Abstract
It is now widely accepted in the field that the normally secreted chaperone clusterin is redirected to the cytosol during endoplasmic reticulum (ER) stress, although the physiological function(s) of this physical relocation remain unknown. We have examined in this study whether or not increased expression of clusterin is able to protect neuronal cells against intracellular protein aggregation and cytotoxicity, characteristics that are strongly implicated in a range of neurodegenerative diseases. We used the amyotrophic lateral sclerosis-associated protein TDP-43 as a primary model to investigate the effects of clusterin on protein aggregation and neurotoxicity in complementary in vitro, neuronal cell and Drosophila systems. We have shown that clusterin directly interacts with TDP-43 in vitro and potently inhibits its aggregation, and observed that in ER stressed neuronal cells, clusterin co-localized with TDP-43 and specifically reduced the numbers of cytoplasmic inclusions. We further showed that the expression of TDP-43 in transgenic Drosophila neurons induced ER stress and that co-expression of clusterin resulted in a dramatic clearance of mislocalized TDP-43 from motor neuron axons, partially rescued locomotor activity and significantly extended lifespan. We also showed that in Drosophila photoreceptor cells, clusterin co-expression gave ER stress-dependent protection against proteotoxicity arising from both Huntingtin-Q128 and mutant (R406W) human tau. We therefore conclude that increased expression of clusterin can provide an important defense against intracellular proteotoxicity under conditions that mimic specific features of neurodegenerative disease.
Electronic supplementary material
The online version of this article (10.1186/s40478-017-0481-1) contains supplementary material, which is available to authorized users.
Keywords: TDP-43, Cytoplasmic inclusions, Proteotoxicity, Chaperone translocation
Introduction
Protein misfolding, aggregation and deposition are unifying features of a wide range of neurodegenerative diseases [1]. The ability of neurons to manage the burden of misfolded proteins and to resist their accumulation into insoluble protein deposits depends critically on the functioning of molecular chaperones. Previous studies have demonstrated that elevation of chaperone levels can protect against neurotoxicity resulting from the effects of pathological protein misfolding in cell culture and in transgenic animal models [2, 3]. Most chaperones are localised within intracellular compartments, although some are secreted into the extracellular environment. Prominent amongst such extracellular chaperones is clusterin (CLU), which is present in both plasma and cerebrospinal fluid (CSF). CLU is a cytoprotective chaperone whose expression level is increased in response to a diverse range of stresses including heat, pro-apoptotic insults, oxidative stress, ionising radiation, and proteotoxicity [4–6]. It has been linked to a correspondingly diverse group of clinical disorders associated with protein misfolding including Alzheimer’s disease (AD) [7], amyloidotic cardiomyopathy [8] and familial amyloidotic polyneuropathy [9]. CLU binds promiscuously to a wide range of misfolded client proteins and either sequesters them into stable, soluble complexes (in the case of proteins forming amorphous aggregates) or inhibits the formation and accumulation of toxic amyloid assemblies [10, 11]. Clusterin is a particularly potent chaperone and can inhibit protein aggregation at molar ratios of chaperone:client protein that are significantly lower than those needed by other chaperones [12, 13]. Extracellular CLU-client protein complexes are susceptible to endocytic clearance by monocytes and hepatocytes, and this action is thought to form a central plank in the operation of a co-ordinated system to maintain extracellular protein homeostasis (proteostasis) [14].
Whilst CLU is predominantly a secreted protein, multiple independent studies have now clearly established that during conditions of ER stress, CLU secretion is inhibited, and full-length CLU exits the secretory system to enter the cytosol. Firstly, the translocation of mature CLU from the ER/Golgi into the cytosol of human astrocytoma U251 cells in response to ER stress was shown by two complementary biochemical and immunofluorescence approaches [15]. Subsequently it was demonstrated that the ER-resident chaperone BiP (GRP78) facilitates the translocation of CLU to the cytosol during ER stress in human prostate cancer LNCaP cells [16]. In addition, it was recently shown that during ER stress in PC3 and LNCaP cells, cytoplasmic CLU interacts with LC3 to promote autophagy [17]. While the mechanism by which CLU translocates from the ER lumen to the cytosol in response to stress is yet to be elucidated, it appears that the release may form a part of a larger strategy in which many different chaperones are released from the ER to the cytosol to defend the cell against intracellular stresses. This has been suggested by the demonstration that, in response to cytosolic foreign DNA or inhibition of the proteasome, BiP is released to the cytosol [18]. Other earlier studies have also suggested that CLU might be involved in stress responses within the cell. For example, CLU accumulates in cells when either lysosomal degradation or the proteasome are specifically inhibited [19], and CLU is itself ubiquitinated during ER stress (a process that can only occur in the cytosol) [15]. Furthermore, other studies have suggested that CLU promotes ubiquitination and proteosomal degradation of the cytosolic proteins, COMMD1 and IkB [20] and the lysosome-mediated degradation of the cell-surface copper transport proteins, ATP7A and ATP7B [21, 22]. Collectively, the previous literature points to a potential involvement of CLU, especially during ER stress, in facilitating the degradation of specific cell proteins via both of the primary cell degradative pathways, the proteasome and autophagy.
The pathologies of many neurodegenerative diseases are known to involve both ER stress and cytotoxic protein aggregation [23]. Consequently, we examined whether increased expression of the unusually potent chaperone CLU could provide protection against intraneuronal protein aggregation and proteotoxicity, especially under conditions of ER stress. As a primary model, we studied the aggregation of the 43 kDa transactive response DNA-binding protein (TDP-43), which is implicated in the pathology of amyotrophic lateral sclerosis (ALS), where it translocates from its normal location in the nucleus and forms insoluble inclusions in the cytoplasm. We investigated the ability of CLU to influence the aggregation and clearance of TDP-43 in vitro, in neuronal cells and in transgenic Drosophila melanogaster. The results indicate that CLU can interact with TDP-43 and potently inhibit its aggregation, and that in a neuronal cell model undergoing ER stress, CLU reduces the numbers of cytoplasmic TDP-43 inclusions. We further show in transgenic Drosophila that co-expression of CLU reduces cytoplasmic mislocalization of TDP-43 in motor neurons, protects against loss of locomotor activity and significantly extends lifespan. Lastly, we showed that in Drosophila photoreceptor cells, CLU gave ER stress-dependent protection against proteotoxicity arising from both Huntingtin-Q128 and mutant (R406W) human tau.
Materials and methods
Cell culture and in vitro methods
Transfection, immunocytochemistry and confocal imaging of mammalian cells
SH-SY5Y, N2a and U251 cells were grown in DMEM/F12 (Life Technologies) supplemented with 10% (v/v) foetal calf serum (Bovogen) either on glass coverslips in 12 well microplates or 8 well glass bottom μ-Slides (Ibidi). When cells were 60-80% confluent they were transfected as indicated with either pCAG-EGFP/RFP (encoding the wild-type TDP-43-tdTomato fusion protein (Addgene plasmid 28,205) using X-tremeGENE HP (Roche), pRc/CMV-HT7 (encoding human CLU; [24]), pEGFP-N1-TDP-CTF (encoding a ~ 20 kDa C-terminal fragment (residues 216-414) of human TDP-43 fused to enhanced green fluorescent protein (EGFP); Addgene plasmid #28197) or pCMV6-AC-(M337 V)TDP-43-tGFP (Origene; M337 V mutation introduced into the wild type human TDP-43 sequence, fused at the C-terminus to turboGFP) using Lipofectamine 2000 (Life Technologies) according to the manufacturer’s instructions. 48 h after transfection, cells were treated (or left untreated) for 10 h with 2.5 μM A23187, 2.75 μM Tg and/or 10 μM MG132 (all from Sigma). The cells were immunostained for CLU as follows. Cells were first chemically fixed by incubation for 15 min on ice in 4% (w/v) paraformaldehyde in phosphate buffered saline (PBS; 135 mM NaCl, 10 mM Na2PO4, 2.7 mM KCl, 1.75 mM KH2PO4, pH 7.4), then permeabilized by a 20 min incubation on ice in 0.5% (v/v) TX-100 in PBS. Mouse hybridoma culture supernatants containing IgG1 G7 (anti-human CLU) or DNP9 (anti-2,4-dinitrophenyl) monoclonal antibodies [11] (both diluted 1:2 in 1% w/v bovine serum albumin (BSA) in PBS) were used as primary antibodies. These were detected using goat anti-mouse Ig conjugated with Alexa Fluor-488 or Alexa Fluor-555 IgG (ab150113 and ab150114, Abcam) (2 μg/ml). The nuclei were then stained with RedDot2 (Biotium) according to the manufacturer’s instructions. The cells were washed with PBS after each staining step. Cells grown on coverslips were mounted on a glass slide using Citifluor™ CFPVOH and AF100 anti-fadent (ProSciTech). Imaging was performed on a Leica TCS SP5 II confocal microscope using Leica Application Suite Advanced Fluorescence version 2.6.1-7314. Sequential excitation was performed using 488 nm, 561 nm and 633 nm lasers and fluorescence emissions collected at 500-540 nm (for the 488 nm laser), 570-620 nm (for the 561 nm laser) and 650-750 nm (for the 633 nm laser). In co-localization analyses, to determine the Manders’ overlap coefficient, images were first background subtracted using ImageJ. Regions of interest were then drawn around the cells to exclude pixels lacking intensity in both fluorescence channels (zero - zero pixels) and the extent of co-localization was quantified using the Coloc 2 function in ImageJ with Costes thresholding.
Immunoprecipitation of CLU from N2a cell lysates
N2a cells were transfected to express TDP-43M337V-tGFP (Origene) and human CLU using Lipofectamine 2000 transfection reagent according to the manufacturer’s instructions (Life Technologies). Some cells were transfected to express only TDP-43M337V-tGFP or CLU. Cells (200,000 in all cases) were harvested with trypsin/EDTA and washed twice with PBS (300 x g, 5 min) before being lysed on ice for 5 min in 150 μl PBS containing 1% (v/v) TX-100 and Complete Protease Inhibitor Cocktail (Roche). Any insoluble material was pelleted (21,000 x g, 15 min, 4 °C) and the cleared lysate (130 μl) was gently mixed overnight at 4 °C with Sepharose beads (~ 20 μl packed volume) coupled with mouse monoclonal G7 anti-CLU antibody. In some cases purified human CLU or BSA (100 nM) was added directly to the lysate immediately prior to the addition of the Sepharose beads. The beads were then washed 4 times in PBS (10 min, 2000 x g, 4 °C) before the bound proteins were eluted from the beads by boiling for 5 min in 25 μl SDS sample buffer. The beads were removed by centrifugation (2000 x g, 10 min) and the eluted proteins (~ 20 μl) analysed by Western blotting for the presence of TDP-43 as described below.
Western blotting
Proteins electrophoresed through a 10% SDS-polyacrylamide gel were transferred onto a nitrocellulose membrane (Sartorius) at 4 °C overnight (20 V) using a Western Transfer Unit (BioRad). The membrane was then blocked for 1 h at room temperature (RT) with blocking buffer (5% (w/v) skim milk in PBS containing 0.1% (v/v) TX-100). The membrane was then incubated with TARDBP monoclonal antibody (clone 2E2-D3, Abnova; 1:500) followed by an HRP-conjugated goat-anti-mouse IgG antibody (DAKO; 1:2000). Each antibody was diluted into blocking buffer, and each incubation was followed by washing the membrane 3X with PBS containing 0.1% (v/v) TX-100, followed by 3X washes with PBS. Bound antibodies were detected using Supersignal West Pico Chemiluminescence Substrate (Thermo Fisher Scientific), according to the manufacturer’s instructions. Bands were detected using X-ray film (Amersham Hyperfilm; GE Life Sciences).
Measurement of TDP-43 inclusions in N2a cells
This was done essentially using the method described in [25], which uses flow cytometric analysis to determine the transfection efficiency, and in cell lysates enumerate (i) fluorescent inclusions formed by the aggregation of fluorescent fusion proteins, and (ii) concurrently, the number of cell nuclei. This allows calculation of the number of fluorescent protein inclusions per 100 transfected cell nuclei. N2a cells grown in 24 well plates were transfected to express TDP-43M337V-GFP and cultured for 10 h with or without thapsigargin (2.75 μM), A23187 (2.5 μM), MG132 (10 μM), chloroquine (50 μM), bafilomycin A1 (100 nM), U0126 (60 nM), rapamycin (1.5 μM), or combinations of the preceding. Cells were then harvested using 0.5% trypsin/EDTA, washed and resuspended in 500 μl of PBS. To estimate transfection efficiency, a 150 μl aliquot of the cell suspension was analysed, together with an aliquot of untransfected N2a cells, by flow cytometry using an LSRFortessa X-20 (BD Bioscience). GFP fluorescence was measured using 488 nm excitation and 525/50 nm emission. The remaining cells were lysed in 0.5% (v/v) Triton X-100 in PBS, containing RedDot2 (1:1000; Biotium) to stain nuclei and Complete protease inhibitor (Roche). The lysate was analyzed by flow cytometry measuring forward and side scatter, and the fluorescence of GFP (acquired as above) and RedDot2 (640 nm excitation, 670/30 nm emission); data was analyzed as described above.
In vitro aggregation assay for TDP-43 peptide
CLU was purified from human plasma obtained from Wollongong Hospital (Wollongong, NSW, Australia) as previously described [11]. The synthetic peptide corresponding to residues 286-331 of TDP-43 (TDP-43286-331) was obtained from China Peptides and Thioflavin T (ThioT) from Sigma. TDP-43286-331 was dissolved in MilliQ water (adjusted with NaOH to pH 11.3) followed by the addition of a one tenth volume of buffer (1 M NaCl, 0.5 M HEPES, pH 7.4), to give a final concentration of approximately 448 μM TDP-43286-331. This solution was then diluted with an equal volume of 100 mM NaCl, 50 mM HEPES, pH 7.4, containing other additions (or not) to give a final concentration of 224 μM TDP-43286-331 with or without 2 μM SOD1 or 0.22, 0.022, 0.011, or 0.008 μM CLU. These solutions were incubated at 37 °C whilst shaking for 16 h in a 384 well microplate. ThioT (20 μM) was added to each well prior to incubation in a FLUOstar OPTIMA (BMG LABTECH) with an excitation filter of 440 +/− 10 nm and an emission filter of 490 +/− 10 nm. SOD1 was also incubated without the addition of TDP-43 and no significant change in fluorescence emission was detected over the time course (data not shown).
In vitro translation and aggregation assay for TDP-43-tGFP
cDNA encoding a human TDP-43-tGFP construct was cloned into pT7CFE1-CHis (Thermo Fisher Scientific). Both the cloning and the verification of insertion by sequencing were performed by GenScript. Full length human TDP-43-tGFP was expressed using the TnT® T7 Quick Coupled Transcription/Translation System (Promega) in a POLARstar Omega plate reader (BMG LABTECH) according to the manufacturer’s instructions (30 °C for 90 min). After the reaction was complete a final yield of 43 nM was estimated (based on manufacturer’s specifications) and the mixture was centrifuged to remove any aggregated material (16,600 x g, 10 min, 4 °C). The supernatant was incubated at either 4 °C or 37 °C for 5 h to induce aggregation with or without the addition of CLU or BSA (both at 43 nM). Following this incubation the samples were centrifuged again as above to pellet any aggregated protein. The supernatant containing any soluble TDP-43-tGFP was collected and 5 μl was diluted 1:1 with 2X SDS sample buffer supplemented with 1% β-mercaptoethanol and boiled for 5 min. The proteins were separated by SDS-PAGE and analysed by Western blot, probing for TDP-43, as described above.
Transgenic Drosophila methods
Generation of transgenic Drosophila
Drosophila expressing HA-tagged human TDP-43 were created previously [26]. Following the same procedure, the human CLU construct was codon optimised for expression in Drosophila and synthesised by Genscript, then cloned into the multiple cloning site of pUAST-AttB. Differences in expression of the constructs that could arise from their integration at different genomic loci were eliminated as the vectors contain sites for exploiting the PhiC31 system for site-specific integration of transgenes [27]. CLU expression was under the control of the UAS-GAL4 system [28]. All injected constructs, including an empty pUAST plasmid for the control line, were incorporated at the same genomic locus (51D) on Chromosome II (Bestgene Inc.). All Drosophila lines were made isogenic by repeated backcrossing. Htt-Q128 and Htt-Q72-GFP flies were a gift of Hyung Don Ryoo (NYU). All other Drosophila stocks used were obtained from Bloomington Stock Centre.
Hemolymph extraction
Drosophila were decapitated and the bodies were placed with the thorax pointing downwards into a p200 pipette tip. Four decapitated Drosophila were placed sequentially into each tip and three tips were placed in to a 1.5 ml Eppendorf tube on ice. The Eppendorf tube was centrifuged for 15 min at 4000 x g in an Eppendorf desk-top microfuge at 4 °C. Approximately 1 μl of straw-coloured hemolymph was collected from each Eppendorf tube and was flash frozen in liquid nitrogen and stored at −80 °C. The total protein concentration of each hemolymph sample was determined by bicinchoninic acid microprotein assay and Western blot detection was carried out using 20 μg of hemolymph protein loaded per lane.
Deglycosylation of head lysate proteins
1-20 μg of protein (hemolymph or Drosophila head homogenate) was diluted in 20 μl H2O. To this solution 2 μl Nonidet P40 (NP40), 2 μl G7 buffer and 1 μl PNGaseF (all reagents from NEB deglycosylation kit) were added according to the manufacturer’s instructions. After gentle agitation for 6 h at RT, the samples were analysed by Western blotting under non-reducing conditions. These conditions are sufficient to remove virtually all sugars [29].
Western blot analysis of fly head homogenates
Five Drosophila per genotype were decapitated, homogenised in RIPA buffer: [50 mM Tris HCl at pH 8, 150 mM NaCl, 1% (v/v) TX-100, 0.5% (w/v) sodium deoxycholate, 0.1% (w/v) SDS and protease inhibitors (Roche), then centrifuged in an Eppendorf desk-top microfuge at 20000 x g for 30 min in order to pellet insoluble proteins. The pellets were dissolved in denaturing buffer (9 M urea, 1% SDS, 25 mM Tris, 1 mM EDTA) at 4 °C, sonicated at 42000 Hz for 30 s, and centrifuged as above for a further 30 min at 20000 x g to pellet any still insoluble proteins. The supernatant (urea-soluble proteins) was used as the insoluble fraction. Protein samples were separated on 4-12% Bis-Tris gels (Invitrogen) and transferred to PVDF membrane (Millipore). Blots were blocked with 5% (w/v) non-fat milk in 0.05% (v/v) TX-100/PBS, and then incubated with the following primary antibodies: rat anti-HA-biotin, High Affinity (3F10) antibody which reacts with the N-terminal HA-tag on the TDP-43 construct (Roche; 1:1000); rabbit polyclonal anti-TDP-43 antibody (Proteintech; 1:2500); mouse anti-CLU G7 and 41D [11] hybridoma culture supernatant (1:10); rabbit anti-phospho-eIF2alpha (Ser51) (Cell Signalling; 1:1000). Blots were washed 6 times for 10 min with gentle agitation at RT in 0.1% (v/v) TX-100/PBS and then incubated with anti-rat, anti-rabbit or anti-mouse secondary antibodies conjugated to HRP (DAKO; 1:5000). All antibodies were diluted in the blocking buffer specified above. Bands were detected using a SuperSignal® West Pico Substrate kit (Thermo Fisher Scientific).
Detection of UPR
A reporter construct, gift of Hyung Don Ryoo (NYU), was created so as to have an enhanced green fluorescent protein (EGFP) inserted after the IRE-1 splice site in XBP1, so that EGFP would only be in frame after XBP1 had been spliced by IRE-1; the splicing of XBP1 by IRE-1 only occurs when there is activation of the UPR, leading to EGFP expression [30]. We co-expressed this construct with each of TDP-43, Htt-Q128 and mutant (R406W) human tau, using a gmr-GAL4 promoter and homogenized the heads of 10 adult Drosophila per experiment, 24 h following eclosion. These samples were then prepared for Western blot as above. The presence or absence of EGFP (and thus UPR activation) was detected by an EGFP specific antibody (mouse monoclonal anti-EGFP antibody, Abcam; 1:2000) and an anti-mouse Ig-HRP secondary antibody as above. In similar experiments, the Western blots were instead probed with rabbit anti-phospho-eIF2alpha (Ser51) antibody (Cell Signaling Technology; 1:1000) to detect phosphorylated eIF2α, an independent marker of the UPR [31].
Drosophila immunohistochemistry
Third-instar larval imaginal eye discs and adult eyes were dissected in PBS, fixed in 4% (w/v) paraformaldehyde (PFA) in 0.05% (v/v) TX-100 in PBS for 20 min at RT then permeabilised for 20 min at RT in 0.5% (v/v) TX-100/PBS, before blocking for 30 min in 5% (w/v) BSA in 0.05% TX-100/PBS. Subsequently, fixed and permeabilised discs were incubated overnight with rat anti-HA-biotin high affinity antibody (Clone 3F10, Roche) and mouse anti-CLU antibody (G7), as above. Samples were then incubated overnight in streptavidin Alexa Fluor 594 conjugate (Invitrogen; 1:10,000) and anti-mouse Alexa Fluor 488 conjugate (Invitrogen; 1:1000). All antibodies were diluted in the blocking buffer described above. Tissue was counterstained with TOTO-3 (Invitrogen; 1:10,000) diluted in 0.05% TX-100/PBS to detect nucleic acids.
Light and scanning electron microscopy of Drosophila eyes
Light and fluorescence microscopy: Drosophila expressing TDP-43 +/− CLU or Htt-Q72-GFP +/− CLU were crossed with gmr-GAL4 Drosophila (Bloomington Stock ID: 1104 & 8121) and maintained in a temperature and humidity controlled incubator at 25°C and 70% humidity. Images were taken of 1-day-old transgenic offspring using 7.5X objective and a Leica epifluorescence microscope; imaginal eye disks prepared as above were imaged using a Leica SP confocal microscope. Scanning electron microscopy: Samples were prepared by fixing whole adult Drosophila overnight in 2.5% glutaraldehyde in 0.1 M PBS (pH 7.4) at 4°C, then dehydrated with an ethanol series. Finally, samples were mounted on stumps and sputter coated using 20 nM Au/Pd in a Polaron E5000. SEM images were collected using a Philips XL30 microscope at 200× magnification.
Climbing assays
Motor function was assessed by a negative geotaxis assay. Drosophila were generated that expressed TDP-43 (with and without CLU), or CLU alone, in motor neurons from the day of hatching. Non-transgenic Drosophila were also tested as a control group. For each treatment group, three vials each containing ten Drosophila were analysed every second day. A climbing index score was calculated as described previously [32]. The average climbing index for the three replicate analyses was calculated for each time point and plotted against time since eclosion (n = 30).
Survival assays
Drosophila were generated at 18°C where Gal80 inhibits GAL4 dependent transcription, thus preventing expression of transgenes in embryos and larvae. Adult Drosophila were moved from 18°C to 29°C (Gal80 is inactive at this temperature and so no longer inhibits expression of transgenes) within 24 h of eclosion; they were then transferred to fresh food and counted every 2-3 days. Gal80; D42-GAL4 is activated by heat shock at 29°C and induces expression in motor neurons. Each treatment group was comprised of 90 non-virgin female Drosophila maintained in glass vials (10 per vial). Median survivals were calculated using Kaplan Meier survival statistics and differences between genotypes were analysed using a Mann-Whitney U test.
Results
ER stress induces CLU to co-localize with cytoplasmic TDP-43-GFP inclusions
Previous studies have shown that ER stress induces release of CLU to the cytosol [15–17]. We confirmed that, as expected, CLU was detected in the cytosol fraction prepared from N2a cells chemically treated to induce ER stress but not in the same fraction prepared from untreated cells (Additional file 1: Figure S1). Furthermore, analysis of the anterior horn cells (motor neurons) in human post-mortem spinal cord sections of three ALS patients and three age-and sex-matched control cases, revealed a striking difference in CLU staining, consistent with the release of CLU from the ER in ALS affected neurons (Additional file 1: Figure S2). In control cases CLU staining mirrored Nissl substance staining, consistent with its localization in the ER (secretory pathway). However, in the anterior horn cells of ALS cases, which exhibited prominent pathological cytoplasmic TDP-43 inclusions, CLU staining was increased and exhibited a more diffuse staining pattern. The pattern of CLU staining in these cells no longer mirrored the distribution of Nissl substance, consistent with a change in the subcellular localization of CLU (Additional file 1: Figure S2).
We next examined the effects of CLU over-expression in a cell model of protein mislocalization relevant to ALS. TDP-43 is a member of the hnRNP family of RNA binding proteins whose functions involve the regulation of RNA splicing and transcription. In almost all cases of sporadic and also in some familial cases of ALS, this predominantly nuclear protein becomes mislocalized in motor neurons to the cytoplasm, where it forms ubiquitinated and hyper-phosphorylated inclusions enriched in proteolytically generated C-terminal fragments of TDP-43 [33]. When N2a (or SH-SY5Y neuroblastoma) cells were transiently transfected to express an aggregation-prone C-terminal fragment of TDP-43 fused to GFP (TDP-43CTF-GFP), which forms aggregates with structural similarity to those in human post-mortem tissue and that is widely used as a tool to study TDP-43 aggregation [33], they developed cytoplasmic inclusions readily detected by confocal microscopy. Furthermore, consistent with ER stress inducing release of CLU to the cytosol, CLU was found co-localized with TDP-43CTF-GFP inclusions in ER stressed N2a cells, but not in unstressed N2a cells (Fig. 1a; Additional file 1: Figure S3). ER stress also induced co-localization of CLU with inclusions formed by a similar fusion protein (mutant M337 TDP-43 fused at the C-terminus with GFP; TDP-43M337V-GFP) in U251 human glioblastoma cells (Additional file 1: Figure S3). We also used co-immunoprecipitation analysis to show that soluble TDP-43M337V-GFP is physically associated with CLU in lysates prepared from co-transfected N2a cells, for both untreated cells (Fig. 1b) and cells treated to induce ER stress (not shown). Interestingly, we also used a monoclonal antibody specific for human CLU to show that exogenous human CLU added (“spiked”) to lysates of transfected N2a cells co-immunoprecipitated with TDP-43M337V-GFP. These results demonstrate that when CLU and TDP-43M337V-GFP are both present in cell lysates they can form complexes, regardless of whether or not the two proteins may have been in different cell compartments prior to lysis. Thus, while it is not possible to use a co-immunoprecipitation approach to verify the occurrence of CLU-TDP-43 complexes in situ inside cells, the results indicate that if the two proteins are present within the same compartment, such as suggested by the confocal images of cytoplasmic co-localization, they are capable of forming complexes.
Fig. 1.
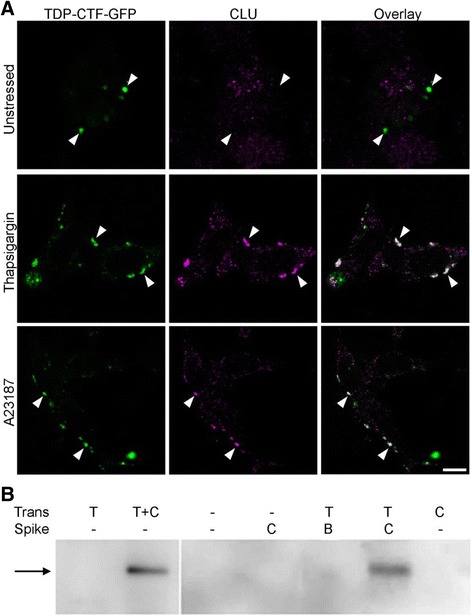
CLU co-localizes with cytoplasmic TDP-43 inclusions in ER stressed cells and co-precipitates with TDP-43-GFP. a N2a cells transfected to express TDP-43CTF-GFP and human CLU (hCLU) were treated as indicated, fixed/permeabilized, then immunostained for hCLU. White arrowheads indicate the positions of inclusions. Overlay panels (right): where there is no colocalization, inclusions appear green; colocalization of TDPCTF-GFP and CLU appears as white. Scale bar is 10 μm; images shown are representative of many. Manders overlap coefficients (TDP-43CTF-GFP and CLU), calculated using at least 20 cells for each treatment, were: untreated, 0.13 +/− 0.14; Tg, 0.62 +/− 0.07; A23187, 0.52 +/− 0.09 (mean +/− SD). See also Additional file 1: Figure S3. b TDP-43 immunoblot showing that TDP-43M337V-GFP co-precipitated with hCLU from lysates of co-transfected N2a cells and also when exogenous purified hCLU (but not BSA) was added (“spiked”) to lysate prepared from N2a cells transfected to express only TDP-43M337V. The black arrow indicates the expected size of TDP-43M337V-GFP. Key above image indicates which protein(s) cells were transfected to express (Trans), and the protein added to the lysate (Spike). The result shown is representative of two independent experiments
CLU potently inhibits TDP-43 aggregation in vitro
Full length TDP-43 is so aggregation-prone that it is extremely difficult to express the full length protein in bacteria as a soluble recombinant product in significant quantities [34]. We therefore initially made use of a well-characterised 46-mer peptide corresponding to a region of the C-terminal domain of TDP-43 (TDP-43286-331), which is known to form fibrillar aggregates [35], to probe the ability of CLU to inhibit the in vitro aggregation of TDP-43. Under the conditions tested, following a 2 h lag phase, the TDP-43286-331 peptide rapidly formed ThioT positive aggregates. Under these same conditions, a CLU:peptide ratio of 1:1000 completely inhibited the appearance of ThioT positive species over a 16 h time course. Even at a 1:25,000 ratio, CLU retarded TDP-43286-331 fibril formation by about 2 h (Fig. 2a). This effect is specific to CLU, as even when used at a ratio of 1:500, a non-chaperone control protein (bovine superoxide dismutase 1, SOD1) had negligible effects on aggregation of the peptide (Fig. 2a). In order to confirm that CLU can also suppress the aggregation of full-length TDP-43, we adapted an in vitro protein expression system for full-length TDP-43-turboGFP (TDP-43-tGFP) described in [34]. Following in vitro expression, a significant amount of full-length TDP-43-tGFP remained soluble for 4 h at 4°C, but after a 4 h incubation at 37°C in the presence of 43 nM BSA (a non-chaperone control protein), very little TDP-43-tGFP remained soluble (Fig. 2b). When incubated for 4 h at 37 °C, the presence of 43 nM CLU resulted in an 8-fold higher level of TDP-43-tGFP remaining in solution (Fig. 2b), demonstrating that CLU is able to very significantly inhibit the aggregation of full-length TDP-43 in vitro.
Fig. 2.
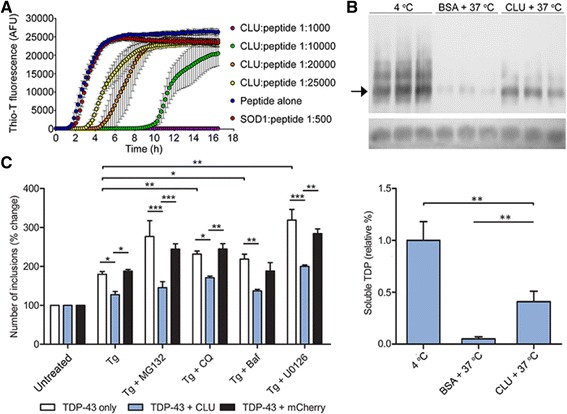
CLU inhibits TDP-43 aggregation and reduces the number of TDP-43-GFP inclusions in neuronal cells during ER stress. a TDP-43286-331 (peptide) was incubated at 37 °C with shaking, in the presence of either CLU or the non-chaperone control protein bovine superoxide dismutase (SOD1), at the molar ratios indicated in the key. Peptide aggregation was monitored by ThioT fluorescence. Data are means +/−SE (n = 4). b Upper panel: Image of Western blot detecting soluble TDP-43-tGFP after incubation at 4 °C, or 37 °C (which increases aggregation), with or without CLU or BSA (a non-chaperone control protein). Black arrow indicates position of monomeric TDP-43-tGFP, upper bands represent soluble oligomers. Loading control (colored dye from Promega transcription/translation kit) shown at bottom. Lower panel: Densitometric quantification of blot results (means + SD, n = 3). c Transfected N2a cells were treated as indicated for 10 h (key below plot), then lysed and TDP-43M337V-tGFP inclusions enumerated by flow cytometry; data plotted as % change relative to untreated cells (means + SEM, n = 4; untreated cells are defined as the 100% value). See also Additional file 1: Figure S5. For all panels, results shown are representative of two or more independent experiments. In (b) and (c), horizontal bars indicate pairwise comparisons, * p < 0.05, ** p < 0.01, *** p < 0.001 (1-way ANOVA with Newman-Keuls post-test (b), or 2-way ANOVA with a Bonferroni post-test (c))
CLU expression specifically reduces the numbers of cytoplasmic TDP-43-GFP inclusions in ER stressed N2a cells
We used a recently developed flow cytometry method, which counts fluorescent particles in cell lysates and normalizes this count against the number of separately enumerated nuclei (FloIT; [25, 36, 37]) to quantify the numbers of TDP-43M337V-GFP inclusions in differently treated N2a cells. The results indicated that (i) in unstressed cells, overexpression of CLU or mCherry had no significant effect on the numbers of TDP-43M337V-GFP inclusions, (ii) the numbers of inclusions were increased in cells treated with either thapsigargin (Tg) alone (to induce ER stress), or with Tg and MG132 (the latter to also inhibit the proteasome), and (iii) overexpression of CLU, but not mCherry, reduced the numbers of TDP-43M337V-GFP inclusions in ER-stressed cells (Fig. 2c). CLU overexpression also specifically reduced the number of TDP-43M337V-GFP inclusions in N2a cells in which ER stress was induced by the addition of Ca2+ ionophore A23187 (Additional file 1: Figure S5A). The overexpression of CLU or mCherry had no significant effect on the numbers of inclusions in unstressed cells (Additional file 1: Figure S5B).
Human CLU is glycosylated, cleaved and secreted when expressed in Drosophila neurons
Given the ability of CLU to reduce the number of TDP-43 inclusions in cultured neuronal cells, we next developed a whole organism model in which the protective effects of CLU against cellular proteotoxicity could be further investigated. Drosophila has no identifiable homologue of clusterin. When expressed in photoreceptor neurons, human CLU was detected in juxtanuclear puncta distributed uniformly across the eye disc (Fig. 3a). This distribution is consistent with the protein being localised in the secretory pathway and analysis of the hemolymph (the circulatory fluid of Drosophila) confirmed that CLU is secreted from the neurons in which it is being expressed (Fig. 3b). Non-reducing SDS-PAGE and Western blot analysis of head lysates from Drosophila expressing CLU revealed that the protein was abundantly expressed and migrated as a single band with an apparent molecular mass of ~ 55 kD; it migrated at a lower molecular mass (~ 50 kD) when the sample was treated with PNGase to remove virtually all glycosylation (Additional file 1: Figure S6A). The molecular mass of CLU detected in the hemolymph and head lysates of Drosophila is, however, less than that of CLU purified from human plasma (~ 55 kD versus 75-80 kD), a difference attributable to the fact that insect cells glycosylate proteins to a lesser extent than mammalian cells [38]. Indeed, enzymatic deglycosylation of CLU expressed in Drosophila results in a smaller loss in apparent mass than for CLU from human plasma, and the deglycosylated forms of both proteins migrate at a very similar position when analyzed by SDS-PAGE (Additional file 1: Figure S6A). Differences in glycosylation do not, however, significantly affect the function of CLU, as indeed the removal of virtually all the attached carbohydrates does not inhibit its chaperone activity [29]. We also confirmed by SDS-PAGE that Drosophila-expressed CLU dissociates into α- and β-subunits when chemically reduced, as seen for CLU from human plasma (Additional file 1: Figure S6B). A faint CLU band detected at about 50 kDa may represent a minor glycoform of CLU, as glyco-variants of CLU are found in human plasma [39]. Taken together, these biochemical studies indicate that Drosophila neurons are able to synthesise and secrete a mature, post-translationally modified form of human CLU.
Fig. 3.
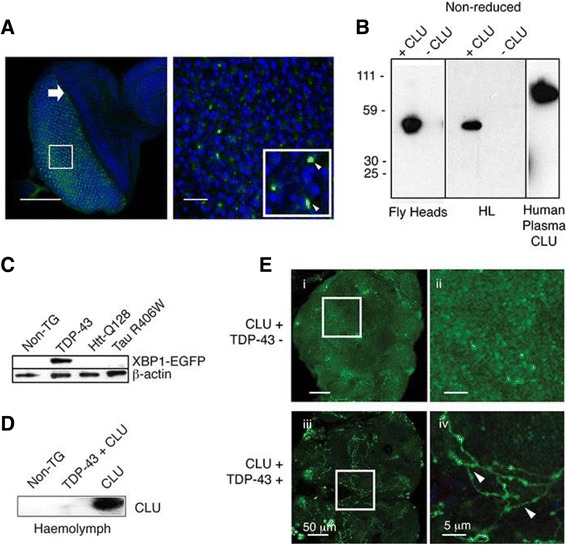
Expression of CLU and TDP-43 in Drosophila. a Confocal microscopy images of imaginal eye discs of third instar larvae expressing human CLU in the photoreceptor cells; CLU (green), TOTO-3 (blue, detects nuclei). Left panel: Large white arrow indicates the morphogenetic furrow (scale bar 100 μm); white box represents the area shown at higher magnification in the large panel to the right. Right panel: Punctate staining of CLU (scale bar 10 μm); the inset white box shows a region with still higher magnification (arrowheads indicate CLU). b Non-reducing Western blots of Drosophila head homogenates, hemolymph (HL), and the positive control (human plasma CLU); expression of CLU indicated above the lanes. c Western blot of whole Drosophila head lysates, prepared from non-transgenic (Non-TG) Drosophila, or Drosophila expressing TDP-43, Htt-Q128 or Tau R406W; detection of XBP1-EGFP indicates activation of the UPR (β-actin was used as a loading control). d Western blot detecting CLU in hemolymph taken from Non-TG Drosophila, and those expressing CLU +/− TDP-43; equal amounts of hemolymph were loaded in each lane under non-reducing conditions. See also Additional file 1: Figure S6. e Confocal microscopy images of the central lobe of 10-day-old, whole adult Drosophila brains expressing CLU +/− TDP-43 in the motor neurons. Images in (i) and (iii) encompass the entire central lobe; the central region in each image contains neuronal process and axons. Images in (ii) and (iv) represent optical zooms of the areas indicated by the white boxes in (i) and (iii), respectively. Scale bars in (i) and (iii) represent 50 μm, while those in (ii) and (iv) represent 5 μm. CLU (green), white arrowheads in (iv) indicate axons. Results shown are representative of two or more independent experiments
TDP-43 expression induces ER stress and cellular retention of CLU in transgenic Drosophila
We first co-expressed both TDP-43 and a fluorescent reporter of the unfolded protein response (an EGFP-based reporter of the alternative splicing of XBP1 [30]) in photoreceptor neurons of the Drosophila eye. For comparison, we also examined the effects of expression both of exon 1 of the Htt gene, containing a 128 glutamine residue expansion (Htt-Q128; [40]), and also of mutant (R406W) human tau, which is associated with frontotemporal dementia with parkinsonism-17 (FTDP-17) [41]. Co-expression of the fluorescent UPR reporter with TDP-43, Htt-Q128 or R406W tau followed by analysis of EGFP levels by Western blotting, revealed that IRE1-dependent splicing of XBP1-EGFP was induced by expression of TDP-43 but not by the expression of either Htt-Q128 or R406W tau (Fig. 3c). The induction of ER stress by TDP-43 expression was also confirmed by Western blot analysis showing that the levels of phosphorylated eIF2α, an independent marker of the UPR [31], were much higher in homogenates of the heads of Drosophila expressing TDP-43 compared to those of control Drosophila (Additional file 1: Figure S6C). Western blotting was also used to confirm that CLU was secreted into the hemolymph when expressed alone in photoreceptor neurons, but that it was barely detectable in this fluid when co-expressed with TDP-43 in the same cells (Fig. 3d). We next expressed CLU in Drosophila motor neurons and examined its distribution in brains by confocal microscopy. Despite the limited resolution of confocal images obtained from the relatively thick adult Drosophila brain, it can be seen that, consistent with its secretion, CLU was diffusely distributed throughout the neuropil (Fig. 3e i & ii), and was not co-localized with neuronal cell bodies or axons. When co-expressed with TDP-43, however, CLU was found at high levels in the axons (Fig. 3e iii & iv).
CLU restores nuclear localization of TDP-43 in adult Drosophila motor neurons
In the motor neurons of 10 day old adult Drosophila brains, when expressed alone, TDP-43 is frequently notably absent from its normal nuclear localization (elipses devoid of TDP-43; Fig. 4a i & ii) and is instead detected in foci localized within the cytoplasm of the motor neurons (white arrowheads; Fig. 4a i) as well as being diffusely distributed throughout the soma, axons and neuronal processes (white arrowheads; Fig. 4b i & ii). This observation is consistent with those made in yeast models in which high levels of expression of wild-type TDP-43, as achieved in the current Drosophila model, result in its mislocalization into the cytosol [42]. When CLU is co-expressed with TDP-43, however, the latter is no longer evident within the axons or neuronal processes and paranuclear foci of TDP-43 are no longer visible. Instead, TDP-43 is predominantly found within the nucleus (Fig. 4a iii & iv) where it strikingly co-localizes with CLU (Fig. 4c). At earlier stages of development, when the two proteins have been co-expressed for shorter periods of time, TDP-43 is largely absent from the nucleus and instead is found mainly in the cytoplasm where it extensively co-localises with CLU (Fig. 4d). Thus CLU co-expression results in the restoration in the adult Drosophila of a predominantly nuclear localization for TDP-43 and reduces the levels of both soluble and insoluble TDP-43 detected in head lysates (Fig. 4e). This effect cannot be attributed to lowered expression of TDP-43 arising from transgene co-expression, as co-expression of TDP-43 with GFP (which has no chaperone activity) has no effect on TDP-43 levels (Additional file 1: Figure S7A).
Fig. 4.
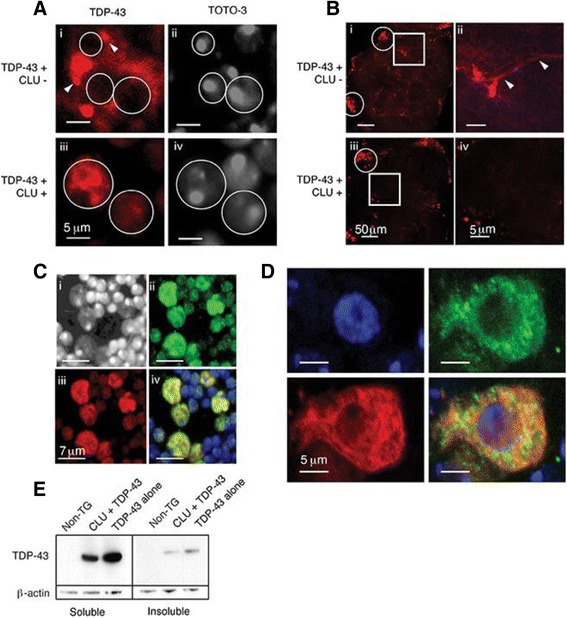
CLU expression reduces TDP-43 mislocalization in Drosophila neurons. Confocal microscopy images of: a Motor neuron cell bodies of 10-day-old adult Drosophila brains expressing TDP-43 alone (i & ii) or together with CLU (iii and iv). Images (i & ii) and (iii & iv) represent the same fields of view; TDP-43 (red; i & iii), TOTO-3 stained nuclei (grey; ii & iv). In (i), white arrowheads indicate paranuclear foci; in all panels white elipses indicate nuclei (scale bars 5 μm). b Central lobe of adult Drosophila brains expressing TDP-43 +/− CLU. Images in (i) and (iii) were collected using a 40X objective and encompass the entire central lobe (scale bars 50 μm); elipses show motor neuron cell bodies and the central region of each image contains neuronal process and axons. Images in (ii) and (iv) represent optical zooms of the areas indicated by the white boxes in (i) and (iii), respectively (scale bars 5 μm). TDP-43 (red), white arrowheads in (ii) indicate axons. c Motor neuron cell bodies of adult Drosophila brains co-expressing CLU (green) and TDP-43 (red); TOTO-3 (white; nucleoli stain intensely, the rest of the nucleus stains less brightly); (iv) overlay of all three images, yellow indicates co-localisation of CLU and TDP-43 (scale bars 7 μm). d Motor neuron cell bodies of third instar larval Drosophila brains co-expressing CLU (green) and TDP-43 (red), TOTO-3 (blue); (iv) overlay of all three images; yellow indicates co-localisation of CLU and TDP-43 (scale bars 5 μm). e Reducing Western blot of the soluble and insoluble fractions of 10-day-old adult Drosophila head homogenate comparing non-transgenic (non-TG) Drosophila and Drosophila expressing TDP-43 +/− CLU. β-actin was used as a loading control. Results shown are representative of several independent experiments
CLU protects Drosophila motor neurons from TDP-43-induced neurotoxicity
We next investigated whether CLU expression could affect the neurodegenerative phenotypes resulting from intracellular expression of TDP-43 in Drosophila motor neurons. Strikingly, co-expression of CLU with TDP-43 in the motor neurons (i) significantly delays the onset of locomotor dysfunction, increasing the time taken for 50% of TDP-43 expressing Drosophila to become immobile from 5 days to 15 days (Fig. 5a), and (ii) increases the median survival time by 33% compared to Drosophila expressing TDP-43 alone (from 15 ± 0.39 days to 20 ± 0.53 days; p = 0.0006, n = 180; Fig. 5b). This rescue is specific to CLU expression, as co-expression of an unrelated protein (GFP; not expected to bind TDP-43) driven by the same UAS-GAL4 system did not affect the median survival of TDP-43-expressing Drosophila (Additional file 1: Figure S7B). Furthermore, CLU expression alone did not cause a significant increase in the longevity of Drosophila. Non-transgenic Drosophila have a median survival of 36 days ±1.09 whereas CLU expressing Drosophila have a median survival of 37 days ±1.18 (p = 0.5056 ns, n = 180; Additional file 1: Figure S7C), indicating that the effects of CLU are on neurotoxicity induced by TDP-43 rather than representing a general effect on ageing.
Fig. 5.
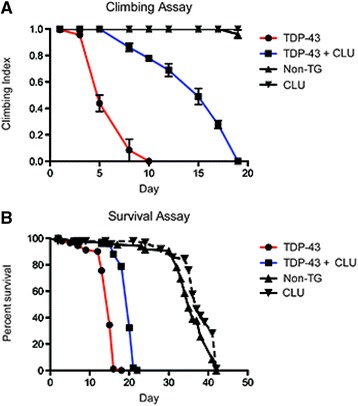
CLU expression reduces TDP-43 toxicity in Drosophila motor neurons. Time-dependent differences between Drosophila expressing TDP-43 +/− CLU in (a) motor function, measured by climbing assay, and (b) survival, compared to non-transgenic (Non-TG) Drosophila. In (a) and (b), values shown are means +/− SEM, and in each case the results shown are representative of three independent experiments. Parent lines used in crosses indicated in key. Differences in climbing index between genotypes were analysed by ANOVA (n = 30). Lifespans were analysed by Kaplan Meier statistics (n = 90). See also Additional file 1: Figure S7
CLU-mediated protection against intracellular proteotoxicity is not restricted to TDP-43 and is dependent on ER stress
We used the “rough eye” assay as a widely accepted tool to assess neurotoxicity in Drosophila models, to test of the effects of CLU expression on a variety of proteotoxic stresses. The gmr-GAL4 promoter was used to express TDP-43 in Drosophila photoreceptors, resulting in neurotoxicity manifested as a depigmentation and structural derrangement of the ommatidia, which was significantly reduced by CLU expression (Fig. 6a). We next expressed two other neurotoxic proteins (Huntingtin-Q128 (Htt-Q128) and mutant R406W human tau), which we had earlier established did not induce ER stress in Drosophila neurons (Fig. 3c). In both these cases, CLU co-expression had no significant effect (Fig. 6a). We reasoned that the lack of protection against proteotoxicity afforded by CLU in these models could relate to its known dependence upon ER stress for release from the ER to the cytosol. To examine this possibility we next expressed in the Drosophila eye the Htt gene (exon 1) with a 72 residue glutamine expansion, which can be readily visualized through its fused EGFP tag (Htt-Q72-EGFP) [43]. We then tested whether CLU co-expression could protect from the resulting aggregation and neurotoxicity during (i) basal conditions, and (ii) chemically-induced ER stress induced by rearing Drosophila on food supplemented with 5 mM DTT.
Fig. 6.
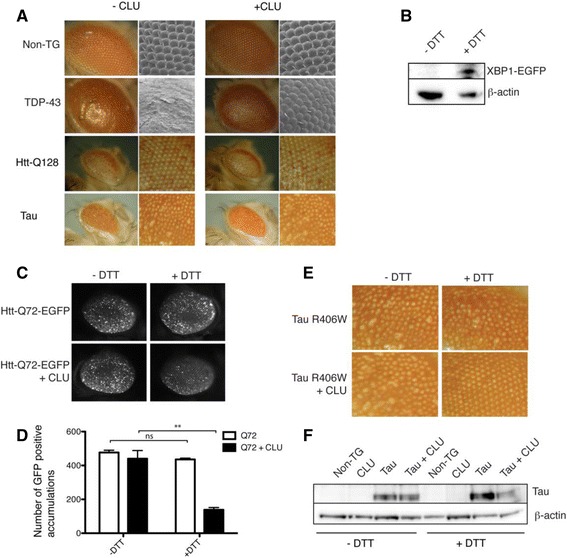
CLU provides ER stress-dependent protection against proteotoxicity. a Light and scanning electron micrographs demonstrating the effects of expression of TDP-43, Htt-Q128 and tau R406W (+/− CLU) in the photoreceptor neurons of adult Drosophila. Light micrographs (left) of Drosophila eyes collected using a 7X objective, electron micrographs (right) taken at 200X magnification. For Htt-Q128 and tau R406W, the images shown on the right are optical zooms of the corresponding images on the left. All images are representative of many experiments. b Western blot of whole non-transgenic Drosophila head lysates prepared from Drosophila fed normal food (−DTT) or food supplemented with DTT (+DTT); detection of XBP1-EGFP indicates activation of the UPR (β-actin was used as a loading control). c Fluorescence micrograph images (collected using a 7X objective) of eyes on Drosophila fed with food +/− DTT (or not), and expressing Htt-Q72-EGFP +/− CLU. d Quantification of the number of individual EGFP accumulations per eye, using images such as those shown in (c) and ImageJ (particle analyser program); **p = 0.0037, n = 9, Student’s t-test. Results shown are representative of several independent experiments. e Light micrographs (collected using a 7X objective) showing the eye phenotype in adult Drosophila fed normal food (−DTT) or food supplemented with DTT (+DTT) resulting from expression of R406W tau +/− CLU in the photoreceptor neurons. f Western blot of Drosophila head homogenates prepared from non-transgenic (Non-TG) Drosophila or Drosophila described in (e)
Western blot analysis of the XBP1-EGFP reporter in Drosophila head homogenates showed that rearing Drosophila on DTT-supplemented food is sufficient to induce ER stress, indicated by induction of the UPR (Fig. 6b). When comparing between Drosophila all co-expressing Htt-Q72-EGFP and CLU, relative to Drosophila fed on standard food, ER stressed Drosophila showed an approximately 70% reduction in the number of fluorescent Htt-Q72-EGFP puncta detected (440.4 ± 47.8 vs 138.4 ± 13.5; respectively; p = 0.0037, n = 9). This effect was absent when Drosophila expressing only Htt-Q72-EGFP were raised on food supplemented with DTT (Fig. 6c&d). Similarly, when Drosophila co-expressing R406W tau and CLU were reared on food supplemented with 5 mM DTT to induce ER stress (but not otherwise), there was a rescue of the rough eye phenotype (Fig. 6e) and a concomitant reduction in the quantity of phosphorylated, insoluble R406W tau deposits detected by SDS PAGE (Fig. 6f). These data indicate that CLU-mediated protection against the aggregation and neurotoxicity of Htt-Q72 and R406W tau is completely dependent upon ER stress (Fig. 6c-f).
Discussion
Although the precise pathway of release remains to be determined, multiple studies now strongly suggest that ER stress triggers the release of post-translationally modified CLU from the ER to the cytoplasm [15–17] (see also Additional file 1: Figure S1). This may well explain the many previous reports that, under a variety of disease conditions, CLU occurs in cellular locations outside the secretory system, including the co-localization of CLU with intracellular tau tangles in Alzheimer’s disease [44], α-synuclein rich Lewy bodies in Parkinson’s disease [45], and inclusion bodies in myofibrillar myopathies [46]. It has also been reported that in response to chemotherapeutic drug treatment, CLU relocates to mitochondria and interacts with Bax to inhibit apoptosis [47]. There is abundant evidence from transgenic mouse models of ALS, neuropathological post mortem studies, and genetic studies that ER stress is a significant component in ALS pathology [48, 49]. While certainly not in themselves definitive, the results of our own immunohistochemical analyses of ALS patient spinal cord tissues are consistent with CLU also being released from the ER in ALS affected neurons (Additional file 1: Figure S2).
Furthermore, the inappropriate mislocalization and accumulation of TDP-43 in cytoplasmic inclusions is a unifying pathology in frontotemporal dementia (FTD) and familial and sporadic ALS cases. It is seen, with the exception of cases resulting from SOD1 mutations [50], in most post-mortem cases of FTD and ALS [51], and correlates well with synaptic pathology and cognitive deficits [52]. Thus, a therapeutic strategy that protects against TDP-43 proteotoxicity could provide benefits in most cases of ALS. Therefore, in the present study we used a variety of complementary approaches, including in vitro, cell and whole animal models, to evaluate whether the unusually potent and multifunctional chaperone CLU can affect the aggregation, cytoplasmic mislocalization and proteotoxicity of TDP-43.
We showed using both a C-terminal peptide corresponding to the most aggregation-prone region of TDP-43, and full length in vitro translated TDP-43-tGFP, that CLU potently inhibits the in vitro aggregation of TDP-43 and maintains it in a soluble form (Fig. 2). These results strongly suggest that CLU can directly interact with TDP-43, as there is no known mechanism by which a chaperone can inhibit protein aggregation without directly interacting with the client protein. The ability of CLU to inhibit the aggregation of the TDP-43286-331 peptide even at molar ratios of CLU:peptide as low as 1:1000 could arise from the fact that only a very small fraction of the soluble peptide molecules may be in a misfolded conformation at any one time, enabling CLU to interact specifically with these species to stabilize them and prevent them acting as aggregation nuclei [10]. We also showed using three different transfected cell models (N2a, SH-SY5Y and U251), that during ER stress, but not otherwise, CLU co-localized with cytoplasmic TDP-43-GFP inclusions (Fig. 1, Additional file 1: Figure S3). The treatments used to induce ER stress in the cell models (MG132, Tg and A23187) have been widely used in other studies [15–18] and we confirmed that even an 18 h exposure of N2a cells to 10 uM MG132 had negligible effects on cell viability (Additional file 1: Figure S4). In N2a cells, we showed that over-expression of CLU specifically and significantly reduced the numbers of inclusions within these cells (Fig. 2c). Co-immunoprecipitation analyses demonstrated that CLU can physically associate with TDP-43CTF-GFP in N2a cell lysates (Fig. 1b), suggesting that once in the cytosol, CLU would be capable of forming stable complexes with TDP-43.
In transgenic Drosophila we showed that overexpression of TDP-43 induced both ER stress and the retention of CLU within both photoreceptor and motor neurons (Fig. 3). Under these conditions, CLU was shown to co-localize with TDP-43 first in the cytoplasm and then, at later stages in development, in the nuclei of adult Drosophila motor neurons, effectively restoring the normal nuclear distribution of TDP-43 (Fig. 4). In this whole animal model, the effects of enforced CLU expression were to generate a dramatic recovery of the loss of locomotor activity associated with TDP-43 proteotoxicity, and a significant extension of lifespan (Fig. 5). These striking effects are specific to CLU expression as we showed that the co-expression of GFP in TDP-43-expressing Drosophila did not alter either the level of TDP-43 expression or the median survival (Additional file 1: Figure S7). Finally, we showed that enforced CLU expression could also protect against the proteotoxicity of Htt-Q72 and R406W tau expressed in Drosophila photoreceptor neurons, but only when ER stress was chemically induced (Fig. 6).
In both the neuronal cell and Drosophila models studied here, ER stress was a consistent requirement for CLU to provide protection against protein mislocalization and toxicity. We cannot exclude the suggestion that over-expression of CLU might act to reduce the levels of experimentally induced ER stress, and this might account in part for the protective effect of CLU expression. However, even if this were correct, our results in transgenic Drosophila strongly suggest that this is not the only mechanism by which CLU over-expression can protect against the cytoplasmic accumulation of toxic proteins. Notably, the level of cytoplasmic accumulation/toxicity of Htt-Q72/R406W tau expressed in cells of the Drosophila eye is unaffected by either DTT treatment alone or CLU co-expression alone, but is significantly reduced only when the two conditions are combined (Fig. 6). Furthermore, in transfected N2a cells, we only measure the release of CLU to the cytosol following treatments to induce ER stress, again strongly suggesting that CLU over-expression does not ablate the induction of ER stress. All our findings are consistent with a model in which ER stress induces release of CLU to the cytosol where it exerts effects that act to reduce net TDP-43 accumulation. There are a several putative mechanisms by which cytosolic CLU could exert the observed effects on TDP-43 mislocalization, aggregation, toxicity and clearance. These include (i) the well characterized chaperone activity of CLU [11, 12, 53] allowing it to directly interact with TDP-43 to form stable, soluble complexes, thereby inhibiting its further aggregation to form cytoplasmic inclusions, (ii) CLU may interact with misfolded species of TDP-43 to neutralize their toxicity, as is the case in its interactions with other toxic protein oligomers [12, 54, 55], and (iii) CLU, which is ubiquitinated once in the cytosol [15], may direct soluble CLU-TDP-43 complexes towards the proteasome or autophagy for degradation. Both the proteasome and autophagy are known to act to clear TDP-43 inclusions [56] and there is a growing body of evidence implicating CLU as a substrate of both these major cellular degradative pathways and a control element of autophagy [15, 19–22]. It is also interesting to note in this context that Cha-Molstad et al. have recently reported that in response to MG132-mediated inhibition of the proteasome, or cytosolic foreign DNA, the ER-resident chaperone BiP is released to the cytosol [18]. We speculate therefore that the release of CLU to the cytosol might form a part of a larger strategy in which multiple chaperones are released from the ER to the cytosol to defend the cell against intracellular stresses.
An important aspect of motor neurons is that they have a very limited heat shock response owing to their impaired ability to activate the key molecular heat shock response molecule, HSF1 [57]. This results in a high threshold for induction of the heat shock response in motor neurons conferring an element of vulnerability. The results presented in this study provide a clear demonstration in a whole organism model that during ALS disease-relevant conditions the enforced expression of CLU provides significant protection against the TDP-43-mediated proteotoxicity underpinning pathology, to substantially enhance motor neuron survival, reduce locomotor deficits and extend lifespan. We have shown previously that constitutively intracellular chaperones are able to rescue TDP-43 associated neurotoxicity [14], however this is the first instance, to our knowledge, where a normally secreted chaperone can exert similar protective effects.
In protein misfolding diseases such as ALS and Alzheimer’s disease (AD) there are multiple proteins misfolding in both the intra- and extracellular environments. For example, pathological, extracellular amyloid beta accumulation is often seen in post-mortem ALS and AD cases exhibiting intracellular accumulation of TDP-43 or tau [58]. It appears feasible that CLU can combat protein misfolding in both the intra- and extracellular compartments and may alter its subcellular localization dependent upon the environmental stress. Previous studies attempting to harness the therapeutic benefits of chaperones in protein misfolding diseases have used Hsp90 inhibitors, such as 17-AAG, which can upregulate small heat shock proteins in the intracellular compartment. 17-AAG can rescue the neurotoxicity associated with the pathological accumulation of TDP-43, polyglutamine, alpha synuclein, mutant androgen receptor, tau and amyloid beta [59–63]. However, 17-AAG also upregulates CLU expression [64], potentially contributing to the neuroprotective effects afforded by the therapeutic intervention in these models. Apart from ALS, many other neurodegenerative diseases including Alzheimer’s, Parkinson’s and Huntington’s diseases, also feature ER stress associated with pathology [23]. Therefore, it is conceivable that further studies investigating temporal changes in CLU levels and localization associated with pathologies could reveal new avenues for developing therapeutic strategies to address currently incurable and highly devastating diseases.
Conclusions
The results presented in this study demonstrate in a whole organism model that during ALS disease-relevant conditions, enforced CLU expression significantly protects against TDP-43-mediated proteotoxicity to substantially enhance motor neuron survival, reduce locomotor deficits and extend lifespan. This is the first instance, to our knowledge, where a normally secreted chaperone has been shown to exert such protective effects in the intracellular context.
Acknowledgements
This work was supported by a Wellcome Trust/MRC strategic award (LML, TMP and CMD), the International Journal of Experimental Pathology (JG), the James Baird fund (JG), the BBSRC (CMD, LML, JRK), Grants-In-Aid from the Motor Neurone Disease Research Institute of Australia (GIA1223 & GIA1335) (MRW, JJY), an Illawarra Health and Medical Research Institute Dementia Research Grant (MRW), a Lloyd Binet PhD Scholarship (Australian Rotary Health) (RAB) and an Australian Government Postgraduate Award (DRW). The authors declare that there are no conflicts of interest.
Additional file
Supplementary Methods. Figure S1. In mouse N2a neuroblastoma cells, endogenous CLU is released from the ER to the cytosol in response to treatment with MG132 and Tg. Figure S2. Spinal cord anterior horn cells of ALS patients contain phospho-TDP-43 inclusions and elevated levels of abnormally distributed intracellular CLU. Figure S3. ER stress-dependent co-localization of CLU with cytoplasmic TDP-43 inclusions in different transfected cell models. Figure S4. Treatment of N2a cells with 10 μM MG132 for 18 h does not cause significant loss of cell viability. Figure S5. (A) CLU reduces the number of TDP-43M337V-GFP inclusions in response to ER stress. Transfected N2a cells were treated as indicated and then lysed and fluorescent protein inclusions were enumerated by flow cytometry. (B) Raw flow cytometry data showing that expression of CLU or mCherry in unstressed (untreated) transfected N2a cells expressing TDP-43M337V-GFP has no significant effect on numbers of inclusions. Figure S6. Western blot data relating to the CLU Drosophila model. Figure S7. Overexpression of GFP does not suppress expression of TDP-43, and additional Drosophila survival data. (DOCX 1057 kb)
Authors’ contributions
JMG, DRW, RAB and SS performed experiments and analyzed the data. KM assisted with human tissue immunohistochemistry procedures; CS assisted with access to human tissues, case selection and image analysis. MRW and LML designed experiments and, together with CMD, oversaw their implementation. TPB supported generation of the transgenic Drosophila and JRK contributed to biochemical analyses. JMG, DRW, JJY, JRK and CMD contributed to authoring of the manuscript; MRW was the co-ordinating author. All authors read and approved the final manuscript.
Ethics approval and consent to participate
All clinical data were collected as part of the Scottish Motor Neurone Disease register (ethics approval from Scotland A Research Ethics Committee 10/MRE00/78 and 15/SS/0216) and all patients consented to the use of their data during life. All post mortem tissue was collected via the Edinburgh Brain Bank, approved by a national ethics committee, in line with the Human Tissue (Scotland) Act. Use of human tissue for post-mortem studies has been reviewed and approved by the Sudden Death Brain Bank ethics committee and the Academic and Clinical Central Office for Research and Development (ACCORD) medical research ethics committee (AMREC). Consent to publish is acquired at consent for post mortem. No identifiable information is provided in this study.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Footnotes
Electronic supplementary material
The online version of this article (10.1186/s40478-017-0481-1) contains supplementary material, which is available to authorized users.
Contributor Information
Christopher M. Dobson, Email: cmd44@cam.ac.uk
Mark R. Wilson, Email: mrw@uow.edu.au
References
- 1.Lindberg I, Shorter J, Wiseman RL, Chiti F, Dickey CA, McLean PJ (2015) Chaperones in neurodegeneration. J Neurosci 35(41):13853–13859 [DOI] [PMC free article] [PubMed]
- 2.Kakkar V, Månsson C, de Mattos EP, Bergink S, van der Zwaag M, van Waarde MAWH, et al. The S/T-rich motif in the DNAJB6 chaperone delays polyglutamine aggregation and the onset of disease in a mouse model. Mol Cell. 2016;62(2):272–283. doi: 10.1016/j.molcel.2016.03.017. [DOI] [PubMed] [Google Scholar]
- 3.Tam S, Geller R, Spiess C, Frydman J. The chaperonin TRiC controls polyglutamine aggregation and toxicity through subunit-specific interactions. Nat Cell Biol. 2006;8(10):1155–1162. doi: 10.1038/ncb1477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Trougakos IP. The molecular chaperone apolipoprotein J/clusterin as a sensor of oxidative stress: implications in therapeutic approaches. Gerontol. 2013;59(6):514–523. doi: 10.1159/000351207. [DOI] [PubMed] [Google Scholar]
- 5.Loison F, Debure L, Nizard P, Le Goff P, Michel D, Le Dréan Y. Up-regulation of the clusterin gene after proteotoxic stress: implication of HSF1-HSF2 heterocomplexes. Biochem J. 2006;395(1):223–231. doi: 10.1042/BJ20051190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Wilson MR, Easterbrook-Smith SB. Clusterin is a secreted mammalian chaperone. Trends Biochem Sci. 2000;25(3):95–98. doi: 10.1016/S0968-0004(99)01534-0. [DOI] [PubMed] [Google Scholar]
- 7.Harold D, Abraham R, Hollingworth P, Sims R, Gerrish A, Hamshere M, et al. Genome-wide association study identifies variants at CLU and PICALM associated with Alzheimer’s disease. Nat Genet. 2009;41(10):1088–1093. doi: 10.1038/ng.440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Greene M, Sam F, Soo Hoo P, Patel R, Seldin D, Connors L. Evidence for a functional role of the molecular chaperone clusterin in amyloidotic cardiomyopathy. Am J Pathol. 2011;178(1):61–68. doi: 10.1016/j.ajpath.2010.11.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Magalhaes J, Saraiva MJ. The heat shock response in FAP: the role of the extracellular chaperone clusterin. Amyloid. 2012;19(SUPPL. 1):3–4. doi: 10.3109/13506129.2012.675370. [DOI] [PubMed] [Google Scholar]
- 10.Narayan P, Orte A, Clarke RW, Bolognesi B, Hook S, Ganzinger KA, et al. The extracellular chaperone clusterin sequesters oligomeric forms of the amyloid-beta1-40 peptide. Nat Struct Mol Biol. 2012;19(1):79–84. doi: 10.1038/nsmb.2191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Humphreys DT, Carver JA, Easterbrook-Smith SB, Wilson MR. Clusterin has chaperone-like activity similar to that of small heat-shock proteins. J Biol Chem. 1999;274(11):6875–6881. doi: 10.1074/jbc.274.11.6875. [DOI] [PubMed] [Google Scholar]
- 12.Yerbury JJ, Poon S, Meehan S, Thompson B, Kumita JR, Dobson CM, et al. The extracellular chaperone clusterin influences amyloid formation and toxicity by interacting with prefibrillar structures. FASEB J. 2007;21(10):2312–2322. doi: 10.1096/fj.06-7986com. [DOI] [PubMed] [Google Scholar]
- 13.Kumita JR, Poon S, Caddy GL, Hagan CL, Dumoulin M, Yerbury JJ, et al. The extracellular chaperone clusterin potently inhibits human lysozyme amyloid formation by interacting with prefibrillar species. J Mol Biol. 2007;369(1):157–167. doi: 10.1016/j.jmb.2007.02.095. [DOI] [PubMed] [Google Scholar]
- 14.Wyatt A, Yerbury J, Ecroyd H, Wilson M. Extracellular chaperones and proteostasis. Annu Rev Biochem. 2013;82:295–322. doi: 10.1146/annurev-biochem-072711-163904. [DOI] [PubMed] [Google Scholar]
- 15.Nizard P, Tetley S, Le Drean Y, Watrin T, Le Goff P, Wilson MR, et al. Stress-induced retrotranslocation of clusterin/ApoJ into the cytosol. Traffic. 2007;8(5):554–565. doi: 10.1111/j.1600-0854.2007.00549.x. [DOI] [PubMed] [Google Scholar]
- 16.Li N, Zoubeidi A, Beraldi E, Gleave ME. GRP78 regulates clusterin stability, retrotranslocation and mitochondrial localization under ER stress in prostate cancer. Oncogene. 2013;32(15):1933–1942. doi: 10.1038/onc.2012.212. [DOI] [PubMed] [Google Scholar]
- 17.Zhang F, Kumano M, Beraldi E, Fazli L, Du C, Moore S, et al. Clusterin facilitates stress-induced lipidation of LC3 and autophagosome biogenesis to enhance cancer cell survival. Nat Commun. 2014;5:6775. doi: 10.1038/ncomms6775. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Cha-Molstad, H, Sung, KS, Hwang, J, Kim, KA, Yu, JE, Yoo, et al. (2015) Amino-terminal arginylation targets endoplasmic reticulum chaperone BiP for autophagy through p62 binding. Nat Cell Biol 17(7):917-929 [DOI] [PMC free article] [PubMed]
- 19.Balantinou E, Trougakos IP, Chondrogianni N, Margaritis LH, Gonos ES. Transcriptional and posttranslational regulation of clusterin by the two main cellular proteolytic pathways. Free Rad Biol Med. 2009;46(9):1267–1274. doi: 10.1016/j.freeradbiomed.2009.01.025. [DOI] [PubMed] [Google Scholar]
- 20.Zoubeidi A, Ettinger S, Beraldi E, Hadaschik B, Zardan A, Klomp LWJ, et al. Clusterin facilitates COMMD1 and I-κB degradation to enhance NF-κB activity in prostate cancer cells. Mol Cancer Res. 2010;8(1):119–130. doi: 10.1158/1541-7786.MCR-09-0277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Materia S, Cater MA, Klomp LWJ, Mercer JFB, La Fontaine S. Clusterin and COMMD1 independently regulate degradation of the mammalian copper ATPases ATP7A and ATP7B. J Biol Chem. 2012;287(4):2485–2499. doi: 10.1074/jbc.M111.302216. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Materia S, Cater MA, Klomp LWJ, Mercer JFB, La Fontaine S. Clusterin (Apolipoprotein J), a molecular chaperone that facilitates degradation of the copper-ATPases ATP7A and ATP7B. J Biol Chem. 2011;286(12):10073–10083. doi: 10.1074/jbc.M110.190546. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Ogen-Shtern N, Ben David T, Lederkremer GZ. Protein aggregation and ER stress. Brain Res. 2016;1648:658–666. doi: 10.1016/j.brainres.2016.03.044. [DOI] [PubMed] [Google Scholar]
- 24.Humphreys D, Hochgrebe TT, Easterbrook-Smith SB, Tenniswood MP, Wilson MR. Effects of clusterin overexpression on TNFalpha- and TGFbeta-mediated death of L929 cells. Biochemistry. 1997;36(49):15233–15243. doi: 10.1021/bi9703507. [DOI] [PubMed] [Google Scholar]
- 25.Whiten D, San Gil R, McAlary L, Yerbury J, Ecroyd H, Wilson MR. Rapid flow cytometric measurement of protein inclusions and nuclear trafficking. Sci Rep. 2016;6:31138. doi: 10.1038/srep31138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Gregory JM, Barros TP, Meehan S, Dobson CM, Luheshi LM. The aggregation and neurotoxicity of TDP-43 and its ALS-associated 25 kDa fragment are differentially affected by molecular chaperones in Drosophila. PLoS One. 2012;7(2):e31899. doi: 10.1371/journal.pone.0031899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Bischof J, Maeda RK, Hediger M, Karch F, Basler K. An optimized transgenesis system for drosophila using germ-line-specific φC31 integrases. Proc Nat Acad Sci USA. 2007;104(9):3312–3317. doi: 10.1073/pnas.0611511104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Brand AH, Perrimon N. Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development. 1993;118(2):401–415. doi: 10.1242/dev.118.2.401. [DOI] [PubMed] [Google Scholar]
- 29.Stewart EM, Aquilina JA, Easterbrook-Smith SB, Murphy-Durland D, Jacobsen C, Moestrup S, et al. Effects of glycosylation on the structure and function of the extracellular chaperone clusterin. Biochemistry. 2007;46(5):1412–1422. doi: 10.1021/bi062082v. [DOI] [PubMed] [Google Scholar]
- 30.Ryoo H, Domingos P, Kang M-J, Steller H. Unfolded protein response in a drosophila model for retinal degeneration. EMBO J. 2007;26(1):242–252. doi: 10.1038/sj.emboj.7601477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Hetz C. The unfolded protein response: controlling cell fate decisions under ER stress and beyond. Nat Rev Mol Cell Biol. 2012;13(2):89–102. doi: 10.1038/nrm3270. [DOI] [PubMed] [Google Scholar]
- 32.Luheshi LM, Tartaglia GG, Brorsson AC, Pawar AP, Watson IE, Chiti F, et al. Systematic in vivo analysis of the intrinsic determinants of amyloid β pathogenicity. PLoS Biol. 2007;5(11):2493–2500. doi: 10.1371/journal.pbio.0050290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Zhang YJ, Xu YF, Cook C, Gendron TF, Roettges P, Link CD, et al. Aberrant cleavage of TDP-43 enhances aggregation and cellular toxicity. Proc Nat Acad Sci USA. 2009;106(18):7607–7612. doi: 10.1073/pnas.0900688106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Huang YC, Lin KF, He RY, Tu PH, Koubek J, Hsu YC, et al. Inhibition of TDP-43 aggregation by nucleic acid binding. PLoS One. 2013;8(5):e64002. doi: 10.1371/journal.pone.0064002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Guo W, Chen Y, Zhou X, Kar A, Ray P, Chen X, et al. An ALS-associated mutation affecting TDP-43 enhances protein aggregation, fibril formation and neurotoxicity. Nat Struct Mol Biol. 2011;18(7):822–831. doi: 10.1038/nsmb.2053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Zeineddine R, Whiten D, Farrawell N, McAlary L, Hanspal M, Wilson MR, et al. Flow cytometric measurement of the cellular propagation of TDP-43 aggregation. Prion. 2017;11(3):195–204. doi: 10.1080/19336896.2017.1314426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.McAlary, L, Aquilina, JA, Yerbury, JJ (2016) Susceptibility of mutant SOD1 to form a destabilized monomer predicts cellular aggregation and toxicity but not in vitro aggregation propensity. Front Neurosci 10(NOV):499 [DOI] [PMC free article] [PubMed]
- 38.Brooks SA. Protein glycosylation in diverse cell systems: implications for modification and analysis of recombinant proteins. Exp Rev Proteom. 2006;3(3):345–359. doi: 10.1586/14789450.3.3.345. [DOI] [PubMed] [Google Scholar]
- 39.Kapron JT, Hilliard GM, Lakins JN, Tenniswood MPR, West KA, Carr SA, et al. Identification and characterization of glycosylation sites in human serum clusterin. Prot Sci. 1997;6(10):2120–2133. doi: 10.1002/pro.5560061007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Lee W-C, Yoshihara M, Littleton J. Cytoplasmic aggregates trap polyglutamine-containing proteins and block axonal transport in a Drosophila model of Huntington’s disease. Proc Natl Acad Sci U S A. 2004;101(9):3224–3229. doi: 10.1073/pnas.0400243101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Wittmann C, Wszolek M, Shulman J, Salvaterra P, Lewis J, Hutton M, et al. Tauopathy in Drosophila: neurodegeneration without neurofibrillary tangles. Science. 2001;293(5530):711–714. doi: 10.1126/science.1062382. [DOI] [PubMed] [Google Scholar]
- 42.Johnson B, McCaffery M, Lindquist S, Gitler A. A yeast TDP-43 proteinopathy model: exploring the molecular determinants of TDP-43 aggregation and cellular toxicity. Proc Natl Acad Sci U S A. 2008;105(17):6439–6444. doi: 10.1073/pnas.0802082105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Zhang S, Binari R, Zhou R, Perrimon N. A genomewide RNA interference screen for modifiers of aggregates formation by mutant Huntingtin in Drosophila. Genetics. 2010;184(4):1165–1179. doi: 10.1534/genetics.109.112516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Giannakopoulos P, Kövari E, French LE, Viard I, Hof PR, Bouras C. Possible neuroprotective role of clusterin in Alzheimer's disease: a quantitative immunocytochemical study. Acta Neuropathol. 1998;95(4):387–394. doi: 10.1007/s004010050815. [DOI] [PubMed] [Google Scholar]
- 45.Sasaki K, Doh-ura K, Wakisaka Y, Iwaki T. Clusterin/apolipoprotein J is associated with cortical Lewy bodies: immunohistochemical study in cases with alpha-synucleinopathies. Acta Neuropathol. 2002;104(3):225–230. doi: 10.1007/s00401-002-0546-4. [DOI] [PubMed] [Google Scholar]
- 46.Ferrer I, Carmona M, Blanco R, Moreno D, Torrejon-Escribano B, Olive M. Involvement of clusterin and the aggresome in abnormal protein deposits in myofibrillar myopathies and inclusion body myositis. Brain Pathol. 2005;15(2):101–108. doi: 10.1111/j.1750-3639.2005.tb00504.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Zhang H, Kim JK, Edwards CA, Xu Z, Taichman R, Wang CY. Clusterin inhibits apoptosis by interacting with activated Bax. Nat Cell Biol. 2005;7(9):909–915. doi: 10.1038/ncb1291. [DOI] [PubMed] [Google Scholar]
- 48.Wang L, Popko B, Roos RP. The unfolded protein response in familial amyotrophic lateral sclerosis. Hum Mol Genet. 2011;20(5):1008–1015. doi: 10.1093/hmg/ddq546. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Kanekura K, Suzuki H, Aiso S, Matsuoka M. ER stress and unfolded protein response in amyotrophic lateral sclerosis. Mol Neurobiol. 2009;39(2):81–89. doi: 10.1007/s12035-009-8054-3. [DOI] [PubMed] [Google Scholar]
- 50.Mackenzie IRA, Bigio EH, Ince PG, Geser F, Neumann M, Cairns NJ, et al. Pathological TDP-43 distinguishes sporadic amyotrophic lateral sclerosis from amyotrophic lateral sclerosis with SOD1 mutations. Ann Neurol. 2007;61(5):427–434. doi: 10.1002/ana.21147. [DOI] [PubMed] [Google Scholar]
- 51.Saberi S, Stauffer JE, Schulte DJ, Ravits J. Neuropathology of amyotrophic lateral sclerosis and its variants. Neurol Clin. 2015;33(4):855–876. doi: 10.1016/j.ncl.2015.07.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Prudlo J, König J, Schuster C, Kasper E, Büttner A, Teipel S, et al. TDP-43 pathology and cognition in ALS. Neurology. 2016;87(10):1019–1023. doi: 10.1212/WNL.0000000000003062. [DOI] [PubMed] [Google Scholar]
- 53.Poon S, Rybchyn MS, Easterbrook-Smith SB, Carver JA, Pankhurst GJ, Wilson MR. Mildly acidic pH activates the extracellular molecular chaperone clusterin. J Biol Chem. 2002;277(42):39532–39540. doi: 10.1074/jbc.M204855200. [DOI] [PubMed] [Google Scholar]
- 54.Narayan, P, Holmström, KM, Kim, DH, Whitcomb, DJ, Wilson, MR, St. George-Hyslop, P, et al. (2014) Rare individual amyloid-β oligomers act on astrocytes to initiate neuronal damage. Biochemistry 53(15):2442-2453 [DOI] [PMC free article] [PubMed]
- 55.Cascella R, Conti S, Tatini F, Evangelisti E, Scartabelli T, Casamenti F, et al. Extracellular chaperones prevent Aβ-induced toxicity in rat brains. Biochim Biophys Acta - Mol Basis Dis. 2013;1832(8):1217–1226. doi: 10.1016/j.bbadis.2013.04.012. [DOI] [PubMed] [Google Scholar]
- 56.Scotter EL, Vance C, Nishimura AL, Lee YB, Chen HJ, Urwin H, et al. Differential roles of the ubiquitin proteasome system and autophagy in the clearance of soluble and aggregated TDP-43 species. J Cell Sci. 2014;127(6):1263–1278. doi: 10.1242/jcs.140087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Batulan Z, Shinder GA, Minotti S, He BP, Doroudchi MM, Nalbantoglu J, et al. High threshold for induction of the stress response in motor neurons is associated with failure to activate HSF1. J Neurosci. 2003;23(13):5789–5798. doi: 10.1523/JNEUROSCI.23-13-05789.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Spires-Jones T, Attems J, Thai D (2017) Interactions of pathological proteins in neurodegenerative diseases. Acta Neuropathol. doi:10.1007/s00401-017-1709-7 [DOI] [PMC free article] [PubMed]
- 59.Sinadinos C, Quraishe S, Sealey M, Samson PB, Mudher A, Wyttenbach A. Low endogenous and chemical induced heat shock protein induction in a 0N3Rtau-expressing Drosophila larval model of Alzheimer's disease. J Alzheimers Dis. 2013;33(4):1117–1133. doi: 10.3233/JAD-2012-121534. [DOI] [PubMed] [Google Scholar]
- 60.Ho SW, Tsui YTC, Wong TT, Cheung SKK, Goggins WB, Yi LM, et al. Effects of 17-allylamino-17-demethoxygeldanamycin (17-AAG) in transgenic mouse models of frontotemporal lobar degeneration and Alzheimer's disease. Transl Neurodegen. 2013;2(1):24. doi: 10.1186/2047-9158-2-24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Rusmini P, Simonini F, Crippa V, Bolzoni E, Onesto E, Cagnin M, et al. 17-AAG increases autophagic removal of mutant androgen receptor in spinal and bulbar muscular atrophy. Neurobiol Dis. 2011;41(1):83–95. doi: 10.1016/j.nbd.2010.08.023. [DOI] [PubMed] [Google Scholar]
- 62.Riedel, M, Goldbaum, O, Schwarz, L, Schmitt, S, Richter-Landsberg, C (2010) 17-AAG induces cytoplasmic α-synuclein aggregate clearance by induction of autophagy. PLoS One 5(1):e8753 [DOI] [PMC free article] [PubMed]
- 63.Waza M, Adachi H, Katsuno M, Minamiyama M, Sang C, Tanaka F, et al. 17-AAG, an Hsp90 inhibitor, ameliorates polyglutamine-mediated motor neuron degeneration. Nat Med. 2005;11(10):1088–1095. doi: 10.1038/nm1298. [DOI] [PubMed] [Google Scholar]
- 64.Zinkie S, Gentil BJ, Minotti S, Durham HD. Expression of the protein chaperone, clusterin, in spinal cord cells constitutively and following cellular stress, and upregulation by treatment with Hsp90 inhibitor. Cell Stress Chaperones. 2013;18(6):745–758. doi: 10.1007/s12192-013-0427-x. [DOI] [PMC free article] [PubMed] [Google Scholar]


