Abstract
The initiating oncogenic event in almost half of human lung adenocarcinomas is still unknown, a fact that complicates the development of selective targeted therapies. Yet these tumours harbour a number of alterations without obvious oncogenic function including BRAF-inactivating mutations. Inactivating BRAF mutants in lung predominate over the activating V600E mutant that is frequently observed in other tumour types1. Here we demonstrate that the expression of an endogenous Braf(D631A) kinase-inactive isoform in mice (corresponding to the human BRAF(D594A) mutation) triggers lung adenocarcinoma in vivo, indicating that BRAF-inactivating mutations are initiating events in lung oncogenesis. Moreover, inactivating BRAF mutations have also been identified in a subset of KRAS-driven human lung tumours. Co-expression of Kras(G12V) and Braf(D631A) in mouse lung cells markedly enhances tumour initiation, a phenomenon mediated by Craf kinase activity2,3, and effectively accelerates tumour progression when activated in advanced lung adenocarcinomas. We also report a key role for the wild-type Braf kinase in sustaining Kras(G12V)/Braf(D631A)-driven tumours. Ablation of the wild-type Braf allele prevents the development of lung adenocarcinoma by inducing a further increase in MAPK signalling that results in oncogenic toxicity; this effect can be abolished by pharmacological inhibition of Mek to restore tumour growth. However, the loss of wild-type Braf also induces transdifferentiation of club cells, which leads to the rapid development of lethal intrabronchiolar lesions. These observations indicate that the signal intensity of the MAPK pathway is a critical determinant not only in tumour development, but also in dictating the nature of the cancer-initiating cell and ultimately the resulting tumour phenotype.
The RAS–MAPK signalling cascade serves as a central node in transducing signals from membrane receptors to the nucleus. This pathway is aberrantly activated in a substantial fraction of human cancers4. Moreover, germline mutations resulting in limited activation of this signalling cascade cause developmental disorders known as RASopathies5. There is also abundant evidence that elevated RAS–MAPK signalling results in cellular toxicity that may serve as a natural barrier to cancer progression early in tumorigenesis6. Finally, genetic abrogation of this pathway in adult mice results in their rapid death7. These findings suggest that defined thresholds of RAS–MAPK activity are required for homeostasis as well as for malignant transformation, but compelling genetic evidence is missing.
In order to augment MAPK signalling in controlled increments we have taken advantage of the expression of an endogenous Braf(D631A) kinase-dead isoform (corresponding to the human BRAF(D594A) mutant) that is known to induce Erk phosphorylation in a Craf-dependent manner2,8. This effect, known as the MAPK paradox, is due to enhanced heterodimerization and activation of the catalytically competent Craf protomer in Braf(D631A)–Craf complexes2,3. In agreement with these observations, lack of wild-type Braf expression in KrasG12V cell lines expressing Braf(D631A) increased the intensity and duration of MAPK signalling (Extended Data Fig. 1), probably as a result of the exclusive formation of Braf(D631A)–Craf heterodimers. Thus, to generate controlled thresholds of MAPK intensity in vivo, we combined wild-type Braf, conditional knockout Braf lox and conditional knock-in Braf LSLD631A with an inducible KrasLSLG12Vgeo allele9 (where LSL indicates a lox-STOP-lox motif). The resulting Kras+/LSLG12Vgeo (hereto designated as K), Kras+/LSLG12Vgeo;Braf +/LSLD631A (designated as KB) and Kras+/LSLG12Vgeo;Braf lox/LSLD631A (designated as KBL) strains were intratracheally infected with adenovirus expressing Cre recombinase (Ad-Cre). Cre-mediated recombination of these alleles results in the induction of distinct levels of Ras–MAPK signalling, with Braf +/+ driving lower activity, Braf +/D631A intermediate intensity and Braf −/D631A maximal activation. This strategy allowed us to investigate the effect of various MAPK activity thresholds on cell transformation, adenocarcinoma development and cellular toxicity in vivo.
Lung cells expressing these conditional alleles were identified by co-expression of the Kras(G12V) oncoprotein with β-geo, a chimaeric bacterial protein with β-galactosidase activity9 (Extended Data Fig. 2). As illustrated in Fig. 1a, small X-gal+ hyperplasias could be readily detected in the lungs of K mice 1 month after Ad-Cre infection. By constrast, Ad-Cre-infected KB mice displayed abundant hyperplastic areas together with adenomas, a lesion that is extremely infrequent in K mice at this early stage10. Remarkably, X-gal+ alveolar hyperplasias and adenomas were nearly absent in Ad-Cre-infected KBL animals. These mice, however, displayed hypertrophic bronchi as their most prominent feature (Fig. 1a). The lack of KrasG12V-driven alveolar lesions in KBL mice is unlikely to be a consequence of the absence of an active Braf, since this kinase is dispensable for the development of KrasG12V-driven lung adenocarcinoma7,11. Thus, we explored the possibility that the absence of alveolar lesions was due to some sort of cellular toxicity such as that caused by the induction of senescence or DNA damage, two phenomena known to act as barriers to tumour development6,12,13. Indeed, examination of lung lysates and sections from KBL mice detected p19Arf and p53 tumour suppressors, active caspase-3 and γ-H2AX, a marker for DNA damage, as early as 1 week following Ad-Cre infection (Fig. 1b, c). We did not observe differences in senescence-associated β-gal staining (data not shown). The concomitant increase in the phosphorylation of Erk1/2 and p90Rsk suggested that elevated MAPK signalling triggered a stress response that impaired tumour cell proliferation. Yet it has been recently described that mutant Kras-driven lung adenocarcinomas display a tonic activation of the DNA damage response to prevent the induction of genotoxic stress14. We hypothesize that the exclusive formation of Braf(D631A)–Craf complexes in KBL mice exceeded such a toxic signalling threshold and induced a stress response incompatible with tumour development. Thus, our results indicate that Ras-oncogenic toxicity is quantitatively dictated by MAPK function. In support of this hypothesis, whereas Mek inhibition prevented tumour growth in Ad-Cre-infected KB mice, it rescued the toxic phenotype in KBL animals in a dose-dependent manner and restored tumour progression at intermediate drug concentrations, most likely by curtailing MAPK activity to levels compatible with cell proliferation (Fig. 1d). This is in agreement with the observation that RAS-induced senescence in primary cells is bypassed by inhibition of MEK and ERK kinases15. These observations suggest that KrasG12V-driven lung tumour cells can only proliferate within a limited range of MAPK activity. Whereas excess MAPK activity induced DNA damage- mediated cellular toxicity, insufficient MAPK signalling cannot sustain tumour growth.
Figure 1. Combinations of KrasG12V, Braf D631A and wild-type Braf alleles establish a MAPK activity window that determines cell transformation and oncogene toxicity.
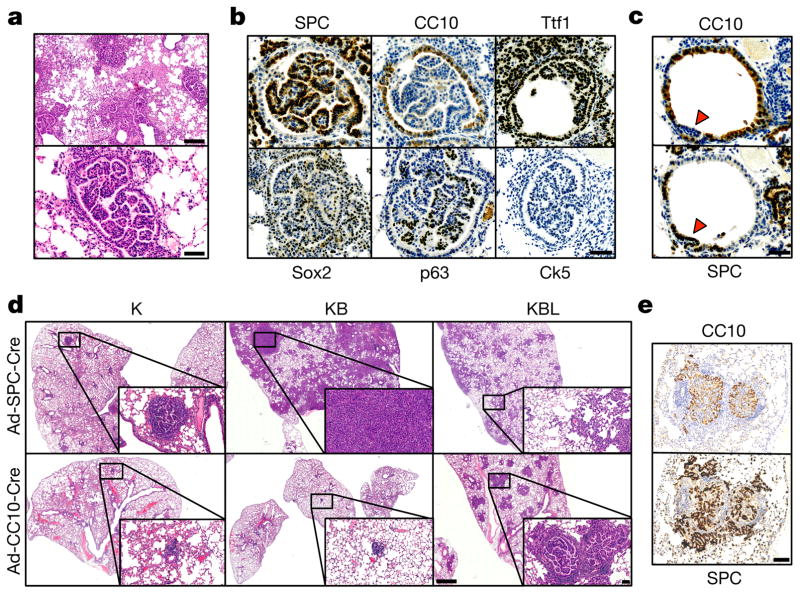
a, Whole-mount X-gal staining of representative lung sections (n = 5 per genotype) from Kras+/LSLG12Vgeo (K), Kras+/LSLG12VgeoBraf ; +/LSLD631A (KB) and Kras+/LSLG12VgeoBraf ; lox/LSLD631A (KBL) mice 1 month after Ad-Cre infection. X-gal staining identifies β-galactosidase expression as a surrogate marker for KrasG12V-expressing cells. Scale bar, 1mm. Insets show high-magnification images. Scale bar, 100 μm. b, Western blot analysis of lung lysates from Kras+/LSLG12Vgeo (K), Kras+/LSLG12Vgeo;Braf +/LSLD631A (KB) and Kras+/LSLG12Vgeo;Braf lox/LSLD631A (KBL) mice 1 week after Ad-Cre infection. Migration of p19ARF, p53, γ-H2AX, cleaved caspase-3 (C3A), p-Erk1/2, Erk1/2, p-p90Rsk and p90Rsk is indicated by arrowheads. Gapdh was used as loading control. Lysates from two independent animals per genotype are shown. c, Representative immunostaining of paraffin-embedded lung sections (n = 5 per genotype) from Kras+/LSLG12Vgeo (K), Kras+/LSLG12Vgeo;Braf +/LSLD631A (KB) and Kras+/LSLG12Vgeo;Braf lox/LSLD631A (KBL) mice 1 week after Ad-Cre infection using the indicated antibodies. Scale bar, 50 μm. d, Whole- mount X-gal staining of representative lung sections (n = 3 per genotype) from Kras+/LSLG12Vgeo (K), Kras+/LSLG12Vgeo;Braf +/LSLD631A (KB) and Kras+/LSLG12VgeoBraf ; lox/LSLD631A (KBL) mice 1 month after infection with 108 Ad-Cre particles. During this period, mice were treated with the indicated doses of the Mek inhibitor PD-0325901. The percentage of KrasG12V-expressing cells (X-gal-positive area) per lung section is indicated in each panel (top left, shown as mean ± s.d.). Scale bar, 200 μm. Insets in each panel show high-magnification images (top right; scale bar, 50 μm) or representative images of p-Erk1/2 immunostaining (bottom right; scale bar, 25 μm).
Next, cohorts of Ad-Cre infected K, KB and KBL mice were followed over time. In agreement with the earlier onset and the more rapid progression of the alveolar lesions, the KB animals displayed significantly shortened survival (Fig. 2a). Histopathological analysis of their lungs at 6 months post Ad-Cre infection revealed a 7.5-fold increase in tumour burden compared to K controls (Fig. 2b). Moreover, KB mice displayed advanced adenocarcinomas, a tumour stage that is extremely infrequent at this time in tumours driven by KrasG12V alone (Fig. 2c). Tumours present in KB mice displayed SPC+CC10− immunostaining, which suggests an alveolar type II (AT2) origin as previously described for adenocarcinomas driven by oncogenic Kras alone10,16 (Fig. 2d). Altogether, these observations suggest that MAPK hyperactivation by coexisting Kras(G12V) and Braf(D631A) mutations resulted in increased transformation of AT2 cells and accelerated tumour progression. The MAPK paradoxical activation model postulates that the observed tumour phenotype is mediated by Craf kinase activity2,8,17. To genetically validate this hypothesis in the lung tumours studied here, we added conditional knock-in Craf (also known as Raf1) kinase-dead alleles (Craf LSLD468A) to KB mice. The resulting strain, Kras+/LSLG12Vgeo;Braf +/LSLD631A;Craf LSLD468A/LSLD468A (designated as KBCKD) was used to determine whether genetic inhibition of the Craf kinase reverted the increased tumorigenic phenotype displayed by KB mice. Expression of the Craf(D468A) kinase-dead isoform led to a substantial decrease in the levels of phosphorylated (p-)Erk1/2 and overall tumour burden (Fig. 2e, f and Extended Data Fig. 3), significantly extending the survival of Ad-Cre-infected KBCKD mice compared to KB animals. Similarly, Craf ablation (KBCL mice) also resulted in prolonged survival (Fig. 2g). Altogether, these results demonstrate that the increased tumour burden and faster adenocarcinoma progression in KB animals is due to Craf-mediated hyper-activation of MAPK signalling.
Figure 2. Mice concomitantly expressing Kras(G12V) and Braf(D631A) show reduced survival and increased tumour development mediated by Craf kinase activity.
a, Survival of Kras+/LSLG12Vgeo (K, open circles, n =14), Kras +/LSLG12Vgeo;Braf +/LSLD631A (KB, solid circles, n =22) and Kras+/LSLG12Vgeo;Braf lox/LSLD631A (KBL, open triangles, n = 17) mice after Ad-Cre intratracheal infection. P < 0.0001, obtained using the log-rank test (Mantel–Cox). b, Quantification of average tumour (adenoma and adenocarcinoma) burden (per cent of total lung area) in lung sections from Kras+/LSLG12Vgeo (K, n =5), Kras+/LSLG12Vgeo;Braf +/LSLD631A (KB, n =7) and Kras+/LSLG12Vgeo;Braf lox/LSLD631A (KBL, n = 7) mice 6 months after Ad-Cre intratracheal infection. **P <0.01, *P <0.05. c, Representative haematoxylin and eosin staining of paraffin-embedded lung sections obtained from Kras+/LSLG12Vgeo (K, n =5), Kras+/LSLG12Vgeo;Braf +/LSLD631A (KB, n =7) and Kras+/LSLG12Vgeo;Braf lox/LSLD631A (KBL, n = 7) mice 6 months after Ad-Cre infection. Scale bar, 5 mm. d, Representative immunostaining of paraffin-embedded sections showing tumours from Ad-Cre-infected Kras+/LSLG12Vgeo;Braf +/LSLD631A (KB, n = 7) mice killed at humane end point using antibodies against SPC and CC10. Scale bar, 1 mm. e, Quantification of average tumour (adenoma and adenocarcinoma) burden (per cent of total lung area) in lung sections from Kras+/LSLG12Vgeo;Braf +/LSLD631A (KB, n = 7) and Kras+/LSLG12VgeoBraf ; +/LSLD631ACraf ; LSLD468A/LSLD468A (KBCKD, n = 5) mice 6 months after Ad-Cre intratracheal infection. **P <0.01. f, Representative haematoxylin and eosin staining of paraffin-embedded lung sections (n = 5 per genotype) from Kras+/LSLG12Vgeo;Braf +/LSLD631A (KB), and Kras+/LSLG12Vgeo;Braf +/LSLD631A;Craf LSLD468A/LSLD468A (KBCKD) mice 6 months after Ad-Cre infection. Scale bar, 5 mm. g, Survival of Kras+/LSLG12Vgeo;Braf +/LSLD631A (KB, solid circles, n =22), Kras+/LSLG12Vgeo;Braf +/LSLD631ACraf ; LSLD468A/LSLD468A (KBCKD, solid triangles, n = 12) and Kras+/LSLG12Vgeo;Braf +/LSLD631A;Craf lox/lox (KBCL, empty squares, n = 7) mice after Ad-Cre intratracheal infection. P <0.0001, obtained using the log-rank test (Mantel–Cox).
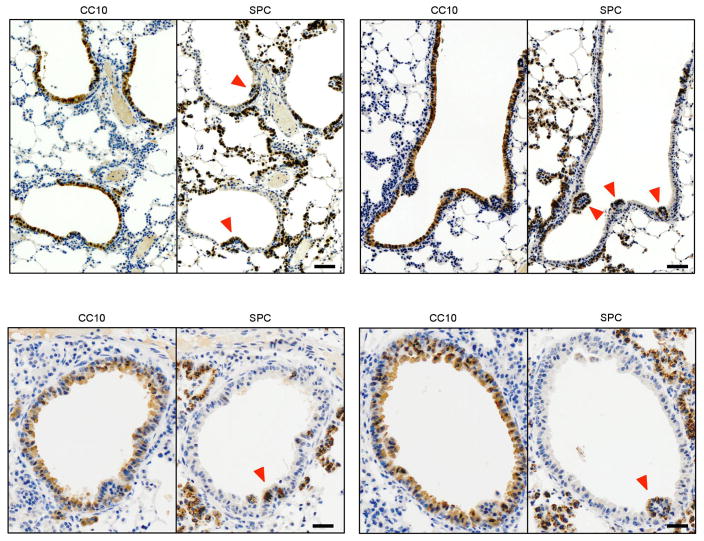
As indicated above, the tumour burden of the KBL cohort was significantly decreased compared to KB mice, suggesting that excessive MAPK activity in the absence of wild-type Braf expression may be detrimental for lung adenocarcinoma development. Yet, in spite of the reduced tumour burden, KBL animals reached humane end point at the same time as KB mice (Fig. 2a, b). Detailed examination of the lungs of KBL mice revealed the presence of intrabronchiolar carcinomas (112 of 250 bronchi, 45%), a rare lesion in K (0 of 162) or KB (20 of 145, 14%) cohorts (Fig. 3a). Immunostaining with a collection of markers including Ttf1 and SPC, characteristic of lung adenocarcinoma and Sox2, p63 and CK5, diagnostic of squamous cell carcinoma, did not clarify the origin of these intrabronchiolar lesions since the resulting expression pattern was not in accordance with the expected profile of either of these tumour types18 (Fig. 3b). Analysis of the initial tumour stages of these lesions revealed protruding papillary structures accompanied by loss of CC10 expression, a marker characteristic of the bronchiolar epithelium. Most of these lesions (95%) acquired expression of the AT2 marker SPC, thus suggesting a transdifferentiation process (Fig. 3c and Extended Data Fig. 4).
Figure 3. Hyperactive MAPK signalling triggers bronchiolar carcinoma by transdifferentiation of club cells.
a, Haematoxylin and eosin staining of paraffin-embedded sections showing the presence of intrabronchiolar carcinoma in Ad-Cre-infected Kras+/LSLG12Vgeo;Braf lox/LSLD631A (KBL) mice killed at humane end point. Two different magnifications are shown. Scale bar, 200 μm (top) and 50 μm (bottom). b, Representative immunostaining of paraffin-embedded sections showing intrabronchiolar tumours from Ad-Cre-infected Kras+/LSLG12Vgeo;Braf lox/LSLD631A (KBL, n = 5) mice using antibodies against, SPC, CC10, Ttf1, Sox2, p63 and Ck5. Scale bar, 50 μm. c, Immunostaining of paraffin-embedded consecutive sections showing early intrabronchiolar lesions (1 week after Ad-Cre infection) in Kras+/LSLG12Vgeo;Braf lox/LSLD631A (KBL) mice using antibodies against SPC and CC10. Arrowheads indicate protruding bronchiolar papillary growth undergoing transdifferentiation. Scale bar, 50 μm. d, Representative haematoxylin and eosin staining of paraffin-embedded lung sections (n = 3 per genotype) from Kras+/LSLG12Vgeo (K), Kras+/LSLG12Vgeo; Braf +/LSLD631A (KB) and Kras+/LSLG12Vgeo;Braf lox/LSLD631A (KBL) mice killed 2 months after infection with cell-type-restricted Ad-Cre viruses that selectively induce Cre-mediated recombination in AT2 (Ad-SPC-Cre) (top) and club cells (Ad-CC10-Cre) (bottom). Scale bar, 500 μm. Insets show high-magnification images. Scale bar, 100 μm. e, SPC and CC10 immunostaining of paraffin-embedded consecutive sections showing intrabronchiolar tumours from Kras+/LSLG12Vgeo;Braf lox/LSLD631A (KBL) mice infected with lineage-specific Ad-CC10-Cre. Scale bar, 100 μm.
To further clarify the origin of these intrabronchiolar lesions we performed in vivo tracing experiments by infecting K, KB and KBL mice with lineage-specific Ad-CC10-Cre or Ad-SPC-Cre viruses that restrict Cre-mediated recombination to club or AT2 cells respectively19. Analysis of lungs of K and KB mice after Ad-SPC-Cre infection revealed the presence of adenomas that, as expected, were larger and more abundant in KB animals. Likewise, KBL mice predominantly developed alveolar hyperplasias that failed to progress to advanced stages. Importantly, infection with Ad-CC10-Cre did not induce tumour growth in K or KB mice but caused abundant SPC+ intrabronchiolar carcinomas in KBL animals (Fig. 3d, e). These results indicate that the threshold of MAPK activity toxic for AT2 cells resulted in the efficient transformation of cells present in bronchial epithelium that are usually refractory to KrasG12V-driven oncogenesis10,20,21. Altogether, these results suggest a quantitative impact of MAPK activity controlling various aspects of Kras oncogene-driven lung carcinogenesis. As such, oncogenic transformation may critically depend on cell- or tissue-specific programs that are hijacked to adjust MAPK activity and avoid senescence or other tumour-suppressive stress responses. This quantitative model is not exclusive to MAPK signalling as a similar threshold response level has been proposed for Wnt/β-catenin-driven22 or PI3K/AKT-driven tumorigenesis23.
Mutational analysis of different human cancers has recently uncovered that among the BRAF hot spots in lung adenocarcinoma, those resulting in inactivating mutations predominate over the V600E activating substitution1. However, the contribution of BRAF-inactive mutants to lung cancer progression is unclear. Interestingly, a percentage of BRAF-inactivating mutations (11%) coexist with upstream RAS alterations (Table 1). Thus, we decided to investigate whether expression of the Braf(D631A) kinase-inactive mutant in pre-existing KrasG12V-driven tumours may enhance tumour progression. To this end, we added the Braf LSLD631A allele to a lung tumour model driven by the Flp recombinase. The resulting strain, Kras+/FSFG12V; Tg.hUb-cre-ERT2+/T;Braf +/LSLD631A (designated as KFB) allows the temporal separation of tumour initiation (Flp-mediated Kras(G12V) expression) from genetic events induced during tumour progression (Cre-mediated Braf(D631A) expression). Induction of the Braf(D631A) kinase-inactive isoform in pre-existing KrasG12V-driven tumours resulted in reduced survival owing to the accelerated progression of these lesions (Fig. 4a and Extended Data Fig. 5). Thus, these results provide an experimental explanation for the concurrence of oncogenic KRAS and BRAF-inactivating mutations in certain human lung tumours. These observations also reinforce the current clinical indication that class I RAF inhibitors (vemurafenib and derivatives targeting the activated form of the BRAF kinase) should not be used in patients with KRAS-mutant lung adenocarcinoma.
Table 1.
BRAF hypoactive mutants with NF1 and RAS mutations
| Human skin cutaneous melanoma | |||||
|---|---|---|---|---|---|
|
| |||||
| Sample | BRAF | NF1 | KRAS | NRAS | HRAS |
| TCGA-EE-A29N-06 | D594N | Y80* | K117N | – | – |
| TCGA-EE-A2GJ-06 | G466E | – | – | G12D | – |
| TCGA-EE-A2GC-06 | G466E | Q2239* | – | – | – |
| TCGA-EE-A19F-06 | G466E | X69_splice, L274F | – | – | – |
| TCGA-EE-A1ZZ-06 | N581S | Q684*, S1599F | – | P34L | – |
| TCGA-EE-A181-06 | S467L | Q519*, R440*, L2604F | – | – | – |
| TCGA-EE-A3JD-06 | S467L | P1851S, R1276Q | – | – | – |
| Human lung adenocarcinoma | |||||
| Sample | BRAF | NF1 | KRAS | NRAS | HRAS |
| TCGA-05-5428-01 | D594N | – | – | – | – |
| TCGA-50-5044-01 | D594H | – | – | – | – |
| TCGA-05-4410-01 | G466V | – | – | – | – |
| TCGA-78-7537-01 | G466V | – | – | – | – |
| TCGA-78-7153-01 | G466A | – | – | – | – |
| TCGA-44-6744-01 | N581S | – | – | – | |
| TCGA-44-2662-01 | N581S | – | – | – | – |
| TCGA-05-4249-01 | A762E | – | G12C | – | – |
| TCGA-55-6979-01 | P367R | – | – | – | – |
The co-occurrence of BRAF hypoactive mutants with NF1 and RAS mutations in human cancer datasets. Data from skin cutaneous melanoma and lung adenocarcinoma patients obtained from The Cancer Genome Atlas cutaneous melanoma and lung adenocarcinoma studies, respectively. Sample ID as well as the precise mutations identified in these patients are indicated. Asterisks indicate that the mutation creates a STOP codon, and therefore a truncated protein. X69_splice represents a mutation affecting the splice site adjacent to amino acid 69.
Figure 4. Activation of endogenous Braf D631A results in lung adenocarcinoma development.
a , Kras+/FSFG12V;Braf +/LSLD631A; Tg.hUb-cre-ERT2+/T (KFB) and control Kras+/FSFG12V;Tg.hUb-cre-ERT2+/T (KF) mice were infected intratracheally with Ad-Flp and tumour formation was monitored by computed tomography (CT). Mice bearing tumours visible on CT (KFB, n = 29 tumours from 14 mice and KF, n =38 tumours from 15 mice) were fed ad libitum a tamoxifen-containing diet resulting in expression of the Braf(D631A) kinase-dead isoform in KFB. Tumour volume increase (measured as fold change) was re-evaluated by CT after 8 weeks in continuous diet. **P <0.01. b, Haematoxylin and eosin (H&E) staining, as well as SPC and CC10 immunostaining, of paraffin-embedded lung sections from Ad-Cre-infected Braf +/LSLD631A mice killed at humane end-point. Scale bar, 500 μm and 50 μm (top right). c, Representative images of p-Erk1/2 immunostaining of paraffin-embedded lung sections (n = 3 per genotype) from Ad-Cre-infected Kras+/LSLG12Vgeo, Kras+/LSLG12Vgeo;Trp53 lox/lox, Braf +/LSLV637E (equivalent to human BRAFV600E) or Braf +/LSLD631A mice. Scale bar, 100 μm.
Yet, the majority of BRAF-inactivating mutations present in human lung cancer (89%) do not coexist with RAS mutations (Table 1). To assess whether Braf inactive mutants could induce lung adenocarcinoma formation in the absence of Kras mutations, we infected Braf +/LSLD631A mice intratracheally with Ad-Cre. Analysis of their lungs 12 months after infection revealed the presence of tumours in 9 of 22 mice (41% incidence) compared to 14 of 18 (78%) in Kras+/LSLG12Vgeo animals. All tumours studied displayed histology characteristic of lung adenocarcinoma with SPC+CC10− immunostaining (Fig. 4b and Extended Data Fig. 6a). Notably, p-Erk1/2 levels in Braf D631A-driven lung adenocarcinomas were higher than those observed in Braf V637E (equivalent to human BRAFV600E), KrasG12V or KrasG12V;Trp53−/− tumours (Fig. 4c). As control, we confirmed that the Braf LSLD631A allele was efficiently recombined in tumour tissue. Moreover, we determined that there were no Ras mutations in these tumours (data not shown).
These results suggest that Braf-inactivating mutations initiate lung adenocarcinoma (Extended Data Fig. 6b). Our results are in good agreement with data from the Rosen laboratory indicating that increased levels of wild-type Ras–GTP complexes can cooperate with Braf hypoactive mutants to trigger tumour development in epithelial cells24. Similarly, increasing the pool of Ras–GTP by a dominant active Sos is sufficient to induce MAPK hyperactivation and the formation of epithelial tumours in response to Raf inhibition25. Likewise, MAPK activation in primary keratinocytes carrying the Braf LSLD631A allele depended both on Craf and RTK signalling. Of note, elimination of the Braf wild-type allele induced a further increase in p-Erk1/2 (Extended Data Fig. 6c). In lung, the high levels of endogenous Ras–GTP present in adult AT2 cells26 may explain the oncogenicity of the Braf(D631A) inactive mutant in the absence of Kras(G12V). By contrast, hypoactive BRAF mutants require the presence of RAS or NF1 mutations to trigger melanoma development, both in experimental GEM models2 as well as in human tumours (Table 1).
In summary, we provide the first genetic evidence demonstrating that a kinase-inactivating Braf mutation induces lung adenocarcinoma development. Importantly, in lung adenocarcinoma patients kinase- inactivating BRAF mutations are more prevalent than the activating V600E allele1. Furthermore, a recent retrospective study identified BRAF(D594G) (affecting the same residue as the mouse mutation used in this study) as the most frequent BRAF alteration in these patients27. Our analysis of human lung adenocarcinomas with hypoactive BRAF revealed co-occurring alterations such as mutations in RTK signalling antagonists that might cooperate in sustaining MAPK activity (Extended Data Fig. 7). The accompanying manuscript by Yao et al.24 suggests that RTK profiling of tumours driven by kinase-impaired BRAF mutants can identify the dominant receptor for the design of specific treatment for individual patients. Alternatively, our results suggest that these patients could benefit from therapies based on selective CRAF inhibitors. The mouse model described here will be useful to evaluate effective drug combinations.
METHODS
Mice
KrasLSLG12Vgeo (ref. 9), Braf LSLD631A (ref. 2, described therein as Braf LSLD594A), Braf lox (ref. 28), Braf LSLV637E (ref. 29, described therein as Braf CA) and Trp53lox/lox (ref. 30) strains have been previously described. The targeting vector for the Craf LSLD468A allele was generated by Gene Bridges GmBH. The D468A miscoding mutation (GAC (Asp) to GCC (Ala)) was engineered within the Craf locus present in the BAC RP23-37K24 by multi-site Red/ET (‘triple recombination’). A loxP-cDNA-STOP-PGK-Neo-loxP cassette containing a partial Craf cDNA sequence encompassing exons 13 to 17 was inserted 150 bp upstream of the 5′ end of the mutated exon13 by Red/ET recombination. The resulting targeting vector was linearized with NotI and SalI restriction enzymes and electroporated into B6129SF1/J ES cells. Clones having undergone proper homologous recombination were identified by Southern blot analysis. Two independent recombinant ES cell clones were microinjected into FVB donor blastocysts and implanted into pseudo-pregnant females. Chimaeric mice were backcrossed to C57BL/6J mice and germline transmission of the Craf LSLD468A allele was confirmed by Southern blot analysis. The targeting vector used to obtain the KrasFSFG12V allele was generated by Taconic Artemis. In brief, the homology arms including the first exon containing the oncogenic G12V mutation were amplified by PCR using as a template a targeting vector previously developed to generate a KrasLSLG12Vgeo allele9. A PGK-Neo-STOP cassette was generated by PCR amplification of the 1,377 bp STOP cassette derived from the KrasLSLG12Vgeo targeting vector with primers that incorporated NdeI restriction sites. The STOP cassette was subsequently cloned into the NdeI restriction site of pBASIC10 (Taconic Artemis) flanked by FRT sequences. The resulting targeting vector was linearized with NotI and electroporated into B6129SF1/J ES cells. Two independent recombinant ES cell clones were microinjected into C57BL/6J blastocysts and transplanted into pseudo-pregnant females. Chimaeric mice were backcrossed to C57BL/6J mice and germ line transmission of the targeted allele was confirmed by Southern blot analysis. All animal experiments were approved by the Ethical Committee of CNIO and performed in accordance with the guidelines stated in the International Guiding Principles for Biomedical Research Involving Animals, developed by the Council for International Organizations of Medical Sciences (CIOMS). All strains were genotyped by Transnetyx.
Tumour induction and drug treatments
Tumours were induced in 8- to 12-week-old mice (both male and female animals were used) by single intratracheal infection with 106 adenoviral particles (unless stated otherwise) after anaesthesia (i.p. injection of ketamine 75 mg kg−1, xylazine 12 mg kg−1) as previously reported7. Ad-Cre, Ad-CC10-Cre, Ad-SPC-Cre and Ad-Flp were purchased from University of Iowa Vector Core Facility. The Mek inhibitor PD-0325901 (Tocris Bioscience) was dissolved in 0.5% hydroxypropyl methyl cellulose, 0.2% Tween-80 water solution and administered by daily gavage. Following the ethical committee guidelines all mice were euthanized when showing respiratory problems.
Histopathology and immunohistochemistry
For routine histological analysis, all lung lobes from each mouse were fixed in 10% buffered formalin (Sigma), embedded in paraffin and evaluated in serial sections by conventional haematoxylin and eosin (H&E) staining according to previously published criteria31. All whole mount X-gal-stained sections were counterstained with Nuclear Fast Red. Antibodies used for immunostaining included those raised against: SPC (Millipore, AB3786); CC10 (Santa Cruz Biotechnology, SC-9772); Ttf1 (Epitomics, 2044-1); Sox2 (Cell Signaling Technology, 3728); p63 (Thermo Scientific, MS-1081-P); Ck5 (Covance, PRB-160P) and p-Erk1/2 (Cell Signaling Technology; 9101).
Cell culture
Kras+/G12V;Araf−/−;Braf lox/lox;Craf lox/lox;Trp53−/−;Tg.hUb-cre-ERT2+/T lung adenocarcinoma cell lines were generated from primary tumours. Cells were infected with lentiviruses expressing Braf, Braf(D631A) and Craf (pLVXpuro) using routine procedures and exposed to 50 MOI (multiplicity of infection) of Ad-Cre. 4-hydroxytamoxifen (4-OHT) (Sigma) was used at 600 nM. Cell lines were confirmed mycoplasma free at every freeze/thaw cycle. Primary keratinocytes were obtained from adult tail skin. The antibodies used for western blotting included those raised against: Araf (Cell Signaling Technology, 4432); Braf (Santa Cruz Biotechnology, SC-5284); Craf (BD Biosciences, 610151); p-Erk1/2 (Cell Signaling Technology, 9101); Erk1 (BD Biosciences, 554100); Erk2 (BD Biosciences, 610103); γ-H2AX (Millipore, 05-636); p-p90Rsk (Cell Signaling Technology, 9341); p90Rsk (Santa Cruz Biotechnology, SC-231); p53 (Cell Signaling Technology, 2524); p19ARF (Abcam, ab80); cleaved caspase-3 (Cell Signaling Technology, 9661) and Gapdh (Sigma, G8795).
Computed tomography
Mice were anaesthetized with 1% to 3% flow of isoflurane/oxygen, and the chest area was visualized with the GE eXplore Locus micro PET-CT scanner (GE Healthcare). The resulting raw data were reconstructed to a final image volume of 875 ×875 × 465 slices at 93 μm3 voxel dimensions. Reconstructed slices were output in the manufacturer’s raw format and corrected equal to Hounsfield units and analysed with MicroView analysis software (GE Healthcare).
Digital image quantification
Digital images of immunostained slides were obtained using a whole slide scanner (Dotslide Olympus) with resolution 0.32 μm per pixel (20×/NA 0.75). For automated image quantification the tumour areas on scanned H&E sections were manually delimited within the normal lung tissue according to histological criteria and quantified by Dotslide viewer software.
Statistical and data analysis
The product limit method of Kaplan and Meier was used for generating the survival curves, which were compared by using the log-rank (Mantel–Cox) test. A P value that was less than 0.05 was considered statistically significant for all datasets. All statistical analysis was performed using GraphPad Prism software. No statistical method was used to predetermine sample size in animal studies. Only mice carrying tumours >1 mm in diameter on CT were enrolled in longitudinal studies assessing tumour volume variation. For all animal experiments block randomization was used to ensure a balance in sample size across groups over time. The investigators were blinded during evaluation of tumour size variations following tamoxifen diet.
Data availability
All data generated or analysed during this study are included in this published article (and its Supplementary Information files).
Extended Data
Extended Data Figure 1. Braf(D631A)-dependent activation of MAPK signalling in vitro.
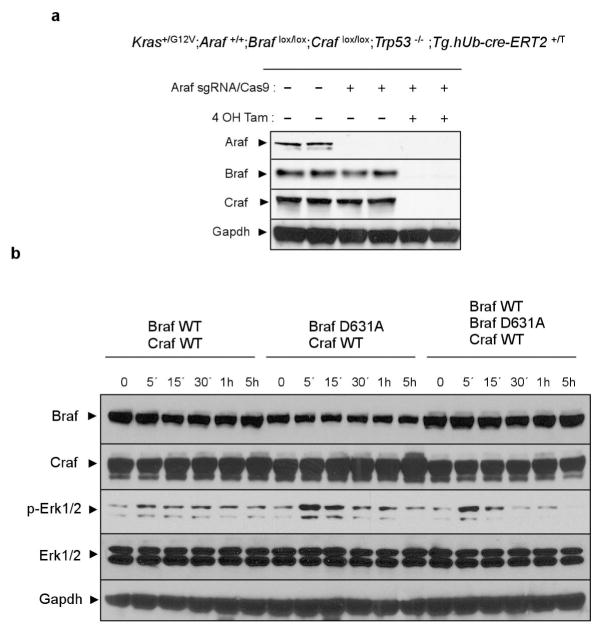
a, Western blot analysis of the expression levels of the three Raf isoforms in lysates derived from Kras+/G12V;Araf+/+;Braf lox/lox; Craf lox/lox;Trp53−/−;Tg.hUb-cre-ERT2+/T lung adenocarcinoma cell lines. The endogenous Araf alleles were eliminated using CRISPR–Cas9 editing where indicated. Braf lox and Craf lox alleles were eliminated by addition of 4-hydroxytamoxifen (4-OHT) to induce their Cre-mediated recombination where indicated. Gapdh is shown as a loading control. b, Kras+/G12V;A-Raf−/−;Braf lox/lox;Craf lox/lox;Trp53−/−;Tg.hUb-cre-ERT2+/T lung adenocarcinoma cell lines were infected with lentiviral particles expressing combinations of wild-type Braf, kinase-inactive Braf(D631A) and wild-type Craf as indicated before the addition of 4-hydroxytamoxifen to induce elimination of the endogenous conditional Braf and Craf alleles. Cells were serum-starved for 24 h and re-stimulated with serum for the indicated times. The activation of MAPK signalling was assessed by western blot analysis of p-Erk1/2. Expression levels of the exogenous Braf and Craf alleles as well as endogenous Erk1/2 and Gapdh are shown as loading controls. For gel source data, see Supplementary Fig. 1.
Extended Data Figure 2. Mouse alleles and strains used in the study.
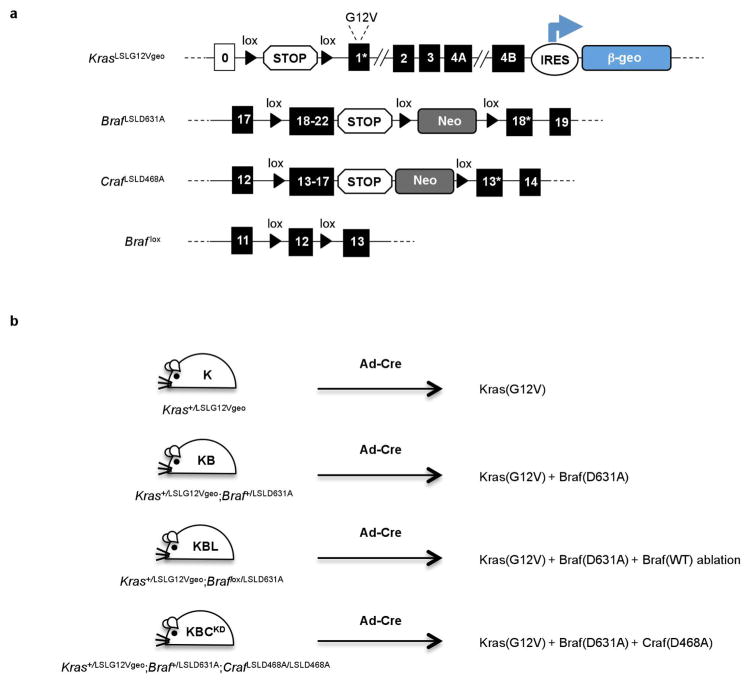
a, Expression of the endogenous KrasG12V oncogene is achieved by Cre-mediated excision of the transcriptional STOP cassette preceding the KrasLSLG12Vgeo allele. Cells expressing the oncogene can be identified by X-gal staining owing to the co-expression of a bicistronic β-galactosidase (β-geo) reporter. Likewise, Cre-mediated excision of the lox-STOP-lox cassette (containing the indicated wild-type exons) induces the expression of the kinase-dead Braf(D631A) and Craf(D468A) mutants instead of the wild-type isoforms. Finally, Cre-mediated excision of the conditional Braf lox allele results in the elimination of the wild-type Braf protein. b, Schematic representation of the proteins expressed in lung cells following intratracheal infection with Ad-Cre virus in the compound strains used in the study.
Extended Data Figure 3. Activation of p-Erk1/2 in lung adenocarcinomas driven by concomitant expression of Kras(G12V) and Braf(D631A) depends on Craf kinase activity.
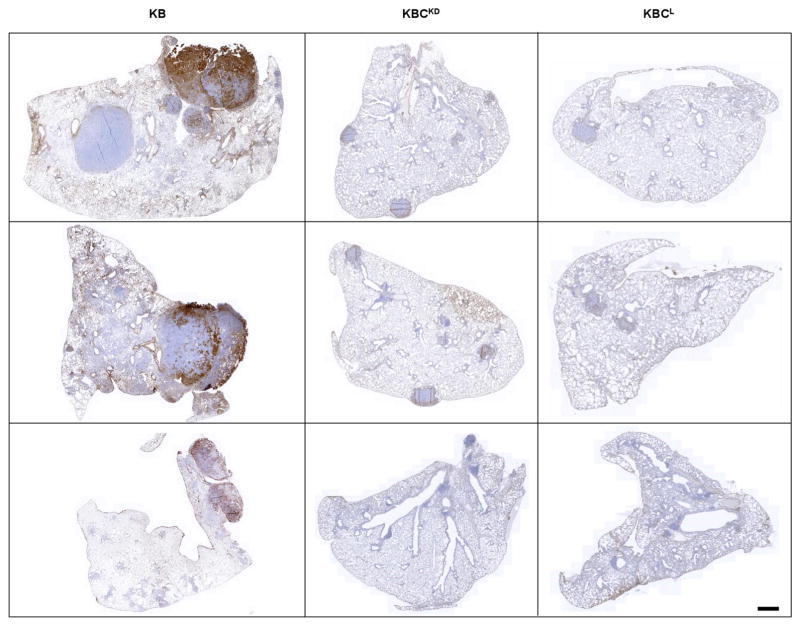
Representative images of p-Erk1/2 immunostaining of paraffin-embedded lung sections (n =3 per genotype) from Ad-Cre-infected Kras+/LSLG12Vgeo;Braf +/LSLD631A (KB), Kras+/LSLG12Vgeo;Braf +/LSLD631A;Craf LSLD468A/LSLD468A (KBCKD) and Kras+/LSLG12Vgeo;Braf +/LSLD631A;Craf lox/lox (KBCL). Tumour samples were collected 6 months after Ad-Cre intratracheal infection coincident with the humane end point of the KB strain. Scale bar, 1 mm.
Extended Data Figure 4. MAPK hyperactivation induces transdifferentiation of the bronchiolar epithelium leading to papillary carcinoma.
Immunostaining of paraffin-embedded sections showing transdifferentiation of the bronchiolar epithelium at early stages (1 week) after Ad-Cre infection of Kras+/LSLG12Vgeo;Braf lox/LSLD631A (KBL). Images display staining using antibodies against CC10 (club cell marker, left panels) and SPC (AT2 marker, right panels). Protruding bronchiolar papillary growth is invariably associated with a transdifferentiation process illustrated by the acquisition of SPC+ staining (red arrowheads). Scale bar, 100 μm (top panels), 50 μm (bottom panels).
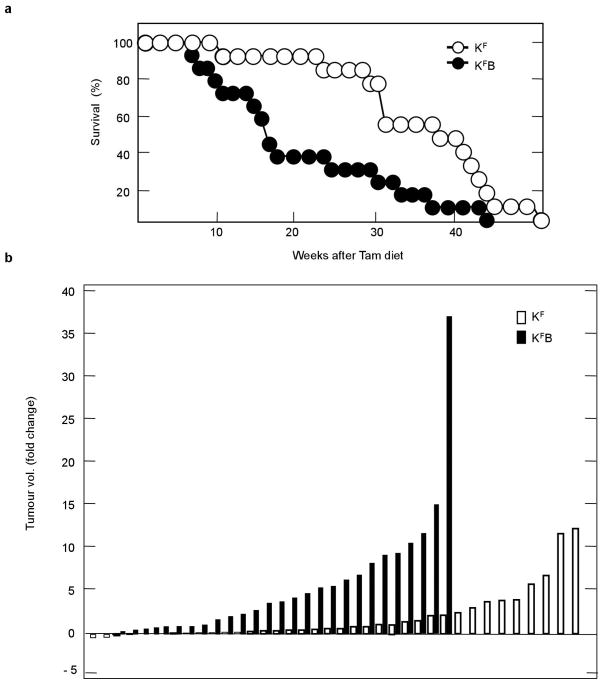
Extended Data Figure 5. Expression of an endogenous Braf(D631A) inactive mutant allele in mice harbouring KrasG12V-driven lung adenocarcinomas detectable by CT increases tumour growth and reduces survival.
a, Survival of Kras+/FSFG12V;Braf +/LSLD631A;Tg.hUb-cre-ERT2+/T (KFB, solid circles n = 14) and control Kras+/FSFG12V;Tg.hUb-cre-ERT2+/T (KF, empty circles n = 13) mice bearing lung adenocarcinomas detectable by CT. Upon tumour detection mice were maintained on a tamoxifen-containing diet to activate expression of the Braf(D631A) kinase-dead isoform. P < 0.0058, obtained using the log-rank test (Mantel–Cox). b, Waterfall plot representation of the tumour volume increase (measured as fold change) when re-evaluated by CT after 8 weeks of continuous tamoxifen-containing diet in Kras+/FSFG12V;Braf +/LSLD631A;Tg.hUb-cre-ERT2+/T (KFB, n = 29 tumours from 14 mice) and control Kras+/FSFG12V; Tg.hUb-cre-ERT2+/T (KF, n = 38 tumours from 15 mice) cohorts. The same dataset is represented in Fig. 4a.
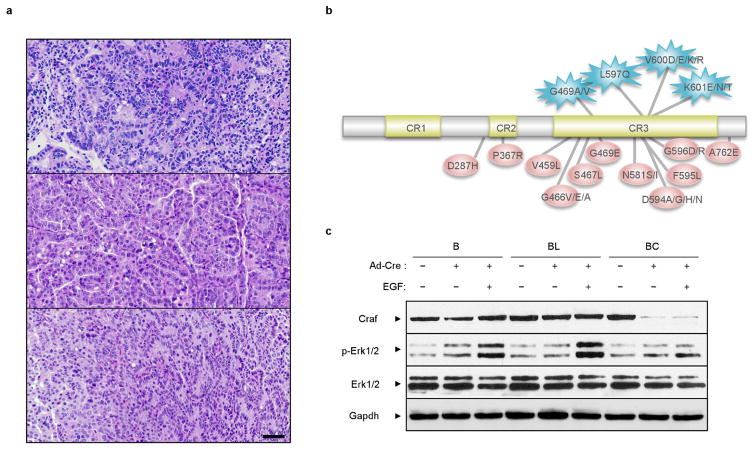
Extended Data Figure 6. Histology of Braf(D631A)-driven lung adenocarcinoma and evaluation of Braf(D631A)-dependent activation of MAPK signalling in vitro.
a, Haematoxylin and eosin staining of paraffin-embedded lung sections from Ad-Cre-infected Braf +/LSLD631A mice killed at humane end point. Sections display moderately to poorly circumscribed tumours occupying nearly an entire lung lobe, with compressed adjacent lung parenchyma and peripherally infiltrated by mononuclear inflammatory cells, mainly macrophages. Tumour masses are composed by papillary outgrowths or less-differentiated areas with increased cellular pleomorphism. Images correspond to three independent mice. Scale bar, 100 μm. b, Schematic representation of BRAF mutations detected in human tumours with high (top) or low/absent (bottom) kinase activity when compared to the wild-type isoform (see also the accompanying manuscript by Yao et al.24). c, Primary keratinocytes derived from Braf +/LSLD631A (B), Braf lox/LSLD631A (BL) and Braf +/LSLD631A; Craf lox/lox (BC) mice were infected with Ad-Cre (5 MOI) and treated with recombinant EGF (150 ng ml−1 for 1 h) where indicated. The activation of MAPK signalling was assessed by western blot analysis of p-Erk1/2, demonstrating that the activation of MAPK signalling by Braf(D631A) in epithelial cells requires upstream RTK activation and is enhanced by elimination of the wild-type Braf allele. Efficient recombination of the Craf conditional alleles in Ad-Cre-infected BC cells is also shown, demonstrating that MAPK activation by Braf(D631A) is Craf-dependent. Expression levels of Erk1/2 and Gapdh are shown as loading controls. For gel source data, see Supplementary Fig. 1.
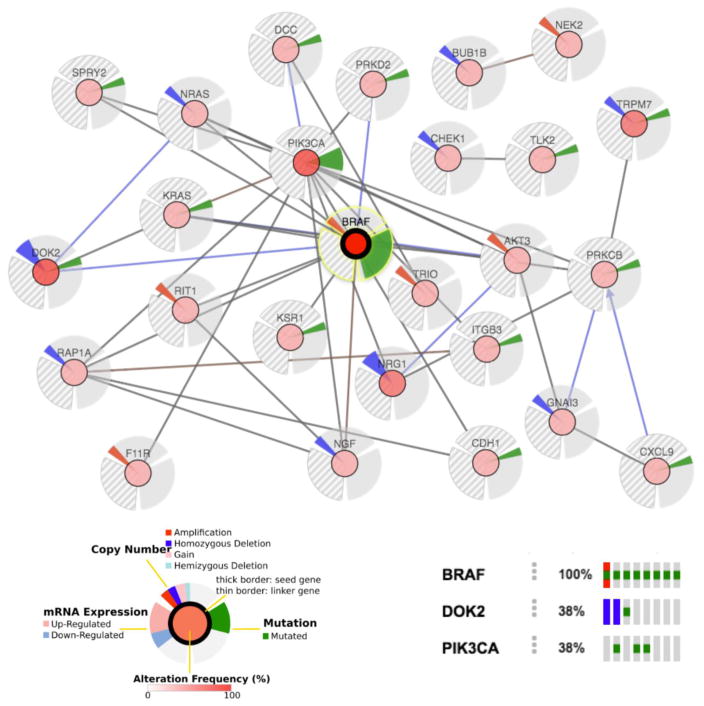
Extended Data Figure 7. Co-occurring alterations in lung adenocarcinoma patients with BRAF-inactivating mutations.
Whole-genome sequencing data available from the TCGA LUAD study at cBioportal (http://www.cbioportal.org) was used to generate a gene network for the analysis of genetic alterations coincident with BRAF hypoactive mutations. PIK3CA mutations (Q296E, E542K and E545K) were the most frequent co-occurring event. In addition, mutations or gene deletions in known RTK signalling antagonists (DOK2, SPRY2), MAPK scaffolds (KSR1), members of the RAS subfamily (RAP1A, RIT1) or the guanine nucleotide exchange factor TRIO might potentially cooperate with inactive BRAF to sustain MAPK activity.
Supplementary Material
Acknowledgments
We thank A. de Martino for histopathological evaluation of murine lung tumours. This work was supported by grants to M.B. from the European Research Council (ERC-AG/250297-RAS AHEAD), EU-Framework Programme (HEALTH-F2-2010-259770/LUNGTARGET and HEALTH-2010-260791/EUROCANPLATFORM) and Spanish Ministry of Economy and Competitiveness (SAF2011-30173 and SAF2014-59864-R). M.B. is the recipient of an Endowed Chair from the AXA Research Fund. Funding was also provided by grants to N.R. from the National Institutes of Health (P01 CA129243; R35 CA210085); the Commonwealth Foundation for Cancer Research, the Center for Experimental Therapeutics at Memorial Sloan Kettering Cancer Center and the Stand Up To Cancer – American Cancer Society Lung Cancer Dream Team Translational Research Grant (SU2C-AACR-DT17-15). Support was also received from the NIH MSKCC Cancer Center Support Grant P30 CA008748. Work in the laboratory of R.C. was supported by grants FP7 ERC-2009-StG (242965-Lunely) and Associazione Italiana per la Ricerca sul Cancro (AIRC) grant IG-12023. P.N. was the recipient of an FPU fellowship from the Spanish Ministry of Education. C.A. was the recipient of a postdoctoral fellowship from the Spanish Association Against Cancer (AECC).
Footnotes
Online Content Methods, along with any additional Extended Data display items and Source Data, are available in the online version of the paper; references unique to these sections appear only in the online paper.
Supplementary Information is available in the online version of the paper.
Author Contributions D.S. and M.B. designed experiments and research aims, analysed data and wrote the manuscript with help from co-authors. P.N. performed experiments and analysed the data with help from C.A., L.E. and M.T.B. R.C. carried out critical interpretation of the tumour phenotype. R.M. provided the Braf+/LSLD631A strain. Z.Y., N.R, R.M. and R.C. contributed critical information and helpful discussions. D.G.P. and G.G.-L. performed the bioinformatic analysis of human datasets.
The authors declare competing financial interests: details are available in the online version of the paper.
Readers are welcome to comment on the online version of the paper.
References
- 1.Chang MT, et al. Identifying recurrent mutations in cancer reveals widespread lineage diversity and mutational specificity. Nat Biotechnol. 2016;34:155–163. doi: 10.1038/nbt.3391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Heidorn SJ, et al. Kinase-dead BRAF and oncogenic RAS cooperate to drive tumor progression through CRAF. Cell. 2010;140:209–221. doi: 10.1016/j.cell.2009.12.040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Poulikakos PI, Zhang C, Bollag G, Shokat KM, Rosen N. RAF inhibitors transactivate RAF dimers and ERK signalling in cells with wild-type BRAF. Nature. 2010;464:427–430. doi: 10.1038/nature08902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Sebolt-Leopold JS, Herrera R. Targeting the mitogen-activated protein kinase cascade to treat cancer. Nat Rev Cancer. 2004;4:937–947. doi: 10.1038/nrc1503. [DOI] [PubMed] [Google Scholar]
- 5.Rauen KA. The RASopathies. Annu Rev Genomics Hum Genet. 2013;14:355–369. doi: 10.1146/annurev-genom-091212-153523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Hills SA, Diffley JFX. DNA replication and oncogene-induced replicative stress. Curr Biol. 2014;24:R435–R444. doi: 10.1016/j.cub.2014.04.012. [DOI] [PubMed] [Google Scholar]
- 7.Blasco RB, et al. c-Raf, but not Braf, is essential for development of K-Ras oncogene-driven non-small cell lung carcinoma. Cancer Cell. 2011;19:652–663. doi: 10.1016/j.ccr.2011.04.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Kamata T, et al. BRAF inactivation drives aneuploidy by deregulating CRAF. Cancer Res. 2010;70:8475–8486. doi: 10.1158/0008-5472.CAN-10-0603. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Guerra C, et al. Tumor induction by an endogenous K-ras oncogene is highly dependent on cellular context. Cancer Cell. 2003;4:111–120. doi: 10.1016/s1535-6108(03)00191-0. [DOI] [PubMed] [Google Scholar]
- 10.Mainardi S, et al. Identification of cancer initiating cells in K-Ras driven lung adenocarcinoma. Proc Natl Acad Sci USA. 2014;111:255–260. doi: 10.1073/pnas.1320383110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Karreth FA, Frese KK, DeNicola GM, Baccarini M, Tuveson DA. C-Raf is required for the initiation of lung cancer by K-Ras(G12D) Cancer Discov. 2011;1:128–136. doi: 10.1158/2159-8290.CD-10-0044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Bartkova J, et al. DNA damage response as a candidate anti-cancer barrier in early human tumorigenesis. Nature. 2005;434:864–870. doi: 10.1038/nature03482. [DOI] [PubMed] [Google Scholar]
- 13.Gorgoulis VG, et al. Activation of the DNA damage checkpoint and genomic instability in human precancerous lesions. Nature. 2005;434:907–913. doi: 10.1038/nature03485. [DOI] [PubMed] [Google Scholar]
- 14.Dietlein F, et al. A synergistic interaction between Chk1- and MK2 inhibitors in KRAS-mutant cancer. Cell. 2015;162:146–159. doi: 10.1016/j.cell.2015.05.053. [DOI] [PubMed] [Google Scholar]
- 15.Deschênes-Simard X, et al. Tumor suppressor activity of the ERK/MAPK pathway by promoting selective protein degradation. Genes Dev. 2013;27:900–915. doi: 10.1101/gad.203984.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Cheung WKC, Nguyen DX. Lineage factors and differentiation states in lung cancer progression. Oncogene. 2015;34:5771–5780. doi: 10.1038/onc.2015.85. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Andreadi C, et al. The intermediate-activity (L597V)BRAF mutant acts as an epistatic modifier of oncogenic RAS by enhancing signaling through the RAF/MEK/ERK pathway. Genes Dev. 2012;26:1945–1958. doi: 10.1101/gad.193458.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Langer CJ, Besse B, Gualberto A, Brambilla E, Soria JC. The evolving role of histology in the management of advanced non-small-cell lung cancer. J Clin Oncol. 2010;28:5311–5320. doi: 10.1200/JCO.2010.28.8126. [DOI] [PubMed] [Google Scholar]
- 19.Sutherland KD, et al. Cell of origin of small cell lung cancer: inactivation of Trp53 and Rb1 in distinct cell types of adult mouse lung. Cancer Cell. 2011;19:754–764. doi: 10.1016/j.ccr.2011.04.019. [DOI] [PubMed] [Google Scholar]
- 20.Xu X, et al. Evidence for type II cells as cells of origin of K-Ras-induced distal lung adenocarcinoma. Proc Natl Acad Sci USA. 2012;109:4910–4915. doi: 10.1073/pnas.1112499109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Sutherland KD, et al. Multiple cells-of-origin of mutant K-Ras-induced mouse lung adenocarcinoma. Proc Natl Acad Sci USA. 2014;111:4952–4957. doi: 10.1073/pnas.1319963111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Buchert M, et al. Genetic dissection of differential signaling threshold requirements for the Wnt/beta-catenin pathway in vivo. PLoS Genet. 2010;6:e1000816. doi: 10.1371/journal.pgen.1000816. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Papa A, et al. Cancer-associated PTEN mutants act in a dominant- negative manner to suppress PTEN protein function. Cell. 2014;157:595–610. doi: 10.1016/j.cell.2014.03.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Yao Z, et al. Tumours with class 3 BRAF mutants are sensitive to the inhibition of activated RAS. Nature. 2017 doi: 10.1038/nature23291. http://dx.doi.org/10.1038/nature23291. [DOI] [PMC free article] [PubMed]
- 25.Doma E, et al. Skin tumorigenesis stimulated by Raf inhibitors relies upon Raf functions that are dependent and independent of ERK. Cancer Res. 2013;73:6926–6937. doi: 10.1158/0008-5472.CAN-13-0748. [DOI] [PubMed] [Google Scholar]
- 26.Kammouni W, et al. Increased K-ras protein and activity in mouse and human lung epithelial cells at confluence. Cell Growth Differ. 2002;13:441–448. [PubMed] [Google Scholar]
- 27.Carter J, et al. Non-p.V600E BRAF mutations are common using a more sensitive and broad detection tool. Am J Clin Pathol. 2015;144:620–628. doi: 10.1309/AJCP85ATMJOZOUDJ. [DOI] [PubMed] [Google Scholar]
- 28.Chen AP, et al. Forebrain-specific knockout of B-raf kinase leads to deficits in hippocampal long-term potentiation, learning, and memory. J Neurosci Res. 2006;83:28–38. doi: 10.1002/jnr.20703. [DOI] [PubMed] [Google Scholar]
- 29.Dankort D, et al. A new mouse model to explore the initiation, progression, and therapy of BRAFV600E-induced lung tumors. Genes Dev. 2007;21:379–384. doi: 10.1101/gad.1516407. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Jonkers J, et al. Synergistic tumor suppressor activity of BRCA2 and p53 in a conditional mouse model for breast cancer. Nat Genet. 2001;29:418–425. doi: 10.1038/ng747. [DOI] [PubMed] [Google Scholar]
- 31.Jackson EL, et al. The differential effects of mutant p53 alleles on advanced murine lung cancer. Cancer Res. 2005;65:10280–10288. doi: 10.1158/0008-5472.CAN-05-2193. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All data generated or analysed during this study are included in this published article (and its Supplementary Information files).