Abstract
Purpose
The poor prognosis of multiple myeloma with t(4;14) is driven by the fusion of genes encoding MMSET and immunoglobulin heavy chain. Specific genes affected by MMSET and their clinical implications in Non-MMSET myeloma remain undetermined.
Experimental design
We obtained gene-expression profiles of 1,032 newly diagnosed myeloma patients enrolled in Total Therapy 2, Total Therapy 3, Myeloma IX, and HOVON65-GMMGHD4 trials, and 156 patients from Multiple Myeloma Resource Collection. Probes most correlated with MMSET myeloma were selected based on a multivariable linear regression and Bonferroni correction, and refined based on the strength of association with survival in Non-MMSET patients.
Results
Ten MMSET-like probes were associated with poor survival in Non-MMSET myeloma. Non-MMSET myeloma patients in the highest quartile of the 10-gene signature (MMSET-like myeloma) had 5-year overall survival similar to that of MMSET myeloma (highest quartile vs. lowest quartile hazard ratio [HR]=2.0, 95% confidence interval [CI] 1.5-2.8 in MMSET-like myeloma; HR=2.3, 95% CI: 1.6-3.3 in MMSET myeloma). Analyses of MMSET-like gene signature suggested the involvement of p53 and MYC pathways.
Conclusion
MMSET-like gene signature captures a subset of high-risk myeloma patients under-represented by conventional risk stratification platforms, and defines a distinct biological subtype.
Keywords: multiple myeloma, high risk myeloma, MMSET, t(4;14), cDNA microarray
Introduction
Multiple myeloma has extremely heterogeneous outcomes. Among many prognostic factors utilized in myeloma, translocation t(4;14)(p16.3;q32.3) is an oncogenic event associated with poor prognosis. (1) The key molecular target of t(4;14) is multiple myeloma SET domain (MMSET) at chromosomal band 4p16.3. (2-5) The detection of MMSET overexpression with gene-expression profiling (GEP) consistently identifies a high-risk subgroup in multiple myeloma. (6) While the prognostic significance of MMSET is well established, the underlying mechanism of its excess risk is poorly understood. Given MMSET encodes histone methyltransferase, its overexpression has been attributed to alter epigenetic regulation of genes involved in cell cycle progression and DNA damage repair. (7) However, downstream gene targets and molecular pathways regulated by MMSET remain unclear.
What is also unknown in myeloma is the presence of biological homology shared between high-risk and non-high-risk subgroups. This question comes within the context of the recent advancement of genetic sequencing, which identified diverse spectrum of disease biology that, at times, redefined conventional risk stratification and management. For instance, “BRCA-ness” was identified in up to 14% of non-small cell lung cancer and 15% of head and neck cancer patients due to epigenetic inactivation of genes responsible for DNA damage repair, such as BRCA1 and FNACF. (8) In breast and ovarian cancers, next-generation sequencing demonstrated the presence of certain genes beyond BRCA1/2, such as PALB2, ATM or CHEK2, was strongly associated with an increased risk of cancer diagnosis and early death. (9-11) Recent discoveries in solid tumor suggest a substantial proportion of cancer patients harbors molecular signatures similar to those of high-risk subtypes.
We hypothesize there is an overlap of disease biology between the established high-risk myeloma and its non-high-risk counterpart. Specifically, the same genes involved in the pathogenesis and adverse outcomes of MMSET myeloma (6) could also be relevant to a subset of Non-MMSET patients with poor clinical outcomes (hereby refer to “MMSET-like myeloma”). To characterize genes and molecular pathways influencing survival across different myeloma subtypes, we assessed expression levels of 54,675 genes in 1,188 newly diagnosed multiple myeloma patients. Among 71 genes significantly altered in MMSET myeloma, 10 genes most strongly associated with survival were selected and combined into a GEP risk score. Patients who did not have detectable MMSET but were at the top quartile of the 10-gene risk score were categorized as MMSET-like myeloma. Five-year survivals were similar between patients with MMSET myeloma and MMSET-like myeloma. Pathway analysis identified MYC and TP53 transcriptional regulators as lead candidates targeted by the observed genes within the risk score. Our findings suggest there is a homology of aggressive disease biology and clinical outcomes shared between MMSET myeloma and a subset of non-MMSET myeloma.
Methods
Study design
From the NCBI Gene Expression Omnibus (GEO), we downloaded unprocessed CEL files from the following datasets: Total Therapy (TT) 2 (N=345, accession number GSE2658, NCT00083551); TT 3 (N=214, accession number GSE2658, NCT00081939); HOVON65/GMMG-HD4, (N=320, accession number GSE19784, ISRCTN64455289); Myeloma IX (N=247, accession number GSE15695, ISRCTN68454111); and Multiple Myeloma Reference Collection (MMRC) (N=288, accession number GSE26760). The sample size of each data set was determined after excluding 8 profiles (accession number GSE19784) that were normal plasma cells and 16 patients (accession number GSE26760) who were smoldering myeloma (n=11), MGUS (n=2), or plasma cell leukemia (n=3). Anonymized patient characteristics of TT trials were obtained from GEO and were identified with the same accession numbers. Anonymized patient characteristics of Myeloma IX and HOVON65/GMMG-HD4 trials were obtained through personal correspondence with Mark van Duin and Ping Wu, respectively. Anonymized patient characteristics of MMRC were obtained from Multiple Myeloma Genome Portal. (12) Selected characteristics of patients from the five studies are shown in Table 1.
Table 1. Patient and study characteristics.
| Induction Therapy | Maintenance | Median Age, Years (range) | Women, N (%) | ISS 1, N (%) | ISS 2, N (%) | ISS 3, N (%) | Maximum Follow-up (years) | |
|---|---|---|---|---|---|---|---|---|
|
TT2 (N=345) TT3 (N=208) |
D(T)-PACE (N=345) | Thalidomide | 57 (24-77) | 148 (43) | 184 (53) | 90 (26) | 71 (21) | 8.2 |
| VTD-PACE (N=208) | Bort-Thal-Dex | 60 (32-75) | 72 (34) | 100 (48) | 64 (31) | 44 (21) | 4.4 | |
| HOVON65/GMMGHD5 (N=296) | VAD (N=143) | Thalidomide | 58 (27-65) | 61 (43) | 55 (38) | 44 (31) | 44 (31) | 6.1 |
| PAD (N=153) | Bortezomib | 56 (31-65) | 58 (38) | 51 (33) | 62 (41) | 40 (26) | 5.8 | |
| Myeloma IX (N=183) | CTD (N=58) | (+/-) Thalidomide (N=30) | 59.5 (45-69) | 11 (37) | 8 (27) | 12 (40) | 10 (33) | 8.1 |
| NULL (28) | 61 (35-68) | 13 (46) | 10 (36) | 10 (36) | 8 (28) | 7.6 | ||
| CVAD (N=48) | (+/-) Thalidomide (N=23) | 57 (39-69) | 6 (26) | 8 (35) | 7 (30) | 8 (35) | 7.6 | |
| NULL (N=25) | 60 (48-68) | 10 (40) | 6 (24) | 9 (36) | 10 (40) | 7.4 | ||
| CTDa (N=41) | (+/-) Thalidomide (N=19) | 73 (67-83) | 8 (42) | 2 (11) | 5 (27) | 12 (63) | 7.5 | |
| NULL (N=22) | 73.5 (61-84) | 12 (55) | 1 (5) | 11 (50) | 10 (46) | 7.7 | ||
| Melphalan (N=36) | (+/-) Thalidomide (N=15) | 70 (63-80) | 8 (53) | 5 (33) | 5 (33) | 5 (33) | 7.2 | |
| NULL (N=21) | 74 (62-89) | 8 (38) | 2 (10) | 7 (34) | 12 (57) | 6.5 | ||
| MMRC* (N=156) | -* | -* | 60 (24-89) | 51 (32) | 74 (48) | 46 (30) | 36 (23) | -* |
| All Studies (N=1188) | -* | -* | 59 (24-89) | 466 (39%) | 506 (43%) | 372 (31%) | 310 (26%) | 8.2 |
D(T)-PACE: Dexamethasone with or without thalidomide, cisplatin P, doxorubicin A, cyclophosphamide C, etoposide E. VTD-PACE: V Bortezomib. PAD: Bortezomib P, Doxorubicin A, Dexamethasone D, VAD: Vincristine V, Doxorubicin A, Dexamethasone D. CTD/CVAD: Cyclophosphamide C, Thalidomide T, Doxorubicin A, Dexamethasone D.
=information not available
All gene-expression data were derived from CD138+ purified plasma cells of newly diagnosed myeloma patients, which were hybridized to Affymetrix Human Genome U133 Plus 2.0 cDNA microarray (Santa Clara, CA). All raw CEL files were processed using the justMAS function in the R statistical programming language, and gene-expression levels were log2 transformed. The final dataset included GEPs of 1,188 myeloma patients with complete data for age, sex, beta-2 microglobulin, and albumin. For the HOVON65/GMMG-HD4 trial, FISH data regarding MMSET status was available for 241 patients; MMSET status by FISH versus gene expression revealed a correlation of 0.81 (Spearman's rho). For the analysis of survival outcomes, we excluded 156 patients from MMRC as it was not a clinical trial, and only used the remaining data from 1,032 patients.
Institutional Review Boards of respective institutions approved all studies. All subjects provided written informed consents approving the use of their samples for research purposes.
Statistical analysis
MMSET myeloma patients, non-MMSET myeloma patients, and genes associated with MMSET myeloma
To classify patients into MMSET or Non-MMSET myeloma, we used the previously reported microarray model using 700 gene probes to assign subjects into one of seven molecular subtypes. (13) We assessed the association of MMSET myeloma with individual expression levels of 54,675 available probes. By using linear regression models for each probe and for each study, gene-expression levels were dependent variables of MMSET status, age (divided by 50 years or less, 51-60 years, 61-70 years, 71 years or older), sex, and International Staging System (ISS) stage. (14) For each probe, study-specific linear regression coefficients for MMSET myeloma were then combined across studies using a random effects meta-analysis. (15) Prior to finalizing the probes that were significantly associated with MMSET myeloma, all 700 gene probes used in the Arkansas model (13) were removed. We performed a random effects meta-analysis after Bonferroni correction for multiple testing (p<0.05/54,675=9.14×10-7). The absolute value of the random effects slope parameter for MMSET myeloma was 2 or greater, indicating MMSET myeloma had 2-fold or greater changes in log expression of a given gene.
Identification of probes associated with survival in Non-MMSET patients
To identify probes relevant to survival of non-MMSET patients, results from the aforementioned meta-analysis were analyzed by a stepwise variable selection (proc phreg, SAS 9.3) in a Cox proportional hazards model. Duration of follow-up was defined by the start of treatment until death or censoring. Censoring occurred when a subject reached 5 years or was lost to follow up. For the initial selection of probes, we included probes that passed Bonferroni correction for multiple testing, and those showed log 2 or greater changes of expression. For each probe, a minimal p<0.1 in a marginal Cox proportional hazards model was set for initial inclusion, and a criteria of p<0.05 was set to retain a probe in the model. All models were adjusted for age, sex, ISS stage (14) and treatment. (16-18) The risk score was calculated based on the adjusted Cox regression model (Appendix 1).
Validation
To assess the unbiased association of the risk score and survival, we conducted a 5-fold cross-validation. (18) Briefly, the original dataset was divided into five equal parts, with equal numbers of patients from individual studies in each part. Four of the five parts were used to develop a gene signature following the aforementioned procedures (training set). The remaining fifth part was used to compute the association of the risk score and survival using Cox regression models (test set). Validation was performed five times with each part serving as a test set once. Risk scores from five test sets were median-centered and combined to form an independently scored measure of risk.
Sensitivity analysis
To assess stability of our results, we conducted three separate sensitivity analyses (Supplemental Tables 1 and 2). First, we used survival outcomes throughout the full follow-up time of up to 98 months instead of censoring at 60 months. Second, we excluded patients who were treated with proteasome inhibitor-based regimens, such as VTD-PACE and PAD, from the analysis. Third, patients on Myeloma IX trial were coded separately if they encountered death or censoring before the second randomization for thalidomide maintenance. In all three sensitivity analyses, the main results remained unchanged.
Pathway analysis
To determine biological functions of the identified gene probes, pathway analysis was performed using Ingenuity Pathway Analysis software package and the molecular signaling database from the Broad Institute (MsigDB). (19) Gene networks were constructed using the upstream regulator analysis to identify transcription factors with the most interactions with selected genes (Figure 1).
Figure 1.
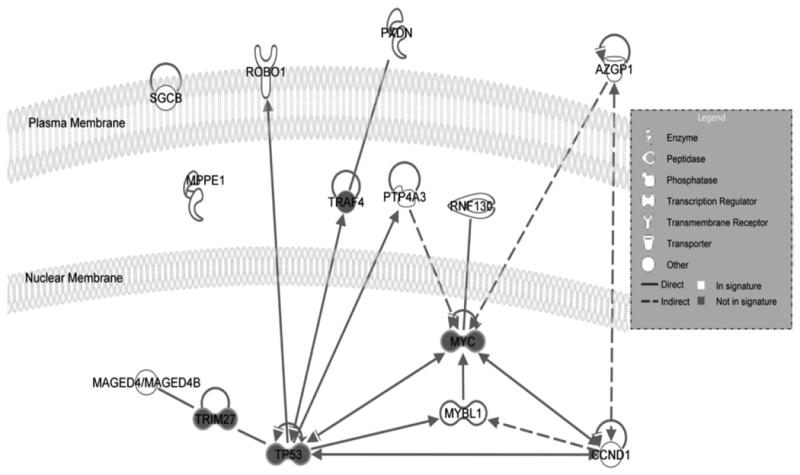
Full lines represent direct interactions while dashed lines indicate indirect interactions. An arrow pointing from one protein to another indicates that the first protein acts on or activates the second protein (at which the arrow is pointing).
Results
Among 1,032 myeloma patients included in this study, 139 (13.4%) had MMSET myeloma defined by GEP (Table 1). (13) In MMSET myeloma, the median age was 59 years (range 24-89), and 68% were males. Distributions of ISS stage I, II, and III were 48%, 30% and 22%, respectively. Similar to prior reports (6), MMSET myeloma was associated with a higher mortality after adjusting for age, sex, ISS stage and treatment (hazard ratio (HR) =1.7, p<0.001).
To determine if the same genes involved in MMSET myeloma were also relevant to survival of Non-MMSET myeloma patients, we took the following analytical approach: First, as described in the Methods, we obtained GEPs of 1,188 newly diagnosed myeloma patients and defined 71 gene probes correlated with MMSET myeloma (Supplemental Table 3). From these probes, we further identified those associated with 5-year survival in Non-MMSET patients and created a 10-gene risk score predictive of survival. Lastly, we conducted a functional pathway analysis.
Gene probes correlated with MMSET myeloma
After the random effects meta-analysis, we identified 71 gene probes (0.13%) correlated with MMSET myeloma. The selected genes showed 2-fold or greater changes in log-expressions (range 2.0 to 3.7 or -2.0 to -3.7) in MMSET patients compared to Non-MMSET patients, and meta-analytic p-values ranged from 1.9×10-11 to 5.2×10-36 (Supplemental Table 3). Genes highly correlated with MMSET myeloma included cyclin D1 (CCND1), cyclin D2 (CCND2), a transcription factor Kruppel-like factor 4 (KLF4), ubiquitin carboxyl-terminal esterase L1 (UCHL1), and alpha-2-glycoprotein (AZGP1) (Supplemental Table 3).
Probes enriched in MMSET myeloma, Non-MMSET myeloma, and survival
From the identified 71 gene probes, 10 genes were strongly associated with 5-year survival of Non-MMSET patients (Table 2). AZGP1 and CCND1 were most significantly associated with survival (probe-specific HRs: 0.89-0.91 for CCND1 and 1.07-1.14 for AZGP1, p<0.001). To define risk scores relevant to survival of non-MMSET myeloma patients, a multivariable Cox proportional hazards model was applied to the 10 genes. Risk score groups of the first quartile (low-risk) and the fourth quartile (high-risk) were compared within Non-MMSET patients in cross-validation. High-risk Non-MMSET patients (here by referred as “MMSET-like myeloma”) had a similarly increased risk of mortality (HR=2.0, 95% confidence interval (CI) 1.5-2.8, p<0.001) comparable to MMSET patients (HR=2.3, 95% CI 1.6-3.3, p<0.001).
Table 2. Genes identified as highly correlated to MMSET myeloma and associated with survival in Non-MMSET patients.
| gene | probe | Lin Reg coef1 | p-value | HR2 (95% CI) | p-value | TP53 Interacting | MYC Interacting | annotation |
|---|---|---|---|---|---|---|---|---|
| CCND1 | 208712_at | -3.6 | 1.70E-33 | 0.90 (0.86-0.94) | <.0001 | Yes | Yes | cyclin D1 |
| AZGP1 | 209309_at | 3.08 | 6.40E-25 | 1.10 (1.05-1.16) | <.0001 | No | Yes | alpha-2-glycoprotein 1 zinc |
| SGCB | 205120_s_at | 2.51 | 3.90E-17 | 1.09 (1.03-1.15) | 0.0014 | No | No | sarcoglycan beta (dystrophin-associated glycoprotein) |
| MAGED4 | 223313_s_at | 2.95 | 5.00E-23 | 0.92 (0.87-0.97) | 0.0033 | No | No | melanoma antigen family D 4 |
| PXDN | 212012_at | 2.21 | 1.30E-13 | 1.09 (1.03-1.15) | 0.0035 | No | No | peroxidasin homolog (Drosophila) |
| RNF130 | 217865_at | 2.47 | 1.40E-16 | 0.93 (0.89-0.98) | 0.0053 | No | Yes | ring finger protein 130 |
| MYBL1 | 213906_at | 2.5 | 5.30E-17 | 1.08 (1.02-1.14) | 0.0086 | Yes | Yes | v-myb myeloblastosis viral oncogene homolog (avian)-like 1 |
| PTP4A3 | 206574_s_at | 2.51 | 4.70E-17 | 0.94 (0.90-0.99) | 0.0104 | Yes | Yes | protein tyrosine phosphatase type IVA member 3 |
| MPPE1 | 213924_at | 2.28 | 2.10E-14 | 0.92 (0.86-0.98) | 0.0109 | No | No | Metallophosphoesterase 1 |
| ROBO1 | 213194_at | 2.23 | 8.60E-14 | 1.06 (1.01-1.11) | 0.0288 | Yes | No | roundabout axon guidance receptor homolog 1 (Drosophila) |
linear regression coefficient associated with MMSET from meta analysis that uses log2 transformed probe levels as outcome
HRs from adjusted Cox regression model that fit each probe separately to 5-year survival
Pathway analysis
To characterize genes and molecular pathways influencing survival across different myeloma subtypes, we conducted analysis of the 10 genes associated with 5-year survival in MMSET-like myeloma by using IPA (Ingenuity® Systems). Pathway analysis identified MYC and TP53 transcriptional regulators as lead candidates for the observed gene expression changes within the gene signature risk score (Figure 1). TP53 was identified as a transcriptional regulator of four genes (CCND1, PTP4A3, MYBL1, and ROBO1) (p=1.9×10-3), and MYC was a transcriptional regulator of five genes (CCND1, AZGP1, PTP4A3, MYBL1, and RNF130) (p=3.1×10-4).
Discussion
To characterize genes and molecular pathways influencing survival across myeloma subtypes, we assessed expression levels of over 55,000 gene probes from tumor cells obtained from 1,188 newly diagnosed myeloma patients. 71 genes were significantly altered in patients with the MMSET molecular subtype. Selecting from these genes, 10-gene risk score demonstrated similar 5-year survivals between MMSET myeloma and Non-MMSET patients categorized as the top quartile risk score (MMSET-like myeloma). A 5-fold cross-validation was conducted to determine the unbiased association of the risk score and survival. Pathway analysis identified MYC and TP53 transcriptional regulators were associated with the observed gene-expresssion changes of 10 genes.
Of clinical relevance, our findings suggest an overlap of disease biology between conventionally divided groups of high-risk and non-high-risk myelomas. The study findings should be interpreted within the context of recent advancement of genetic sequencing, which refined tumor subtypes based on recurrent genetic alterations. In ovarian cancer, approximately half of the patients were found to have homologous recombination deficiency (HRD) mimicking the genetic phenotype of BRCA mutation (i.e. “BRCA-ness”). (20) Intriguingly, the presence of HRD in BRCA wild-type patients predicted striking sensitivity to PARP inhibition in a prospective trial (overall response rate: 32% with HRD vs. 11% without HRD), albeit less than the true BRCA mutated group (66%). Evolving knowledge in biological homology across different tumor subtypes proposes a new therapeutic strategy is required to improve the outcome of patients with MMSET-like gene signature. As seen in differential responsiveness to PARP inhibition in cancers with BRCA-ness, MMSET-like subgroup may also benefit from established or investigational regimens developed for high-risk myeloma, rather than those developed for standard-risk population. Such regimens tested in high-risk myeloma include proteasome inhibitors (21-23) and other investigational agents aimed at novel targets such as FGFR3, (5) CD38, (24) and MEK pathway. (25) Further research needs to validate the role of genomic risk stratification tools to capture high-risk population, and to prospectively assess clinical outcome to potential treatment options within the identified subgroup.
Another important observation of this study is the demonstration of TP53 and MYC as downstream targets of MMSET gene signature. t(4;14) accounts for 15% of myeloma population and is linked to universal overexpression of MMSET gene. (3, 4) Histone methyltransferase encoded at catalytic SET domain methylates lysine residue of histone, leading to epigenetic regulation of genes involved in cell cycle progression, p53 pathway, and integrin signaling. (7) The role of MMSET as a myeloma oncogene is supported by an experimental knock-down of MMSET in myeloma cell-lines, which led to decreased proliferation and increased apoptosis. (2, 26, 27), Among many targets altered by MMSET overexpression, c-myc is an important downstream pathway enhanced by MMSET through down-regulation of miR-126. (28, 29) The overlap of p53 and MYC pathways has also been described in model systems of other malignancies. In vitro, p53 represses c-myc transcription by deacetylation of histone located at c-myc promoter (30) and by miR-145-mediated gene silencing, (31) and arrests cell cycle. These findings support the primary role of MMSET as a regulator of epigenetic machineries, rather than genetic instability, and is corroborated by findings from whole exome sequencing which demonstrated only a few mutational changes in the t(4;14) subgroup. (32) Taken together, an aggressive clinical phenotype of MMSET overexpression is attributable to the fine-tuning of selected genes. Functional studies are required to assess direct binding or indirect modulation of 10 genes by MSMET and to validate downstream activity of MMSET-like signature converging into selected signaling pathways, such as MYC and p53.
Gene-expression profiling is a mature and robust technology with many validated platforms in multiple myeloma reported to date. (33) Compared to previously established platforms, MMSET-like signature has several unique aspects. First, MMSET-like gene signature was developed from a biologically homogeneous population with a single genetically defined abnormality, and was applied to the overall population with an aim to select patients influenced by similar pathobiology. This sequence of development is reversed from what had been done in conventional studies, which performed hierarchical gene clustering among biologically heterogeneous population. (34) By using the latter method, a given gene-expression group can contain several different genetic abnormalities within the subtype, (6) which may have led to inconsistent results in predicting therapeutic responses. (35) The 10-gene signature proposed by the current study was developed from a homogenous subgroup, hence may be more representative of a single biological entity and can serve a useful risk stratification tool for treatment trials. Second, with the exception of one gene, 10-gene signature did not overlap with previously reported platforms such as EMC 92-gene, (34) UAMS 70-gene (6) and IFM 15-gene signatures. (36) This finding further supports that MMSET-like gene signature represents a distinct biological subtype utilizing a selected set of genes. Interestingly, ROBO1 was the only gene within our 10-gene platform that was previously reported in another gene expression profile (37) and in a sequencing study as a candidate gene in myeloma. (32) Downstream of ROBO1 is associated with E-cadherin mediated regulation of WNT signaling in pancreatic cancer, and its functional role in myeloma remains to be studied.
We demonstrated 10-gene signature that were significantly altered in MMSET myeloma and associated with inferior survival in Non-MMSET myeloma patients. Pathway analysis of the MMSET-like gene signature recapitulated clustering of important signaling pathways in myeloma, specifically TP53 and MYC pathways. MMSET-like gene expression profile was able to capture a distinct biological subtype under-represented by conventional platforms, and was strongly linked poor clinical outcome. The proposed gene signature can serve as a reliable screening platform representative of high-risk disease biology, as we move towards personalized therapy for myeloma.
Supplementary Material
Statement of Translational Relevance.
Multiple myeloma is a biologically and clinically heterogeneous disease. The presence of biological homology shared between conventional high-risk and non-high-risk myeloma subgroups has not been reported to date. We hypothesized that molecular risk stratification can capture biological homology between patients with or without MMSET overexpression, and be used as a prognostic tool. We identified 10-gene signature associated with MMSET myeloma. We obtained gene-expression profiles of 1,032 newly diagnosed myeloma patients enrolled in Total Therapy 2, Total Therapy 3, Myeloma IX, and HOVON65-GMMGHD4 trials, and 156 patients from Multiple Myeloma Resource Collection. Expression of MMSET-like gene signature in Non-MMSET subgroup was associated with similarly poor survival. Pathway analysis of MMSET-like gene signature revealed the involvement of p53 and MYC signaling pathways. MMSET-like gene signature captures a subset of high-risk myeloma patients under-represented by conventional risk stratification platforms, and defines a distinct biological subtype.
Acknowledgments
None
Grant Support: This work was supported by the intramural research program of the National Institutes of Health. Research support for S.P.W. was made possible through the National Institutes of Health (NIH) Medical Research Scholars Program, a public-private partnership supported jointly by the NIH and generous contributions to the Foundation for the NIH from Pfizer Inc, The Leona M. and Harry B. Helmsley Charitable Trust, and the Howard Hughes Medical Institute, as well as other private donors. For a complete list, please visit the Foundation website at http://www.fnih.org/work/programs-development/medical-research-scholars-program).
Footnotes
Disclosure of Potential Conflict of Interest: Dr. Landgren has given scientific presentations at meeting funded by Onyx Pharmaceuticals/AMGEN, Celgene, BMS and Jansen; has served on the Independent Data Monitoring Committee for clinical trials by Millennium Pharmaceuticals/Takeda; and has served on the advisory committee for Medscape Educations, Myeloma program.
Authors' Contributions: Contribution: S.P.W, R.M.P., and O.L. designed the study and performed bioinformatics analysis; S.P.W, R.M.P., I.E.A., S.M., P.S., M.D., N.M., B.W., and O.L. reviewed data analysis and wrote manuscript.
Supplementary information contains 3 tables and 1 figure, which are available at Clinical Cancer Research Online (http://clincancerres.aacrjournals.org/).
References
- 1.Cavo M, Terragna C, Renzulli M, Zamagni E, Tosi P, Testoni N, et al. Poor outcome with front-line autologous transplantation in t(4;14) multiple myeloma: low complete remission rate and short duration of remission. Journal of clinical oncology : official journal of the American Society of Clinical Oncology. 2006;24:e4–5. doi: 10.1200/JCO.2005.04.7506. [DOI] [PubMed] [Google Scholar]
- 2.Chesi M, Nardini E, Lim RS, Smith KD, Kuehl WM, Bergsagel PL. The t(4;14) translocation in myeloma dysregulates both FGFR3 and a novel gene, MMSET, resulting in IgH/MMSET hybrid transcripts. Blood. 1998;92:3025–34. [PubMed] [Google Scholar]
- 3.Santra M, Zhan F, Tian E, Barlogie B, Shaughnessy J., Jr A subset of multiple myeloma harboring the t(4;14)(p16;q32) translocation lacks FGFR3 expression but maintains an IGH/MMSET fusion transcript. Blood. 2003;101:2374–6. doi: 10.1182/blood-2002-09-2801. [DOI] [PubMed] [Google Scholar]
- 4.Keats JJ, Maxwell CA, Taylor BJ, Hendzel MJ, Chesi M, Bergsagel PL, et al. Overexpression of transcripts originating from the MMSET locus characterizes all t(4;14)(p16;q32)-positive multiple myeloma patients. Blood. 2005;105:4060–9. doi: 10.1182/blood-2004-09-3704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Mirabella F, Wu P, Wardell CP, Kaiser MF, Walker BA, Johnson DC, et al. MMSET is the key molecular target in t(4;14) myeloma. Blood Cancer J. 2013;3:e114. doi: 10.1038/bcj.2013.9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Shaughnessy JD, Jr, Zhan F, Burington BE, Huang Y, Colla S, Hanamura I, et al. A validated gene expression model of high-risk multiple myeloma is defined by deregulated expression of genes mapping to chromosome 1. Blood. 2007;109:2276–84. doi: 10.1182/blood-2006-07-038430. [DOI] [PubMed] [Google Scholar]
- 7.Martinez-Garcia E, Popovic R, Min DJ, Sweet SM, Thomas PM, Zamdborg L, et al. The MMSET histone methyl transferase switches global histone methylation and alters gene expression in t(4;14) multiple myeloma cells. Blood. 2011;117:211–20. doi: 10.1182/blood-2010-07-298349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Marsit CJ, Liu M, Nelson HH, Posner M, Suzuki M, Kelsey KT. Inactivation of the Fanconi anemia/BRCA pathway in lung and oral cancers: implications for treatment and survival. Oncogene. 2004;23:1000–4. doi: 10.1038/sj.onc.1207256. [DOI] [PubMed] [Google Scholar]
- 9.Antoniou AC, Casadei S, Heikkinen T, Barrowdale D, Pylkas K, Roberts J, et al. Breast-cancer risk in families with mutations in PALB2. The New England journal of medicine. 2014;371:497–506. doi: 10.1056/NEJMoa1400382. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Weischer M, Nordestgaard BG, Pharoah P, Bolla MK, Nevanlinna H, Van't Veer LJ, et al. CHEK2*1100delC heterozygosity in women with breast cancer associated with early death, breast cancer-specific death, and increased risk of a second breast cancer. Journal of clinical oncology : official journal of the American Society of Clinical Oncology. 2012;30:4308–16. doi: 10.1200/JCO.2012.42.7336. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Easton DF, Pharoah PD, Antoniou AC, Tischkowitz M, Tavtigian SV, Nathanson KL, et al. Gene-panel sequencing and the prediction of breast-cancer risk. The New England journal of medicine. 2015;372:2243–57. doi: 10.1056/NEJMsr1501341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Multiple Myeloma Genome Portal. [cited; Available from: http://www.broadinstitute.org/mmgp.
- 13.Zhan F, Huang Y, Colla S, Stewart JP, Hanamura I, Gupta S, et al. The molecular classification of multiple myeloma. Blood. 2006;108:2020–8. doi: 10.1182/blood-2005-11-013458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Dimopoulos M, Gika D, Zervas K, Kyrtsonis M, Symeonidis A, Anagnostopoulos A, et al. The international staging system for multiple myeloma is applicable in symptomatic Waldenstrom's macroglobulinemia. Leuk Lymphoma. 2004;45:1809–13. doi: 10.1080/10428190410001687512. [DOI] [PubMed] [Google Scholar]
- 15.Pfeiffer RM, Gail MH, Pee D. On combining data from genome-wide association studies to discover disease-associated SNPs. Statistical Science. 2009;24:547–60. [Google Scholar]
- 16.Zangari M, van Rhee F, Anaissie E, Pineda-Roman M, Haessler J, Crowley J, et al. Eight-year median survival in multiple myeloma after total therapy 2: roles of thalidomide and consolidation chemotherapy in the context of total therapy 1. Br J Haematol. 2008;141:433–44. doi: 10.1111/j.1365-2141.2008.06982.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Morgan GJ, Gregory WM, Davies FE, Bell SE, Szubert AJ, Brown JM, et al. The role of maintenance thalidomide therapy in multiple myeloma: MRC Myeloma IX results and meta-analysis. Blood. 2012;119:7–15. doi: 10.1182/blood-2011-06-357038. [DOI] [PubMed] [Google Scholar]
- 18.Molinaro AM, Simon R, Pfeiffer RM. Prediction error estimation: a comparison of resampling methods. Bioinformatics. 2005;21:3301–7. doi: 10.1093/bioinformatics/bti499. [DOI] [PubMed] [Google Scholar]
- 19.Molecular Signaling Database from the Broad Institute. [cited; Available from: http://www.broadinstitute.org/gsea/msigdb.
- 20.McNeish I, Oza A, Coleman RL, Scott CL, Konecny G, O'Malley DM, et al. Results of ARIEL2: A Phase 2 trial to prospectively identify ovarian cancer patients likely to respond to rucaparib using tumor genetic analysis. Journal of clinical oncology : official journal of the American Society of Clinical Oncology. 2015;33 (abstract 5508) [Google Scholar]
- 21.Avet-Loiseau H, Leleu X, Roussel M, Moreau P, Guerin-Charbonnel C, Caillot D, et al. Bortezomib plus dexamethasone induction improves outcome of patients with t(4;14) myeloma but not outcome of patients with del(17p) Journal of clinical oncology : official journal of the American Society of Clinical Oncology. 2010;28:4630–4. doi: 10.1200/JCO.2010.28.3945. [DOI] [PubMed] [Google Scholar]
- 22.Neben K, Lokhorst HM, Jauch A, Bertsch U, Hielscher T, van der Holt B, et al. Administration of bortezomib before and after autologous stem cell transplantation improves outcome in multiple myeloma patients with deletion 17p. Blood. 2012;119:940–8. doi: 10.1182/blood-2011-09-379164. [DOI] [PubMed] [Google Scholar]
- 23.Jakubowiak AJ, Siegel DS, Martin T, Wang M, Vij R, Lonial S, et al. Treatment outcomes in patients with relapsed and refractory multiple myeloma and high-risk cytogenetics receiving single-agent carfilzomib in the PX-171-003-A1 study. Leukemia. 2013;27:2351–6. doi: 10.1038/leu.2013.152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Lokhorst HM, Plesner T, Laubach JP, Nahi H, Gimsing P, Hansson M, et al. Targeting CD38 with Daratumumab Monotherapy in Multiple Myeloma. N Engl J Med. 2015;373:1207–19. doi: 10.1056/NEJMoa1506348. [DOI] [PubMed] [Google Scholar]
- 25.Holkova B, Zingone A, Kmieciak M, Bose P, Badros AZ, Voorhees PM, et al. A Phase II Trial of AZD6244 (Selumetinib, ARRY-142886), an Oral MEK1/2 Inhibitor, in Relapsed/Refractory Multiple Myeloma. Clin Cancer Res. 2015 doi: 10.1158/1078-0432.CCR-15-1076. [Epub ahead of print] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Lauring J, Abukhdeir AM, Konishi H, Garay JP, Gustin JP, Wang Q, et al. The multiple myeloma associated MMSET gene contributes to cellular adhesion, clonogenic growth, and tumorigenicity. Blood. 2008;111:856–64. doi: 10.1182/blood-2007-05-088674. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Brito JL, Walker B, Jenner M, Dickens NJ, Brown NJ, Ross FM, et al. MMSET deregulation affects cell cycle progression and adhesion regulons in t(4;14) myeloma plasma cells. Haematologica. 2009;94:78–86. doi: 10.3324/haematol.13426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Min DJ, Ezponda T, Kim MK, Will CM, Martinez-Garcia E, Popovic R, et al. MMSET stimulates myeloma cell growth through microRNA-mediated modulation of c-MYC. Leukemia. 2013;27:686–94. doi: 10.1038/leu.2012.269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Popovic R, Martinez-Garcia E, Giannopoulou EG, Zhang Q, Zhang Q, Ezponda T, et al. Histone methyltransferase MMSET/NSD2 alters EZH2 binding and reprograms the myeloma epigenome through global and focal changes in H3K36 and H3K27 methylation. PLoS Genet. 2014;10:e1004566. doi: 10.1371/journal.pgen.1004566. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Ho JS, Ma W, Mao DY, Benchimol S. p53-Dependent transcriptional repression of c-myc is required for G1 cell cycle arrest. Mol Cell Biol. 2005;25:7423–31. doi: 10.1128/MCB.25.17.7423-7431.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Sachdeva M, Zhu S, Wu F, Wu H, Walia V, Kumar S, et al. p53 represses c-Myc through induction of the tumor suppressor miR-145. Proc Natl Acad Sci U S A. 2009;106:3207–12. doi: 10.1073/pnas.0808042106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Bolli N, Avet-Loiseau H, Wedge DC, Van Loo P, Alexandrov LB, Martincorena I, et al. Heterogeneity of genomic evolution and mutational profiles in multiple myeloma. Nat Commun. 2014;5:2997. doi: 10.1038/ncomms3997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Chng WJ, Dispenzieri A, Chim CS, Fonseca R, Goldschmidt H, Lentzsch S, et al. IMWG consensus on risk stratification in multiple myeloma. Leukemia. 2014;28:269–77. doi: 10.1038/leu.2013.247. [DOI] [PubMed] [Google Scholar]
- 34.Kuiper R, Broyl A, de Knegt Y, van Vliet MH, van Beers EH, van der Holt B, et al. A gene expression signature for high-risk multiple myeloma. Leukemia. 2012;26:2406–13. doi: 10.1038/leu.2012.127. [DOI] [PubMed] [Google Scholar]
- 35.Amin SB, Yip WK, Minvielle S, Broyl A, Li Y, Hanlon B, et al. Gene expression profile alone is inadequate in predicting complete response in multiple myeloma. Leukemia. 2014;28:2229–34. doi: 10.1038/leu.2014.140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Decaux O, Lode L, Magrangeas F, Charbonnel C, Gouraud W, Jezequel P, et al. Prediction of survival in multiple myeloma based on gene expression profiles reveals cell cycle and chromosomal instability signatures in high-risk patients and hyperdiploid signatures in low-risk patients: a study of the Intergroupe Francophone du Myelome. J Clin Oncol. 2008;26:4798–805. doi: 10.1200/JCO.2007.13.8545. [DOI] [PubMed] [Google Scholar]
- 37.Zhan F, Hardin J, Kordsmeier B, Bumm K, Zheng M, Tian E, et al. Global gene expression profiling of multiple myeloma, monoclonal gammopathy of undetermined significance, and normal bone marrow plasma cells. Blood. 2002;99:1745–57. doi: 10.1182/blood.v99.5.1745. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


