Abstract
Although average survival rates for lung cancer have improved, earlier and better diagnosis remains a priority. One promising approach to assisting earlier and safer diagnosis of lung lesions is bronchoalveolar lavage (BAL), which provides a sample of lung tissue as well as proteins and immune cells from the vicinity of the lesion, yet diagnostic sensitivity remains a challenge. Reproducible isolation of lung epithelia and multianalyte extraction have the potential to improve diagnostic sensitivity and provide new information for developing personalized therapeutic approaches. We present the use of a recently developed exclusion-based, solid-phase-extraction technique called SLIDE (Sliding Lid for Immobilized Droplet Extraction) to facilitate analysis of BAL samples. We developed a SLIDE protocol for lung epithelial cell extraction and biomarker staining of patient BALs, testing both EpCAM and Trop2 as capture antigens. We characterized captured cells using TTF1 and p40 as immunostaining biomarkers of adenocarcinoma and squamous cell carcinoma, respectively. We achieved up to 90% (EpCAM) and 84% (Trop2) extraction efficiency of representative tumor cell lines. We then used the platform to process two patient BAL samples in parallel within the same sample plate to demonstrate feasibility and observed that Trop2-based extraction potentially extracts more target cells than EpCAM-based extraction.
Keywords: lab on a chip, microtechnology, lung cancer, bronchoalveolar lavage, solid-phase extraction, clinical automation
Introduction
Lung cancer is the number one cause of cancer-related deaths in the United States.1 Improved early diagnosis and ability to stratify patients and match them to effective treatment options is critical to improving patient outcomes. Currently lung cancer is identified using a computed tomography scan, and specific diagnosis is made via biopsy and histology. One type of biopsy, percutaneous needle biopsy, provides precise sampling of a tumor for histology to differentiate between the various subtypes of lung cancer. However, percutaneous lung biopsies are associated with significant risk to the patient, with some studies finding that patients have an 8% to 35% chance of pneumothorax (collapsed lung).2–5 Bronchoscopy methods, such as the bronchoalveolar lavage (BAL), are considered to be safer than percutaneous techniques because they avoid puncturing the pleural membrane, but they are limited by sensitivity.6 During a BAL, saline is instilled in a region of the lung where the lesion is and is collected back, providing cells, proteins, and lipids (Fig. 1A). The BAL can sometimes yield a diagnosis. However, there are multiple challenges to successful diagnosis via BAL.
Figure 1.
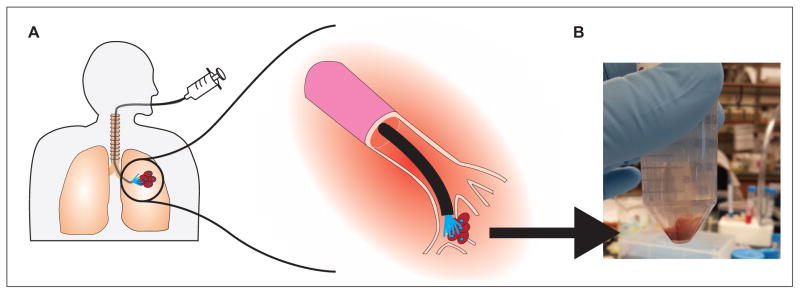
Bronchoalveolar lavage sampling (BAL) is used to sample lung lesions in a less invasive manner than traditional percutaneous needle biopsy method. (A) A bronchoscope is guided down the bronchial tree, where it is used to wash the lesion site with saline. The saline is collected and its contents are analyzed. (B) BAL sample after centrifugation in a 50 mL conical tube. BAL samples are very heterogenous, containing cells, proteins, and extracellular matrix, making it difficult to process and analyze.
First, it is often difficult to sample small lesions in the periphery of the lung using normal bronchoscopes because of their size and difficulty of accurate navigation to the lesion site. Thus, smaller bronchoscopes,7 endobronchial ultrasound, and more advanced navigation procedures such as electromagnetic navigation are now being used.8 Although it is safer than percutaneous biopsies, the BAL sampling procedure is rarely used for accurate diagnosis of cancer. However, if performed immediately after the regularly scheduled bronchoscopic sampling, it may be able to provide a significant amount of additional tissue (e.g., normal epithelium, stroma, and immune cells) than typically acquired via BAL (Fig. 1B). This additional tissue may be important in lung cancer studies. Lastly, we currently lack robust approaches to examine these cells and other factors within the saline wash solution that results from a BAL.
Recent advances in exclusion-based sample preparation (ESP) could allow more information to be obtained from a BAL sample. ESP extraction makes use of paramagnetic particles (PMPs) in a different way than traditional PMP-based solid-phase extraction (SPE) assays. In traditional PMP-based SPE, PMPs are coupled with magnetic force to hold a sample fraction in place while the remaining fluid is removed or washed away, but with ESP, it is the analyte that is removed from the bulk fluid. This allows highly efficient and gentle sample processing as well as multianalytical extraction from a single sample.9,10 Here we use one embodiment of ESP, called Sliding Lid for Immobilized Droplet Extraction (SLIDE), to leverage these advantages for the processing of BAL samples from lung cancer patients.
The SLIDE multianalyte extraction platform was specifically designed to facilitate automation.11 The SLIDE platform is applied here to allow parallel extraction of lung epithelial cells from patient BAL samples. Broader, more efficient processing using this platform can allow more clinically useful information to be gleaned from the less invasive BAL. Further, the method helps to address current challenges in downstream histological analysis given it is well-suited for cell isolation from complex samples.
Methods
Device Fabrication
A custom extraction plate was fabricated from cyclic olefin copolymer resin using hot embossing techniques described by Young et al.12 An aluminum mold was made using an automated CNC mill.13 The plate is designed with a 270 μL input well, 60 μL wash well, and 40 μL output well. Given the SLIDE automation has standard well-plate geometries preprogrammed into its software, the dimensions of the extraction plate were designed to resemble that of a 96-well plate.
Gilson Pipetmax Custom Automation
The Gilson Pipetmax was fitted with a special pipette head that houses neodymium magnets (K&J Magnetics). The magnets can be moved up and down by giving the robot the commands to aspirate or dispense. The magnets are put in the lowered position to attract magnetic beads to the pipette head to remove the beads from a sample well. The magnets are raised to allow a separate magnet below the sample plate to re-collect the beads into a desired sample well downstream (Fig. 2B). The operation of this specialized pipette head was programmed using the TRILUTION micro Protocol Builder software provided by Gilson Inc.
Figure 2.
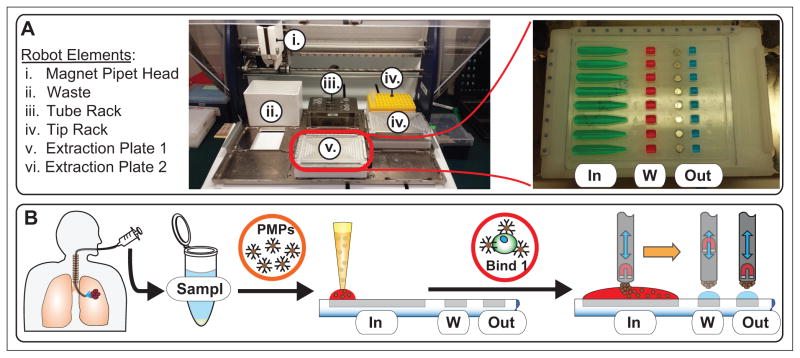
SLIDE platform overview. (A) The robot deck elements include a waste bin, a tube rack for samples and buffers, a tip rack, and two extraction plates. Each extraction plate consists of an input (In), a wash (W), and an output well (Out). (B) Samples are extracted via bronchoalveolar lavage and preprocessed with buffers and paramagnetic particles (PMPs). Samples are then placed in the tube rack, where the robot protocol distributes them to an input well where they are allowed to bind for 30 min at room temperature. Afterward, the modified magnetic pipet head is moved into position at the input well with its magnets in the bottom position. The magnet head is then moved slowly along the length of the well, extracting PMPs along with any cells bound to them. After collecting PMPs from the input, the magnet head moves them to the wash well, which has magnets embedded just below it in the plate platform. The magnets in the modified pipet head are then raised into the top position, allowing the PMPs to be pulled down into the well by the now stronger magnetic force of the embedded magnets. The plate is slid until the output well is positioned above the embedded magnets and the wash well is briefly mixed. Next, the magnets in the pipette head are lowered back to the bottom position, and the magnet head is positioned at the wash well to collect the PMPs, which are then moved to the output well and mixed in the same fashion as previously used for the wash well. Further details of SLIDE processing are contained in ref. 11. Downstream processing is performed manually off chip.
SLIDE Protocol
The protocol consists of four steps: 1) loading of the sample, 2) binding of PMPs to target analytes, 3) extraction of target analytes, and 4) washing of the extracted sample. Briefly, all samples were prepared in a solution of 1× phosphate-buffered saline (PBS), 1% bovine serum albumin (BSA), 1 mM EDTA, and PBS with Tween 20 (PBST) to bring the sample to a volume of 300 μL. Immediately prior to processing, PMPs are added to the sample tubes and placed in the tube rack of the robot for SLIDE processing (Fig. 2A). The sample is incubated in the input well with the beads for 30 min. The magnetic head is then used to transfer the cells to the wash well. The wash well is pipetted to mix and resuspend the sample, allowing cells that were carried over nonspecifically to fall to the bottom of the well. The bound cells are then transferred to the output well, where the sample can be collected for downstream analysis. The protocol takes approximately 60 min to complete. A step-by-step illustration of the protocol is provided in Figure 2B.
PMP Preparation
PMPs (ThermoFisher, 11205D) were washed with PBST and coated in a 1.5 μg/mL solution of capture antibody (Biotinylation Kit, Thermo Fisher D20655; EpCAM, R&D Systems AF960; or Trop2, R&D Systems AF650) for 30 to 60 min at room temperature (RT) in a benchtop shaker, then they were washed again with PBST to remove any unbound antibody.
Cell Line Culture
H358, A549, and H226 human lung cancer and THP-1 human monocyte cell lines were cultured according to ATCC recommendations in RPMI 1640 media with 10% fetal bovine serum (FBS) and 1% penicillin/streptomycin added. When passaging, the adherent lung cancer cells were dissociated with either Trypsin-EDTA (0.05%) or using a cell scraper. Culturing in suspension, the THP-1 cells were simply split as they became confluent.
Cell Line Labeling and Extraction
For extraction studies, human lung cancer cell lines and THP-1 human monocyte cell line were fluorescently labeled with 1.6 μM Calcein AM (ThermoFisher, C3100MP) and CellTracker Red (ThermoFisher, C34552), respectively, at 37 °C for 15 min followed by one wash. It has been shown that epithelial cells make up between 0.05% and 1.5% of the cells recovered in BAL.14 To replicate this cell fraction, 10,000 cells of the cancer cell line population were spiked into a sample of 1,500,000 THP-1 monocytes, giving an experimental epithelial fraction of 0.67%. These cell line samples were then processed according the SLIDE protocol.
Patient Sample Extraction and Target Cell Identification
Primary samples were acquired via BAL, guided by electromagnetic navigation, from two patients suspected to have lung cancer. The bronchoscope was navigated to within millimeters of the lesion site. First, forceps and brush biopsies were collected and sent to pathology and cytology, respectively. A BAL was then performed. During the BAL, four 20 mL aliquots of saline were instilled and collected, each into a syringe; approximately 50% of the instilled aliquot was returned upon collection. Samples were then processed in a similar fashion as detailed in Pollock et al.14 Patient BAL saline samples were twice centrifuged at 500 rcf for 10 min and resuspended with DMEM containing 10% FBS and HEPES buffer. Some BAL samples contained excess mucous and extracellular matrix to the point where pipetting was severely hindered, even with large orifice pipettes. In cases such as these, the sample was washed through a 100 μm cell strainer (Corning Inc., 352360). The samples were then fixed in 4% paraformaldehyde (PFA) for 15 min, washed, and resuspended in 1 mL of PBS + 1% BSA. Each of the two patient samples used in this study contained a large cell/tissue pellet, so a 200 μL aliquot of each was used for extraction, and the remaining sample was saved at 4 °C for later studies. The 200 μL aliquot was then used for processing according the SLIDE protocol. In total, processing, imaging, and analyzing of patient samples takes approximately 24 h to complete.
Calculation of Cell Line Extraction Efficiency
Extraction efficiency is calculated as the number of cells that were identified via microscopy in the input and output wells of the SLIDE extraction plate then dividing the output count by the combined total. Similarly, the ability to extract cell aggregates, as opposed to individual cells, was also quantified by counting the number of aggregates in the input and output wells of the device. Tumor cells were identified after extraction using FIJI image analysis software15 by automated detection and counting of fluorescently labeled cells (see the Cell Line Labeling and Extraction section).
Immunohistochemistry
To verify EpCAM expression, all three cell lines were fixed (15 min in 4% PFA) and fluorescently stained for EpCAM expression. To stain, approximately 25 × 103 cells were incubated at RT for 60 min with a 10 μg/mL solution of primary goat anti-human EpCAM IgG (R&D Systems, AF960) in PBS. The cells were washed and then incubated at RT in a 10 μg/mL Alexa Fluor 568 donkey anti-goat IgG (Thermo Fisher, A-11057) and Hoechst (Molecular Probes, H1399) in PBS solution for 30 min and then washed and imaged on a fluorescent microscope. Trop2 expression in cell lines was verified in the same fashion using a 1:50 concentration of mouse anti-human Trop2 PE (Fisher Scientific, BDB564837).
All extracted cancer cells underwent the same intracellular staining protocol for lung cancer markers, TTF1 and p40. Extracted cancer samples were fixed with 4% PFA and permeabilized with 0.2% Triton X-100. Mouse anti-human TTF1 (Abcam, ab72876) and rabbit anti-human p40 (EMD Millipore, ABS552) were diluted to a 10 μg/mL concentration in PBS. Samples were incubated at 4 °C overnight in the diluted antibody solution. They were then washed and incubated for 60 min at RT in a 10 μg/mL secondary antibody solution of Alexa-fluor 488 anti-mouse (ThermoFisher, A-11029) and Alexa-fluor 568 anti-rabbit (ThermoFisher, A-11036). The samples were washed again and incubated for 30 min at RT in a solution of 1:120 Hoechst and 1:30 of Alexa-fluor 647 anti-human CD45 (Biolegend, HI30). Samples were washed a final time and then fluorescently imaged.
Immunostaining Fluorescence Quantification
Cells were quantified using multichannel fluorescence microscopy followed by image analysis in the JEX image analysis program.16,17 Briefly, for cell lines, images were background subtracted and cell locations were determined by finding fluorescence maxima corresponding to the channel for Hoechst nuclear staining within an image. These coordinates were then used to quantify the intensity in the other imaging channels by measuring the mean fluorescent intensity within a 6.5 μm diameter circular region around each maxima. Given the additional complexity of the patient samples compared with cell lines, a modified JEX workflow was used. Following background subtraction, filtering and thresholding were performed on the nuclear channel to define separable regions for each cell. The integrated fluorescence intensity of these regions was quantified in each fluorescence channel. During analysis of fluorescence, “Ln” refers to the natural logarithm.
Results
Cell lines provided reproducible samples with which to characterize the performance of the SLIDE platform in terms of cell extraction and immunostaining. The protocol was then applied to patient samples to demonstrate feasibility.
Cell Line Extraction
The efficiency and specificity of epithelial cell extraction were quantified using H358, A549, and H226 cell lines, which represent high, moderate, and low/negligible EpCAM-expressing tumor cells, as shown in Figure 3A and 3B. The fluorescence intensity density plots show three distinct populations of cells. Figure 3C shows that even cells with moderate EpCAM expression such as the A549 cells can be robustly extracted. However, a slight drop in capture efficiency is observed when compared with the high-expressing H358 cells. The non–EpCAM-expressing H226 cells had an average extraction efficiency of 2.6% (Fig. 3C). In terms of purity, the samples started with 150 monocytes per tumor cell, whereas afterward, the H358 sample had 0.8 ± 0.14 monocytes per tumor cell and the A549 sample had 0.9 ± 0.3 monocytes per tumor cell. Thus, tumor cell sample purity was enriched by greater than 150-fold.
Figure 3.
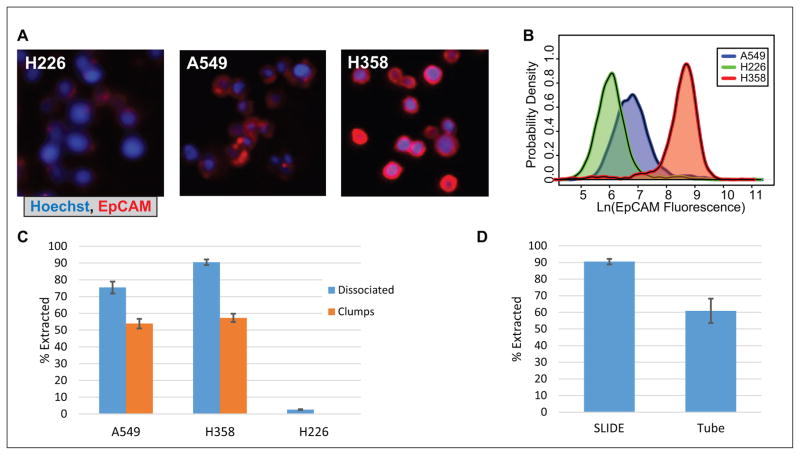
Cell extraction via EpCAM. (A) Representative images (identically treated) of EpCAM expression in H358, A549, and H226 cells. (B) A probability density plot of the natural log of EpCAM fluorescence in each of the three cell lines. (C) Extraction efficiency of cell lines using EpCAM. The EpCAM-positive cell lines (H358 and A549) were spiked into separate fractions of a cell suspension of THP-1 monocytes for a total of eight technical replicates of each condition. Extraction efficiency was compared with that of the H226 cell line, which moderately expresses EpCAM. Extraction efficiency of cell clumps is also shown. (D) Comparison of extraction efficiency of H358 cells in phosphate-buffered saline using SLIDE versus standard tube-based paramagnetic particle extraction.
A common characteristic of BAL samples is the presence of large clumps of target cells that have been sloughed off the tumor during the BAL procedure. To test how the SLIDE platform performs when target cells are part of larger aggregates, the cell lines were scraped off their culture plate rather than being dissociated with Trypsin to promote suspension of cell aggregates or fragments of cell sheets. Figure 3C shows that greater than 50% of the clumps in samples were extracted. Clumps were defined as aggregates of more than two cells.
The extraction efficiency of the SLIDE platform was also compared with manual extraction using the traditional approach PMP-bound cells in tubes, magnetic tube racks, and pipet-based washes. The SLIDE platform isolated H358 cells with ~90% efficiency compared with ~60% efficiency with the traditional method (Fig. 3D).
As part of characterizing capture using Trop2, Trop2 expression of the H226 and H358 cells was measured via fluorescence microscopy. H226 cells exhibited low fluorescence compared with the H358 cells (Fig. 4A, B). In concordance with Trop2 expression, almost 85% of the H358 cells (high Trop2) were captured, whereas less than 2% of the H226 cells (low Trop2) were captured (Fig. 4C).
Figure 4.
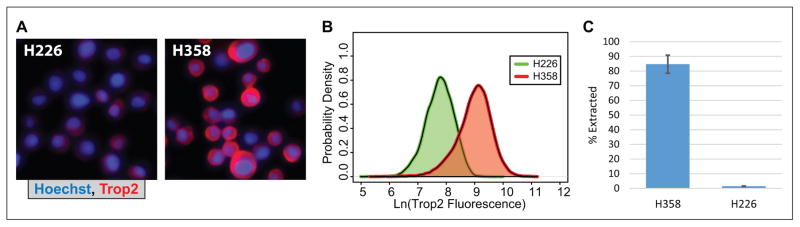
Cell extraction via Trop2. (A) Representative images (identically treated) of Trop2 expression in H358 and H226 cells. (B) A probability density plot of the natural log of Trop2 fluorescence in each of the two cell lines. (C) Extraction efficiency of the cell lines using Trop2.
Biomarker Staining
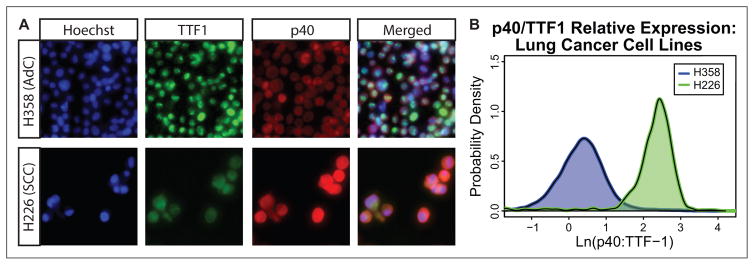
The staining protocol for analyzing p40 and TTF1 expression in patient samples was characterized using the H358 and H226 non–small-cell lung cancer (NSCLC) cell lines. Fluorescent immunohistochemistry (IHC) shows that H358 cells have a higher TTF1:p40 expression ratio than H226 cells, mirroring what is seen clinically (Fig. 5A). This difference is enough that two distinct curves are produced when the expression ratio is illustrated in a density plot (Fig. 5B). Note the localization of TTF1 to the nucleus of the H358 cells compared with the H226 cells.
Figure 5.
Cancer diagnostic cell line controls. (A) Fluorescent immunohistochemistry images of H358 and H226 cells with channels split and merged. (B) The ratio of TTF1 and p40 fluorescent intensity was determined for each cell; the log of the ratio was then taken and plotted on a density curve.
Patient Samples
This study looked at two patient samples acquired via BAL and processed as described above. These samples, referred to as patient 1 and patient 2, were strained using a 100 μm cell strainer to remove unwanted mucous. The samples were then fixed in 4% PFA for 15 min, washed, and resuspended in 1 mL of PBS + 1% BSA. Both samples contained a large cell/tissue pellet, so a 200 μL aliquot of each was used for extraction, and the remaining sample was saved at 4 °C for later studies. Each sample aliquot was split in half, adding 25 μL of anti-EpCAM–coated PMPs to the first half of each sample and adding 25 μL of anti-Trop2–coated PMPs to the remaining 100 μL, respectively. The two samples split into two extraction conditions, resulting in four parallel extractions. The patient samples were processed with the SLIDE and stained just as the cell line controls were in Figure 6.
Figure 6.
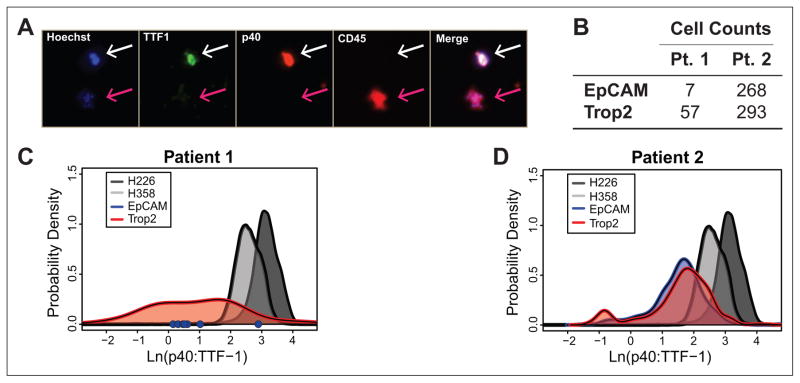
Patient bronchoalveolar lavage cell extraction and staining. (A) An example image from a patient sample that has been processed with the automated SLIDE protocol and stained for TTF1, p40, CD45, and Hoechst. A cancer cell (TTF1 and p40 positive, white arrow) and a clump of leukocytes (CD45 positive, pink arrow) can be seen in this image. (B) Number of target cells extracted from each patient (Pt.) sample aliquot using EpCAM and Trop2 capture antigens. (C, D) Density histograms of log(p40:TTF1) for patient target cells and associated cell line controls. EpCAM capture for Pt. 1 produced too few cells to make a histogram. Instead, the seven cells are plotted as blue dots on the x-axis.
Given the BAL sample contains cell debris and nontarget cells, even after extraction, multiple criteria were used to define “target” lung epithelial cells for subsequent analysis. The target cells must be Hoechst+ and CD45− and generally exhibit cell-like morphology in bright field. For example, a representative cancer cell that is TTF1+, p40+, and CD45− is shown in Figure 6A and indicated using a white arrow. Although the cancer cells were purified, there were still leukocytes that were carried along with the extracted sample fraction as it was moved from the input to the output, much like the clump of CD45+ leukocytes identified by the pink arrow in Figure 6A. There were noticeably more cells captured from patient 2 BAL than patient 1. The CD45− cell counts in Figure 6B show that over 230 more target cells were extracted from patient 2 than patient 1 for both capture antibodies. What is also interesting is that in both cases, Trop2 isolation captured more target cells than EpCAM.
H358 and H226 cells were stained in parallel with the patient samples as lung adenocarcinoma (AdC) and squamous cell carcinoma (SCC) controls, respectively. As before, a density plot was generated from the log of the fluorescence intensity ratio of p40:TTF1. Because so few cells were captured from patient 1 using EpCAM extraction, the individual cell ratios were plotted as blue dots on the x-axis of Figure 6C, concentrating between a 0 to 1 log ratio. The bulk of the cells captured using either technique on patient 2 are centered around a log ratio of ~1.8, but there was a small subpopulation of cells captured using Trop2 that express greater TTF1 than p40 (Fig. 6D).
Given the limited number of patients, conclusions cannot be made regarding a relationship between patient/disease characteristics and results. However, a short summary of the patient information is as follows. Patient 1 was diagnosed with NSCLC, not otherwise specified. Bronchoscopic biopsy was used to verify diagnosis of the 33 mm diameter mass in the right upper lobe. In patient 2, a bronchoscopic biopsy of an 18 mm diameter left upper lobe nodule revealed AdC.
Discussion
The SLIDE platform offers the potential to improve the quality and quantity of diagnostic information that can be obtained from highly complex and heterogenous BAL samples from lung cancer patients. However, the potential benefits of the SLIDE technology for BAL sample analysis have yet to be highlighted. Toward this goal, we first demonstrated and characterized an automated SLIDE protocol for the extraction and staining of cell lines and applied it to analyze BAL samples.
Extraction characterization was performed using conditions and cell lines intended to recapitulate important aspects of BAL samples and the disease. Target cells were spiked into and extracted from suspensions of THP-1 monocytes to mimic the prevalence of monocytes often observed in BAL samples. A typical BAL sample is made up of anywhere from 80% to 95% alveolar macrophages.18,19 We chose to explore the use of two different capture antigens for cell extraction. Although the standard approach for capturing epithelial cells is to target EpCAM, it can be expressed moderately in normal lung epithelia and variably in different types of lung cancer.20,21 Thus, not all target cells are expected to express EpCAM. A potential alternative capture antigen for lung epithelia is Trop2. Much like EpCAM, Trop2 expression has also been shown to be moderate in normal lung tissue while variably present in malignant tissues.22 Another reason these two capture proteins are significant is that they appear to be related to the disease in slightly different manners, as evidenced by their different prognostic values.20,23–26 Thus, EpCAM and Trop2 were chosen for our initial demonstration of the platform because they are not only useful capture targets but also could have significant impact as prognostic biomarkers. We also chose cell lines representative of the two most frequent types of lung cancer. NSCLC makes up more than 80% of lung cancer cases.27 Within that group, AdC and SCC are the most common subtypes. The H358 cell line is an established lung AdC, and H226 is an established lung SCC. Clinically, lung AdC and SCC are differentiated from one another using transcription factors TTF1 and p40 as IHC markers, thus informing our choice of antibodies for staining characterization. AdC overexpresses TTF1, whereas p40 is overexpressed in SCC (Fig. 5).28
We first demonstrated the specificity and efficiency of the SLIDE platform for EpCAM-based tumor cell extraction using cell lines. We performed EpCAM immunostaining and single-cell fluorescence analysis to establish the relative abundance of the capture antigens on each of the target cell lines. Results demonstrate a large difference in the average expression among the different cell lines as well as the single-cell heterogeneity of the expression (Fig. 3A, B). It is expected that expression level should correspond well with extraction efficiency. The platform was able to extract the highly and moderately EpCAM-expressing cell lines with roughly 90% and 75% efficiency, respectively, compared with 2.6% for the H226 cell line that had very low relative EpCAM expression (Fig. 3C). We also quantified the ability to capture cell clumps using EpCAM, given that, in our experience, BAL samples contain many cell aggregates or fragments of epithelial sheets of variable size. Also, the biomarker expression of cells within such aggregates could spur interesting observations; thus, avoiding unnecessary dissociation of aggregates is preferred. Although there was a significant drop as compared with the capture of individual cells, it remains encouraging that more than half of the aggregates were isolated (Fig. 3C). Lastly, we compared the extraction efficiency of the SLIDE-based platform to a standard tube-based PMP protocol in which PMP-bound samples are placed in a tube in a magnetic tube rack to perform washes. Although the standard method is well established and effective in many scenarios, the SLIDE protocol described here exhibited a 20% to 30% increase in cell extraction along with reduced variability from one sample to the next (Fig. 3D). We suspect this is due to the gentler extraction and washing process of the SLIDE technique.11 Overall, the high capture efficiencies for H358 and A549 cells and the correspondence of the EpCAM expression levels and capture efficiencies suggests EpCAM capture is both efficient and specific using the SLIDE platform.
The second capture antigen, Trop2, was also tested using lung cancer cell lines. The platform performed as expected, extracting the high Trop2-expressing H358 cells with almost 85% efficiency and the low Trop2-expressing H226 cells with 1.52% efficiency (Fig. 4). The Trop2 extraction of both the H358 and H226 lung cancer cell lines using Trop2 appeared to be nearly as efficient as EpCAM extraction. Again, we observe a high capture efficiency and a high correspondence between expression levels and capture efficiency, indicating Trop2 capture is also specific and efficient using the SLIDE platform.
After establishing the performance of EpCAM and Trop2 extraction, we demonstrated the ability of the SLIDE platform to process actual patient BAL samples (Fig. 6). In this brief demonstration, we processed two patient samples in parallel on the same day, on the same sample plate, performing both EpCAM and Trop2 capture on each sample. After extraction, we manually stained the cells for Hoechst (nuclear stain), CD45 (a marker of hematopoietic lineage), p40, and TTF1. In parallel, we also stained H358 and H226 cells to account for day-to-day variability in the manual staining procedure and to enable more robust interpretation of results. Both samples showed a noticeably lower p40:TTF1 ratio than the cell line controls. Both samples also displayed a higher number of target cells when extracted using Trop2 compared with EpCAM. Patient 2 had enough cells with both EpCAM and Trop2 capture for creating comparison density histograms of the two capture methods. Overall, the histograms are largely consistent, in keeping with EpCAM and Trop2 capture results for H358 cells during characterization with cell lines. However, there may also be an additional small population of low p40:TTF1 ratio target cells that were extracted using Trop2 that were not extracted using EpCAM. This observation is consistent with clinical observations that individual patients can have cancers made up of both AdC and SCC cells. However, to see if such a trend continues would require a much larger data set and is complicated by the fact that normal epithelium will also most likely be present in samples as well. Lastly, the distributions of the Trop2-captured cells of patient 1 and patient 2 appeared to show different levels of heterogeneity. The distribution for patient 1 was relatively well distributed, whereas for patient 2, peaks were noticeable in the distribution, highlighting the heterogeneity that is likely to be expected from patient to patient. Overall, results demonstrate that the automated platform is able to perform specific and efficient parallel cell extractions from complex and heterogeneous patient BAL samples, enabling isolation of different subsets of target cells for downstream analysis.
Although results of the patient sample are promising, they also highlight topics to address moving forward. One topic is the choice of capture antigens. For example, as part of extraction characterization with cell lines, we observed that EpCAM and Trop2 are not effective at extracting H226 cells; thus, it is likely that additional capture antigens will need to be explored. One such protein is CAIX, which has been found to be expressed in a high percentage of lung SCC.29,30 Capture antigens might also be linked more specifically to treatment options. For instance, therapeutic targets such as epidermal growth factor receptor and fibroblast growth factor receptor mutations are exclusively expressed in lung AdC and SCC, respectively.31 Furthermore, whether such extractions should be performed in parallel on separate fractions of a sample or in sequence on a single sample should also be considered.
Automation allows for more objective and repeatable parallel processing, which allows for a more nuanced interpretation of parallel sample results. However, the SLIDE methodology is also amenable to sequential extraction; hence, the additional extraction plate shown in Figure 2A. Therefore, future development to automate sequential capture could allow different cell populations to be isolated in variable sequence from the same input well, to tease apart the nature and overlap of the populations captured with each antigen. Development of such approaches will need to address the potential for target cells to express multiple capture target antigens and require optimization of the order in which the different capture antibodies are introduced. Currently, sequential capture would require multiple plates; however, alteration of the SLIDE apparatus, such as additional pairs of wash and output wells, could enable sequential capture on a single plate. It is also possible that multiple antibodies either on the same beads or on different beads mixed together can be used to capture multiple populations simultaneously, such as for removal of background cells.
Another topic to address moving forward is the automation of both extraction and downstream readouts. Currently, only the extraction of cells has been automated, yet the platform is also well suited for extraction of other analytes such as proteins and nucleic acids. The platform is also amenable to automation of immunostaining procedures, which could prove important for continued development of the immunostaining analysis initiated here. In the present study, manual immunostaining was used for downstream analysis; however, there was a noticeable increase in the p40:TTF1 ratio in the cell line controls (Figs. 5 and 6). This increase could indicate the presence of significant biological variability in the cell lines or the need for further optimizing the staining protocol to remove day-to-day variability in the manual staining process. Although, it is still possible to perform a relative qualitative comparison of the parallel-processed patient results obtained here; optimization will be needed to make immunostaining and patient comparisons more quantitative to improve confidence in the biological interpretation of results. Moving forward, we plan to sample a normal and diseased portion of each patient. This will provide a more direct and patient-specific normal control for assay endpoints. Further, integration of a staining procedure into the automated protocol could not only aid in optimization but also improve repeatability. Similarly, integration of additional endpoints, such as extraction of soluble factors and nucleic acids, will also strengthen this automated platform for BAL analysis, allowing comparison of concordant metrics for more robust interpretation of results and an increased ability to leverage BAL samples for safer and more robust diagnosis of lung cancer.
In conclusion, BAL is a unique and promising cancer sampling technique but is currently less precise than percutaneous techniques for purposes of diagnosis, in part because of the complexity and heterogeneity of the sample itself. However, being heterogeneous in nature, the sample also provides other analytes such as immune cells, cytokines, and lipids that might provide additional clinically relevant information as well as basic insights into the tumor microenvironment. Here, we have demonstrated a new approach to tumor cell extraction from BAL samples that has the potential to leverage these additional sample components. We characterized the ability of the platform to provide specific and efficient capture of cell lines from monocyte-rich backgrounds and showed feasibility of performing automated parallel processing of patient BALs for downstream immunostaining endpoints. We envision that future iterations of the technology will involve exploration of alternative capture antigens that could potentially be coupled with informative markers of diagnosis and therapy. Likewise, we imagine that further integration of downstream processing will enable more robust analysis from this promising and rich yet complex and heterogeneous type of tumor sample for lung cancer patients.
Acknowledgments
We would like to thank all patients who participated in this study. We are also grateful for the help of the UWCCC Lung Cancer clinical research group.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported via the University of Wisconsin NCI Comprehensive Cancer Center as an Investigator-Initiated Pilot Project and NIH/NCI P30 CA014520 (JJT, JWW, JSF, DJB). Funding support for JML, DJG, and JMS was provided by NIH-NCI 5R01 CA181648, Prostate Cancer Foundation–Movember Challenge Award, and the UWCCC Lung Cancer Research Fund.
Footnotes
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: David J. Guckenberger holds equity in Salus Discovery, LLC and Tasso, Inc. Joshua M. Lang holds equity in Salus Discovery, LLC. David J. Beebe holds equity in Bellbrook Labs LLC, Tasso Inc., Stacks to the Future LLC, Lynx Biosciences LLC, Onexio Biosystems LLC, and Salus Discovery LLC.
References
- 1.Siegel R, Rebecca S, Jiemin M, et al. Cancer Statistics, 2014. CA Cancer J Clin. 2014;64:9–29. doi: 10.3322/caac.21208. [DOI] [PubMed] [Google Scholar]
- 2.Gildea TR, Mazzone PJ, Demet K, et al. Electromagnetic Navigation Diagnostic Bronchoscopy. Am J Respir Crit Care Med. 2006;174:982–989. doi: 10.1164/rccm.200603-344OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Tomiyama N, Yasuhara Y, Nakajima Y, et al. CT-Guided Needle Biopsy of Lung Lesions: A Survey of Severe Complication Based on 9783 Biopsies in Japan. Eur J Radiol. 2006;59:60–64. doi: 10.1016/j.ejrad.2006.02.001. [DOI] [PubMed] [Google Scholar]
- 4.Wang Y, Jiang F, Tan X, et al. CT-Guided Percutaneous Transthoracic Needle Biopsy for Paramediastinal and Nonparamediastinal Lung Lesions: Diagnostic Yield and Complications in 1484 Patients. Medicine. 2016;95:e4460. doi: 10.1097/MD.0000000000004460. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Azrumelashvili T, Mizandari M, Dundua T, et al. Ultrasound and CT Guided Thoracic Biopsy Approaches: Effectiveness and Complications. Georgian Med News. 2016;255:32–39. [PubMed] [Google Scholar]
- 6.Gould MK, Donington J, Lynch WR, et al. Evaluation of Individuals with Pulmonary Nodules: When Is It Lung Cancer? Diagnosis and Management of Lung Cancer, 3rd ed: American College of Chest Physicians Evidence-Based Clinical Practice Guidelines. Chest. 2013;143(5 suppl):e93S–120S. doi: 10.1378/chest.12-2351. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Wang Memoli JS, Nietert PJ, Silvestri GA. Meta-Analysis of Guided Bronchoscopy for the Evaluation of the Pulmonary Nodule. Chest. 2012;142:385–393. doi: 10.1378/chest.11-1764. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Leong S, Ju H, Marshall H, et al. Electromagnetic Navigation Bronchoscopy: A Descriptive Analysis. J Thorac Dis. 2012;4:173–185. doi: 10.3978/j.issn.2072-1439.2012.03.08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Strotman L, Lindsay S, Rachel O, et al. Selective Nucleic Acid Removal via Exclusion (SNARE): Capturing mRNA and DNA from a Single Sample. Anal Chem. 2013;85:9764–9770. doi: 10.1021/ac402162r. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Berry SM, Regehr KJ, Casavant BP, et al. Automated Operation of Immiscible Filtration Assisted by Surface Tension (IFAST) Arrays for Streamlined Analyte Isolation. J Lab Autom. 2013;18:206–211. doi: 10.1177/2211068212462023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Casavant BP, Guckenberger DJ, Beebe DJ, et al. Efficient Sample Preparation from Complex Biological Samples Using a Sliding Lid for Immobilized Droplet Extractions. Anal Chem. 2014;86:6355–6362. doi: 10.1021/ac500574t. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Young EWK, Berthier E, Guckenberger DJ, et al. Rapid Prototyping of Arrayed Microfluidic Systems in Polystyrene for Cell-Based Assays. Anal Chem. 2011;83:1408–1417. doi: 10.1021/ac102897h. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Guckenberger DJ, de Groot TE, Wan AMD, et al. Micromilling: A Method for Ultra-Rapid Prototyping of Plastic Microfluidic Devices. Lab Chip. 2015;15:2364–2378. doi: 10.1039/c5lc00234f. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Pollock K, Albares L, Wendt C, et al. Isolation of Fibroblasts and Epithelial Cells in Bronchoalveolar Lavage (BAL) Exp Lung Res. 2013;39:146–154. doi: 10.3109/01902148.2013.781720. [DOI] [PubMed] [Google Scholar]
- 15.Schindelin J, Arganda-Carreras I, Frise E, et al. Fiji: An Open-Source Platform for Biological-Image Analysis. Nat Methods. 2012;9:676–682. doi: 10.1038/nmeth.2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Warrick JW, Timm A, Swick A, et al. Tools for Single-Cell Kinetic Analysis of Virus-Host Interactions. PLoS One. 2016;11:e0145081. doi: 10.1371/journal.pone.0145081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. [accessed Jan 13, 2017];JEX Image Databasing and Analysis Software. www.github.com/jaywarrick/JEX.
- 18.Pollock K, Albares L, Wendt C, et al. Isolation of Fibroblasts and Epithelial Cells in Bronchoalveolar Lavage (BAL) Exp Lung Res. 2013;39:146–154. doi: 10.3109/01902148.2013.781720. [DOI] [PubMed] [Google Scholar]
- 19.van Haarst JM, Hoogsteden HC, de Wit HJ, et al. Dendritic Cells and Their Precursors Isolated from Human Bronchoalveolar Lavage: Immunocytologic and Functional Properties. Am J Respir Cell Mol Biol. 1994;11:344–350. doi: 10.1165/ajrcmb.11.3.8086170. [DOI] [PubMed] [Google Scholar]
- 20.Pak MG, Shin DH, Lee CH, et al. Significance of EpCAM and TROP2 Expression in Non-Small Cell Lung Cancer. World J Surg Oncol. 2012;10:53. doi: 10.1186/1477-7819-10-53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. [accessed Nov 16, 2016];Tissue Expression of EPCAM—Summary: The Human Protein Atlas. http://www.proteinatlas.org/ENSG00000119888-EPCAM/tissue.
- 22. [accessed Nov 16, 2016];Tissue Expression of TACSTD2—Summary: The Human Protein Atlas. http://www.proteinatlas.org/ENSG00000184292-TACSTD2/tissue.
- 23.Li Z, Jiang X, Zhang W. TROP2 Overexpression Promotes Proliferation and Invasion of Lung Adenocarcinoma Cells. Biochem Biophys Res Commun. 2016;470:197–204. doi: 10.1016/j.bbrc.2016.01.032. [DOI] [PubMed] [Google Scholar]
- 24.Lin JC, Wu YY, Wu JY, et al. TROP2 Is Epigenetically Inactivated and Modulates IGF-1R Signalling in Lung Adenocarcinoma. EMBO Mol Med. 2012;4:472–485. doi: 10.1002/emmm.201200222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Shvartsur A, Bonavida B. Trop2 and Its Overexpression in Cancers: Regulation and Clinical/Therapeutic Implications. Genes Cancer. 2015;6(3–4):84–105. doi: 10.18632/genesandcancer.40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Zhou N, Wang H, Liu H, et al. MTA1-Upregulated EpCAM Is Associated with Metastatic Behaviors and Poor Prognosis in Lung Cancer. J Exp Clin Cancer Res. 2015;34:157. doi: 10.1186/s13046-015-0263-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Planque C, Kulasingam V, Smith CR, et al. Identification of Five Candidate Lung Cancer Biomarkers by Proteomics Analysis of Conditioned Media of Four Lung Cancer Cell Lines. Mol Cell Proteomics. 2009;8:2746–2758. doi: 10.1074/mcp.M900134-MCP200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Nobre AR, André A, Fernando S. p40: A p63 Isoform Useful for Lung Cancer Diagnosis—A Review of the Physiological and Pathological Role of p63. Acta Cytol. 2013;57:1–8. doi: 10.1159/000345245. [DOI] [PubMed] [Google Scholar]
- 29.Ramsey ML, Yuh BJ, Johnson MT, et al. Carbonic Anhydrase IX Is Expressed in Mesothelioma and Metastatic Clear Cell Renal Cell Carcinoma of the Lung. Virchows Arch. 2012;460:89–93. doi: 10.1007/s00428-011-1178-7. [DOI] [PubMed] [Google Scholar]
- 30.McDonald PC, Winum JY, Supuran CT, et al. Recent Developments in Targeting Carbonic Anhydrase IX for Cancer Therapeutics. Oncotarget. 2012;3:84–97. doi: 10.18632/oncotarget.422. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Bishop JA, Julie TF, Westra WH, et al. p40 (ΔNp63) Is Superior to p63 for the Diagnosis of Pulmonary Squamous Cell Carcinoma. Mod Pathol. 2011;25:405–415. doi: 10.1038/modpathol.2011.173. [DOI] [PubMed] [Google Scholar]