Abstract
Although XPO5 has been characterized to have tumor-suppressor features in the miRNA biogenesis pathway, the impact of altered expression of XPO5 in cancers is unexplored. Here we report a novel “oncogenic” role of XPO5 in advanced prostate cancer. Using prostate cancer models, we found that excess levels of XPO5 override the inhibitory effect of the canoncial miRNA-mRNA regulation, resulting in a global increase in proteins expression. Importantly, we found that decreased expression of XPO5 could promote an increase in proteasome degradation, whereas overexpression of XPO5 leads to altered protein posttranslational modification via hyperglycosylation, resulting in cellular protein stability. We evaluated the therapeutic advantage of targeting XPO5 in prostate cancer and found that knocking down XPO5 in prostate cancer cells suppressed cellular proliferation and tumor development without significantly impacting normal fibroblast cells survival. To our knowledge, this is the first report describing the oncogenic role of XPO5 in overriding the miRNAs regulation control. Furthermore, we believe that these findings will provide an explanation as to why, in some cancers that express higher abundance of mature miRNAs, fail to suppress their potential protein targets.
Introduction
While significant advances have been made in the diagnosis and treatment of prostate cancer, each year, thousands of men still die from prostate cancer in the United States from metastatic disease [1]. Current treatments provide temporary relief with hormone deprivation [2]; however, inevitably, the cancer develops androgen independence for which only modest results have been achieved by our current arsenal of chemotherapeutics. Thus, greater understanding of cellular pathways and molecular basis of prostate cancer progression is needed to treat this lethal phenotype.
MicroRNAs (miRNAs) have been previously demonstrated to play an important role in the development and progression of many cancers [3], [4], [5]. Several studies have confirmed that they also play an important role in prostate cancer progression including the development of androgen-independent disease [3]. miRNAs are small 19- to 22-nt RNA moieties that are initially transcribed as long, 5′-capped, and polyadenylated, known as primary miRNAs (pri-miRNAs) [6]. Within these transcripts, the mature miRNA sequences are locked in ~60- to 80-nucleotide hairpin structures. The canonical processing by the RNase III enzyme Drosha and its cofactor DGCR8 removes the precursor hairpin (pre-miRNA) from the pri-miRNA transcript. The resulting pre-miRNA is then exported by a uniquely involved XPO5 protein out of the nucleus to the cytoplasm where the RNase III enzyme Dicer performs a second cleavage to generate a double-stranded 21- to 23-nucleotide RNA molecule [7]. Once the dicer cleaves, the hairpin loops the RNA-induced silencing complex (RISC) and then associates with the RNA duplex, retaining one RNA strand, the mature miRNA which then sequence-specifically targets complementary messenger RNAs, leading to mRNA cleavage or translational silencing [8]. It is currently believed that the degree of complementarity between the miRNA and its target mRNA determines the method of silencing, i.e., perfect identity leads to cleavage and imperfect matching to translational repression [5].
The nuclear export step for these hairpin looped miRNAs is predominantly facilitated by the XPO5 in a Ran-GTP–dependent manner [9]. There is accumulating evidence that XPO5 is dysregulated in cancers [10], with recent evidence indicating that XPO5 directly interacts with Dicer mRNA, regulating the expression of dicer levels in some cancer cells [11]. In this study, we evaluated the impact of XPO5 levels in prostate cancer by using integrated global miRNAs and proteomic approaches identifying several oncoproteins that were significantly upregulated in the XPO5-overexpressed prostate cancer models. Importantly, we found that overexpression of XPO5 in prostate cancer cells overrides the inhibitory effect of miRNAs regulation and facilitates protein stabilization. Finally, we provide evidence that overexpression of XPO5 has a role in androgen and drug resistance in prostate cancer, and shows promise as a candidate drug target. Taken together, these novel functional properties of XPO5 contradict the previous notion of XPO5’s tumor-suppressive role in cancer, specifically prostate cancer.
Materials and Methods
Protein and Peptide Extraction from Cells for Proteomic Analysis
The cell pellet for each LAPC4 Ctr or shRNA XPO5 or overexpressed wt XPO5 from three T-75-cm flasks was first denatured in 1 ml of 8 M urea and 0.4 M NH4HCO3 and sonicated thoroughly. The protein concentration was measured using a BCA protein assay kit (Thermo Fisher Scientific Inc.). The proteins were then reduced by incubating in 12 mM Tris (2-carboxyethyl) phosphine (TCEP) for 30 minutes and alkylated by addition of 16 mM iodoacetamide at room temperature for 30 minutes in the dark. Sample was diluted with buffer (100 mM Tris-HCl, pH 7.5) containing 0.5 μg/μl trypsin and incubated at 37°C overnight. The digested proteins were checked for completion of trypsin digestion using SDS-PAGE and silver staining. Peptides were purified with C18 desalting columns and dried using SpeedVac (Savant SPD; Thermo Scientific).
iTRAQ Labeling of Global Tryptic Peptides from Cell Lines
Each iTRAQ (isobaric tags for relative and absolute quantitation) 4-plex reagent was dissolved in 70 μl of methanol. One milligram of each tryptic peptide sample was added into 250 μl of iTRAQ dissolution buffer (Sciex), mixed with iTRAQ 4-plex reagent, and incubated for 1 hour at room temperature. iTRAQ channel 114 was used to label LAPC4 stably selected with control plasmid, iTRAQ 115 was used for stably selected LAPC4 XPO5 shRNA, and iTRAQ 116 was used to label LAPC4-overexpressing XPO5. iTRAQ channel 117 was used as a replicate for LAPC4 stably selected control samples in order to determine the analytical reproducibility. The four sets of tagged peptides were combined and purified by Strong Cation Exchange column. Then, 10% of the labeled peptides were dried and resuspended into 0.4% acetic acid solution prior to fractionation for mass spectrometry analysis.
Analysis of Proteins and N-Glycans
The detailed procedures are described in our previous studies [12], [13]. Proteins were extracted from cell pellets by sonication in RIPA lysis buffer (1× Tris HCl, 1% NP-40, 0.5% sodium deoxycholate, 0.1% SDS, 2 mM EDTA, and 50 mM NaF). The supernatant was taken for protein immobilization after buffer exchange using pH 9.0 buffer (sodium citrate and sodium carbonate) using desalting column (Zeba; Life Technologies). Protein concentration was measured by a BCA assay. Samples were conjugated to Aminolink Resin (beads hereafter) by Glycoprotein Immobilization for Glycan extract (GIG). Protein immobilization was carried via reductive amination; the remaining active aldehyde sites on beads were blocked by 50 mM NaCNBH3 in 1 M Tris-HCl for 30 minutes; the sialic acids were stabilized by carbodiimide coupling using p-toluidine-EDC (42.8 mg pT in 400 μl 1 M HCl, 40 μl EDC, and 25 μl concentrated HCl). The beads were washed with 10% formic acid, 10% ACN (0.1% TFA), 1 M NaCl, and DI water (500 μl; 3×) [13].
The N-glycans were first digested from the immobilized glycoproteins using 2 μl PNGase F, 40 μl G7, and 358 μl DI water (37°C, 2 hours). The released N-glycans were purified using Carbograph [12]. The N-glycans were dried in a SpeedVac for isobaric labeling using 4-plex QUANTITY. The labeled N-glycans were pooled for Carbograph cleanup. Beads were washed subsequently with 10% ACN and DI water (500 μl; 3×) before reduction with 10 mM TCEP in 8 M urea/0.2 M NH4HCO3 (pH 8.0) (500 μl) (1 hour at 37°C) followed by alkylation (50 μl of 200 mM iodoacetamide; 30 minutes). The beads were centrifuged, and proteins on beads were digested with 50 μl trypsin (0.5 μg/μl) in 450 μl of 2 M urea/0.2 M NH4HCO3 overnight at 37°C with mixing. Peptides were purified by C18 cartridge (Waters Corporation; Milford, MA) and eluted with 500 μl 60% ACN (0.1% TFA) twice.
Expression Profiles Data Analysis
Pre-MicroRNA/Mature MicroRNA Profiling
To avoid separate sample preparation, we profiled pre-microRNA and mature microRNA expression on the same Affymetrix microarray chip (Affymetrix GeneChip miRNA 3.0 arrays) for each sample. This gives a glimpse into the relationship between profiled microRNA pre- and matured forms. We read our raw files, utilized the “robust multiarray average” for background correction, and quantile normalized our data utilizing functions defined in the “oligo” R package [14], [15], [16]. We identified premature and mature microRNA probes according to Affymetrix annotations and performed a paired t test on the median expression values between the miRNAs (premature versus mature microRNAs) in each of our sample sets.
Differential MicroRNA Expression
We employed the generalized linear model approach for differential gene expression detection which involved fitting a mixed-effects linear model for each feature to estimate micro-RNA expression differences between our groups of samples (i.e., XPO5OVEXP versus XPO5CTRL, XPO5KDOWN versus XPO5CTRL, and XPO5OVEXP versus XPO5KDOWN). This extends the simple hierarchical parametric model for general microarray experiments with arbitrary numbers of treatments and RNA samples. The method applies an empirical Bayes approach to moderate standard errors of log2 differential fold-change. In summary, for each analyzed feature, moderated t statistics, log-odds ratios of differential expression (B-statistics), and raw and adjusted P values [false discovery rate (FDR) control by the Benjamini and Hochberg method] were obtained. We performed all analyses using software packages available from the R/Bioconductor for statistical computing “Limma”.
Protein Expression Data Analysis
MSMS Data and Differential Expression Analysis
We searched our tandem mass spectrometry–derived raw data against the RefSeq protein database using the SEQUEST search engine in Proteome Discoverer v1.4. We specified oxidation of methionine, carbarmidomethylation of cysteine, and N-terminal iTRAQ modification as fixed residue modifications. We specified lysine (K) and tyrosine (Y) iTRAQ modifications as dynamic modifications. Peptide identification FDR was specified as 0.01. Parsimonious protein grouping was specified to allow at least one peptide per protein. High-confidence peptide spectrum matches (i.e, PSMs better than prespecified false discovery rate cutoff) were used for protein grouping. Peptide (PSM) and protein quantifications were based on ratios of iTRAQ reporter ions: 114, 115, 116, 117. From our sample preparation, wild-type (control) samples were labeled with the iTRAQ 114 and iTRAQ 117; XPO5 knocked-down samples were labeled with iTRAQ 115, while the overexpressed XPO5 samples were labeled with iTRAQ 116.
Our specified reporter ion quantification ratios were 115/114 (XPO5 shRNA versus control samples), 116/114 (XPO5 overexpressed versus control sample), and 117/114 (control versus control). To derive 116/115 (XPO5 overexpressed versus XPO5 knockdown) differential expression, we multiplied 116/114 values by inverse of 115/114 values (e.g., 116/114/ [115/114] − 1). For protein differential expression estimation between sample groups, Proteome Discoverer defaults to median quantification ratio of PSMs (peptides) unique to inferred proteins (protein group representative protein).
We exploited reported ratios of 117/114 for a first-level quality control of reporter ion ratios. We filtered out PSMs and associated proteins for which reported 117/114 ratios do not fall within ±2 SD of the mean of the distribution of all 117/114 ratios. For quality reporter ion ratios, we expect concordant 117/114 ratios to be about one. The further from Unison this is, we assume the less reliable the reported ratios are.
Proteome Discoverer reports coefficient of variations (CVs) of unique PSMs' reporter ion ratios. As a second-level quality control measure, we set an acceptable limit at less than or equal to 30%. We filtered out PSMs and associated protein with reporter ion ratio CVs greater than 30%.
Integrated Data Analysis
MicroRNA Targets Identification
We identified regulatory microRNAs targets using the multiMiR R package [17]. The multiMiR package and database contain miRNA-target information from 14 known databases. multiMiR subclassifies these databases as “predicted,” “validated,” and “disease-drug related.” It consists of a compilation of about 50 million records from the 14 different microRNA databases and represents about the largest compendium of microRNA-targets relationship information source. The 14 databases include 3 validated miRNA-target databases (miRecords [18], miRTarBase [19], and TarBase [20]), the 8 predicted miRNA-target databases (DIANA-microT [21], ElMMo [22], MicroCosm [23], miRanda [24], miRDB [25], PicTar [26], PITA [27], and TargetScan [28]), and the 3 disease-/drug-related miRNA databases (miR2Disease [29], Pharmaco-miR [30], and PhenomiR [31]. Their respective meta-information can be found at http://multimir.ucdenver.edu/. The multiMiR R provides an interface to the multiMiR database on the remote server. It is a collection of R functions to display information, build query, submit query to the web server, parse, and summarize results returned by the server. For a more confident and comprehensive miRNA-target interaction results, multiMiR provides a control of the score cutoff for predictions. We utilized the package default, i.e., the top 20 percent of predictions.
Results
Overexpression of miRNA Processing Machinery in Prostate Cancer Cells
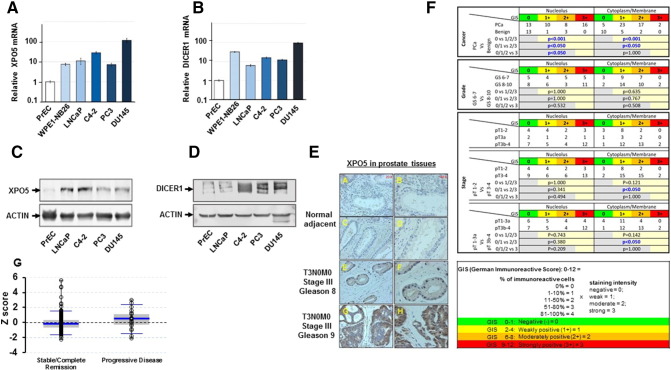
To evaluate the differential expression of Exportin-5 (XPO5) and Dicer in normal and prostate cancer cell lines, we first examined the relative mRNA expression pattern of XPO5 and Dicer using qRT-PCR. Assessing the relative mRNA expression between these cells showed a significantly higher expression of XPO5 and the Dicer mRNA in all prostate cancer cell lines (LNCaP, C4-2, PC3, and DU145) and the HPV18 genome-transformed prostate cell line (WPE1-NB26) compared to the normal epithelial PrEC cells on (Figure 1, A and B). To further validate the differential altered expression of XPO5 and Dicer1 in normal versus prostate cancer cells, we performed Western blot analysis. As shown in Figure 1, C and D and Supplemental Figure 1A, a significantly higher expression of XPO5 and Dicer1 proteins was detected in all prostate cancer cells compared to normal PrEcs or NIH3T3 cells (Figure 1, C and D and Supplemental Figure 1A). These data give us the confidence to further explore this pathway in prostate patient specimens.
Figure 1.
Quantitative RT-PCR for XPO5 (A) and DICER (B) relative mRNA expression in prostate cancer cells and in normal primary PrEC prostate epithelial cell line. Western blot analysis showing overexpression of XPO5 and DICER1 in several prostate cancer cell lines compared to the normal epithelial PrEc cells (C and D). Tissue immunohistrochemistry (IHC) staining for XPO5 protein using specimens from stage III Gleason 8 and 9 prostate cancer and normal adjacent prostate tissue (E). German immunoreactive scoring (GIS) to evaluate the correlation between the XPO5 expression and Gleason scores in prostate cancer TMA (F). Comparision of Z-scores between stable and progressive disease from the TCGA prostate cancer (PRAD) data set (G). Error bars represent the mean ± SEM.
Supplemental Figure 1.

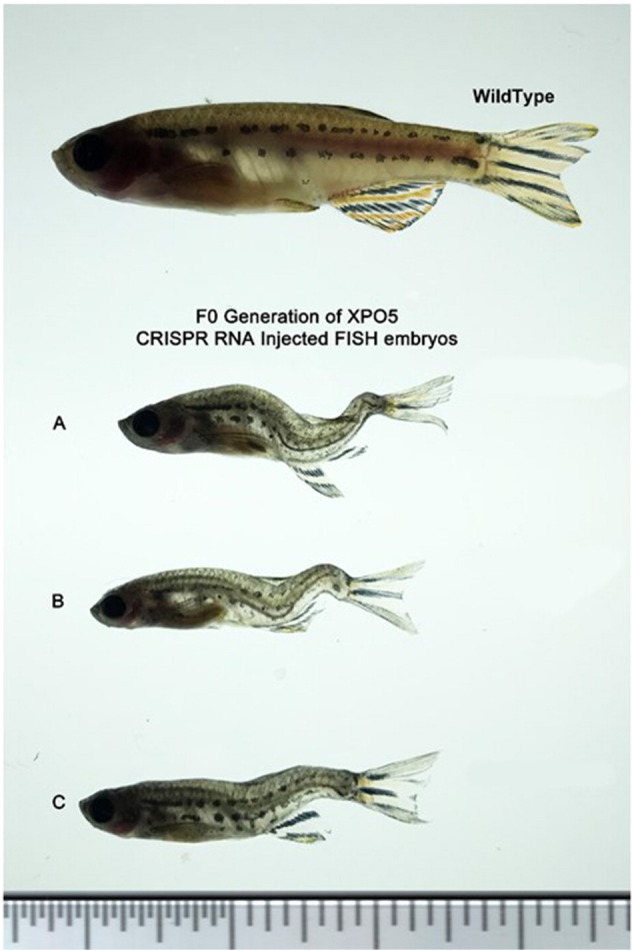
F0 generation for XPO5 zebra fish model with CRISPR RNA (A, B, C) compared to the mock-injected wild-type zebrafish suggesting embryonic deformities associated with XPO5 knockouts.
To determine whether the miRNA processing machinery was overexpressed in prostate patients, we evaluated prostate cancer specimens stage III (Gleason 8 and 9) and found a significantly higher expression of XPO5 in cancer compared to the normal adjacent prostate tissues (Figure 1E). Locally advanced, stage III prostate cancer is characterized by the invasion of the tumor cells outside the capsule to one or either side of prostate gland or to the seminal vesicles. To study whether XPO5 overexpression was correlated to Gleason scores and aggressive phenotype, we evaluate a prostate tissue microarray (TMA) using German immunoreactive scoring described in the Materials and Methods section. Briefly, tumor cells were scored based on their intensity (0 = negative to 3 = strong) in the cytoplasm or nuclear immunoreactivity. All cores in the TMAs were analyzed and scored by a clinical pathologist. These results indicated a statistically significant difference in the nuclear and cytoplasmic reactivity of XPO5 between cancer and benign disease. However, no statistical difference was observed between Gleason 6 to 7 and Gleason 8 to 10, while, on the other hand, a statistical different in the cytoplasmic expression of XPO5 between stage pT1 to 2 and pT3 to 4 disease was observed, confirming stage III disease to have a higher XPO5 expression (Figure 1F). To further evaluate whether XPO5 overexpression might be playing a role in prostate cancer progression, we interrogate the previously published TCGA (PRAD data set) for the XPO5 mRNA in prostate cancer patients with progressive (n = 21) or stable disease (n = 215), where we found a differential higher expression of XPO5 in progressive disease (P = .04) compared to patients in remission or with stable disease (Figure 1G).
Regulation of DICER and Mature miRNAs by XPO5
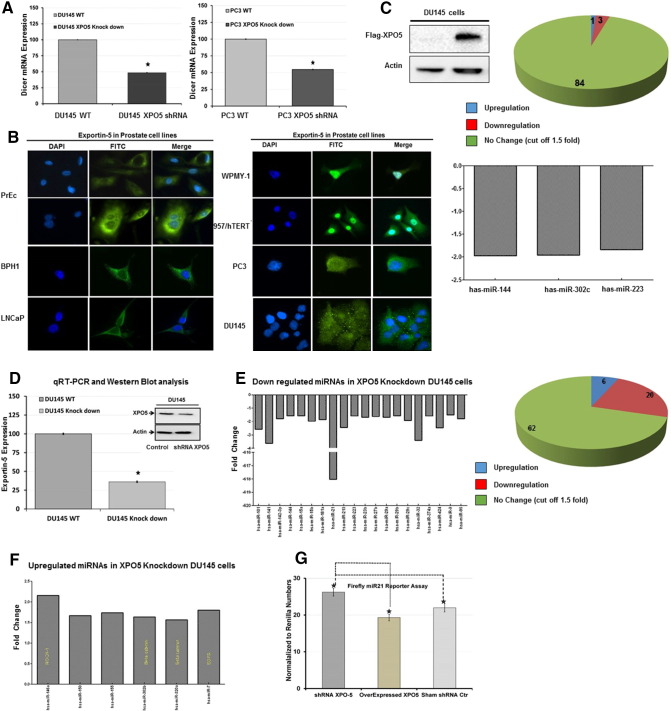
In order to evaluate the significance of XPO5 in the microRNA biogenesis pathway, we knocked down the XPO5 protein using shRNA approach in two different (DU145 and PC3) prostate cancer cell lines and examined the expression of the downstream DICER protein by qRT-PCR analysis. As shown in Figure 2A, cells that were transfected with XPO5 shRNA displayed significantly lower expression of DICER compared to the mock-transfected prostate cancer cells. During the preparation of this manuscript, several publications [32], [33] pointed toward genetic alterations in XPO5 in solid tumors impacting cellular localization, trapping the protein in the nucleus, and resulting in lower miRNA cargo activity. To explore the nucleocytoplasmic function of XPO5 in prostate cancer, we performed the immunofluorescent assay for XPO5 in normal (PrECs), benign (BPH-1), and prostate cancer (LNCaP, WPMY-1, 957/hTERT, PC3, and DU145) cells (Figure 2B). All cell lines including the two transformed cells WPMY-1 (SV40 large-T antigen-immortalized cells) and 957/hTERT (retroviral carrying LXSN-hTERT transformed cells) demonstrated both cytoplasmic and nuclear staining of XPO5. Interestingly, we found that both the transformed WPMY-1 and 957/hTERT cells exhibited significant retention of nuclear XPO5. To further confirm the functional status of XPO5 in prostate cancer cells, we overexpressed wt-Flag tagged-XPO5 in DU145 cells (Figure 2C) and subsequently measured and quantified the expression pattern of 88 mature miRNAs using a qRT-PCR array. As shown in Figure 2C, cells that were ectopically overexpressed with wild-type XPO5 failed to alter the levels of mature miRNAs, confirming that the endogenous XPO5 was functional in these cells. There was only 1 out of 88 mature miRNAs (miR-7c) that was slightly elevated (1.5-fold), while three other miRNAs (miR-144, miR-302c, and miR-223) were downregulated in the DU145 cells that were transiently overexpressed with wild-type XPO5 plasmid. We also examined mature miRNAs in DU145 cells by knocking down the XPO5 expression with shRNA (Figure 2D). Compared to XPO5-overexpressing DU145 cells, XPO5 knockdown in DU145 cells resulted in a significant change in the mature miRNAs’ expression profile. Twenty mature miRNAs were suppressed at least 1.5-fold or more when compared to the mock-transfected DU145 cells (Figure 2E). One of the oncogenic microRNA the miR-21 was found to be 618-fold downregulated in XPO5 knockdown cells. Similarly, around six miRNAs were found to be overexpressed in these XPO5 knockdown DU145 cells (Figure 2F), which may suggest the differences in the binding affinities of some miRNAs for the XPO5 protein. We further confirmed these observations of lower miR-21 levels in XPO5 knockdown cells by using the miR-21 Luciferase reporter assay. DU145 cells grown at a density of 1 × 105 cells in 96-well plate were co-transfected with miR-21 luciferase reporter plasmid (pMR-Luc) in the presence of either shRNA against XPO5 or by overexpressing XPO5 (pUC-Flag-XPO5) or mock shRNA plasmids together with Renilla luciferase plasmid (pRL-CMV) for normalization. As shown in Figure 2G, cells that were knocked down with an XPO5 shRNA had a significantly higher luciferase activity compared to the cells that were overexpressed with XPO5 plasmid. Using RNA interference in cell lines to knock down genes has been previously known to have some off-target effects [34]. To evaluate if there were any off-target effects associated with our shRNA constructs, we utilized the qRT-PCR and measured 84 different mRNAs in the XPO5 shRNA knockdown and control DU145 cells. As shown in the Supplemental Table S2, no significant changes [35] were found in the mRNA levels between the shRNA XPO5 knockdown and control DU145 cells, confirming the specificity of RNA interference for XPO5 gene.
Figure 2.
Quantitative RT- PCR analysis for DICER mRNA expression in XPO5 knockdown DU145 and PC3 cells (A). XPO5 protein localization by immunofluorescence microscopy in several different prostate cell lines (B). Evaluating the impact of XPO5 overexpression by qRT-PCR on the mature miRNA profiles in DU145 cells (C). Relative qRT-PCR analysis of XPO5 in wild-type and XPO5 knockdown DU145 cells (D). Impact of XPO5 knockdown using miRNA qRT-PCR miroarray analysis on mature miRNAs profile in DU145 cells showing lower expression for 20 mature miRNAs (E) and overexpression of 6 mature miRNAs in the XPO5 knock down DU145 cells (F). miR-21 firefly reporter assay in XPO5 knockdown and overexpressed DU145 cells compared to mock-transfected cells (G). Error bars represent the mean ± SEM where * indicates P ≤ .05.
Knocking Down XPO5 Suppresses the Proliferation of Prostate Cancer Cells
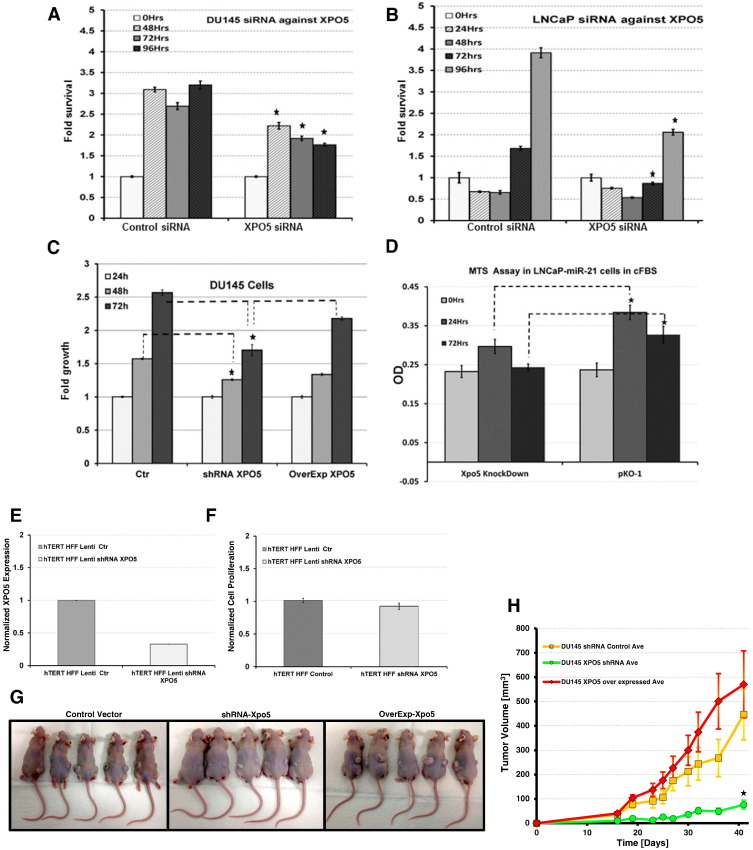
We next sought to determine whether targeting XPO5 might have any effects on the proliferation of prostate cancer cells. We utilized the RNA interference approach to knock down XPO5 and the downstream Dicer proteins (data not shown) in two different prostate cancer (DU145 and LNCaP) cell line models. Cells plated at the density of 1 × 105 cells into 96-well plates were transfected with 2 pmol of synthetic siRNA against XPO5 or mock siRNA in serum-free media according to the manufacturer recommendations. After 6 hours, transfected cells were replaced with normal serum-containing media. MTS assay was performed to evaluate cell proliferation at different time points as indicated in Figure 3, A and B. Cell transfected with XPO5 siRNA showed a significantly lower proliferation (P ≤ .05) compared to the mock siRNA transfected DU145 and LNCaP cells (Figure 3, A and B). Similar results of decreased cellular proliferation were also observed by MTS assay in DU145 cells that were stably selected to express an shRNA against XPO5 compared to the mock or overexpressed XPO5 cells (Figure 3C).
Figure 3.
MTS assay at indicated time points on DU145 cells (A) and LNCaP cells (B) that were either mock trasfected or transfected with siRNA against XPO5 or DICER (A and B). MTS assay using stably selected DU145 cells with XPO5 shRNA or overexpressed flag tagged XPO5 compared to vector control DU145 cells (C). MTS assay using the LNCaP-miR21 castration-resistant prostate cancer cell model that was mock transfected using pKO-1 vector or XPO5 shRNA containing plasmid (*P < .05) (D). Relative qRT-PCR analysis for the XPO5 mRNA expression in human HFF hTERT cells that were knocked down or wild-type for XPO5 using lentiviral vectors (E). Cell proliferation assay showing no significant difference between the wild-type type and XPO5 knockdown HFF hTERT cells (F). Tumor xenografts in athymic nude mice for DU145 cells that were stably selected to express an shRNA against XPO5 or plasmid overexpressing XPO5 or control vectors (G). Comparision of tumor growth kinetics between DU145 cells that were either knocked down or overexpressed for XPO5 expression; all error bars represent mean ± SE, and * indicates P ≤ .05 (H).
To further establish the function of XPO5 in the biology of prostate cancer, we utilized the previously characterized LNCaP-miR21 castration-resistant prostate cancer cell line model using MTS assay. LNCaP-miR21 cells grown in charcoal-stripped media were either with transfected shRNA against XPO5 or control plasmid followed by MTS assay at given time point (0, 24, and 72 hours). As shown in Figure 3D, cells that were transfected with plasmid alone (pKO-1) continue to grow in charcoal-stripped media, while on the other hand, cells that were transfected with shRNA against XPO5 showed a significantly lower proliferation at any given time, suggesting a positive role of XPO5 in prostate cancer cell growth.
While XPO5 remained the main transporter of many miRNAs, there was a strong possibility that knocking down the expression of XPO5 in normal cells might have deleterious cytotoxic effects. To evaluate whether suppressing XPO5 expression might be toxic to normal cells, we performed knockdown of XPO5 in normal human foreskin fibroblast (HFF) cells using lentiviral vectors. Quantitative PCR analysis was performed to confirm the XPO5 knockdown in these stably selected HFF cells (Figure 3E). Following confirmation of reduced XPO5 mRNA expression, MTS assay was performed that showed no statistical difference in proliferation between the XPO5 knockdown or control HFF cells (Figure 3F), confirming the selective phenotypic effect of XPO5 knockdown on prostate cancer cells. Similarly, we also studied the disruption of XPO5 in zebrafish model via CRISPR/Cas9 technology to evaluate embryonic toxicity of XPO5. Although we did not find significant differences in the survival of F0 progenies between XPO5 CRISPR/Cas embryos to mock injected, there were predominant developmental deformities in the Spine of F0 XPO5 disrupted zebrafish model (Supplemental Figure 1B).
To further investigate the effect of XPO5 knockdown on the in vivo growth kinetics of prostate cancer cells, we generated stable knockdown (DU145 XPO5 shRNA), scrambled control shRNA (DU145 shRNA Control), and overexpressed wild-type XPO5 (DU145 OverExp XPO5) cells using lentiviral vectors. Briefly, DU145 cells (DU145 shRNA control, DU145 XPO5 shRNA, and DU145 XPO5 OverExp) grown in exponential phase were implanted subcutaneously into the dorsal rear flank region of five athymic nude mice per experimental condition. Tumor volumes were measured every 3 days after implantation. The experiment was terminated for all groups after 6 weeks when animals in the overexpressed XPO5 group started displaying signs of distress due to large tumor burden from s.c. implantation (Figure 3G). Tumor volumes were plotted against time as shown in Figure 3H. The graph showed that the tumor size of DU145 OverExp XPO5 group was always higher than that of mock or DU145 shRNA XPO5 group. The control group (DU145 shRNA control) exhibited similar growth pattern as that of the XPO5-overexpressed group (P > .05); however, on the other hand, the XPO5 knockdown group (DU145 XPO5 shRNA) had significantly lower tumor volumes at any given time. The smaller tumor volume throughout the length of the experiment corroborates the antiproliferative role of XPO5 shRNA in prostate cancer biology.
Targeting XPO5 with shRNA in Androgen-Sensitive Prostate Cancer Cells Suppresses the AR Activity
To understand whether dysregulated expression of XPO5 in prostate cancer cells might be affecting the AR-PSA nexus in advance prostate cancers, nuclear and cytoplasmic AR and PSA protein expression was assessed in LAPC4 cells that were stably knocked down for XPO5 or overexpressed using Flag-XPO5. As shown in Figure 4A, knocking down XPO5 in LAPC4 cells induced the expression of AR compared to the control cells. This expression, however, was not functional as the downstream PSA expression was significantly compromised in both cytoplasmic and nuclear fractionated lysate. In contrast, overexpression of the XPO5 gene in LAPC4 cells causes the induction of AR and PSA expression in both nuclear and cytoplasmic fraction that was further confirmed by immunofluorescence assay (Figure 4B).
Figure 4.
Nuclear and cytoplasmic fractionation for DU145 cells that were stably selected for XPO5 knockdown and XPO5-overexpressed LAPC4 cells showing higher levels of PSA in XPO5-overexpressing cells compared to the control and XPO5 knockdown cells (A). Immunofluorescence microscopy of PSA in control, XPO5 shRNA knockdown, and XPO5-overexpressed LAPC4 cells (B). Clinical ELISA assay measuring total secreted PSA into the media from control, XPO5-shRNA, and XPO5-overexpressing LAPC4 cells (C). Colony formation assay for LAPC4 control and LAPC4-shRNA XPO5 knockdown cells (D). Western blot analysis on cell lysates from control and knockdown XPO5 shRNA LAPC4 cells (E). Error bars represent mean ± SE, and * indicates P ≤ .05.
To further endorse that overexpression of XPO5 induces PSA production in prostate cancer cells, we utilized the clinical ELISA-based PSA assay to measure total secreted PSA in condition media. LAPC4 cells in our three experimental conditions (control, shRNA XPO5, and overexpressing XPO5) were analyzed for PSA production in media at indicated time points. As expected, LAPC4 cells that were stably selected to overexpress XPO5 have the highest PSA levels in media when compared to LAPC4 control cells, whereas shRNA-mediated knockdown of XPO5 in LAPC4 cells was found to have the least secreted PSA levels (Figure 4C). We also validated these data in the LNCaP model that was stably selected with XPO5 shRNA or control vector using the AR reporter firefly luciferase assay [36]. LNCaP-shRNA XPO5 or control cells were co-transfected with the AR reporter vector (pBk-PSE-PBN-Luc) together with pRL-CMV for transfection normalization in the presence and absence of R1881 (androgens) (Supplemental Figure 2). XPO5 knockdown LNCaP cells displayed lower luciferase numbers compared to the control LNCaP cells. Taken together, these data demonstrate that knocking down XPO5 in androgen-sensitive prostate cancer cells can suppress PSA expression.
Supplemental Figure 2.
Androgen receptor reporter assay was utilized to study the activity of AR in XPO5 knockdown and control LNCaP cells. LNCaP cells grown in charcoal-stripped FBS-containing media was co-transfected with firefly pRL-PSE/PBN-F.Luc and Renilla luciferase (pRL-luc) plasmids in the presence and absence of R1881. Error bars represent mean ± SE.
Lower proliferation has been previously shown to decrease the PSA production in prostate cancer cells. In order to investigate whether lower PSA in LAPC4 shRNA XPO5 cells was due to suppressed growth, we performed a clonogenic assay followed by crystal violet staining using the knockdown XPO5 and control LAPC4 cells. As shown in Figure 4D, cells that were knockdown for XPO5 have relatively smaller and fewer numbers of clones compared to the control LAPC4 cells.
Previously, targeting DICER expression via knockdown has been shown to elicit a DNA damage response in cancer cells [37]. Similarly, based on our results (Figure 2, A and B), we have shown that DICER levels were regulated by the XPO5 expression. To investigate the mechanisms that might be responsible for lower proliferation in XPO5 knockdown LAPC4 cells, we hypothesized that XPO5 knockdown could induce a DNA damage response, impacting cell growth rates. To confirm this hypothesis, we compared the expression profile of known DNA damage response proteins (PCNA and gamma H2AX) from LAPC4-WT cells and XPO5-shRNA knockdown cells. As shown in Figure 4E, LAPC4 cells that were knocked down for XPO5 protein had higher levels of gamma H2AX expression when compared to the control cells. The upregulated expression of gamma H2AX, which is an established marker for double-standard DNA assault [38], might be responsible for the slower proliferation rate of LAPC4 shRNA XPO5 cells as indicated by the downregulation of PCNA in XPO5 knockdown LAPC4 cells (Figure 4E).
Overexpression of XPO5 Overrides the Inhibitory Effect of miRNAs Regulation Control
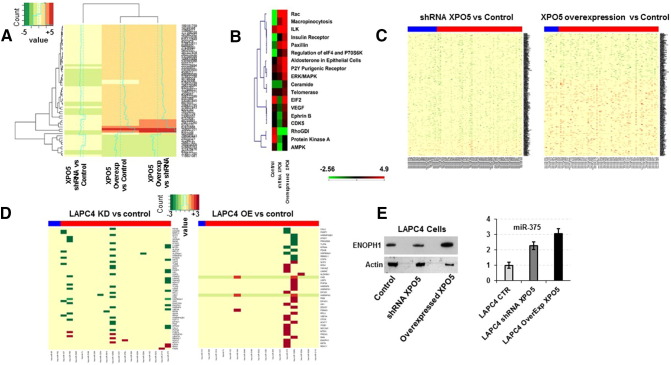
XPO5 plays a major role in the miRNA biogenesis, and overexpression of XPO5 has been shown to enhance the production of mature miRNAs [39]. Therefore, we sought to determine the status of miRNA profiles in LAPC4 cells that were either knocked down or overexpressed for XPO5. We analyzed (n = 3) biological replicates for each cell line (LAPC4 control, LAPC4-shRNA XPO5, and LAPC4-OverExp XPO5) using Affymetrix GeneChip miRNA microarrays. We employed the generalized linear model for differential gene expression that involves fitting of mixed effects in linear models to estimate miRNA expression differences between samples. To avoid separate sample preparation, we profiled pre-microRNA and mature micro-RNA expression on the same Affymetrix microarray chip (Affymetrix GeneChip miRNA 3.0 arrays) for each sample. This gives a glimpse into the relationship between profiled microRNA pre- and matured forms. The raw files were read utilizing the “robust multiarray average” as described in our Methods section. Paired t test was performed on the median expression values between the miRNAs (premature versus mature microRNAs) in each of the sample sets. Using a volcano plot, we evaluated fold-changes between the differentially expressed pre- and mature miRNAs in the XPO5 knockdown and overexpressed LAPC4 cells (Supplemental Figure 3). While there was no statistical difference in the pre-miRNA profiles, a profound affect was observed in the expression profiles of mature miRNAs in both knockdown and overexpressed XPO5 LAPC4 cells (Supplemental Figure 3). To further investigate whether these differentially regulated miRNAs in these cells have any functional consequences on the protein expression profiles, we performed quantitative global proteomics analysis using state-of-the-art mass spectrometry on the LAPC4 cells that were either knocked down or overexpressed for XPO5. Briefly, LAPC4 WT cells, LAPC4-shRNA XPO5, and LAPC4-OverExp XPO5 (n = 3) were harvested in 8 M urea and 1 M NH4HCO3 as shown in the schematic (Supplemental Figure 4). Cells from each flask were harvested individually at the confluency of 80% to 85% and were subjected to iTRAQ labeling as described in the Material and Methods section. After fractionation by basic reverse-phase liquid chromatography, each fraction was analyzed by liquid chromatography–tandem mass spectrometry (LC-MS/MS) to obtained quantitative data. At a 1% FDR, we were able to identify 1351 protein groups with a minimum of 2 peptides per protein (Supplemental Table S3). Heat maps were generated between the differentially expressed proteins that have shown at least two-fold change between the cell types. As shown in Figure 5A, there was a significant increase in the global protein expression profiles from the LAPC4 cells that were stably selected to overexpressed XPO5 gene; on the other hand, a large number of proteins in the LAPC4-shRNA XPO5 cells were downregulated when compared to the control cell lysate. The global iTRAQ proteomics data for the XPO5 overexpressing, knockdown, or control cells were further subjected to Ingenuity pathway analyses to evaluate which proteins were affected by XPO5 in these cells. As shown in Figure 5B, LAPC4 cells that were stably selected to overexpress XPO5 showed several cellular proliferation proteins including insulin receptor, ERK/MARK, VEGF, EIF2, and telomerase that were upregulated by overexpressing XPO5 in LAPC4 cells compared to the knockdown or control cells.
Supplemental Figure 3.

Volcano plot showing fold-changes between differentially expressed pre- and mature miRNAs in XPO5 knockdown or XPO5-overexpressing LAPC4 cells compared to control LAPC4 cells (A). Venn diagrams for differentially expressed pre- and mature miRNAs between XPO5-overexpressed LAPC4 versus LAPC4 control cells, LAPC4 knockdown versus mock control cells, and XPO5-overexpressed LAPC4 cells versus XPO5 knockdown LAPC4 prostate cancer cells (B).
Supplemental Figure 4.
A schematic representation of the workflow using LC-MS/MS for global proteomic analyses for LAPC4 prostate cancer cell lines that were stably selected with shRNA against XPO5, vector control, or with overexpressed flag tag wt XPO5 containing plasmid.
Figure 5.
Global proteomics using iTRAQ ratios of mass spectrometry data showing fold changes between LAPC4 cells that were stably selected with shRNA against XPO5 versus control or overexpressing LAPC4 cells with wild-type XPO5 (pcDNA-3.1 Flag-XPO5) plasmid versus control or LAPC4 cells overexpressed with XPO5 versus LAPC4 cells that were knocked down for XPO5 (A). Global iTRAQ proteomics data for the XPO5-overexpressing, knockdown, or control cells were subjected to Ingenuity pathway analyses showing overexpression of several cellular proliferation proteins including insulin receptor, ERK/MARK, VEGF, EIF2, and telomerase (B). Integrated analysis for mature miRNAs and global proteomics expression in LAPC4 cells using heat maps showing the dysregulation of miRNA regulation control with a global upregulation of protein in overexpressing XPO5 cells (C). Integrated analysis for miRNA-protein expression with P value ≤ .05 for the differentially expressed proteins and miRNAs in XPO5 knockdown or overexpressing LAPC4 cells (D). Western Blot for ENOPH1 protein and Taqman qRT-PCR analysis for mature miR375 showing higher expression ENOPH1 in XPO5-overexpressing LAPC4 cells even in the presence of higher levels of mature miRNA-375, while similar levels of miR-375 in XPO5 knockdown LAPC4 cells show lower expression of ENOPH1 protein, suggesting a dysregulation of miRNAs in XPO5 overexpression LAPC4 cells. Error bars represent mean ± SE (E).
To further explore whether miRNAs were differentially regulating the expression of proteins in the knockdown and overexpressing XPO5 cells, we performed an integrated data analysis on the mature miRNAs from the miRNA array that were identified and mapped to the targets using the multiMIR database and proteins. As shown in the heat map (Figure 5C), the differentially expressed miRNAs from low to higher expression levels (heat bar, blue: low and red: high) are plotted against their protein targets on the y-axis between knockdown (LAPC4-shRNA XPO5) versus control and LAPC4-OverExp-XPO5 versus LAPC4 control cells (Figure 5D, P < .05). The green box on the heat map represents the suppressed or downregulated proteins, while the red symbolizes the overexpression for a particular protein in the presence of mature miRNA. Interestingly, comparing the two heat maps for the integrated miRNA/protein analysis, we found that overexpression of XPO5 caused a significant increase in the proteins’ expression regardless of the presence of specific mature miRNAs (Figure 5, C and D). For instance, enolase-phosphatase 1 protein which is a well-known target for miR-375 was detected to express at higher levels in the LAPC4-OverExp-XPO5 cells compared to the XPO5 knockdown cells, suggesting that XPO5 overexpression may be responsible for the dysregulated function of mature miRNAs which was further confirmed by Western blot and by qRT-PCR analyses (Figure 5E).
Overexpression of XPO5 May Contribute to the Posttranslational Modifications (PTMs) of Proteins
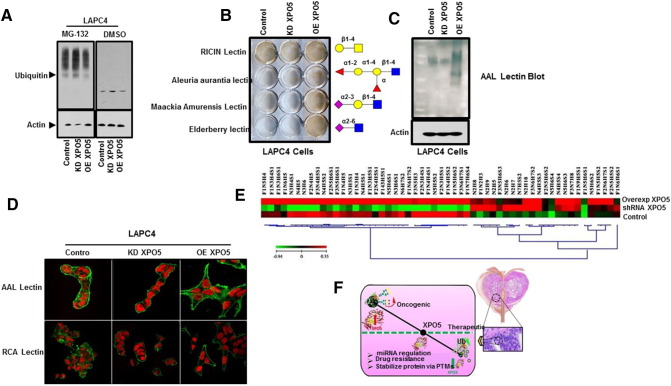
Discerning the global proteomics data (Figure 5C) and the integrated analysis of miRNAs and protein expression between the knockdown and overexpressing XPO5 LAPC4 cells (Figure 5, E and F), we noticed that majority of the proteins in the LAPC4-overxpressing XPO5 cells displayed higher protein abundance when compared to the XPO5 knockdown or control. One possibility for this phenomenon might be that the XPO5 protein may interact and stabilize many cellular mRNAs or mediate nuclear export of some unspliced mRNA that lacks hairpin structure. Alternatively, reduced protein degradation and increased overall protein stability could be responsible. To evaluate the later hypothesis, we investigated the impact of XPO5 expression on the proteasome degradation pathway. Both XPO5-shRNA LAPC4 cells and LAPC4 overexpressed cells were subjected to the 26S proteasome inhibitor MG132 and analyzed via Western blot using a polyclonal ubiquitin antibody. As shown in Figure 6A, treatment with MG132 resulted in a profound increase in bound ubiquitin relative to treatment with DMSO. When compared to our control LAPC4 cells, XPO5 knockdown resulted in a higher level of ubiquitin signaling. Furthermore, LAPC4 cells with increased expression of XPO5 displayed reduced levels of ubiquitin signaling, suggesting an overall lower rate of ubiquitin-mediated proteosomal degradation in the XPO5-overexpressing LAPC4 cells (LAPC4-OverExp XPO5).
Figure 6.
Western blot analysis on whole cell lysate with anti-ubiquitin antibody for LAPC4 control, LAPC4 shRNA XPO5, and LAPC4-overexpressing XPO5 cells in the presence of IMG132 or DMSO mock-treated control showed higher binding of ubiquitin in the XPO5 knockdown and control cells compared to the XPO5-overexpressing LAPC4 cells (A). Biotinylated lectin assay for LAPC4 cells (control, shRNA XPO5, and overexpressing XPO5) showing higher staining for Maackiaamurensis Lectin1 (MAL1), Aleuria aurantia lectin (AAL) and Elderberry lectin (EL) in XPO5-overexpressing LAPC4 cells (B). Lectin blot for AAL indicating higher fucosylation in XPO5-overexpressing LAPC4 cells, while knocking down XPO5 significantly suppresses AAL binding, suggesting lower fucosylation in XPO5 knockdown LAPC4 cells (C). Confocal microscopy for AAL and RCA lectins in LAPC4 control, XPO5 knockdown, and XPO5-overexpressing LAPC4 cells (D). Heat map from quantitative glycomics (GIG) method showing log fold-changes between different glycans in LAPC4 cells that were knocked down or overexpressed for XPO5 expression (E).
PTMs including glycosylation have been previously shown to play critical roles in the stability of many proteins [40], [41], [42]. In order to evaluate whether XPO5 expression in LAPC4 cells may be impacting glycosylation of proteins and thus influence the overall stability of proteins, we performed a biotinylated lectin binding assay using four different lectins (Ricin Lectin, Aleuria aurantia lectin, Macckia Amurensus Lectin I, and Elderberry Lectin) in LAPC4 cells that were either wild-type knockdown or overexpressed for XPO5 protein. As each lectin has a specificity of binding to a particular cell membrane glycan, the binding assay helped us to speculate the overall abundance of glycan in a specific cell type. Experiment was performed in the 6-well plate format on adherent cells that were plated at density of 2 × 105 cells before incubating them with different lectins followed by HRP-conjugated streptavidin-based DAB detection. As shown in Figure 6B, there is no change in the staining pattern of Ricinus communis Agglutinin I (RCA) lectin which is known to bind to galactose or N-acetylgalactosamine residues of membrane glycoconjugates between all cell types. However, on the other hand, a strong staining pattern for Aleuria aurantia lectin (AAL), Maackia amurensis Lectin1 (MAL-1), and Elderberry lectin (Eb) was only detected in the XPO5-overexpressing LAPC4 cells, suggesting the abundance of fucosylation (AAL lectin) and sialyation (MAL1 and Eb) in these cells. We further supported this results by performing both an AAL lectin blot (Figure 6C) and confocal microscopy (Figure 6D) and found higher abundance for fucosylated proteins in XPO5-overexpressing cells compared to the wild-type or XPO5 knockdown cells. To confirm these observations, we next sought to determine the global glycosylation pattern of these cell types ((LAPC4 control, LAPC4-shRNA XPO5, and LAPC4-OverExp XPO5) by using a more robust high-throughput quantitative glycomics analysis. Using the GIG methodology as described in our Materials and Methods section, we were able to analyze and quantify glycans at the global level for each cell type. Comparing the log fold changes in glycan profiles using heat map between the knock down (LAPC4-shRNA-XPO5) to the overexpressing XPO5 LAPC4 cells (LAPC4-OverExp XPO5), we found an overall increase in the fucosylated and sialyated glycan species of the XPO5-overexpressing LAPC4 cells (Figure 6E), which was consistent with the biotinylated lectin assay (Figure 6B). We further confirm these findings in another prostate cancer PC3 cell line model and found higher staining of AAL lectin in PC3 cell lysate that also expressed higher abundance of XPO5 proteins compared to the normal PrEC prostate cell line (Supplemental Figure 5).
Supplemental Figure 5.
XPO5 overexpression and higher fucosylation in PC3 cells.
Discussion
The nuclear export of pre-miRNAs into the cytoplasm of cells is a crucial step in the maturation biology of miRNAs and its regulation control. Aberrant expression of proteins involved in these processes results in pleiotropic effects on many genes with functional consequences including the risk of carcinogenesis [32]. Although Exportin 5 (XPO5) remains the main passenger protein for all of the miRNA hairpin structures and a rate-limiting step in miRNA biogenesis [9], very few reports have studied and characterized its function in the context of miRNA-protein regulation. Using prostate cancer specimens and cell line models, we have identified overexpression of XPO5 in advanced prostate cancer and herein report that overexpression of XPO5 in prostate cancer can cause aberrant canonical miRNA regulation, resulting in the global upregulation of proteins. While XPO5 has been previously characterized as haploinsufficent tumor suppressor [43], in this report, we revealed that overexpression of the wild-type XPO5 is tumorigenic in prostate cancer cells. Similarly, suppressing the expression of XPO5 was found to have a therapeutic effect in these models.
Several studies have identified genetic alterations including polymorphisms for XPO5 that rendered the potential of XPO5 in the transport mechanism of pre-miRNAs [10], [33], [44]. We however found that this was not the case in prostate cancer cells after confirming it with several different approaches including fluorescence microscopy, overexpressing of the wild-type XPO5 gene in prostate cancer cells and subsequently quantifying the mature miRNA by qRT-PCR analysis (Figure 2E), and genomic DNA sequencing (data not shown). While global downregulation of mature miRNAs has been generalized in cancers, miRNAs in prostate cancer have been shown to be upregulated with the highest levels of expression in high-grade cancers [45]. The importance of XPO5 protein in the miRNA biogenesis pathway can be realized from the fact that XPO5 expression regulates levels of the DICER proteins (Figure 2A) and has been shown to interact directly with the DICER mRNA [11]. XPO5 was also reported to protect pre-miRNAs from nucleases degradation which was observed after XPO5 knockdown failed to result in the accumulation of pre-miRNA inside the nucleus [9], [46]. We were also interested in investigating the mechanism by which XPO5 knockdown may cause growth arrest. Beside checkpoints and mitotic catastrophic arrest, one of the major pathways associated with cell cycle arrest is the DNA damage response. It has been previously shown that decreased expression of Dicer elicits DNA damage response and induces senescence in Drosophila or primary cells by activating the DNA damage checkpoint or p19(Arf)-p53 signaling [47], [48]. Our data support the notion that knocking down XPO5 results in suppression of DICER expression (Figure 2A), which may lead to the hyperactivation of gamma H2AX which in turn reduced proliferation of LAPC4 cells (Figure 4E). While XPO5 suppression was mostly therapeutic in prostate cancer cell models, we also explored whether XPO5 overexpression might have any functional consequences in prostate cancer biology and more specifically on the AR-PSA nexus. We found that overexpressing of XPO5 enhanced the PSA expression when compared to the wild-type controls, whereas knocking down XPO5 significantly suppressed the PSA expression in prostate cancer cell models (Figure 4A). These data were further supported by our global proteomic analyses data where PSA expression was positively altered in XPO5-overexpressed prostate cancer cells. PSA is a highly glycosyalated protein [49], [50] and has been shown to be highly fucosylated [51] and sialyated [52] in prostate cancer. We believe that besides the dysregulation mechanism of miRNAs, glycosylation of PSA might be playing role in the stability of PSA in LAPC4 overexpressing XPO5 cells.
During the preparation of our manuscript, Kim et al. published their seminal work by showing a very modest effect of XPO5 in the biology of miRNAs and revealed that the function of XPO5 is necessary but not critical in miRNA maturation [53]. Our results were largely in accordance with them as our miRNA microarray analysis and qRT PCR data did not demonstrate a significant depletion of mature miRNAs in knockdown XPO5 cells. To better understand the impact of XPO5 expression on global protein profiles, we performed an integrated analysis of miRNA expression and quantitative global proteomics analysis in an attempt to relate miRNA profiles to their target protein expression profiles. Of interest, we observed a global upregulation of many proteins (P < .05) in XPO5-overexpressing cells even in the presence of higher levels of miRNAs targeted to their respective mRNA. In contrast, in XPO5 knockdown prostate cancer cells, miRNAs still retained their inhibitory properties as indicated by suppressed protein expression profiles (Figure 5, E and F). These data indicate that overexpression of XPO5 can significantly alter the canonical miRNA-mRNA-protein regulation, with the dysregulated mechanism of the miRNA processing machinery playing a role in the progression to advanced disease.
To further elucidate the mechanisms and explain our observation of increased protein abundance in overexpressing XPO5 cells, we evaluated the 26S proteosomal degradation pathway by blocking the proteolytic activity of the 26S proteasome with MG-132 or vehicle control and compared the ubiquitination patterns in XPO5-overexpressed or knockdown LAPC4 cells. We detected higher levels of ubiquitin-bounded proteins in the knockdown and control cells compared to the overexpressing XPO5 cells, suggesting an overall lower degradation of proteins which might be responsible for the fate of protein in the XPO5-overexpressing LAPC4 cells (Figure 6A). Besides reduced proteasome degradation, other cellular mechanisms that may be involved in higher protein levels observed in XPO5-overexpressed cells, including increase in the stability of proteins mediated by increased PTMs specifically glycosylation. We hypothesized whether hyperglycosylation might be one reason for lower ubiquitination levels observed in XPO5-overexpressing LAPC4 cells. To investigate these phenomena, we first utilized several lectins to analyze the glycosylation patterns of cells with varying protein levels of XPO5, revealing higher N-glycosylation patterns (fucosylation and sialyation) for XPO5 overexpression cells. This result was validated by a more robust approach where we employed quantitative N-linked glycan analysis, confirming a global increase in fucosylated and sialyated proteins in overexpressing XPO5 cells.
Conclusion
These data suggest a novel role of XPO5 in prostate cancer biology, where overexpression of XPO5 causes dysregulation in the miRNA processing mechanism, resulting in a shift in protein stability via lower ubiquitination and hyper N-linked glycosylation, as well as conferring a growth advantage by suppressing the DNA damage response. These results have significance in the biology of prostate cancer and the future development of therapeutics to treat advanced disease.
The following are the supplementary data related to this article.
Supplemental Figure 6.

Relative mRNA in wiltype and XPO5 knockdown DU145 cells.
Supplementary Material iTRAQ LC-MS/MS proteoimcs Data for LAPC4 cells.
qRT-PCR Primer pairs.
Fold mRNA in XPO5 shRNA and Control DU145 cells.
Global proteomics LC-MS/MS iTRAQ data in LAPC4 cells.
Acknowledgments
Acknowledgement
This study was supported in part by grants from Flight Attendant Medical Research Institute (FAMRI) to Naseruddin Höti, and the National Cancer Institute, the Early Detection Research Network grant (EDRN, U01CA152813) to Dr. Hui Zhang. We are very much thankful to Dr. Jeff S Mum and his lab for sharing their resources in the completion of this manuscript.
Footnotes
Conflict of Interest: None.
Financial Support: This study was supported in part by grants from Flight Attendant Medical Research Institute (FAMRI) to N. Höti and the National Cancer Institute, the Early Detection Research Network grant (EDRN, U01CA152813) to H. Zhang & RO1 (R01CA121153-01A2) award to R. Rodriguez.
References
- 1.Siegel RL, Miller KD, Jemal A. Cancer statistics, 2017. CA Cancer J Clin. 2017;67:7–30. doi: 10.3322/caac.21387. [DOI] [PubMed] [Google Scholar]
- 2.Merseburger AS, Alcaraz A, von Klot CA. Androgen deprivation therapy as backbone therapy in the management of prostate cancer. Onco Targets Ther. 2016;9:7263–7274. doi: 10.2147/OTT.S117176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Ribas J, Ni X, Haffner M, Wentzel EA, Salmasi AH, Chowdhury WH, Kudrolli TA, Yegnasubramanian S, Luo J, Rodriguez R. miR-21: an androgen receptor-regulated microRNA that promotes hormone-dependent and hormone-independent prostate cancer growth. Cancer Res. 2009;69:7165–7169. doi: 10.1158/0008-5472.CAN-09-1448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Baffa R, Fassan M, Volinia S, O'Hara B, Liu CG, Palazzo JP, Gardiman M, Rugge M, Gomella LG, Croce CM. MicroRNA expression profiling of human metastatic cancers identifies cancer gene targets. J Pathol. 2009;219:214–221. doi: 10.1002/path.2586. [DOI] [PubMed] [Google Scholar]
- 5.Macfarlane LA, Murphy PR. MicroRNA: biogenesis, Function and Role in Cancer. Curr Genomics. 2010;11:537–561. doi: 10.2174/138920210793175895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Ambros V. microRNAs: tiny regulators with great potential. Cell. 2001;107:823–826. doi: 10.1016/s0092-8674(01)00616-x. [DOI] [PubMed] [Google Scholar]
- 7.Ambros V, Chen X. The regulation of genes and genomes by small RNAs. Development. 2007;134:1635–1641. doi: 10.1242/dev.002006. [DOI] [PubMed] [Google Scholar]
- 8.Petersen CP, Bordeleau ME, Pelletier J, Sharp PA. Short RNAs repress translation after initiation in mammalian cells. Mol Cell. 2006;21:533–542. doi: 10.1016/j.molcel.2006.01.031. [DOI] [PubMed] [Google Scholar]
- 9.Yi R, Qin Y, Macara IG, Cullen BR. Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs. Genes Dev. 2003;17:3011–3016. doi: 10.1101/gad.1158803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Melo SA, Moutinho C, Ropero S, Calin GA, Rossi S, Spizzo R, Fernandez AF, Davalos V, Villanueva A, Montoya G. A genetic defect in exportin-5 traps precursor microRNAs in the nucleus of cancer cells. Cancer Cell. 2010;18:303–315. doi: 10.1016/j.ccr.2010.09.007. [DOI] [PubMed] [Google Scholar]
- 11.Bennasser Y, Chable-Bessia C, Triboulet R, Gibbings D, Gwizdek C, Dargemont C, Kremer EJ, Voinnet O, Benkirane M. Competition for XPO5 binding between Dicer mRNA, pre-miRNA and viral RNA regulates human Dicer levels. Nat Struct Mol Biol. 2011;18:323–327. doi: 10.1038/nsmb.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Yang S, Hoti N, Yang W, Liu Y, Chen L, Li S, Zhang H. Simultaneous analyses of N-linked and O-linked glycans of ovarian cancer cells using solid-phase chemoenzymatic method. Clin Proteomics. 2017;14:3. doi: 10.1186/s12014-017-9137-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Yang S, Wang M, Chen L, Yin B, Song G, Turko IV, Phinney KW, Betenbaugh MJ, Zhang H, Li S. QUANTITY: an isobaric tag for quantitative glycomics. Sci Rep. 2015;5:17585. doi: 10.1038/srep17585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Irizarry RA, Bolstad BM, Collin F, Cope LM, Hobbs B, Speed TP. Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res. 2003;31:e15. doi: 10.1093/nar/gng015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U, Speed TP. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics. 2003;4:249–264. doi: 10.1093/biostatistics/4.2.249. [DOI] [PubMed] [Google Scholar]
- 16.Bolstad BM, Irizarry RA, Astrand M, Speed TP. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics. 2003;19:185–193. doi: 10.1093/bioinformatics/19.2.185. [DOI] [PubMed] [Google Scholar]
- 17.Ru Y, Kechris KJ, Tabakoff B, Hoffman P, Radcliffe RA, Bowler R, Mahaffey S, Rossi S, Calin GA, Bemis L. The multiMiR R package and database: integration of microRNA-target interactions along with their disease and drug associations. Nucleic Acids Res. 2014;42:e133. doi: 10.1093/nar/gku631. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Xiao F, Zuo Z, Cai G, Kang S, Gao X, Li T. miRecords: an integrated resource for microRNA-target interactions. Nucleic Acids Res. 2009;37:D105–110. doi: 10.1093/nar/gkn851. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Hsu SD, Lin FM, Wu WY, Liang C, Huang WC, Chan WL, Tsai WT, Chen GZ, Lee CJ, Chiu CM. miRTarBase: a database curates experimentally validated microRNA-target interactions. Nucleic Acids Res. 2011;39:D163–169. doi: 10.1093/nar/gkq1107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Vergoulis T, Vlachos IS, Alexiou P, Georgakilas G, Maragkakis M, Reczko M, Gerangelos S, Koziris N, Dalamagas T, Hatzigeorgiou AG. TarBase 6.0: capturing the exponential growth of miRNA targets with experimental support. Nucleic Acids Res. 2012;40:D222–229. doi: 10.1093/nar/gkr1161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Maragkakis M, Vergoulis T, Alexiou P, Reczko M, Plomaritou K, Gousis M, Kourtis K, Koziris N, Dalamagas T, Hatzigeorgiou AG. DIANA-microT Web server upgrade supports Fly and Worm miRNA target prediction and bibliographic miRNA to disease association. Nucleic Acids Res. 2011;39:W145–148. doi: 10.1093/nar/gkr294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Gaidatzis D, van Nimwegen E, Hausser J, Zavolan M. Inference of miRNA targets using evolutionary conservation and pathway analysis. BMC Bioinformatics. 2007;8:69. doi: 10.1186/1471-2105-8-69. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Griffiths-Jones S, Saini HK, van Dongen S, Enright AJ. miRBase: tools for microRNA genomics. Nucleic Acids Res. 2008;36:D154–158. doi: 10.1093/nar/gkm952. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Betel D, Wilson M, Gabow A, Marks DS, Sander C. The microRNA.org resource: targets and expression. Nucleic Acids Res. 2008;36:D149–153. doi: 10.1093/nar/gkm995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Wang X. miRDB: a microRNA target prediction and functional annotation database with a Wiki interface. RNA. 2008;14:1012–1017. doi: 10.1261/rna.965408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Anders G, Mackowiak SD, Jens M, Maaskola J, Kuntzagk A, Rajewsky N, Landthaler M, Dieterich C. doRiNA: a database of RNA interactions in post-transcriptional regulation. Nucleic Acids Res. 2012;40:D180–186. doi: 10.1093/nar/gkr1007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Kertesz M, Iovino N, Unnerstall U, Gaul U, Segal E. The role of site accessibility in microRNA target recognition. Nat Genet. 2007;39:1278–1284. doi: 10.1038/ng2135. [DOI] [PubMed] [Google Scholar]
- 28.Grimson A, Farh KK, Johnston WK, Garrett-Engele P, Lim LP, Bartel DP. MicroRNA targeting specificity in mammals: determinants beyond seed pairing. Mol Cell. 2007;27:91–105. doi: 10.1016/j.molcel.2007.06.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Jiang Q, Wang Y, Hao Y, Juan L, Teng M, Zhang X, Li M, Wang G, Liu Y. miR2Disease: a manually curated database for microRNA deregulation in human disease. Nucleic Acids Res. 2009;37:D98–104. doi: 10.1093/nar/gkn714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Rukov JL, Wilentzik R, Jaffe I, Vinther J, Shomron N. Pharmaco-miR: linking microRNAs and drug effects. Brief Bioinform. 2014;15:648–659. doi: 10.1093/bib/bbs082. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Ruepp A, Kowarsch A, Schmidl D, Buggenthin F, Brauner B, Dunger I, Fobo G, Frishman G, Montrone C, Theis FJ. PhenomiR: a knowledgebase for microRNA expression in diseases and biological processes. Genome Biol. 2010;11:R6. doi: 10.1186/gb-2010-11-1-r6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Melo SA, Esteller M. Disruption of microRNA nuclear transport in human cancer. Semin Cancer Biol. 2014;27:46–51. doi: 10.1016/j.semcancer.2014.02.012. [DOI] [PubMed] [Google Scholar]
- 33.Geng JQ, Wang XC, Li LF, Zhao J, Wu S, Yu GP, Zhu KJ. MicroRNA-related single-nucleotide polymorphism of XPO5 is strongly correlated with the prognosis and chemotherapy response in advanced non–small-cell lung cancer patients. Tumour Biol. 2016;37:2257–2265. doi: 10.1007/s13277-015-3980-3. [DOI] [PubMed] [Google Scholar]
- 34.Jackson AL, Linsley PS. Recognizing and avoiding siRNA off-target effects for target identification and therapeutic application. Nat Rev Drug Discov. 2010;9:57–67. doi: 10.1038/nrd3010. [DOI] [PubMed] [Google Scholar]
- 35.Caffrey DR, Zhao J, Song Z, Schaffer ME, Haney SA, Subramanian RR, Seymour AB, Hughes JD. siRNA off-target effects can be reduced at concentrations that match their individual potency. PLoS One. 2011;6:e21503. doi: 10.1371/journal.pone.0021503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Hoti N, Li Y, Chen CL, Chowdhury WH, Johns DC, Xia Q, Kabul A, Hsieh JT, Berg M, Ketner G. Androgen receptor attenuation of Ad5 replication: implications for the development of conditionally replication competent adenoviruses. Mol Ther. 2007;15:1495–1503. doi: 10.1038/sj.mt.6300223. [DOI] [PubMed] [Google Scholar]
- 37.Tang KF, Ren H, Cao J, Zeng GL, Xie J, Chen M, Wang L, He CX. Decreased Dicer expression elicits DNA damage and up-regulation of MICA and MICB. J Cell Biol. 2008;182:233–239. doi: 10.1083/jcb.200801169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Turinetto V, Giachino C. Multiple facets of histone variant H2AX: a DNA double-strand-break marker with several biological functions. Nucleic Acids Res. 2015;43:2489–2498. doi: 10.1093/nar/gkv061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Yi R, Doehle BP, Qin Y, Macara IG, Cullen BR. Overexpression of exportin 5 enhances RNA interference mediated by short hairpin RNAs and microRNAs. RNA. 2005;11:220–226. doi: 10.1261/rna.7233305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Watanabe I, Zhu J, Recio-Pinto E, Thornhill WB. Glycosylation affects the protein stability and cell surface expression of Kv1.4 but Not Kv1.1 potassium channels. A pore region determinant dictates the effect of glycosylation on trafficking. J Biol Chem. 2004;279:8879–8885. doi: 10.1074/jbc.M309802200. [DOI] [PubMed] [Google Scholar]
- 41.Huskens D, Princen K, Schreiber M, Schols D. The role of N-glycosylation sites on the CXCR4 receptor for CXCL-12 binding and signaling and X4 HIV-1 viral infectivity. Virology. 2007;363:280–287. doi: 10.1016/j.virol.2007.01.031. [DOI] [PubMed] [Google Scholar]
- 42.Walsh MT, Foley JF, Kinsella BT. Characterization of the role of N-linked glycosylation on the cell signaling and expression of the human thromboxane A2 receptor alpha and beta isoforms. J Pharmacol Exp Ther. 1998;286:1026–1036. [PubMed] [Google Scholar]
- 43.Iwasaki YW, Kiga K, Kayo H, Fukuda-Yuzawa Y, Weise J, Inada T, Tomita M, Ishihama Y, Fukao T. Global microRNA elevation by inducible Exportin 5 regulates cell cycle entry. RNA. 2013;19:490–497. doi: 10.1261/rna.036608.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Horikawa Y, Wood CG, Yang H, Zhao H, Ye Y, Gu J, Lin J, Habuchi T, Wu X. Single nucleotide polymorphisms of microRNA machinery genes modify the risk of renal cell carcinoma. Clin Cancer Res. 2008;14:7956–7962. doi: 10.1158/1078-0432.CCR-08-1199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Chiosea S, Jelezcova E, Chandran U, Acquafondata M, McHale T, Sobol RW, Dhir R. Up-regulation of dicer, a component of the MicroRNA machinery, in prostate adenocarcinoma. Am J Pathol. 2006;169:1812–1820. doi: 10.2353/ajpath.2006.060480. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Zeng Y, Cullen BR. Structural requirements for pre-microRNA binding and nuclear export by Exportin 5. Nucleic Acids Res. 2004;32:4776–4785. doi: 10.1093/nar/gkh824. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Tang KF, Ren H. The role of dicer in DNA damage repair. Int J Mol Sci. 2012;13:16769–16778. doi: 10.3390/ijms131216769. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Mudhasani R, Zhu Z, Hutvagner G, Eischen CM, Lyle S, Hall LL, Lawrence JB, Imbalzano AN, Jones SN. Loss of miRNA biogenesis induces p19Arf-p53 signaling and senescence in primary cells. J Cell Biol. 2008;181:1055–1063. doi: 10.1083/jcb.200802105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Vermassen T, Speeckaert MM, Lumen N, Rottey S, Delanghe JR. Glycosylation of prostate specific antigen and its potential diagnostic applications. Clin Chim Acta. 2012;413:1500–1505. doi: 10.1016/j.cca.2012.06.007. [DOI] [PubMed] [Google Scholar]
- 50.Belanger A, van Halbeek H, Graves HC, Grandbois K, Stamey TA, Huang L, Poppe I, Labrie F. Molecular mass and carbohydrate structure of prostate specific antigen: studies for establishment of an international PSA standard. Prostate. 1995;27:187–197. doi: 10.1002/pros.2990270403. [DOI] [PubMed] [Google Scholar]
- 51.Li QK, Chen L, Ao MH, Chiu JH, Zhang Z, Zhang H, Chan DW. Serum fucosylated prostate-specific antigen (PSA) improves the differentiation of aggressive from non-aggressive prostate cancers. Theranostics. 2015;5:267–276. doi: 10.7150/thno.10349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Li Y, Tian Y, Rezai T, Prakash A, Lopez MF, Chan DW, Zhang H. Simultaneous analysis of glycosylated and sialylated prostate-specific antigen revealing differential distribution of glycosylated prostate-specific antigen isoforms in prostate cancer tissues. Anal Chem. 2011;83:240–245. doi: 10.1021/ac102319g. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Kim YK, Kim B, Kim VN. Re-evaluation of the roles of DROSHA, Export in 5, and DICER in microRNA biogenesis. Proc Natl Acad Sci U S A. 2016;113:E1881–1889. doi: 10.1073/pnas.1602532113. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary Material iTRAQ LC-MS/MS proteoimcs Data for LAPC4 cells.
qRT-PCR Primer pairs.
Fold mRNA in XPO5 shRNA and Control DU145 cells.
Global proteomics LC-MS/MS iTRAQ data in LAPC4 cells.