Abstract
Really interesting new gene (RING) finger protein is a type of zinc-binding motif found in a large family of functionally distinct proteins. RING finger proteins are involved in diverse cellular processes including apoptosis, DNA repair, cell cycle, signal transduction, tumour suppressor, vesicular transport, and peroxisomal biogenesis. RING finger protein 38 (RNF38) is a member of the family whose functions remain unknown. To gain insight into the putative effects of RNF38 in the central nervous system, we localised its expression. The aim of this study was to identify the neuroanatomical location(s) of rnf38 mRNA and its peptide, determine the type of RNF38-expressing cells, and measure rnf38 gene expression in the brain of male tilapia. The distributions of rnf38 mRNA and its peptide were visualised using in situ hybridisation with digoxigenin-labelled RNA antisense and immunocytochemistry, respectively. Both were identically distributed throughout the brain, including the telencephalon, preoptic area, optic tectum, hypothalamus, cerebellum, and the hindbrain. Double-labelling immunocytochemistry for RNF38 and the neuronal marker HuC/D showed that most but not all RNF38 protein was expressed in neuronal nuclei. Quantitative real-time polymerase chain reaction showed the highest level of rnf38 mRNA in the midbrain, followed by the preoptic area, cerebellum, optic tectum, telencephalon, hindbrain and hypothalamus. These findings reveal a differential spatial pattern of RNF38 in the tilapia brain, suggesting that it has potentially diverse functions related to neuronal activity.
Keywords: immunocytochemistry, in situ hybridisation, midbrain, preoptic area, brain
Introduction
Really interesting new gene (RING) finger domain is a zinc-binding motif found in a large family of functionally distinct proteins (Eisenberg et al., 2002). RING finger proteins are involved in diverse cellular functions such as regulating apoptosis (cIAP1, MDM2, and SIAH1) (Roperch et al., 1999; Hu and Yang, 2003; Kojima et al., 2005), cell cycle (APC11 and RBX1) (Gmachl et al., 2000; Jia et al., 2009), DNA repair (BRCA1, RAD5, and RAD18) (Gowen et al., 1998; Ulrich and Jentsch, 2000), signal transduction (TRAF2) (Vallabhapurapu et al., 2008), tumour suppression (BRCA1, CBL, and SIAH1) (Roperch et al., 1999; Shakya et al., 2008; Sanada et al., 2009), vesicular transport (PEP5 and RMA1) (Srivastava et al., 2000; Matsuda et al., 2001), and peroxisomal biogenesis (PEX2 and PEX12) (Platta et al., 2009). Proteins containing the RING finger motif are capable of mediating ubiquitination and thus, act as E3 ubiquitin ligases. Ubiquitination is achieved when ubiquitin is covalently attached to proteins via an enzymatic cascade involving ubiquitin-activating enzyme (E1), ubiquitin-conjugating enzyme (E2), and ubiquitin-ligating enzyme (E3) (Wilkinson, 1987; Deshaies and Joazeiro, 2009). However, not all RING-containing proteins are E3 ubiquitin ligases.
RING finger protein 38 (RNF38) possesses a conserved RING-H2 domain among species, which consists of Cys-X2-Cys-X(9−39)-Cys-X(1−3)-His-X(2−3)-His-X2-Cys-X(4−48)-Cys-X2-Cys, where Cys is a cysteine residue, His is a histidine residue and X is any amino acid residue and binds two zinc atoms (Eisenberg et al., 2002). Rnf38 mRNA is widely expressed in the brain, heart, liver, spleen, placenta, and testis (Eisenberg et al., 2002), and its protein is a nuclear protein that can bind to and ubiquitinate tumour-suppressor protein p53 (Sheren and Kassenbrock, 2013) through its E3 ubiquitin ligase activity (Sheren and Kassenbrock, 2013; Buetow et al., 2015). RNF38 is often associated with ageing and cancerous diseases (Fu et al., 2006; Nacheva et al., 2010; Sato et al., 2013). For examples, rnf38 is up-regulated in the hypothalamic area during ageing (Fu et al., 2006) and concomitantly deleted during the lymphoid blast transformation of chronic myeloid leukaemia (Nacheva et al., 2010).
RING finger proteins such as antidepressant-related gene 34 (ADRG34) and RING finger protein 180 (RNF180/RINES) are modulated by the serotonergic system (Yamada et al., 2000; Kabayama et al., 2013), which in turn regulates emotional and social behaviour (Kabayama et al., 2013). Recently, chronic treatment of the medaka fish with the antidepressant citalopram up-regulated brain expression of rnf38 (Moriya et al., 2015), while similar treatment of male mice yielded comparable results (Soga et al., unpublished observations). Thus, there is evidence of a relationship between RNF38 and the vertebrate serotonin system, which could influence social behaviour.
The Nile tilapia, Oreochromis niloticus, is an excellent model organism to investigate the role of RING finger proteins in social behaviour as each individual male fish exhibits a distinctive phenotype in the form of dominance and subordination. To gain insight into the putative regional influence of RNF38 in the central nervous system, we localised its expression in the brain. Rnf38 mRNA and protein were identified using in situ hybridisation (ISH) and immunocytochemistry (ICC), respectively, and double-labelling immunocytochemistry (d-ICC) was performed to determine the type of RNF38-expressing cells. Absolute rnf38 mRNA levels were examined using real-time polymerase chain reaction (PCR) to quantitatively verify the observed regional expression of ISH.
Materials and methods
Animals
Sexually mature male Nile tilapia were maintained in freshwater aquaria at 28 ± 0.5°C under a controlled natural light regime (14-h light; 10-h dark cycle) in standard fish tanks (size: 450 × 295 × 300 mm) equipped with circulating water systems and constant aeration. The fish were fed twice a day with cichlid pellets (Ziegler, Gardners, PA, USA). All experimental procedures were conducted in compliance with the guidelines of Monash University Animal Ethics Committee, AEC (MARP/2015/109).
Localisation of rnf38 mRNA
RNA probe synthesis
The primers for RNA probe synthesis were designed to amplify a 300 bp fragment of tilapia rnf38 (GenBank accession number XM 005467588.3) from whole brain cDNA. The primer sequences were as follows: forward primer, 5′-CTTGGCGAGCATCTGTCTT-3′, and reverse primer, 5′-TTCCACTTCACCATCATCGA-3′. The resulting fragment was cloned into a pGEM-T Easy Vector (Promega, Madison, WI, USA) to obtain pGEM T-Easy/rnf38 plasmid and the sequence of the cDNA-containing plasmid was confirmed by sequencing using BigDye Terminator v3.1 Cycle Sequencing kit (Applied Biosystems, Foster City, CA, USA) and 3310 Genetic Analyzer (Applied Biosystems). After confirming the sequence of the cloned tilapia rnf38 fragment (300 bp; position 2490-2789), the cDNA-containing plasmid was linearised with either Sal1 or Bsp191 restriction endonuclease and purified using Wizard SV Gel and PCR Clean-up System (Promega). Anti-sense and sense tilapia rnf38 riboprobes were synthesised from the purified linearised plasmid using MAXIscript In Vitro Transcription Kit (Ambion, Austin, TX, USA) and labelled using digoxigenin (DIG)-RNA labelling mix (Roche Diagnostics, Basel, Switzerland). The transcription mixture (10 μL) consisted of 0.5 μg linearised plasmid, ATP, CTP, and GTP (1 mM each), 0.65 mM UTP, 0.35 mM DIG-UTP, 10 mM DTT, 1 U/μL RNase Inhibitor, and T7 or SP6 RNA polymerase (0.9 or 0.95 U/μL respectively) was incubated at 37°C for 2 h. The digestive reactions were halted by the addition of 20 mM EDTA (pH 8.0), and synthesised riboprobes were purified through two precipitation steps by adding 400 mM and 37.5 μL of 100% ethanol. The riboprobes were stored at −80°C until use.
Brain tissue preparation for ISH
Sexually mature males (n = 3) were anaesthetised by immersion into 0.02% benzocaine solution (Sigma, St. Louis, MO, USA) prior to decapitation. The brains then were harvested and fixed in 4% paraformaldehyde in 0.1 M phosphate buffer (pH 7.3) for 6 h at 4°C, cryoprotected in 20% sucrose in 0.1 M phosphate buffer (pH 7.3), and embedded in frozen section compound (Leica, Wetzlar, Germany). Brain sections were sectioned in coronal plane (15 μm) using a cryostat (Leica CM1860), thaw-mounted onto silane-coated slide glass (Muto Pure Chemicals, Tokyo, Japan) and stored at −80°C until use for DIG ISH.
ISH
Brain sections were permeabilised with 0.2M HCl for 10 min, treated with proteinase K (1 μg/mL) for 15 min at 37°C, and hybridised with DIG-labelled anti-sense RNA probe (50 ng/mL) for rnf38 overnight at 55°C in a closed moist chamber. An equivalent amount of DIG-labelled sense probe was applied onto alternate sections. The sections were washed stringently after hybridisation with 2X saline sodium citrate (SSC) for 30 min at room temperature, followed by 2X SSC for 30 min at 60°C and 0.1X SSC for 30 min at 60°C, and blocked with 2% normal sheep serum. Alkaline phosphatase-conjugated anti-DIG antibody (1:500, 11093274910, RRID:AB_514497, Roche Diagnostics) was then used to detect the hybridised DIG-labelled RNA probes and 4-nitroblue tetrazolium chloride/5-bromo-4-chloro-3-indolyl-phosphate (NBT/BCIP, Roche Diagnostics) was used to develop the chromogenic reaction.
Image analysis
Section images were scanned and captured with a MIRAX MIDI slide scanner (Carl Zeiss, Oberkochen, Germany) and computer software (Pannoramic Scanner; 3DHISTECH, Budapest, Hungary). The staining density of rnf38 mRNA was subjectively scored on a four-point scale as follows: + + + (high), + + (moderate), + (low), and - (absent). Nomenclature for the brain area was adopted from Parhar (1990); Soga et al. (2005), and Ogawa et al. (2016).
Localisation of RNF38 protein in the brain
Brain tissue preparation for ICC
Sexually mature males (n = 2) were anaesthetised by immersion into 0.02% benzocaine solution (Sigma) prior to decapitation. The brains were then harvested and fixed in 4% paraformaldehyde in 0.1 M phosphate buffer (pH 7.3) for 6 h at 4°C, cryoprotected in 20% sucrose in 0.1 M phosphate buffer (pH 7.3), and embedded in frozen section compound (Leica). The brains were sectioned in coronal plane (15 μm) using a cryostat (Leica CM1860) and mounted on silane-coated glass slides (Muto Pure Chemicals) and stored at −80°C until use for ICC.
ICC
Primary polyclonal rabbit anti-RNF38 antiserum (1:100, ab121487, RRID:AB_11128227, Abcam, Cambridge, UK) including 0.5% Triton-X and 2% normal goat serum was applied to each section, and the slides were incubated in a closed moist chamber for 48 h at 4°C. Pre-absorption of the primary antiserum with 5 μg/ml RNF38 antigen (VVFSGQHLPVCSVPPPMLQACSVQHLPVPY AAFPPLISSDPFLIHPPHLSPHHPPHLPPPGQFVPFQTQQSRSPLQRIENEVELLGEHLPVGGFTYPPSAHPPTLPPSAPL, ab165637, Abcam) was applied onto alternate sections. The sections were then incubated in biotinylated anti-rabbit immunoglobulin IgG (1:200) for 30 min and avidin-biotinylated horseradish peroxidase complex (1:50) for 45 min at room temperature (PK-6101, RRID:AB_2336820, Vectastain ABC Elite Kit, Vector Laboratories, Burlingame, CA, USA). Antigen-antibody complexes were visualised with Alexa Fluor 488 Streptavidin (1:500; S32354, RRID:AB_2315383, Invitrogen Corporation, Carlsbad, CA, USA).
Image analysis
Section images were scanned and captured with a MIRAX MIDI slide scanner (Carl Zeiss) with an appropriate excitation filter for Alexa Fluor 488 and computer software (Pannoramic Scanner). RNF38 peptide staining density was subjectively scored on a four-point scale as follows: + + + (high), + + (moderate), + (low), and – (absent). Nomenclature for the brain area was adopted from Parhar (1990); Soga et al. (2005), and Ogawa et al. (2016).
Type of RNF38-expressing cells
Brain tissue preparation for d-ICC
A sexually mature male (n = 1) was anaesthetised by immersion into 0.02% benzocaine solution (Sigma, St. Louis, MO, USA) prior to decapitation. The brain was then harvested and fixed in 4% paraformaldehyde in 0.1 M phosphate buffer (pH 7.3) for 6 h at 4°C, cryoprotected in 20% sucrose in 0.1 M phosphate buffer (pH 7.3), and embedded in frozen section compound (Leica). The brain was sectioned in sagittal plane (15 μm) using a cryostat (Leica CM1860) and mounted on silane-coated glass slides (Muto Pure Chemicals) and stored at −80°C until use for d-ICC.
d-ICC of RNF38 with neuronal marker HuC/D or glial fibrillary acidic protein (GFAP)
Primary polyclonal rabbit anti-RNF38 antiserum (1:100, ab121487, RRID:AB_11128227, Abcam) including 0.5% Triton-X and 2% normal goat serum was applied to each section, and the slides were incubated in a closed moist chamber for 48 h at 4°C. The sections then were visualised with goat anti-rabbit IgG secondary antibody, Alexa Fluor 594 conjugated (1.400, A11037, RRID:AB_2534095, Thermo Fisher Scientific, Waltham, MA, USA). Then, either primary monoclonal mouse anti-HuC/D antiserum (1:500, A21271, RRID:AB_221448, Thermo Fisher Scientific) or primary polyclonal rabbit anti-GFAP antiserum (1:500, Z0334, RRID:AB_10013382, Dako, Glostrup, Denmark) including 0.5% Triton-X and 2% normal goat serum was applied to alternate sections, and the slides were incubated in a humidified chamber for 24 h at 4°C. The sections were visualised with goat anti-mouse IgG secondary antibody, Alexa Fluor 488 conjugated (1.500, A11001, RRID:AB_2534069, Thermo Fisher Scientific) or goat anti-rabbit IgG secondary antibody, Alexa Fluor 488 conjugated (1:500, A11008, RRID:AB_143165, Thermo Fisher Scientific).
Image analysis
Section images were captured using a Nikon fluorescent microscope (Eclipse 90i, Nikon, Tokyo, Japan), equipped with a Nikon DXM 1200C camera and NIS-Element 3.0 software.
Absolute quantification of rnf38 mRNA in the brain using real-time PCR
Cloning of the rnf38 gene
Primers for real-time gene expression were designed to amplify a 101 bp fragment of tilapia rnf38 (GenBank accession number XM 005467588.3) from whole brain cDNA. The primer sequences were as follows: forward primer, 5′-TCTGTGGTCTTCAGTGGTCAA-3′, and reverse primer, 5′- GGGAATGGGTAGGGCATT-3′. The resulting fragment was cloned into a pGEM-T Easy Vector (Promega) by means of RT-PCR cloning method. The primer set was designed to avoid genomic DNA amplification and achieved 97.7% amplification efficiency.
Brain tissue preparation for quantitative real-time PCR
Sexually mature males (n = 8) were anaesthetised by immersion into 0.02% benzocaine solution (Sigma, St. Louis, MO, USA) prior to decapitation. The brains were rapidly harvested and fresh frozen in frozen section compound on a block of dry ice and stored at −80°C.
Brain micro-dissection for quantitative real-time PCR
The brain micro-dissection was performed as previously described with modifications (Maruska et al., 2013). Fresh frozen whole brains were sectioned in coronal plane (60 μm) using a cryostat (Leica CM1860) and briefly thaw-mounted onto uncoated glass microscope slides (Sail Boat Lab Co., Zhejiang, China) and stored at −80°C until microdissection. Brain tissues were collected with a modified 200-μL pipette tip (ExtraGene, Taichung, Taiwan) and 25G needle (Terumo, Tokyo, Japan), and transferred into 200 μL TRIzol reagent (Thermo Fisher Scientific). To prevent cross-contamination and RNA degradation, the needle was cleaned with RNAse-Away (Thermo Scientific), ethanol, and diethyl pyrocarbonate (DEPC)-treated water in successive order between each sample. Tilapia brain atlases (Parhar, 1990; Soga et al., 2005; Ogawa et al., 2016) were used to identify the following brain regions: the telencephalon, preoptic area (including minimal portions of ventral telencephalon), optic tectum, midbrain (including minimal portions of the medullary nuclei of the hindbrain), hypothalamus (including portions of the glomerular, preglomerular, subglomerular, and posterior thalamic nuclei), cerebellum, and hindbrain (Figure 7A).
RNA extraction and cDNA synthesis
Total RNA was extracted from homogenised microdissected brain tissue with TRIzol reagent according to the manufacturer's instructions. Briefly, 40 μL chloroform was added to the homogenised samples and centrifuged at 12,000 g for 15 min at 4°C to allow separation of the biphasic mixture into a lower red, phenol-chloroform phase and upper colourless, aqueous phase. RNA was precipitated from the aqueous phase by mixing with 100 μL isopropyl alcohol. The samples were then mixed vigorously and centrifuged at 12,000 g for 15 min at 4°C. Upon centrifugation, the supernatant was removed, and the pellet was washed twice with 75% ethanol. After removal of the 75% ethanol and 15 min of drying, the RNA was resolved in 20 μL DEPC-treated water. The total RNA sample (500 ng) was reverse transcribed to cDNA in a reaction mixture of 20 μL containing 1X RT Random Primer, 1X RT buffer, 30U RNase Inhibitor, 75U Multiscribe Reverse Transcriptase with High Capacity cDNA Reverse Transcription Kit (Applied Biosystems) and oligo d(T) primer. The reaction was initiated at 25°C for 10 min for primer annealing, and 37°C for 120 min for reverse transcription, followed by 85°C for 5 min for enzyme inactivation. The cDNA samples were stored at −20°C until use.
Absolute quantitative real-time PCR
The amounts of rnf38 mRNA were determined using quantitative real-time PCR. The PCR reaction was performed in 10-μL duplicate reactions using a SensiFAST SYBR Hi-ROX Kit (Bioline, Taunton, MA, USA) with 0.2 μM each of forward and reverse primers and 1 μL cDNA. The plasmid containing rnf38 cDNA was serially diluted to concentrations of 108, 107, 106, 105, 104, 103, and 102 copy/μL as standard cDNAs for quantification. Real-time PCR was carried out using 7500 Fast Real-Time PCR system (Applied Biosystems) with conditions of 95°C for 10 min, followed by 40 cycles of 95°C for 15 s and 60°C for 1 min followed by a dissociation step. Single PCR products were verified by melting curve analysis, and the sizes of PCR product were confirmed with agarose gel electrophoresis.
Statistical analysis
Data are expressed as the mean values with standard error (SEM) per group.
Results
Partial sequence of rnf38 cDNA
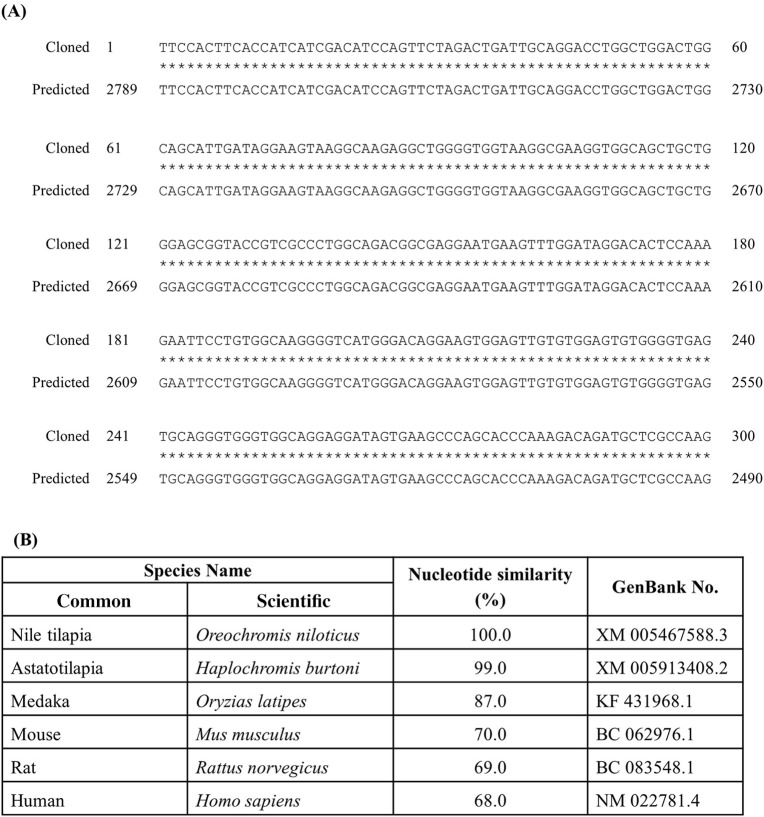
The identified partial length of rnf38 cDNA consisted of 300 nucleotides spanning 2 exons when compared to the tilapia genome (Figure 1A). The identified partial sequence of rnf38 cDNA cloned from Nile tilapia showed 100% homology to the predicted sequence in the GenBank (XM 005467588.3) (Figure 1B), and there was a high degree of homology between the partial sequence with the rnf38 genes of other species such as astatotilapia (Haplochromis burtoni), medaka (Oryzias latipes), mouse (Mus musculus), rat (Rattus norvegicus), and human (Homo sapiens) (Figure 1B).
Figure 1.
Cloned sequence of tilapia rnf38 mRNA. (A) Alignment of cloned rnf38 partial sequence with predicted sequence from GenBank (XM 005467588.3; position 2,490–2,789). (B) Nucleotide similarity between identified tilapia rnf38 partial sequences with astatotilapia (Haplochromis burtoni), medaka (Oryzias latipes), mouse (Mus musculus), rat (Rattus norvegicus), and human (Homo sapiens).
ISH and ICC controls
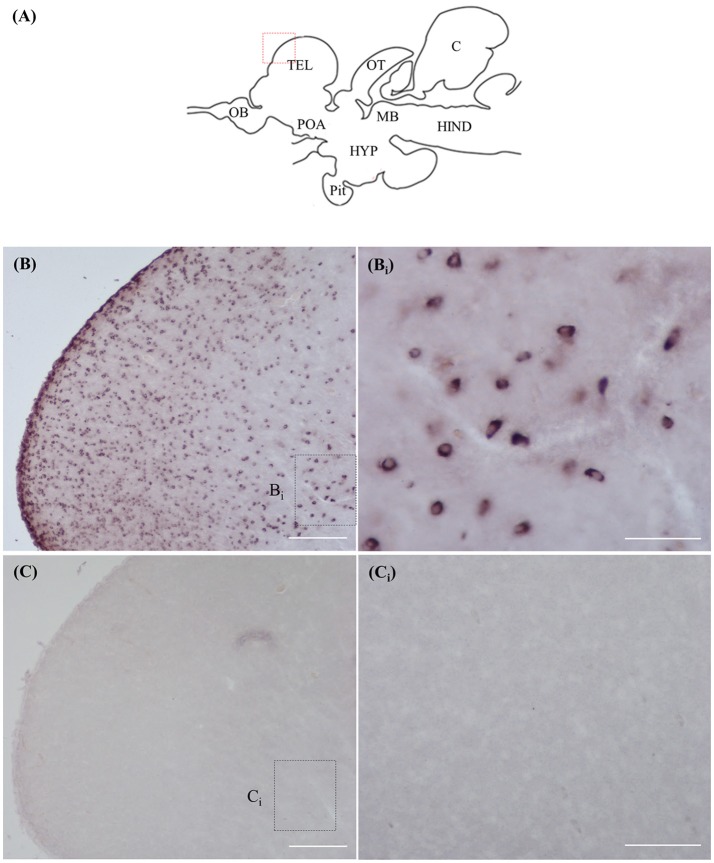
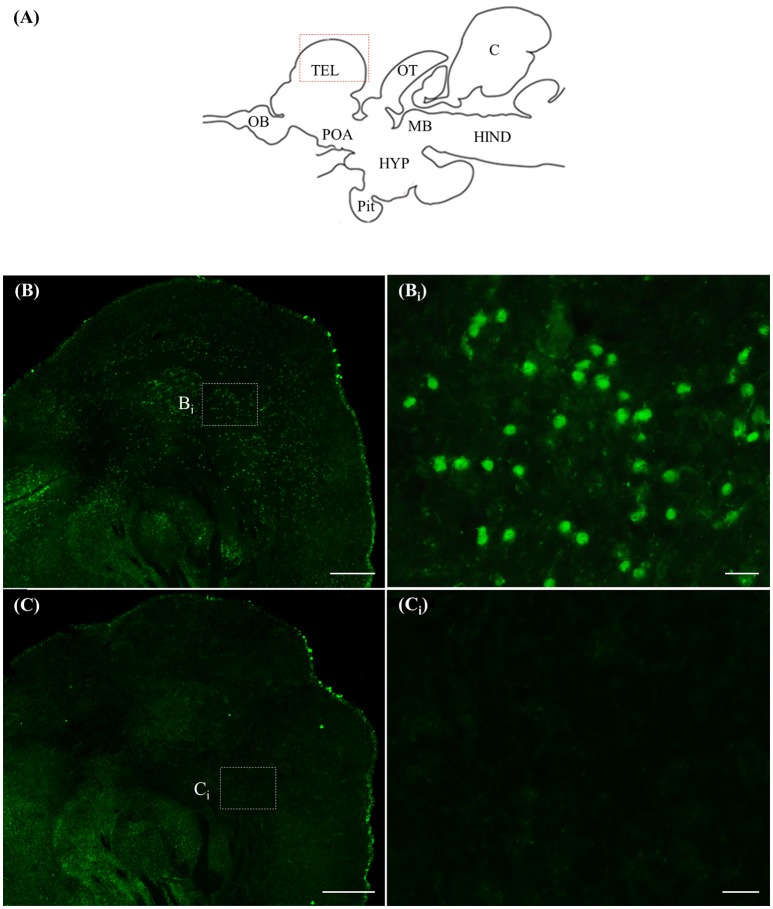
Staining with the rnf38 anti-sense RNA probe showed distinct cell labelling (Figures 2A,B,Bi), whereas staining with the rnf38 sense RNA probe showed no hybridisation signal (Figures 2C,Ci). As for ICC, staining with RNF38 antibody revealed clear nuclear labelling (Figures 3A,B,Bi), whereas there was no staining with antibody pre-absorbed with the RNF38 antigen (Figures 3C,Ci).
Figure 2.
In situ hybridisation of rnf38 in tilapia brain. (A) Schematic sagittal diagram of tilapia brain. (B,Bi) In situ hybridisation staining with rnf38 anti-sense RNA probe shows distinct cell-labelling. (Bi) shows a higher magnification of the boxed area in (B). (C,Ci). In situ hybridisation staining with rnf38 sense RNA probe shows no hybridisation signal. Ci shows a higher magnification of the boxed area in (C). Scale bars: (B,C), 200 μm; (Bi,Ci), 50 μm.
Figure 3.
Immunocytochemistry of RNF38 in tilapia brain. (A) Schematic sagittal diagram of tilapia brain. (B,Bi) Immunocytochemistry staining with RNF38 antibody shows distinct nuclear-labelling. (Bi) shows a higher magnification of the boxed area in (B). (C,Ci) Nuclear staining is completely blocked when the antibody is pre-absorbed with the RNF38 antigen prior to immunocytochemical procedure. (Ci) shows a higher magnification of the boxed area in (C). Scale bars: (B,C), 20 μm; (Bi,Ci), 20 μm.
Localisation of rnf38 mRNA and RNF38 protein
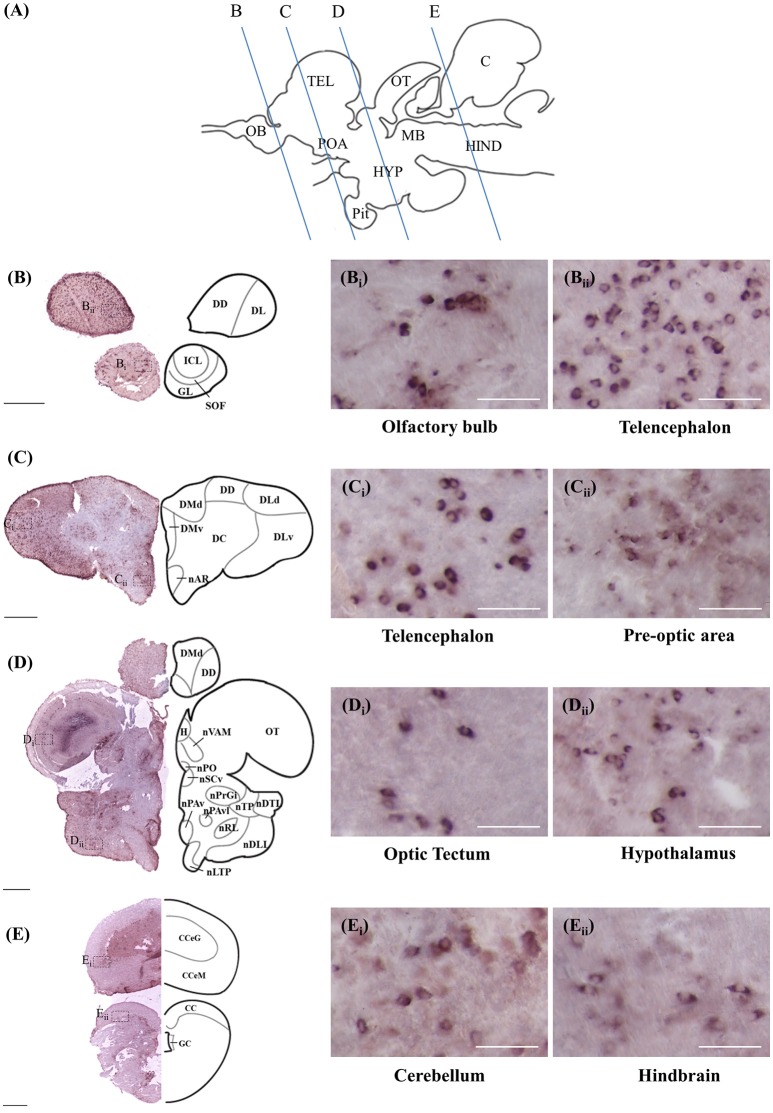
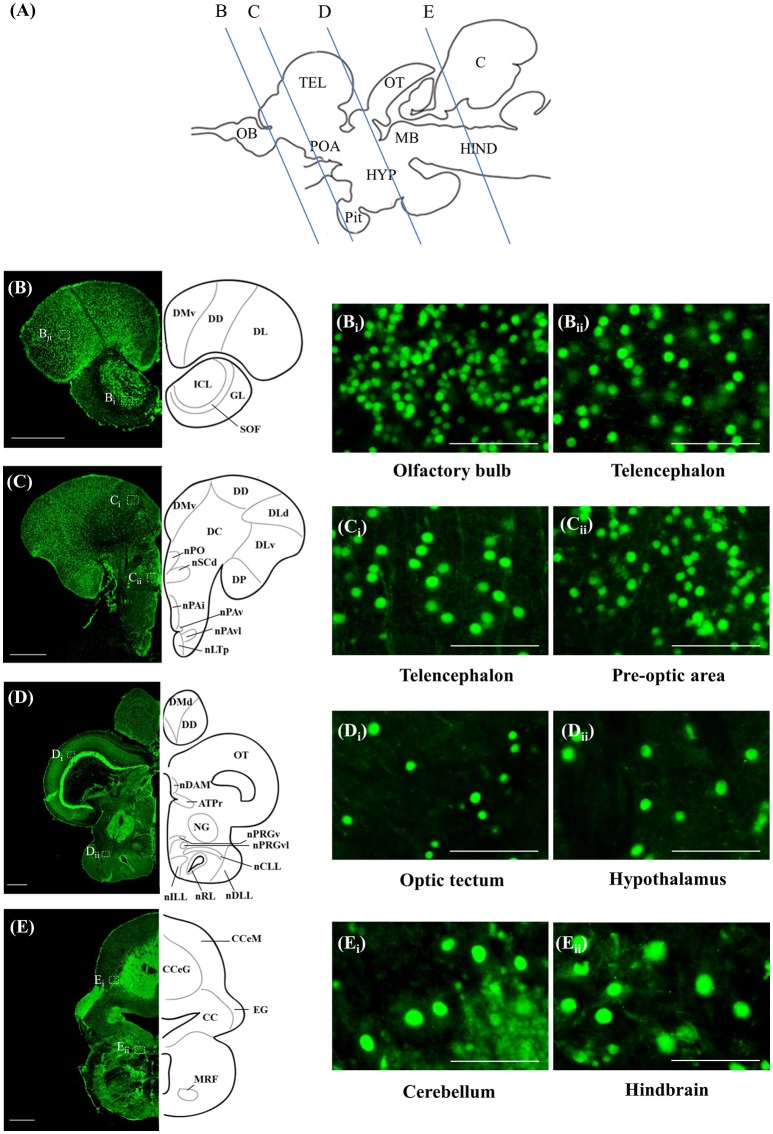
ISH and ICC showed that rnf38 mRNA and RNF38 protein were expressed throughout the tilapia brain. Figures 4A–Eii, 5A–Eii provide an overview of the distributions of rnf38 hybridisation signals and RNF38 immunoreactivity in the brain (see Table 1).
Figure 4.
Rnf38 mRNA expression in the brain of tilapia. (A) Schematic sagittal diagram of tilapia brain with approximate locations of coronal sections (B–E). (B–E) Representative low magnification photomicrographs of coronal sections are shown on the left and outlined as a mirror image on the right. (Bi,Bii,Ci,Cii,Di,Dii,Ei,Eii) show a higher magnification of the boxed area in (B–E) respectively. Scale bars: (B–E), 500 μm; (Bi,Bii,Ci,Cii,Di,Dii,Ei,Eii), 50 μm. For abbreviations, see Table 1.
Figure 5.
RNF38 protein expression in the brain of tilapia. (A) Schematic sagittal diagram of tilapia brain with approximate locations of coronal sections (B–E). (B–E) Representative low magnification photomicrographs of coronal sections are shown on the left and outlined as a mirror image on the right. (Bi,Bii,Ci,Cii,Di,Dii,Ei,Eii) show a higher magnification of the boxed area in (B–E) respectively. Scale bars: (B–E), 500 μm; (Bi,Bii,Ci,Cii,Di,Dii,Ei,Eii), 50 μm. For abbreviations, see Table 1.
Table 1.
Distributions of rnf38 mRNA and protein in the brain of Nile tilapia, Oreochromis niloticus.
| Brain region | Abbr. | Densities of staining | |
|---|---|---|---|
| mRNA | Protein | ||
| TELENCEPHALON | |||
| Olfactory bulbs | |||
| Internal cellular layer | ICL | + + + | + + + |
| Glomerular layer | GL | + + | + + |
| Secondary olfactory fibre layer | SOF | + | + |
| Dorsal telencephalon | |||
| Dorsal zone | DD | + + + | + + + |
| Lateral zone | DL | + + + | + + + |
| Latero-dorsal area | DLd | + + + | + + + |
| Latero-ventral area | DLv | + + + | + + + |
| Central zone | DC | + + + | + + + |
| Medial zone | DM | + + + | + + + |
| Medio-anterior area | DMa | + + + | + + + |
| Medio-dorsal area | DMd | + + + | + + + |
| Medio-ventral area | DMv | + + + | + + + |
| Ventral telencephalon | |||
| Dorsal nucleus | Vd | + + + | + + + |
| Ventral nucleus | Vv | + + + | + + + |
| Supracommissural nucleus | Vs | + + + | + + + |
| Entopeduncular nucleus | nE | + + + | + + + |
| DIENCEPHALON | |||
| Preoptic area | |||
| Anterior preoptic recess nucleus | nAR | + + + | + + + |
| Nucleus preopticus | nPO | + + + | + + + |
| Suprachiasmatic nucleus | nSC | + + + | + + + |
| Dorsal zone | nSCd | + + + | + + + |
| Ventral zone | nSCv | + + + | + + + |
| Epithalamus | |||
| Habenula | H | + + + | + + + |
| Epiphysis | EP | + | + |
| Dorsal thalamus | |||
| Posterior zone | DP | + + + | + + + |
| Medial zone | nDAM | + + + | + + + |
| Lateral zone | nDVL | + + | + + |
| Glomerular nucleus | NG | + + | + + |
| Ventral thalamus | |||
| Lateral zone | nVAL | + + | + + |
| Medial zone | nVAM | + + + | + + + |
| Posterior tuberculum | |||
| Posterior tuberal nucleus | nPT | + + + | + + + |
| Posterior tuberal area | |||
| Rostral zone | ATPr | + + + | + + + |
| Lateral zone | ATPl | + + | + + |
| Preglomerular nucleus | |||
| Anterior zone | nPrGa | + + | + + |
| Dorso-lateral subdivision | nPrGdl | − | − |
| Inferior zone | nPrGi | + + + | + + + |
| Medial subdivision | nPrGm | + + + | + + + |
| Superior subdivision | nPrGs | + + + | + + + |
| Ventral subdivision | nPrGv | + + + | + + + |
| Ventro-lateral subdivision | nPrGvl | + + | + + |
| Posterior thalamic nucleus | PT | + + + | + + + |
| Subglomerular nucleus | nSbG | + + | + + |
| Posterior thalamic nucleus | nTP | + + + | + + + |
| Periventricular area | |||
| Anterior periventricular nucleus | |||
| Dorsal zone | nPAd | + + + | + + + |
| Dorso-lateral subdivision | nPAdl | + + | + + |
| Intermedial zone | nPAi | + + + | + + + |
| Ventral zone | nPAv | + + | + + |
| Ventro-lateral subdivision | nPAvl | + + + | + + + |
| Posterior periventricular nucleus | |||
| Dorsal zone | nPPd | + + + | + + + |
| Medial zone | nPPm | + + + | + + + |
| Ventral zone | nPPv | + + + | + + + |
| Hypothalamus | |||
| Anterior tuberal nucleus | nAT | + + | + + |
| Lateral tuberal nucleus | |||
| Anterior zone | nLTa | + + + | + + + |
| Lateral zone | nLTl | + + + | + + + |
| Posterior zone | nLTp | + + + | + + + |
| Rostral zone | nLTr | + + + | + + + |
| Ventral zone | nLTv | + + | + + |
| Periventricular hypothalamus | |||
| Ventral zone | Hv | + + + | + + + |
| Dorsal zone | Hd | + + + | + + + |
| Paraventricular organ | PVo | + + + | + + + |
| Diffuse nucleus of the lateral lobe | nDLL | + + + | + + + |
| Diffuse nucleus of the lateral tori | nDTL | + + + | + + + |
| Intermediate nucleus of the lateral lobe | nILL | + + + | + + + |
| Central nucleus of the lateral lobe | nCLL | + + + | + + + |
| Lateral recess nucleus | nRL | + + + | + + + |
| Posterior recess nucleus | nRP | + + + | + + + |
| Synencephalon | |||
| Nucleus of the medial longitudinal fascicle | nMLF | + + + | + + + |
| Pretectum | |||
| Pretectal area | AP | + + + | + + + |
| Pretectal nucleus | |||
| Dorsal zone | nPCd | + + + | + + + |
| Ventral zone | nPCv | + + | + + |
| Fasciculus retroflexus | FR | + | + |
| MESENCEPHALON | |||
| Optic tectum | OT | + + + | + + + |
| Semicircular torus | |||
| Semicircular torus (layer 2) | TS2 | + + + | + + + |
| Semicircular torus (layer 3) | TS3 | + + | + + |
| Tegmentum | |||
| Dorsal tegmental nucleus | DTN | + + | + + |
| Rostral tegmental nucleus | RT | + + | + + |
| Perilemniscal nucleus | pL | + + + | + + + |
| Oculomotor nucleus | NIII | + + + | + + + |
| Medial nucleus | MR | + + + | + + + |
| Superior nucleus | SR | + + + | + + + |
| RHOMBENCEPHALON | |||
| Cerebellum | |||
| Valvula cerebelli | |||
| Lateral division | Val | + | + |
| Medial division | Vam | + | + |
| Granular zone of the corpus cerebellum | CCeG | − | − |
| Molecular zone of the corpus cerebellum | CCeM | + + | + + |
| Eminentia granularis | EG | + + | + + |
| Medullary nuclei | |||
| Crista cerebellaris | CC | − | + |
| Superior reticular formation | SRF | + + + | + + + |
| Lateral reticular formation | LRF | + + | + + |
| Medial or intermediate reticular formation | MRF | + + + | + + + |
| Central gray | GC | + + | + + |
| Lateral valvular nucleus | nLV | + + + | + + + |
| Lateral longitudinal fascicle | LLF | − | − |
| Vascular lacuna of area postrema | VAS | − | − |
+ + +, high; + +, moderate; +, low; −, absent.
Telencephalon
In the olfactory bulb, there was very dense staining of positively labelled cells in the internal cell layer (ICL) (Figures 4B, 5B). Staining of positively labelled cells was also observed in the glomerular layer (GL) and secondary olfactory fibre layer (SOF) of the olfactory bulb (Figures 4B, 5B). In the dorsal telencephalon, there was robust staining of positively labelled cells in almost all subdivisions including the dorsal, lateral, and medial zones (DD, DL, and DM, respectively) (Figures 4B–D, 5B–D). However, staining intensity in the medio-ventral area of the dorsal telencephalon (DMv) was weaker compared to other telencephalic areas (Figures 4C, 5B–C). Although strong staining of positively labelled cells was observed in the central area of the dorsal telencephalon (DC), the distribution was quite diffuse (Figures 4C, 5C). Heavy staining was observed in all subdivisions of the ventral telencephalon including the dorsal, ventral, supracommissural, and entopeduncular nuclei (Vd, Vv, Vs, and nE, respectively) (Table 1).
Diencephalon
The preoptic area (POA) had very intense staining, specifically in the anterior preoptic recess nucleus (nAR), nucleus preopticus (nPO), and dorsal and ventral zones of the suprachiasmatic nucleus (nSCd and nSCv) (Table 1, Figures 4C–D, 5C). Caudal to the POA, several subdivisions of the dorsal thalamus showed positively staining including the posterior, medial, and lateral zones (DP, nDAM, and nDAL, respectively) and glomerular nucleus (NG) (Table 1, Figure 5D). Strong staining was present in the lateral and medial zones of the ventral thalamus (nVAL and nVAM) (Table 1, Figure 4D). In the epithalamus, intense signals were present in the habenula (H) (Table 1, Figure 4D). Other subdivisions of the epithalamus, such as the epiphysis (EP), also contained positively labelled cells (Table 1).
Positively labelled cells were identified in almost all subdivisions of the posterior tuberculum, which includes the posterior tuberal nucleus (nPT), rostral and lateral zones of posterior tuberal area (ATPr and ATPl), and anterior, inferior, medial, superior, ventral, and ventro-lateral subdivisions of the preglomerular nucleus (nPrGa, nPrGi, nPrGm, nPrGs, nPrGv, and nPrGvl, respectively) (Table 1, Figures 4D, 5D). Strong staining was also observed in other subdivisions of posterior tuberculum, which includes the posterior thalamic nucleus (PT), subglomerular nucleus (nSbG), and posterior thalamic nucleus (nTP) (Table 1). Hybridisation signals and immunoreactivity were absent in the dorso-lateral subdivision of preglomerular nucleus (nPrGdl) (Table 1).
Strong staining of positively labelled cells was found throughout the hypothalamus. Positively labelled cells were present in the anterior tuberal nucleus (NAT), and anterior, lateral, posterior, and rostral zones of the lateral tuberal nucleus (nLTa, nLTl, nLTp, and nLTr, respectively) of the hypothalamus (Table 1, Figure 4D). Within the periventricular hypothalamus, intense staining was observed in the ventral and dorsal zones (Hv and Hd) (Table 1). Strong staining was also found in diffuse nuclei of the lateral lobe and lateral tori (nDLL and nDTL), intermediate and central nucleus of lateral lobe (nILL and nCLL), and lateral and posterior recess nuclei (nRL and nRP) (Table 1, Figures 4D, 5D). Positively labelled cells were also present in the anterior periventricular nuclei including the dorsal, dorso-lateral, intermedial, ventral, and ventro-lateral parts (nPAd, nPAdl, nPAi, nPAv, and PAvl, respectively) (Table 1, Figure 4D). Staining was also observed in the posterior periventricular nuclei including the dorsal, medial, and ventral parts (nPPd, nPPm, and nPPv) (Table 1).
Strong staining was present in the nucleus of the medial longitudinal fascicle (nMLF), pretectal area (AP), and pretectal nucleus including the dorsal and ventral regions (nPCd and nPCv, respectively) (Table 1). Signals were also present along the border of the fasciculus retroflexus fibre tract (FR) (Table 1).
Mesencephalon
Intense staining of positively labelled cells was observed in the optic tectum (OT) (Figures 4D, 5D). Strong signals were observed in the layers 2 and 3 of the semicircular torus (TS2 and TS3), dorsal and rostral tegmental nucleus (DTN and RT), and perilemniscal nucleus (pL) (Table 1). Positive cells were also present in the oculomotor nucleus (NIII), which includes the medial and superior nuclei of the nervus oculomotorius (MR and SR) (Table 1).
Rhombencephalon
There were relatively fewer positively labelled cells in the rhombencephalon compared to telencephalon, diencephalon, and mesencephalon. Within the cerebellum, staining was observed in the ganglionic layer between the molecular and granular layers of the corpus cerebellum (CCeM and CCeG, respectively) (Figures 4E, 5E). Staining was also observed in other subdivisions of the cerebellum: the lateral and medal division of the valvula cerebelli and the eminentia granularis (Val, Vam, and EG, respectively) (Table 1). However, in the crista cerebellaris (CC) of the medullary nuclei, despite minimal RNF38 immunoreactive labelling, no hybridisation signals were observed (Figures 4E, 5E). Caudally in the rhombencephalon, other medullary nuclei also had patches of strong staining of positively labelled cells, specifically in the superior reticular formation, lateral reticular formation, and medial or intermediate reticular formation (SRF, LRF, and MRF, respectively) (Table 1). Strong staining was also observed in the central gray (CG) and lateral valvular nucleus (nLV) (Table 1, Figure 4E). No signal was observed in the lateral longitudinal fascicle (LLF) or vascular lacuna of the area postrema (VAS) (Table 1).
Type of RNF38-expressing cells
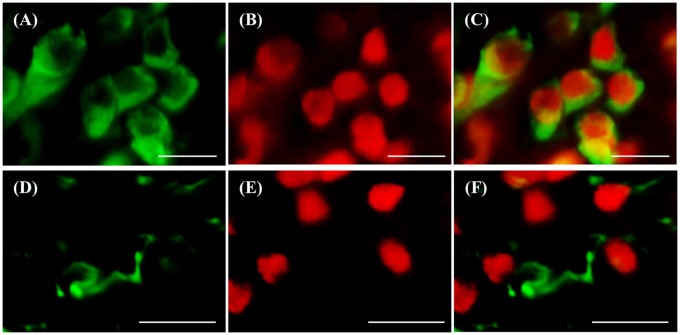
d-ICC for RNF38 and the neuronal marker HuC/D showed that most RNF38 protein, although not all, was expressed in neuronal nuclei (Figures 6A–C). Correspondingly, d-ICC for RNF38 and GFAP showed that RNF38 was absent in astroglial cells (Figures 6D–F).
Figure 6.
Double-labelling immunocytochemistry of RNF38 protein with neuronal marker HuC/D or glial marker GFAP. (A–C) Neuronal marker HuC/D (A, green), RNF38 peptide (B, red), merged image (C). (D–F) Glial marker GFAP (D, green), RNF38 protein (E, red) and merged image (F). Scale bars: (A–F), 10 μm.
Levels of rnf38 mRNA in the brain
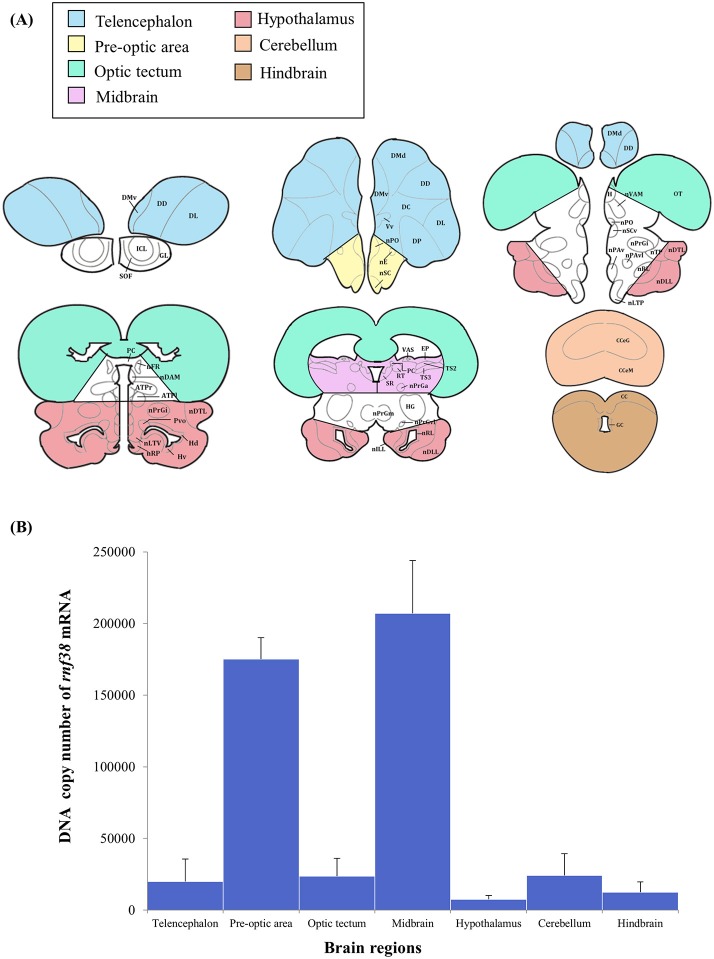
Real-time PCR was performed to examine rnf38 gene expression in seven brain regions: the telencephalon, preoptic area, optic tectum, midbrain, hypothalamus, cerebellum and hindbrain. The highest levels of rnf38 mRNA were detected in the midbrain, followed by the preoptic area, cerebellum, optic tectum, telencephalon, hindbrain and hypothalamus (Figure 7B). The regional distribution of rnf38 mRNA detected using real-time PCR corresponded well with that of rnf38 mRNA visualised by ISH.
Figure 7.
Distribution of rnf38 mRNA in the telencephalon, preoptic area, optic tectum, midbrain, hypothalamus, cerebellum and hindbrain of tilapia. (A) Brain areas collected during microdissection (telencephalon, n = 5; preoptic area, n = 6; optic tectum, n = 8; midbrain, n = 7; hypothalamus, n = 6; cerebellum, n = 5; and hindbrain, n = 5) for rnf38 gene expression study. (B) DNA copy number of rnf38 mRNA in telencephalon, optic tectum, hypothalamus, cerebellum and hindbrain of male tilapia. Data represent the means ± SEM.
Discussion
rnf38 gene sequence
To the best of our knowledge, we used genomic cloning to successfully obtain a partial sequence of rnf38 cDNA in the Nile tilapia. It showed 100% homology to the predicted sequences in GenBank (XM 005467588.3), thus confirming the partial sequence of tilapia rnf38. Comparison with rnf38 genes of other species such as Haplochromis burtoni (99%, GenBank XM 005913408.2), Oryzias latipes (87%, GenBank KF 431968.1), Mus musculus (70%, GenBank BC 062976.1), Rattus norvegicus (69%, GenBank BC 083548.1), and Homo sapiens (68%, GenBank NM 022781.4) showed high homology. This indicates that the gene is significantly conserved from an evolutionary perspective, suggesting functional relevance.
Expression of rnf38 mRNA and RNF38 protein in the brain
The present study examined the distribution of positively labelled cells in the brain of the Nile tilapia using ISH and ICC. Positively labelled cells in ISH are indicative of rnf38 expression, while those from ICC are indicative of the presence of the protein. Several lines of evidence show that the RNA probe and antibody can specifically identify the expression of rnf38 mRNA and its peptide, respectively. Firstly, the labelled cells were not observed in sections labelled with the sense probe, and pre-absorption of the antibody with antigen blocked all immunostaining. Second, there was good correspondence between rnf38 mRNA expression patterns and RNF38 protein immunoreactivity. As specificity has been demonstrated for the RNF38 probe and antibody, they will be useful for future investigations of the roles of RNF38 in social behaviour. Such studies may include examining changes in rnf38 mRNA levels during social defeat in Nile tilapia. The antibody will also be useful for double-labelling experiments with serotonergic neurons to clarify the neurochemical properties of RNF38 cells.
ISH and ICC showed that rnf38 mRNA and protein are widely distributed throughout the brain. In the quantitative assay, real-time PCR demonstrated that the rnf38 mRNA expression corresponded to the distribution of rnf38 mRNA as visualised by ISH. Via real-time PCR, the expression of rnf38 mRNA was previously reported in the telencephalon, optic tectum, hypothalamus, cerebellum, and hindbrain of the medaka fish (Moriya et al., 2015), as well as in the hypothalamus of male mice (Fu et al., 2006). Widespread RNF38 expression throughout the brain indicates its potential involvement in a variety of cellular functions.
ISH revealed substantially more positively labelled cells in the telencephalon compared to the hindbrain; however, real-time PCR data showed little difference between the two regions. This discrepancy may be due to that the real-time PCR data reflects rnf38 mRNA copy number per 500 ng of RNA, whereas ISH indicates the cells expressing rnf38 mRNA rather than the amount of mRNA. It is highly possible that to compensate for the smaller number of cells compared to the telencephalon, each rnf38-positive cell in the hindbrain expresses more mRNA per cell. Notably, the discrepancy is not due to absence of normalisation: expression of the housekeeping gene B-actin was stable in all brain regions (data not shown).
In the present study, real-time PCR showed high rnf38 mRNA expression in the midbrain, an area implicated in monoamine synthesis by serotonergic neurons (Loveland et al., 2014). Furthermore, chronic treatment of the medaka fish with the antidepressant citalopram up-regulated brain expression of rnf38 (Moriya et al., 2015), as did similar treatment of male mice (Soga et al., unpublished observations). This suggests a relationship between RNF38 and the serotonergic system. Therefore, it is possible that rnf38 could potentially regulate the serotonergic system or vice versa. Rnf38 mRNA was highly expressed in the preoptic area, which suggests its potential role in behaviour and reproduction (Liu et al., 1997; Foran and Bass, 1999; Goodson and Bass, 2000; Floody et al., 2011; Floody, 2014).
Functional considerations
The widespread expression of RNF38 throughout the brain indicates its potential involvement in a variety of cellular functions. Previous studies have shown that RNF38 is an E3 ubiquitin ligase, and protein ubiquitination is a highly regulated cellular process (Deshaies and Joazeiro, 2009; Sheren and Kassenbrock, 2013). Furthermore, RNF38 can interact with and activate the ubiquitin-conjugating enzyme E2 (Buetow et al., 2015), thus potentially acting as an intrinsic factor that mediates ubiquitination via the RING finger domain. Since RNF38 is a nuclear protein capable of ubiquitinating p53 (Sheren and Kassenbrock, 2013), it is therefore a potential regulator of p53 signalling. In addition, our results show that RNF38 protein is localised in the nucleus. The presence of a coiled-coil domain near the RING finger domain of rnf38, a region involved in protein-protein/DNA interaction, predicts that RNF38 possesses DNA-binding and transcriptional activities (Morett and Bork, 1999; Moriya et al., 2015). Our results also provide the first evidence that RNF38 is expressed in neurons, suggesting that it possesses neuronal activity.
Conclusion
In summary, we successfully identified the partial sequence of rnf38, and localised rnf38 mRNA and RNF38 protein in the brain of Nile tilapia using ISH and ICC respectively. The rnf38 mRNA and its peptide were distributed throughout the tilapia brain. RNF38 expression was highest in the preoptic area and midbrain. We also successfully showed that the majority of RNF38 protein, although not all, was expressed in neuronal nuclei. This work established the neuroanatomical distribution of RNF38 in the tilapia fish brain. Based on its widespread, differential localisation, RNF38 appears to have a broad, and possibly diverse influence on central nervous system function.
Author contributions
KC conducted all experiments and analysed the data together with TS. KC wrote this manuscript. TS and IP designed the experiment, did data analysis and edited the manuscript together. TS and IP received a research grant from Malaysia government for this project.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Footnotes
Funding. This work was supported by research grant from Monash University, Malaysia and Fundamental Research Grant Scheme (FRGS), FRGS/2/2014/STO3/MUSM/02/01 from the Ministry of Higher Education, Malaysia.
References
- Buetow L., Gabrielsen M., Anthony N. G., Dou H., Patel A., Aitkenhead H., et al. (2015). Activation of a primed RING E3-E2-ubiquitin complex by non-covalent ubiquitin. Mol. Cell 58, 297–310. 10.1016/j.molcel.2015.02.017 [DOI] [PubMed] [Google Scholar]
- Deshaies R. J., Joazeiro C. A. (2009). RING domain E3 ubiquitin ligases. Annu. Rev. Biochem. 78, 399–434. 10.1146/annurev.biochem.78.101807.093809 [DOI] [PubMed] [Google Scholar]
- Eisenberg I., Hochner H., Levi T., Yelin R., Kahan T., Mitrani-Rosenbaum S. (2002). Cloning and characterization of a novel human gene RNF38 encoding a conserved putative protein with a RING finger domain. Biochem. Biophys. Res. Commun. 294, 1169–1176. 10.1016/S0006-291X(02)00584-3 [DOI] [PubMed] [Google Scholar]
- Floody O. R. (2014). Role of acetylcholine in control of sexual behavior of male and female mammals. Pharmacol. Biochem. Behav. 120, 50–56. 10.1016/j.pbb.2014.02.007 [DOI] [PubMed] [Google Scholar]
- Floody O. R., Katin M. J., Harrington L. X., Schassburger R. L. (2011). Cholinergic control of male mating behavior in hamsters: effects of central oxotremorine treatment. Pharmacol. Biochem. Behav. 100, 299–310. 10.1016/j.pbb.2011.08.030 [DOI] [PubMed] [Google Scholar]
- Foran C. M., Bass A. H. (1999). Preoptic GnRH and AVT: axes for sexual plasticity in teleost fish. Gen. Comp. Endocrinol. 116, 141–152. 10.1006/gcen.1999.7357 [DOI] [PubMed] [Google Scholar]
- Fu C., Hickey M., Morrison M., McCarter R., Han E.-S. (2006). Tissue specific and non-specific changes in gene expression by aging and by early stage CR. Mech. Ageing Dev. 127, 905–916. 10.1016/j.mad.2006.09.006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gmachl M., Gieffers C., Podtelejnikov A. V., Mann M., Peters J. M. (2000). The RING-H2 finger protein APC11 and the E2 enzyme UBC4 are sufficient to ubiquitinate substrates of the anaphase-promoting complex. Proc. Natl. Acad. Sci. U.S.A. 97, 8973–8978. 10.1073/pnas.97.16.8973 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goodson J. L., Bass A. H. (2000). Forebrain peptides modulate sexually polymorphic vocal circuitry. Nature 403, 769–772. 10.1038/35001581 [DOI] [PubMed] [Google Scholar]
- Gowen L. C., Avrutskaya A. V., Latour A. M., Koller B. H., Leadon S. A. (1998). BRCA1 required for transcription-coupled repair of oxidative DNA damage. Science 281, 1009–1012. 10.1126/science.281.5379.1009 [DOI] [PubMed] [Google Scholar]
- Hu S., Yang X. (2003). Cellular inhibitor of apoptosis 1 and 2 are ubiquitin ligases for the apoptosis inducer Smac/DIABLO. J. Biol. Chem. 278, 10055–10060. 10.1074/jbc.M207197200 [DOI] [PubMed] [Google Scholar]
- Jia L., Soengas M. S., Sun Y. (2009). ROC1/RBX1 E3 ubiquitin ligase silencing suppresses tumor cell growth via sequential induction of G2-M arrest, apoptosis, and senescence. Cancer Res. 69, 4974–4982. 10.1158/0008-5472.CAN-08-4671 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kabayama M., Sakoori K., Yamada K., Ornthanalai V. G., Ota M., Morimura N., et al. (2013). Rines E3 ubiquitin ligase regulates MAO-A levels and emotional responses. J. Neurosci. 33, 12940–12953. 10.1523/JNEUROSCI.5717-12.2013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kojima K., Konopleva M., Samudio I. J., Shikami M., Cabreira-Hansen M., McQueen T., et al. (2005). MDM2 antagonists induce p53-dependent apoptosis in AML: implications for leukemia therapy. Blood 106, 3150–3159. 10.1182/blood-2005-02-0553 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Y.-C., Salamone J. D., Sachs B. D. (1997). Lesions in medial preoptic area and bed nucleus of stria terminalis: differential effects on copulatory behavior and noncontact erection in male rats. J. Neurosci. 17, 5245–5253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Loveland J. L., Uy N., Maruska K. P., Carpenter R. E., Fernald R. D. (2014). Social status differences regulate the serotonergic system of a cichlid fish, Astatotilapia burtoni. J. Exp. Biol. 217, 2680–2690. 10.1242/jeb.100685 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maruska K. P., Zhang A., Neboori A., Fernald R. D. (2013). Social opportunity causes rapid transcriptional changes in the social behaviour network of the brain in an African cichlid fish. J. Neuroendocrinol. 25, 145–157. 10.1111/j.1365-2826.2012.02382.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsuda N., Suzuki T., Tanaka K., Nakano A. (2001). Rma1, a novel type of RING finger protein conserved from Arabidopsis to human, is a membrane-bound ubiquitin ligase. J. Cell Sci. 114, 1949–1957. [DOI] [PubMed] [Google Scholar]
- Morett E., Bork P. (1999). A novel transactivation domain in parkin. Trends Biochem. Sci. 24, 229–231. 10.1016/S0968-0004(99)01381-X [DOI] [PubMed] [Google Scholar]
- Moriya S., Khel N. B., Parhar I. S. (2015). Cloning and serotonergic regulation of RING finger protein38 (rnf38) in the brain of medaka (Oryzias latipes). Neuroscience 294, 109–115. 10.1016/j.neuroscience.2015.03.012 [DOI] [PubMed] [Google Scholar]
- Nacheva E. P., Brazma D., Virgili A., Howard-Reeves J., Chanalaris A., Gancheva K., et al. (2010). Deletions of immunoglobulin heavy chain and T cell receptor gene regions are uniquely associated with lymphoid blast transformation of chronic myeloid leukemia. BMC Genomics 11:41. 10.1186/1471-2164-11-41 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ogawa S., Sivalingam M., Biran J., Golan M., Anthonysamy R. S., Levavi-Sivan B. S., et al. (2016). Distribution of LPXRFa, a gonadotropin-inhibitory hormone ortholog peptide, and LPXRFa receptor in the brain and pituitary of the tilapia. J. Comp. Neurol. 524, 2753–2775. 10.1002/cne.23990 [DOI] [PubMed] [Google Scholar]
- Parhar I. S. (1990). Neuroendocrinology of Oreochromis mossambicus. Ph.D. thesis. [Google Scholar]
- Platta H. W., El Magraoui F., Bäumer B. E., Schlee D., Girzalsky W., Erdmann R. (2009). Pex2 and pex12 function as protein-ubiquitin ligases in peroxisomal protein import. Mol. Cell. Biol. 29, 5505–5516. 10.1128/MCB.00388-09 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roperch J.-P., Lethrone F., Prieur S., Piouffre L., Israeli D., Tuynder M., et al. (1999). SIAH-1 promotes apoptosis and tumor suppression through a network involving the regulation of protein folding, unfolding, and trafficking: identification of common effectors with p53 and p21Waf1. Proc. Natl. Acad. Sci. U.S.A. 96, 8070–8073. 10.1073/pnas.96.14.8070 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanada M., Suzuki T., Shih L.-Y., Otsu M., Kato M., Yamazaki S., et al. (2009). Gain-of-function of mutated C-CBL tumour suppressor in myeloid neoplasms. Nature 460, 904–908. 10.1038/nature08240 [DOI] [PubMed] [Google Scholar]
- Sato T., Tran T. H., Peck A. R., Liu C., Ertel A., Lin J., et al. (2013). Global profiling of prolactin-modulated transcripts in breast cancer in vivo. Mol. Cancer 12:59. 10.1186/1476-4598-12-59 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shakya R., Szabolcs M., McCarthy E., Ospina E., Basso K., Nandula S., et al. (2008). The basal-like mammary carcinomas induced by Brca1 or Bard1 inactivation implicate the BRCA1/BARD1 heterodimer in tumor suppression. Proc. Natl. Acad. Sci. U.S.A. 105, 7040–7045. 10.1073/pnas.0711032105 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sheren J. E., Kassenbrock C. K. (2013). RNF38 encodes a nuclear ubiquitin protein ligase that modifies p53. Biochem. Biophys. Res. Commun. 440, 473–478. 10.1016/j.bbrc.2013.08.031 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Soga T., Ogawa S., Millar R. P., Sakuma Y., Parhar I. S. (2005). Localization of the three GnRH types and GnRH receptors in the brain of a cichlid fish: insights into their neuroendocrine and neuromodulator functions. J. Comp. Neurol. 487, 28–41. 10.1002/cne.20519 [DOI] [PubMed] [Google Scholar]
- Srivastava A., Woolford C. A., Jones E. W. (2000). Pep3p/Pep5p complex: a putative docking factor at multiple steps of vesicular transport to the vacuole of Saccharomyces cerevisiae. Genetics 156, 105–122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ulrich H. D., Jentsch S. (2000). Two RING finger proteins mediate cooperation between ubiquitin-conjugating enzymes in DNA repair. EMBO J. 19, 3388–3397. 10.1093/emboj/19.13.3388 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vallabhapurapu S., Matsuzawa A., Zhang W., Tseng P.-H., Keats J. J., Wang H., et al. (2008). Nonredundant and complementary functions of TRAF2 and TRAF3 in a ubiquitination cascade that activates NIK-dependent alternative NF-κB signaling. Nat. Immunol. 9, 1364–1370. 10.1038/ni.1678 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilkinson K. (1987). Protein ubiquitination: a regulatory post-translational modification. Anticancer. Drug Des. 2, 211–229. [PubMed] [Google Scholar]
- Yamada M., Yamazaki S., Takahashi K., Nishioka G., Kudo K., Ozawa H., et al. (2000). Identification of a novel gene with RING-H2 finger motif induced after chronic antidepressant treatment in rat brain. Biochem. Biophys. Res. Commun. 278, 150–157. 10.1006/bbrc.2000.3773 [DOI] [PubMed] [Google Scholar]