Abstract
Background
Duchenne muscular dystrophy (DMD) is a lethal X-linked muscle wasting disorder caused by the absence of dystrophin, a large cytoskeletal muscle protein. Increasing the levels of the dystrophin-related-protein utrophin is a highly promising therapy for DMD and has been shown to improve pathology in dystrophin-deficient mice. One contributing factor to muscle wasting in DMD is mitochondrial pathology that contributes to oxidative stress and propagates muscle damage. The purpose of this study was to assess whether utrophin could attenuate mitochondria pathology and oxidative stress.
Methods
Skeletal muscles from wildtype C57BL/10, dystrophin-deficient mdx, dystrophin/utrophin double knockout (dko) and dystrophin-deficient mdx/utrophin over-expressing mdx-Fiona transgenic mice were assessed for markers of mitochondrial damage.
Results
Using transmission electron microscopy, we show that high levels of utrophin ameliorate the aberrant structure and localisation of mitochondria in mdx mice whereas absence of utrophin worsened these features in dko mice. Elevated utrophin also reverts markers of protein oxidation and oxidative stress, elevated in mdx and dko mice, to wildtype levels. These changes were observed independently of a shift in oxidative phenotype.
Conclusion
These findings show that utrophin levels influence mitochondrial pathology and oxidative stress. While utrophin deficiency worsens the pathology, utrophin over-expression in dystrophic muscle benefits mitochondria and attenuates the downstream pathology associated with aberrant mitochondrial function.
Electronic supplementary material
The online version of this article (10.1186/s13395-017-0139-5) contains supplementary material, which is available to authorized users.
Keywords: Duchenne muscular dystrophy, Mitochondria defects, Oxidative stress, Utrophin
Background
Duchenne muscular dystrophy (DMD) is a severe and progressive muscle wasting disorder [1, 2] affecting 1:5000 boys [3]. DMD is first evident early in childhood when boys exhibit ambulatory and balance difficulties. Muscle degeneration leads to loss of ambulation at 8–12 years, and patients succumb to respiratory and/or cardiac insufficiencies in their second or third decade of life [4]. The primary cause of pathology is the lack of the protein dystrophin, a large structural protein located at the sarcolemma, connecting the internal cytoskeleton to the surrounding extracellular matrix that confers membrane stability [5]. In its absence, the sarcolemma is susceptible to contraction-induced injury, propagating numerous secondary pathologies. These include extensive loss of calcium (Ca2+) homeostasis, calpain activation, oxidative stress and myofibre degeneration [6–11]. One contributing factor to pathology is aberrant mitochondrial function which drives, in part, oxidative stress and propagates muscle damage [12].
Mitochondria are cytoplasmic double membrane-bound organelles involved in energy conversion, generation of reactive oxygen species (ROS) and Ca2+ handling [13, 14]. Involvement of mitochondria in the dystrophic pathology has been well characterised with reduced mitochondrial respiration found in skeletal muscles from dystrophin-deficient (mdx) mice [15, 16] and DMD [17] patients. Percival and colleagues also observed a reduced density of sub-sarcolemmal mitochondria and aberrant localisation of inter-myofibrillar mitochondria in mdx mice [16]. In vitro, dystrophin-deficient myotubes have been shown to be more susceptible to free radical-induced injury compared to wildtype myotubes [10] and in vivo studies showed elevated lipid peroxidation and antioxidant gene expression [11]. These studies highlight the involvement of oxidative stress in the pathophysiology of mdx mice, thought to be due to Ca2+-induced mitochondrial dysfunction. In support of this, drug-mediated desensitisation of mitochondria to Ca2+ overload improves aspects of dystrophic pathology in mdx mice [18]. Studies by Vila and colleagues also implicate mitochondria in membrane repair as they accumulate at the site of rupture; a process that is diminished in mdx myofibres (which demonstrate impaired sarcolemmal repair) [19]. These studies highlight that loss of dystrophin leads to mitochondrial dysfunction which in turn plays a significant role in dystrophic pathogenesis.
Utrophin is a structural and functional autosomal parologue of dystrophin [20, 21] that is localised to the sarcolemma during foetal development and confined to the neuromuscular junction (NMJ) in mature muscle [22]. Pre-clinical studies have shown that both transgenic utrophin over-expression and pharmacological modulation using small compounds can prevent pathology in mdx mice [23–27]. Thus, modulation of utrophin is a highly promising therapy for DMD patients, currently under investigation in a phase II clinical trial [28]. Given the ability of utrophin to act as a surrogate to compensate for dystrophin deficiency at the sarcolemma and the contribution of aberrant mitochondrial function to the dystrophic pathology, the purpose of this study was to test whether utrophin over-expression or absence could influence mitochondrial pathology caused by dystrophin deficiency. As oxidative stress propagates the dystrophic phenotype and is a downstream effect of mitochondrial dysfunction, markers of oxidative stress were also assessed.
The capacity of utrophin to improve the dystrophic pathology has also been implicated independently from membrane stabilisation. Upregulation of a key metabolic regulator proliferator-activated receptor gamma coactivator (PGC)-1α has been shown to improve mdx pathology, likely through stimulation of a slower oxidative phenotype [29, 30]. These improvements were observed alongside elevated utrophin and Sirtuin (Sirt) 1 protein expression levels. Therefore, we have also assessed PCG-1α signalling to determine whether over-expression of utrophin can impact the dystrophic pathology outside of its recognised role as a dystrophin surrogate.
Our overall aim was to better understand the molecular impact of utrophin upregulation and deficiency on dystrophic muscle to inform the optimal clinical application of utrophin-modulation strategies.
Methods
Mice
Wildtype C57BL/10ScSnOlaHsd (C57BL/10), dystrophin-deficient C57BL/1010ScSn-Dmdmdx/J (mdx), dystrophin/utrophin double knockout (dko) and dystrophin-deficient/utrophin over-expressing C57/Bl10ScSn-Dmdmdx/J-Tg (ACTA1-Utrn)2Ked (mdx-Fiona) mice were assessed. C57BL/10 mice were obtained from Envigo (UK) and all other mouse strains were bred in the Biomedical Services facility, University of Oxford. Ten-week-old male mice were sacrificed and muscles were immediately excised and snap frozen (with the exception of those taken for transmission electron microscopy assessment) in liquid nitrogen or embedded in OCT and frozen in thawing isopentane. Samples were stored at − 80 °C until further analysis.
Histological assessment
Serial sections (10 μm) were cut transversely through the extensor digitorum longos (EDL) using a refrigerated (− 20 °C) cryostat (Bright OTF5000 Cryostat, Casa Alvarez Scientific Material, Spain). To assess muscle architecture fresh frozen sections were reacted with haematoxylin and eosin (H&E). Cytochrome oxidase (COX) staining was performed by incubating sections for 1 h at 37 °C in a phosphate buffer (pH 7.6) containing 50 mM sodium dibasic phosphate heptahydrate, 50 mM sodium monobasic phosphate monohydrate, 2.2 M sucrose, 1.3 mM 3,3′-diaminobenzidine tetrahydrochloride and 80 mM cytochrome C. Sites of COX activity were coloured brown [31, 32]. Succinate dehydrogenase (SDH) staining was performed by incubating sections for 15 min in SDH solution containing 50 mM disodium succinate, 0.7 M sodium azide, 6.5 mM disodium EDTA, 13.5 mM NaH2PO4.H2O, 6.8 mM Na2HPO4.7H2O, 1.5 mM nitroblue tetrazolium and 1.1 mM 1-methoxyphenzine methosulfate. Sites of SDH activity were coloured blue [33]. Gomori’s trichome staining was performed by incubating slides for 5 min in 1% glacial acetic acid, 12.8 mM chromotrope 2R, 3.7 mM fast green FCF and 2.1 mM phosphotungstic acid prior to dehydration in ascending concentrations of ethanol. Mitochondria were coloured red. All sections were air dried before application of coverslip and mounting medium and acquisition of digital images (Axioplan 2 Microscope System; Carl Zeiss, Germany).
Western blot analysis
Muscle samples were homogenised (Polytron 2100; Lucerne, Switzerland) for 3–15 s on ice in RIPA buffer (Sigma-Aldrich) supplemented with protease inhibitors (1:100; Sigma-Aldrich) and sodium orthovanadate (1 mM; Sigma-Aldrich). Following normalisation of protein concentration, homogenates were resolved in Laemmli buffer before heating to 95 °C (with the exception of those assessed using MitoProfile which were heated to 37 °C) for 5 min and were separated by sodium dodecyl sulphate-polyacrylamide gel electrophoresis (SDS-PAGE). For larger molecular weight proteins, utrophin and dystrophin, samples were loaded onto NuPAGE 3–8% TRIS Acetate Midi Gels (Novex, Life Technologies) and transferred overnight to PVDF membranes (Millipore). Membranes were blocked for 1 h with 5% skimmed milk in 0.1% PBS-Tween20 (T) and then incubated with primary antibodies in 0.1% PBS-T for 1 h at room temperature. Primary antibody and dilutions used were: utrophin, (MANCHO3; 84A) 1:50, a gift from G.E. Morris and dystrophin (Abcam; ab15277) 1:50. For all other targets, samples were loaded onto Criterion TGX (Tris-Glycine eXtended) pre-cast gels (Bio-Rad Laboratories, CA, USA) and were transferred for 4 h to PVDF membranes (Millipore). Membranes were blocked for 1 h with 5% skimmed milk in 0.1% PBS-T and then incubated with primary antibodies in 0.1% PBS-T overnight at 4 °C. Primary antibodies and dilutions used were Sirt1 (Millipore; 07–131) 1:500, AMPK (Cell Signaling Technologies; 9272S) 1:1000, phosphorylated AMPK (Cell Signaling Technologies; 9271S) 1:1000 and MitoProfile® Total OXPHOS Rodent WB Antibody Cocktail (Abcam; 110,413) 1:1000. For all targets, antibody binding was detected with horseradish peroxidase (HRP)-conjugated immunoglobulin and visualised by chemiluminescent detection (ECL Prime Western Blotting detection reagent Amersham) and imaging system ImageQuant LAS 4000 (GE Healthcare Life Sciences). Band densities were quantified with the Fiji ImageJ 1.49i software and normalised to total protein content of samples. Total protein was assessed by incubating membrane in Ponceau S solution (Sigma-Aldrich) followed by imaging using the ImageQuant LAS 4000 (GE Healthcare Life Sciences). Lane densities were quantified with the Fiji ImageJ 1.49i software. Total protein staining was used as the loading control as expression of housekeeping genes are not stable across genotypes.
Transmission electron microscopy (TEM)
Ultrastructural analysis was performed on tibialis anterior (TA) muscle samples fixed in 2.5% glutaraldehyde and 2–4% paraformaldehyde in 0.1 mM cacodylate buffer (pH 7.4) at room temperature for 1–2 h and then at 4 °C for 24 h. Samples were washed with 0.1 mM sodium cacodylate buffer 4 × 15 min, with the third wash also containing 25 mM glycine. Samples were post-fixed with 2% osmium, 1.5% potassium ferrocyanide in 0.1 M sodium cacodylate buffer for 1 h with rotation at 4 °C. Samples were washed 4 × 15 min with water, en bloc stained with 0.5% uranyl acetate overnight at 4 °C, washed with water and dehydrated 3–4 h using a graded ethanol series. Samples were infiltrated with TAAB TLV epoxy resin according to the following schedule: 1:3 resin:ethanol for 2 h, 1:1 resin:ethanol for 3 h, 3:1 resin:ethanol for 1 h, overnight in 100% resin, followed by 6 more changes of 100% resin over 36 h before embedding in beem capsules and polyermising for 48 h at 60 °C. Ultrathin sections (60 nm) were post-stained for 5 min with lead citrate, washed with water and examined using a FEI Tecnai 12 transmission electron microscope (OR, USA) operated at 120 kV with a Gatan OneView CMOS camera (CA, USA).
Immunofluorescence
Fresh frozen muscle cross-sections (10 μm thick) assessed of utrophin and dystrophin were blocked for 30 min with 10% foetal bovine serum (FBS)/PBS and then incubated with primary antibodies in 5% FBS/PBS overnight at 4 °C. Primary antibodies and dilutions used were utrophin (developed in-house as previously described [34]) 1:2000 and dystrophin (Abcam; ab15277) 1:2000. Sections were reacted with Alexa Fluor ® 488 donkey anti-goat IgG antibody (ThermoFisher Scientific; A-11055) or Alexa Fluor ® 594 donkey anti-rabbit IgG antibody (ThermoFisher Scientific; R37117) 1:2000 for 1 h at room temperature and then rinsed in PBS before air drying and application of cover slip with fluorescent mounting medium. Intensity was detected using a fluorescence microscope (Axioplan 2 Microscope System; Carl Zeiss, Germany).
Immunoglobulin (Ig) G staining for assessment of membrane integrity
Fresh frozen muscle cross-sections were blocked for 30 min with 10% FBS/PBS and then incubated with Alexa Fluor ® 488 goat anti-mouse IgG (Life Technologies; ab150117) antibody, 1:750 in 5% FBS/PBS overnight at 4 °C. Sections were rinsed in PBS before air drying and application of cover with fluorescent mounting medium, intensity was detected using a fluorescence microscope (Axioplan 2 Microscope System; Carl Zeiss, Germany).
Gene expression analysis
PGC-1α and PGC-1β mRNA expression were determined by quantitative real-time polymerase chain reaction (qPCR) as previously described [27]. Total RNA was extracted from 10 to 20 mg of TA muscle using TRIzol reagent as per manufacturer’s recommendations. Two hundred fifty nanogram RNA was transcribed into cDNA using the QuantiTect Reverse Transcription kit (Qiagen 205313), and the resulting cDNA was stored at − 20 °C for subsequent analysis. qPCR was performed using the StepOnePlus™ Real-Time PCR system (Applied Biosystems) with SYBR® Fast Master Mix (Thermofisher 4385612). Primer sequences used were PGC-1α (FWD, 5′-AAGTGTGGAACTCTCTGGAACTG-3′; REV, 5′-GGGTTATCTTGGTTGGCTTTATG-3′; PGC-1β (FWD, 5′-TGCTGCTGTCCTCAAATACG-3′, REV, 5′-TGGAGACTGCTCTGGAAGGT-3′). Gene expression was normalised to ribosomal protein S13 (FWD; 5′-CCCCGAGGATCTCTACCATT-3′, REV; 5′-GCCACTAGACAGAGGCTGT-3′). Results were analysed using the ΔΔCT method.
Spectrophotometric enzyme assays
Quadriceps muscles were crushed in liquid nitrogen and suspended in homogenisation buffer (pH 7.4) containing 100 mM KCl, 50 mM MOPS and 0.5 mM EGTA. For assessment of complex I activity, 2.5 mg of tissue was added to 1 mL assay buffer heated to 30 °C containing 0.25 M potassium phosphate buffer (pH 7.4; potassium phosphate dibasic and potassium phosphate monobasic), 5 mM MgCl2, 0.13 mM NADH, 65 μM Coenzyme Q1, 2.5 mg fatty acid-free bovine serum albumin per reaction and 3.6 mM Antimycin A. Change in absorbance was read at 340 nm. For assessment of complex II activity, 2.5 mg of protein was added to 0.25 M potassium phosphate buffer (pH 7.4; potassium phosphate dibasic and potassium phosphate monobasic), 5 mM MgCl2 and 2 mM sodium succinate dibasic hexahydrate followed by a 10 min incubation at 30 °C prior to the addition of 2 μg/mL Antimycin A, 2 μg/mL rotenone and 50 uM dichlorophenolindoprol (DCIP). Change in absorbance was read at 600 nm. For both assays, absorbance was read using a Pharmaspec UV-1700 UV visible spectrophotometer (Shimadzu, Kyoto Japan) and UV 2.10 software (Shimadzu). Activity of complex I and II was determined by the gradient of change in absorbance relative to the extinction coefficient of NADH and DCIP, respectively. Assays were validated by the addition of specific complex inhibitors, rotenone (complex I) and oxaloacetate (complex II). All reagents used for these assays were obtained from Sigma.
Oxidation indicator dihydroethidium (DHE) intensity
Fresh frozen muscle cross-sections (10 μm thick) were incubated in 2 μM DHE (Cayman Chemical, MI, USA) in PBS with 0.1% DMSO at 37 °C for 30 mins. Sections were rinsed in PBS before air drying and application of cover with fluorescent mounting medium containing DAPI. DHE intensity was detected as red fluorescence using a fluorescence microscope (Axioplan 2 Microscope System; Carl Zeiss, Germany).
Protein oxidation detection
OxyBlot™ Protein Oxidation Detection Kit (Millipore S7150) was utilised to perform immunoblot detection of carbonyl groups introduced into proteins by oxidative reactions. Samples were prepared and assessed as per manufacturer’s instructions. Briefly, fresh frozen muscle tissue was homogenised, as described previously, and protein concentrations normalised. Carbonyl groups in protein side chains were derivatised by reaction with 2, 4-dinitrophenylhydrazine (DNPH) and samples separated using SDS-PAGE. For separation, samples were loaded onto Criterion TGX (Tris-Glycine eXtended) pre-cast gels (Bio-Rad Laboratories, CA, USA) and were transferred for 4 h to PVDF membranes (Millipore). Membranes were blocked for 1 h with 5% skimmed milk in 0.1% PBS-T and then incubated with primary antibody (provided) in 0.1% PBS-T overnight at 4 °C. Antibody binding was detected with HRP-conjugated immunoglobulin and visualised by chemiluminescent detection (ECL Prime Western Blotting detection reagent Amersham) and imaging system ImageQuant LAS 4000 (GE Healthcare Life Sciences). Band densities were quantified with the Fiji ImageJ 1.49i software and normalised to total protein content of samples. Lanes with higher intensity band densities indicate a greater degree of protein carbonylation (a type of oxidation that can be promoted by ROS) [35].
Statistical analysis
Results were analysed using Prism (GraphPad Software, Inc.). One-way analysis of variance (ANOVA) was used to compare groups and Tukey’s post hoc test was used to determine significant differences between individual groups. The level of significance was set at P < 0.05 for all comparisons. All values are presented as mean ± S.E.M.
Results
Loss of dystrophin and utrophin from the sarcolemma leads to membrane rupture
Prior to the investigation of mitochondrial pathology, we first confirmed the phenotypes of the mouse strains used. Assessment of muscle architecture via H&E staining of EDL muscles revealed the expected phenotypes, with dystrophic legions evident in mdx muscle, worsened in dko muscle and ameliorated in mdx-Fiona muscle (Fig. 1a). Utrophin immunofluorescence (IF) revealed the presence of utrophin at the NMJ in C57BL/10 muscle, the NMJ and in regenerating fibres in mdx muscle and uniformly localised to the sarcolemma of mdx-Fiona muscle. As expected, utrophin was absent from dko muscle (Fig. 1b). Dystrophin was present only in C57BL/10 mice and absent from all other strains (Fig. 1c). IgG infiltration was observed in some mdx mice, was evident in all dko mice and not present in C57BL/10 and mdx-Fiona animals (Fig. 1d). Western blot assessment revealed the expected results. Dystrophin was present only in C57BL/10 mice. Utrophin levels were elevated in mdx-Fiona mice compared to mdx mice, detected in low levels in C57BL/10 mice and absent from dko mice (Fig. 1e–f).
Fig. 1.
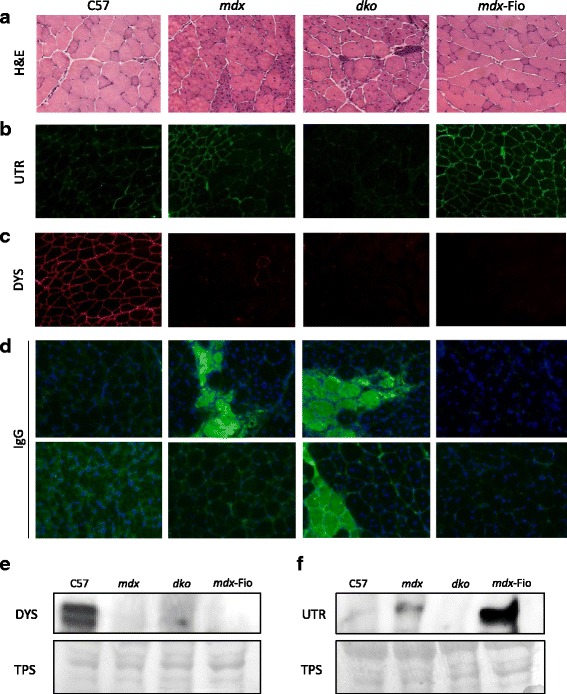
Muscle architecture and dystrophin and utrophin expression in extensor digitorum longus (EDL) muscles from C57BL/10, mdx, dko and mdx-Fiona mice. Representative images of (a) muscle architecture (assessed by H&E staining), (b) utrophin (UTR) immunofluorescence, (c) dystrophin (DYS) immunofluorescence and (d) membrane integrity (assessed via IgG infiltration and indicated by green) of EDL muscle cross-sections from C57BL/10 (C57), mdx, dko and mdx-Fiona (mdx-Fio) mice (n = 5–6/group). Protein expression levels, detected by western blot, of (e) dystrophin and (f) utrophin in quadriceps muscles of C57, mdx, dko and mdx-Fio mice and corresponding loading control (total protein stain; TPS; each lane contains 3 pooled samples from each genotype)
Aberrant mitochondrial structure and localisation are evident in mdx and dko mice and rescued in mdx-Fiona mice
In order to assess mitochondrial sub-structure and localisation, TA muscles from each mouse strain were investigated using TEM. Longitudinal sections of TA muscles from C57BL/10 mice exhibited localisation of mitochondria to the I band of myofibres and sound cristae formation within the mitochondria. Healthy structure was also observed in cross-sectional images (Fig. 2 and Additional file 1: Figure S1). Mitochondria were present at the I band in mdx muscle; however they also displayed abnormal aggregation, localisation to areas of necrosis and compromised structure (Fig. 2, Additional file 2: Figure S2). The aberrant features present in mdx mice were exacerbated in dko mice. Clear mitochondrial delocalisation, aggregation in regions of necrosis and severe disruption of internal membrane structure were all evident (Fig. 2, Additional file 3: Figure S3). TEM analysis of mdx-Fiona muscle did not reveal markers of pathology, with correct localisation and intact internal mitochondria structure (Fig. 2, Additional file 4: Figure S4). Histological assessment further supported TEM results with Gomori’s trichrome staining revealing increased mitochondrial aggregation in regions of necrosis in the EDL muscles from mdx and dko mice (Fig. 3a). Muscles from mdx-Fiona mice were comparable to C57BL/10 mice (Fig. 3a). These results show that utrophin is able to prevent the damage to mitochondria that results from dystrophin deficiency.
Fig. 2.
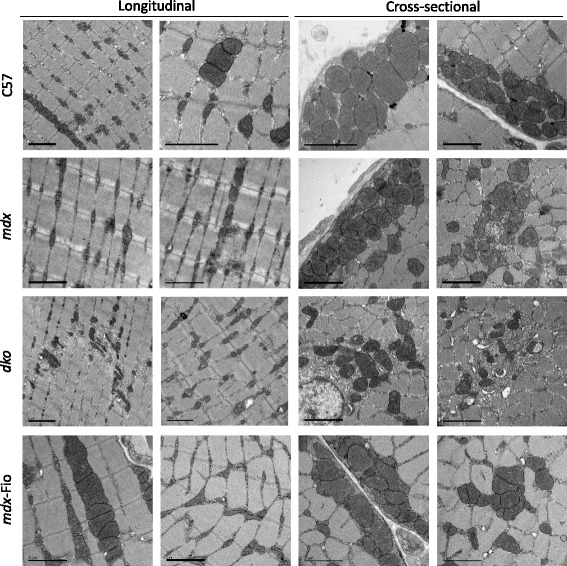
Transmission electron micrographs of tibialis anterior muscles from C57BL/10, mdx, dko and mdx-Fiona mice. Representative images of longitudinal sections and cross-sections of tibialis anterior muscles from C57BL/10 (C57), mdx, dko and mdx-Fiona (mdx-Fio) mice (n = 3/group). All scale bars = 2 μm, magnification varies
Fig. 3.
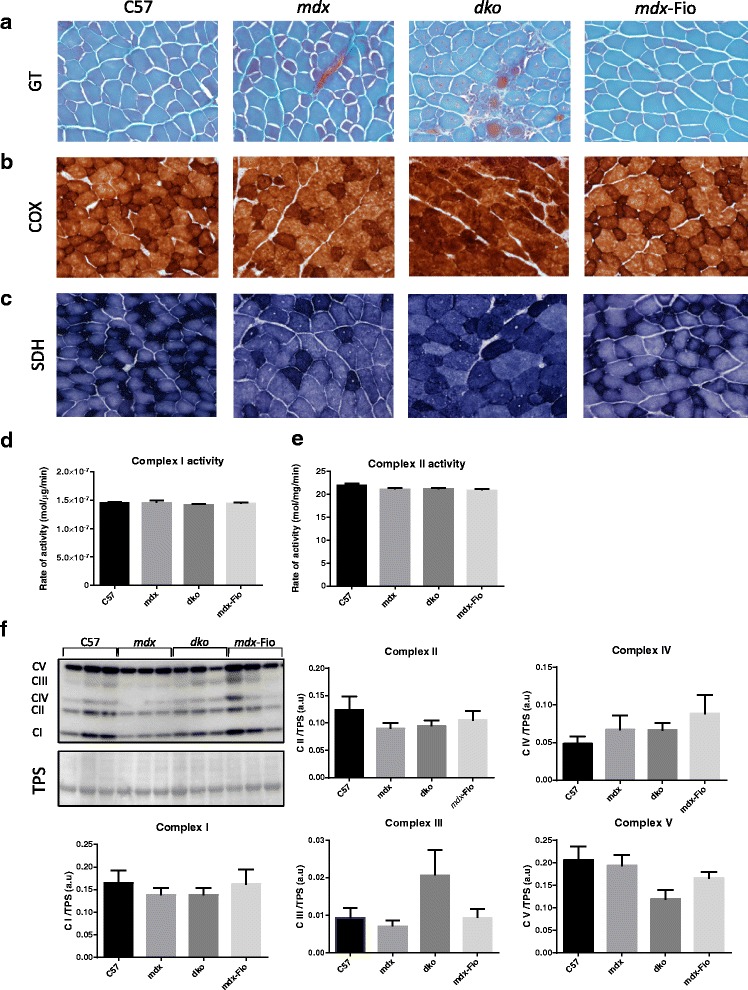
Assessment of markers of oxidative capacity in C57BL/10, mdx, dko and mdx-Fiona mice. Representative images of extensor digitorum longos muscles from C57BL/10 (C57), mdx, dko and mdx-Fiona (mdx-Fio) mice assessed for (a) mitochondrial aggregation (Via Gomori’s trichrome; GT), (b) cytochrome oxidase (COX) activity and (c) succinate dehydrogenase (SDH) activity indicated by increased intensity of red, brown and blue staining, respectively (n = 5–6/group). Enzyme activity of mitochondria respiratory chain complexes (d) I and (e) II (n = 4/group). f Representative images and quantification of protein expression in mitochondria respiratory chain complexes I–V in quadriceps muscles from C57, mdx, dko and mdx-Fio mice, normalised to loading control (total protein stain (TPS)). Data are represented as mean ± SEM, no differences were observed between groups. Representative image n = 3/group, quantification generated from n = 9/group
Mitochondrial benefits are independent from oxidative phenotype in mdx-Fiona mice
COX and SDH staining and enzyme activity assessment of mitochondrial respiratory chain complexes I and II were employed to test oxidative capacity of EDL muscles. COX and SDH staining, indicated by brown and blue colouring respectively, were not altered in mdx-Fiona mice compared to C57BL/10 animals. Muscle cross-sections from dko mice did not exhibit discrete staining of myofibres but rather, for both SDH and COX activity, overall darker staining compared to C57BL/10, mdx and mdx-Fiona mice (Fig. 3b–c). In support of these findings, specific enzymes activity of respiratory chain complexes I and II (Fig. 3d, e) and protein expression of complexes I–V (Fig. 3f) showed no significant difference between groups. Due to high levels of variation in protein expression, the sample number was increased to 9 for assessment of mitoprofile. However, statistical differences were still not resolved. These findings suggest oxidative capacity does not impact mitochondrial health in this context.
Mitochondria-associated cell signalling was altered in mdx and dko mice
Assessment of the Sirt1/PGC-1α/AMP-activated protein kinase (AMPK) signalling axis, implicated in metabolic control and mitochondrial biogenesis [36] revealed elevated expression in mdx and dko mice compared to C57BL/10 and mdx-Fiona mice. Western blot analysis of quadriceps muscles revealed significantly elevated Sirt1 protein levels in mdx mice compared to C57/BL10, dko and mdx-Fiona mice (Fig. 4a). AMPK activation was assessed by comparisons between phosphorylated and total protein expression levels. Western blot analysis revealed that total AMPK was elevated in mdx and dko mice. However, the relative levels of phosphorylation were only significantly different in muscle homogenates from dko mice compared to mdx-Fiona mice (Fig. 4b).Given the capacity of PGC-1β to compensate for PGC-1α [37], gene expression analysis of both was next assessed. PGC-1α was slightly elevated in dko mice compared to mdx-Fiona mice. No significant differences were observed between remaining genotypes (Fig. 4c). Assessment of PGC-1β revealed slightly elevated levels in dko mice compared to C57BL/10 mice, no significant differences were observed between remaining genotypes (Fig. 4c). Components of this signalling axis are elevated in mdx and dko mice and maintained at wildtype levels in mdx-Fiona mice; however this signalling does not appear to be involved in the preservation of mitochondrial health observed in mdx-Fiona mice.
Fig. 4.
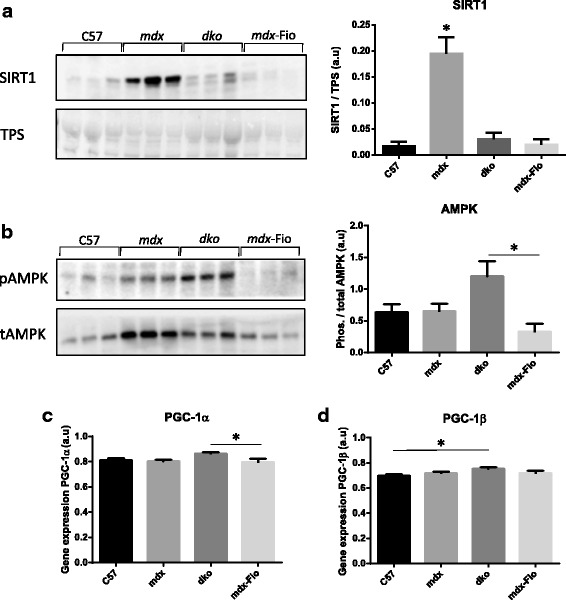
Assessment of mitochondria-associated cell signalling in C57BL/10, mdx, dko and mdx-Fiona mice. Representative images and quantification of a Sirtuin 1 (Sirt 1), normalised to loading control (total protein stain; TPS), and b phosphorylation status of AMP-activated protein kinase (AMPK), normalised to total protein expression of AMPK, in quadriceps muscles from C57BL/10 (C57), mdx, dko and mdx-Fiona (mdx-Fio) mice. Gene expression of c proliferator-activated receptor gamma coactivator (PGC) -1α and d PGC-1β in tibialis anterior muscles from C57, mdx, dko and mdx-Fio mice. Data are represented as mean ± SEM, * denotes significance set at P < 0.05. Representative image n = 3/group, quantification generated from n = 5-6/group
Markers of oxidative stress are elevated in mdx and dko mice and reduced in mdx- Fiona mice
Assessment of carbonylated protein content in quadriceps muscle homogenates were assessed using the oxyblot assay. Carbonylated protein levels were elevated in mdx mice compared to C57BL/10 mice. mdx-Fiona mice presented a very similar profile to wildtype animals. In contrast, dko mice exhibited elevated levels of protein carbonylation compared to mdx-Fiona mice, but levels were not significantly different to protein carbonylation in C57BL/10 mice. Protein carbonylation levels were not different between mdx and dko mice (Fig. 5a). These findings were supported by assessment of the oxidation indicator DHE, elevated in EDL cross-sections from mdx and dko mice compared to C57BL/10 and mdx-Fiona mice (Fig. 5b). Overall, markers of oxidative stress were increased in dko and mdx mice and maintained at wildtype levels in mdx-Fiona mice.
Fig. 5.
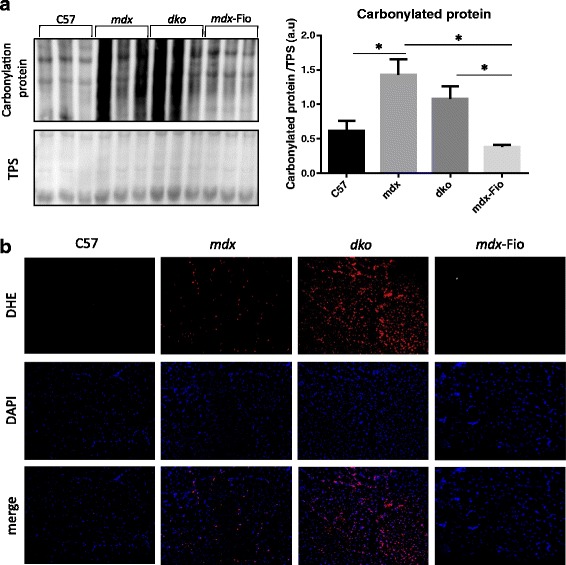
Protein carbonylation and oxidative indicator dihydroethidium (DHE) intensity in C57BL/10, mdx, dko and mdx-Fiona mice. a Representative images and quantification of carbonylated proteins (indicated by more intense banding), normalised to loading control (total protein stain (TPS)) in quadriceps muscles from C57BL/10 (C57), mdx, dko and mdx-Fiona (mdx-Fio) mice. b Representative images of extensor digitorum longos muscle cross-sections reacted with DHE and DAPI from C57, mdx, dko and mdx-Fiona mice. Data are represented as mean ± SEM, * denotes significance set at P < 0.05. Representative image n = 3/group, quantification generated from n = 6/group
Discussion
The data presented here show that increased utrophin expression ameliorates the aberrant mitochondrial structure and mitochondrial localisation observed in mdx and dko mice. Elevated utrophin in dystrophic muscle also reverted markers of protein carbonylation and oxidative stress to wildtype levels. These changes were observed independently from a shift in oxidative phenotype. Together, these findings show that utrophin over-expression benefits mitochondria in dystrophic muscle and attenuates the downstream pathology associated with aberrant mitochondrial function. Utrophin is expressed from early development in the transgenic models used here whereas utrophin modulation using viral vectors or small drugs can only be administered from birth [23]. However, these data reinforce the view that utrophin has the potential to compensate for dystrophin deficiency in DMD patients.
Previously, we have shown that both transgenic and pharmacological modulation of utrophin improves the dystrophic pathology [23, 26, 27]. Over-expression of the full-length utrophin ameliorated the dystrophic histopathology and improved membrane integrity. In order to assess the impact of utrophin on the sub-cellular structure of dystrophic muscle, TEM was utilised. Using similar techniques, Percival and colleagues showed that mitochondria in mdx mice exhibit aberrant structure, delocalisation from the I band and cluster in regions of necrosis [16]. Our findings emulate these observations, with mitochondria in mdx mice frequently localised to regions of necrosis and loss of cristae formation. We also observed that additional loss of utrophin, in dko mice, worsened these parameters with more extensive delocalisation and breakdown of internal structure. TEM images of muscles from mdx-Fiona mice were comparable to healthy muscles from C57BL/10 mice. These observations were further supported by Gomori’s trichrome staining which revealed aggregation of mitochondria in muscles from mdx and dko mice but not in those from mdx-Fiona and C57BL/10 mice. Our findings support previous studies that demonstrate dystrophin efficiency impairs spatial control of mitochondria and elicits mitochondrial damage [16]. Furthermore, we show for the first time these features are exacerbated by utrophin deficiency. Transgenic over-expression of utrophin was able to attenuate the above parameters, indicating utrophin can effectively replace dystrophin at the sarcolemma preventing mitochondrial damage.
Given the improved mitochondrial structure, it might be expected that the PGC-1α signalling pathway would be elevated in muscles from mdx-Fiona mice compared to mdx and dko mice. However, the opposite was observed, indicating the elevated signalling of this pathway is associated with the dystrophic phenotype and not mitochondrial health. As mdx mice undergo prolonged bouts of degeneration and regeneration, elevated Sirt1 signalling may be associated with mitophagy of damaged myofibres and mitochondrial biogenesis of regenerating myofibres. This may also be the case for elevated AMPK activation and PGC-1α/β expression in muscles from dko mice, although regeneration may contribute to a lesser extent. Overall, we see altered signalling associated with mitochondria in mdx and dko mice, and this was restored to wildtype levels with utrophin over-expression in dystrophic muscle. However, the changes observed in the PGC-1α signalling pathway are unlikely to contribute to the improvements observed in mitochondrial pathology.
High levels of utrophin are associated with a slower, more oxidative, muscle fibre type phenotype which is protective of dystrophic muscle [29, 38]. To determine whether a shift in fibre type profile contributed to the improved mitochondrial parameters observed using TEM, markers of oxidative capacity were tested across genotypes. No differences in COX staining, SDH staining, enzyme activity of respiratory chain complexes I and II or mitochondrial respiratory chain complex protein expression were observed in mdx-Fiona mice compared to mdx mice. However, similar to a previous study, SDH and COX staining of dko muscle appeared more intense compared to wildtype muscle [39]. This is likely to be a product of the severe phenotype in which fast twitch fibres are preferentially lost and not due a direct action on mitochondria [40]. Therefore, it does not appear that increasing utrophin expression impacts oxidative capacity of dystrophic muscle. Although we see no difference in COX or SDH staining in mdx mice compared to C57BL/10 mice, which is consistent with previous findings [7, 16, 17], transgenic upregulation of PGC-1α is able to elevate the proportion of slow twitch myofibres in mdx mice to above that of C57BL/10 mice [29]. Hence, the lack of reduced activity in mdx mice should not impede a transition to a slower fibre type profile in mdx-Fiona mice.
Although our observations regarding COX and SDH staining were concordant with previous studies, expression patterns of mitochondrial respiratory chain complex proteins were not. Percival and colleagues have previously observed elevated expression of complex III in mdx muscle compared to C57BL/10 muscle [16]; however, Ryu and colleagues saw reduced levels of complexes I, II, IV and V in mdx muscles compared to C57BL/10 mice [41]. The discrepancies between these studies and our own may be accounted for by the highly dynamic nature of mitochondria [42]. It is possible the turnover of these proteins may mask disease impact upon mitochondria. Therefore, measurements of the mitochondria respiratory chain complexes may not be a reliable indicator of the metabolic state of mitochondria. Taken together, these findings suggest transgenic upregulation of utrophin does not impact oxidative capacity in the dystrophic context. Therefore, oxidative phenotype is not contributing factor to improved mitochondrial pathology observed in mdx-Fiona mice.
Oxidative stress is a prominent feature of the dystrophic pathology. mdx mice were previously described as more susceptible to ROS-mediated cell damage and exhibited higher levels of lipid peroxidation compared to wildtype mice [10]. Oxidative injury was also found to precede myofibre necrosis in mdx mice, indicating it is likely one to be of the initial phases of pathology [11]. Consequently, the development of antioxidants for the treatment of DMD has been a substantial line of research with numerous compounds investigated, e.g. coenzyme Q10, catalase, green tea extract, Epigallocatechin gallate, BN82270, Idebenone, melatonin and N-acetylcysteine [14, 43–49]. Mitochondrial Ca2+ overload is known to promote ROS generation and membrane permeability in healthy muscle [13, 50]. As mitochondrial dysfunction is prominent in dystrophic muscle, it is likely to be one major mechanism of oxidative stress in DMD. Here, we show oxidative stress and protein carbonylation were elevated in mdx and dko mice compared to C57BL/10 mice. This effect was prevented in mdx-Fiona mice where parameters were comparable with wildtype.
Given utrophin is localised to the sarcolemma and not the cytosol, reductions in oxidative stress are not likely to be due to a scavenging action but rather a downstream effect of improved membrane integrity. Dystrophic myofibres are more susceptible to membrane rupture compared to healthy muscle fibres, allowing for excessive influx of Ca2+ rich extracellular fluid into the cytosol [7–9]. Given the link between Ca2+ overload and mitochondrial ROS production and prevention of IgG infiltration observed in mdx-Fiona mice, this is likely to be the mechanism of the attenuation of oxidative stress. Although IgG infiltration was not present in all mdx muscles, histology can only capture that state of the muscle at the time of harvest. It may be that these mice experience sarcolemmal rupture prior to muscle excision and this elicited mitochondrial stress. Although speculative, this seems the most likely mechanism of elevated oxidative stress. Irrespective of mechanism, our findings demonstrate the antioxidant effect of utrophin in dystrophic muscle. DMD is recognised as having multiple secondary pathologies and of these oxidative stress features prominently. A fuller understanding of the capacity of a specific treatment to target different aspects of pathology will inform the strongest approach for combinatorial treatment.
We recently identified numerous serological biomarkers that are elevated in mdx mice compared to wildtype mice; many of these are mitochondria-associated proteins [51]. Heat shock protein (HSP) 60, cytochrome C (CYCS), Sirt 2, HtrA serine peptidase 2 (HTRA2) and mitochondrial import inner membrane translocase subunit TIM14 (DNAJC19) were all elevated in serum from mdx mice and importantly normalised to wildtype levels in mdx-Fiona mice. These findings further support the capacity of high levels of utrophin to impact mitochondria in dystrophic muscle. Taken in conjunction with our current findings, mitochondrial biomarkers are highlighted as a valuable tool for assessing pre-clinical studies and potentially clinical benefits following treatment with small utrophin-modulating drugs.
Conclusions
Here, we show that utrophin over-expression in dystrophic muscle improves mitochondrial structure/localisation and reduces oxidative stress and that these benefits are potentially due to improved membrane integrity. We also show a detrimental impact on these parameters in dko mice, emphasising the importance of utrophin. Our results highlight the effectiveness of utrophin in alleviating mitochondrial pathology resulting from dystrophin deficiency. Additionally, we demonstrate the antioxidant effect of utrophin modulation, further elucidating its molecular impact on dystrophic muscle and relevance in multi-faceted treatment approaches for DMD patients.
Additional files
Transmission electron micrographs of tibialis anterior (TA) muscles from C57BL/10 (C57) mice. Cross-sections of TA muscles from C57 mice. Scale bars = (A) 10 μm, (B) 2 μm and (C) 1000 nm. (n = 3; C57 1, C57 2, C57 3). (PPTX 787 kb)
Transmission electron micrographs of tibialis anterior (TA) muscles from mdx mice. Cross-sections of TA muscles from mdx mice. Scale bars = (A) 10 μm, (B) 2 μm and (C) 1000 nm. (n = 3; mdx 1, mdx 2, mdx 3). (PPTX 805 kb)
Transmission electron micrographs of tibialis anterior (TA) muscles from dko mice. Cross-sections of TA muscles from dko mice. Scale bars = (A) 10 μm, (B) 2 μm and (C) 1000 nm. (n = 3; dko 1, dko 2, dko 3). (PPTX 841 kb)
Transmission electron micrographs of tibialis anterior (TA) muscles from mdx-Fiona (mdx-Fio) mice. Cross-sections of TA muscles from mdx-Fio mice. Scale bars = (A) 10 μm, (B) 2 μm and (C) 1000 nm. (n = 3; Fio 1, Fio 2, Fio 3). (PPTX 771 kb)
Acknowledgements
We acknowledge G.E. Morris (Oswestry, UK) for the MANCHO3 antibody, Dr. Errin Johnson from the Electron Microscopy Facility, The Sir William Dunn School of Pathology, University of Oxford, for generation of electron micrograph images. Also, Dr. Lisa Heather and Mr. Kenny Roberts for their expertise in mitochondrial function and use of Spectrophotometer.
Funding
This work was supported by grants from the Medical Research Council, Muscular Dystrophy UK and Duchenne UK.
Availability of data and materials
The datasets used and/or analysed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- AMPK
AMP-Activated protein kinase
- ANOVA
One-way analysis of variance
- C57BL/10
C57BL/10ScSnOlaHsd
- Ca2+
Calcium
- COX
Cytochrome oxidase
- CYCS
Cytochrome C
- DCIP
Dichlorophenolindoprol
- DHE
Dihydroethidium
- dko
Dystrophin/utrophin double knockout
- DMD
Duchenne muscular dystrophy
- DNAJC19
Mitochondrial import inner membrane translocase subunit TIM14
- DNPH
2, 4-dinitrophenylhydrazine
- EDL
Extensor digitorum longos
- FBS
Foetal bovine serum
- H&E
Haematoxylin and eosin
- HRP
Horseradish peroxidase
- HSP
Heat shock protein
- HTRA2
HtrA serine peptidase 2
- IF
Immunofluorescence
- Ig
Immunoglobulin
- mdx
Dystrophin-deficient C57BL/1010ScSn-Dmdmdx/J
- mdx-Fiona
Dystrophin-deficient/utrophin over-expressing C57/Bl10ScSn-Dmdmdx/J-Tg (ACTA1-Utrn)2Ked
- NMJ
Neuromuscular junction
- PGC
Proliferator-activated receptor gamma coactivator
- qPCR
Quantitative real time-polymerase chain reaction
- ROS
Reactive oxygen species
- SDH
Succinate dehydrogenase
- SDS-PAGE
Sodium dodecyl sulphate-polyacrylamide gel electrophoresis
- Sirt
Sirtuin
- T
Tween20
- TA
Tibialis anterior
- TEM
Transmission electron microscopy
- UK
United Kingdom
Authors’ contributions
The study was conceived and designed by TLK, SG and KED. Experimental work was performed by TLK, LM, BE, SH, SES and SG. Data were analysed by TLK, SG and KED. The manuscript was written by TLK, SG and KED. All authors have read and approved the final manuscript.
Ethics approval
All animal procedures were performed in accordance with United Kingdom (UK) Home Office regulations in compliance with the European Community Directive published in 1986 (86/609/ EEC). The study was performed under certificate of designation number 30/2306 and project license number 30/3104 following approval by the University of Oxford Departments of Physiology, Anatomy & Genetics and Experimental Psychology Joint Departmental Ethics Review Committee.
Competing interests
The authors declare that they have no competing interests
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Footnotes
Electronic supplementary material
The online version of this article (10.1186/s13395-017-0139-5) contains supplementary material, which is available to authorized users.
Contributor Information
Tahnee L. Kennedy, Email: tahnee.kennedy@dpag.ox.ac.uk
Lee Moir, Email: lee.moir@dpag.ox.ac.uk.
Sarah Hemming, Email: sarah.hemming@dpag.ox.ac.uk.
Ben Edwards, Email: benjamin.edwards@dpag.ox.ac.uk.
Sarah Squire, Email: sarah.squire@dpag.ox.ac.uk.
Kay Davies, Phone: +44 (0)1865 285880, Email: kay.davies@dpag.ox.ac.uk.
Simon Guiraud, Email: simon.guiraud@dpag.ox.ac.uk.
References
- 1.Mendell JR, et al. Evidence-based path to newborn screening for Duchenne muscular dystrophy. Ann Neurol. 2012;71(3):304–313. doi: 10.1002/ana.23528. [DOI] [PubMed] [Google Scholar]
- 2.Guiraud S, et al. The pathogenesis and therapy of muscular dystrophies. Annu Rev Genomics Hum Genet. 2015;16:281–308. doi: 10.1146/annurev-genom-090314-025003. [DOI] [PubMed] [Google Scholar]
- 3.Mah JK, et al. A systematic review and meta-analysis on the epidemiology of Duchenne and Becker muscular dystrophy. Neuromuscul Disord. 2014;24(6):482–491. doi: 10.1016/j.nmd.2014.03.008. [DOI] [PubMed] [Google Scholar]
- 4.Bach JR, et al. Management of end stage respiratory failure in Duchenne muscular dystrophy. Muscle Nerve. 1987;10(2):177–182. doi: 10.1002/mus.880100212. [DOI] [PubMed] [Google Scholar]
- 5.Dellorusso C, et al. Tibialis anterior muscles in mdx mice are highly susceptible to contraction-induced injury. J Muscle Res Cell Motil. 2001;22(5):467–475. doi: 10.1023/A:1014587918367. [DOI] [PubMed] [Google Scholar]
- 6.Spencer MJ, Croall DE, Tidball JG. Calpains are activated in necrotic fibers from mdx dystrophic mice. J Biol Chem. 1995;270(18):10909–10914. doi: 10.1074/jbc.270.18.10909. [DOI] [PubMed] [Google Scholar]
- 7.Rezvani M, Cafarelli E, Hood DA. Performance and excitability of mdx mouse muscle at 2, 5, and 13 wk of age. J Appl Physiol. 1995;78(3):961–967. doi: 10.1152/jappl.1995.78.3.961. [DOI] [PubMed] [Google Scholar]
- 8.Head S, Williams D, Stephenson G. Increased susceptibility of EDL muscles from mdx mice to damage induced by contraction with stretch. J Muscle Res Cell Motil. 1994;15(4):490. [PubMed] [Google Scholar]
- 9.Glesby MJ, et al. Serum CK, calcium, magnesium, and oxidative phosphorylation in mdx mouse muscular dystrophy. Muscle Nerve. 1988;11(8):852–856. doi: 10.1002/mus.880110809. [DOI] [PubMed] [Google Scholar]
- 10.Rando TA, et al. Muscle cells from mdx mice have an increased susceptibility to oxidative stress. Neuromuscul Disord. 1998;8(1):14–21. doi: 10.1016/S0960-8966(97)00124-7. [DOI] [PubMed] [Google Scholar]
- 11.Disatnik M-H, et al. Evidence of oxidative stress in mdx mouse muscle: studies of the pre-necrotic state. J Neurol Sci. 1998;161(1):77–84. doi: 10.1016/S0022-510X(98)00258-5. [DOI] [PubMed] [Google Scholar]
- 12.Allen DG, Whitehead NP, Froehner SC. Absence of dystrophin disrupts skeletal muscle signaling: roles of Ca2+, reactive oxygen species, and nitric oxide in the development of muscular dystrophy. Physiol Rev. 2016;96(1):253–305. doi: 10.1152/physrev.00007.2015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Brookes PS, Darley-Usmar VM. Role of calcium and superoxide dismutase in sensitizing mitochondria to peroxynitrite-induced permeability transition. Am J Phys Heart Circ Phys. 2004;286(1):H39–H46. doi: 10.1152/ajpheart.00742.2003. [DOI] [PubMed] [Google Scholar]
- 14.Chahbouni M, et al. Melatonin treatment counteracts the hyperoxidative status in erythrocytes of patients suffering from Duchenne muscular dystrophy. Clin Biochem. 2011;44(10):853–858. doi: 10.1016/j.clinbiochem.2011.04.001. [DOI] [PubMed] [Google Scholar]
- 15.Kuznetsov AV, et al. Impaired mitochondrial oxidative phosphorylation in skeletal muscle of the dystrophin-deficient mdx mouse. Mol Cell Biochem. 1998;183(1–2):87–96. doi: 10.1023/A:1006868130002. [DOI] [PubMed] [Google Scholar]
- 16.Percival JM, et al. Defects in mitochondrial localization and ATP synthesis in the mdx mouse model of Duchenne muscular dystrophy are not alleviated by PDE5 inhibition. Hum Mol Genet. 2013;22(1):153–167. doi: 10.1093/hmg/dds415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Sperl W, et al. High resolution respirometry of permeabilized skeletal muscle fibers in the diagnosis of neuromuscular disorders. Mol Cell Biochem. 1997;174(1–2):71–78. doi: 10.1023/A:1006880529195. [DOI] [PubMed] [Google Scholar]
- 18.Millay DP, et al. Genetic and pharmacologic inhibition of mitochondrial-dependent necrosis attenuates muscular dystrophy. Nat Med. 2008;14(4):442–447. doi: 10.1038/nm1736. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Vila MC, et al. Mitochondria mediate cell membrane repair and contribute to Duchenne muscular dystrophy. Cell Death Diff. 2016;24(2):330–42. [DOI] [PMC free article] [PubMed]
- 20.Love DR, et al. An autosomal transcript in skeletal muscle with homology to dystrophin. 1989. [DOI] [PubMed] [Google Scholar]
- 21.Tinsley JM, et al. Primary structure of dystrophin-related protein. Nature. 1992;360(6404):591–593. doi: 10.1038/360591a0. [DOI] [PubMed] [Google Scholar]
- 22.Lin S, Burgunder J-M. Utrophin may be a precursor of dystrophin during skeletal muscle development. Dev Brain Res. 2000;119(2):289–295. doi: 10.1016/S0165-3806(99)00165-0. [DOI] [PubMed] [Google Scholar]
- 23.Tinsley J, et al. Expression of full-length utrophin prevents muscular dystrophy in mdx mice. Nat Med. 1998;4(12):1441–1444. doi: 10.1038/4033. [DOI] [PubMed] [Google Scholar]
- 24.Krag TO, et al. Heregulin ameliorates the dystrophic phenotype in mdx mice. Proc Natl Acad Sci U S A. 2004;101(38):13856–13860. doi: 10.1073/pnas.0405972101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Amenta AR, et al. Biglycan recruits utrophin to the sarcolemma and counters dystrophic pathology in mdx mice. Proc Natl Acad Sci. 2011;108(2):762–767. doi: 10.1073/pnas.1013067108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Tinsley JM, et al. Daily treatment with SMTC1100, a novel small molecule utrophin upregulator, dramatically reduces the dystrophic symptoms in the mdx mouse. PLoS One. 2011;6(5):e19189. doi: 10.1371/journal.pone.0019189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Guiraud S, et al. Second-generation compound for the modulation of utrophin in the therapy of DMD. Hum Mol Genet. 2015;24:4212–24. [DOI] [PMC free article] [PubMed]
- 28.Summit Therapeutics. First patient enrolled in summit’s phaseOut DMD, a phase 2 clinical trial of ezutromid in boys with DMD. 2016; Available from: http://www.summitplc.com/media/press-releases/.
- 29.Selsby JT, et al. Rescue of dystrophic skeletal muscle by PGC-1α involves a fast to slow fiber type shift in the mdx mouse. PLoS One. 2012;7(1):e30063. doi: 10.1371/journal.pone.0030063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Handschin C, et al. PGC-1α regulates the neuromuscular junction program and ameliorates Duchenne muscular dystrophy. Genes Dev. 2007;21(7):770–783. doi: 10.1101/gad.1525107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Seligman AM, et al. Nondroplet ultrastructural demonstration of cytochrome oxidase activity with a polymerizing osmiophilic reagent, diaminobenzidine (DAB) J Cell Biol. 1968;38(1):1–14. doi: 10.1083/jcb.38.1.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Pearse, A . Histochemistry: theoretical and applied. 3. Boston: Little, Brown and Co.; 1968. [Google Scholar]
- 33.Blanco C, Sieck GC, Edgerton V. Quantitative histochemical determination of succinic dehydrogenase activity in skeletal muscle fibres. Histochem J. 1988;20(4):230–243. doi: 10.1007/BF01747468. [DOI] [PubMed] [Google Scholar]
- 34.Weir AP, et al. A-and B-utrophin have different expression patterns and are differentially up-regulated in mdx muscle. J Biol Chem. 2002;277(47):45285–45290. doi: 10.1074/jbc.M205177200. [DOI] [PubMed] [Google Scholar]
- 35.Suzuki YJ, Carini M, Butterfield DA. Protein carbonylation. Antioxid Redox Signal. 2010;12(3):323–325. doi: 10.1089/ars.2009.2887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Tang BL. Sirt1 and the mitochondria. Mol Cells. 2016;39(2):87. doi: 10.14348/molcells.2016.2318. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Wareski P, et al. PGC-1α and PGC-1β regulate mitochondrial density in neurons. J Biol Chem. 2009;284(32):21379–21385. doi: 10.1074/jbc.M109.018911. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Gramolini AO, et al. Increased expression of utrophin in a slow vs. a fast muscle involves posttranscriptional events. Am J Phys Cell Phys. 2001;281(4):C1300–C1309. doi: 10.1152/ajpcell.2001.281.4.C1300. [DOI] [PubMed] [Google Scholar]
- 39.Rafael JA, et al. Dystrophin and utrophin influence fiber type composition and post-synaptic membrane structure. Hum Mol Genet. 2000;9(9):1357–1367. doi: 10.1093/hmg/9.9.1357. [DOI] [PubMed] [Google Scholar]
- 40.Webster C, et al. Fast muscle fibers are preferentially affected in Duchenne muscular dystrophy. Cell. 1988;52(4):503–513. doi: 10.1016/0092-8674(88)90463-1. [DOI] [PubMed] [Google Scholar]
- 41.Ryu D, et al. NAD+ repletion improves muscle function in muscular dystrophy and counters global PARylation. Sci Transl Med. 2016;8(361):361ra139. doi: 10.1126/scitranslmed.aaf5504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Mishra P, Chan DC. Mitochondrial dynamics and inheritance during cell division, development and disease. Nat Rev Mol Cell Biol. 2014;15(10):634–646. doi: 10.1038/nrm3877. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Bjelakovic G, et al. Antioxidant supplements for prevention of mortality in healthy participants and patients with various diseases. Cochrane Libr. 2012;133(2):164–5. [DOI] [PMC free article] [PubMed]
- 44.Buetler TM, et al. Green tea extract decreases muscle necrosis in mdx mice and protects against reactive oxygen species. Am J Clin Nutr. 2002;75(4):749–753. doi: 10.1093/ajcn/75.4.749. [DOI] [PubMed] [Google Scholar]
- 45.Nakae Y, et al. Subcutaneous injection, from birth, of epigallocatechin-3-gallate, a component of green tea, limits the onset of muscular dystrophy in mdx mice: a quantitative histological, immunohistochemical and electrophysiological study. Histochem Cell Biol. 2008;129(4):489–501. doi: 10.1007/s00418-008-0390-2. [DOI] [PubMed] [Google Scholar]
- 46.Selsby JT. Increased catalase expression improves muscle function in mdx mice. Exp Physiol. 2011;96(2):194–202. doi: 10.1113/expphysiol.2010.054379. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Terrill JR, et al. N-Acetylcysteine treatment of dystrophic mdx mice results in protein thiol modifications and inhibition of exercise induced myofibre necrosis. Neuromuscul Disord. 2012;22(5):427–434. doi: 10.1016/j.nmd.2011.11.007. [DOI] [PubMed] [Google Scholar]
- 48.Burdi R, et al. First evaluation of the potential effectiveness in muscular dystrophy of a novel chimeric compound, BN 82270, acting as calpain-inhibitor and anti-oxidant. Neuromuscul Disord. 2006;16(4):237–248. doi: 10.1016/j.nmd.2006.01.013. [DOI] [PubMed] [Google Scholar]
- 49.Buyse GM, Gueven N, McDonald CM. Idebenone as a novel therapeutic approach for Duchenne muscular dystrophy. Eur Neurological Rev. 2015;10(2):189–194. doi: 10.17925/ENR.2015.10.02.189. [DOI] [Google Scholar]
- 50.Nethery D, et al. PLA2 dependence of diaphragm mitochondrial formation of reactive oxygen species. J Appl Physiol. 2000;89(1):72–80. doi: 10.1152/jappl.2000.89.1.72. [DOI] [PubMed] [Google Scholar]
- 51.Guiraud S, et al. Identification of serum protein biomarkers for utrophin based DMD therapy. Sci Rep. 2017;7:43697. doi: 10.1038/srep43697. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Transmission electron micrographs of tibialis anterior (TA) muscles from C57BL/10 (C57) mice. Cross-sections of TA muscles from C57 mice. Scale bars = (A) 10 μm, (B) 2 μm and (C) 1000 nm. (n = 3; C57 1, C57 2, C57 3). (PPTX 787 kb)
Transmission electron micrographs of tibialis anterior (TA) muscles from mdx mice. Cross-sections of TA muscles from mdx mice. Scale bars = (A) 10 μm, (B) 2 μm and (C) 1000 nm. (n = 3; mdx 1, mdx 2, mdx 3). (PPTX 805 kb)
Transmission electron micrographs of tibialis anterior (TA) muscles from dko mice. Cross-sections of TA muscles from dko mice. Scale bars = (A) 10 μm, (B) 2 μm and (C) 1000 nm. (n = 3; dko 1, dko 2, dko 3). (PPTX 841 kb)
Transmission electron micrographs of tibialis anterior (TA) muscles from mdx-Fiona (mdx-Fio) mice. Cross-sections of TA muscles from mdx-Fio mice. Scale bars = (A) 10 μm, (B) 2 μm and (C) 1000 nm. (n = 3; Fio 1, Fio 2, Fio 3). (PPTX 771 kb)
Data Availability Statement
The datasets used and/or analysed during the current study are available from the corresponding author on reasonable request.


