Abstract
Many studies have shown that variation in transcription is associated with changes in behavioral state, or with variation within a state, but little has been done to address if the same genes are involved in both. Here we investigate the transcriptional basis of variation in parental provisioning using two species of burying beetle, Nicrophorus orbicollis and Nicrophorus vespilloides. We used RNA-seq to compare transcription in parents that provided high amounts of provisioning behavior versus low amounts in males and females of each species. We found no overarching transcriptional patterns distinguishing high from low caring parents, and no informative transcripts that displayed particularly large expression differences in either sex. However, we did find subtler gene expression differences between high and low provisioning parents that are consistent across both sexes and species. Furthermore, we show that transcripts previously implicated in transitioning into parental care in N. vespilloides had high variance in the levels of transcription and were unusually likely to display differential expression between high and low provisioning parents. Thus, quantitative behavioral variation appears to reflect many transcriptional differences of small effect. Furthermore, the same transcripts required for the transition between behavioral states are also related to variation within a behavioral state.
Keywords: burying beetle, gene set enrichment analysis, Nicrophorus, parental care, quantitative trait transcripts, RNA-seq
Intermediate between a variant allele and the quantitative effect it produces is some variable mechanistic process (Maynard Smith et al. 1985). One such mechanism is variation in gene expression (Albert and Kruglyak 2015), the study of which has been enhanced by the development of transcriptomic methods (Todd et al. 2016) leading to a plethora of evolutionary studies using this technique. However, most evolutionary transcriptomic studies examine the changes in transcription associated with transitions in phenotypic state: transitions from one state to another. In evolution, however, it is variation within a phenotypic state – typically quantitative variation (Lynch and Walsh 1998) – that is exposed to selection and evolves. If variation in transcription is in fact a mediator of phenotypic effects, then two things must be true: allelic variation must influence transcriptional variation, and transcriptional variation must influence phenotypic variation. Through research mapping QTLs for gene expression phenotypes (eQTL), evidence for the first of these is rapidly mounting (Albert and Kruglyak 2015). However, empirical work addressing the latter is less common. Research searching for quantitative trait transcripts (QTT; Passador-Gurgel et al. 2007) in Drosophila (Passador-Gurgel et al. 2007, Ayroles et al. 2009), human disease (Petretto et al. 2008, Lodder et al. 2012), and crops (Yang et al. 2012) has suggested that many genes are transcriptionally linked to phenotype (Mackay et al. 2009). Furthermore, work in Drosophila has suggested that predicting candidate transcripts with phenotypic associations may be relatively difficult (Nuzhdin et al. 2009). However, to date, QTT-like studies have only been performed in a handful of organisms, and none in natural populations, limiting the evolutionary conclusions that can be drawn. Here we begin to address this void by examining how transcriptional variation is related to behavioral variation in a complex quantitative behavior, parental provisioning of food by regurgitation to begging offspring.
Behavioral phenotypes can be among the most complex traits found in animals. Behavior is typically highly quantitative in nature, and is produced variably but reliably in response to environmental conditions (Székely et al. 2010). As such, behaviors are expected to be underpinned by an especially complicated set of genetic influences (Boake et al. 2002). Furthermore, the genetic basis of behavior can be logically split into two processes: first, a process that produces a qualitative change in behavioral state; and second, a process that produces a specific quantitative value of the new behavior. Many studies of transcription and behavior have focused on how transcription changes with changes in behavioral state (reviewed in Harris and Hofmann 2014). For example, in honeybees, differential expression is associated with the transition between nursing behavior and foraging behavior (Ben-Shahar et al. 2002, Whitfield et al. 2003, Grozinger et al. 2007, Ament et al. 2008). Other studies investigate how transcription relates to quantitative variation in behavior (QTT) or behavioral type in Drosophila (Dierick and Greenspan 2006, Edwards et al. 2006), fish (Cummings et al. 2008, Sanogo et al. 2012, Rey et al. 2013, Fraser et al. 2014 Wong et al. 2015, Bell et al. 2016) and honey bees (Alaux et al. 2009), and have ascribed a strong and multigenic transcriptional basis to behavioral variation.
Information linking transcriptional associations with behavioral transitions to transcriptional associations with within-behavior variation is lacking. This requires data on levels of transcription in the same behavior using both types of comparisons. If there is overlap between these two transcriptional mechanisms, it suggests that the production of specific behavioral phenotypes is a single process; transcripts are adjusted to produce behavioral change and the specific level of that adjustment determines the quantitative phenotype. In this scenario, the transcriptional changes required to produce behavioral transitions are loose, only needing to follow a broad pattern. If, on the other hand, there is little or no overlap between these mechanisms, it instead suggests that two distinct processes are needed to produce a specific behavioral phenotype; one tightly controlled process to produce a transition, and a separately regulated process producing the quantitative phenotype. It has been predicted that regulatory mechanisms for different types of behavioral change will be different (Cardoso et al. 2015), though one study has found transcriptional overlap between behavioral transitions and behavioral variation in sticklebacks (Bell et al. 2016).
In this study, we examine how variation in transcription is associated with variation in parental provisioning behavior in burying beetles (Nicrophorus spp.), an insect taxon with unusually well-developed and well-described parenting behavior (Eggert and Müller 1997, Eggert et al. 1998, Scott 1998). Recent studies in N. vespilloides investigated the transcriptional underpinnings of transitions between behavioral states, from solitary to parenting and back to a solitary state, using transcriptomics (Parker et al. 2015, Palmer et al. 2016), candidate gene studies (Cunningham et al 2014, 2016, Roy-Zokan et al. 2015) and proteomics (Cunningham et al. 2017). Parenting and specifically the act of provisioning food by regurgitation to begging offspring is an ideal phenotype for contrasting the mechanisms causing behavioral transitions versus within-state variation because it is quantifiable, predictable, shows continuous variation, and is heritable (Walling et al. 2008, Benowitz et al. 2016).
To examine the prediction that quantitative gene expression is related to the quantitative expression of provisioning behavior in Nicrophorus, we performed RNA-seq comparing gene expression of 20 individuals a priori classified as having high or low levels of provisioning behavior based on extensive behavioral observation (Benowitz et al. 2016). Furthermore, we repeated this in both sexes as both males and females are competent parents, performing separate RNA-seq experiments in males and females of two species, N. vespilloides and N. orbicollis. We broadly expected to find many transcripts associated with variation in provisioning behavior in each of the four samples. Given the strong behavioral (Benowitz et al. 2016) but imperfect genetic (Parker et al. 2015) similarity between sexes, we expected moderate but not complete overlap in the transcripts underlying variation in provisioning between males and females. We predicted we would find even less overlap between species, given the nuanced behavioral differences between them (Benowitz et al. 2016, Benowitz and Moore 2016).
Additionally, following Cardoso et al. (2015), we predicted that the transcripts related to quantitative expression within parenting behavior should be different than those related to transitions between parental and non-parental states in N. vespilloides (Parker et al. 2015). Moreover, given the trait is quantitative and heritable (Walling et al. 2008), we predicted that more moderate differential expression is involved, as there may be many genes of small effect. Our results show that very nuanced changes in gene expression are associated with provisioning behavior and are shared to a degree across sexes and species. Contrary to our predictions, however, we found that the transcripts related to changes between behavioral states are more likely than random to be also related to within-state variation in provisioning behavior.
Materials and Methods
PHENOTYPING PARENTING
We collected Nicrophorus vespilloides and Nicrophorus orbicollis from Cornwall, UK and Georgia, USA, respectively, and maintained outbred laboratory colonies for multiple generations as described in Benowitz et al. (2015). Detailed methods of behavioral experimentation are described in Benowitz et al. (2016). All observations were of uniparental adults that were reared and observed in common and controlled laboratory conditions. Briefly, we made concurrent observations of female N. vespilloides (n = 57) and N. orbicollis (n = 61) during the hours of peak parental care, approximately one day after larval hatching. We scan-sampled each family 80 times over an eight-hour period, recording each time the adult was observed having mouth-to-mouth contact with larvae. We then repeated the experiment at a later date under the same conditions for males of N. vespilloides (n = 79) and N. orbicollis (n = 78). After we finished observations of each parent, we removed their heads, flash-froze the heads in liquid nitrogen, and stored them at -80°C for later transcriptome sequencing.
SAMPLE COLLECTION, PREPARATION, AND RNA SEQUENCING
After we completed all behavioral observations, we collected (flash freezing) the heads of the 10 highest and 10 lowest caring parents from each species and sex for our RNA-seq experiment. Sampling from the extremes of the phenotypic distribution in this way increases the power to detect associations with quantitative traits (Van Gestel et al. 2000). We collected only heads to limit our tissue types to brain and head fat body, the latter being an endocrine organ in insects. We extracted RNA from these 80 heads by first homogenizing them in liquid nitrogen and Qiazol (Qiagen, Venlo, Netherlands), then adding chloroform (Roy-Zokan et al. 2015). After this step, extractions followed standard protocols from a Qiagen RNeasy Lipid kit. RNA libraries were then prepared using a TruSeq Stranded RNA LT Kit (Illumina, San Diego, CA) with one-third of the normal reaction volume (Reynolds et al. 2015). Samples were then sequenced on an Illumina NextSeq Mid Output Flow Cell at the Georgia Genomics Facility (GGF), generating paired-end 75 base pair reads. We first sequenced females of each species together, then both species of males together at a later date.
NICROPHORUS ORBICOLLIS GENOME SEQUENCING AND ASSEMBLY
We assembled the N. orbicollis genome de novo from Illumina sequencing to produce comparable mapping resources for transcriptome analysis. The N. vespilloides genome was recently sequenced, assembled, and annotated (Cunningham et al. 2015). For genome sequencing, we chose a single larva that was the product of two generations of inbreeding. We extracted its DNA using a sodium dodecyl sulfate (SDS) lysis buffer (Chen et al. 2010) and a phenol chloroform extraction. Two libraries, one 350 bp PCR-free library and one 6.5kb mate-pair library were made using Illumina manufacturer’s instructions, and sequenced on an Illumina NextSeq Mid Output Flow Cell at GGF generating paired-end 75 base pair reads. We next trimmed reads for quality using Trimmomatic 0.32 (Bolger et al. 2014), removing 4bp windows with average quality less than 15 and removing reads under 25bp. We assembled the trimmed reads with Platanus (Kajitani et al. 2014) under the same parameters as for N. vespilloides (Cunningham et al. 2015). We then used DeconSeq (Schmieder and Edwards 2011) to remove any possible contaminants from the genome assembly. Genome assembly was 193 mb, with a scaffold N50 of 178.7 kb. BUSCO analysis (Simão et al. 2015) showed the assembly had complete sequence for 82.9% of conserved insect orthologs, and partial sequence for 88.6%.
TRANSCRIPTOME ASSEMBLY AND DIFFERENTIAL EXPRESSION ANALYSIS
We trimmed all RNA reads for quality using Trimmomatic 0.32 as above (Bolger et al. 2014) and mapped them to their respective reference genome using Tophat 2 (Kim et al. 2013). We then assembled reads for each species into transcripts using Cufflinks (Roberts et al. 2011), taking the longest isoform provided. We then remapped reads to the assembled and reduced transcriptome and generated gene level counts using RSEM (Li and Dewey 2011). Descriptive statistics for each assembled transcriptome are provided in Table S1. We then used edgeR (Robinson et al. 2010) to filter transcripts by abundance and perform normalization. We used normalized counts to perform hierarchical clustering analysis using the R package heatmap under default parameters. We then performed differential expression (DE) analysis in edgeR between high and low caring parents. We analyzed each species and sex separately, thus creating four separate DE analyses, assessing significance after multiple testing using FDR (Benjamini and Hochberg 1995) with a cutoff of 0.05 to match a closely related RNA-seq study (Parker et al. 2015). Our DE analysis in N. orbicollis females uncovered three samples with relatively high expression values for some transcripts (see Table S2). We therefore checked to determine if they were outliers by comparing these genes to overall expression profiles. Only the single gene was unusually highly expressed in each sample, and so these individuals were retained in the analysis because their overall expression profiles were not unusual (Fig. S1).
GENE SET ENRICHMENT ANALYSIS
We used Gene Set Enrichment Analysis (GSEA) to examine the similarity between DE transcripts in our four independent datasets. GSEA has the ability to test a predicted set of transcripts across the entire transcriptome, as opposed to similar methods such as hypergeometric statistics that compare the overlap between specified subsets of the transcriptome. Following Subramanian et al. (2005), we selected the top 100 differentially expressed transcripts from one dataset (N. vespilloides females) to serve as a predictive set of transcripts. These 100 transcripts intentionally included both significantly and not significantly DE transcripts, as a major part of the rationale behind our GSEA was to assess the biological significance of genes with low-fold changes (and thus not called significant in edgeR). To examine this set of genes in another dataset, we calculated a running-sum statistic going down the entire list of genes (ordered from most DE to least DE). This statistic increases each time a gene in the set is encountered (weighted by the amount of DE) and decreases by a constant amount each time a gene outside of the set is encountered. The test statistic, or E-score, is the global maximum of this running sum statistic. The significance of the E-score was calculated by comparing it against 10,000 E-scores generated from gene lists with randomized phenotypic permutations. Therefore, this analysis asks whether the predictive gene set displays more DE using the real phenotypic information than random phenotypic information, a conservative GSEA methodology (Subramanian et al. 2005, Ackermann and Strimmer 2009). Directionality of DE was not considered to preserve consistency between real and permuted datasets (which do not contain directionality due to the lack of phenotypic information). For the set of 100 genes taken from N. vespilloides females, we performed this analysis in N. vespilloides males, N. orbicollis females, and N. orbicollis males. We also repeated the same analysis using the top 100 differentially expressed genes from N. orbicollis females. This predictive gene set was analyzed in N. orbicollis males, N. vespilloides females, and N. vespilloides males. We performed all GSEA analysis in R using homemade scripts.
MAPPING AND DIFFERENTIAL EXPRESSION USING PREVIOUSLY PUBLISHED TRANSCRIPTOME
Previous research on N. vespilloides compared gene expression profiles of individuals before they were parents, to individuals that were actively parenting (regurgitating food to begging offspring), to individuals who were no longer parenting (Parker et al. 2015). In order to compare our results directly to those of Parker et al. (2015) we mapped our N. vespilloides RNA-seq reads directly to their published transcriptome using Tophat2 (Kim et al. 2013) and generated read counts using RSEM (Li and Dewey 2011). Gene level counts were then filtered and normalized in edgeR as above. We then specifically examined the 867 (752 after abundance filtering) genes differentially expressed during parenting in N. vespilloides (Parker et al. 2015). First, we examined whether these transcripts displayed greater variability among parenting than non-caring transcripts by comparing the average squared coefficient of variance (CV2) to that of two random transcript sets of equal size and average abundance using analysis of variance. We then performed GSEA using the same methodology as above to examine whether these genes displayed unusual patterns of differential expression between parents with high versus low provisioning phenotypes.
Results and Discussion
BEHAVIORAL PHENOTYPE
The complete description of behavioral variation in both species and both sexes is given in Benowitz et al. (2016). For N. orbicollis, the top 10 females caring for offspring selected for further analysis provided (mean ± SD) 39.8 ± 5.7 bouts of care during the observation period. The lowest 10 N. orbicollis females provided 11.7 ± 4.5, bouts of care during the observation period. For N. vespilloides, the top 10 females caring for offspring provided 72.5 ± 2.5 bouts of care during the observation period. The lowest 10 N. vespilloides females provided 13.6 ± 9.7 bouts of care during the observation period. In males, in N. orbicollis, the top 10 parents provided 33.9 ± 4.4 bouts of care while the lowest 10 provided 11.8 ± 1.3 bouts of care. For N. vespilloides males the top 10 provided 72.9 ± 3.1 bouts of care while the lowest 10 provided 12.6 ± 8.6 bouts of care. These individuals provided the samples for analysis of transcriptional variation.
TRANSCRIPTOME-WIDE ANALYSIS OF PARENTAL VARIATION
We first examined whether an individual’s overall transcriptomic profile predicted its parental provisioning phenotype. Therefore, we used hierarchical clustering to group all 20 samples in each of the four RNA-seq experiments per the similarity of their genome-wide expression profile. In none of our four experiments did samples of similar phenotype group together (Fig. S1–S4), thus, there is no obvious overall transcriptional signature of being a high or low provisioning parent. We next examined two possibilities: that variation in provisioning behavior is either associated with a few genes of large effect or with many genes of small effect.
To test the first of these possibilities, that variation in provisioning is associated with a few large effect transcripts, we performed standard analysis for differential expression (DE) in edgeR. DE analysis turned up a small number of significantly DE transcripts in three of our four RNA-seq experiments but with no obvious patterns (Table S2). Consistent with Parker et al. (2015), more variation in transcription was associated with females but there was no overlap between the species. In males, there was only one transcript that was significantly differentially expressed between high and low provisioning parents N. orbicollis and none in N. vespilloides. Because uniparental provisioning behavior is remarkably similar between sexes at both the phenotypic (Benowitz et al. 2016) and molecular (Parker et al. 2015) levels, we expected the DE transcripts found in males and females of a species to at least partially replicate each other. Thus, there is no suggestion that these transcripts are evolutionarily associated with variation in provisioning behavior.
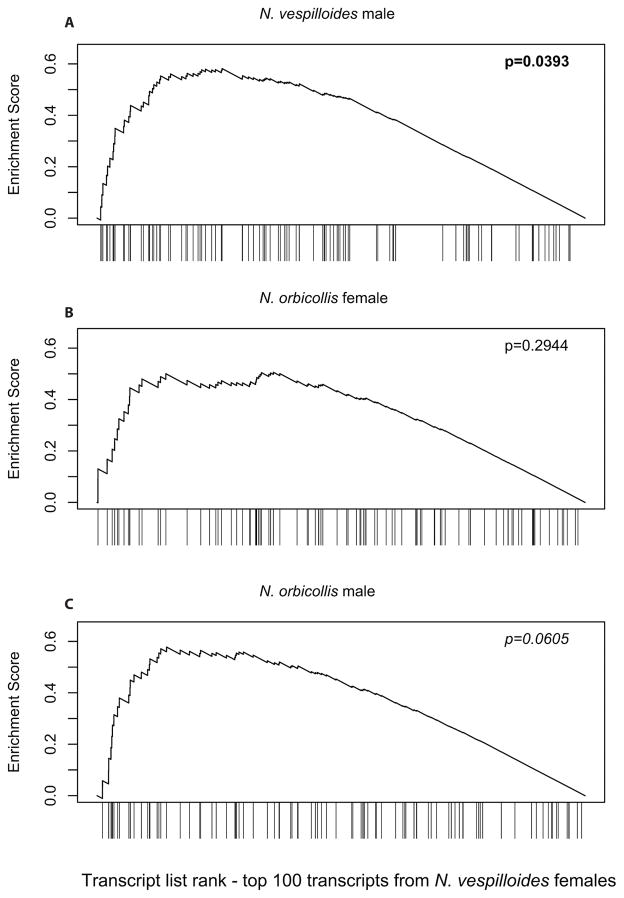
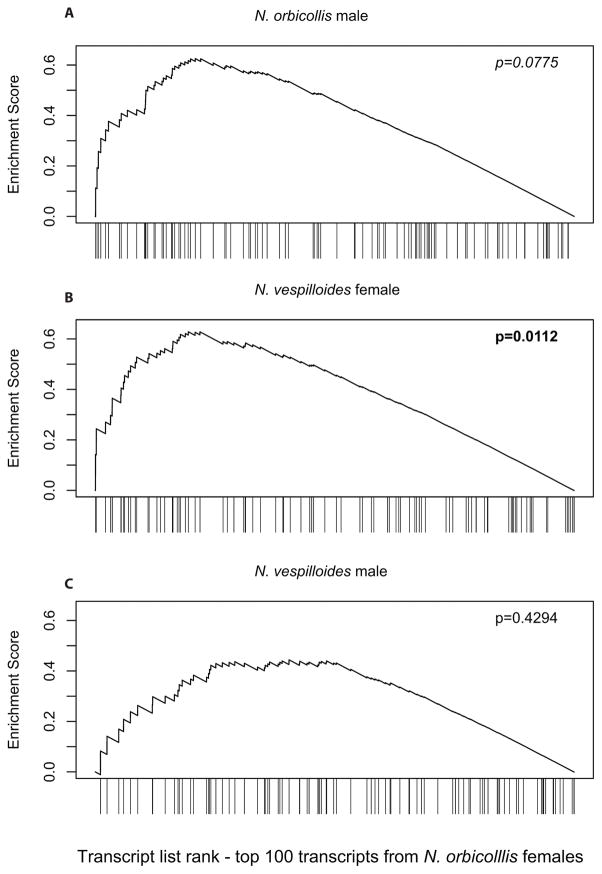
Standard analyses of differential expression, as performed above, often do a poor job of finding transcripts with small expression differences between treatments because of multiple testing across the entire transcriptome (Bullard et al. 2010). Therefore, to test whether such subtle differences might be associated with behavioral phenotype, we asked whether broader patterns of differential expression were similar across the four experiments. To answer this question, we used Gene Set Enrichment Analysis (GSEA; Subramanian et al. 2005) to assess whether the top DE genes from the standard analysis above displayed unusual patterns of DE in the other three RNA-seq datasets. This should only be the case if two things are true: that there are real transcriptional effects associated with provisioning behavior, and that those effects were at least partially shared across sexes and/or species. Therefore, the GSEA tested our hypothesis that even transcripts not considered statistically significant by edgeR might still be biologically related to parenting. Because females have more robust gene expression changes during parenting (Parker et al. 2015) we used the most DE transcripts from females of each species as a set of “predictor” transcripts, and following Subramanian et al. (2005) we a priori restricted the analysis to the top 100 DE transcripts. The top 100 DE transcripts from N. vespilloides females showed a high degree of DE in N. vespilloides males (E = 0.582, p = 0.0393; Fig. 1A), no unusual DE in N. orbicollis females (E = 0.506, p = 0.2944; Fig. 1B), and a moderate degree of DE in N. orbicollis males (E = 0.578, p = 0.0605; Fig. 1C). The top 100 DE transcripts from N. orbicollis females showed a moderate degree of DE in N. orbicollis males (E = 0.625, p = 0.0775; Fig. 2A), significant DE in N. vespilloides females (E = 0.628, p = 0.0112; Fig. 2B), and no unusual DE in N. vespilloides males (E = 0.444, p = 0.4294; Fig. 2C). N. vespilloides males and females display much greater behavioral variation than N. orbicollis parents (Benowitz et al. 2016), which may account for the greater transcriptional similarity we observed within a species.
Figure 1.
Gene set enrichment analysis of transcripts predicted from N. vespilloides females. The top 100 differentially expressed transcripts from N. vespilloides females are (A) significantly DE in N. vespilloides males, (B) not DE in N. orbicollis females, and (C) moderately DE in N. orbicollis males.
Figure 2.
Gene set enrichment analysis of transcripts predicted from N. orbicollis females. The top 100 differentially expressed transcripts from N. orbicollis females are (A) moderately DE in N. orbicollis males, (B) significantly DE in N. vespilloides females, and (C) not DE in N. vespilloides males.
Our data largely recapitulate the results of other QTT-like studies, which typically demonstrate hundreds of transcripts with significant associations with phenotype (Mackay et al. 2009). It is unclear why we saw weak statistical associations between phenotype and individual transcripts, though that may simply be a consequence of the messiness of parental care as a phenotype with low but non-zero heritability (Walling et al. 2008), moderate repeatability (Benowitz et al. 2016), the inability of RNA-seq to measure potentially important classes of transcripts such as neuropeptides (Valcu and Kempenaers 2014), or the possibility that large and causal transcriptional differences might have occurred prior to our sampling time. Despite this, we suggest that the transcriptomic basis of variation in Nicrophorus provisioning behavior is multifaceted yet extremely subtle, with many transcripts showing slight associations with phenotype.
The differential expression of any transcript related to parenting may reflect either environmental plasticity or genetic variation. Our dataset likely includes transcripts of both types, and cannot distinguish between them. However, given that beetles were reared in common and controlled environments, we suggest that plasticity is at most playing a minor role in generating phenotypic and associated transcriptional differences between individuals. Instead, we argue that it is more likely that many of the transcriptional associations we find reflect genetic differences between individuals. Such genetic differences may be causing variation in behavior or may simply be correlated with it, perhaps reflecting variation in energy demands. Again, it is probable that our results capture transcripts of both types. However, either way, our data reflect the idea that even large differences in behavioral phenotype are associated with modest differences in the mechanisms related to that behavior.
These results have implications for the longstanding question of whether phenotypic variation is underpinned by a few genes of large effect or many genes of small effect (Rockman 2012). Though our RNA-seq approach cannot directly answer the question of how many loci are responsible for producing phenotypic variation, it is suggestive of how many variables might be involved. eQTL studies suggest two broad possible explanations for our data. One is that the many expression differences we see could be caused by an eQTL hotspot, a single locus with causal effects on hundreds of transcriptional phenotypes (Schadt et al. 2003, Mehrabian et al. 2005, West et al. 2007). However, hotspots are not a ubiquitous feature of eQTL studies (Monks et al. 2004, Morley et al. 2004, Smith and Kruglyak 2008, Rockman et al. 2010), which are more likely to uncover cis-acting variants affecting individual genes (Gilad et al. 2008). Therefore, we suggest that a more likely possibility is that the transcriptional phenotypes in our study themselves display polygenic inheritance (Petretto et al. 2006).
Furthermore, we find no evidence of any genes whose transcription has major effects on phenotype. Thus, we argue that our results support an infinitesimal model of quantitative inheritance (Fisher 1918, Bulmer 1972), with many genes contributing an individually undetectable effect on phenotypic expression.
DIRECT COMPARISON OF TRANSCRIPTIONAL VARIATION WITHIN AND BETWEEN BEHAVIORS
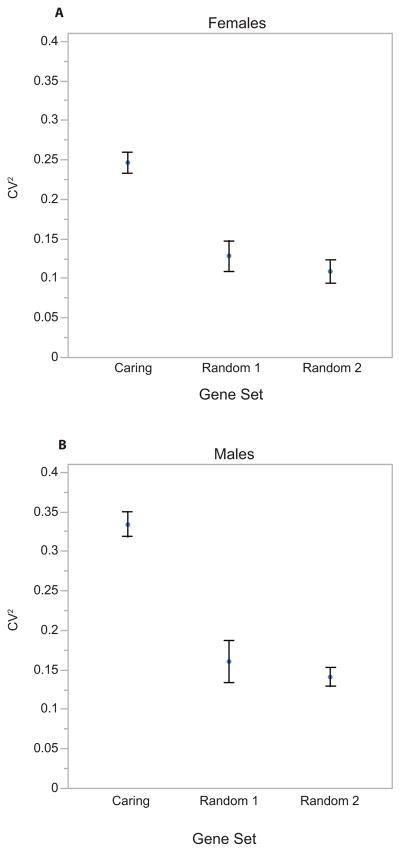
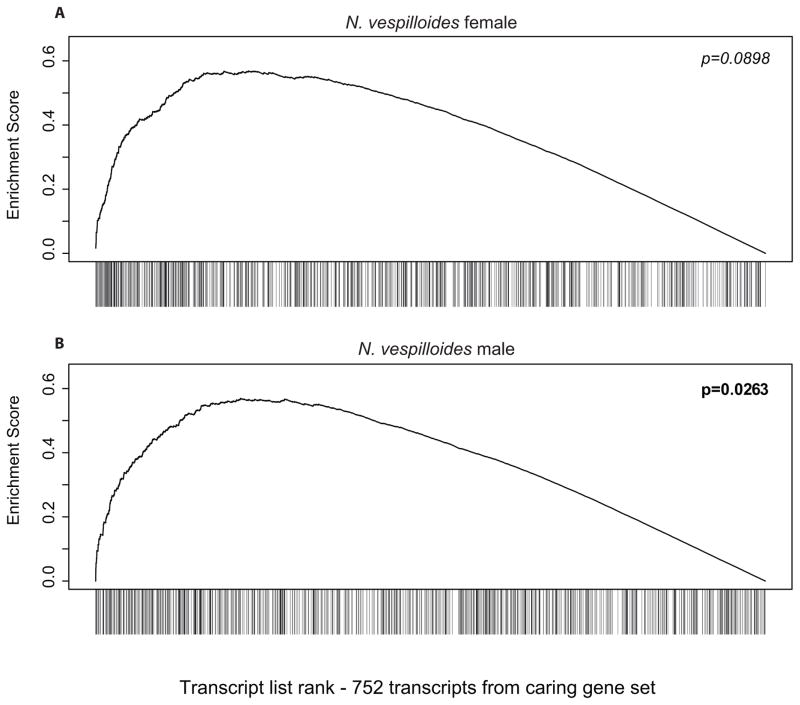
We next asked whether the transcripts involved in transitioning into parental care behavior are also likely to be involved in generating quantitative variance in provisioning behavior. Parker et al. (2015) identified a set of 867 transcripts that were significantly differentially expressed in N. vespilloides between parents and non-parents. Using the published transcriptome from Parker et al. (2015), we asked whether this transcript set displayed unusual patterns of differential expression between high and low provisioning N. vespilloides. First, we examined the variability of these transcripts by comparing the average squared coefficient of variance of transcripts in this set from that of two random transcript sets of equal abundance. We found that, for males (F2,2250 = 31.595, p < 0.0001) as well as females (F2,2250 = 22.337, p < 0.0001), the caring transcript set displays higher variability between samples than random genes (Fig. 3). Next, to see if this high level of variation was related to parental behavior, we used the same GSEA methodology as above. We found that these caring transcripts were significantly DE in males (E = 0.569, p = 0.0263; Fig. 4A) and moderately DE in females (E = 0.570, p = 0.0898; Fig. 4B).
Figure 3.
The variance of transcripts implicated in caring versus random genes. Transcripts differentially expressed between caring and non-caring N. vespilloides (Parker et al. 2015) show higher expression variance among caring parents than random transcripts in both (A) N. vespilloides females and (B) N. vespilloides males.
Figure 4.
Gene set enrichment analysis of transcripts previously implicated in caring. The 752 transcripts found in our data set that were differentially expressed between caring and non-caring N. vespilloides from Parker et al. (2015) are (A) moderately DE in N. vespilloides females, and (B) significantly DE in N. vespilloides males.
Thus, it appears that a subset of the transcripts required to turn on parenting behavior are also modified further in association with quantitative expression of that same behavior. This suggests that parenting fits a model wherein specific behavioral phenotypes are obtained by simply modulating the extent of the initial response creating the behavior. In turn, this supports the idea that transitions between behaviors do not rely on specific transcriptional changes. Instead, it appears that only broad expression changes are required, and that these genes retain a high degree of potential variability after initiation of behavior. This may be further related to the relatively high flexibility of behavior; if the genes involved in producing behaviors are relatively unconstrained, they may be able to produce a broad range of phenotypes in response to environmental variation.
CONCLUSIONS
We present three fundamental insights into the transcriptional architecture of variation in behavior, using parental care in Nicrophorus beetles as our model. First, many transcripts appear to be related to variation in parenting behavior, each with subtle effects. Second, the mechanisms governing variation in parenting share some commonality across both sexes and species, as expected. Third, the transcripts related to transitions into and out of parenting behavior are likely to be related to producing variation within parenting behavior. Together, these results not only provide basic mechanistic information on the construction of specific behavioral phenotypes, but may also lead to insight as to why behavior is uniquely flexible and evolvable.
Supplementary Material
Supplementary Figure 1. Hierarchical clustering of N. orbicollis female samples by transcriptional profile.
Supplementary Figure 2. Hierarchical clustering of N. orbicollis male samples by transcriptional profile.
Supplementary Figure 3. Hierarchical clustering of N. vespilloides female samples by transcriptional profile.
Supplementary Figure 4. Hierarchical clustering of N. vespilloides male samples by transcriptional profile.
Supplementary Table 1. Descriptive statistics for Cufflinks produced transcriptome assemblies for N. orbicollis and N. vespilloides. Means indicate the average of the 20 samples sequenced for each species/sex.
Supplementary Table 2. Transcripts significantly differentially expressed between high and low provisioning parents. A list of the significantly DE transcript IDs from each experiment and their blast homologies. Asterisks next transcripts from N. orbicollis females indicate transcripts whose differential expression is explained by extremely high expression values in the same 3 individuals (B47, B91, and B92), and therefore may bias results for these specific transcripts.
Acknowledgments
B. Hsu and V. Anderson assisted with Nicrophorus colony maintenance. B. Schmitz, D. Neumann, N. Rohr, K. Sandlin, J. Wagner, and M. Alabady assisted with library preparation and Illumina sequencing. D. Hall, K. Ross, T. Linksvayer, A. Wiberg, D. Parker, and M. Ritchie provided helpful discussions and suggestions for statistical analysis. T. Moore, J. Slate, P. Tiffin, T. Flatt, and two anonymous reviewers provided valuable comments on the manuscript. Our research was funded by an NSF grant (IOS-1354358) to A.J.M., and an NIH Training Grant Fellowship (T32GM007103) to K.M.B.
Footnotes
Sequence data associated with this project will be available through NCBI (Bioproject # PRJNA371654). Behavioral data from Benowitz et al. (2016) is available through Dryad (doi:10.5061/dryad.25rm2).
LITERATURE CITED
- Ackermann M, Strimmer K. A general modular framework for gene set enrichment analysis. BMC Bioinf. 2009;10:47. doi: 10.1186/1471-2105-10-47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alaux C, Sinha S, Hasadsri L, Hunt GJ, Guzmán-Novoa E, DeGrandi-Hoffman G, Uribe-Rubio JL, Southey BR, Rodriguez-Zas S, Robinson GE. Honey bee aggression supports a link between gene regulation and behavioral evolution. Proc Natl Acad Sci USA. 2009;106:15400–15405. doi: 10.1073/pnas.0907043106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Albert FW, Kruglyak L. The role of regulatory variation in complex traits and disease. Nat Rev Genet. 2015;16:197–212. doi: 10.1038/nrg3891. [DOI] [PubMed] [Google Scholar]
- Ament SA, Corona M, Pollock HS, Robinson GE. Insulin signaling is involved in the regulation of worker division of labor in honey bee colonies. Proc Natl Acad Sci USA. 2008;105:4226–4231. doi: 10.1073/pnas.0800630105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ayroles JF, Carbone MA, Stone EA, Jordan KW, Lyman RF, Magwire MM, Rollmann SM, Duncan LH, Lawrence F, Anholt RR, et al. Systems genetics of complex traits in Drosophila melanogaster. Nat Genet. 2009;41:299–307. doi: 10.1038/ng.332. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bell AM, Bukhari SA, Sanogo YO. Natural variation in brain gene expression profiles of aggressive and nonaggressive individual sticklebacks. Behaviour. 2016;153:1723–1743. doi: 10.1163/1568539X-00003393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc B. 1995;57:289–300. [Google Scholar]
- Benowitz KM, Moore AJ. Biparental care is predominant and beneficial to parents in the burying beetle Nicrophorus orbicollis (Coleoptera: Silphidae) Biol J Linn Soc. 2016;119:1082–1088. doi: 10.1111/bij.12830. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benowitz KM, Moody KJ, Moore AJ. Are species differences in maternal effects arising from maternal care adaptive? J Evol Biol. 2015;28:503–509. doi: 10.1111/jeb.12573. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benowitz KM, McKinney EC, Moore AJ. Difference in parenting in two species of burying beetle, Nicrophorus orbicollis and Nicrophorus vespilloides. J Ethol. 2016;34:315–319. doi: 10.1007/s10164-016-0477-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ben-Shahar Y, Robichon A, Sokolowski MB, Robinson GE. Influence of gene action across different time scales of behavior. Science. 2002;296:741–744. doi: 10.1126/science.1069911. [DOI] [PubMed] [Google Scholar]
- Boake CRB, Arnold SJ, Breden F, Meffert LM, Ritchie MG, Taylor BJ, Wolf JB, Moore AJ. Genetic tools for studying adaptation and the evolution of behavior. Am Nat. 2002;160:S143–S159. doi: 10.1086/342902. [DOI] [PubMed] [Google Scholar]
- Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30:2114–2120. doi: 10.1093/bioinformatics/btu170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bullard JH, Purdom E, Hansen KD. Evaluation of statistical methods for normalization and differential expression in mRNA-Seq experiments. BMC Bioinf. 2010;11:94. doi: 10.1186/1471-2105-11-94. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bulmer MG. The genetic variability of polygenic characters under optimizing selection, mutation, and drift. Genet Res. 1972;19:17–25. doi: 10.1017/s0016672300014221. [DOI] [PubMed] [Google Scholar]
- Cardoso SD, Teles MC, Oliveira RF. Neurogenomic mechanisms of social plasticity. J Exp Biol. 2015;218:140–149. doi: 10.1242/jeb.106997. [DOI] [PubMed] [Google Scholar]
- Chen H, Rangasamy M, Tan SY, Wang H, Siegfried BD. Evaluation of five methods for total DNA extraction from western corn rootworm beetles. PLoS ONE. 2010;5:e11963. doi: 10.1371/journal.pone.0011963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cummings ME, Larkins-Ford J, Reilly CR, Wong RY, Ramsey M, Hofmann HA. Sexual and social stimuli elicit rapid and contrasting genomic responses. Proc R Soc Lond B. 2008;275:393–402. doi: 10.1098/rspb.2007.1454. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cunningham CB, Douthit MK, Moore AJ. Octopimanergic gene expression and flexible social behaviour in the subsocial burying beetle Nicrophorus vespilloides. Ins Mol Biol. 2014;23:391–404. doi: 10.1111/imb.12090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cunningham CB, Ji L, Wiberg RAW, Shelton J, McKinney EC, Parker DJ, Meagher RB, Benowitz KM, Roy-Zokan EM, Ritchie MG, et al. The genome and methylome of a beetle with complex social behavior, Nicrophorus vespilloides (Coleoptera: Silphidae) Genom Biol Evol. 2015;7:3383–3396. doi: 10.1093/gbe/evv194. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cunningham CB, VanDenHeuvel K, Khana DB, McKinney EC, Moore AJ. The role of neuropeptide F in a transition to parental care. Biol Letts. 2016;12:20160158. doi: 10.1098/rsbl.2016.0158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cunningham CB, Badgett M, Meagher RB, Orlando R, Moore AJ. Ethological principles predict the neuropeptides co-opted to influence parenting. Nat Comm. 2017;8:14225. doi: 10.1038/ncomms14225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dierick HA, Greenspan RJ. Molecular analysis of flies selected for aggressive behavior. Nat Gen. 2006;38:1023–1031. doi: 10.1038/ng1864. [DOI] [PubMed] [Google Scholar]
- Edwards AC, Rollmann SM, Morgan TJ, Mackay TFC. Quantitative genomics of aggressive behavior in Drosophila melanogaster. PLoS Genet. 2006;2:e154. doi: 10.1371/journal.pgen.0020154. https://doi.org/10.1371/journal.pgen.0020154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eggert A-K, Müller JK. Biparental care and social evolution in burying beetles: lessons from the larder. In: Choe JC, Crespi BJ, editors. The evolution of social behavior in insects and arachnids. Cambridge University Press; Cambridge: 1997. pp. 216–236. [Google Scholar]
- Eggert AK, Reinking M, Müller JK. Parental care improves offspring survival and growth in burying beetles. Anim Behav. 1998;55:97–107. doi: 10.1006/anbe.1997.0588. [DOI] [PubMed] [Google Scholar]
- Fisher RA. The correlation between relatives under the supposition of Mendelian inheritance. Trans R Soc Edinburgh. 1918;52:399–433. [Google Scholar]
- Fraser BA, Janowitz I, Thairu M, Travis J, Hughes KA. Phenotypic and genomic plasticity of alternative male reproductive tactics in sailfin mollies. Proc R Soc Lond B. 2014;281:20132310. doi: 10.1098/rspb.2013.2310. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gilad Y, Rifkin SA, Pritchard JK. Revealing the architecture of gene regulation: the promise of eQTL studies. Trends Genet. 2008;24:408–415. doi: 10.1016/j.tig.2008.06.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grozinger CM, Fan Y, Hoover SE, Winston ML. Genome-wide analysis reveals differences in brain gene expression patterns associated with caste and reproductive status in honey bees (Apis mellifera) Mol Ecol. 2007;16:4837–4848. doi: 10.1111/j.1365-294X.2007.03545.x. [DOI] [PubMed] [Google Scholar]
- Harris RM, Hofmann HA. Neurogenomics of behavioral plasticity. In: Landry CR, Aubin-Horth N, editors. Ecological genomics. Springer Science; Dondrecht: 2014. pp. 149–168. [Google Scholar]
- Kajitani R, Toshimoto K, Noguchi H, Toyoda A, Ogura Y, Okuno M, Yabana M, Harada M, Nagayasu E, Maruyama H, et al. Efficient de novo assembly of highly heterozygous genomes from whole-genome shotgun short reads. Genom Res. 2014;24:1384–1395. doi: 10.1101/gr.170720.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions, and gene fusions. Genom Biol. 2013;14:R36. doi: 10.1186/gb-2013-14-4-r36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li B, Dewey CN. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinf. 2011;12:323. doi: 10.1186/1471-2105-12-323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lodder EM, Scicluna BP, Milano A, Sun AY, Tang H, Remme CA, Moerland PD, Tanck MW, Pitt GS, Marchuk DA, et al. Dissection of a quantitative trait locus for PR interval duration identifies Tnni3k as a novel modulator of cardiac conduction. PLoS Genet. 2012;8:e1003113. doi: 10.1371/journal.pgen.1003113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lynch M, Walsh B. Genetics and analysis of quantitative traits. Sinauer; Sunderland MA: 1998. [Google Scholar]
- Mackay TFC, Stone EA, Ayroles JF. The genetics of quantitative traits: challenges and prospects. Nat Rev Genet. 2009;10:565–57. doi: 10.1038/nrg2612. [DOI] [PubMed] [Google Scholar]
- Maynard Smith J, Burian R, Kauffman S, Alberch P, Campbell J, Goodwin B, Lande R, Raup D, Wolpert L. Developmental constraints and evolution: a perspective from the Mountain Lake conference on development and evolution. Q Rev Biol. 1985;60:265–287. [Google Scholar]
- Mehrabian M, Allayee H, Stockton J, Lum PY, Drake TA, Castellani LW, Suh M, Armour C, Edwards S, Lamb J, et al. Integrating genotypic and expression data in a segregating mouse population to identify 5-lipoxygenase as a susceptibility gene for obesity and bone traits. Nat Genet. 2005;37:1224–1233. doi: 10.1038/ng1619. [DOI] [PubMed] [Google Scholar]
- Monks SA, Leonardson A, Zhu H, Cundiff P, Pietrusiak P, Edwards S, Phillips JW, Sachs A, Schadt EE. Genetic inheritance of gene expression in human cell lines. Am J Hum Genet. 2004;75:1094–1105. doi: 10.1086/426461. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morley M, Molony CM, Weber TM, Devlin JL, Ewens KG, Spielman RS, Cheung VG. Genetic analysis of genome-wide variation in human gene expression. Nature. 2004;430:743–747. doi: 10.1038/nature02797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nuzhdin SV, Brisson JA, Pickering A, Wayne ML, Harshman LG, McIntyre LM. Natural genetic variation in transcriptome reflects network structure inferred with major effect mutations: insulin/TOR and associated phenotypes in Drosophila melanogaster. BMC Genom. 2009;10:124. doi: 10.1186/1471-2164-10-124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Palmer WJ, Duarte A, Schrader M, Day JP, Kilner R, Jiggins FM. A gene associated with social immunity in the burying beetle Nicrophorus vespilloides. Proc R Soc Lond B. 2016;283:20152733. doi: 10.1098/rspb.2015.2733. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parker DJ, Cunningham CB, Walling CA, Stamper CE, Head ML, Roy-Zokan EM, McKinney EC, Ritchie MG, Moore AJ. Transcriptomes of parents identify parenting strategies and sexual conflict in a subsocial beetle. Nat Comm. 2015;6:8449. doi: 10.1038/ncomms9449. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Passador-Gurgel G, Hsieh WP, Hunt P, Deighton N, Gibson G. Quantitative trait transcripts for nicotine resistance in Drosophila melanogaster. Nat Genet. 2007;39:264–268. doi: 10.1038/ng1944. [DOI] [PubMed] [Google Scholar]
- Petretto E, Mangion J, Dickens NJ, Cook SA, Kumaran MK, Lu H, Fischer J, Maatz H, Kren V, Pravenec M, et al. Heritability and tissue specificity of expression quantitative trait loci. PLoS Genet. 2006;2:e172. doi: 10.1371/journal.pgen.0020172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Petretto E, Sarwar R, Grieve I, Lu H, Kumaran MK, Muckett PJ, Mangion J, Schroen B, Benson M, Punjabi PP, et al. Integrated genomic approaches implicate osteoglycin (Ogn) in the regulation of left ventricular mass. Nat Genet. 2008;40:546–552. doi: 10.1038/ng.134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rey S, Boltana S, Vargas R, Roher N, MacKenzie S. Combining animal personalities with transcriptomics resolves individual variation within a wild-type zebrafish population and identifies underpinning molecular differences in brain function. Mol Ecol. 2013;22:6100–6115. doi: 10.1111/mec.12556. [DOI] [PubMed] [Google Scholar]
- Reynolds D, Hofmeister BT, Cliffe E, Alabady M, Siegel TN, Schmitz RJ, Sabatini R. Histone H3 variant regulates RNA polymerase II transcription termination and dual strand transcription of siRNA loci in Trypanosoma brucei. PLoS Genet. 2016;12:e1005758. doi: 10.1371/journal.pgen.1005758. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roberts A, Pimentel H, Trapnell C, Pachter L. Identification of novel transcripts in annotated genomes using RNA-seq. Bioinformatics. 2011;27:2325–2329. doi: 10.1093/bioinformatics/btr355. [DOI] [PubMed] [Google Scholar]
- Robinson MD, McCarthy DJ, Smyth GK. edgeR: a Bioconductor package for differential expression analysis of digital gene expression. Bioinformatics. 2010;26:139–140. doi: 10.1093/bioinformatics/btp616. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rockman MV. The QTN program and the alleles that matter for evolution: all that’s gold does not glitter. Evolution. 2012;66:1–17. doi: 10.1111/j.1558-5646.2011.01486.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rockman MV, Skrovanek SS, Kruglyak L. Selection at linked sites shapes heritable phenotypic variation in C. elegans. Science. 2010;330:372–376. doi: 10.1126/science.1194208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roy-Zokan EM, Cunningham CB, Hebb LE, McKinney EC, Moore AJ. Vitellogenin and vitellogenin receptor gene expression is associated with male and female parenting in a subsocial insect. Proc R Soc Lond B. 2015;282:20150787. doi: 10.1098/rspb.2015.0787. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanogo YO, Band MA, Blatti C, Sinha S, Bell AM. Transcriptional regulation of brain gene expression in response to a territorial intrusion. Proc R Soc Lond B. 2012;279:4929–4938. doi: 10.1098/rspb.2012.2087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schadt EE, Monks SA, Drake TA, Lusis AJ, Che N, Colinayo V, Ruff TG, Milligan SB, Lamb JR, Cavet G, et al. Genetics of gene expression surveyed in maize, mouse, and man. Nature. 2003;422:297–302. doi: 10.1038/nature01434. [DOI] [PubMed] [Google Scholar]
- Schmieder R, Edwards R. Fast identification and removal of sequence contamination from genomic and metagenomic datasets. PLoS ONE. 2011;6:e17288. doi: 10.1371/journal.pone.0017288. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scott MP. The ecology and behavior of burying beetles. Annu Rev Entomol. 1998;43:595–618. doi: 10.1146/annurev.ento.43.1.595. [DOI] [PubMed] [Google Scholar]
- Simão FA, Waterhouse RM, Ioannidis P, Kriventseva EV, Zdobnov EM. BUSCO: assessing genome assembly and annotation completeness with single copy orthologs. Bioinformatics. 2015;31:3210–3212. doi: 10.1093/bioinformatics/btv351. [DOI] [PubMed] [Google Scholar]
- Smith EN, Kruglyak L. Gene-environment interaction in yeast gene expression. PLoS Biol. 2008;6:e83. doi: 10.1371/journal.pbio.0060083. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102:15545–15550. doi: 10.1073/pnas.0506580102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Székely T, Moore AJ, Komdeur J. Social behaviour: genes, ecology, and evolution. Cambridge University Press; Cambridge: 2010. [Google Scholar]
- Todd EV, Black MA, Gemmell NJ. The power and promise of RNA-seq in ecology and evolution. Mol Ecol. 2016;25:1224–1241. doi: 10.1111/mec.13526. [DOI] [PubMed] [Google Scholar]
- Valcu CM, Kempenaers B. Proteomics in behavioral ecology. Behav Ecol. 2014;26:1–15. [Google Scholar]
- Van Gestel S, Houwing-Duistermaat JJ, Adolfsson R, van Duijn CM, Van Broeckhoven C. Power of selective genotyping in genetic association analyses of quantitative traits. Behav Genet. 2000;30:141–146. doi: 10.1023/a:1001907321955. [DOI] [PubMed] [Google Scholar]
- Walling CA, Stamper CE, Smiseth PT, Moore AJ. The quantitative genetics of sex differences in parenting. Proc Natl Acad Sci USA. 2008;105:18430–18435. doi: 10.1073/pnas.0803146105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- West MA, Kim K, Kliebenstein DJ, Van Leeuwen H, Michelmore RW, Doerge RW, Clair DAS. Global eQTL mapping reveals the complex genetic architecture of transcript-level variation in Arabidopsis. Genetics. 2007;175:1441–1450. doi: 10.1534/genetics.106.064972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Whitfield CW, Cziko AM, Robinson GE. Gene expression profiles in the brain predict behavior in individual honey bees. Science. 2003;302:296–299. doi: 10.1126/science.1086807. [DOI] [PubMed] [Google Scholar]
- Wong RY, Lamm MS, Godwin J. Characterizing the neurotranscriptomic states in alternative stress coping. BMC Genom. 2015;16:425. doi: 10.1186/s12864-015-1626-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang D, Ye C, Ma X, Zhu Z, Zhou X, Wang H, Meng Q, Pei X, Yu S, Zhu J. A new approach to dissecting complex traits by combining quantitative trait transcript (QTT) mapping and diallel cross analysis. Chin Sci Bull. 2012;57:2695–2700. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary Figure 1. Hierarchical clustering of N. orbicollis female samples by transcriptional profile.
Supplementary Figure 2. Hierarchical clustering of N. orbicollis male samples by transcriptional profile.
Supplementary Figure 3. Hierarchical clustering of N. vespilloides female samples by transcriptional profile.
Supplementary Figure 4. Hierarchical clustering of N. vespilloides male samples by transcriptional profile.
Supplementary Table 1. Descriptive statistics for Cufflinks produced transcriptome assemblies for N. orbicollis and N. vespilloides. Means indicate the average of the 20 samples sequenced for each species/sex.
Supplementary Table 2. Transcripts significantly differentially expressed between high and low provisioning parents. A list of the significantly DE transcript IDs from each experiment and their blast homologies. Asterisks next transcripts from N. orbicollis females indicate transcripts whose differential expression is explained by extremely high expression values in the same 3 individuals (B47, B91, and B92), and therefore may bias results for these specific transcripts.