Abstract
Background
Malaria programmes use Plasmodium falciparum histidine-rich protein-2 (PfHRP2) based rapid diagnostic tests (RDTs) for malaria diagnosis. The deletion of this target antigen could potentially lead to misdiagnosis, delayed treatment and continuation of active transmission.
Methods
Plasmodium falciparum isolates (n = 1162) collected in Southern Mozambique were assessed by RDTs, microscopy and/or 18SrRNA qPCR. pfhrp2 and pfhrp3 deletions were investigated in isolates from individuals who were negative by RDT but positive by microscopy and/or qPCR (n = 69) using gene-specific PCRs, with kelch13 PCR as the parasite DNA control.
Results
Lack of pfhrp2 PCR amplification was observed in one of the 69 isolates subjected to molecular analysis [1.45% (95% CI 0.3–7.8%)].
Conclusions
The low prevalence of pfhrp2 deletions suggests that RDTs will detect the vast majority of the P. falciparum infections. Nevertheless, active surveillance for changing deletion frequencies is required.
Keywords: Malaria, Deletion, RDT, Mozambique, Pfhrp2
Background
Malarial parasites exhibit striking genetic plasticity that allows their rapid adaption to new drugs [1] and detection methods [2, 3]. This adaptability of the parasite endangers preventive and therapeutic measures against malaria, as the success of control programmes largely relies on early diagnosis and effective treatment. Rapid diagnostic tests (RDTs) are commonly used in malaria case management and elimination programmes, particularly in remote areas where facilities for microscopy are not available [4].
Plasmodium falciparum histidine-rich protein-2 (PfHRP2), together with Plasmodium lactate dehydrogenase and aldolase, are the key target antigens in commercially available RDTs [5]. Evidence from South America, India and Africa [2, 3] suggest that “deletion” of the target epitope within the parasite PfHRP2 antigen could adversely impact the life of an affected individual as a consequence of delayed or no treatment. Besides pfhrp2, pfhrp3 also affects the performance of RDT, as it has sequence homology with the pfhrp2 and can be detected by the monoclonal antibodies used against PfHRP2 in RDTs [6].
With increasing false negative RDT reports in African countries, WHO has considered the need of rigorous monitoring of malaria parasites that lack the pfhrp2 gene [2, 3, 7]. RDTs were introduced in Mozambique in 2007 and national wide use started in 2010 [8]. However, there is no information available about the extent of pfhrp2 and pfhrp3 deletions in P. falciparum parasites circulating in Mozambique. In this context, this study aimed to assess the presence of pfhrp2 and pfhrp3 deletions in P. falciparum isolates from Manhiça and Magude districts of Southern Mozambique.
Methods
Study site and design
Between 2010 and 2016, a total of 9124 blood samples were collected onto filter papers during cross-sectional studies conducted at the beginning (November) or end (May) of the malaria season in Southern Mozambique (Manhiça and Magude; Table 1). In Mozambique, a peak in transmission is usually seen during the rainy season, from November to April. Transmission intensity in southern Mozambique is generally low, although areas of high transmission may still be observed [9].
Table 1.
P. falciparum isolates collected during cross-sectionals with diagnostic results
| Years | Place | Samples collected | P. falciparum positive samples | RDT negative, microscopy and qPCR positives samples | RDT negative and only qPCR positives samples |
|---|---|---|---|---|---|
| 2010 | Manhiça | 969 | 105 | 1 | – |
| 2011 | Manhiça | 842 | 138 | 3 | – |
| 2012 | Manhiça | 924 | 116 | 3 | – |
| 2013 | Manhiça | 829 | 166 | 8 | – |
| 2014 | Manhiça | 908 | 211 | 7 | – |
| 2015 | Manhiça | 770 | 93 | 9 | – |
| 2015 | Magude | 981 | 101 | 7 | – |
| 2015 | Magude | 1322 | 174 | – | 125 |
| 2016 | Magude | 1579 | 58 | 1 | – |
Malaria diagnosis was conducted using microscopy, HRP2-based RDT and qPCR; or only RDTs and qPCRs. The inclusion criteria for the deletion analysis were: 1) a negative HRP2-based RDT (SD BIOLINE Malaria Antigen P. f—05FK50) but positive by microscopy and qPCR (18S rRNA) or 2) a negative HRP2-based RDT but positive qPCR (18S rRNA) if microscopy was not performed. First, a nested PCR targeting single copy k13 gene (nPCRk13) was performed to verify the presence of parasite DNA in the sample [10]. Second, pfhrp2 and pfhrp3 genes were amplified using standard primers as described elsewhere [5, 11]. Finally, pfhrp2 and pfhrp3 deletions were concluded if kelch13 gene PCR was positive, but PCRs for pfhrp2 and pfhrp3 failed to amplify the respective gene. The laboratory-adapted culture lines 3D7 as a positive control for both pfhrp2 and pfhrp3, and HB3 and DD2 as negative controls for pfhrp3 and pfhrp2, respectively, were amplified simultaneously. The National Mozambican Ethics Review Committee and the Hospital Clínic of Barcelona Ethics Review Committee approved the collection of samples and molecular analysis. Informed consent and permission (in the case of children under 18 years of age) were also obtained from each participant or a parent/legal guardian during the cross-sectional studies.
Microscopy
Thin and thick blood smears were air-dried, stained with Giemsa and examined using a light microscope fitted with a 100 × oil immersion lens and a 10 × eyepiece to quantify parasitaemia in the Centro de Investigação em Saúde de Manhiça (CISM) laboratory [9]. Slides were read twice by two different qualified microscopists, and if there was discordance in the results, a third reading was performed by an additional microscopist.
Rapid diagnostic test
A trained laboratory technician collected approximately 10 μL of blood from an individual by finger-prick to perform an RDT. The PfHRP2-based RDT (SD BIOLINE Malaria Antigen P. f—05FK50) was used as per the manufacturer’s instructions.
DNA extraction and Plasmodium falciparum detection by real time PCR (qPCR)
DNA was extracted from a half of the filter paper (Whatman, 903TM), containing a 25 μL blood drop by using QIAamp DNA Mini kit (Qiagen). The ABI PRISM 7500 HT Real-Time System (Applied Biosystems) was used to amplify purified parasite DNA templates, using a previously described method [12, 13]. Parasitaemia in the clinical samples was quantified by extrapolation against the standard curve prepared from an in vitro culture of 3D7 strain.
kelch13 nested PCR (nPCRk13)
Purified DNA templates were amplified using 2720 Thermal Cycler (Applied Biosystems) following a previously described method for the kelch13 gene [10].
pfhrp2 and pfhrp3 PCRs
Samples with intact parasite DNA confirmed by nPCRk13 were used for further amplification of region covering exon 1 and 2, as well as exon 2 of pfhrp2 and pfhrp3 genes [5, 11], following previously described methods with minor changes. These changes include the use of 1× HOT FirePol Master Mix, annealing temperatures of 63 °C of 1 min for across regions of exon 1 and 2 of pfhrp2 gene and 60 °C of 1 min for exon 2 amplification for both pfhrp2 and pfhrp3 genes. PCR products were visualized using 2% agarose (Invitrogen) and a UV trans-illuminator.
Results
Among the 9124 blood samples collected from participants in cross-sectional studies conducted in Southern Mozambique between 2010 and 2016, 1162 were P. falciparum positive by qPCR and/or by microscopy and RDTs. Among these 1162 P. falciparum isolates, 164 samples were found eligible for the pfhrp2 and pfhrp3 deletion analysis based on a RDT negative, microscopy positive and qPCR positive results (MO+/RDT−/qPCR+; n = 39), or an RDT negative and qPCR positive result (RDT−/qPCR+ ; n = 125). Filter papers and corresponding DNAs were available for 155 (95%) of these 164 P. falciparum isolates. Among these, 70 (45%) were positive by nPCRK13 (849 bp amplicon size; Fig. 1). Median qPCR parasite densities of the P. falciparum isolates that were negative by nPCR targeting kelch13 gene was 2.17 parasites/µL (interquartile range 1.2–4.4 parasites/µL).
Fig. 1.
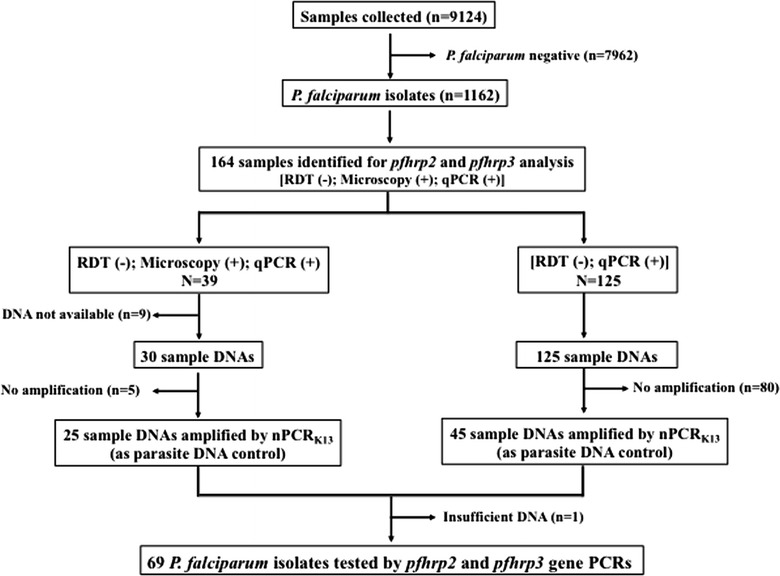
Schematic representation of sample selection for pfhrp2 and pfhrp3 deletion analysis
69 samples were analysed for pfhrp2 and pfhrp3 deletions, as one DNA sample was not enough for the analysis (Fig. 1). Parasite densities in these samples ranged from the 3 to 330,214 parasites/µL by qPCR (Table 2). The laboratory 3D7 strain returned all the expected PCR products of pfhrp2 (exon 1–2 = 303 bp and exon 2 = 816 bp) and pfhrp3 (exon 1–2 = 301 bp and exon 2 = 719 bp). As expected, laboratory strains DD2 and HB3 lacked pfhrp2 and pfhrp3 amplifications respectively (Fig. 2).
Table 2.
Parasite densities (parasites/µL of blood), age, sex and year of sample collection of the samples included in the study
| Years | Place | Parasitemia by microscopy | Parasitemia by qPCR | Sex | Age (in years) |
|---|---|---|---|---|---|
| 2010 | Manhiça | 1670 | 3858.5 | Male | 4 |
| 2013 | Manhiça | 56 | 33.5 | Male | 12 |
| 2013 | Manhiça | 546 | 600.3 | Male | 2 |
| 2013 | Manhiça | 143603 | 25250 | Male | 3 |
| 2013 | Manhiça | 203 | 1500 | Male | 14 |
| 2013 | Manhiça | 39 | 13.2 | Female | 24 |
| 2014 | Manhiça | 232 | 996.6 | Male | 4 |
| 2014 | Manhiça | 44594 | 23469.6 | Female | 15 |
| 2014 | Manhiça | 140814 | 84817 | Female | 3 |
| 2014 | Manhiça | 5721 | 3827.8 | Female | 7 |
| 2014 | Manhiça | 52 | 657.5 | Female | 3 |
| 2015 | Manhiça | 386 | 167.58659 | Female | 14 |
| 2015 | Manhiça | 1657 | 3407.6235 | Female | 8 |
| 2015 | Manhiça | 100 | 98.169426 | Female | 2 |
| 2015 | Manhiça | 325 | 156.13457 | Male | 2 |
| 2015 | Manhiça | 51 | 32.917961 | Male | 11 |
| 2015 | Manhiça | 99 | 108.99004 | Male | 17 |
| 2015 | Manhiça | 5648 | 1463.4641 | Female | NA |
| 2015 | Magude | 3610 | 1610.6819 | Female | 4 |
| 2015 | Magude | 2950.5 | 676.75568 | Male | 9 |
| 2015 | Magude | 303.5 | 224.56026 | Female | 50 |
| 2015 | Magude | 928.5 | 384.57782 | Male | 2 |
| 2015 | Magude | 2637 | 95.924171 | Female | 11 |
| 2015 | Magude | NA | 14.8498 | Male | 7 |
| 2015 | Magude | NA | 2.44336 | Female | 10 |
| 2015 | Magude | NA | 27.025 | Male | NA |
| 2015 | Magude | NA | 2.70269 | Female | 15 |
| 2015 | Magude | NA | 20.8876 | Female | 47 |
| 2015 | Magude | NA | 330214 | Female | 3 |
| 2015 | Magude | NA | 99.4996 | Female | 4 |
| 2015 | Magude | NA | 14.9045 | Female | 12 |
| 2015 | Magude | NA | 8.46764 | Female | 7 |
| 2015 | Magude | NA | 9.3634 | Male | 3 |
| 2015 | Magude | NA | 69.8859 | Male | 43 |
| 2015 | Magude | NA | 4.21137 | Female | 17 |
| 2015 | Magude | NA | 4.75359 | Female | 9 |
| 2015 | Magude | NA | 314.461 | Male | 4 |
| 2015 | Magude | NA | 104.915 | Male | 19 |
| 2015 | Magude | NA | 25.1698 | Female | 11 |
| 2015 | Magude | NA | 27.3208 | Male | 28 |
| 2015 | Magude | NA | 20.9188 | Female | NA |
| 2015 | Magude | NA | 1184.68 | Male | 15 |
| 2015 | Magude | NA | 4.68228 | Female | 9 |
| 2015 | Magude | NA | 612.026 | Female | 12 |
| 2015 | Magude | NA | 182.307 | Female | 7 |
| 2015 | Magude | NA | 61.8145 | Female | 5 |
| 2015 | Magude | NA | 6.33165 | Female | 2 |
| 2015 | Magude | NA | 50.2857 | Female | 40 |
| 2015 | Magude | NA | 90.4941 | Male | 1 |
| 2015 | Magude | NA | 14.7296 | Male | 3 |
| 2015 | Magude | NA | 539.879 | Female | 35 |
| 2015 | Magude | NA | 8.85961 | Male | 12 |
| 2015 | Magude | NA | 231.339 | Female | 8 |
| 2015 | Magude | NA | 19.9863 | Male | 15 |
| 2015 | Magude | NA | 55.4191 | Female | 2 |
| 2015 | Magude | NA | 73.8065 | Male | 12 |
| 2015 | Magude | NA | 24.4031 | Female | 29 |
| 2015 | Magude | NA | 419.439 | Female | 27 |
| 2015 | Magude | NA | 158.95 | Male | 12 |
| 2015 | Magude | NA | 6.46944 | Female | 45 |
| 2015 | Magude | NA | 9.88753 | Female | 40 |
| 2015 | Magude | NA | 2.68125 | Male | 18 |
| 2015 | Magude | NA | 78.1784 | Female | 13 |
| 2015 | Magude | NA | 20.7993 | Male | 15 |
| 2015 | Magude | NA | 576.566 | Female | 11 |
| 2015 | Magude | NA | 54.7864 | Male | 9 |
| 2015 | Magude | NA | 19.1635 | Female | 35 |
| 2015 | Magude | NA | 491.664 | Male | 45 |
| 2016 | Magude | 609 | 645.284 | Male | 31 |
NA not available
Fig. 2.
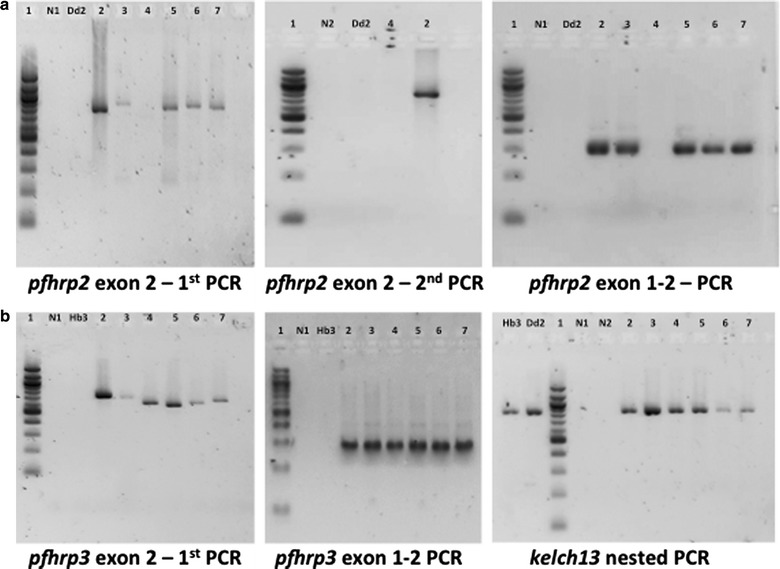
Molecular analysis of P. falciparum field isolates along with reference strains 3D7, Dd2 and HB3. a Amplification of regions covering exon 1 and 2 (exon 1–2) as well as exon2 of pfhrp2 in P. falciparum field isolates and reference strains. a Also show lack of amplification of region exons 1–2 and exon 2 of pfhrp2 in the field isolate (H1.3), Dd2 strain and negative controls whereas amplification was present in other isolates and 3D7 strain. b Amplification of kelch13 gene, region exon 1–2 and exon 2 of pfhrp3 in both P. falciparum field isolates and reference strains. (Lane 1—100 bp ladder; Lane 2—3D7; Lane 3—H1308; Lane 4—H1.3; Lane 5—H2.4; Lane 6—H2.7; Lane 7—H6.4; N1—negative control first PCR; N2—negative control nested PCR)
No amplification was noticed in negative controls (with water and human genomic DNA), which confirms the P. falciparum specificity of all the primer sets used in this study. Expected PCR products were observed upon the amplification of regions across exon 1 and exon 2, as well as exon 2 of pfhrp2 and pfhrp3 genes in all the samples except one sample (H1.3). PCR amplification of region covering pfhrp2 exon 1 and exon 2, as well as exon 2 was not observed in isolate H1.3 (Fig. 2a), while obtaining a positive amplification product for kelch13 gene. This lack of amplification was confirmed in a second and independent PCR test. The microscopy and qPCR parasite density of sample H1.3 were 2950.5 and 676.75 parasites/µL, respectively. Upon case investigation, this sample was found to correspond to a 30 months old male child from Magude who reported previous episodes of fever (during last 30 days), lived in a fumigated household and slept under a bed net the night before the sample was collected. Apart from this, varying pfhrp2 and pfhrp3 exon 2 PCR products lengths (600–1000 bp) were also observed in the analysed samples.
Discussion
This study provides the first evidence of pfhrp2 deletion in P. falciparum isolates circulating in Southern Mozambique. The prevalence of 1.45% (95% CI 0.3–7.8%) pfhrp2 deletion among analyzed samples is low as compared to the prevalence observed in other malaria endemic countries such as India (2.4%), Senegal (2.4%), Mali (5%) and Ghana (30.3%) [14–17]. As per WHO guidelines, 5% prevalence of pfhrp2 deletion has been considered as a minimum threshold to change the type of RDTs [3]. Therefore, PfHRP2-based RDTs are likely to detect the vast majority of the malaria parasites in southern Mozambique, but careful periodic monitoring for changes in deletion frequencies may be required to identify cases such as the single mutant detected in this study.
Previous reports have shown that pfhrp3 deletion could be an early warning signal for pfhrp2 deletion [11]. However, the pfhrp3 deletion has not been observed in the present study. Since only blood spots on filter paper were available in this study, plasma PfHRP2 protein levels or RNA based assays for the same sample could not be performed. However, a number of independent pfhrp2 PCR based investigation was done to confirm the lack of pfhrp2 gene in the P. falciparum isolate. Moreover, as significant amount of P. falciparum isolates (n = 164) were detected by real time PCR but not by nested PCR, given lower sensitivity of the latter [18], and consequently were not eligible for hrp2/hrp3 assessment. Finally, the varying length of exon 2 of pfhrp2 and pfhrp3 PCR products indicates the presence of different numbers of previously identified amino acid repeats [5].
According to 2016 WHO world malaria day fact sheet, the use of RDT has significantly increased globally from 46 million sold in 2008 to 314 million in 2014. In 2014, 53% of global RDTs (P. falciparum-specific tests) were delivered to African countries [3]. The excessive use of PfHRP2 based RDTs might enhance the selection of P. falciparum isolates with pfhrp2 deletion, especially in endemic areas where pfhrp2 deletion is present. Thus, it is important to monitor the presence of parasites with pfhrp2 and pfhrp3 deletions to avoid false negative results by RDT. Limitation of the study is that the sample’s material was not available for amplification of flanking genes of pfhrp2 and pfhrp3 genes.
Conclusions
The low prevalence of pfhrp2 deletions suggests that RDTs will detect the vast majority of the P. falciparum infections in Mozambique. However, active surveillance to detect increases in pfhrp2 deletion frequencies is required towards the common goal to eliminate malaria.
Authors’ contributions
HG carried out deletion PCRs analysis, interpretation of results and wrote the draft of this manuscript. GM, BG, PC, LN and WS participated in fieldwork, collected clinical and epidemiological data and laboratory analyses. AM, JC, FS, NRR, PA and PA participated in the study design, interpretation of results and writing of this article. All authors read and approved the final manuscript.
Acknowledgements
We would like to thank all the study participants, field workers and those who worked in the lab, as well as everyone who supported this study directly or indirectly.
Competing interests
The authors declare that they have no competing interests.
Availability of data and materials
The datasets analyzed in this study are available from the corresponding author on request.
Consent for publication
All authors have given their consent for publication.
Ethics approval and consent to participate
The National Mozambican Ethics Review Committee and the Hospital Clínic of Barcelona Ethics Review Committee approved the collection of samples and molecular analysis. Informed consent and permission (in the case of children under 18 years of age) were also obtained from each participant or a parent/legal guardian during the cross-sectional studies.
Funding
We would also like to thank the La Caixa and Bill and Melinda Gates Foundations for providing the funds for this study (Registration Number—OPP111526), as well as Instituto de Salud Carlos III [PI13/01478 cofunded by the Fondo Europeo de Desarrollo Regional (FEDER), and CES10/021-I3SNS to AM]. HG has a fellowship from the Overseas Postdoctoral Fellowship programme by Science and Engineering Research Board, Department of Science & Technology, Government of India (SB/OS/PDF-043/2015-16). The Centro de Investigaçao em Saude de Manhica (CISM) receives major core funding from the Spanish Agency for International Cooperation (AECI). ISGlobal is a member of the CERCA Programme, Generalitat de Catalunya.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Himanshu Gupta, Email: himanshu.gupta@isglobal.org.
Gloria Matambisso, Email: Gloria.Matambisso@manhica.net.
Beatriz Galatas, Email: beatriz.galatas@isglobal.org.
Pau Cisteró, Email: pau.cistero@isglobal.org.
Lidia Nhamussua, Email: lidia.nhamussua@manhica.net.
Wilson Simone, Email: Wilson.Simone@manhica.net.
Jane Cunningham, Email: cunninghamj@who.int.
N. Regina Rabinovitch, Email: regina.rabinovich@isglobal.org.
Pedro Alonso, Email: alonsop@who.int.
Franciso Saute, Email: francisco.saute@manhica.net.
Pedro Aide, Email: pedro.aide@manhica.net.
Alfredo Mayor, Phone: +34 93 227 5400, Email: alfredo.mayor@isglobal.org.
References
- 1.Bloland P. Drug resistance in malaria. Geneva, World Health Organization, 2001. http://www.who.int/csr/resources/publications/drugresist/malaria.pdf.
- 2.WHO. False-negative RDT results and implications of new reports of Plasmodium falciparum histidine-rich protein 2/3 gene deletions. Geneva: World Health Organization; 2016. http://www.finddx.org/wp-content/uploads/2016/06/WHO-information-note-false-neg-rdt-results-May2016.pdf.
- 3.WHO. Plasmodium falciparum hrp2/3gene deletions. Malaria Policy Advisory Committee Meeting. Geneva: World Health Organization; 2016. http://www.who.int/malaria/mpac/mpac-sept2016-hrp2-consultation-short-report-session7.pdf.
- 4.Wongsrichanalai C, Barcus MJ, Muth S, Sutamihardja A, Wernsdorfer WH. A review of malaria diagnostic tools: microscopy and rapid diagnostic test (RDT) Am J Trop Med Hyg. 2007;77:119–127. [PubMed] [Google Scholar]
- 5.Baker J, McCarthy J, Gatton M, Kyle DE, Belizario V, Luchavez J, et al. Genetic diversity of Plasmodium falciparum histidine-rich protein 2 (PfHRP2) and its effect on the performance of PfHRP2-based rapid diagnostic tests. J Infect Dis. 2005;192:870–877. doi: 10.1086/432010. [DOI] [PubMed] [Google Scholar]
- 6.Lee N, Baker J, Andrews KT, Gatton ML, Bell D, Cheng Q, et al. Effect of sequence variation in Plasmodium falciparum histidine-rich protein 2 on binding of specific monoclonal antibodies: implications for rapid diagnostic tests for malaria. J Clin Microbiol. 2006;44:2773–2778. doi: 10.1128/JCM.02557-05. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Berhane A, Russom M, Bahta I, Hagos F, Ghirmai M, Uqubay S. Rapid diagnostic tests failing to detect Plasmodium falciparum infections in Eritrea: an investigation of reported false negative RDT results. Malar J. 2017;16:105. doi: 10.1186/s12936-017-1752-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.President’s malaria initiative-FY 2014 Mozambique malaria operational plan. https://www.pmi.gov/docs/default-source/default-document-library/malaria-operational-plans/fy14/mozambique_mop_fy14.pdf.
- 9.Galatas B, Guinovart C, Bassat Q, Aponte JJ, Nhamússua L, Macete E, et al. A prospective cohort study to assess the micro-epidemiology of Plasmodium falciparum clinical malaria in Ilha Josina Machel (Manhiça, Mozambique) Malar J. 2016;15:444. doi: 10.1186/s12936-016-1496-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Ménard D, Khim N, Beghain J, Adegnika AA, Shafiul-Alam M, Amodu O, et al. A worldwide map of Plasmodium falciparum K13-propeller polymorphisms. N Engl J Med. 2016;374:2453–2464. doi: 10.1056/NEJMoa1513137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Gamboa D, Ho M, Bendezu J, Torres K, Chiodini P, Barnwell J, et al. A large proportion of P. falciparum isolates in the Amazon region of Peru lack pfhrp2 and pfhrp3: implications for malaria rapid diagnostic tests. PLoS ONE. 2010;5:e8091. doi: 10.1371/journal.pone.0008091. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Mayor A, Serra-Casas E, Bardají A, Sanz S, Puyol L, Cisteró P, et al. Sub-microscopic infections and long-term recrudescence of Plasmodium falciparum in Mozambican pregnant women. Malar J. 2009;8:9. doi: 10.1186/1475-2875-8-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Taylor SM, Mayor A, Mombo-Ngoma G, Kenguele HM, Ouédraogo S, Ndam NT, et al. A quality control program within a clinical trial Consortium for PCR protocols to detect Plasmodium species. J Clin Microbiol. 2014;52:2144–2149. doi: 10.1128/JCM.00565-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Bharti PK, Chandel HS, Ahmad A, Krishna S, Udhayakumar V, Singh N. Prevalence of pfhrp2 and/or pfhrp3 gene deletion in Plasmodium falciparum population in eight highly endemic states in India. PLoS ONE. 2016;11:e0157949. doi: 10.1371/journal.pone.0157949. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Wurtz N, Fall B, Bui K, Pascual A, Fall M, Camara C, et al. Pfhrp2 and pfhrp3 polymorphisms in Plasmodium falciparum isolates from Dakar, Senegal: impact on rapid malaria diagnostic tests. Malar J. 2013;12:34. doi: 10.1186/1475-2875-12-34. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Koita OA, Doumbo OK, Ouattara A, Tall LK, Konaré A, Diakité M, et al. False-negative rapid diagnostic tests for malaria and deletion of the histidine-rich repeat region of the hrp2 gene. Am J Trop Med Hyg. 2012;86:194–198. doi: 10.4269/ajtmh.2012.10-0665. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Amoah LE, Abankwa J, Oppong A. Plasmodium falciparum histidine rich protein-2 diversity and the implications for PfHRP 2: based malaria rapid diagnostic tests in Ghana. Malar J. 2016;15:101. doi: 10.1186/s12936-016-1159-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Hofmann N, Mwingira F, Shekalaghe S, Robinson LJ, Mueller I, Felger I. Ultra-sensitive detection of Plasmodium falciparum by amplification of multi-copy subtelomeric targets. PLoS Med. 2015;12:e1001788. doi: 10.1371/journal.pmed.1001788. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The datasets analyzed in this study are available from the corresponding author on request.


