Abstract
Currently, there are 20,197 human protein-coding genes in the most expertly curated database (UniProtKB/Swiss-Pro). Big efforts have been made by the international consortium, the Chromosome-Centric Human Proteome Project (C-HPP) and independent researchers, to map human proteome. In brief, anno 2017 the human proteome was outlined. The male factor contributes to 50% of infertility in couples. However, there are limited human spermatozoa proteomic studies. Firstly, the development of the mapping of the human spermatozoa was analyzed. The human spermatozoa have been used as a model for missing proteins. It has been shown that human spermatozoa are excellent sources for finding missing proteins. Y chromosome proteome mapping is led by Iran. However, it seems that it is extremely challenging to map the human spermatozoa Y chromosome proteins based on current mass spectrometry-based proteomics technology. Post-translation modifications (PTMs) of human spermatozoa proteome are the most unexplored area and currently the exact role of PTMs in male infertility is unknown. Additionally, the clinical human spermatozoa proteomic analysis, anno 2017 was done in this study.
Keywords: Human, Proteome, Proteomics, Spermatozoa, Y chromosome
Introduction
Anno 2017; from proteome to proteoform:
There was a revelation obtained from human genome project results showing a smaller number of analyzed genes that was approximately 20,300 rather than ∼100,000 (1). This finding determined that the complication in our biological system can be due to variation at the level of protein rather than a large number of distinct genes (2). The diversity among well related protein molecules which are chemically different can be driven from variation within populaces, cell and tissue and their subcellular localization. The intricacy of allelic variations, alternative splicing of RNA transcripts and post-translational modifications can be due to DNA, RNA and protein structures and levels. These machineries build distinct protein molecules which are able to affect cell signaling, gene regulation and activation of protein complexes. Although the complexity of protein structures was first identified by using two-dimensional gel electrophoresis, novel proteomic technologies have been demonstrated to develop the decisive architectures of proteome (3, 4). The word proteome was first introduced in 1995 by Wilkins. The term has become quite popular due to its simple and easy definition (5). The global protein analysis idea somehow is not new. It was first appeared in late 1970s (6). Nowadays, this scientific field is called proteomics, which is the study of the large-scale number of proteins expressed in a cell, tissue or biological fluids (7, 8). In general, proteomics is divided in five central pillars: mass spectrometry-based proteomics, array-based proteomics, structural proteomics, clinical proteomics and informatics (9).
Mass spectrometry has developed as a key platform for proteomic analyses, with two contrasting approaches which are bottom-up and top-down proteomics. In the bottom-up approach, typically the enzyme trypsin is used to digest the proteome to small peptides. The complex peptide mixture is separated and analyzed by liquid chromatography tandem-mass spectrometry (LC-MS/MS) (10, 11). In top-down proteomics, the proteins of the proteome in an unbroken fashion are directly separated and fragmented by LC-MS/MS. The top-down approach has been reported to provide the most useful data for accurate identification and characterization of molecular composition (12, 13). On the other hand, executing bottom-up approach can be difficult due to the complexity of produced results and limitation of technical knowledge.
MS-based proteomics is in constant movement and new terms (such as proteoform), new instrumentation and software are under development which help to identify more proteins (14). The newest term, proteoform is defined as all different molecular forms in which the protein product of a single gene can be found, encompassing all forms of genetic variation, alternative splicing of RNA transcripts, and post-translational modifications (PTMs) (15, 16).
First maps of the human proteome:
It has been indicated that mass spectrometry was able to develop the analysis of human proteome in a way which is comparable to influence of next-generation sequencing on genomics and analysis of transcriptomics (7, 17, 18). The international consortium for the Chromosome-Centric Human Proteome Project (C-HPP) was established in 2011. The propose of C-HHP is to identify and characterize each of the 20,300 human protein coding genes including single amino acid polymorphisms (SAPs), splice variant isoforms and post-translational modifications (PTMs) (19–21). The guidelines for C-HPP are set, and each of international teams has selected a specific chromosome (22, 23).
More than 150 posters and papers have been published by C-HPP since 2013 (www.thehpp.org). Using combined MS-based proteomics and antibody technology followed by bioinformatics, less than 14,000 protein-coding genes have been identified (24).
Based on two independent studies from HUPO organization, draft maps of human proteome were published in 2014 using MS-based proteomics platform. They were able to identify 17,294 and 19,629 protein encoding genes, respectively (25, 26). The draft maps of the human proteomics were in larger scale than the multinational Human Proteome Project effort. In their studies, Kim et al. and Wilhelm et al. have used different healthy biological samples from human including testis. However, none of them used the human spermatozoa to map the human proteome. Finalizing the human proteome is more challenging than genome. It will have much more greater impact on understanding human diseases. The first analysis of the portray of human proteome clearly shows this challenge (27). Ezkurdia et al. analyzed the data from the human proteome and showed that Kim et al. and Wilhelm et al. have overestimated their identification of proteins in case of trans-membrane proteins. Furthermore, they showed both studies have an abundance of poor spectra, low-scoring peptide-spectrum matches and incorrectly identified proteins (28). Additionally, a reanalyzation of Kim et al.’s study by HUPO team showed that they could identify only 11,000 not 17,000 genes as they hypothesized (29). However, despite these critical analyses of the first mapping of the human proteome, it does not undervalue these studies, since they have more than 500 citations two years after publication.
MS-based proteomics technology has improved during years and it has become a useful biomedical research instrument. MS-based technology can help in diagnosis of disease-related mutations and biomarkers. However, lack of genomic data in infertility or other diseases associated with proteomics data is still an undetermined issue.
The main goal of the medical proteomics is to find disease biomarkers including proteins or peptides that are specific to a disease. This goal has not been reached yet. However, Human Proteome Project and other independent studies are inspiring concerning medical proteomics (30–35).
The size of the human spermatozoa proteome:
It has been reported that the recognition and evaluation of expressed proteins in cells and tissues could develop a better approach to understand the cell dynamics and tissue purposes in a variety of fields (36). Human spermatozoa can provide optimal cells to be investigated from a proteomic perspective because they do not represent physiologically active transcription and translation. As such, proteomics has the potential to transform our understanding of the workings of the mature cell. Such a leap in knowledge is necessary as spermatozoa are very specialized cells (37).
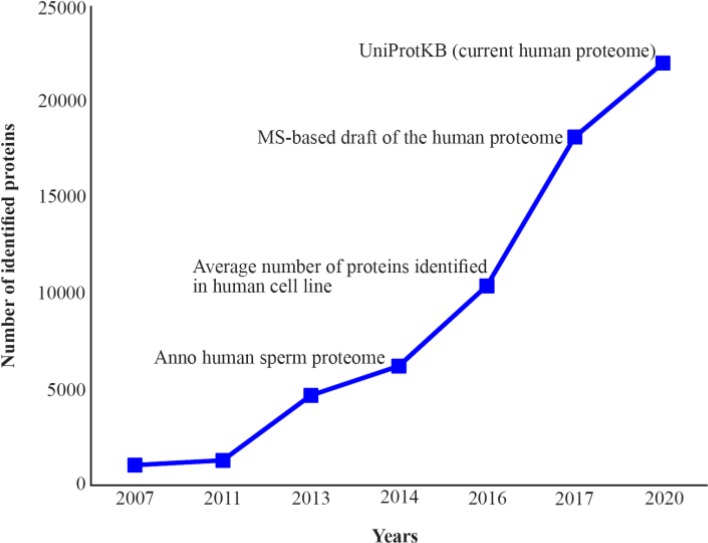
After a moderate initiation for the proteome analysis of the human spermatozoa, a quick development has been created in the last couples of years (Figure 1) (38–41).
Figure 1.
The human spermatozoa proteome mapping development compared to the human cell line proteome and the current known human proteome
An earlier attempt to map the human spermatozoa proteome was published in 2005 (42). The authors claimed that they have identified over 1,700 human spermatozoa proteins, however, no specified list of correlated proteins has been declared by them (42). In 2007, the first large scale analysis of the human spermatozoa proteome was published with a protein list that identified 1,053 proteins (38). The first attempt to organize and catalogue the human spermatozoa proteome was done in 2011 by the authors (39). The collection of 1,300 proteins was reported. Following development of MS-based proteomics technology, the human spermatozoa proteome was further subjected to proteomic analysis. Wang et al. identified 4,675 human spermatozoa proteins, of which 227 were testis-specific (40). The latest investigation that catalogued the human spermatozoa proteome was done by Amaral et al. (41). They were able to collect 6,198 unique human spermatozoa proteins. Finally, the question is: how big is the size of the human spermatozoa proteome?
For a long, due to transcriptional inactivity of sperm cells, human spermatozoa proteome is believed to be restricted to a couple of thousand proteins (37, 39). By looking at the development of the human spermatozoa proteome analysis during the last years, it seems that the depth of understanding in human spermatozoa proteome has not been reached yet (Figure 1). It is reported that MS-based proteomics technology is not limited by sensitivity. It is rather limited by dynamic range and effective sequencing speed (43). Furthermore, it has been shown that human cell line proteome analysis by the MS-based technology has reached saturation level for quantification and identification in 2011 (Average number of protein in the human cell line, figure 1) although an improved resolution and sequencing speed of mass analyzer was achieved (44).
MS-based proteomics technology has further developed leading to have deeper view of the human proteome. It seems that by the development of MS-based proteomics technology and software, it becomes easier to analyze the current human proteome (MS-based draft of the human proteome and UniProtKB current human proteome, figure 1) (25, 26, 45). The current size of the human proteome is based on protein-coding genes (21,931 proteins), while the isoforms, PTM or alternative splicing are not included in the mentioned version. The Proteomics DB database, which is the outcome of the first draft of human proteome has 86,771 isoforms (46). Another database, which solely developed for the mass spectrometry (MS) identification of human proteins, is neXtProt. This database demonstrates proteins existence, their related isoforms, post-translational modifications as well as subcellular localization (47). Almost 400 proteins are found in the neXtProt by searching the term of “spermatozoa”.
One of the challenges of the proteomics study is to analyze the proteome quantitatively. Estimating protein concentration in a cell is significant. Understanding cell biology depends on the knowledge of the cellular protein quantities. For example, system biology approach which describes behavior of a cell depends on the knowledge of protein copy number per cell whereas estimation of absolute protein concentration of human protein is technically challenging and limited (48, 49). It is believed that no study has used quantitative tandem MS strategy to estimate the cellular concentration of the human spermatozoa proteome yet.
Several researchers in a variety of studies have used the bottom-up approach for proteomic analysis of the human spermatozoa. Peptide Atlas has been reported as a database that is under investigation for gathering peptides in MS-based proteomics of bottom-up approaches (50). It explains the genome via peptide identification of proteins. One of the interesting points of Peptide Atlas is collection of proteomic analysis for the male reproductive system; however, there are no studies regarding the human spermatozoa by now, to the best of our knowledge.
The analysis of human spermatozoa proteome is becoming more proteomic. However, the most recent proteomics technology approach which is top-down proteomics has not been well studied yet. It would be interesting to investigate whether unknown proteins or missing proteins (see below) of the human spermatozoa proteome can be identified or not using top-down proteomics.
Missing proteins:
The human gene product variety should not be miscalculated according to the alternative mRNA splicing, post-translational modifications (PTMs) and polymorphism of single amino acid. To the best of our knowledge, no body exactly knows how many proteins are expressed in the ∼230 cell types that build our body while between 1 to 2 million proteoforms have been recommended (39, 51).
Among ∼20,197 human protein-coding genes, about 3834 (10%) of them lack any experimental evidence at the protein level. These proteins are named “missing proteins” (51). There are many reasons why these proteins lack an evidence of experimental protein expression in the data bases e.g. UniProtKB or neXtProt. Lack of the evidence for protein expression level has been reported to be due to: 1) Protein specificity for each organ; 2) Expression of specified proteins in the early development stage, particularly embryonic and fetal; 3) Being below our current limits for detection of protein expression which itself can be due to short rates of synthesis or rapid degradation; and 4) Expression of proteins under stress condition (23, 52, 53).
A great number of proteins are identified by the first draft of the human proteome using MS-based proteomics technology. However, it is clear that missing proteins can be considered as one of the main challenges for the future work. MS-based proteomics is not able to identify all proteins, e.g. Neuroglobin, a protein that needs to be identified by in-depth research lab, and cannot be detected by current MS-based proteomics technology (54). That can be due, at least in part, to the detection limits of mass spectrometry.
The ProteomicsDB database does include a section, “Adopt a protein”, which calls protein experts in the world to fill the gap in the human proteome missing proteins (46).
The C-HPP has provided a scheme to investigate the information associated with the missing proteins (55, 56). One of the exciting studies, published in the annual report of C-HPP, was the use of human spermatozoa as a model for detecting missing proteins (56). The argumentation to use the human spermatozoa for detecting the missing proteins is that the human spermatozoa proteins are cell specific. Surprisingly, the authors were able to identify 89 of the missing proteins in the human spermatozoa. They showed that the genes of these proteins were located on 20 different chromosomes. The chromosomes that do not carry the genes encoding any of these proteins were 21, 22 and Y (57).
Y-chromosome:
In the Chromosome-centric Human Proteome Project (C-HPP), the proteome mapping of the human Y chromosome is considered to be conducted by Iran (58). The human Y chromosome is a type of sex chromosome that exists in male mammalian species basically. The human Y chromosome is about three times smaller than the human X chromosome, and its male sex-determining function is exclusively located on the short arm. Male sex determination has been reported as an outcome of gonadal sex purpose during embryonic development. In the presence of the human Y chromosome, the embryonic gonads turn into testes, however, in the absence of human Y chromosome, the gonads develop into ovaries (59, 60). It has been demonstrated that deletions or mutations, particularly in the long arm of human Y chromosome, may lead to male infertility, and also influence the reproductive performances of related sons (61–63). In relation to C-HPP, Jangravi demonstrated a current revise of the malespecific region (MSY) in the human Y chromosome protein-encoding genes. The human Y chromosome proteins were analyzed corresponding to each disease. They also indicated protein-protein interactions and post-translational modifications of protein-coding genes in the MSY (64). Most recently, Rengaraj et al. analyzed the human Y chromosome-encoded proteins (66 Y chromosome-encoded proteins retrieved from NCBI database), their related pathways, and their related interactions using bioinformatics tools (65). It is very important to understand the UniProtKB database that shows only 47 human Y chromosome-encoded proteins which have evidence on protein level (http://www.uniprot.org/docs/humchry). Regarding neXtProt and PeptideAtlas databases, the retrieved human Y chromosome-encoded proteins showed 44 and 40, respectively (65). The MS-based proteomics draft of the human proteome conducted by Wilhelm et al. showed a 57% coverage of the human Y chromosome-encoded proteins (46).
The protein pathways analysis of the 66 proteins encoded by human Y chromosome demonstrated 4 major pathways, including cell signaling pathways, receptor signaling pathways, cellular processes, and metabolic pathways (65).
An analysis of the current catalogue of the human spermatozoa proteome for human Y chromosome-encoded proteins retrieved from UniProtKB database determined the following encoded Y chromosome proteins as detected by current MS-based proteomics technology including ATP-dependent RNA helicase (O15523), Heat shock transcription factor, Y-linked (Q96LI6), Protocadherin-11 Y-linked precursor (Q9BZA8), 40S ribosomal protein S4, Y isoform 1 (P22090), Testisspecific Y-encoded protein 1 (Q01534) and ubiquitin carboxyl-terminal hydrolase FAF-Y (O00507). However, these numbers of proteins cover only ∼10% of the current human Y chromosome-encoded proteins.
The protein abundance can be roughly estimated by MS using the number of identified unique peptides of a protein (66). The six identified human Y chromosome-encoded proteins in the human spermatozoa proteome are identified with the following number of unique peptides: O15523 (4), Q96LI6 (4), P22090 (4), Q01534 (2), O00507 (3), and Q9BZA8 (Not reported). It is clear that human Y chromosome-encoded proteins are not highly expressed proteins even in the human spermatozoa cell based on the number of the unique peptides identified from the mentioned proteins.
These six human Y chromosome-encoded proteins in the human spermatozoa were analyzed for protein-protein interactions using STRING database (Figure 2) showing that in a medium confidence search (score 0.4), there is a strong interaction between O1523 (DDX3Y) and P22090 (RPS4Y1) in the human spermatozoa cell (67).
Figure 2.
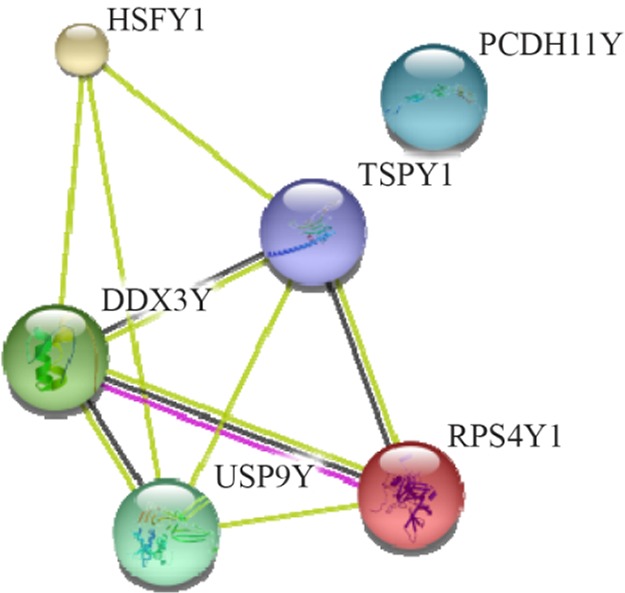
Interactions among the six human Y chromosome-encoded proteins in the human spermatozoa cell. A medium confidence view (score 0.4) of the interaction among human Y chromosome-encoded proteins was prepared using the STRING database program. Thicker lines represent stronger associations, and thinner lines represent medium associations. The UniportKB associate code and corresponding gene code are O1523 (DDX3Y), P22090 (RPS4Y1), Q96LI6 (HSFY1), Q01534 (TSPY1), O00507 (USP9Y), and Q9BZA8 (PCDH11Y)
To determine the cellular pathways involving these six human Y chromosome-encoded proteins in the human spermatozoa, the Reactome pathway knowledge base database was searched (68). The six human Y chromosome-encoded proteins activate the following pathways in the human spermatozoa cell including gene expression, metabolism of proteins and signal transduction.
Post-translational modifications in the human spermatozoa:
Spermatozoa can be considered as an ideal model for investigation of post-translational modifications since the transcriptional and translational activities are almost inactive (69, 70). It has been reported that spermatozoa functions can be mostly regulated at the protein level while its post-translational modifications (PTMs) are particularly vital. Regulating spermatozoa functions such as maturation and acquisition of fertilizing potential can be affected by PTMs on existing proteins (71).
It is very complicated to understand which PTMs are the most frequent inside the cells. Recently, UniProt database collected 307 diverse types of PTM (http://www.uniprot.org/docs/ptmlist). PTM leads to a change in total mass of the related protein and can alter the residue nature. Though the current proteome-wide statistics analysis of UniProtKB database for experimental PTMs shows the following PTMs are dominated by Phosphorylation, Acetylation, N-linked glycosylation, Amidation and Hydroxylation, the putative PTMs are first dominated by N-linked glycosylation (72).
There is a significant challenge and a required expertise in the large-scale MS-based PTMs proteomics analysis compared to conventional MS-based proteome analysis is needed.
Therefore, there is only a handful of large-scale MS-based proteomics studies of the human spermatozoa PTMs. These studies have focused on the phopshoproteome, N-linked glycoproteome, acetylproteome and ubiquitination (71, 73–75).
The full potential of large-scale MS-based proteomics technology in order to better understand PTMs functions in the human spermatozoa is not well defined. To the best of our knowledge, the only study that used high-throughput technology was carried out by Ficarro et al. They showed that the phosphorylation plays an important role and specifically valosin-containing protein was phosphorylated during capacitation. However, phosphorylated sites of this protein were not identified (74).
This paper was not about the PTM function in the human spermatoza. The purpose was just to attract attention to PTM study regarding the spermatozoa since there are limited studies avialabe related to this subject.
Proteomic analysis of spermatozoa:
Semen analysis screening information might indicate male infertility factor while, not reflecting reproductive potential constantly (76). Therefore, screen of sperm DNA damage and oxidative stress can be recommended to forecast reproduction (77–82).
The global protein analysis/proteomics has been investigated for more than 40 years. However, only during the last 10 years, the studies regarding male infertility and spermatozoa have gained momentum (Table 1).
Table 1.
Proteomics studies of the human spermatozoa
| Proteomics methodologies | Type of patients/studies | Outcome | Year (Reference) |
|---|---|---|---|
| 2DE, in-gel digestion, LC-MS/MS and MALDI-TOF MS | To compare the sperm protein expression profile (proteome map) from a patient who experienced failed fertilization at IVF with fertile controls | First proteome comparison of different qualities of sperm. Identification of 4 proteins differentially expressed | 2004 (83) |
| 2DE, in-gel digestion MALDI-TOF MS | To compare sperm protein expression profiles in asthenozoospermic patients with that of normozoospermic donors | Identification of 10 differentially expressed proteins | 2007 (84) |
| 2DE, in-gel digestion MALDI-TOF MS | A comparison of asthenozoospermic sperm proteins to the fertile controls | Identification of 17 differentially expressed proteins | 2008 (85) |
| 2DE, in-gel digestion MALDI-TOF MS | Asthenozoospermic sperm of patients were compared to fertile controls | Identification of 12 differentially phosphorylated proteins | 2009 (86) |
| 2DE-DIGE, in-gel digestion LC-MS/MS and MALDI-TOF MS | The spermatic proteomic profiles of patients, with a complete failure of fertilization and no spermatozoa bound to the zona pellucida, compared to controls | Identification of 12 differentially expressed proteins | 2009 (87) |
| 2DE-DIGE, in-gel digestion MALDI-TOF/TOF-MS | To investigate the differences in protein expression between human round-headed and normal spermatozoa | Identification of 35 proteins differentially in round-headed sperm compared with normal sperm | 2009 (88) |
| 2DE, in-gel digestion MALDI-TOF MS and MS/MS | Protein profile of capacitated versus ejaculated human sperm | Identification of 29 proteins differentially expressed in capacitated sperm, swim-up selected capacitated sperm and ejaculated sperm | 2009 (89) |
| 2DE-DIGE, in-gel digestion MALDI-TOF/TOF-MS | Identification of diabetes- and obesity-associated proteomic changes in human spermatozoa | Identification of seven and nine proteins associated with type-1 diabetes and obesity, respectively | 2009 (90) |
| 2DE, in-gel digestion MALDI-MS/MS | To understand the molecular basis of sperm motility using asthenozoospermic sperm versus controls | Identification of eight proteins showing different abundance | 2010 (91) |
| 2DE-DIGE, in-gel digestion MALDI-TOF/TOF MS and LC-MS | The sperm protein profile was compared between fertile and oligoasthenozoospermic men | Identification of four proteins showing different abundance | 2011 (92) |
| 2DE-DIGE, in gel digestion MALDI-TOF/TOF MS and LC-MS/MS | To unveil disease-associated proteomic changes potentially affecting male fertility, the proteomes of sperm cells from type-1 diabetic, type-2 diabetic, non-diabetic obese and clinically healthy individuals, comparatively analyzed | Identification of 12 (type-1 diabetic), 71 (type-2 diabetic) and 13 (nondiabetic obese) proteins showing different abundance. Eppin protein complex components increased in sperm from the three groups of patients | 2011 (93) |
| 2DE, in-gel digestion MALDI-TOF MS | To screen and investigate the differentially expressed proteins in the sperm of infertile patients, whose sperm parameters met the WHO guidelines | Identification of 24 proteins showing different abundance | 2012 (94) |
| Washing IMAC with a phosphoprotein enrichment kit, in-solution digestion LC-MS | To investigates the phosphoproteins involved in sperm motility in an attempt to identify the key pathways regulating sperm motility and likely to be altered in spermatozoa of asthenozoospermic individuals | Identification of 66 differentially regulated phosphoproteins | 2012 (95) |
| In-solution digestion LC-MS/MS | To compare the proteomic profiles of spermatozoa exhibiting an impaired capacity for sperm-egg recognition with normal cells | Identification of seven proteins showing different abundance in sperm unable to bind to the ZP | 2012 (96) |
| 2DE-DIGE, in-gel digestion LC-MS | Spermatozoa suspensions from ROS+ and ROS− groups analyzed | Identification of 31 protein spots showing different abundance (25 increased and 6 decreased) | 2013 (80) |
| 2DE-DIGE, in-gel digestion MALFI-TOF/TOF MS | To explore the differentially expressed proteins in normal sperm motility and idiopathic asthenozoospermia | Identification of 15 proteins showing different abundance | 2013 (97) |
| SDS-PAGE in-gel digestion LC-MS/MS | To screen for associations between sperm protein profiles and sperm concentration, motility, and DNA fragmentation index | Identification of 4 protein groups that correlate with DNA fragmentation and/or motility | 2013 (98) |
| SDS-PAGE in-gel digestion LC-MS/MS | Analysis of the human sperm profile with high sperm counts | Significant inter-individual variation in head sperm protein profiles | 2013 (99) |
| In-solution digestion LC-MS/MS | To examine if elevated levels of reactive oxygen species cause an alteration in the proteomic profile of spermatozoa | Identification of 15 proteins showing different abundance (10 increased and five decreased abundance) | 2013 (81) |
| In-solution digestion 2D LC-MS/MS | The human sperm proteome profile of law and high DNA fragmentation of normozoospermic men | Identification of 71 (low DNA fragmentation) and 23 (high DNA fragmentation) proteins showing different abundance | 2013 (100) |
| In-solution digestion, 6-plex TMT labeling, LC-MS/MS | Proteomic profiling of the human spermatozoa following successful or unsuccessful pregnancy via assisted reproductive technology (ART) | Identification of 21 proteins showing different abundance | 2013 (101) |
| 2-plex TMT labeling, SDS-PAGE in gel digestion LC-MS/MS | Normozoospermic sperm proteome samples with different IVF outcomes (pregnancy versus no pregnancy) compared | Identification of 64 proteins showing different abundance | 2014 (102) |
| 2D-DIGE, in-gel digestion MALDI-TOF MS | Proteomics profiling of the human sperm cryopreservation | 27 proteins showing different abundance | 2014 (103) |
| 2D-DIGE, in-gel digestion MALDI-TOF MS/MS | Proteome analysis of sperm samples collected by swim-up from control and acute epididymitis patients analyzed | 35 proteins showing different abundance | 2014 (104) |
| 2D-DIGE in-gel digestion MALDI-TOF MS or LC-MS/MS | Proteomic profiles of spermatozoa in patients with a complete failure of fertilization and no spermatozoa bound to the zona pellucida compared with those of controls | Identification of 12 proteins showing different abundance | 2014 (105) |
| SDS-PAGE in-gel digestion LC-MS/MS | To investigate the human sperm proteome and its relation to blastocyst development and reproductive success | Identification of 49 proteins showing different abundance in sperm resulting in bad embryo development | 2014 (106) |
| In-solution digestion LC-MS/MS | Differential proteomic analysis was performed on spermatozoa from both obesity-associated asthenozoospermia and clinically healthy individuals | Identification of 127 proteins showing different abundance | 2014 (107) |
| 6-plex TMT labeling, SDS-PAGE in gel digestion LC-MS/MS | The proteomics study was based on a comparison between sperm samples differing in motility (asthenozoospermic versus normozoospermic) and comparison between sperm subpopulations of fractionated normozoospermic samples differing in motility (non-migrated versus migrated) | Similar proteomic alterations detected in asthenozoospermic and nonmigrated sperm | 2014 (108) |
| SDS-PAGE in-gel digestion LC-MS/MS | Study on immature and mature ejaculated sperm from fertile men | 98 of them showed an increasing trend in expression levels | 2016 (109) |
The primary studies of the human spermatozoa proteome which used the differential proteomics approaches were focused on failure in the in vitro fertilization (IVF) due to male factor (83, 87). The authors identified 32 proteins which could improve the understanding of IVF failure due to male factor. They used gel-based proteomics technology (2DE followed by MALDI-TOF-MS protein identification). More recently, two other studies have used gel-free applications of proteomics approaches (6-plex TMT labeling followed by LC-MS/MS) on the human sperm to dig deeper on understanding IVF failure due to male factor (101, 102). Altogether, the mentioned studies have reported 85 deregulated proteins suggesting that epigenetic alterations may contribute to failure of assisted reproduction. Another interesting published study is based on frozen–thawed versus fresh human spermatozoa proteome that showed a malfunction of spermatozoon after cryopreservation (103).
On the other hand, several studies have focused on the asthenozoospermic patients. The importance of these patients is the high number of them and identification of proteins which are involved in the sperm motility. Furthermore, a sufficient amount of spermatozoa proteins can be easily extracted from asthenozoospermic sperm (84–86, 91, 95, 97, 107, 110).
Taken together, all deregulated identified proteins which have used MS-based proteomics technology shared protein involved in the cytoskeleton, metabolism or energy production (41).
Some studies have focused on reactive oxygen species (ROS) effect on the human spermatozoa. An imbalance in oxidative stress caused by a high generation of ROS by mitochondria has an effect on DNA of the human spermatozoa. Furthermore, it has an effect on the spermatozoa proteome (80, 81, 98, 100).
Very few studies have focused on the globozoospermic and oligoasthenozoospermic sperm. Both studies showed an altered proteome compared to fertile human sperm proteome (88, 92).
Two different studies have revealed the harmful consequence of the metabolic diseases including diabetes or obesity on the human sperm; however, the damaging effect on male fertility is not well identified at the molecular level. In their studies, they found the significant changes in the composition of the human sperm proteome (90, 93).
Finally, Cui et al. applied the proteomics to a relevant human fertility model and identified proteins which were critical for sperm maturation, motility and fertilization capacity (109).
Conclusion
Great efforts have been done to explore the human proteome after identification of the human genome. Fifteen years after the first draft of the human genome, it is obvious today that the complexity of the human lays on the human proteome. A network of scientific collaboration has investigated human proteome mapping using advanced mass spectrometry-based proteomics. Regarding the proteome mapping of the human spermatozoa, the research is still in its infancy in spite of knowing the fact that male factor contributes 50% to infertility. In this review, the human proteome information was assessed with the specific focus on the human sperm proteome anno 2017. The most precise human protein database shows ∼21,931 proteins. Furthermore, some researchers have been able to identify on average 10,361 proteins from cell lines using advance mass spectrometry-based proteomics. However, the number of identified proteins from the human spermatozoa is limited to ∼6,500. This can be caused by either reaching mass-spectrometry current limitations or not reaching the depth of human spermatozoa proteome.
In order to go deeper in identification of the human proteome, the proteomics researchers have formed the international consortium for the Chromosome-Centric Human Proteome Project. Iran is leading the mapping of Y chromosome. Accordingly, by looking at the human Y chromosome-encoding proteins, it is clear that these proteins are low expressed in the human sperm. It is, furthermore, recognized that the human sperm proteins are also low expressed compared to other cells. However, with the development of mass spectrometry and miniaturization of sample preparations, it seems there is still work to do regarding identification and quantification of the human Y chromosome-encoded proteins.
To conclude, despite several publications that have focused on many comparative and functional sperm proteomic studies and providing putative biomarkers for male (in) fertility, some points are still unclear. The use of higher throughput techniques coupled to various up-to-date options for differential proteomics might provide further light toward knowledge of sperm (dys) functions at molecular level.
Footnotes
Conflict of Interest
The authors declare no conflict of interest. This article has not received financial support.
References
- 1.Pruitt KD, Tatusova T, Maglott DR. NCBI reference sequences (RefSeq): a curated non-redundant sequence database of genomes, transcripts and proteins. Nucleic Acids Res. 2007;35(Database issue): D61–5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Schlüter H, Apweiler R, Holzhütter HG, Jungblut PR. Finding one’s way in proteomics: a protein species nomenclature. Chem Cent J. 2009;3:11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.O’Farrell PH. High resolution two-dimensional electrophoresis of proteins. J Biol Chem. 1975;250(10): 4007–21. [PMC free article] [PubMed] [Google Scholar]
- 4.Crutchfield CA, Thomas SN, Sokoll LJ, Chan DW. Advances in mass spectrometry-based clinical biomarker discovery. Clin Proteomics. 2016;13:1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Wilkins MR, Sanchez JC, Gooley AA, Appel RD, Humphery-Smith I, Hochstrasser DF, et al. Progress with proteome projects: why all proteins expressed by a genome should be identified and how to do it. Biotechnol Genet Eng Rev. 1996;13:19–50. [DOI] [PubMed] [Google Scholar]
- 6.Bray D. The protein explosion. Nature. 1977;267 (5611):481–2. [Google Scholar]
- 7.Aebersold R, Mann M. Mass spectrometry-based proteomics. Nature. 2003;422(6928):198–207. [DOI] [PubMed] [Google Scholar]
- 8.Hondermarck H. Proteomics: biomedical and pharmaceutical applications. 1st ed Dordrecht: Kluwer Academic Publishers; 2004. 407 p. [Google Scholar]
- 9.Tyers M, Mann M. From genomics to proteomics. Nature. 2003;422(6928):193–7. [DOI] [PubMed] [Google Scholar]
- 10.Gilany K, Van Elzen R, Mous K, Coen E, Van Dongen W, Vandamme S, et al. The proteome of the human neuroblastoma cell line SH-SY5Y: an enlarged proteome. Biochim Biophys Acta. 2008; 1784(7–8):983–5. [DOI] [PubMed] [Google Scholar]
- 11.Gilany K, Moens L, Dewilde S. Mass spectrometry-based proteomics in the life sciences: a review. J Paramed Sci. 2010;1(1). [Google Scholar]
- 12.Tran JC, Zamdborg L, Ahlf DR, Lee JE, Catherman AD, Durbin KR, et al. Mapping intact protein isoforms in discovery mode using top-down proteomics. Nature. 2011;480(7376):254–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Chait BT. Chemistry. Mass spectrometry: bottomup or top-down? Science. 2006;314(5796):65–6. [DOI] [PubMed] [Google Scholar]
- 14.Smith LM, Kelleher NL, Consortium for Top Down Proteomics Proteoform: a single term describing protein complexity. Nat Methods. 2013;10 (3):186–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Patrie SM. Top-down mass spectrometry: proteomics to proteoforms. Adv Exp Med Biol. 2016; 919:171–200. [DOI] [PubMed] [Google Scholar]
- 16.Toby TK, Fornelli L, Kelleher NL. Progress in Top-Down Proteomics and the Analysis of Proteoforms. Annu Rev Anal Chem (Palo Alto Calif). 2016;9(1):499–519. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Cravatt BF, Simon GM, Yates JR., 3rd The biological impact of mass-spectrometry-based proteomics. Nature. 2007;450(7172):991–1000. [DOI] [PubMed] [Google Scholar]
- 18.Bensimon A, Heck AJ, Aebersold R. Mass spectrometry-based proteomics and network biology. Annu Rev Biochem. 2012;81:379–405. [DOI] [PubMed] [Google Scholar]
- 19.Legrain P, Aebersold R, Archakov A, Bairoch A, Bala K, Beretta L, et al. The human proteome project: current state and future direction. Mol Cell Proteomics. 2011;10(7):M111.009993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Paik YK, Jeong SK, Omenn GS, Uhlen M, Hanash S, Cho SY, et al. The chromosome-centric human proteome project for cataloging proteins encoded in the genome. Nat Biotechnol. 2012;30(3):221–3. [DOI] [PubMed] [Google Scholar]
- 21.Hancock W, Omenn G, Legrain P, Paik YK. Proteomics, human proteome project, and chromosomes. J Proteome Res. 2011;10(1):210. [DOI] [PubMed] [Google Scholar]
- 22.Paik YK, Omenn GS, Uhlen M, Hanash S, Marko-Varga G, Aebersold R, et al. Standard guidelines for the chromosome-centric human proteome project. J Proteome Res. 2012;11(4):2005–13. [DOI] [PubMed] [Google Scholar]
- 23.Omenn GS. The strategy, organization, and progress of the HUPO human proteome project. J Proteomics. 2014;100:3–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Marko-Varga G, Omenn GS, Paik YK, Hancock WS. A first step toward completion of a genome-wide characterization of the human proteome. J Proteome Res. 2013;12(1):1–5. [DOI] [PubMed] [Google Scholar]
- 25.Kim MS, Pinto SM, Getnet D, Nirujogi RS, Manda SS, Chaerkady R, et al. A draft map of the human proteome. Nature. 2014;509(7502):575–81. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Wilhelm M, Schlegl J, Hahne H, Gholami AM, Lieberenz M, Savitski MM, et al. Mass-spectrometry-based draft of the human proteome. Nature. 2014;509(7502):582–7. [DOI] [PubMed] [Google Scholar]
- 27.Ezkurdia I, Vázquez J, Valencia A, Tress M. Analyzing the first drafts of the human proteome. J Proteome Res. 2014;13(8):3854–5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Ezkurdia I, Calvo E, Del Pozo A, Vázquez J, Valencia A, Tress ML. The potential clinical impact of the release of two drafts of the human proteome. Expert Rev Proteomics. 2015;12(6):579–93. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Omenn GS, Lane L, Lundberg EK, Beavis RC, Nesvizhskii AI, Deutsch EW. Metrics for the human proteome project 2015: progress on the human proteome and guidelines for high-confidence protein identification. J Proteome Res. 2015;14(9): 3452–60. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Uhlén M, Fagerberg L, Hallström BM, Lindskog C, Oksvold P, Mardinoglu A, et al. Proteomics. tissue-based map of the human proteome. Science. 2015;347(6220):1260419. [DOI] [PubMed] [Google Scholar]
- 31.Zhang B, Wang J, Wang X, Zhu J, Liu Q, Shi Z, et al. Proteogenomic characterization of human colon and rectal cancer. Nature. 2014;513(7518):382–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Hathout Y. Proteomic methods for biomarker discovery and validation. Are we there yet? Expert Rev Proteomics. 2015;12(4):329–31. [DOI] [PubMed] [Google Scholar]
- 33.Aebersold R, Bader GD, Edwards AM, van Eyk JE, Kussmann M, Qin J, et al. The biology/disease-driven human proteome project (B/D-HPP): enabling protein research for the life sciences community. J Proteome Res. 2013;12(1):23–7. [DOI] [PubMed] [Google Scholar]
- 34.Gilany K, Minai-Tehrani A, Savadi-Shiraz E, Rezadoost H, Lakpour N. Exploring the human seminal plasma proteome: an unexplored gold mine of biomarker for male infertility and male reproduction disorder. J Reprod Infertil. 2015;16(2):61–71. [PMC free article] [PubMed] [Google Scholar]
- 35.Kabiri M, Amoozegar MA, Tabebordbar M, Gilany K, Salekdeh GH. Effects of selenite and tellurite on growth, physiology, and proteome of a moderately halophilic bacterium. J Proteome Res. 2009; 8(6):3098–108. [DOI] [PubMed] [Google Scholar]
- 36.Kahn P. From genome to proteome: looking at a cell’s proteins. Science. 1995;270(5235):369–70. [DOI] [PubMed] [Google Scholar]
- 37.Ainsworth C. Cell biology: the secret life of sperm. Nature. 2005;436(7052):770–1. [DOI] [PubMed] [Google Scholar]
- 38.Baker MA, Reeves G, Hetherington L, Müller J, Baur I, Aitken RJ. Identification of gene products present in Triton X-100 soluble and insoluble fractions of human spermatozoa lysates using LC-MS/MS analysis. Proteomics Clin Appl. 2007;1(5): 524–32. [DOI] [PubMed] [Google Scholar]
- 39.Gilany K, Lakpour N, Vafakhah M, Sadeghi MR. The profile of human sperm proteome; a minireview. J Reprod Infertil. 2011;12(3):193–9. [PMC free article] [PubMed] [Google Scholar]
- 40.Wang G, Guo Y, Zhou T, Shi X, Yu J, Yang Y, et al. In-depth proteomic analysis of the human sperm reveals complex protein compositions. J Proteomics. 2013;79:114–22. [DOI] [PubMed] [Google Scholar]
- 41.Amaral A, Castillo J, Ramalho-Santos J, Oliva R. The combined human sperm proteome: cellular pathways and implications for basic and clinical science. Hum Reprod Update. 2014;20(1):40–62. [DOI] [PubMed] [Google Scholar]
- 42.Johnston DS, Wooters J, Kopf GS, Qiu Y, Roberts KP. Analysis of the human sperm proteome. Ann N Y Acad Sci. 2005;1061:190–202. [DOI] [PubMed] [Google Scholar]
- 43.de Godoy LM, Olsen JV, de Souza GA, Li G, Mortensen P, Mann M. Status of complete proteome analysis by mass spectrometry: SILAC labeled yeast as a model system. Genome Biol. 2006;7(6): R50. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Geiger T, Wehner A, Schaab C, Cox J, Mann M. Comparative proteomic analysis of eleven common cell lines reveals ubiquitous but varying expression of most proteins. Mol Cell Proteomics. 2012;11(3): M111.014050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Breuza L, Poux S, Estreicher A, Famiglietti ML, Magrane M, Tognolli M, et al. The UniProtKB guide to the human proteome. Database (Oxford). 2016;2016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.ProteomicsDB [Internet] Technische Universität München (TUM) and SAP SE: Germany; 2014 [cited 2017 July 6]. Available from: https://www.proteomicsdb.org/.
- 47.Gaudet P, Michel PA, Zahn-Zabal M, Cusin I, Duek PD, Evalet O, et al. The neXtProt knowledgebase on human proteins: current status. Nucleic Acids Res. 2015;43(Database issue):D764–70. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Vogel C, Abreu Rde S, Ko D, Le SY, Shapiro BA, Burns SC, et al. Sequence signatures and mRNA concentration can explain two-thirds of protein abundance variation in a human cell line. Mol Syst Biol. 2010;6:400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Beck M, Schmidt A, Malmstroem J, Claassen M, Ori A, Szymborska A, et al. The quantitative proteome of a human cell line. Mol Syst Biol. 2011;7: 549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Farrah T, Deutsch EW, Omenn GS, Campbell DS, Sun Z, Bletz JA, et al. A high-confidence human plasma proteome reference set with estimated concentrations in PeptideAtlas. Mol Cell Proteomics. 2011;10(9):M110.006353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Gaudet P, Argoud-Puy G, Cusin I, Duek P, Evalet O, Gateau A, et al. neXtProt: organizing protein knowledge in the context of human proteome projects. J Proteome Res. 2013;12(1):293–8. [DOI] [PubMed] [Google Scholar]
- 52.Lane L, Bairoch A, Beavis RC, Deutsch EW, Gaudet P, Lundberg E, et al. Metrics for the human proteome project 2013–2014 and strategies for finding missing proteins. J Proteome Res. 2014;13 (1):15–20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Reddy PJ, Ray S, Srivastava S. The quest of the human proteome and the missing proteins: digging deeper. OMICS. 2015;19(5):276–82. [DOI] [PubMed] [Google Scholar]
- 54.Fago A, Hundahl C, Dewilde S, Gilany K, Moens L, Weber RE. Allosteric regulation and temperature dependence of oxygen binding in human neuroglobin and cytoglobin. molecular mechanisms and physiological significance. J Biol Chem. 2004;279(43):44417–26. [DOI] [PubMed] [Google Scholar]
- 55.Horvatovich P, Lundberg EK, Chen YJ, Sung TY, He F, Nice EC, et al. Quest for missing proteins: update 2015 on chromosome-centric human proteome project. J Proteome Res. 2015;14(9):3415–31. [DOI] [PubMed] [Google Scholar]
- 56.Paik YK, Omenn GS, Overall CM, Deutsch EW, Hancock WS. Recent advances in the chromosome-centric human proteome project: missing proteins in the spot light. J Proteome Res. 2015;14 (9):3409–14. [DOI] [PubMed] [Google Scholar]
- 57.Jumeau F, Com E, Lane L, Duek P, Lagarrigue M, Lavigne R, et al. Human spermatozoa as a model for detecting missing proteins in the context of the chromosome-centric human proteome project. J Proteome Res. 2015;14(9):3606–20. [DOI] [PubMed] [Google Scholar]
- 58.Hosseini Salekdeh Gh. I-49: Human Y chromosome proteomeproject. Int J Fertil Steril. 2010;4 (Suppl 1):49. [Google Scholar]
- 59.Goodfellow P, Darling S, Wolfe J. The human Y chromosome. J Med Genet. 1985;22(5):329–44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.German J, Simpson JL, Chaganti RS, Summitt RL, Reid LB, Merkatz IR. Genetically determined sex-reversal in 46,XY humans. Science. 1978;202 (4363):53–6. [DOI] [PubMed] [Google Scholar]
- 61.Pryor JL, Kent-First M, Muallem A, Van Bergen AH, Nolten WE, Meisner L, et al. Microdeletions in the Y chromosome of infertile men. N Engl J Med. 1997;336(8):534–9. [DOI] [PubMed] [Google Scholar]
- 62.Cooke HJ. Y chromosome and male infertility. Rev Reprod. 1999;4(1):5–10. [DOI] [PubMed] [Google Scholar]
- 63.Krausz C, McElreavey K. Y chromosome and male infertility. Front Biosci. 1999;4:E1–8. [DOI] [PubMed] [Google Scholar]
- 64.Jangravi Z, Alikhani M, Arefnezhad B, Sharifi Tabar M, Taleahmad S, Karamzadeh R, et al. A fresh look at the male-specific region of the human Y chromosome. J Proteome Res. 2013;12(1):6–22. [DOI] [PubMed] [Google Scholar]
- 65.Rengaraj D, Kwon WS, Pang MG. Bioinformatics annotation of human Y chromosome-encoded protein pathways and interactions. J Proteome Res. 2015;14(9):3503–18. [DOI] [PubMed] [Google Scholar]
- 66.Gilany K, Aerts M, Dewilde S, Devreese B, Moens L, Vafakhah M. Proteome analysis of mouse brain exposed to chronic hypoxia. Cell. 2011;12(4):503–10. [Google Scholar]
- 67.Szklarczyk D, Franceschini A, Wyder S, Forslund K, Heller D, Huerta-Cepas J, et al. STRING v10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 2015;43 (Database issue):D447–52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Croft D, Mundo AF, Haw R, Milacic M, Weiser J, Wu G, et al. The Reactome pathway knowledgebase. Nucleic Acids Res. 2014;42(Database issue): D472–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Premkumar E, Bhargava PM. Transcription and translation in bovine spermatozoa. Nat New Biol. 1972;240(100):139–43. [DOI] [PubMed] [Google Scholar]
- 70.Baker MA. The ‘omics revolution and our understanding of sperm cell biology. Asian J Androl. 2011;13(1):6–10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Zhou T, Xia X, Liu J, Wang G, Guo Y, Guo X, et al. Beyond single modification: reanalysis of the acetylproteome of human sperm reveals widespread multiple modifications. J Proteomics. 2015; 126:296–302. [DOI] [PubMed] [Google Scholar]
- 72.Khoury GA, Baliban RC, Floudas CA. Proteomewide post-translational modification statistics: frequency analysis and curation of the swiss-prot database. Sci Rep. 2011;1:90. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Wang G, Wu Y, Zhou T, Guo Y, Zheng B, Wang J, et al. Mapping of the N-linked glycoproteome of human spermatozoa. J Proteome Res. 2013;12(12): 5750–9. [DOI] [PubMed] [Google Scholar]
- 74.Ficarro S, Chertihin O, Westbrook VA, White F, Jayes F, Kalab P, et al. Phosphoproteome analysis of capacitated human sperm. evidence of tyrosine phosphorylation of a kinase-anchoring protein 3 and valosin-containing protein/p97 during capacitation. J Biol Chem. 2003;278(13):11579–89. [DOI] [PubMed] [Google Scholar]
- 75.Sun G, Jiang M, Zhou T, Guo Y, Cui Y, Guo X, et al. Insights into the lysine acetylproteome of human sperm. J Proteomics. 2014;109:199–211. [DOI] [PubMed] [Google Scholar]
- 76.World Health Organization, Department of Reproductive Health and Research WHO laboratory manual for the examination and processing of human semen. 5th ed Switzerland: World Health Organization; 2010. 286 p. [Google Scholar]
- 77.Oliva R, Castillo J, editors. Sperm nucleoproteins. USA: Springer; 2011. 504 p. (Zini A, Agarwal A, editors. Sperm chromatin: Biological and clinical applications in male infertility and assisted reproduction). [Google Scholar]
- 78.de Mateo S, Martínez-Heredia J, Estanyol JM, Domínguez-Fandos D, Vidal-Taboada JM, Ballescà JL, et al. Marked correlations in protein expression identified by proteomic analysis of human spermatozoa. Proteomics. 2007;7(23):4264–77. [DOI] [PubMed] [Google Scholar]
- 79.Aitken RJ, Smith TB, Jobling MS, Baker MA, De Iuliis GN. Oxidative stress and male reproductive health. Asian J Androl. 2014;16(1):31–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Hamada A, Sharma R, du Plessis SS, Willard B, Yadav SP, Sabanegh E, et al. Two-dimensional differential in-gel electrophoresis-based proteomics of male gametes in relation to oxidative stress. Fertil Steril. 2013;99(5):1216–26.e2. [DOI] [PubMed] [Google Scholar]
- 81.Sharma R, Agarwal A, Mohanty G, Hamada AJ, Gopalan B, Willard B, et al. Proteomic analysis of human spermatozoa proteins with oxidative stress. Reprod Biol Endocrinol. 2013;11:48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Jafarzadeh N, Mani-Varnosfaderani A, Minai-Therani A, Savadi-Shiraz E, Sadeghi MR, Gilany K. Metabolomics fingerprinting of seminal plasma from unexplained infertile men: a need for novel diagnostic biomarkers. Mol Reprod Dev. 2015;82 (3):150. [DOI] [PubMed] [Google Scholar]
- 83.Pixton KL, Deeks ED, Flesch FM, Moseley FL, Björndahl L, Ashton PR, et al. Sperm proteome mapping of a patient who experienced failed fertilization at IVF reveals altered expression of at least 20 proteins compared with fertile donors: case report. Hum Reprod. 2004;19(6):1438–47. [DOI] [PubMed] [Google Scholar]
- 84.Zhao C, Huo R, Wang FQ, Lin M, Zhou ZM, Sha JH. Identification of several proteins involved in regulation of sperm motility by proteomic analysis. Fertil Steril. 2007;87(2):436–8. [DOI] [PubMed] [Google Scholar]
- 85.Martínez-Heredia J, de Mateo S, Vidal-Taboada JM, Ballescà JL, Oliva R. Identification of proteomic differences in asthenozoospermic sperm samples. Hum Reprod. 2008;23(4):783–91. [DOI] [PubMed] [Google Scholar]
- 86.Chan CC, Shui HA, Wu CH, Wang CY, Sun GH, Chen HM, et al. Motility and protein phosphorylation in healthy and asthenozoospermic sperm. J Proteome Res. 2009;8(11):5382–6. [DOI] [PubMed] [Google Scholar]
- 87.Frapsauce C, Pionneau C, Bouley J, de Larouzière V, Berthaut I, Ravel C, et al. [Unexpected in vitro fertilization failure in patients with normal sperm: a proteomic analysis]. Gynecol Obstet Fertil. 2009; 37(10):796–802. French. [DOI] [PubMed] [Google Scholar]
- 88.Liao TT, Xiang Z, Zhu WB, Fan LQ. Proteome analysis of round-headed and normal spermatozoa by 2-D fluorescence difference gel electrophoresis and mass spectrometry. Asian J Androl. 2009;11 (6):683–93. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Secciani F, Bianchi L, Ermini L, Cianti R, Armini A, La Sala GB, et al. Protein profile of capacitated versus ejaculated human sperm. J Proteome Res. 2009;8(7):3377–89. [DOI] [PubMed] [Google Scholar]
- 90.Kriegel TM, Heidenreich F, Kettner K, Pursche T, Hoflack B, Grunewald S, et al. Identification of diabetes- and obesity-associated proteomic changes in human spermatozoa by difference gel electrophoresis. Reprod Biomed Online. 2009;19(5):660–70. [DOI] [PubMed] [Google Scholar]
- 91.Siva AB, Kameshwari DB, Singh V, Pavani K, Sundaram CS, Rangaraj N, et al. Proteomics-based study on asthenozoospermia: differential expression of proteasome alpha complex. Mol Hum Reprod. 2010;16(7):452–62. [DOI] [PubMed] [Google Scholar]
- 92.Thacker S, Yadav SP, Sharma RK, Kashou A, Willard B, Zhang D, et al. Evaluation of sperm proteins in infertile men: a proteomic approach. Fertil Steril. 2011;95(8):2745–8. [DOI] [PubMed] [Google Scholar]
- 93.Paasch U, Heidenreich F, Pursche T, Kuhlisch E, Kettner K, Grunewald S, et al. Identification of increased amounts of eppin protein complex components in sperm cells of diabetic and obese individuals by difference gel electrophoresis. Mol Cell Proteomics. 2011;10(8):M110.007187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Xu W, Hu H, Wang Z, Chen X, Yang F, Zhu Z, et al. Proteomic characteristics of spermatozoa in normozoospermic patients with infertility. J Proteomics. 2012;75(17):5426–36. [DOI] [PubMed] [Google Scholar]
- 95.Parte PP, Rao P, Redij S, Lobo V, D’Souza SJ, Gajbhiye R, et al. Sperm phosphoproteome profiling by ultra performance liquid chromatography followed by data independent analysis (LC-MS(E)) reveals altered proteomic signatures in asthenozoospermia. J Proteomics. 2012;75(18):5861–71. [DOI] [PubMed] [Google Scholar]
- 96.Redgrove KA, Nixon B, Baker MA, Hetherington L, Baker G, Liu DY, et al. The molecular chaper-one HSPA2 plays a key role in regulating the expression of sperm surface receptors that mediate sperm-egg recognition. PLoS One. 2012;7(11): e50851. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Shen S, Wang J, Liang J, He D. Comparative proteomic study between human normal motility sperm and idiopathic asthenozoospermia. World J Urol. 2013;31(6):1395–401. [DOI] [PubMed] [Google Scholar]
- 98.Behrouzi B, Kenigsberg S, Alladin N, Swanson S, Zicherman J, Hong SH, et al. Evaluation of potential protein biomarkers in patients with high sperm DNA damage. Syst Biol Reprod Med. 2013;59(3): 153–63. [DOI] [PubMed] [Google Scholar]
- 99.Kichine E, Di Falco M, Hales BF, Robaire B, Chan P. Analysis of the sperm head protein profiles in fertile men: consistency across time in the levels of expression of heat shock proteins and peroxiredoxins. PLoS One. 2013;8(10):e77471. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Intasqui P, Camargo M, Del Giudice PT, Spaine DM, Carvalho VM, Cardozo KH, et al. Unraveling the sperm proteome and post-genomic pathways associated with sperm nuclear DNA fragmentation. J Assist Reprod Genet. 2013;30(9): 1187–202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Zhu Y, Wu Y, Jin K, Lu H, Liu F, Guo Y, et al. Differential proteomic profiling in human spermatozoa that did or did not result in pregnancy via IVF and AID. Proteomics Clin Appl. 2013;7 (11–12):850–8. [DOI] [PubMed] [Google Scholar]
- 102.Azpiazu R, Amaral A, Castillo J, Estanyol JM, Guimerà M, Ballescà JL, et al. High-throughput sperm differential proteomics suggests that epigenetic alterations contribute to failed assisted reproduction. Hum Reprod. 2014;29(6):1225–37. [DOI] [PubMed] [Google Scholar]
- 103.Wang S, Wang W, Xu Y, Tang M, Fang J, Sun H, et al. Proteomic characteristics of human sperm cryopreservation. Proteomics. 2014;14(2–3):298–310. [DOI] [PubMed] [Google Scholar]
- 104.Pilatz A, Lochnit G, Karnati S, Paradowska-Dogan A, Lang T, Schultheiss D, et al. Acute epididymitis induces alterations in sperm protein composition. Fertil Steril. 2014;101(6):1609–17. e1–5. [DOI] [PubMed] [Google Scholar]
- 105.Frapsauce C, Pionneau C, Bouley J, Delarouziere V, Berthaut I, Ravel C, et al. Proteomic identification of target proteins in normal but nonfertilizing sperm. Fertil Steril. 2014;102(2):372–80. [DOI] [PubMed] [Google Scholar]
- 106.McReynolds S, Dzieciatkowska M, Stevens J, Hansen KC, Schoolcraft WB, Katz-Jaffe MG. Toward the identification of a subset of unexplained infertility: a sperm proteomic approach. Fertil Steril. 2014;102(3):692–9. [DOI] [PubMed] [Google Scholar]
- 107.Liu Y, Guo Y, Song N, Fan Y, Li K, Teng X, et al. Proteomic pattern changes associated with obesity-induced asthenozoospermia. Andrology. 2015;3(2):247–59. [DOI] [PubMed] [Google Scholar]
- 108.Amaral A, Castillo J, Estanyol JM, Ballescà JL, Ramalho-Santos J, Oliva R. Human sperm tail proteome suggests new endogenous metabolic pathways. Mol Cell Proteomics. 2013;12(2):330–42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Cui Z, Sharma R, Agarwal A. Proteomic analysis of mature and immature ejaculated spermatozoa from fertile men. Asian J Androl. 2016;18(5): 735–46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Amaral A, Paiva C, Attardo Parrinello C, Estanyol JM, Ballescà JL, Ramalho-Santos J, et al. Identification of proteins involved in human sperm motility using high-throughput differential proteomics. J Proteome Res. 2014;13(12):5670–84. [DOI] [PubMed] [Google Scholar]