will match the same (left) position. Reporting the distribution of matching reads across the genome the program also evaluates reads with multiple occurrence and their positions. Only ideal correspondence to the genome was permitted. The profiles obtained for experimental and control samples were normalized by the scaling method initially offered by Affymetrix for microarray data analysis [44] and later implemented in several other approaches [45]. This method assumes unaffected protein occupancy at most genomic positions and quantifies the scaling factor on the basis of corrected mean values obtained after removal of 2% signals with highest and lowest intensities from both control and experimental sets.
After normalization, the read counts were estimated in the running windows of 25, 35 or 75 bp, and the ratios R between values obtained for experimental and control libraries were calculated. Peaks were localized requiring R≥1.5 for at least 50% positions of at least 60 bp genomic region in length (corresponds to the assumed size of the Dps binding site [18]). Genomic regions with R<1.0 for all positions in both experiments, spanning for at least 60 bp that did not overlap with peaks found by CLC GW with p-values < 1e-4 were collected into the set “unbound regions” (UR). Regions separated by less than 30 bp were combined if R-values in all intermediate positions did not exceed 1.1.
Search for genomic units that overlap the Dps binding sites
More than a hundred sets of genomic regions with different functional loads were compared with the selected set of the Dps binding sites. These genomic units included binding sites of different proteins obtained by ChIP-chip [46–48] or ChIP-seq [49–51], promoter islands found computationally [43, 52], REP-elements annotated in the E. coli genome, and direct or inverted repeats identified by the Unipro UGENE software package [53].
Before analysis genomic coordinates of all data sets were translated into current version (U00096.3) of the E. coli MG1655 genome. Then, the total areas in the genome occupied by Dps and each protein of interest or a specific element were estimated. Thereafter, an expected number of overlapping base pairs in the two analyzed sets, in the case of their independent distribution, was calculated and compared with the observed intersection. All used ChIP-chip and ChIP-seq data were processed by their authors. These data, in which protein binding sites are listed with left and right borders [48–51], were directly used for positional comparison. In ChIP-chip data sets [46, 47] the lists of 60 bp microarray probes with averaged and normalized enrichment ratio were provided. Thus, the probes with at least 2-fold enrichment were selected, and 60 bp regions were assessed for overlap with genomic regions bound and unbound by Dps. An overlap with RNAP σ70 binding sites (aerobic growth) obtained by ChIP-seq and presented as point coordinates of peak maxima for 36 bp reads [50] was evaluated by the same way.
Search for dominant sequence motifs
Nucleotide sequences over presented within the Dps binding sites were searched by the MEME suite programs (version 4.11.2, http://meme-suite.org/tools/meme, [54]) and DMINDA (http://csbl.bmb.uga.edu/DMINDA/annotate.php#tabs-1, [55]) as described in the text. Both software packages were run in the discriminative mode so as to discover motifs enriched in the experimental library compared to the set of sequences that are not bound by Dps.
qRT-PCR
The total fraction of cellular RNAs was isolated from the wild type E. coli K12 MG1655 and the dps-null mutant cultures using TRIzol reagent (Ambion, USA) and then treated with DNAse I (New England Biolabs) following the manufacturers’ protocols. Primers used are listed in S1 Table. cDNAs were obtained in 25 μl of the reaction mixture, containing 1 or 2 μg of RNA and 80U of RevertAid reverse transcriptase (Thermo Scientific, Lithuania) according to the manufacturer’s protocol. Two microliters of this mixture were subsequently used for PCR performed in 20 μl with 4 μl of qPCRmix-HS SYBR (Evrogen, Russia). qPCR was run on DTlite thermal cycler (DNA Technology, Russia). The PCR program included 1 minute of initial melting at 95°C, followed by 35 cycles: 95°C for 20 s, annealing at 56°C for 20 s, and synthesis at 72°C for 25 s. The fluorescence of SYBR Green I was measured at the end of each cycle for 15 s. Samples without reverse transcriptase were used as negative controls. Two RNA products were used as a reference to estimate expression efficiency of chromosomal genes. The first one, transcribed in antisense direction to ysaA, was used previously [56]. The second one (lacZ-mRNA) was selected from the set of regions unbound by Dps (page 4 in S2 Table). The data obtained with two reference products were averaged. In qRT-PCR experiments for GFP-reporter assays (see below), mRNA of kanamycin resistance gene (kan) was used as a reference product. Specificity of the synthesized products in all cases was confirmed by electrophoresis in 5% polyacrylamide gel. The threshold cycles calculated by the qPCR software (DNA Technology, Russia) for the target and the reference genes in both the wild type and the dps-null mutant samples were used to estimate the mutation-mediated changes. Relative abundance was calculated using 2-ΔΔCT method. The computed values were expressed as fold ratios and averaged between 4–8 biological samples with 2, 3 or 6 technical replicates for each of them. Statistical significance of differences was assessed using the Student's t-test.
GFP-reporter assays
To assess the ability of Dps to affect expression of its own gene, the dps regulatory region (primers dps_F1 and dps_R2) was cloned upstream of the reporter gfp gene in the plasmid pET28b-eGFP [57]. It was then transformed into the wild-type and the dps-null mutant cells. Fluorescence intensity was registered in colonies grown on LB agar supplemented with kanamycin (20 μg/ml) using a Leica microscope (2.5x objective). Plasmid with no promoters inserted upstream the dps gene was used as a control. Data were processed with ImageJ software [58].
Results
Dps binds genomic DNA in a non-random manner
Two data sets from the ChIP-seq experiments were obtained on HiSeq 2000 and MiSeq Illumina systems, providing sequence libraries with a substantially different sequencing depth. From the first experiment, 27,850,801 reads of the control set and 42,133,025 reads of immunoprecipitated sample were aligned to the E. coli MG1655 genome, with default settings in CLC GW. Stringent criteria gave 25,682,627 and 39,199,442 mapped reads, respectively. In the MiSeq experiment, we obtained 5,952,682 and 7,102,289 aligned reads with the default settings of CLC GW, while stringent criteria gave 4,341,979 and 5,788,335 hits, respectively. Depending on the settings, 14,875 or 14,309 peaks at least 60 bp in length were found in the first experiment with average p-values 5.9E-03 or 6.4E-03. If the peaks located at a 0–90 bp distance were combined (fused peaks are shaded gray in Pages 1 and 2 of S2 Table), the number of targeted areas decreased, but was still large (10,516 and 10,241 peaks, respectively). The second experiment gave, respectively, 5,350 and 4,772 combined peaks (Pages 3 and 4 of S2 Table) with average p-values 7.5E-03 and 9.2E-03. About 85% of the peaks detected in the second experiment (both settings) overlap those from the first experiment (the overlapping regions are marked bold in S2 Table). Given that the experiments were done in different laboratories and the samples were sequenced on different platforms, this similarity can be considered good, providing a chance to detect the core set of preferred Dps binding sites in the genome.
To validate them, we analyzed the bar chart representing the genome-wide profile of Dps targets detected in a SELEX experiments in vitro (TEC DataBase (https://shigen.nig.ac.jp/ecoli/tec/top/, [35]). From these data 371 peaks were localized with 100 bp accuracy, of which 326 overlapped with the peaks revealed in one or both our experiments. Thus, the data obtained in vivo (S2 Table) and in vitro (https://shigen.nig.ac.jp/ecoli/tec/top/) assume the presence of preferred targets for Dps in the genome of E. coli.
Nevertheless, before a detailed analysis of the received output data, at least two issues should be addressed. First, based on the automatic peak calling, an average size of the sites occupied by Dps in the two experiments appeared to be different: 102–104 bp and 214–220 bp (S2 Table). This discrepancy could reflect some functional features that were differentially realized in the two experiments or arise due to different length of sequence reads that have been mapped and subjected to peak calling (standard 50 nt in the first experiment and 35–150 nt in the second). For that reason, the next screen was made for 50 nt sequences only. In the case of the second experiment they were taken from the centre of longer reads.
Second, both large-scale distributions of revealed Dps-binding sites show a similar bias towards the first half of replicores (Fig 1A). This bias may indicate specific enrichment of this region with the preferred Dps targets or, otherwise, reflect the natural difference in the DNA copy number that is higher near the origin of replication (Figs 2B and S1 central circles, [59]). CLC GW relies on the peak shape scores, which are calculated for the complete genome and identifies peaks as genomic regions where these scores are greater than a given threshold. To compensate the copy-number mediated bias, the peak calling in the next screen has been done using the R parameter estimated as a fold ratio between the normalized number of sequence reads registered in the immunoprecipitated and control sequence libraries.
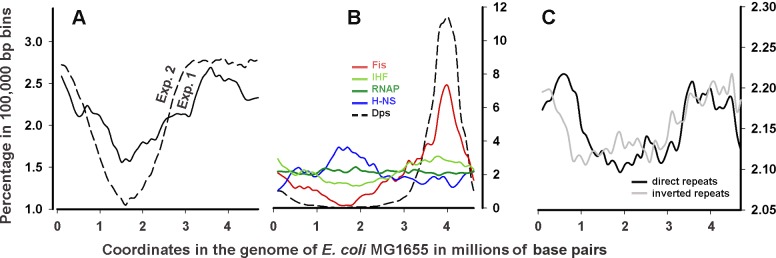
Fig 1. Large-scale profiles of the Dps targets correlate with the landscape of direct and inverted repeats and the pattern of Fis binding sites.
A: Distribution of the Dps contact regions along the E. coli MG1655 genome identified by CLC GW in two experiments (the default settings). The areas covered by Dps are combined in 100,000 bp bins and plotted as percentage to the total length of all sites occupied by Dps. B: The same for the contact sites of Fis [49], IHF [51], H-NS [49] and RNAP [50] from the cells grown in conditions similar to those used in our experiments. The plot for Dps shows the distribution of the sites from the combined set (CS) (Page 3, S3 Table). C: The same for direct (5–24 bp separated by 1–15 bp) and inverted (5–18 bp separated by 3–20 bp) repeats collected from the genome of E. coli MG1655 using Unipro UGENE [53].
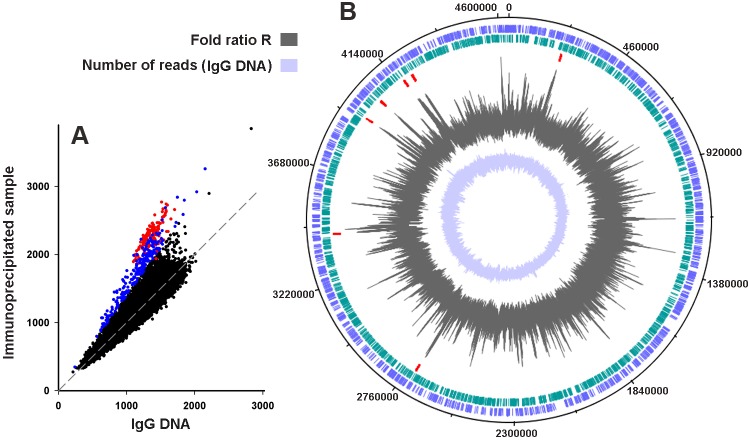
Fig 2. Dps binds genomic DNA in a non-random manner.
A: Correlation between the number of reads from immunoprecipitated and control sequence libraries calculated for 100 bp bins (first experiment). All points, corresponding to rRNAs operons are marked in red. All other points with R-values higher than 1.4 are plotted in blue. Dashed line shows the bisectrix of the plot. B: Distribution of the Dps-binding sites in the genome. Two outer circles represent the gene map of the top and the bottom strands of the E. coli MG1655 genome. The red ticks on the third circle mark positions of rRNA operons. The profile of R-values and the distribution of reads registered in the control library calculated for a 35 bp running window are plotted on the fourth and fifth circles, respectively.
The reads from both experiments were aligned to the genome using Matcher program [43] and analyzed as described in Materials and Methods. Correlation plots for control and immunoprecipitated samples are shown in Figs 2A and S1A. They demonstrate a high quality of normalization for both experiments and also indicate the presence of outliers. Many windows with R-ratio ≥ 1.4, found in the first experiment (blue dots in Fig 2A) were also found among outliers of the second experiment (blue symbols in S1A Fig), but the preferential interaction with ribosomal operons was detected only in the first one (red dots in Figs 2A and S1A). Thus, providing a chance to identify a common set of regions bound by Dps in both experiments, the data also revealed certain rearrangements in their profiles, which were caused by some untraceable differences in growth conditions.
Peak calling was made using the R-ratio of read counts found in 25, 35 or 75 bp running windows in the libraries of immunoprecipitated and control samples aligned to the genome. Two shorter windows gave contact sites better corresponding to each other and to the peaks found by CLC GW (see Figs 2B and S1B for 35 and 25 bp running windows, respectively). Of them 35 bp window also decreased the number of “empty” areas. It was, therefore, used to obtain Dps binding sites for further analysis.
Only 193 binding sites for Dps were found in the first library (Page 1, S3 Table), including seven very long regions (over 5,000 bp) with genes of ribosomal RNAs (red outliers in Fig 2A) and many tRNAs genes (Page 1, S3 Table). These genomic regions were poorly presented or absent in the set obtained by CLC GW, since reads with multiple matching to the genome were ignored. Yet, 178 out of 193 (92%) sites, found by the Matcher-based approach, at least partially overlap with peaks collected by CLC GW (indicated in bold in Page 1, S3 Table). Approximately 50-fold difference in the number of peaks revealed by the two approaches (Pages 1–2, S2 Table and Page 1, S3 Table) was not too surprising. Searching for peaks exceeding threshold for a given number of sequence reads, CLC GW can find more regions with high shape scores than Matcher does requiring higher than given cut off level fold-ratio between control and experimental samples. The values of R-ratio in the selected peaks of the first experiment varied from 1.52 to 2.56, thus the scale of fold changes was rather narrow and roughly similar across the genome (Fig 2B). Dps targets found by the Matcher-based approach appeared to be longer than those found by CLC GW (average size even without rRNA operons equals to 292 bp). Though, the size distribution histogram has a maximum in the range of 110–130 bp (S2B Fig) that nicely corresponds to the values detected by the CLC GW-mediated screening (100–110 bp, S2A Fig).
In the second experiment, Matcher revealed 1647 Dps binding sites (Page 2, S2 Table, S1B Fig). This is only three times fewer than in the case of the CLC GW screening, mainly because of lower coverage and, correspondingly, a lower background as compared to the first experiment. The range of the R-value variations in the peak maxima in this case was slightly wider: from 1.67 to 9.63, but still narrower, than it is typical for the binding sites of transcription factors (see, for instance ref. [47] and [50] for FNR). Again, a large part of sites, found by the Matcher-based mapping (83%), at least partly overlapped with the peaks collected by CLC GW (indicated in bold in Page 2, S3 Table). An average length of the Dps binding sites estimated on the basis of the second screen (195 bp, Page 2 in S3 Table) was approximately the same as assessed previously (214–220 bp, Pages 3 and 4 in S2 Table). However, the peak maxima in the size distribution histogram that was in the range of 160–190 bp (S1A Fig), shifted to 110–130 bp, coinciding with the first experiment (S1B Fig). As such, eleven-thirteen DNA helix turns can be suggested as a tentative estimate for the size of the site occupied by Dps in vivo, although it is twice as long as the estimate made in vitro on the basis of titration assays [18].
The set of the Dps targets detected in both experiments contains 451 genomic regions (Page 3, S3 Table). It was composed on the basis of 1647 peaks found in the second experiment (Page 2, S3 Table), from which the sites that were unbound in the first experiment were removed. Although the bias conditioned by an increased DNA copy number near the replication origin was, at least partially, compensated by the R-ratio, the binding sites detected in both experiments were mainly found in the last quarter of the genome (Fig 1B, dashed line). The most stable pattern of interaction is, therefore, located in this region. These genomic regions are further referred as CS (combined set). They were compared with 1227 genomic loci unbound by Dps in both ChIP-seq experiments and SELEX screening (Page 3, S3 Table), hereinafter referred as UR.
Dps-binding sites are enriched with inverted repeats
Using electrophoretic mobility shift assays we recently found that in vitro Dps effectively binds artificial DNA molecules with a 3-way junction, and the direct contact with these polynucleotides was observed in their branching point [19]. Thus, we assumed that stronger binding may take place at the DNA loci that provide a platform for the simultaneous interaction with several N-tails of the Dps particle due to locally increased DNA concentration. If so, it might be expected that the Dps binding sites are enriched with inverted and/or direct repeats that can form helix-loop structures in the negatively supercoiled DNA due to alternative interactions with neighboring complementary sequences on the same or opposite strand.
Genomic distribution of 5–24 bp-long direct repeats separated by 0–20 bp, and 5–18 bp inverted repeats separated by 3–20 bp was obtained with the integrated Unipro UGENE toolkit (http://ugene.net/) [53, 60]. To estimate the difference between the expected and the observed overlap of these ubiquitous genomic elements with the Dps binding sites, each pair of repeats was considered as a holistic unit, and the total intersection of each type of repeats with the combined set of Dps targets (Page 3, S3 Table) was evaluated with a 1 bp resolution.
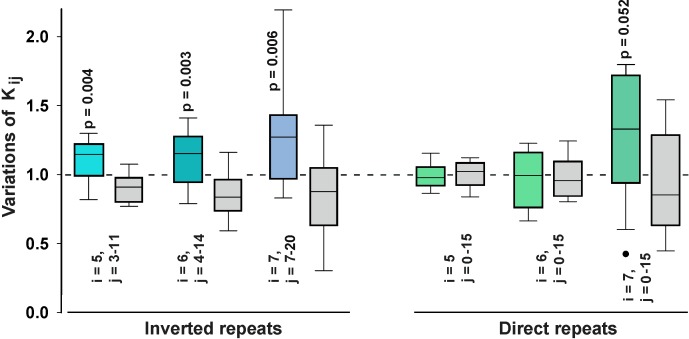
The number of overlapping nucleotides was then compared with the value expected by chance and the ratio between these two numbers (Kij) was used as a measure of deviation from the expected co-occurrences for pairs of different lengths (i) and spacers (j). For instance, the bacterial genome contains 2993 inverted pentanucleotides separated by 3 bp. Their total length in the genome is 38909 bp (0.838%), assuming 916 bp overlap with 109330 bp of CS. In reality we found 1134 common base pairs, which gave 1.24-fold excess over expected ratio (Ki-5,j-3 = 1) and one (i = 5, j = 3) of nine values (i = 5, j = 3–11) to the first box-plot in Fig 3. Using this approach, we estimated Kij values for all pairs of repeats, but only those that were 5–7 bp long gave statistically reliable values (Fig 3) without zero occurrences in both sets.
Fig 3. Dps-binding sites are enriched with inverted repeats.
The overlap between sequences of CS (colored box-plots) and UR (gray boxes) with direct or inverted repeats was characterized by the parameter Kij as described in the text. Black dot on the right panel shows one outlier. Box-plots with statistically significant differences are provided with corresponding p-values. Regions of bound and unbound sets overlapping with repeated sequences of both types are indicated in S3 Table.
We found that pairs of inverted repeats tend to be overrepresented at the Dps binding sites and underrepresented in the set of unbound regions. The statistical significance of the difference was maximal in the ranges of distances (j) shown in Fig 3. Thus, we conclude that Dps-binding sites are enriched with inverted repeats, which may facilitate or stabilize the Dps-DNA interaction. The situation with direct repeats, which large-scale distribution in the genome follows the profile of inverted repeats (Fig 1C), is less clear. The numbers of penta- and hexanucleotides in CS and UR vary in approximately the same range, and Kij are close to the expected values (Fig 3). The number of heptanucleotides within Dps binding sites tends to be higher than in UR, but the difference is statistically insignificant (p = 0.056).
Dps has high affinity to REP-elements and promoter islands
The E. coli genome has two types of structure-specific elements with repeated sequences. They include interspersed across the genome REP-elements (Repetitive Extragenic Palindromic sequences) [61] and promoter islands [62]. REP-elements are 25–35 bp in length and usually occur in multiple copies at one genomic location [61]. Nearby copies are always in inverted orientations potentiating their folding in complex secondary structures with some participation of short palindromes present in a single copy. We analyzed the overlap of CS and UR with all 355 REP-sequences annotated in the genome of E. coli [63], and found a 1.65-fold excess in the CS intersection level with 302 REP-elements containing 1–3 copies of the repeating sequences (Fig 4A). Their occurrence in the UR set, on the contrary, was significantly below normal (Fig 4B). Short REP-elements can thus be considered as suitable targets for interaction with Dps.
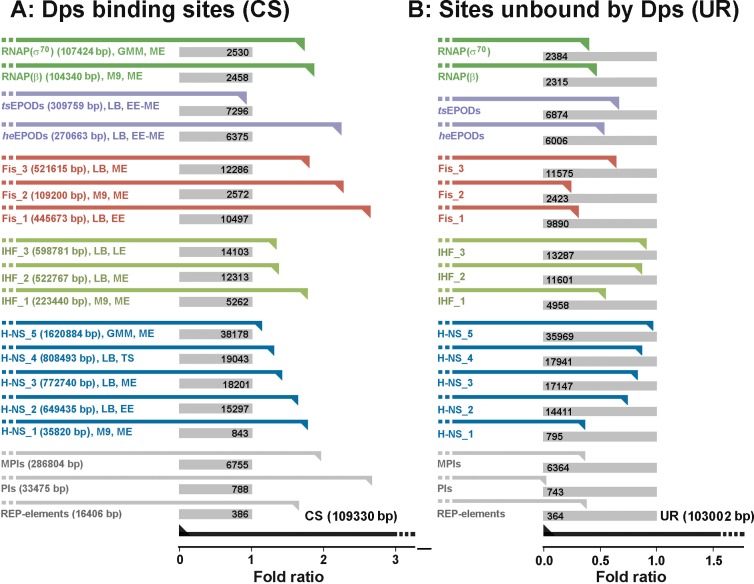
Fig 4. Dps shares its binding sites with other proteins of bacterial nucleoid and has affinity to REP-elements and promoter islands.
Intersection of Dps targets (A) and sites unbound by Dps (B) with structural and functional elements of bacterial genome was estimated as described above for repeated sequences and plotted as fold ratio to the expected values. Bent black arrows on the bottom schematically show areas occupied by CS or UR. Gray rectangles and numerals inside indicate the expected number of common base pairs if compared modules are independently distributed along the genome. Gray and colored bent arrows show registered overlap calculated in 1 bp resolution. Numerals in parenthesis indicate the size of compared sets. Genomic locations of REP elements were taken from KEGG DataBase (http://www.genome.jp/kegg/, [63]), and fold ratios obtained for 302 REP-sequences containing 1–3 REP-modules (14–100 bp) were plotted. Analyzed ChIP-chip and ChIP-seq data sets were obtained from [46–51] for cells grown in LB medium (LB), M9 medium with fructose (M9) or MOPS minimal medium with glucose (GMM), harvested at early (EE), middle (ME) or late (LE) exponential phase or upon transition to the steady growth (TS).
Promoter islands have a high density of closely located or even overlapping promoters in both orientations [43, 52] creating repeats of both type. Two sets of islands were used for comparison. The first one, PIs, contained 78 promoter-rich regions initially found by σ70-specific promoter finder PlatProm [52]. Another one was composed of 434 mixed promoter islands (MPIs) found by the unified version of PlatProm searching for promoter-like signals on the basis of structural information, which is similar for promoters of different σ-factors [43]. Both sets of promoter islands significantly overlapped with the sites occupied by Dps (Fig 4A), while their presence in the UR set was much lower than expected by chance (Fig 4B). Dps, therefore, has enhanced affinity for promoter islands.
Dps shares binding sites with structural proteins of bacterial nucleoid
The interest to promoter islands was stipulated by their specific structure and association with horizontally acquired genes [43, 64]. Such genes are usually subjected to silencing by H-NS [65, 66] that occupies up to 90% of PIs during exponential growth [64]. As estimated above, about 13% of the islands were in contact with Dps (Page 3, S3 Table), allowing a potentiality for simultaneous interaction with these two proteins. Of 57 Dps binding sites, overlapping with promoter islands, 54 can also bind H-NS (Page 3, S3 Table). In CS, there are 181 Dps-bound sequences targeted by H-NS (Page 3, S3 Table). Maximal excess (1.76-fold) was obtained for cells grown in M9 media and harvested at exponential phase (Fig 4A, [46]), but the overlap of Dps and H-NS binding sites is likely to decrease upon transition to a steady growth (Fig 4A). The intersection of CS with IHF was approximately the same (Fig 4A). It was found for 177 Dps targets, of which 91 can also interact with H-NS (Page 3, S3 Table). The maximal overlap of CS was detected for Fis binding sites (1.8–2.6 fold excess over expected value). That was anticipated, as Fis and Dps are the most abundant nucleoid proteins in bacterial cell during exponential and steady growth, respectively, and Fis inhibits nucleoid condensation at logarithmic phase [33]. More than a half (230) of the Dps binding sites overlapped with Fis contact regions, and there were 62 genomic regions with an average length of 352 bp occupied by all four proteins considered here. Dps, therefore, shares its binding sites with at least three other structural proteins of the bacterial nucleoid.
Sequences overrepresented in the Dps binding sites have a common motif with consensuses of other nucleoid proteins
The overlap of genomic regions occupied by Dps with binding sites of other nucleoid proteins makes the identification of a specific motif for Dps challenging and problematic. A preliminary search for over presented sequences was undertaken with MEME suite programs [54] for the 304 shortest samples from CS (total size—49789 bp, i.e. less than the 50000 bp limit of the software). Programs of this suite integrated into MEME-chip [67] were run in a discriminative mode with the 739 shortest sequences of UR (total size—49988 bp) and predicted three dominant motifs with E-values in the range of 4.6E-021–1.0E-011, including T(G/A)A(t/c)A, TA(c/t/g)(T/A), TT(T/a)(g/a) and one long sequence: (t/c)(g/c)(t/c)AGGCC(g/t)GATAAG(g/a)CG, which, however, was found only in 10 samples. DMINDA [55] predicted more potential candidates as consensuses of grouped overrepresented sequences. However, many of them contained A/T-rich motifs typical for nucleoid proteins, H-NS (consensuses: (a/g)ATA(A/t)(t/a) [49] or (T/g/c)(C/g/a)G(A/t)T(A/T)a(A/t)(t/a)t [68]), Fis (tG-t(g/t)(a/g)tTTTTT(c/g/a)-Ca [49]), IHF ((t/c)-(A/T)—(g/c)(t/c)—(A/t)(T/a)(t/a)(t/a) [51]), or some other transcription factors. For instance, TGAT, is a part of sequences recognized by H-NS, Fis and also FNR (aaa-tTGAt-ta-(a/g)TCAAtta(a/t)t [46] or tTGAt—-aTCAa [50]). During exponential, growth FNR has a limited number of binding sites, though their overlap with regions occupied by Dps is 1.9- and 2.7-fold higher than expected by chance (data from [47] and [50], respectively; S3 Table for FNR-bound regions). We, therefore, removed 336 genomic regions bound by other proteins from CS and used the remaining 115 sites occupied by Dps (average size 207 bp) for sequence analysis. In this case, the best motif found by MEME suite was (t/c)GATA (E = 4.8e-007), while DMINDA generated six groups with longer consensuses corresponding to this sequence for at least four nucleotides. They included: TG(G/a)(T/c)GAT (E = 4.7e-025), CTG(A/G)(T/c)AA (E = 1.7e-011), GATA-CG (E = 3.6e-009), cgCCTGATGC (E = 9.9e-009), C-GGCGAT (E = 3.2e-007) and CGATa-CG (E = 3.6e-006). When 84 of 115 sequences containing at least one of those motifs were again used to find overrepresented motifs in the Dps binding sites, the dominant sequences revealed by MEME appeared to be GATA (E = 1.0e-007), while DMINDA suggested TG-tGAT as the most statistically significant (E = 4.6e-018) motif. Thus, the presence of overrepresented motifs in the Dps-bound genomic regions can not be ignored and they may be important for complex formation. However, a lack of a typical DNA-binding domain in the Dps protein and presence of the revealed motifs in the sites specifically recognized by other nucleotide proteins and transcription factors, also allows a suggestion that Dps prefers to bind DNA near the regions occupied by other proteins.
Deletion of Dps differentially affects gene expression
Almost 50% of Dps-binding sites are immersed in the coding sequence of genes. The others at least partially encompass promoters or/and terminators, where different repeating sequences act as components of binding modules for the transcription factors, or transcription stop signals (Page 3, S3 Table). These targets include 66 sites located between divergently transcribed genes and 33 sites in the spacers between convergent genes. The genome has an equal number of spacers separating divergent and convergent genes, so this bias toward the promoter regions may indicate the involvement of Dps in transcription regulation. Consistent with this interpretation, Dps targets strongly overlap with RNA polymerase (RNAP) binding sites (Fig 4) and of the two types of genomic domains with extensive protein occupancy, only those that are enriched with RNA polymerase molecules were preferentially occupied by Dps (Fig 4F). Those domains (EPODs, Extensive Protein Occupancy Domains) were identified without the use of any specific antibody and were classified by the authors [48] into two functional categories: highly expressed (he) and transcriptionally silent (ts) EPODs. Excessive overlap with Dps contacts was obtained only for heEPODs (Fig 4A). We, therefore, tested the propensity of Dps to affect expression efficiency of 8 genes using the dps-null mutant.
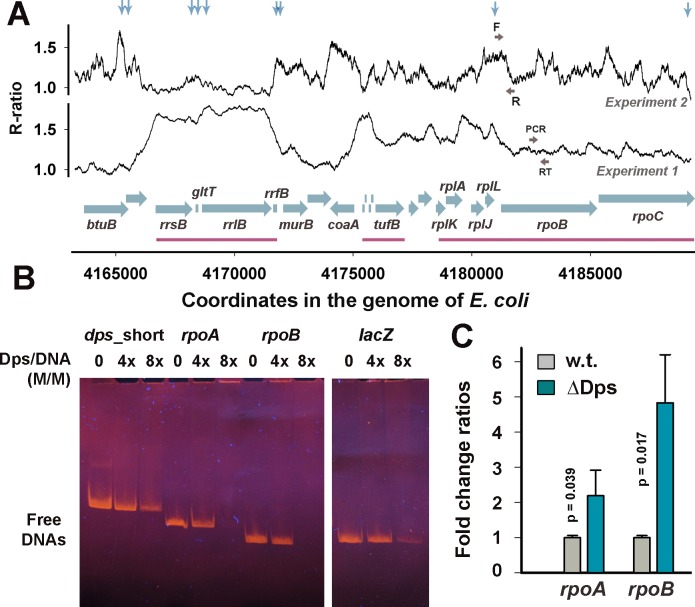
Since the two ChIP-seq experiments revealed a fundamental difference in the interaction of Dps with ribosomal operons, it was reasonable to test the dependence of their expression on the presence of Dps. However, seven copies of rRNA genes complicate the data evaluation, so we took rpoA and rpoB genes, encoding α- and β–subunits of RNAP as representatives of genes from the operons of ribosomal proteins. Fig 5A shows the profiles of Dps contacts in the rpoB genomic region. There are three operons with ribosomal genes. All of them were fully occupied by Dps in the first experiment, including the part of the rpoB coding sequence, while in the second experiment the base line remained at the level of the control sample (R ≈ 1.0) and even fell below it within and nearby the rrsB operon. Almost the same was true for rpoA, transcribed as a polycistronic unit rpsMKD-rpoA-rplQ (S3A Fig). In the first experiment this and the neighboring rplNXE-rpsNH-rplFR-rpsE-rpmD-rplO-secY-rpmJ operon were extensively occupied by Dps (S3A Fig, Page 1 in S3 Table), while in the second assay the R-values only slightly exceeded the control level, and were below it in the rpoA region (S3B Fig). This remodeling was also detected for some other genes with growth-dependent expression. For instance, the oppABCDF operon (S4A Fig), encoding inner membrane proteins of ABC transporter and following stringent control reprogramming similar to ribosomal genes [69], was covered by Dps in the first experiment, while in the second one only two small peaks have maxima exceeding R = 1 (S4A Fig). In contrast, the binding of Dps to the fis genomic locus was detected only in the second experiment (S5A Fig). The rho gene, which encodes a transcription terminator and is expressed independent from ribosomal genes together with its neighborhood (S6A Fig), as well as the dps gene, selected as potential target for autoregulation (Fig 6A), represent examples of the genomic loci bound by Dps in both experiments, while the lacZYA, mhpR-lacI and mhpABCDFE operons exemplify regions remained unbound in both experiments (S3B Fig).
Fig 5. Deletion of the dps gene affects rpoA and rpoB expression.
A: Profiles of the Dps binding sites obtained in two experiments (indicated) for the genomic region with three operons of ribosomal genes (running window of nine 35 bp bins). Genes are represented by blue horizontal arrows; magenta lines show ribosomal operons. Vertical arrows mark locations of inverted repeats (if longer than 7 bp). B: Band shift assays performed for indicated genomic loci. The regulatory region of the dps gene was used as a positive control for all band shift assays in this study. Fragment from the lacZ coding sequence was used as a reference gene for qRT-PCR experiments. Positioning of primers for amplification (F and R) is indicated in panel A of this figure, Fig 6A (for the dps regulatory region) and in S3 Fig (for rpoA and lacZ). C: Changes in the expression efficiency of selected genes in response to dps deletion. Primers used for reverse transcription and consecutive PCR are designated as RT and PCR, respectively, here and all other figures. Expression levels were estimated based on 3 and 5 biological samples (3–18 technical repeats in each) for rpoA and rpoD, respectively. Error bars show an average deviation. Statistical significance was assessed using Student’s t-test.
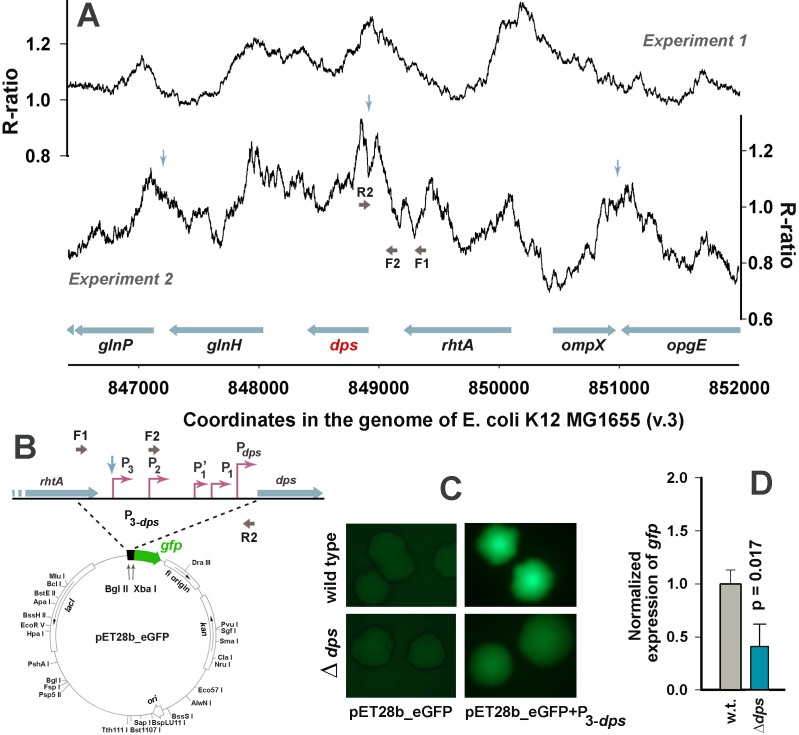
Fig 6. qRT-PCR and reporter assays revealed apparent positive autoregulation of the dps gene.
A: Profiles of the Dps binding sites obtained in two experiments for the dps genomic locus (running window of nine 35 bp bins). Genes are displayed as blue horizontal arrows. Vertical arrows show locations of inverted repeats, if longer than 7 bp (blue). Primer pair F1-R2 was used for cloning the whole regulatory region in pET28b_eGFP. The dps_short fragment (positive control in all band-shift assays) was amplified with F2-R2. B: Scheme of the pET28b-eGFP reporter vector with the insert of the dps promoter region. C: Images of E. coli MG1655 cell colonies transformed by pET28b-eGFP plasmid with or without the promoter insert. Images were obtained by fluorescent microscope Leica with 2 or 0.5 sec exposition for control and experimental cells, respectively. D: Changes in the expression efficiency of gfp in response to dps deletion measured in qRT-PCR using kan as a reference gene. Expression levels were estimated based on 3 biological samples with 3 technical repeats in each. Error bars show an average deviation. Statistical significance was assessed using Student’s t-test.
Demonstrating all possible types of similarity or difference between two experiments, the selected profiles confirmed certain correlation of detected peaks with disposition of long inverted repeats and also revealed some periodicity in the pattern of Dps binding. Quasi-regular alternation of the Dps-occupied and free DNA segments can be a specific feature in the mode of its interaction with the genome. Like nucleosomal fazing, this may be important for maintaining optimal conformation of active chromatin and for its rapid remodeling under stress conditions.
Band-shift assays were performed to characterize the ability of Dps to interact with linear DNA fragments representing the Dps-bound (Figs 5B, S4B, S5B and S6B) and unbound (Figs 5 and S5) regions. In all experiments the previously characterized DNA fragment, amplified with primers F2 –R2 from the regulatory region of the dps gene (dps_short) [19], was used as a positive control. Interaction efficiency was estimated based on the amount of DNA remained unbound. As a result, we observed the ability of Dps to interact with DNA fragments taken from the regions bound by Dps in vivo (Figs 5, S4–S6) and also confirmed its different affinity to the linear DNA fragments [19], but we did not find obvious correlation with ChIP-seq data in the efficiency of binding. For instance, from the two sequences unbound in vivo, taken from fis regulatory region (fis_up) and the lacZ coding area, first remained unbound in vitro (S5B Fig), while the second interacted with Dps even stronger than the control fragment “dps_short” (Fig 5B, profile in S3 Fig). Moreover, the DNA fragment from rho genomic loci representing the peak well reproduced in vivo, demonstrated rather weak binding in vitro (S6B Fig). Most likely this low correlation between in vivo and in vitro experiments is due to the ability of Dps to form complexes with partially melted ends of linear fragments [19] that can mask other types of interaction, if any. On the other hand, Dps-DNA binding in the cell might be significantly dependent from other proteins of bacterial nucleoid.
Functional dependence on Dps was detected for 5 out of 7 genes tested by qRT-PCR using dps-null mutant (Figs 5C, S4C–S6C). As expected, removal of Dps had a similar effect on rpoA and rpoB (Fig 5C). However, the level of their expression varied, and for rpoB it was unchanged in the mutant strain in one of 5 biological replicates, which probably reflects observed unstable interaction of Dps with ribosomal operons (Fig 5A). The increase in the amount of rpoA and rpoB mRNAs assumes a negative impact of Dps on their expression, which probably can be extended to ribosomal genes as well. Response of rho was the opposite (S6C Fig). The proteins of transcriptional machinery are, therefore, under the differential control of growth-dependent regulatory networks and it is possible that direct or indirect Dps-mediated stimulation of the ρ-dependent transcription termination is an important step in the process of genome packaging.
The strongest inhibition was detected for fis (S5C Fig). This was quite unexpected, since a stimulatory rather than inhibitory effect of the dps deletion would have been more anticipated if one takes into account the opposite changes in the concentrations of Fis and Dps during the growth of bacteria [4]. The fis gene is transcribed together with the upstream dusB, and the promoter of the latter is strongly inhibited by Fis [70, 71]. If Dps interferes with Fis, then its deletion can strengthen Fis-mediated suppression, explaining the observed inhibitory effect by the relieved competition.
We did not detect any changes in the amount of oppA or oppB mRNAs, but Dps appeared to be required for expression of oppD (S4C Fig). Taken into account operonic organization of this locus, competition with other transcription factors in the region between unaffected oppA—oppB and suppressed oppD seemed unlikely. Though, the analysis of available data sets revealed extended Fis contacts within oppBCD (wavy lines in S4 Fig). Thus, the detected inhibitory response to dps deletion in this case can be also explained by competitive interactions with Fis.
To test the possibility of feedback regulation for dps, the region containing all experimentally verified promoters [72, 73] was incorporated into pET28b-eGFP in front of gfp (Fig 6B). Removal of dps decreased the fluorescence of cells transformed with a resulting plasmid 1.5 fold (from 211.1±21.8 to 140.5±16.7 relative units, Fig 6C). Approximately the same inhibitory response was registered by qRT-PCR (primers gfp_F and gfp_R, S1 Table) for exponentially growing cells (Fig 6D). Promoter region of dps has two sites of primary contacts with Dps revealed by DNAse I footprinting [19]. One of them (65–120 bp upstream of Pdps) fully covers the binding site for activator IHF and is located nearby repressor Fis contact regions (92–104 bp and 19–33 bp from Pdps, respectively [74]). Another one (216–281 bp from Pdps) has a 30 bp inverted repeat in the center (indicated in Fig 6A and 6B). Relieved interference with Fis in the first binding site perfectly explains inhibitory effect of dps deletion (Fig 6D), while lost interaction with inverted repeat can break topological state of the promoter region. The dps regulatory region can, thus, be suggested as a model for further studies of the Dps ability to bind structural elements and interfere with other regulatory proteins.
Discussion
During exponential growth bacterial cells contain at least 6,000 monomers of Dps [3, 75], which are distributed along the entire uncondensed genome [76]. Upon starvation, Dps co-crystallizes with DNA [30] by hypothetical mechanism that does not require any sequence or structural specificity. To understand if Dps has any binding preferences we, therefore, selected cells harvested at exponential rather than stationary phase, although the intracellular concentration of Dps increases dramatically during steady growth [3, 75]. Automatic evaluation by CLC WB revealed several thousand of Dps targets with a total length of about 1,100,000–1,500,000 bp (S2 Table), while the cumulative size of areas occupied by Dps provided by the Matcher-based approach was 90,000–300,000 bp (Pages 1 and 2, S3 Table). In the exponential growing culture there are 500–700 Dps dodecamers per cell. If 60 bp are sufficient for their interaction with DNA (estimated in the titration experiments made in vitro [18]), then only 30,000–42,000 bp can be occupied. If our estimate of ~120 bp (based on the maxima positioning in the size distribution curves (S2B Fig)) reflects the topology of natural binding in the cell, then in 60,000–84,000 bp of the genome can be covered by Dps. These ranges are significantly smaller, than the total size of binding sites revealed by both methods in both experiments. Therefore, to maintain functional integrity of the genome during exponential growth Dps most likely uses different combinations of binding sites in different cells.
At stationary phase there are 7000–15,000 Dps dodecamers per cells, which can occupy 420,000–900,000 bp or 840,000–1,800,000 bp if, respectively, 60 or 120 bp are required for interaction with a single Dps particle. This roughly corresponds to the total length of Dps targets found by CLC GW (1,100,000–1,500,000 bp), suggesting a possibility that they are used as primary anchor sites for the genome packaging. But the question about the size of a single binding site still remains open, because both 60 bp and 120 bp are much longer than the physical size of the Dps dodecamer, ~7 nm [5], and analyzing the available AFM images, we have not found convincing evidence of DNA wrapping on the protein globule.
The large number of contact sites found for Dps is not very surprising, since the nucleoid proteins Fis, IHF, H-NS and even transcription factor FNR (during anaerobic growth) also bind to many locations in the genome, especially in stress conditions (500,000–1,600,00 bp) [46, 47, 49, 51]. This ubiquity assumes a substantial overlap between the contact sites of different proteins, and the crucial role of nucleoid-associated proteins in FNR occupancy at many of its target sites has been discussed [50]. Quantitatively estimating the superposition of Dps contact sites with the targets of three nucleoid proteins, we found the maximum overlap with the sites occupied by Fis. This is also not surprising, as it is believed that condensing the genome during steady growth; Dps replaces Fis that maintains the optimal structural state of the nucleoid during exponential growth. Less expected was an overlap with the binding sites of H-NS, a histone-like protein functioning as a transcription factor, or a structural protein [77, 78]. Unlike other nucleoid proteins, H-NS has an exclusively high specificity to horizontally acquired genes [65, 66] and promoter islands often located nearby [43, 62, 64]. We previously found that deletion of hns significantly activated transcriptional activity of the islands, while deletion of dps caused a differential effect of approximately the same scale as found in this study [56]. Now it became clear that both proteins (Fig 4A for Dps) have a strong affinity to the islands, where their functional interplay can be used to keep expression of foreign genes at the optimal level.
Although the genome has a large number of potential Dps binding sites, their distribution, even at a large scale was not uniform (Fig 1). The largest occupancy was observed for approximately 800,000 bp of the genome, including the origin of replication and 6 of 7 rRNA operons, where the most significant differences between the two ChIP-seq experiments were detected. This bias can be only partially explained by the gradient in the DNA copy number, since using R-ratio between read counts detected in the anti-Dps immunoprecipitated sequence libraries and the control sets, we obtained increase, rather than decrease in this bias for CS (Fig 1B, dashed plot). Thus, it is likely that the presence of a large number of Dps molecules in the last quarter of the genome is biologically meaningful permitting fast remodeling under stress conditions by turning off expression of ribosomal genes or by affecting replication initiation.
Both experiments indicated an interaction of Dps within or nearby oriC (positions 3925744–3925975) (S2 Table), which is important as it is known that Dps can delay replication by direct interaction with DnaA [79]. The genomic regions bound by Dps in this area contain many repeated sequences, including three pairs of inverted heptanucleotides within the origin (underlined below). One pair overlaps with two binding sites for IciA (in bold): AGAGATCTGTTCTATTGTGATCTCTTATTAG (position 3925757). This protein inhibits replication by blocking the DNA opening by DnaA. Four other sequences can form two alternative complementary pairs: AGGATCATTAACTGTGAATGATCGGTGATCCT, or GATCATTAACTGTGAATGATC (position 3925844–46, respectively) overlapping the DnaA binding site (in bold). Thus, it is possible that inverted repeats, forming a 4-way junction create a platform for interaction with the Dps-DnaA submolecular complex.
Enrichment of the Dps targets with inverted repeats (Fig 3) supports a previous suggestion that they are important for binding [19]. A total of 380 peak regions (84%) from CS possess 5–7 bp inverted repeats (Page 3, S3 Table), while in the set of unbound sequences this percentage is only 54.8% (Page 4, S3 Table). The role of inverted repeats in Dps binding is also supported by some correlation between their positioning and the peaks in the Dps binding profiles (Figs 5A, 6A and S3–S6). However, a large number of inverted repeats in the genome limits the possibility of their use for predictive mapping.
For direct repeats, which have the same large-scale profile as inverted repeats (Fig 1C), this ubiquity is even higher, and may explain why CS and UR differ in their presence only for long repeats (Fig 3). Almost all Dps contact regions (94%, Page 3, S3 Table) have tandem repeats, but they are also found in 78.6% of unbound sequences (Pages 4, S3 Table). For all that, one of the Dps primary contacts in the promoter region of its own gene covers the cluster with 5 bp direct repeats [19]. The ability of tandem repeats to facilitate or stabilize interaction with Dps is, therefore, unclear.
A similar conclusion can be reached regarding the role of sequence motifs over-represented in Dps binding sites GATA and TG-tGAT. Their presence can not be ignored, and direct interaction with Dps deserves special study by methods intended for this purpose. Currently it is likely that Dps can bind DNA near the regions occupied by other proteins. Then, revealed sequence motifs may belong to their specific targets. REP-elements can also contribute to the detected enrichment. Providing a platform for the structure-specific interaction they also “contaminate” bound regions with overrepresented motifs (consensus: GCC(g/t)GATG-CG(a/g)CG(t/c)——(g/a)CG(c/t)CTTATC(c/a)GGCCTAC, inverted repeats are underlined).
The fact that 6 of 8 genes with tested expression efficiency responded to the removal of dps, suggests that Dps can function as a transcription factor. This feature has not been previously described for this protein, though in many cases a “regulatory” effect can still be explained by changed availability of the promoter regions for transcriptional activators and/or inhibitors. Even specifically reduced amount of mRNA transcribed from intraoperonic oppD can be explained by the canceled competition for Fis in the dps-null mutant (S4 Fig). Whatever the mechanism of registered changes is, it became clear that variations in the intracellular Dps concentration may modulate the profile of mRNAs transcribed in exponentially growing cells. By now it has only been detected for the profiles of cellular proteins upon starvation [1].
The transcriptional activity of the dps promoters appeared to be also suppressed in the dps-null mutant (Fig 6D). Even though this suppression is most probably mediated by relieved interference with any of three regulatory proteins—Fis, H-NS and MntR-Mn2+ able to repress the dps promoter activity [80, 81], the data obtained indicate positive dependence of the dps expression on the cellular concentration of its protein product. Reporter assays revealed the same dependence for steady growing bacteria in cell colonies (Fig 6C). They strongly support the qRT-PCR data, though do not indicate that positive feedback also takes place during steady growth, because registered fluorescence can arise from GFP accommodated in cell colonies before they reached the stationary phase. It is clear, that positive autoregulation, either direct or competition-mediated, is favorable for the rapid genome packaging in case of emergency, but some cellular components are required to stop it when necessary and to remove Dps from the condensed chromosome. Some nutrients can fulfill this role during the exit from starvation. We recently found, for instance, that D-galacturonate and D-glucuronate, but not glucose, are capable of altering the oligomeric form of Dps [82], which may be crucial for remodeling. In any case, it became clear that interacting with genomic DNA Dps can affect expression of some genes. In most cases, including apparent positive autoregulation, this influence can be attributed to Dps competition with Fis. Functional interplay of these two proteins, which share common binding sites (Fig 4) and substitute each other depending on the environmental conditions, on promoters of genes from Fis regulon became thus of particular interest.
Supporting information
A: Correlation between the number of reads from immunoprecipitated and control libraries calculated for 100 bp bins (second experiment). All points, corresponding to rRNAs operons are marked in red. Blue dots correspond to the regions containing bins with R>1.4 in the first experiment (Fig 2A). Dashed line shows bisectrix of the plot. B: Distribution of the Dps binding sites the E. coli MG1655 genome. Two outer circles represent the gene map of the top and the bottom strands of the E. coli MG1655 genome. The red ticks on the third circle mark positions of rRNA operons. The profile of R-values and the distribution of reads registered in the control library were calculated for a 25 bp running window and plotted on the fourth and the fifth circles, respectively.
(TIF)
Histograms demonstrating the size-dependent distribution of the Dps-binding sites, revealed in two ChIP-seq experiments by CLC GW (A) and the Matcher (B) (blue and green plots, respectively). Solid and dashed lines correspond to the peaks found with default and stringent settings, respectively. The magenta plot in panel B corresponds to CS.
(TIF)
Profiles of the Dps binding sites obtained in the two experiments (indicated) for the genomic region containing two operons of genes encoding ribosomal proteins (A) and operons of metabolic genes (B) (running window of nine 35 bp bins). Genes are shown by blue horizontal arrows, operons are indicated by magenta lines. Vertical arrows mark locations of inverted repeats (if longer than 7 bp). Positioning of primers used for amplification (F and R) and qRT-PCR (RT and PCR) are indicated.
(TIF)
A: Profiles of the Dps binding sites obtained in the two experiments (indicated) for the genomic region associated with the oppABCDF operon (running window of nine 35 bp bins). Disposition of genes is indicated by blue horizontal arrows. Genes transcribed as a polycistronic unit are underlined by magenta line. Vertical arrows mark locations of inverted repeats (if longer than 7 bp). Wavy lines indicate sites bound by H-NS, Fis and IHF in the data sets obtained in [46, 47, 49–51] and used for intersection analysis (Fig 4). Positioning of primers used for amplification (F and R) and qRT-PCR (RT and PCR) are indicated. B: Band shift assays performed for indicated DNA fragments. C: Dependence of expression efficiency in response to dps deletion. Expression levels were estimated based on 5, 7 and 6 biological samples (3 technical repeats in each) for oppA, oppB and oppD, respectively. Error bars show an average deviation. Statistical significance was assessed using Student’s t-test.
(TIF)
A: Profiles of the Dps binding sites obtained in the two experiments (indicated) for the genomic region associated with the dusB-fis operon (running window of nine 35 bp bins). Disposition of genes is indicated by blue horizontal arrows. Genes transcribed as a polycistronic unit are underlined by magenta line. Vertical arrows mark locations of inverted repeats (if longer than 7 bp). Positioning of primers used for amplification (F and R) and qRT-PCR (RT and PCR) are indicated. B: Band shift assays, performed for indicated DNA fragments. C: Dependence of expression efficiency of fis in response to dps deletion. Expression levels were estimated based on 4 biological samples with 3 technical repeats in each. Error bars show an average deviation. Statistical significance was assessed using Student’s t-test.
(TIF)
A: Profiles of the Dps binding sites obtained in two experiments (indicated) for the genomic region associated with the rho gene. Disposition of genes is indicated by blue horizontal arrows. Genes transcribed as a polycistronic unit are underlined by magenta line. Vertical arrows mark locations of inverted repeats (if longer than 7 bp). Positioning of primers used for amplification (F and R) and qRT-PCR (RT and PCR) are indicated. B: Band-shift assays, performed for the indicated DNA fragments. C: Dependence of expression efficiency of rho in response to dps deletion. Expression levels were estimated based on 3 biological samples with 6 technical repeats in each. Error bars show an average deviation. Statistical significance was assessed using Student’s t-test.
(TIF)
(DOC)
(XLSX)
(XLSX)
Acknowledgments
We thank M.D. Logacheva for cooperation in the whole-genome sequencing of dps-null mutant and S.S. Kiselev for extracting the direct and inverted repeats from the genome of E. coli, as well as for assistance in their preliminary evaluation.
Data Availability
The data discussed in this publication have been deposited in NCBI's Gene Expression Omnibus [42] and are accessible through GEO Series accession number GSE102091 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE102091).
Funding Statement
This work was funded by: 1. Russian Scientific Foundation (grant 14-14-00985, http://rscf.ru/node/8, PI: ONO, team members: SSA, MNT, USS (sample preparation, MiSeq sequencing, data analysis and evaluation)); 2. Russian Foundation for basic research (grant 16-34-01044, http://www.rfbr.ru/rffi/eng, PI: USS, reporter assays); 3. HHMI International Early Career Scientist Program (55007424, FAK) (http://www.hhmi.org/, FAK, HiSeq Illumina sequencing); 4. European Research Council grant agreement (335980_EinME, https://erc.europa.eu/, FAK, HiSeq Illumina sequencing); 5. Russian Foundation for Basic Research 16-04-01570 (http://www.rfbr.ru/rffi/eng) (PI ONO). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1.Almirón M, Link AJ, Furlong D, Kolter R. A novel DNA-binding protein with regulatory and protective roles in starved Escherichia coli. Genes Dev. 1992;6: 2646–2654. Available: http://www.ncbi.nlm.nih.gov/pubmed/1340475. [DOI] [PubMed] [Google Scholar]
- 2.Talukder AA, Ishihama A. Dps is a stationary phase—specific protein of Escherichia coli nucleoid. Adv Microbiol. 2014;4: 1095–1104. doi: doi: 10.4236/aim.2014.415120 [Google Scholar]
- 3.Azam TA, Iwata A, Nishimura A, Ueda S, Ishihama A. Growth phase-dependent variation in protein composition of the Escherichia coli nucleoid. J Bacteriol. 1999;181: 6361–6370. Available: http://www.ncbi.nlm.nih.gov/pmc/articles/PMC103771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Ishihama A, Kori A, Koshio E, Yamada K, Maeda H, Shimada T, et al. Intracellular concentrations of 65 species of transcription factors with known regulatory functions in Escherichia coli. J Bacteriol. 2014;196: 2718–2727. doi: 10.1128/JB.01579-14 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Grant RA, Filman DJ, Finkel SE, Kolter R, Hogle JM. The crystal structure of Dps, a ferritin homolog that binds and protects DNA. Nat Struct Biol. 1998;5: 294–303. doi: 10.1038/nsb0498-294 [DOI] [PubMed] [Google Scholar]
- 6.Jeong KC, Hung KF, Baumler DJ, Byrd JJ, Kaspar CW. Acid stress damage of DNA is prevented by Dps binding in Escherichia coli O157:H7. BMC Microbiology. 2008;8: 181–193. doi: 10.1186/1471-2180-8-181 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Martinez A, Kolter R. Protection of DNA during oxidative stress by the nonspecific DNA-binding protein Dps. J Bacteriol. 1997;179: 5188–5194. Available: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC179379/. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Gundlach J, Winter J. Evolution of Escherichia coli for maximum HOCl resistance through constitutive expression of the OxyR regulon. Microbiology. 2014;160: 1690–1704. doi: 10.1099/mic.0.074815-0 [DOI] [PubMed] [Google Scholar]
- 9.Nair S, Finkel SE. Dps protects cells against multiple stresses during stationary phase. J. Bacteriol. 2004;186: 4192–4198. doi: 10.1128/JB.186.13.4192-4198.2004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Algu K, Choi VSC, Dhami RS, Duncan DAK. Dps confers protection of DNA sequence integrity in UV-irradiated Escherichia coli. J Exp Microbiol Immunol. 2007;11: 60–65. https://www.microbiology.ubc.ca/sites/default/files/roles/drupal_ungrad/JEMI/11/11-60.pdf. [Google Scholar]
- 11.Calhoun LN, Liyanage R, Lay JO Jr, Kwon YM. Proteomic analysis of Salmonella enterica serovar Enteriditis following propionate adaptation. BMC Microbiology. 2010;10: 249–259. doi: 10.1186/1471-2180-10-249 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Theoret JR, Cooper KK, Zekarias B, Roland KL, Law BF, Curtiss III R, et al. The Campylobacter jejuni Dps homologue is important for in vitro biofilm formation and cecal colonization of poultry and may serve as a protective antigen for vaccination. Clin Vaccine Immunol. 2012;19: 1426–1431. doi: 10.1128/CVI.00151-12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Lacqua A, Wanner O, Colangelo T, Martinotti MG, Landini P. Emergence of biofilm-forming subpopulations upon exposure of Escherichia coli to environmental bacteriophages. Appl Environ Microbiol. 2006;72: 956–959. doi: 10.1128/AEM.72.1.956-959.2006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Karas VO, Westerlaken I, Meyer AS. The DNA-binding protein from starved cells (Dps) utilizes dual functions to defend cells against multiple stresses. J Bacteriol. 2015;197: 3206–3215. doi: 10.1128/JB.00475-15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Zhao G, Ceci P, Ilari A, Giangiacomo L, Laue T, Chiancone E, et al. Iron and hydrogen peroxide detoxification properties of DNA-binding protein from starved cells. A ferritin-like DNA-binding protein of Escherichia coli. J Biol Chem. 2002;277: 27689–27696. doi: 10.1074/jbc.M202094200 [DOI] [PubMed] [Google Scholar]
- 16.Zeth K. Dps biomineralizing proteins: multifunctional architects of nature. Biochem J. 2012;445: 297–311. doi: 10.1042/BJ20120514 [DOI] [PubMed] [Google Scholar]
- 17.Ilari A, Stefanini S, Chiancone E, Tsermoglou D. The dodecameric ferritin from Listeria innocua contains a novel intersubunit iron-binding site. Nat Struct Biol. 2000;7: 38–43. doi: 10.1038/71236 [DOI] [PubMed] [Google Scholar]
- 18.Ceci P, Cellai S, Falvo E, Rivetti C, Rossi GL, Chiancone E. DNA condensation and self-aggregation of Escherichia coli Dps are coupled phenomena related to the properties of the N-terminus. Nucl Acids Res. 2004;32: 5935–5944. doi: 10.1093/nar/gkh915 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Melekhov VV, Shvyreva US, Timchenko AA, Tutukina MN, Preobrazhenskaya EV, Burkova DV, et al. Modes of Escherichia coli Dps interaction with DNA as revealed by atomic force microscopy. PLoS ONE. 2015;10: e01265041 doi: 10.1371/journal.pone.0126504 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Bozzi M, Mignogna G, Stefanini S, Barra D, Longhi C, Valenti P, et al. A novel non-heme iron-binding ferritin related to the DNA-binding proteins of the Dps family in Listeria innocua. J Biol Chem. 1997;272: 3259–3265. doi: 10.1074/jbc.272.6.3259 [DOI] [PubMed] [Google Scholar]
- 21.Papinutto E, Dundon WG, Pitulis N, Battistutta R, Montecucco C, Zanotti G. Structure of two iron-binding proteins from Bacillus anthracis. J Biol Chem. 2002;277: 15093–15098. doi: 10.1074/jbc.M112378200 [DOI] [PubMed] [Google Scholar]
- 22.Ceci P, Mangiarotti L, Rivetti C, Chiancone E. The neutrophil-activating Dps protein of Helicobacter pylori, HP-NAP, adopts a mechanism different from Escherichia coli Dps to bind and condense DNA. Nucl Acids Res. 2007;35: 2247–2256. doi: 10.1093/nar/gkm077 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Stillman TJ, Upadhyay M., Norte VA, Sedelnikova SE, Carradus M, Tzokov S, et al. The crystal structures of Lactococcus lactis MG1363 Dps proteins reveal the presence of an N-terminal helix that is required for DNA binding. Mol Microbiol. 2005;57: 1101–1112. doi: 10.1111/j.1365-2958.2005.04757.x [DOI] [PubMed] [Google Scholar]
- 24.Gupta S, Chatterji D. Bimodal protection of DNA by Mycobacterium smegmatis DNA-binding protein from stationary phase cells. J Biol Chem. 2003;278: 5235–5241. doi: 10.1074/jbc.M208825200 [DOI] [PubMed] [Google Scholar]
- 25.Roy S, Saraswathi R, Chatterji D, Vijayan M. Structural studies on the second Mycobacterium smegmatis Dps: invariant and variable features of structure, assembly and function. J Mol Biol. 2008;375: 948–959. doi: 10.1016/j.jmb.2007.10.023 [DOI] [PubMed] [Google Scholar]
- 26.Chowdhury RP, Saraswathi R, Chatterji D. Mycobacterial stress regulation: The Dps ‘‘twin sister” defense mechanism and structure-function relationship. IUBMB Life. 2010;62: 67–77. doi: 10.1002/iub.285 [DOI] [PubMed] [Google Scholar]
- 27.Saraswathi R, Chowdhury RP, Williams SM, Ghatak P, Chatterji D. The mycobacterial MsDps2 protein is a nucleoid-forming DNA binding protein regulated by sigma factors sA and sB. PLoS One. 2009;4: e8017 doi: 10.1371/journal.pone.0008017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Ghatak P, Karmakar K, Kasetty S, Chatterji D. Unveiling the role of Dps in the organization of mycobacterial nucleoid. PLoS One. 2011;6: e16019 doi: 10.1371/journal.pone.0016019 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Wolf SG, Frenkiel D, Arad T, Finkel SE, Kolter R, Minsky A. DNA protection by stress-induced biocristallization. Nature. 1999;400: 83–85. doi: 10.1038/21918 [DOI] [PubMed] [Google Scholar]
- 30.Frenkiel-Krispin D, Levin-Zaidman S, Shimoni E, Wolf SG, Wachtel EJ, Arad T. Regulated phase transitions of bacterial chromatin: a non-enzymatic pathway for generic DNA protection. EMBO J. 2001;20: 1184–1191. doi: 10.1093/emboj/20.5.1184 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Frenkiel-Krispin D, Ben-Avraham I, Englander J, Shimoni E, Wolf SG, Minsky A. Nucleoid restructuring in stationary-state bacteria. Mol Microbiol. 2004;51: 395–405. doi: 10.1046/j.1365-2958.2003.03855.x [DOI] [PubMed] [Google Scholar]
- 32.Kim J, Yoshimura SH, Hizume K, Ohniwa RL, Ishihama A, Takeyasu K. Fundamental structural units of the Escherichia coli nucleoid revealed by atomic force microscopy. Nucl Acids Res. 2004;32: 1982–1992. doi: 10.1093/nar/gkh512 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Ohniwa RL, Morikawa K, Kim J, Ohta T, Ishihama A, Wada C, et al. Dynamic state of DNA topology is essential for genome condensation in bacteria. EMBO J. 2006;25; 5591–5602. doi: 10.1038/sj.emboj.7601414 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Morikawa K, Ohniwa RL, Kim J, Maruyama A, Ohta T, Takeyasu K. Bacterial nucleoid dynamics: oxidative stress response in Staphylococcus aureus. Genes Cells. 2006;11: 409–423. doi: 10.1111/j.1365-2443.2006.00949.x [DOI] [PubMed] [Google Scholar]
- 35.Ishihama A, Shimada T, Yamazaki Y. Transcription profile of Escherichia coli: Genomic SELEX search for regulation targets of transcription factors. Nucl Acids Res. 2016;44: 2058–2074. doi: 10.1093/nar/gkw051 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Datsenko KA, Wanner BL. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci USA. 2000;97: 6640–6645. doi: 10.1073/pnas.120163297 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Thomason LC, Constantino N, Court DL. E. coli genome manipulation by P1 transduction. Curr Protoc Mol Biol Chapter 1:Unit 1.17. 2007. doi: 10.1002/0471142727.mb0117s79 [DOI] [PubMed] [Google Scholar]
- 38.Igarashi K, Ishihama A. Bipartite functional map of the E. coli RNA polymerase alpha subunit: involvement of the C-terminal region in transcription activation by cAMP-CRP. Cell. 1991;65: 1015–1022. doi: 10.1016/0092-8674(91)90553-B [DOI] [PubMed] [Google Scholar]
- 39.Oppermann M. Anion exchange chromatography for purification of monoclonal IgG antibodies In: Peters JP, Baumgarten H (Eds), Monoclonal antibodies. 271–275, Springer-Verlag Berlin Heidelberg; (1992). [Google Scholar]
- 40.Singh SS, Grainger DC. H-NS Can Facilitate Specific DNA-binding by RNA Polymerase in AT-rich Gene Regulatory Regions. PLoS Genet. 2013;9: e1003589 doi: 10.1371/journal.pgen.1003589 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Afgan E, Baker D, van den Beek M, Blankenberg D, Bouvier D, Čech M, et al. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2016 update. Nucl Acids Res. 2016;44: W3–10. doi: 10.1093/nar/gkw343 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Edgar R, Domrachev M, Lash AE. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucl Acids Res. 2002;30: 207–210. doi: https://doi.org/10.1093/nar/30.1.207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Panyukov VV, Kiselev SS, Shavkunov KS, Masulis IS, Ozoline ON. Mixed promoter islands as genomic regions with specific structural and functional properties. Mathem Biol Bioinf. 2013;8: t12–t26. doi: 10.17537/2013.8.432 [Google Scholar]
- 44.Affymetrix I Statistical Algorithms Description Document. 2002, http://www.affymetrix.com/support/technical/whitepapers.affx.
- 45.Aleksic J, Carl S, Frye1 M. Beyond library size: a field guide to NGS normalization. bioRxiv preprint first posted online. 2014; doi: http://dx.doi.org/10.1101/006403.
- 46.Grainger DC, Hurd D, Goldberg MD, Busby SJW. Association of nucleoid proteins with coding and non-coding segments of the Escherichia coli genome. Nucl Acids Res. 2006;34: 4642–4652. doi: 10.1093/nar/gkl542 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Grainger DC, Aiba H, Hurd D, Browning DF, Busby SJW. Transcription factor distribution in Escherichia coli: studies with FNR protein. Nucl Acids Res. 2007;35: 269–278. doi: 10.1093/nar/gkl1023 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Vora T, Hottes AK, Tavazoie S. Protein Occupancy Landscape of a Bacterial Genome. Mol Cell. 2009;35: 247–253. doi: 10.1016/j.molcel.2009.06.035 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Kahramanoglou C, Seshasayee ASN, Prieto AI, Ibberson D, Schmidt S, Zimmermann J et al. Direct and indirect effects of H-NS and Fis on global gene expression control in Escherichia coli. Nucl Acids Res. 2011;39: 2073–2091. doi: 10.1093/nar/gkq934 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Myers KS, Yan H, Ong IM, Chung D, Liang K, Tran F et al. Genome-scale Analysis of Escherichia coli FNR Reveals Complex Features of Transcription Factor Binding. PLoS Genet. 2013;9: e1003565 doi: 10.1371/journal.pgen.1003565 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Prieto AI, Kahramanoglou C, Ali RM, Fraser GM, Seshasayee ASN, Luscombe NM. Genomic analysis of DNA binding and gene regulation by homologous nucleoid-associated proteins IHF and HU in Escherichia coli K12. Nucl Acids Res. 2012;40: 3524–3537 doi: 10.1093/nar/gkr1236 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Shavkunov KS, Masulis IS, Tutukina MN, Deev AA, Ozoline ON. Gains and unexpected lessons from genome-scale promoter mapping. Nucl Acids Res. 2009; 37: 4919–4931. doi: 10.1093/nar/gkp490 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Okonechnikov K, Golosova O, Fursov M, UGENE team. Unipro UGENE: a unified bioinformatics toolkit. Bioinformatics. 2012;28: 1166–1167. doi: 10.1093/bioinformatics/bts091 [DOI] [PubMed] [Google Scholar]
- 54.Bailey TL, Bodén M, Buske FA, Frith M, Grant CE, Clementi L et al. MEME SUITE: tools for motif discovery and searching, Nucl Acids Res. 2009;37: W202–W208. doi: 10.1093/nar/gkp335 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Ma Q, Zhang H, Mao X, Zhou C, Liu B, Chen X et al. DMINDA: an integrated web server for DNA motif analyses. Nucl Acids Res. 2014; 42: W12–W19. doi: 10.1093/nar/gku315 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Purtov YA, Glazunova OA, Antipov SS, Pokusaeva VO, Fesenko EE, Preobrazhenskaya EV et al. Promoter islands as a platform for interaction with nucleoid proteins and transcription factors. J Bioinf Comput Biol. 2013;12: 1441006 doi: 10.1142/S0219720014410066 [DOI] [PubMed] [Google Scholar]
- 57.Masulis IS., Babaeva ZS, Chernyshov SV, Ozoline ON. Visualizing the activity of Escherichia coli divergent promoters and probing their dependence on superhelical density using dual-colour fluorescent reporter vector. Sci Rep. 2015;5: 11449, doi: 10.1038/srep11449 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Schneider CA, Rasband WS, Eliceiri KW. NIH Image to ImageJ: 25 years of image analysis. Nat meth. 2012;9: 671–675. doi: 10.1038/nmeth.2089 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Ivanova D, Taylor T, Smith SL, Dimude JU, Upton A, Mehrjouy MM et al. Shaping the landscape of the Escherichia coli chromosome: replication-transcription encounters in cells with an ectopic replication origin. Nucl Acids Res. 2015;43: 7865–7877. doi: 10.1093/nar/gkv704 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Golosova O, Henderson R, Vaskin Y, Gabrielian A, Grekhov G, Nagarajan V et al. Unipro UGENE NGS pipelines and components for variant calling, RNA-seq and ChIP-seq data analyses. Peer J. 2014;2: e644 doi: 10.7717/peerj.644 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Stern MJ, Ames GFL., Smith NH, Robinson EC, Higgins CF. Repetitive extragenic palindromic sequences: a major component of the bacterial genome. Cell. 1984;37: 1015–1026. doi: 10.1016/0092-8674(84)90436-7 [DOI] [PubMed] [Google Scholar]
- 62.Glazunova OA, Kiselev SS, Shavkunov KS, Bykov AA, Panyukov VV, Ozoline ON. Promoter islands in the genome of E. coli: comparative analysis against AT-rich sequences. Mat. Biolog. Bioinform. 2015;10: t29–t38. doi: 10.17537/2015.10.t29 [Google Scholar]
- 63.Kanehisa M, Sato Y, Kawashima M, Furumichi M, Tanabe M. KEGG as a reference resource for gene and protein annotation. Nucl Acids Res. 2016;44: D457–D462. doi: 10.1093/nar/gkv1070 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Panyukov VV, Ozoline ON. Promoters of Escherichia coli versus promoter islands: function and structure comparison. PLoS ONE, 2013;8: e62601 doi: 10.1371/journal.pone.0062601 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Lucchini S, Rowley G, Goldberg MD, Hurd D, Harrison M, Hinton JCD. H-NS mediates the silencing of laterally acquired genes in bacteria. PLoS Pathog. 2006;2: e81 doi: 10.1371/journal.ppat.0020081 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Dorman CJ. H-NS, the genome sentinel. Nat Rev Microbiol. 2007;5: 157–161. doi: 10.1038/nrmicro1598 [DOI] [PubMed] [Google Scholar]
- 67.Machanick P, Bailey TL. MEME-ChIP: motif analysis of large DNA datasets. Bioinformatics. 2011;27: 1696–1697. doi: 10.1093/bioinformatics/btr189 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Münch R, Hiller K, Grote A, Scheer M, Klein J, Schobert M, et al. Virtual Footprint and PRODORIC: an integrative framework for regulon prediction in prokaryotes. Bioinformatics. 2005;21: 4187–4189. doi: 10.1093/bioinformatics/bti635 [DOI] [PubMed] [Google Scholar]
- 69.Chang D-E, Smalley DJ, Conway T. Gene expression profiling of Escherichia coli growth transitions: an expanded stringent response model. Mol Microbiol. 2002;45: 289–306. doi: 10.1046/j.1365-2958.2002.03001.x [DOI] [PubMed] [Google Scholar]
- 70.Ninnemann O, Koch C, Kahmann R. The E. coli fis promoter is subject to stringent control and autoregulation. EMBO J. 1992;11: 1075–1083. Available at: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC556548/. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Pratt TS, Steiner T, Feldman LS, Walker KA, Osuna R. Deletion analysis of the fis promoter region in Escherichia coli: antagonistic effects of integration host factor and Fis. J Bacteriol. 1997;179: 6367–6377. doi: 10.1128/jb.179.20.6367-6377.1997 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Altuvia S, Almirón M, Huisman G, Kolter R, Storz G. The dps promoter is activated by OxyR during growth and by IHF and sigma S in stationary phase. Mol Microbiol. 1994;13: 265–272. doi: 10.1111/j.1365-2958.1994.tb00421.x [DOI] [PubMed] [Google Scholar]
- 73.Shvyreva US, Tutukina MN, Ozoline ON. Bacterioferritin: properties, structural and functional organization of the dps gene regulatory region. Biophysics. 2011;56: 795–802. doi: 10.1134/S0006350911050204 [PubMed] [Google Scholar]
- 74.Gama-Castro S, Salgado H, Peralta-Gil M, Santos-Zavaleta A, Muniz-Rascado L, Solano-Lira H, et al. RegulonDB version 7.0: transcriptional regulation of Escherichia coli K-12 integrated within genetic sensory response units (Gensor Units). Nucl Acids Res. 2011;39: D98–D105. doi: 10.1093/nar/gkq1110 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Azam TA, Ishihama A. Twelve species of the nucleoid-associated protein from Escherichia coli. Sequence recognition specificity and DNA binding affinity. J Biol Chem. 1999;274: 33105–33113. doi: 10.1074/jbc.274.46.33105 [DOI] [PubMed] [Google Scholar]
- 76.Azam TA, Hiraga S, Ishihama A. Two types of localization of the DNA-binding proteins within the Escherichia coli nucleoid. Genes Cells. 2000;5: 613–626. doi: 10.1046/j.1365-2443.2000.00350.x [DOI] [PubMed] [Google Scholar]
- 77.Oshima T, Ishikawa S, Kurokawa K, Aiba H, Ogasawara N. Escherichia coli histone-like protein H-NS preferentially binds to horizontally acquired DNA in association with RNA polymerase. DNA Res. 2006;13: 141–153. doi: 10.1093/dnares/dsl009 [DOI] [PubMed] [Google Scholar]
- 78.Wang W, Li GW, Chen C, Xie XS, Zhuang X. Chromosome organization by a nucleoid-associated protein in live bacteria. Science. 2011;333: 1445–1449. doi: 10.1126/science.1204697 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Chodavarapu S, Gomez R, Vicente M, Kaguni JM. Escherichia coli Dps interacts with DnaA protein to impede initiation: a model of adaptive mutation. Mol Microbiol. 2008;67: 1331–1346. doi: 10.1111/j.1365-2958.2008.06127.x [DOI] [PubMed] [Google Scholar]
- 80.Grainger DC, Goldberg MD, Lee DJ, Busby SJW. Selective repression by Fis and H-NS at the Escherichia coli dps promoter. Mol Microbiol. 2008;68: 1366–1377. doi: 10.1111/j.1365-2958.2008.06253.x [DOI] [PubMed] [Google Scholar]
- 81.Yamamoto K, Ishihama A, Busby SJ, Grainger DC. The Escherichia coli K-12 MntR miniregulon includes dps, which encodes the major stationary-phase DNA-binding protein. J Bacteriol. 2011;193: 1477–1480. doi: 10.1128/JB.01230-10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Bessonova TA, Shumeiko SA, Purtov YA, Antipov SS, Preobrazhenskaya EV, Tutukina MN, Ozoline ON. Hexuronates influence the oligomeric form of the Dps structural protein of bacterial nucleoid and its ability to bind to linear DNA fragments. Biophysics. 2016;61: 825–832. doi: 10.1134/S0006350916060075 [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
A: Correlation between the number of reads from immunoprecipitated and control libraries calculated for 100 bp bins (second experiment). All points, corresponding to rRNAs operons are marked in red. Blue dots correspond to the regions containing bins with R>1.4 in the first experiment (Fig 2A). Dashed line shows bisectrix of the plot. B: Distribution of the Dps binding sites the E. coli MG1655 genome. Two outer circles represent the gene map of the top and the bottom strands of the E. coli MG1655 genome. The red ticks on the third circle mark positions of rRNA operons. The profile of R-values and the distribution of reads registered in the control library were calculated for a 25 bp running window and plotted on the fourth and the fifth circles, respectively.
(TIF)
Histograms demonstrating the size-dependent distribution of the Dps-binding sites, revealed in two ChIP-seq experiments by CLC GW (A) and the Matcher (B) (blue and green plots, respectively). Solid and dashed lines correspond to the peaks found with default and stringent settings, respectively. The magenta plot in panel B corresponds to CS.
(TIF)
Profiles of the Dps binding sites obtained in the two experiments (indicated) for the genomic region containing two operons of genes encoding ribosomal proteins (A) and operons of metabolic genes (B) (running window of nine 35 bp bins). Genes are shown by blue horizontal arrows, operons are indicated by magenta lines. Vertical arrows mark locations of inverted repeats (if longer than 7 bp). Positioning of primers used for amplification (F and R) and qRT-PCR (RT and PCR) are indicated.
(TIF)
A: Profiles of the Dps binding sites obtained in the two experiments (indicated) for the genomic region associated with the oppABCDF operon (running window of nine 35 bp bins). Disposition of genes is indicated by blue horizontal arrows. Genes transcribed as a polycistronic unit are underlined by magenta line. Vertical arrows mark locations of inverted repeats (if longer than 7 bp). Wavy lines indicate sites bound by H-NS, Fis and IHF in the data sets obtained in [46, 47, 49–51] and used for intersection analysis (Fig 4). Positioning of primers used for amplification (F and R) and qRT-PCR (RT and PCR) are indicated. B: Band shift assays performed for indicated DNA fragments. C: Dependence of expression efficiency in response to dps deletion. Expression levels were estimated based on 5, 7 and 6 biological samples (3 technical repeats in each) for oppA, oppB and oppD, respectively. Error bars show an average deviation. Statistical significance was assessed using Student’s t-test.
(TIF)
A: Profiles of the Dps binding sites obtained in the two experiments (indicated) for the genomic region associated with the dusB-fis operon (running window of nine 35 bp bins). Disposition of genes is indicated by blue horizontal arrows. Genes transcribed as a polycistronic unit are underlined by magenta line. Vertical arrows mark locations of inverted repeats (if longer than 7 bp). Positioning of primers used for amplification (F and R) and qRT-PCR (RT and PCR) are indicated. B: Band shift assays, performed for indicated DNA fragments. C: Dependence of expression efficiency of fis in response to dps deletion. Expression levels were estimated based on 4 biological samples with 3 technical repeats in each. Error bars show an average deviation. Statistical significance was assessed using Student’s t-test.
(TIF)
A: Profiles of the Dps binding sites obtained in two experiments (indicated) for the genomic region associated with the rho gene. Disposition of genes is indicated by blue horizontal arrows. Genes transcribed as a polycistronic unit are underlined by magenta line. Vertical arrows mark locations of inverted repeats (if longer than 7 bp). Positioning of primers used for amplification (F and R) and qRT-PCR (RT and PCR) are indicated. B: Band-shift assays, performed for the indicated DNA fragments. C: Dependence of expression efficiency of rho in response to dps deletion. Expression levels were estimated based on 3 biological samples with 6 technical repeats in each. Error bars show an average deviation. Statistical significance was assessed using Student’s t-test.
(TIF)
(DOC)
(XLSX)
(XLSX)
Data Availability Statement
The data discussed in this publication have been deposited in NCBI's Gene Expression Omnibus [42] and are accessible through GEO Series accession number GSE102091 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE102091).