Abstract
Compelling evidence suggests that epigenetic mechanisms such as DNA methylation play a role in stress regulation and in the etiologic basis of stress related disorders such as Posttraumatic Stress Disorder (PTSD). Here we describe the purpose and methods of an international consortium that was developed to study the role of epigenetics in PTSD. Inspired by the approach used in the Psychiatric Genomics Consortium, we brought together investigators representing seven cohorts with a collective sample size of N=1147 that included detailed information on trauma exposure, PTSD symptoms, and genome-wide DNA methylation data. The objective of this consortium is to increase the analytical sample size by pooling data and combining expertise so that DNA methylation patterns associated with PTSD can be identified. Several quality control and analytical pipelines were evaluated for their control of genomic inflation and technical artifacts with a joint analysis procedure established to derive comparable data over the cohorts for meta-analysis. We propose methods to deal with ancestry population stratification and type I error inflation and discuss the advantages and disadvantages of applying robust error estimates. To evaluate our pipeline, we report results from an epigenome-wide association study (EWAS) of age, which is a well-characterized phenotype with known epigenetic associations. Overall, while EWAS are highly complex and subject to similar challenges as genome-wide association studies (GWAS), we demonstrate that an epigenetic meta-analysis with a relatively modest sample size can be well-powered to identify epigenetic associations. Our pipeline can be used as a framework for consortium efforts for EWAS.
Keywords: EWAS, meta-analysis, trauma, stress
Introduction
Traumatic events are reported by over 70% of individuals during their lifetime [Benjet and others 2016]. They have been associated with a number of deleterious outcomes, including posttraumatic stress disorder (PTSD), a psychiatric disorder characterized by cognitive intrusions, avoidance, negative alterations in thoughts and mood, and alterations in physiological arousal and reactivity [Association 2000]. PTSD can be severe and disabling and is often associated with a range of comorbid psychiatric conditions such as depression and substance use disorders [Brady and others 2000; Najt and others 2011]. PTSD has also been associated with a 2.8-fold increase in suicidal thoughts and behaviors [Sareen and others 2005] as well as with a number of chronic medical conditions [Boscarino 2008; Coughlin 2011; David and others 2014; Heppner and others 2009; Jakovljevic and others 2008; Kubzansky and others 2009; Kubzansky and others 2007; Luft and others 2012]. While the risk of developing PTSD depends in part on the nature of the trauma [Kessler 2000], only a minority of those exposed to trauma develop PTSD. As a result, despite the high prevalence of lifetime trauma, the overall lifetime prevalence of PTSD in the United States is 6.8%, [Breslau and others 1998; Kessler and others 2005; Resnick and others 1993], which suggests there are individual differences in resilience that, if better understood, might inform the development of new approaches to prevention and treatment.
Genetic epidemiological studies suggest that both genetic and environmental factors contribute to PTSD risk. Twin studies estimate the heritability of PTSD to be between 30–70%, [Sartor and others 2012; Sartor and others 2011; Stein and others 2002; True and others 1993; Xian and others 2000] with the remaining variance being attributed to environmental factors. Genetic research, based on both candidate gene and genome-wide association studies (GWAS), has provided support for the role of genetics in the development and severity of PTSD and has begun to identify variants that account for some of the genetic influence on PTSD. The genetic loci identified in the extant GWAS have been implicated in a variety of processes, including neuroprotection, actin polymerization, neuronal function, and immune function [Almli and others 2014; Guffanti and others 2013; Logue and others 2013; Xie and others 2013]. Although promising, no robust genetic variants associated with PTSD have been identified and much work remains to be done to understand the biological basis of PTSD risk [Logue and others 2015].
A growing body of work has explored the role of environmental influences on an individual’s response to trauma. Given the dependence of PTSD development on exposure to environmental (i.e., traumatic) events, clarifying the ways in which environmental influences might affect biological function are critical to understanding the etiology of PTSD. In this regard, epigenetic mechanisms, which can mediate environmental influences on gene function, are particularly relevant. Epigenetic modifications, such as DNA methylation at cytosine-guanine dinucleotides (CpG sites), induce changes in gene expression in part through structural alterations of DNA that are maintained through each round of cell division; they respond to changes in the environment, are potentially reversible, and can be targeted for disease therapies [Feinberg 2007]. DNA methylation regulates gene expression by influencing the recruitment and binding of regulatory proteins to DNA. Typically, higher methylation at gene promoter regions correlates with decreased expression of that gene, while intragenic methylation can regulate alternative promoters and enhancers [Bonasio and others 2010; Maunakea and others 2010].
Animal studies have demonstrated that epigenetic changes—particularly alterations in DNA methylation in response to nurturing—are related to altered responses to stress [Jirtle and Skinner 2007; Weaver and others 2004]. Similar alterations have been reported in the human literature, in both central and peripheral tissues [McGowan and others 2009; Tyrka and others 2012]. Considering the influence of traumatic stress on DNA methylation seen in some studies [Vinkers and others 2015], epigenetic-based investigations may extend genetic research findings. For example, research reporting an association of PTSD with a genetic variant in the PAC1 receptor (ADCYAP1R1; rs2267735) went beyond this finding to observe that PTSD severity was also correlated with methylation levels of the gene [Ressler and others 2011]. Other work has suggested an interactive effect between trauma burden and DNA methylation in the serotonin transporter locus (SLC6A4) on PTSD risk, independent of the widely studied length polymorphism at this same locus [Koenen and others 2011] and a potentially interacting effect of genetic and epigenetic variation at the dopamine receptor (SLC6A3) on PTSD risk [Chang and others 2012]. Indeed, stress exposure itself has been shown to alter epigenetic patterns in both animal and human studies [Moser and others 2015; Roth and others 2009; Sipahi and others 2014]. In addition to candidate gene methylation studies, a small number of studies have examined genome-wide DNA methylation patterns in PTSD [Mehta and others 2013; Smith and others 2011; Uddin and others 2010]. In these first genome wide studies of DNA methylation, immune dysregulation figured prominently among the biological networks associated with PTSD and, at a CpG site level, DNA methylation levels in several CpGs showed suggestive evidence of replication between these studies [Mehta and others 2013; Smith and others 2011; Uddin and others 2010].
Although the emerging literature on epigenetic influences on PTSD is promising, the majority of research to date has been conducted with modest sample sizes, with inherent limited statistical power. Furthermore, studies of DNA methylation have been hampered by technical issues including batch effects [Harper and others 2013] and blood cell composition [Houseman and others 2012]. Experience from large-scale genetic studies, such as the Psychiatric Genomics Consortium (PGC), underscores how large collaborative endeavors can provide the adequate sample sizes and the statistical power necessary to produce significant and replicable results [Logue and others 2015]. Based on this experience, the PGC-PTSD formed the epigenetics working group to organize an expansive network of investigators and their collection of samples with genome-wide DNA methylation data available for joint analyses.
Although there are many advantages to this collaborative approach, there are also challenges. First, methods for assessment of trauma exposure and PTSD symptoms/diagnosis differ among cohorts, requiring harmonization of the phenotypic data. Participating cohorts have assessed PTSD using clinical interviews and self-report measures with possible case diagnoses of lifetime PTSD, current PTSD, or “probable” PTSD available in across different studies. The larger PGC-PTSD has made substantial progress harmonizing phenotypes across studies and a similar approach will be adopted by the PGC-PTSD epigenetics group in its analyses. A second challenge of consortia is to address complications in the data sharing that is a prerequisite to increasing the power and reproducibility of the study. The trend in psychiatric genetics has been to establish large-scale consortia for the purpose of expanding sample sizes beyond what is possible based on the data from any one group. Almost universally used in current large-scale genomic consortia is meta-analysis, through which the data from individual sites are analyzed separately and combined based on summary statistics. In many cases, meta-analysis of individual-level results yields results comparable to those of a “mega-analysis” of pooled data from different studies, [Lin and Zeng 2010; Mathew and Nordstrom 1999; Olkin and Sampson 1998]. However, there have been important distinctions in the way the meta-analysis strategy has been implemented across consortia, including variations in the degree to which the cleaning and analysis of the individual-level data occurs (i.e., centralized versus distributed). The analysis of individual data in a centralized manner allows a high degree of control over the quality control (QC) process and an ability to quickly perform follow-up analyses, but poses difficulties of requiring a larger degree of computational resources and storage at the consortium level and permission from all groups to share data.
The PGC-PTSD epigenetics group uses many of the same protocols and tools developed by the PGC [Logue and others 2015], with the difference that not all data are centrally stored and managed since some constituent samples that originate from US military, US Veteran (VA), or foreign countries are subject to additional regulatory oversight, which do not allow the sharing of individual-level genomic data. To enable participation for these studies, the PGC-PTSD epigenetics group follows a strategy similar to that of the ENIGMA consortium [Thompson and others 2014], in which a set of protocols and scripts are created, in this case to implement standardized QC and analysis pipelines for the Illumina HumanMethylation450 BeadChip. These scripts are performed at each participating site and analysis results are submitted to the consortium where they are assessed, collated, and meta-analyzed. In this study, we compare the performance of two QC and two analytical pipelines to control for genomic inflation, present the final PGC-PTSD epigenetics pipeline, and assess the performance of the PGC-PTSD epigenetics pipeline in a meta-analysis of age.
Materials and Methods
The PGC EWAS Cohorts
The participating cohorts, presented in Table 1, consisted of four military cohorts (MRS, PRISMO, VA-M, and VA-NCPTSD) and three civilian cohorts (DNHS, GTP, and WTC) that all measured DNA methylation (DNAm) with the Illumina HumanMethylation450K BeadChip. Descriptions of the cohorts are in the supplemental information. Each cohort consists of PTSD cases as well as trauma-exposed controls. A total of 1,147 subjects (~50% cases) were selected for inclusion in the EWAS and were subjected to the quality control and analytical pipelines.
Table 1.
PGC-PTSD EWAS Cohorts
| Civilian | Military | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Total | DNHS | GTP | WTC | MRS | PRISMO | VA-M | VA- NCP |
||
| N | 1147 | 100 | 270 | 180 | 126 | 62 | 176 | 233 | |
| Mean Age (SD) | 38.0 (−) | 53.6 (14.0) | 41.7 (12.4) | 49.7 (8.3) | 22.2 (3.0) | 27.1 (9.2) | 34.9 (9.9) | 32.0 (8.4) | |
| Current PTSD, % | 50% | 40% | 39% | 47% | 50% | 50% | 49% | 69% | |
| Male, % | 73% | 38% | 30% | 100% | 100% | 100% | 78% | 90% | |
| Race, % | |||||||||
| White | 56% | 15% | 5% | 76% | 57% | 100% | 100% | 74% | |
| Hispanic | 6% | 0% | 0% | 0% | 25% | 0% | 0% | 14% | |
| Black | 33% | 85% | 94% | 4% | 8% | 0% | 0% | 9% | |
| Asian | 1% | 0% | 0% | 0% | 3% | 0% | 0% | 2% | |
| Other | 3% | 0% | 0% | 20% | 0% | 0% | 0% | 0% | |
Posttraumatic Stress Disorder Assessment
Similar to other analyses conducted by the PGC-PTSD, our analysis required consistently defining and harmonizing PTSD diagnoses across cohorts that used different instruments and methods of diagnosis [Logue and others 2015]. We used a diagnosis of current PTSD based on the diagnostic criteria defined by each cohort’s principal investigator (see supplemental information). Individuals with lifetime diagnoses of PTSD but not current PTSD were excluded from analysis.
Quality Control Procedures
We tested two quality control protocols: the eventually proposed PGC pipeline and a Functional Normalization (Funnorm) pipeline. In the PGC pipeline (Supplemental Figure 1), study investigators first conducted a visual inspection of control-probes designed to report on each step of the Infinium protocol such as bisulfite conversion and hybridization efficiency. In addition, samples with probe detection call rates <90% and those with an average intensity value of either <50% of the experiment-wide sample mean or <2,000 arbitrary units (AU) were excluded. Probes with detection p-values >0.001 or those based on less than three beads were set to missing as were probes that cross-hybridized between autosomes and sex chromosomes [Teschendorff and others 2013]. CpG sites with missing data for >10% of samples within cohorts were excluded from analysis. Probes containing single nucleotide polymorphisms (SNPs; based on 1000 Genomes) within 10 base pairs of the target CpG were maintained in each dataset, but flagged and tracked throughout the analysis pipeline. This decision was based on the growing recognition that sequence variants can influence DNA methylation patterns throughout the genome [Smith and others 2014]. Even if an associated CpG site is influenced by genetics, such as is the case for SKA2 [Boks and others 2016; Rice and others 2008], maintaining such probes is informative to our overall goal of identifying genes important for PTSD [Gibbs and others 2010; Guintivano and others 2014; Heyn and others 2013]. Normalization of probe distribution and background differences between Type I and Type II probes was conducted using Beta Mixture Quantile Normalization (BMIQ) [Teschendorff and others 2013] after background correction. We chose BMIQ after comparing distributions of BMIQ normalized Type II probes in the Detroit Neighborhood Health Study (DNHS) with the raw distributions and distributions after applying the DASEN procedure in the R package wateRmelon (Supplemental Figure 2) [Pidsley and others 2013]. Following normalization, batch effect removal as implemented in the ComBat procedure of the SVA package in bioconductor was used to account for sources of technical variations including batch and positional effects, which can cause spurious associations [Johnson and others 2007]. Individual cohorts also controlled for additional covariates that may not have been balanced within chips but that were of interest in downstream analyses, such as case designation and sex (if relevant). Following completion of this QC pipeline, each cohort confirmed that there were no remaining sources of technical variation by examining the association of PCs of the methylation levels with chip and position using multivariate linear regression, bar plots, and heat maps.
The second QC protocol used the functional normalization procedure implemented in the R package minfi, which has been reported to remove technical variation more effectively than ComBat or other supervised methods [Aryee and others 2014; Fortin and others 2014]. Briefly, raw IDATs were loaded into R and functional normalization applied using the default of two principal components (PCs) as covariates. The resulting normalized beta matrix was then extracted and used for analysis.
Cellular Heterogeneity
DNA methylation is known to vary by cell type which impacts the measured whole blood methylation as a result of the amalgamation of the cell type proportions in each individual’s sample. To control for possible confounding by individuals’ underlying cell type heterogeneity, proportions of CD8, CD4, NK, B cells, monocytes and granulocytes were estimated using each individual’s DNA methylation data, publicly available reference data (GSE36069), and the method described by Jaffe and colleagues [Jaffe and Irizarry 2014; Reinius and others 2012], based on the Houseman algorithm [Houseman and others 2012]. CD8, CD4, NK, B cell, and monocyte cell proportions were included as covariates in our statistical analyses.
Ancestry
Accounting for population stratification has become routine practice for genetic association studies, and most recently has also been shown to be of importance in DNA methylation studies [Barfield and others 2014; Nielsen and others 2010]. GWAS methods such as principal components (PCs) derived from SNPs can be incorporated into EWAS, but were not always available for all cohorts, or all samples within a cohort. An alternative based on methylation probes that proxy nearby SNPs was developed by Barfield et al. for use in European and African American subjects [Barfield and others 2014]. Here we evaluated and extended this approach to other ancestral populations as part of the PGC-PTSD EWAS pipeline.
A subset of ancestry-diverse subjects (N=128, including European Americans, African Americans, Latinos/Native Americans and ‘others’ including East Asians) from the Marine Resiliency study (MRS) were selected based on available genome-wide genotype data (Illumina HumanOmniExpressExome array) and matching Illumina 450K methylation data [Nievergelt and others 2015]. Ancestry using GWAS data was inferred as described in Nievergelt et al. [Nievergelt and others 2013]. In brief, genotypes of 1783 ancestry-informative markers (AIMs) were used to determine a subject’s ancestry at the continental level using STRUCTUREv2.3.2.1, including prior population information of the HGDP reference set [Falush and others 2003; Li and others 2008]. Based on these ancestry estimates, subjects were placed into one of 4 groups: European Americans, African Americans, Latinos/Native Americans and ‘Others’. PCs were derived using Eigenstrat [Price and others 2006].
Ancestry estimates using methylation data were derived using subsets of methylation probes in close proximity to SNPs identified by Barfield et al. [Barfield and others 2014]. Probe sets with 0bp distance (N=7,703 CpG probes), within 1bp distance (17,995 CpG probes), and within 10bp distance (N=50,319 CpG probes) were compared. GWAS-derived PCs were visually compared to methylation-probe derived PCs and genotypes of SNPs in proximity of CpG sites were compared with respective CpG methylation values using Pearson correlation (r).
Statistical Analysis
Within each cohort, logit transformed ß values (M-values) [Du and others 2010] were modeled by linear regression as a function of PTSD, adjusting for sex, age, the estimated cell proportions, and ancestry using PCs. For cohorts with available GWAS data, the first three PCs from the GWAS were used. For cohorts without GWAS data, the method described by Barfield and colleagues was used to generate ancestry PCs directly from the EWAS data. Consistent with the original paper and our analysis (full results below), the second through fourth PCs were used as covariates in the model to control for ancestry. Note that while ancestry is a primary source for variation in GWAS, other potentially confounding factors such as cellular heterogeneity are a primary source for variation in EWAS data. Comparison with SNP data showed that ancestry inference is strongest when excluding EWAS-derived PC1. QQ-plots of the PTSD p-values were examined for evidence of genomic inflation due to unaccounted technical variation or other confounders. In addition, the genomic inflation factors (λ) were calculated for each study. Two adjustments were considered to improve the precision of the estimated variances. First, moderated t-statistics were calculated using the empirical Bayes method implemented in the R package limma [Smyth 2005]. Second, HC3 robust standard errors, which have been shown to be the most effective in samples smaller than 250, were calculated using the R package sandwich [Long and Ervin 2000; Zeileis 2004].
Cohort-level analysis results were combined using the inverse normal method [Marot and others 2009]. Briefly, one-sided p-values for each CpG site in each study were calculated from the t-statistics. Next z-scores were calculated from the one-sided p-values and weighted by the number of subjects in each study relative to the total in the meta-analysis. Two-sided p-values of the z-score were then calculated and genomic inflation examined. Finally, p-values were adjusted for multiple-testing using the False Discovery Rate (FDR) procedure at the type I error rate level of 5 percent [Benjamini and Hochberg 1995].
Sensitivity analysis
Numerous robust associations between age and DNAm have been reported [Bocklandt and others 2011; Hannum and others 2013; Horvath 2013; Horvath and others 2012; Weidner and others 2014]. Because we expect that age will be associated similarly with DNAm in each cohort, we can leverage this highly reproducible relationship to evaluate the pipeline’s performance. In each cohort, we modeled methylation as a function of age along with covariates for ancestry and gender (if applicable) and meta-analyzed the results as outlined above. In addition, we measured concordance between studies by estimating the correlation between the t-statistics of the age variable.
Power
Power for EWAS is more favorable compared to GWAS as a result of the continuous nature of the DNA methylation measures, but instead suffers from poor distribution including low variances and heteroscedasticity [Du and others 2010]. The power to detect a differentially methylated CpG site depends on the percent difference in methylation between cases and controls, the pooled variation in methylation (σ) across CpG sites, and the number of cases and controls [Liu and Hwang 2007]. We conducted simulations to test the smallest mean difference between PTSD cases and controls we could detect based on our projected sample size and a σ=0.43, which represents the highest variation reported in one of our cohorts, and thus a very conservative estimate [Orr and Liu 2009].
Results
Participating cohorts
Sample characteristics for studies that have contributed data to this first PGC-PTSD EWAS study are listed in Table 1 (N=1,147). Three of the seven studies are composed of civilians, while the remaining studies include active duty and veteran military populations. The majority of participants, especially from the military cohorts, were male (73%) and of European American (EA) descent (56%). All participants were exposed to trauma and half of participants suffered from current PTSD (50%). Data collection occurred across the United States (e.g., Atlanta, Detroit, San Diego, Durham, Boston, and New York) and Europe. While a few studies used clinical interviews, the majority of studies used self-report ratings of PTSD symptoms that relied on established cutoffs to assign caseness. A detailed description of participating cohorts is provided in the supplementary information.
Power
The power analysis shows that with our sample of 573 cases and 574 controls (N = 1147), we are sufficiently powered to find at least one CpG site with a mean methylation difference of 0.08 between cases and controls (Figure 1).
Figure 1.
Sample size vs. power to to detect differentially methylated CpG sites. The black curve indicates the number of cases and controls necessary to find a differentially methylated if only one CpG site exists, while the gray line indicates the size necessary if 10 differentially methylated sites exist.
Ancestry
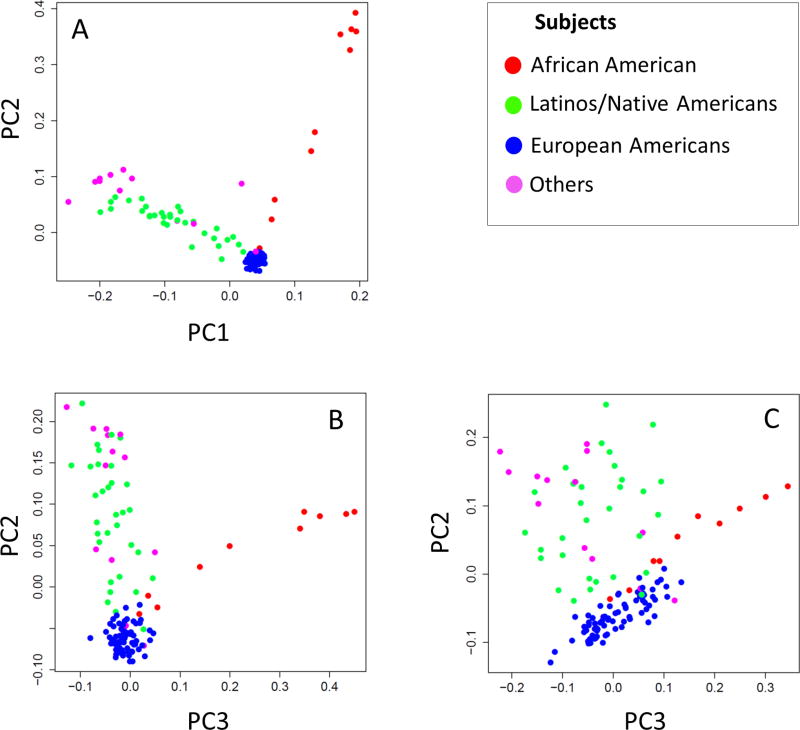
We investigated the utility of DNA methylation-based ancestry estimates based on CpGs with nearby genetic variants in cis as proposed by Barfield et al. [Barfield and others 2014]. A comparison of CpG probes with SNPs within 1bp distance (N= 17,995) and CpG probes with SNPs within 10bp distance (N= 50,319) showed a higher genotype-methylation correlation for the 1bp probes (r=0.29, p=1.8 × 10−15) than the 10bp probes (r=0.06, p=0.0015). Figure 3 shows a SNP derived PC plot based on available GWAS data including PCs 1 and 2 (panel A), PCs 2 and 3 from the methylation-based CpGs with a SNP 1bp from the probes (panel B), and the PCs from CpG probes within 10bp of a SNP (panel C). Supplemental Figure 5 shows PCs 2, 3, 4, and PCs 2, 3 and 6 for, respectively, CpGs within 1bp and 10bp of a SNP. These results along with the genotype-methylation correlations show that the use of DNA methylation ancestry estimates (PCs 2–4) using probes within 1bp of a SNP provide reliable results and are suitable as ancestry covariates in our analyses.
Figure 3.
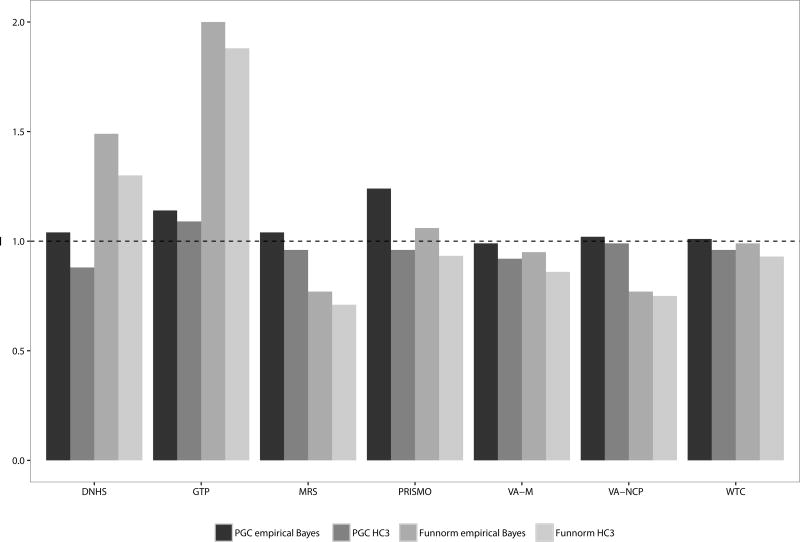
PTSD genomic inflation factors (λ) by quality control pipeline (PGC vs. Funnorm) and standard error adjustment method (empirical Bayes vs. HC3).
Quality Control Results
The number of samples and probes, not including cross-reactive probes) removed in our proposed PGC-EWAS pipeline ranged from 677 to 10,218 across studies (Supplemental Table 3). Figure 2 presents the genomic inflation factors from the analysis of PTSD for each individual study using two different quality control methods as well as two different analysis pipelines: (i) our proposed PGC-EWAS pipeline, described in detail in the Methods section above, as well as (ii) the Functional Normalization (Funnorm) QC pipeline. Resulting data from each QC pipeline were then subjected to linear regression analysis, performed with empirical Bayes and HC3 standard errors, respectively. In studies using the Funnorm pipeline there were large variations in the genomic inflation factor with two studies showing high inflation (DNHS, GTP) and two studies showing substantial deflation (MRS, VA-NCP) regardless of whether empirical Bayes or HC3 standard errors were used. Using the PGC-EWAS pipeline and HC3 standard errors, six of the seven studies showed genomic deflation with (λ < 1.0), while only one study was deflated when using empirical Bayes standard errors (Supplemental Table 1). These results indicate that the PTSD-PGC EWAS pipeline, combined with empirical Bayes standard errors, is the preferred method for cohorts participating in our meta-analysis.
Figure 2.
Ancestry inference using SNPs versus methylation probes in 128 participants of the Marine Resiliency Study (MRS). (a) Principal component (PC) plot showing ancestry inferred using SNPs from a genome-wide association study (GWAS). PC plots based on CpG probes with SNPs within 1 bp distance (b) and with SNPs within 10 bp distance (c), respectively. Subject are placed into four ancestral groups based on ancestry estimates using ancestry-informative SNPs and a reference panel (see methods).
Sensitivity analysis: Age Meta-Analysis Results
Results for the age analysis using our pipeline and no standard error adjustment are presented in Table 3. All studies reported numerous FDR-significant CpG sites but substantially more significant results were reported for the combined meta-analysis. MRS and PRISMO reported the fewest significant sites. However the age range for participants in these studies was narrower as both studies included active military personnel. The correlations of the t-statistics ranged from 0.171 to 0.692 when all sites were analyzed and from 0.441 to 0.886 among the FDR significant sites (Supplemental Figure 3). The strong correlations of the most significant sites indicate that each cohort retained the biological signal of age after QC. Of the 494 CpG sites reported to have been associated with age, 326 were significant after FDR-correction (Supplemental Table 2). In addition, a forest plot of the most significant CpG site representative of the FDR significant sites, shows a consistent direction of effect in each study (Supplemental Figure 4).
Discussion
PTSD is unique among psychiatric disorders in that its occurrence requires exposure to a significant traumatic event. With an environmental exposure embedded into the etiology of the disorder, the PTSD diagnosis affords an unusual opportunity to identify individual differences in the biological response to trauma to increase risk for, or resilience to, the disorder. Here we have introduced an international collaboration that has been established to identify epigenetic associations - specifically, DNA methylation - related to risk for, or resilience to, PTSD. We presented the development of a consistent pipeline for processing and quality-control of epigenome-wide association data comparing two quality control approaches and statistical pipelines. In our analysis of PTSD, we found that our proposed PGC-EWAS pipeline controlled for genomic inflation and deflation more consistently than functional normalization, regardless of the standard error correction used. In light of these findings, we encourage consortia with epigenome-wide methylation data to implement our quality control pipeline including checks for genomic inflation and strengthened associations with age before meta-analyzing across studies.
Through these collaborative efforts to analyze existing DNA methylation data from blood obtained from both military and civilian cohorts, we are poised to collectively address one of the main challenges of psychiatric genomics, namely the need for large, harmonized samples to adequately power genome-scale analyses. The current collaborative dataset allows detection of methylation differences around seven percent, larger than most reported methylation differences, [Vinkers and others 2015]. Additional EWAS datasets that are forthcoming will likely prove essential to detecting PTSD-associated DNA methylation differences in our planned meta-analyses. Moving forward, we anticipate that our collaborative efforts will grow to include additional cohorts from around the world; indeed, in the last year alone, several new studies have expressed interest in participating in future EWAS analyses as their data become available. In addition to the DNA methylation analysis the close allegiance with the PGC-PTSD group has laid the foundation for integrating data from GWAS, EWAS, and gene expression/transcriptome analyses. In combination with other biological measures and coordinated neuroimaging efforts [Logue and others 2015] that may become accessible through this collaboration, these system-wide integrations will facilitate a more complete understanding of the molecular architecture and biological underpinnings of PTSD.
The harmonization of some study characteristics paired with the demographic and clinical diversity of the samples, including the differences between military and civilian trauma, allows us the opportunity to identify DNA methylation patterns predictive for specific groups of individuals and types of trauma. This will not only provide insight into the heterogeneity of PTSD, but may also help explain mechanisms for the variation in conditional effects of different types of trauma on PTSD [Wisco and others 2014]. Additionally, it will also provide a framework from which DNA methylation may be informative for early risk prediction and treatment stratification.
Looking ahead, we are optimistic that our PTSD EWAS collaboration will identify blood-based DNA methylation signatures that associate reliably with PTSD. Identification of robust peripheral biomarkers is an important first step and has potential for early detection and prevention. The ultimate goal is to provide new insights into the etiology of PTSD. To truly understand the mechanistic basis of PTSD, it will be critical to compare our blood-derived epigenetic biomarkers with those from other tissues, in particular brain tissue. As a first pass, DNA methylation-based biomarkers that associate with PTSD at particular CpG sites in blood can, at this time, be compared to CpG site derived from brain tissues, thanks to the Epigenomic Roadmap datasets [Bernstein and others 2010; Kundaje and others 2015]. However, these comparisons will be limited to a comparison of DNA methylation levels in brain and blood in general, as the current data are not representative of PTSD. Over time, however, the development of PTSD brain biobanks of brain and other tissue types including blood cells, will help us to pinpoint whether blood-derived, DNA methylation biomarkers of PTSD reflect similar alterations in brain tissue, as recent work suggests may be possible for certain pathways [Daskalakis and others 2014]. Collectively, these cross-tissue efforts will provide insight into the biological pathways underlying PTSD vulnerability and will ultimately facilitate new treatment and modes of prevention.
Supplementary Material
Table 2.
Age associations using the PGC-PTSD epigenetics QC and analysis pipeline.
| Study | Sites | Sites with (FDR < 0.05) |
Sites with (p < 5×10−5) |
Sites with (p < 5×10−6) |
Sites with (p < 5×10−7) |
|---|---|---|---|---|---|
| DNHS | 455,079 | 4,766 | 1,744 | 678 | 299 |
| GTP | 453,351 | 59,100 | 21,562 | 14,299 | 9,586 |
| MRS | 455,601 | 210 | 311 | 99 | 34 |
| PRISMO | 446,688 | 246 | 316 | 121 | 41 |
| VA-M | 455,641 | 42,474 | 12,913 | 7,213 | 4,159 |
| VA-NCPTSD | 453,747 | 35,217 | 10,522 | 6,331 | 3,991 |
| WTC | 455,340 | 14,239 | 5,013 | 2,730 | 1,525 |
| Meta-Analysis | 444,164 | 119,308 | 57,332 | 46,629 | 38,656 |
Acknowledgments
This work was supported by the U.S. Army Medical Research and Materiel Command and the National Institute of Mental Health (NIMH; R01MH108826) as well as the Biomedical and Laboratory Research and Development (#I01BX002577). We appreciate the technical support of all of the staff, volunteers and participants from the Grady Trauma Project, supported by the National Institutes of Mental Health (MH096764 and MH071537). DNHS, which is grateful to all of the participants and staff for their contributions, was funded by NIH Awards R01DA022720, R01DA022720-S1, and RC1MH088283. The Marine Corps, Navy Bureau of Medicine and Surgery (BUMED) and VA Health Research and Development (HSR&D) provided funding for MRS data collection and analysis and NIH R01MH093500 funded the GWAS assays and analysis. Acknowledged are Victoria B. Risbrough Ph.D (VA San Diego Healthcare System & UCSD), Mark A. Geyer (UCSD), Daniel T. O’Connor (UCSD), all MRS investigators, and all MRS investigators, as well as the MRS administrative core and data collection staff listed in the Methods article (Baker et al, Prev Chronic Dis. 2012;9(10):E97). The authors also thank the Marine and Navy Corpsmen volunteers for military service and participation in MRS. Data collection of PRISMO was funded by the Dutch Ministry of Defence, and DNA methylation analyses were funded by the VENI Award fellowship from the Netherlands Organisation for Scientific Research (NWO, grant number 916.11.086). The VA Boston-National Center for PTSD Study research was supported in part by National Institute of Mental Health Award RO1MH079806, Department of Veterans Affairs, Clinical Science Research & Development Program Award 5I01CX000431-02, Department of Veterans Affairs, Biomedical Laboratory Research & Development Program Award 1I01BX002150-01, the Translational Research Center for TBI and Stress Disorders (TRACTS), a VA Rehabilitation Research and Development Traumatic Brain Injury Center of Excellence (B9254-C), and the Cooperative Studies Program, Department of Veterans Affairs. This research is the result of work supported with resources and the use of facilities at the Pharmacogenomics Analysis Laboratory, Research and Development Service, Central Arkansas Veterans Healthcare System, Little Rock, Arkansas. This work was also supported by a Career Development Award to E. J. Wolf from the Department of Veterans Affairs, Clinical Sciences Research, and Development Program. Dr. Kimbrel was supported by a Career Development Award (#IK2CX000525) from the Clinical Science Research and Development (CSR&D). Dr. Beckham was supported by a Research Career Scientist Award (#11S-RCS-009) from the CSR&D Service of VA ORD. This research was also supported, in part, by a Merit Award (#I01BX002577) from the Biomedical Laboratory Research and Development (BLR&D) Service of VA ORD. The VA Mid-Atlantic Mental Illness Research, Education, and Clinical Center Workgroup includes John A. Fairbank, Mira Brancu, Patrick S. Calhoun, Eric A. Dedert, Eric B. Elbogen, Kimberly T. Green, Robin A. Hurley, Angela C. Kirby, Jason D. Kilts, Christine E. Marx, Gregory McCarthy, Scott D. McDonald, Marinell Miller-Mumford, Scott D. Moore, Rajendra A. Morey, Jennifer C. Naylor, Treven C. Pickett, Jared Rowland, Jennifer J. Runnals, Cindy Swinkels, Steven T. Szabo, Katherine H. Taber, Larry A. Tupler, Elizabeth E. Van Voorhees, H. Ryan Wagner, Richard D. Weiner, and Ruth Yoash-Gantz. Data collection from the World Trade Center (WTC) cohort was supported by CDC awards U01OH010416, U01OH010718, and 200-2011-39410.
The views expressed in this article are those of the authors and do not necessarily reflect the position or policy of the VA, NIH, or the United States government.
Footnotes
Conflict of Interest
Dr. Youssef’s disclosures include Speaker CME honoraria from the Georgia Department of Behavioral Health and Developmental Disabilities (DHBDD). Dr. Youssef received research support from the Department of Veteran Affairs and The Augusta Biomedical Research Corporation. Dr. Stein has in the past 3 years received payments for editorial work from UpToDate, Biological Psychiatry, and Depression and Anxiety. He has also in the past 3 years been paid as a consultant for Actelion Pharmaceuticals, Janssen, Pfizer, Resilience Therapeutics, and Tonix Pharmaceuticals. No other author declares any conflict of interest.
References
- Almli LM, Srivastava A, Fani N, Kerley K, Mercer KB, Feng H, Bradley B, Ressler KJ. Follow-up and Extension of a Prior Genome-wide Association Study of Posttraumatic Stress Disorder: Gene x Environment Associations and Structural Magnetic Resonance Imaging in a Highly Traumatized African-American Civilian Population. Biological psychiatry. 2014 doi: 10.1016/j.biopsych.2014.01.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aryee MJ, Jaffe AE, Corrada-Bravo H, Ladd-Acosta C, Feinberg AP, Hansen KD, Irizarry RA. Minfi: a flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics. 2014:btu049. doi: 10.1093/bioinformatics/btu049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Association AP. DSM-IV-TR: Diagnostic and statistical manual of mental disorders, text revision. Washington, DC: American Psychiatric Association; 2000. p. 75. [Google Scholar]
- Barfield RT, Almli LM, Kilaru V, Smith AK, Mercer KB, Duncan R, Klengel T, Mehta D, Binder EB, Epstein MP, Ressler KJ, Conneely KN. Accounting for population stratification in DNA methylation studies. Genetic epidemiology. 2014;38(3):231–241. doi: 10.1002/gepi.21789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. Journal of the Royal Statistical Society Series B (Methodological) 1995:289–300. [Google Scholar]
- Benjet C, Bromet E, Karam EG, Kessler RC, McLaughlin KA, Ruscio AM, Shahly V, Stein DJ, Petukhova M, Hill E, Alonso J, Atwoli L, Bunting B, Bruffaerts R, Caldas-de-Almeida JM, de Girolamo G, Florescu S, Gureje O, Huang Y, Lepine JP, Kawakami N, Kovess-Masfety V, Medina-Mora ME, Navarro-Mateu F, Piazza M, Posada-Villa J, Scott KM, Shalev A, Slade T, ten Have M, Torres Y, Viana MC, Zarkov Z, Koenen KC. The epidemiology of traumatic event exposure worldwide: results from the World Mental Health Survey Consortium. Psychological medicine. 2016;46(2):327–343. doi: 10.1017/S0033291715001981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bernstein BE, Stamatoyannopoulos JA, Costello JF, Ren B, Milosavljevic A, Meissner A, Kellis M, Marra MA, Beaudet AL, Ecker JR, Farnham PJ, Hirst M, Lander ES, Mikkelsen TS, Thomson JA. The NIH Roadmap Epigenomics Mapping Consortium. Nat Biotechnol. 2010;28(10):1045–1048. doi: 10.1038/nbt1010-1045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bocklandt S, Lin W, Sehl ME, Sánchez FJ, Sinsheimer JS, Horvath S, Vilain E. Epigenetic predictor of age. 2011 doi: 10.1371/journal.pone.0014821. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boks MP, Rutten BP, Geuze E, Houtepen LC, Vermetten E, Kaminsky Z, Vinkers CH. SKA2 Methylation is Involved in Cortisol Stress Reactivity and Predicts the Development of Post-Traumatic Stress Disorder (PTSD) After Military Deployment. Neuropsychopharmacology. 2016;41(5):1350–1356. doi: 10.1038/npp.2015.286. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bonasio R, Tu S, Reinberg D. Molecular signals of epigenetic states. science. 2010;330(6004):612–616. doi: 10.1126/science.1191078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boscarino JA. A prospective study of PTSD and early-age heart disease mortality among Vietnam veterans: implications for surveillance and prevention. Psychosomatic medicine. 2008;70(6):668–676. doi: 10.1097/PSY.0b013e31817bccaf. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brady KT, Killeen TK, Brewerton T, Lucerini S. Comorbidity of psychiatric disorders and posttraumatic stress disorder. Journal of Clinical Psychiatry. 2000 [PubMed] [Google Scholar]
- Breslau N, Kessler RC, Chilcoat HD, Schultz LR, Davis GC, Andreski P. Trauma and posttraumatic stress disorder in the community: the 1996 Detroit Area Survey of Trauma. Archives of general psychiatry. 1998;55(7):626–632. doi: 10.1001/archpsyc.55.7.626. [DOI] [PubMed] [Google Scholar]
- Chang S-C, Koenen KC, Galea S, Aiello AE, Soliven R, Wildman DE, Uddin M. Molecular variation at the SLC6A3 locus predicts lifetime risk of PTSD in the Detroit Neighborhood Health Study. PLoS One. 2012;7(6):e39184. doi: 10.1371/journal.pone.0039184. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coughlin SS. Post-traumatic Stress Disorder and Cardiovascular Disease. The open cardiovascular medicine journal. 2011;5:164–170. doi: 10.2174/1874192401105010164. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Daskalakis NP, Cohen H, Cai G, Buxbaum JD, Yehuda R. Expression profiling associates blood and brain glucocorticoid receptor signaling with trauma-related individual differences in both sexes. Proc Natl Acad Sci U S A. 2014;111(37):13529–13534. doi: 10.1073/pnas.1401660111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- David D, Woodward C, Esquenazi J, Mellman TA. Comparison of comorbid physical illnesses among veterans with PTSD and veterans with alcohol dependence. Psychiatric Services. 2014 doi: 10.1176/appi.ps.55.1.82. [DOI] [PubMed] [Google Scholar]
- Du P, Zhang X, Huang C-C, Jafari N, Kibbe WA, Hou L, Lin SM. Comparison of Beta-value and M-value methods for quantifying methylation levels by microarray analysis. BMC bioinformatics. 2010;11(1):587. doi: 10.1186/1471-2105-11-587. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Falush D, Stephens M, Pritchard JK. Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics. 2003;164(4):1567–1587. doi: 10.1093/genetics/164.4.1567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feinberg AP. Phenotypic plasticity and the epigenetics of human disease. Nature. 2007;447(7143):433–440. doi: 10.1038/nature05919. [DOI] [PubMed] [Google Scholar]
- Fortin J-P, Labbe A, Lemire M, Zanke BW, Hudson TJ, Fertig EJ, Greenwood CM, Hansen KD. Functional normalization of 450k methylation array data improves replication in large cancer studies. Genome Biol. 2014;15(11):503. doi: 10.1186/s13059-014-0503-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gibbs JR, van der Brug MP, Hernandez DG, Traynor BJ, Nalls MA, Lai S-L, Arepalli S, Dillman A, Rafferty IP, Troncoso J. Abundant quantitative trait loci exist for DNA methylation and gene expression in human brain. PLoS Genet. 2010;6(5):e1000952. doi: 10.1371/journal.pgen.1000952. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guffanti G, Galea S, Yan L, Roberts AL, Solovieff N, Aiello AE, Smoller JW, De Vivo I, Ranu H, Uddin M, Wildman DE, Purcell S, Koenen KC. Genome-wide association study implicates a novel RNA gene, the lincRNA AC068718.1, as a risk factor for post-traumatic stress disorder in women. Psychoneuroendocrinology. 2013;38(12):3029–3038. doi: 10.1016/j.psyneuen.2013.08.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guintivano J, Brown T, Newcomer A, Jones M, Cox O, Maher BS, Eaton WW, Payne JL, Wilcox HC, Kaminsky ZA. Identification and replication of a combined epigenetic and genetic biomarker predicting suicide and suicidal behaviors. American journal of psychiatry. 2014 doi: 10.1176/appi.ajp.2014.14010008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hannum G, Guinney J, Zhao L, Zhang L, Hughes G, Sadda S, Klotzle B, Bibikova M, Fan J-B, Gao Y. Genome-wide methylation profiles reveal quantitative views of human aging rates. Molecular cell. 2013;49(2):359–367. doi: 10.1016/j.molcel.2012.10.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harper KN, Peters BA, Gamble MV. Batch effects and pathway analysis: two potential perils in cancer studies involving DNA methylation array analysis. Cancer Epidemiology Biomarkers & Prevention. 2013;22(6):1052–1060. doi: 10.1158/1055-9965.EPI-13-0114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heppner PS, Crawford EF, Haji UA, Afari N, Hauger RL, Dashevsky BA, Horn PS, Nunnink SE, Baker DG. The association of posttraumatic stress disorder and metabolic syndrome: a study of increased health risk in veterans. BMC Med. 2009;7:1. doi: 10.1186/1741-7015-7-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heyn H, Moran S, Hernando-Herraez I, Sayols S, Gomez A, Sandoval J, Monk D, Hata K, Marques-Bonet T, Wang L. DNA methylation contributes to natural human variation. Genome research. 2013;23(9):1363–1372. doi: 10.1101/gr.154187.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horvath S. DNA methylation age of human tissues and cell types. Genome biology. 2013;14(10):R115. doi: 10.1186/gb-2013-14-10-r115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horvath S, Zhang Y, Langfelder P, Kahn RS, Boks M, van Eijk K, van den Berg LH, Ophoff RA. Aging effects on DNA methylation modules in human brain and blood tissue. Genome Biol. 2012;13(10):R97. doi: 10.1186/gb-2012-13-10-r97. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Houseman EA, Accomando WP, Koestler DC, Christensen BC, Marsit CJ, Nelson HH, Wiencke JK, Kelsey KT. DNA methylation arrays as surrogate measures of cell mixture distribution. BMC bioinformatics. 2012;13(1):1. doi: 10.1186/1471-2105-13-86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jaffe AE, Irizarry RA. Accounting for cellular heterogeneity is critical in epigenome-wide association studies. Genome Biol. 2014;15(2):R31. doi: 10.1186/gb-2014-15-2-r31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jakovljevic M, Babic D, Crncevic Z, Martinac M, Maslov B, Topic R. Metabolic syndrome and depression in war veterans with post-traumatic stress disorder. Psychiatr Danub. 2008;20(3):406–410. [PubMed] [Google Scholar]
- Jirtle RL, Skinner MK. Environmental epigenomics and disease susceptibility. Nature Reviews: Genetics. 2007;8(4):253–262. doi: 10.1038/nrg2045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson WE, Li C, Rabinovic A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics. 2007;8(1):118–127. doi: 10.1093/biostatistics/kxj037. [DOI] [PubMed] [Google Scholar]
- Kessler RC. Posttraumatic stress disorder: the burden to the individual and to society. The Journal of clinical psychiatry. 2000;61(suppl 5):4–12. [PubMed] [Google Scholar]
- Kessler RC, Berglund P, Demler O, Jin R, Merikangas KR, Walters EE. Lifetime prevalence and age-of-onset distributions of DSM-IV disorders in the National Comorbidity Survey Replication. Archives of general psychiatry. 2005;62(6):593–602. doi: 10.1001/archpsyc.62.6.593. [DOI] [PubMed] [Google Scholar]
- Koenen KC, Uddin M, Chang SC, Aiello AE, Wildman DE, Goldmann E, Galea S. SLC6A4 methylation modifies the effect of the number of traumatic events on risk for posttraumatic stress disorder. Depression and anxiety. 2011;28(8):639–647. doi: 10.1002/da.20825. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kubzansky LD, Koenen KC, Jones C, Eaton WW. A prospective study of posttraumatic stress disorder symptoms and coronary heart disease in women. Health Psychol. 2009;28(1):125–130. doi: 10.1037/0278-6133.28.1.125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kubzansky LD, Koenen KC, Spiro A, 3rd, Vokonas PS, Sparrow D. Prospective study of posttraumatic stress disorder symptoms and coronary heart disease in the Normative Aging Study. Arch Gen Psychiatry. 2007;64(1):109–116. doi: 10.1001/archpsyc.64.1.109. [DOI] [PubMed] [Google Scholar]
- Kundaje A, Meuleman W, Ernst J, Bilenky M, Yen A, Heravi-Moussavi A, Kheradpour P, Zhang Z, Wang J, Ziller MJ, Amin V, Whitaker JW, Schultz MD, Ward LD, Sarkar A, Quon G, Sandstrom RS, Eaton ML, Wu YC, Pfenning AR, Wang X, Claussnitzer M, Liu Y, Coarfa C, Harris RA, Shoresh N, Epstein CB, Gjoneska E, Leung D, Xie W, Hawkins RD, Lister R, Hong C, Gascard P, Mungall AJ, Moore R, Chuah E, Tam A, Canfield TK, Hansen RS, Kaul R, Sabo PJ, Bansal MS, Carles A, Dixon JR, Farh KH, Feizi S, Karlic R, Kim AR, Kulkarni A, Li D, Lowdon R, Elliott G, Mercer TR, Neph SJ, Onuchic V, Polak P, Rajagopal N, Ray P, Sallari RC, Siebenthall KT, Sinnott-Armstrong NA, Stevens M, Thurman RE, Wu J, Zhang B, Zhou X, Beaudet AE, Boyer LA, De Jager PL, Farnham PJ, Fisher SJ, Haussler D, Jones SJ, Li W, Marra MA, McManus MT, Sunyaev S, Thomson JA, Tlsty TD, Tsai LH, Wang W, Waterland RA, Zhang MQ, Chadwick LH, Bernstein BE, Costello JF, Ecker JR, Hirst M, Meissner A, Milosavljevic A, Ren B, Stamatoyannopoulos JA, Wang T, Kellis M. Integrative analysis of 111 reference human epigenomes. Nature. 2015;518(7539):317–330. doi: 10.1038/nature14248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li JZ, Absher DM, Tang H, Southwick AM, Casto AM, Ramachandran S, Cann HM, Barsh GS, Feldman M, Cavalli-Sforza LL, Myers RM. Worldwide human relationships inferred from genome-wide patterns of variation. Science. 2008;319(5866):1100–1104. doi: 10.1126/science.1153717. [DOI] [PubMed] [Google Scholar]
- Lin DY, Zeng D. On the relative efficiency of using summary statistics versus individual-level data in meta-analysis. Biometrika. 2010;97(2):321–332. doi: 10.1093/biomet/asq006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu P, Hwang JG. Quick calculation for sample size while controlling false discovery rate with application to microarray analysis. Bioinformatics. 2007;23(6):739–746. doi: 10.1093/bioinformatics/btl664. [DOI] [PubMed] [Google Scholar]
- Logue MW, Amstadter AB, Baker DG, Duncan L, Koenen KC, Liberzon I, Miller MW, Morey RA, Nievergelt CM, Ressler KJ. The Psychiatric Genomics Consortium Posttraumatic Stress Disorder Workgroup: Posttraumatic stress disorder enters the age of large-scale genomic collaboration. Neuropsychopharmacology. 2015 doi: 10.1038/npp.2015.118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Logue MW, Baldwin C, Guffanti G, Melista E, Wolf EJ, Reardon AF, Uddin M, Wildman D, Galea S, Koenen KC. A genome-wide association study of post-traumatic stress disorder identifies the retinoid-related orphan receptor alpha (RORA) gene as a significant risk locus. Molecular psychiatry. 2013;18(8):937–942. doi: 10.1038/mp.2012.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Long JS, Ervin LH. Using heteroscedasticity consistent standard errors in the linear regression model. The American Statistician. 2000;54(3):217–224. [Google Scholar]
- Luft B, Schechter C, Kotov R, Broihier J, Reissman D, Guerrera K, Udasin I, Moline J, Harrison D, Friedman-Jimenez G. Exposure, probable PTSD and lower respiratory illness among World Trade Center rescue, recovery and clean-up workers. Psychological medicine. 2012;42(05):1069–1079. doi: 10.1017/S003329171100256X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marot G, Foulley J-L, Mayer C-D, Jaffrézic F. Moderated effect size and P-value combinations for microarray meta-analyses. Bioinformatics. 2009;25(20):2692–2699. doi: 10.1093/bioinformatics/btp444. [DOI] [PubMed] [Google Scholar]
- Mathew T, Nordstrom K. On the equivalence of meta-analysis using literature and using individual patient data. Biometrics. 1999;55(4):1221–1223. doi: 10.1111/j.0006-341x.1999.01221.x. [DOI] [PubMed] [Google Scholar]
- Maunakea AK, Nagarajan RP, Bilenky M, Ballinger TJ, D’Souza C, Fouse SD, Johnson BE, Hong C, Nielsen C, Zhao Y. Conserved role of intragenic DNA methylation in regulating alternative promoters. Nature. 2010;466(7303):253–257. doi: 10.1038/nature09165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McGowan PO, Sasaki A, D'Alessio AC, Dymov S, Labonté B, Szyf M, Turecki G, Meaney MJ. Epigenetic regulation of the glucocorticoid receptor in human brain associates with childhood abuse. Nature neuroscience. 2009;12(3):342–348. doi: 10.1038/nn.2270. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mehta D, Klengel T, Conneely KN, Smith AK, Altmann A, Pace TW, Rex-Haffner M, Loeschner A, Gonik M, Mercer KB. Childhood maltreatment is associated with distinct genomic and epigenetic profiles in posttraumatic stress disorder. Proceedings of the National Academy of Sciences. 2013;110(20):8302–8307. doi: 10.1073/pnas.1217750110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moser DA, Paoloni-Giacobino A, Stenz L, Adouan W, Manini A, Suardi F, Cordero MI, Vital M, Rossignol AS, Rusconi-Serpa S. BDNF methylation and maternal brain activity in a violence-related sample. PLoS ONE. 2015;10(12):1. doi: 10.1371/journal.pone.0143427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Najt P, Fusar-Poli P, Brambilla P. Co-occurring mental and substance abuse disorders: a review on the potential predictors and clinical outcomes. Psychiatry Research. 2011;186(2):159–164. doi: 10.1016/j.psychres.2010.07.042. [DOI] [PubMed] [Google Scholar]
- Nielsen DA, Hamon S, Yuferov V, Jackson C, Ho A, Ott J, Kreek MJ. Ethnic diversity of DNA methylation in the OPRM1 promoter region in lymphocytes of heroin addicts. Human genetics. 2010;127(6):639–649. doi: 10.1007/s00439-010-0807-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nievergelt CM, Maihofer AX, Mustapic M, Yurgil KA, Schork NJ, Miller MW, Logue MW, Geyer MA, Risbrough VB, O’Connor DT. Genomic predictors of combat stress vulnerability and resilience in US Marines: A genome-wide association study across multiple ancestries implicates PRTFDC1 as a potential PTSD gene. Psychoneuroendocrinology. 2015;51:459–471. doi: 10.1016/j.psyneuen.2014.10.017. [DOI] [PubMed] [Google Scholar]
- Nievergelt CM, Maihofer AX, Shekhtman T, Libiger O, Wang X, Kidd KK, Kidd JR. Inference of human continental origin and admixture proportions using a highly discriminative ancestry informative 41-SNP panel. Investigative genetics. 2013;4(1):13. doi: 10.1186/2041-2223-4-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olkin I, Sampson A. Comparison of meta-analysis versus analysis of variance of individual patient data. Biometrics. 1998;54(1):317–322. [PubMed] [Google Scholar]
- Orr M, Liu P. Sample size estimation while controlling false discovery rate for microarray experiments using the ssize. fdr package. RJ. 2009;1:47–53. [Google Scholar]
- Pidsley R, Wong CC, Volta M, Lunnon K, Mill J, Schalkwyk LC. A data-driven approach to preprocessing Illumina 450K methylation array data. BMC genomics. 2013;14(1):293. doi: 10.1186/1471-2164-14-293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Price AL, Patterson NJ, Plenge RM, Weinblatt ME, Shadick NA, Reich D. Principal components analysis corrects for stratification in genome-wide association studies. Nature genetics. 2006;38(8):904–909. doi: 10.1038/ng1847. [DOI] [PubMed] [Google Scholar]
- Reinius LE, Acevedo N, Joerink M, Pershagen G, Dahlén S-E, Greco D, Söderhäll C, Scheynius A, Kere J. Differential DNA methylation in purified human blood cells: implications for cell lineage and studies on disease susceptibility. PloS one. 2012;7(7):e41361. doi: 10.1371/journal.pone.0041361. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Resnick HS, Kilpatrick DG, Dansky BS, Saunders BE, Best CL. Prevalence of civilian trauma and posttraumatic stress disorder in a representative national sample of women. Journal of consulting and clinical psychology. 1993;61(6):984. doi: 10.1037//0022-006x.61.6.984. [DOI] [PubMed] [Google Scholar]
- Ressler KJ, Mercer KB, Bradley B, Jovanovic T, Mahan A, Kerley K, Norrholm SD, Kilaru V, Smith AK, Myers AJ, Ramirez M, Engel A, Hammack SE, Toufexis D, Braas KM, Binder EB, May V. Post-traumatic stress disorder is associated with PACAP and the PAC1 receptor. Nature. 2011;470(7335):492–497. doi: 10.1038/nature09856. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rice L, Waters CE, Eccles J, Garside H, Sommer P, Kay P, Blackhall FH, Zeef L, Telfer B, Stratford I. Identification and functional analysis of SKA2 interaction with the glucocorticoid receptor. Journal of Endocrinology. 2008;198(3):499–509. doi: 10.1677/JOE-08-0019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roth TL, Lubin FD, Funk AJ, Sweatt JD. Lasting Epigenetic Influence of Early-Life Adversity on the BDNF Gene. Biological psychiatry. 2009 doi: 10.1016/j.biopsych.2008.11.028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sareen J, Houlahan T, Cox BJ, Asmundson GJ. Anxiety disorders associated with suicidal ideation and suicide attempts in the National Comorbidity Survey. The Journal of nervous and mental disease. 2005;193(7):450–454. doi: 10.1097/01.nmd.0000168263.89652.6b. [DOI] [PubMed] [Google Scholar]
- Sartor CE, Grant JD, Lynskey MT, McCutcheon VV, Waldron M, Statham DJ, Bucholz KK, Madden PA, Heath AC, Martin NG, Nelson EC. Common heritable contributions to low-risk trauma, high-risk trauma, posttraumatic stress disorder, and major depression. Archives of General Psychiatry. 2012;69(3):293–299. doi: 10.1001/archgenpsychiatry.2011.1385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sartor CE, McCutcheon VV, Pommer NE, Nelson EC, Grant JD, Duncan AE, Waldron M, Bucholz KK, Madden PA, Heath AC. Common genetic and environmental contributions to post-traumatic stress disorder and alcohol dependence in young women. Psychological medicine. 2011;41(7):1497–1505. doi: 10.1017/S0033291710002072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sipahi L, Wildman DE, Aiello AE, Koenen KC, Galea S, Abbas A, Uddin M. Longitudinal epigenetic variation of DNA methyltransferase genes is associated with vulnerability to post-traumatic stress disorder. Psychological medicine. 2014;44(15):3165–3179. doi: 10.1017/S0033291714000968. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith AK, Conneely KN, Kilaru V, Mercer KB, Weiss TE, Bradley B, Tang Y, Gillespie CF, Cubells JF, Ressler KJ. Differential immune system DNA methylation and cytokine regulation in post-traumatic stress disorder. American Journal of Medical Genetics Part B: Neuropsychiatric Genetics. 2011;156(6):700–708. doi: 10.1002/ajmg.b.31212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith AK, Kilaru V, Kocak M, Almli LM, Mercer KB, Ressler KJ, Tylavsky FA, Conneely KN. Methylation quantitative trait loci (meQTLs) are consistently detected across ancestry, developmental stage, and tissue type. BMC genomics. 2014;15(1):145. doi: 10.1186/1471-2164-15-145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smyth GK. Bioinformatics and computational biology solutions using R and Bioconductor. Springer; 2005. Limma: linear models for microarray data; pp. 397–420. [Google Scholar]
- Stein MB, Jang KL, Taylor S, Vernon PA, Livesley WJ. Genetic and environmental influences on trauma exposure and posttraumatic stress disorder symptoms: a twin study. Am J Psychiatry. 2002;159(10):1675–1681. doi: 10.1176/appi.ajp.159.10.1675. [DOI] [PubMed] [Google Scholar]
- Teschendorff AE, Marabita F, Lechner M, Bartlett T, Tegner J, Gomez-Cabrero D, Beck S. A beta-mixture quantile normalization method for correcting probe design bias in Illumina Infinium 450 k DNA methylation data. Bioinformatics. 2013;29(2):189–196. doi: 10.1093/bioinformatics/bts680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thompson PM, Stein JL, Medland SE, Hibar DP, Vasquez AA, Renteria ME, Toro R, Jahanshad N, Schumann G, Franke B, Wright MJ, Martin NG, Agartz I, Alda M, Alhusaini S, Almasy L, Almeida J, Alpert K, Andreasen NC, Andreassen OA, Apostolova LG, Appel K, Armstrong NJ, Aribisala B, Bastin ME, Bauer M, Bearden CE, Bergmann O, Binder EB, Blangero J, Bockholt HJ, Boen E, Bois C, Boomsma DI, Booth T, Bowman IJ, Bralten J, Brouwer RM, Brunner HG, Brohawn DG, Buckner RL, Buitelaar J, Bulayeva K, Bustillo JR, Calhoun VD, Cannon DM, Cantor RM, Carless MA, Caseras X, Cavalleri GL, Chakravarty MM, Chang KD, Ching CR, Christoforou A, Cichon S, Clark VP, Conrod P, Coppola G, Crespo-Facorro B, Curran JE, Czisch M, Deary IJ, de Geus EJ, den Braber A, Delvecchio G, Depondt C, de Haan L, de Zubicaray GI, Dima D, Dimitrova R, Djurovic S, Dong H, Donohoe G, Duggirala R, Dyer TD, Ehrlich S, Ekman CJ, Elvsashagen T, Emsell L, Erk S, Espeseth T, Fagerness J, Fears S, Fedko I, Fernandez G, Fisher SE, Foroud T, Fox PT, Francks C, Frangou S, Frey EM, Frodl T, Frouin V, Garavan H, Giddaluru S, Glahn DC, Godlewska B, Goldstein RZ, Gollub RL, Grabe HJ, Grimm O, Gruber O, Guadalupe T, Gur RE, Gur RC, Goring HH, Hagenaars S, Hajek T, Hall GB, Hall J, Hardy J, Hartman CA, Hass J, Hatton SN, Haukvik UK, Hegenscheid K, Heinz A, Hickie IB, Ho BC, Hoehn D, Hoekstra PJ, Hollinshead M, Holmes AJ, Homuth G, Hoogman M, Hong LE, Hosten N, Hottenga JJ, Hulshoff Pol HE, Hwang KS, Jack CR, Jr, Jenkinson M, Johnston C, Jonsson EG, Kahn RS, Kasperaviciute D, Kelly S, Kim S, Kochunov P, Koenders L, Kramer B, Kwok JB, Lagopoulos J, Laje G, Landen M, Landman BA, Lauriello J, Lawrie SM, Lee PH, Le Hellard S, Lemaitre H, Leonardo CD, Li CS, Liberg B, Liewald DC, Liu X, Lopez LM, Loth E, Lourdusamy A, Luciano M, Macciardi F, Machielsen MW, Macqueen GM, Malt UF, Mandl R, Manoach DS, Martinot JL, Matarin M, Mather KA, Mattheisen M, Mattingsdal M, Meyer-Lindenberg A, McDonald C, McIntosh AM, McMahon FJ, McMahon KL, Meisenzahl E, Melle I, Milaneschi Y, Mohnke S, Montgomery GW, Morris DW, Moses EK, Mueller BA, Munoz Maniega S, Muhleisen TW, Muller-Myhsok B, Mwangi B, Nauck M, Nho K, Nichols TE, Nilsson LG, Nugent AC, Nyberg L, Olvera RL, Oosterlaan J, Ophoff RA, Pandolfo M, Papalampropoulou-Tsiridou M, Papmeyer M, Paus T, Pausova Z, Pearlson GD, Penninx BW, Peterson CP, Pfennig A, Phillips M, Pike GB, Poline JB, Potkin SG, Putz B, Ramasamy A, Rasmussen J, Rietschel M, Rijpkema M, Risacher SL, Roffman JL, Roiz-Santianez R, Romanczuk-Seiferth N, Rose EJ, Royle NA, Rujescu D, Ryten M, Sachdev PS, Salami A, Satterthwaite TD, Savitz J, Saykin AJ, Scanlon C, Schmaal L, Schnack HG, Schork AJ, Schulz SC, Schur R, Seidman L, Shen L, Shoemaker JM, Simmons A, Sisodiya SM, Smith C, Smoller JW, Soares JC, Sponheim SR, Sprooten E, Starr JM, Steen VM, Strakowski S, Strike L, Sussmann J, Samann PG, Teumer A, Toga AW, Tordesillas-Gutierrez D, Trabzuni D, Trost S, Turner J, Van den Heuvel M, van der Wee NJ, van Eijk K, van Erp TG, van Haren NE, van 't Ent D, van Tol MJ, Valdes Hernandez MC, Veltman DJ, Versace A, Volzke H, Walker R, Walter H, Wang L, Wardlaw JM, Weale ME, Weiner MW, Wen W, Westlye LT, Whalley HC, Whelan CD, White T, Winkler AM, Wittfeld K, Woldehawariat G, Wolf C, Zilles D, Zwiers MP, Thalamuthu A, Schofield PR, Freimer NB, Lawrence NS, Drevets W Alzheimer's Disease Neuroimaging Initiative ECICSYSG. The ENIGMA Consortium: large-scale collaborative analyses of neuroimaging and genetic data. Brain imaging and behavior. 2014;8(2):153–182. doi: 10.1007/s11682-013-9269-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- True WR, Rice J, Eisen SA, Heath AC, Goldberg J, Lyons MJ, Nowak J. A twin study of genetic and environmental contributions to liability for posttraumatic stress symptoms. Arch Gen Psychiatry. 1993;50(4):257–264. doi: 10.1001/archpsyc.1993.01820160019002. [DOI] [PubMed] [Google Scholar]
- Tyrka AR, Price LH, Marsit C, Walters OC, Carpenter LL. Childhood adversity and epigenetic modulation of the leukocyte glucocorticoid receptor: preliminary findings in healthy adults. PloS one. 2012;7(1):e30148. doi: 10.1371/journal.pone.0030148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Uddin M, Aiello AE, Wildman DE, Koenen KC, Pawelec G, de Los Santos R, Goldmann E, Galea S. Epigenetic and immune function profiles associated with posttraumatic stress disorder. Proc Natl Acad Sci U S A. 2010;107(20):9470–9475. doi: 10.1073/pnas.0910794107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vinkers CH, Kalafateli AL, Rutten BP, Kas MJ, Kaminsky Z, Turner JD, Boks MP. Traumatic stress and human DNA methylation: a critical review. Epigenomics. 2015;7(4):593–608. doi: 10.2217/epi.15.11. [DOI] [PubMed] [Google Scholar]
- Weaver ICG, Cervoni N, D'Alessio AC, Champagne FA, Seckl JR, Szyf M, Meaney MJ. Epigenetic programming through maternal behavior. Nature Neuroscience. 2004;7:847–854. doi: 10.1038/nn1276. [DOI] [PubMed] [Google Scholar]
- Weidner CI, Lin Q, Koch CM, Eisele L, Beier F, Ziegler P, Bauerschlag DO, Jöckel K-H, Erbel R, Mühleisen TW. Aging of blood can be tracked by DNA methylation changes at just three CpG sites. Genome biology. 2014;15(2):1. doi: 10.1186/gb-2014-15-2-r24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wisco BE, Marx BP, Wolf EJ, Miller MW, Southwick SM, Pietrzak RH. Posttraumatic stress disorder in the US veteran population: results from the National Health and Resilience in Veterans Study. The Journal of clinical psychiatry. 2014;75(12):1338–1346. doi: 10.4088/JCP.14m09328. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xian H, Chantarujikapong SI, Scherrer JF, Eisen SA, Lyons MJ, Goldberg J, Tsuang M, True WR. Genetic and environmental influences on posttraumatic stress disorder, alcohol and drug dependence in twin pairs. Drug & Alcohol Dependence. 2000;61(1):95–102. doi: 10.1016/s0376-8716(00)00127-7. [DOI] [PubMed] [Google Scholar]
- Xie P, Kranzler HR, Yang C, Zhao H, Farrer LA, Gelernter J. Genome-wide association study identifies new susceptibility loci for posttraumatic stress disorder. Biological psychiatry. 2013;74(9):656–663. doi: 10.1016/j.biopsych.2013.04.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zeileis A. Econometric computing with HC and HAC covariance matrix estimators 2004 [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.