Abstract
Aptamers are short non-coding, single-stranded oligonucleotides (RNA or DNA) developed through Systematic Evolution of Ligands by Exponential enrichment (SELEX) in vitro. Similar to antibodies, aptamers can bind to specific targets with high affinity, and are considered promising therapeutic agents as they have several advantages over antibodies, including high specificity, stability, and non-immunogenicity. Furthermore, aptamers can be produced at a low cost and easily modified, and are, therefore, called “chemical antibodies”. In the past years, a variety of aptamers specifically bound to both breast cancer biomarkers and cells had been selected. Besides, taking advantage of nanomaterials, there were a number of aptamer-nanomaterial conjugates been developed and widely investigated for diagnostics and targeted therapy of breast cancer. In this short review, we first present a systematical review of various aptamer selection methods. Then, various aptamer-based diagnostic and therapeutic strategies of breast cancer were provided. Finally, the current problems, challenges, and future perspectives in the field were thoroughly discussed.
Keywords: Aptamer, SELEX, Breast cancer, Diagnosis, Targeted therapy
Background
Breast cancer is currently the second leading cause of death among women in the world [1]. Although the mortality of breast cancer has decreased owing to the development of modern therapeutic approaches, the morbidity is still very high [2]. About 1,700,000 new cases of breast cancer and 521,900 deaths every year have been reported worldwide [3]. In China, especially in rural areas where people have lower income and less access to health care facilities, the morbidity of breast cancer has reached 24.20 per 100,000 [4]. In the United States, 231,840 women were diagnosed with and 40,290 died from breast cancer in 2015 [5]. As a highly heterogeneous malignancy, breast cancer can be classified into four subtypes based on the expression of specific biomarkers estrogen receptor (ER), progesterone receptor (PR), human epidermal growth factor receptor-2 (HER2), and proliferation marker Ki-67. That are luminal A (ER+ or PR+, HER2−), luminal B (ER+ or PR+, HER2+), HER2-overexpressing, and triple-negative (ER−, PR−, HER2−), whereas Ki-67 expression marks early-relapsing tumors [6–9]. In clinics, breast cancer subtypes usually show different response to therapy; therefore, treatment strategies are always chosen based on the molecular classification of tumors [10]. Early diagnosis and timely therapy are of a paramount importance in breast cancer prognosis, as the chances of survival significantly increase. However, current diagnostic technologies, including imaging, molecular detection, and immunohistochemistry (IHC) have inherent limitations and may provide uncertain results [11, 12]. The most common treatment strategies today are endocrine therapy, chemotherapy, and targeted therapy using antibodies recognizing cancer biomarkers; however, they are not always successful and may cause adverse effects and drug tolerance [13]. Therefore, there is a constant need to develop novel effective methods for breast cancer diagnosis and therapy.
Aptamers provide a potential alternative source of molecules for targeted therapy. Aptamers represent functional short single-stranded non-coding RNA or DNA that can specifically bind to targets with high affinity [14–20]; they are developed by the SELEX (Systematic Evolution of Ligands by EXponential enrichment) method in vitro and can recognize various targeting molecules by forming several secondary structures such as stem, loop, hairpin, bulge, pseudoknot, and G-quadruplex [21]. Compared with antibodies, aptamers have such advantages as high specificity and affinity, stability, and non-immunogenicity. In addition, they can be easily modified at a low cost to target a variety of molecules, and are, therefore, called “chemical antibodies” [22–25]. Because of these characteristics, in the past decade coupling with various nanomaterials, aptamers have been extensively studied in the fields of biological detection [26–30], molecular diagnosis [31, 32], bioimaging [33–35], tumor molecular classification [36], and drug delivery [37, 38].
Recently, applications of aptamers and aptamer-nanomaterial conjugates for cancer detection and therapy have attracted increasing attention. Aptamers with affinity to tumor molecular biomarkers are widely used for the detection of cancer markers and circulating tumor cells (CTCs), providing timely diagnosis, prognosis monitoring, and targeted therapy. Thus, a series of aptamers optimized for breast cancer have been selected and applied in diagnostics and treatment (Table 1), opening a new avenue for personalized medicine of breast cancer. A previously published review about aptamer-based diagnostics and targeted therapy in breast cancer focused on aptamer applications [39]; however, systematic analysis of aptamer selection methods has not yet been performed. Here, we evaluated various SELEX methods, discuss the applications of aptamers in diagnostics and treatment of breast cancer, and provide an outlook of current problems and future perspectives in this field.
Table 1.
Aptamers specific for breast cancer
| Aptamer | Sequence (5′–3′) | Target | References |
|---|---|---|---|
| H2 | GGGCCGTCGAACACGAGCATGGTGCGTGGACCTAGGATGACCTGAGTACTGTCC | HER2 | [40] |
| AS1411 | GGTGGTGGTGGTTGTGGTGGTGGTGG | Nucleolin | [41, 42] |
| ERaptD4 | ATACCAGCTTATTCAATTCGTTGCATTTAGGTG CATTACGGGGGTTATCCGCTCTCTCAGATAGTA TGTGCAATCA |
ERα | [43] |
| S6 | TGGATGGGGAGATCCGTTGAGTAAGCGGGCGTGTCTCTCTGCCGCCTTGCTATGGGG | SK-BR-3 cells | [44] |
| XLX-1-A | GAATTCAGTCGGACAGCGAAGTAGTTTTCCTT CTAACCTAAGAACCCGCGGCAGTTTAATGTAG ATGGACGAA |
MDA-MB-231 cells | [45] |
| MUC1 | GCAGTTGATCCTTTGGATACCCTGG | Mucin-1 | [46] |
| HB5 | AACCGCCCAAATCCCTAAGAGTCTGCACTTGT CATTTTGTATATGTATTTGGTTTTTGGCTCTCAC AGACACACTACACACGCACA |
HER2 | [47] |
| KMF2-1a | AGGCGGCAGTGTCAGAGTGAATAGGGGATGTA CAGGTCTGCACCCACTCGAGGAGTGACTGAGCGACGAAGACCCC |
MCF-10AT1 cells | [48] |
| SYL3C | CACTACAGAGGTTGCGTCTGTCCCACGTTGTCATGGGGGGTTGGCCTG | EpCAM | [49] |
Selection of specific aptamers using SELEX
In 1990, Tuerk and Gold [14] were the first to select two RNA sequences with the binding affinity to T4 DNA polymerase from a random RNA library in vitro, and designated the method Systematic Evolution of Ligands by EXponential enrichment (SELEX). At the same time, Ellington and Szostak [50] isolated RNA subpopulations from a large combinatorial library of 1010 random-sequence RNA molecules, which can specifically bind various organic dyes, and coined the term “aptamer”. Since then, the SELEX method has been widely applied for aptamer selection in vitro. As shown in Fig. 1, conventional SELEX procedures usually include: (1) random single-stranded (ss)DNA or RNA library design and synthesis (2) target binding, (3) elution, (4) PCR amplification, (5) preparation of a single-stranded oligonucleotide library, (6) multiple rounds of selection, (7) cloning and sequencing, and (8) aptamer identification [51]. As a rule, negative selection is often needed before incubation with the targets to remove non-specific sequences. However, there are differences between selection of DNA and RNA aptamers. Thus, to obtain DNA aptamers, the amplification of bound sequences is performed by a regular PCR, followed by ssDNA preparation to generate the secondary library, whereas in case of RNA aptamers, reverse transcription PCR should be used prior to the preparation of the secondary random RNA library.
Fig. 1.
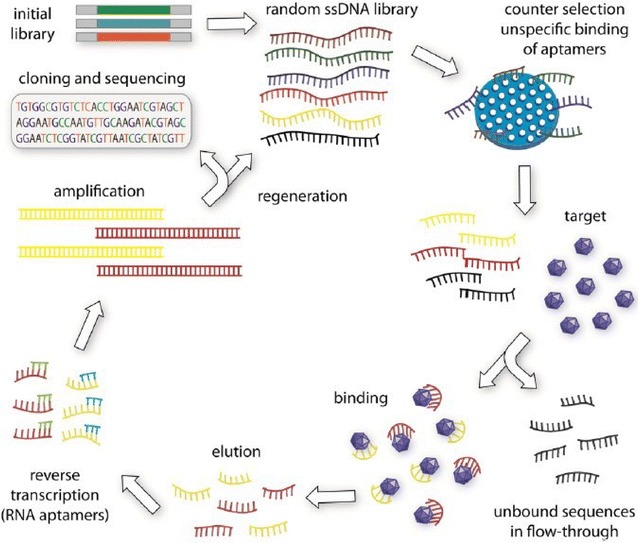
Schematic illustration of conventional SELEX. Usually, an initial random library should be synthesized for selection. Then, counter selection is always needed before incubate the library with the targets to remove non-specific binding sequences. After incubation the target-bound sequeneces are collected and amplificated by PCR method. For selecting DNA aptamers, general PCR is enough for the amplification of the secondary library. While, as for selecting RNA aptamers, reverse transcription PCR is introduced before amplification and preparation of secondary library. After several successive rounds of selection, the library is cloned and sequenced followed by evaluation and identification of the enriched aptamers
(Reprinted with permission from Ref. [51]. Copyright © 2015, Elsevier)
The past decades have witnessed fast development of SELEX methods and applications of aptamers in various areas. Although the conventional SELEX can successfully screen aptamers in vitro, the procedures are tedious, laborious, and time-consuming. Therefore, a number of modified SELEX methods of high efficiency have been developed to select aptamers against various targets, including proteins, peptides, small chemical compounds, metal ions, bacteria, cultured cells, and even heterogeneous in vivo targets [52–60]. According to the target type, SELEX methods can be divided into purified target-based SELEX, Cell-SELEX, and in vivo SELEX [61, 62].
Purified target-based SELEX
Purified targets of aptamers include metal ions, small molecules, proteins, and many others [62–64]. In its general flow, SELEX of purified target-specific aptamers follow the original procedure (Fig. 2) [65], where the key step is separation of the target-bound oligonucleotides. Therefore, various solid-phase carriers are used in purified target-based SELEX, such as magnetic nanoparticles (MNPs), agarose beads, nitrocellulose membrane, and adsorptive microplates. Block et al. [66] first selected thrombin-specific ssDNA aptamers by immobilizing thrombin on a concanavalin A-agarose column. In our laboratory, Xi et al. [65] selected three DNA aptamers specifically bound to hepatitis B virus (HBV) surface antigen (HBsAg) using carboxylated MNPs for HBsAg immobilization. As a result, they generated a chemiluminescence aptasensor based on aptamer H01, which could detect HBsAg in spiked samples or clinical serum specimens with a detection limit of 0.1 ng/mL HBsAg in serum, which was lower compared to the 0.5 ng/mL limit of a clinically used ELISA. In a similar study, Huang et al. [67] obtained a DNA aptamer against HBV e antigen (HBeAg) by MNP-based SELEX and developed an aptamer-based fluorescence biosensor for highly efficient quantitative detection of up to 609 ng/mL HBeAg in serum, which could be completed within 2 min.
Fig. 2.
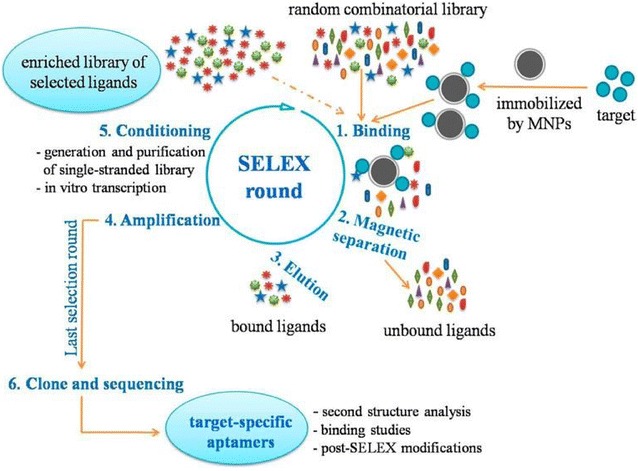
Schematic illustration of purified-target based SELEX procedures using magnetic particles as solid phase carrier for immobilization of targets
(Reprinted with permission from Ref. [65]. Copyright © 2015, American Chemical Society)
Cell-SELEX
Although the achievements in aptamer selection against purified targets, especially proteins, have been significant, there are some limitations regarding the application of such aptamers, mainly due to a synthetic nature of the target proteins which differ from their native counterparts in conformation and higher order structure, and bioactivity. Consequently, the aptamers selected against artificial proteins may have much lower sensitivity in recognizing native targets. Therefore, whole living cells have been considered and successfully used for aptamer generation by the method known as Cell-SELEX [68]. The production of aptamers in Cell-SELEX generally includes both positive and negative selection. As shown in Fig. 3, 2–4 cycles of positive selection are first conducted by incubating a random-sequence oligonucleotide library with target cells to remove unbound sequences. Then, negative selection is performed by incubating the secondary library with non-target cells, after which unbound sequences are collected, applied to the target cells, eluted, amplified by PCR, and used to prepare an ssDNA secondary library for the next round of selection. Finally, when the selection process is completed, target cell-specific aptamers are PCR-amplified, cloned, and sequenced. Compared with purified target-based SELEX, Cell-SELEX can generate novel highly specific aptamers against native complex functional molecules expressed on the cell surface, without previous identification of the target. Therefore, since targets are unknown, the selected aptamers can be used to discover new molecular biomarkers [69–71].
Fig. 3.
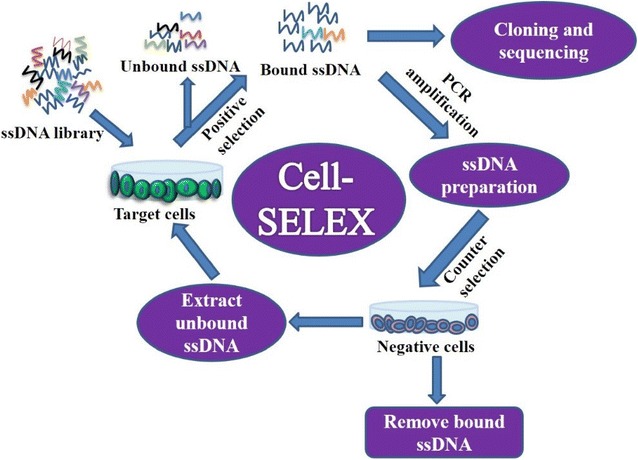
Schematic illustration of Cell-SELEX procedures
Over the past years, hundreds of cell-specific aptamers have been obtained by Cell-SELEX. Thus, Bayrac et al. [72] successfully used a Cell-SELEX-based procedure without negative selection to generate glioblastoma multiforme-specific DNA aptamers, which showed high specificity and affinity to glioblastoma cell lines with a K d in the nanomolar range. Li et al. [73] developed a panel of DNA aptamers against colon cancer SW620 cells after 14 rounds of selection using Cell-SELEX. The finally selected aptamer XL-33 showed high binding affinity (K d = 0.7 nM), and its truncated counterpart XL-33-1 containing only 45 nt also displayed excellent selectivity to the target cells. Furthermore, tissue imaging indicated that XL-33-1 could distinguish between metastatic tumor tissues or lymph nodes and benign tissues with a detection rate of 81.7%. In our laboratory, we have recently successfully applied Cell-SELEX to develop a series of DNA aptamers specifically binding to cultured human hepatocellular carcinoma HepG2 cells [74]. The selected aptamers showed high binding affinity to HepG2 cells with K d values ranging from 46.3 to 199.4 nM and could distinguish HepG2 cells from normal human liver cells.
In vivo SELEX
Currently, most aptamers are selected in in vitro conditions, which provide a simple and controllable binding environment. However, considering that the ultimate goal is the application of aptamers in vivo, i.e., in a very complex physiological environment, in vitro-selected aptamers may not have sufficient stability and half-life to exert the desirable effects [75]. Therefore, generation of aptamers with physiological stability is a task of a paramount importance. In vivo SELEX is a new aptamer selection approach based on using animal models to obtain tissue- and organ-specific aptamers (Fig. 4) [61]. The detailed protocol used in in vivo SELEX is as follows: intravenous injection of a random oligonucleotide library, harvesting the tissue or organ of interest, extraction and amplification of the bound molecules, and preparation of a secondary random library for the next selection cycle. Mi et al. [76] tested a nuclease-resistant RNA aptamer against hepatic colon cancer metastases in tumor-bearing mice using in vivo SELEX, and identified the target molecule as p68, an RNA helicase upregulated in colorectal cancer. Wang et al. [77] applied in vivo SELEX to select RNA molecules specific for human non-small cell lung cancer using cultured NCI-H460 cancer cells and tumor-bearing xenograft mice, and obtained an aptamer with high specificity and affinity to both cancer cell line and mouse tumor tissues.
Fig. 4.
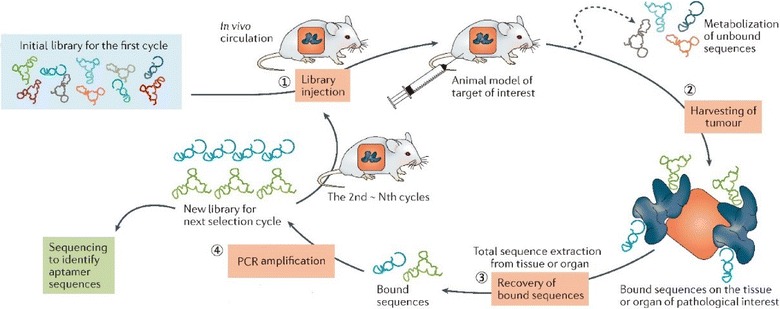
Schematic illustration of in vivo SELEX procedures
(Reprinted with permission from Ref. [61]. Copyright © 2017, Nature Publishing Group)
Highly efficient SELEX
In order to improve SELEX efficiency, various approaches have been recently developed, including capillary electrophoresis SELEX (CE-SELEX) [78], microfluidic SELEX [79], high-throughput sequencing-assisted SELEX (HT-SELEX) [80, 81], monoclonal surface display SELEX (MSD-SELEX) [82], and automated SELEX [83]. Zhu et al. [82] designed a novel MSD-SELEX method for rapid and efficient selection and identification of aptamers (Fig. 5). They combined an initial library with primer-modified beads to produce a library of monoclonal DNA-displaying beads via highly parallel single-molecule emulsion PCR, which they incubated with the target. This new aptamer selection approach was afterwards successfully applied to identify high-affinity aptamers against various targets. Compared to conventional SELEX methods, the newly developed MSD-SELEX approach is simple, rapid, efficient, and cost-effective. Dong et al. [84] screened an alpha-fetoprotein-bound ssDNA aptamer using CE-SELEX technology with only four selection cycles. The aptamer could not only distinguish HepG2 cells from A549 cells by immunofluorescence imaging but also efficiently inhibited the migration and invasion of hepatocellular carcinoma cells in vivo. Moreover, Lin et al. [85] developed a microfluidic SELEX chip based on magnetic beads to select hemoglobin (Hb)- and HbA1c-specific ssDNA aptamers (Fig. 6). They coated magnetic beads with HbA1c and Hb, performed several rounds of selection and enrichment with an ssDNA library, and selected specific oligonucleotides, which were sequenced and identified. Compared with conventional SELEX methods, the developed microfluidic SELEX system dramatically decreased the incubation and partitioning time, thus simplifying the entire SELEX process. In addition, various newly developed separation and amplification technologies, including flow cytometry [86, 87], biacore surface plasmon resonance [88], atomic force microscopy [89–91], and digital PCR [92] have been integrated into SELEX to obtain aptamers with high affinity and specificity to targets.
Fig. 5.
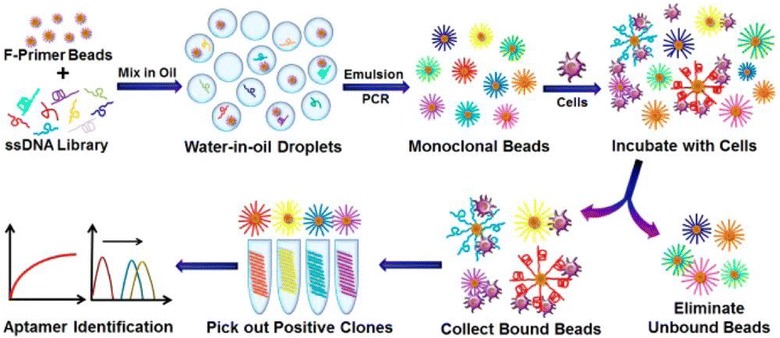
Schematic illustration of monoclonal surface display SELEX (MSD-SELEX) procedures
(Reprinted with permission from Ref. [82]. Copyright © 2014, American Chemical Society)
Fig. 6.
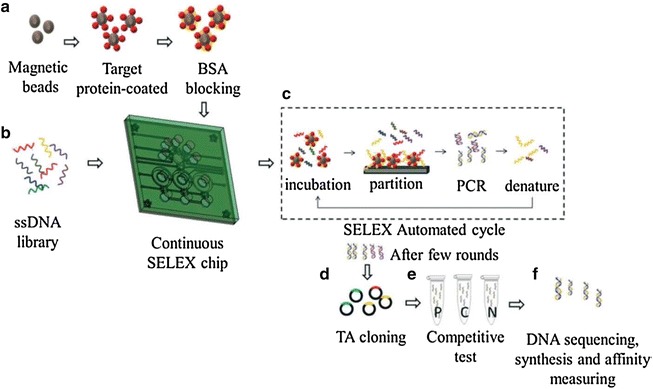
Schematic illustration of microfluidic SELEX procedures
(Reprinted with permission from Ref. [85]. Copyright © 2014, Royal Society of Chemistry)
Applications of aptamers for breast cancer diagnostics
Detection of breast cancer biomarkers using aptamers
Biomarker detection plays a very important role in early diagnosis, monitoring of curative effects, and prognosis in breast cancer. Among the identified breast cancer-specific biomarkers, HER2 is one of the most important and commonly used not only for molecular classification but also for the targeted therapy of breast cancer in clinics. Niazi et al. [40] used Cell-SELEX to select an anti-HER2 ssDNA aptamer H2 which showed high binding affinity to HER2 with a K d of 270 nM. As a next step, they developed a fluorescence-based magnetic microbead-carbon nanotube-H2 hybrid system for HER2 detection in vitro and demonstrated its potential for identification of the native HER2 protein in solution and biological samples. Qureshi et al. [93] have developed a label-free capacitive aptasensor by functionalizing anti-HER2 ssDNA aptamers onto interdigitated microelectrodes of the capacitor for HER2 detection in serum (Fig. 7). The aptasensor was very sensitive and could be applied to detect HER2 in human serum within a dynamic range of 0.2–2 ng/mL, demonstrating versatility and potential as a tool for early diagnosis of breast cancer.
Fig. 7.
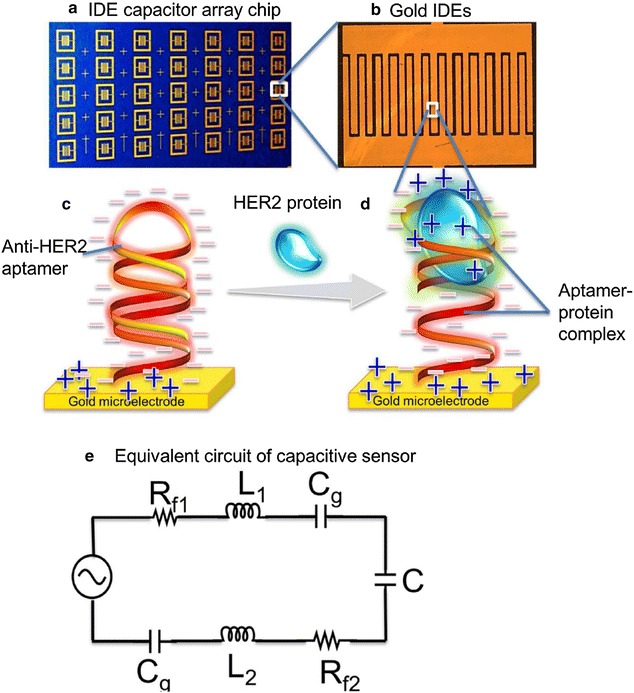
Schematic illustration of label-free capacitive aptasensor for HER2 protein detection in serum. a capacitor array chip, b capacitor, c gold IDE functionalized with anti-HER2 aptamer whose phosphodiester backbone contributes abundant negatively charged species on capacitors, d formation of aptamer-HER2 complex that induce charge distribution and thus influence specific changes in capacitance that can be measured and e equivalent circuit of the interdigitated capacitor sensor
(Reprinted with permission from Ref. [93]. Copyright © 2015, Elsevier)
Besides HER2, other breast cancer biomarkers were used to select aptamers for breast cancer diagnostics [94–98]. Thus, Ahirwar et al. [43] used the HT-SELEX approach to identify an ERα-specific DNA aptamer, which could be internalized by ERα-positive breast cancer cells and localize specifically in the nucleus. The aptamer was applied to reveal ERα expression in breast cancer, and the results were consistent with IHC detection of ERα in breast cancer tissues strongly and moderately positive, or negative for ERα. Li et al. [99] designed a microcantilever biosensor for label-free detection of nucleolin, a protein regulating the stability of Bcl-2 mRNA in cancer cells. In their system, nucleolin-specific aptamer AS1411 was functionalized on sensor cantilevers in the microcantilever array, while reference cantilevers were modified with 6-mercapto-1-hexanol to eliminate environmental disturbances. In this aptasensor, aptamer AS1411 interaction with nucleolin in the sample would induce surface stress change, resulting in differential deflection between the sensor and reference cantilevers. The system demonstrated high sensitivity with a detection limit of about 1.0 nM.
Detection of breast cancer cells using aptamers
Identification of breast cancer cells, especially CTCs in patient serum is important for early cancer diagnosis, monitoring of treatment effects, and prognosis. Aptamers evolved from both purified target-based SELEX and Cell-SELEX provide a powerful tool for detecting breast cancer cells [100–104]. Cai et al. [105] have developed a signal-on fluorescence aptasensor using a mucin 1 (MUC1)-specific aptamer and ssDNA-sensitized luminescent terbium (TbIII) for label-free, facile, and sensitive detection of MCF-7 breast cancer cells. The principle of the method is that in the presence of MCF-7 cells the aptamer would bind to MUC1 on the cell membrane. After centrifugation, aptamer-cell conjugates would be separated from supernatant. The signal probe is then added, and in the absence of aptamer-cell conjugates, the emission of Tb3+ would be enhanced, whereas in their presence, the signal probe would hybridize with the aptamer and lose the ability to induce Tb3+ emission. The proposed aptasensor exhibited excellent sensitivity towards MCF-7 cells with a detection limit as low as 70 cells/mL. Furthermore, Jo et al. [106] synthesized dual aptamer-functionalized dye-doped silica nanoparticles based on MUC1 and HER2 aptamers for highly sensitive detection of breast cancer cells (Fig. 8). In their method, the aptamers modified on magnetic beads would capture the target cells, which would be precipitated by magnetic separation, and the dual aptamer-functionalized dye-doped silica nanoparticles are added as signal generators. Then, magnetic separation and fluorescence excitation are conducted and fluorescence signal would be detected if target cells were captured; otherwise, no fluorescence would be observed. Compared with single aptamer-based detection, this newly developed technique provides very sensitive diagnostics of breast cancer cells with a detection limit of 10 cells/mL, which is dramatically lower compared to other methods. We have recently synthesized aptamer-functionalized silver nanoclusters (AgNCs) based on a MUC1 aptamer for imaging of MCF-7 breast cancer cells [107], which could efficiently differentiate MCF-7 cells from more aggressive MDA-MB-231 breast cancer cells and A549 human lung cancer cells.
Fig. 8.
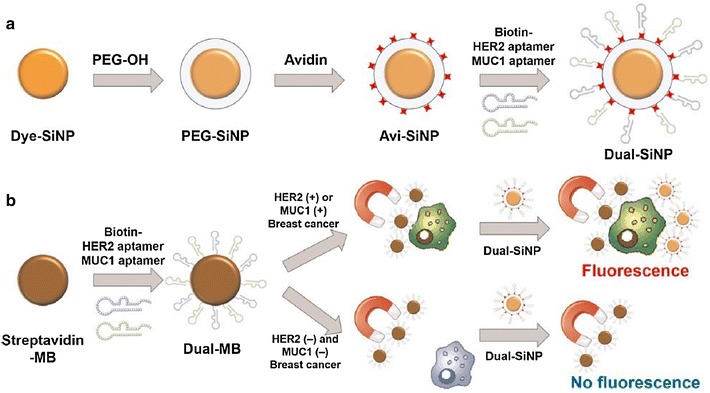
a Modification processes of the silica nanoparticles. Dye-SiNPs were synthesized via the reverse microemulsion technique, followed by being coated with PEG and avidin immobilization. Then, dual aptamer-functionalized dye-doped silica nanoparticles (Dual-SiNPs) were synthesized by modifying MUC1 aptamer and HER2 aptamer onto Dye-SiNPs through biotin-avidin interactions. b Schematic illustration of Dual-SiNPs for highly sensitive detection of MUC1(+) and HER2(+) breast cancer cells. MUC1(+) or HER2(+) breast cancer cells were selectively enriched and separated by dual aptamer-modified magnetic beads followed by being detected with Dual-SiNPs
(Reprinted with permission from Ref. [106]. Copyright © 2015, Elsevier)
Detection of breast cancer tissues using aptamers
Currently, IHC is still the gold standard in breast cancer detection used to confirm definite diagnosis. However, antibodies used in IHC are expensive, diffuse slowly, and always need signal amplification. In this respect, aptamers have attracted increasing attention as the means to overcome these limitations [108–111]. Thus, Gupta et al. [112] have developed a fluorescent HER2-targeting aptamer for breast cancer tissue imaging, which opened a door to aptamer application in histochemistry. Li et al. [45] used Cell-SELEX to identify a DNA aptamer LXL-1-A specifically binding to MDA-MB-231 breast cancer cells, which showed high affinity and excellent selectivity towards MDA-MB-231 cells and was utilized for breast cancer histochemistry. The results were very promising, as aptamer LXL-1-A was highly specific to breast cancer tissues, demonstrating a 76% detection rate for tumor metastasis in regional lymph nodes.
Applications of aptamers in breast cancer therapy
Aptamers can be used not only as diagnostic tools but also as therapeutic agents, because they can, after specific binding to target proteins, either directly inhibit/activate protein function or modulate protein–protein interactions [61, 75]. Furthermore, aptamers can also be used as recognition ligands or carriers for targeted drug delivery [113, 114]. To date, although there have been a couple of aptamers clinically tested for disease treatments; only one of them has already been approved by the FDA for the treatment of all types of neovascular age-related macular degeneration [115]. The limited in vivo therapeutic applications of aptamers mainly result from the limitations of nuclease degradation, metabolism and toxicity of aptamers in the physiological environment [75, 116]. Thus, in order to decrease the degradation and improve efficiency of aptamer-based therapeutics in vivo, the original selected aptamers always need chemical modification, remoulding or conjugation with other biomaterials. In this part, we focused on the therapeutic applications of aptamers for targeted therapy of breast cancer.
Aptamers as therapeutics
Aptamers can bind to and regulate the function of the target protein or gene expression, and may signal cells to start apoptosis or enhance their sensitivity to chemical drugs [117–121]. Bala et al. [122] selected two glutathione-binding RNA aptamers with respective K d values of 41.8 and 48.9 nM, which could induce apoptosis of MCF-7 breast cancer cells through the accumulation of reactive oxygen species (ROS) and modulation of intracellular glutathione and caspase-3 activation, suggesting that glutathione-binding RNA aptamers could be developed into effective therapeutic agents for breast cancer treatment. In another study, aptamer AS1411 targeting a Bcl-2 mRNA-binding protein nucleolin was examined for the ability to induce Bcl-2 mRNA instability and cytotoxicity in MCF-7 and MDA-MB-231 breast cancer cells [123]. The results indicated that aptamer AS1411 could efficiently inhibit the growth of MCF-7 and MDA-MB-231 cells (but not that of normal mammary epithelial cells), decrease the half-life of Bcl-2 mRNA in cancer cells, and inhibit nucleolin binding to the AU-rich element 1 in Bcl-2 mRNA, promoting mRNA destabilization. These findings suggest that aptamer AS1411 is a promising molecular decoy which competes with Bcl-2 mRNA for binding to cytoplasmic nucleolin in breast cancer cells, eventually inducing cell death. In another recent study, Varshney et al. [124] selected an RNA aptamer specific for the RID2 domain of human telomerase reverse transcriptase (hTERT), which could tightly and specifically bind to the hTERT peptide and regulate telomerase activity in MCF-7 breast cancer cells, suggesting this RNA aptamer a potential telomerase-blocking agent to be used for breast cancer treatment.
Aptamer-guided drug delivery
Up to date, chemotherapy is still the major treatment approach in cancer. However, since most drugs have low selectivity towards cancer cells and can also kill normal cells, chemotherapy always causes various side effects. Therefore, targeted delivery of chemotherapeutic agents may overcome this limitation and improve their efficacy and specificity. In the past years, a number of targeted strategies have been developed for the treatment of breast cancer. Among them, aptamers can represent ideal targeting ligands with high specificity towards various cancer-related molecules; therefore, aptamer-guided drug delivery systems for breast cancer treatment have attracted considerable attention. Liu et al. [47] developed a new HER2-specific aptamer using SELEX and designed an aptamer-doxorubicin complex (Apt-Dox) by incorporating Dox into the aptamer DNA structure for targeted drug delivery to HER2-positive breast cancer cells. The results demonstrated that the aptamer-guided Dox delivery could elicit selective cytotoxic effects in HER2-positive breast cancer cells, while only weak toxicity was observed for HER2-negative cells. Moreover, Dai et al. [125] have assembled a MUC1-targeting drug delivery system using a MUC1 aptamer and a DNA tetrahedron (Td) carrying Dox within its DNA structure. As a result, the Aptamer-Td complex could selectively bind and deliver Dox to MUC1-positive breast cancer cells, demonstrating significantly higher cytotoxicity towards MUC1-positive MCF-7 breast cancer cells compared to MUC1-negative control cells in vitro (P < 0.01), suggesting a promising drug delivery system for the targeted therapy of MUC1-positive breast cancer. Most recently, Tao et al. [126] had generated novel polydopamine (pD)-modified nanoparticle-aptamer bioconjugates (Apt-pD-DTX/NPs) for in vivo targeted therapy of breast cancer (Fig. 9). Both in vitro cell experiments and in vivo animal studies demonstrated that the Apt-pD-DTX/NPs could enhance targeting efficiency and therapeutic effects, reduce the adverse effects of anti-tumor drugs, and improve survival quality during treatment, indicating the Apt-pD-DTX/NPs a potentially qualified drug delivery system for breast cancer therapy.
Fig. 9.
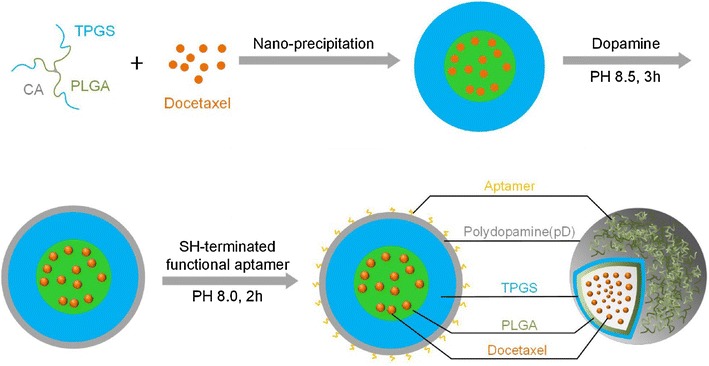
Schematic illustration of the preparation procedure of the targeted Apt-pD-DTX/NPs
(Adapted with permission from Ref. [126]. Copyright © 2016, Ivyspring International Publisher)
Aptamer-nanomaterial conjugates have also been extensively studied in relation to cancer treatment [127]. During the past years, a series of aptamer-nanomaterial conjugates based targeted therapies of breast cancer have also been widely investigated, showing great potential for clinical transformation. Beqa et al. [128] designed a novel hybrid nanomaterial consisting of gold nano popcorn-attached single wall carbon nanotubes for targeted photothermal treatment of breast cancer. The results demonstrated that the targeted SK-BR-3 human breast cancer cells could be killed within 10 min using laser irradiation at 785 nm with 1.5 W/cm2 power. In another study, researchers took advantage of gold nanorods (GNRs), which have strong broad-band longitudinal surface plasmon absorption in the range of 600–1100 nm [129], and synthesized a novel DNA aptamer-tethered GNRs for efficient photothermal therapy of breast cancer [130] (Fig. 10). They used Cell-SELEX to obtain a novel DNA aptamer KW16-13 specific for human breast ductal carcinoma MCF10CA1h cells, which was then conjugated to GNRs and applied for targeted photothermal therapy. The results indicated that the aptamer-GNR conjugates had more than 70-fold specificity to MCF10CA1h tumor cells over normal cells and could be specifically internalized by and cause death in 96% of tumor cells after near-infrared light irradiation. Malik et al. [131] conjugated aptamer AS1411 to gold nanospheres (GNSs) and synthesized AS1411-linked GNSs for targeted therapy of human breast cancer. AS1411-GNSs showed high stability both in solutions and serum, and could be easily internalized by target cells in higher doses and cause increased anti-proliferative/cytotoxic effects compared with pure AS1411 or GNSs modified with control oligonucleotides. Furthermore, the administration of AS1411-GNSs in vivo could dramatically inhibit the growth of xenografted tumors in mice without any toxic side effects, suggesting AS1411-GNSs a promising drug candidate for clinical application in breast cancer treatment.
Fig. 10.
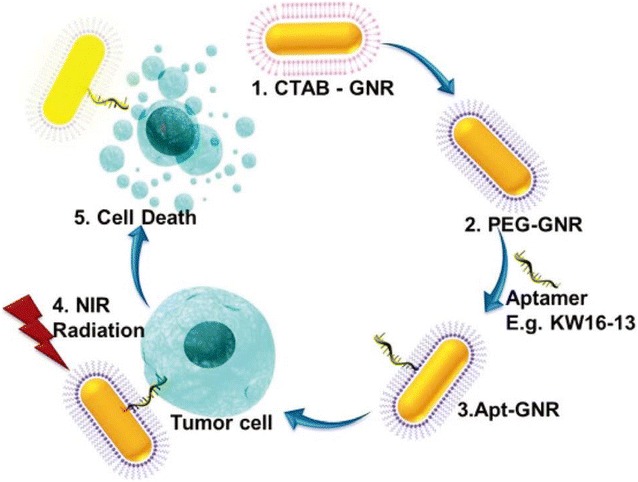
Schematic illustration of the synthesis and use of aptamer functionalized gold nanorods (Apt-GNRs) for targeted photothermal therapy of breast cancer. First, cetyl trimethylammonium bromide-GNRs (CTAB-GNRs) were synthesized via a seed-mediated approach (1). Secondly, the CTAB was replaced by PEG (2) for functionalizing aptamer (3). Then, the Apt-GNRs were incubated with the targeted breast cancer cells followed by photothermal ablation in near infrared range (4), which eventually caused targeted cell death specifically (5)
(Reprinted with permission from Ref. [130]. Copyright © 2015, Royal Society of Chemistry)
Besides chemical drugs and nanomaterials, aptamers can also deliver small interfering (si)RNA for gene therapy. Thiel et al. [132] covalently linked a HER2-specific RNA aptamer to siRNA targeting the anti-apoptotic Bcl-2 gene for use in HER2-positive breast cancer. The aptamer-siRNA conjugates could be selectively taken by HER2-positive breast cancer cells and silence Bcl-2 expression, enhancing the sensitivity of HER2-positive breast cancer cells to chemotherapy. Wang et al. [133] designed an aptamer-siRNA-based system targeting negative apoptosis regulator survivin, which is overexpressed in cancer stem cells of Dox-resistant breast cancer tumors and plays a very important role in drug resistance. They used an RNA aptamer specific for epithelial cell adhesion molecule (EpCAM) and a Dicer substrate survivin siRNA for the treatment of Dox-resistant breast cancer in vivo. The results demonstrated that the aptamer-guided and CSC-targeted in vivo RNAi could deliver a high siRNA dose to breast cancer stem cells in xenograft tumours, which resulted in silencing of survivin and the increase in cancer cell chemo-sensitivity, and eventually suppressed tumour growth, and prolonged the survival of mice bearing xenograft tumours.
Conclusions and future perspectives
In recent years, aptamers have become a focus of research related to diagnostics and treatment of breast cancer because of their advantages over conventional cancer-inhibiting antibodies, such as high target specificity and affinity, good stability, non-immunogenicity, easy modification, and low production cost. In the past decades, a number of aptamers selected against breast cancer biomarkers and/or cells showed great potential for improving diagnostic and treatment rates in breast cancer. In the first part of the review, we discussed and compared various aptamer selection methods, including purified-target based SELEX, Cell-SELEX, in vivo SELEX, and high-efficiency SELEX techniques. Although aptamers evolved via purified target-based SELEX and Cell-SELEX are very promising as detection agents in vitro, they are very easily degraded in physiological environment together with bad efficiency. Therefore, selecting aptamers with good stability and efficiency in physiological environment is very important. Thus, in vivo SELEX was subsequently developed for selection of novel aptamers stable and effective enough in physiological conditions, which can be combined with chemical modification to further enhance the stability and therapeutic efficiency of selected aptamers in vivo. Although the above three SELEX methods can screen aptamers successfully, the reproducibility is not very clear and the selection efficiency is very low. Nowadays, a series of highly efficient SELEX methods have also been developed, but they have the worse stability and reproducibility in comparison with the traditional SELEX methods. Finally, some highly efficient SELEX methods need sophisticated equipment, which is difficult to meet in most laboratories in the world.
In the second part, we specifically focused on aptamer-based detection of breast cancer biomarkers, cells, and tissues, and targeted therapy. Aptamers selected using breast cancer biomarkers and/or cells can be developed into a series of aptasensors for fast, sensitive, and highly specific detection of breast cancer, representing promising candidate agents for early cancer diagnosis. Furthermore, aptamers can be utilized as therapeutics, drug carriers, or recognition ligands for targeted drug delivery in breast cancer treatment. Up to date, various aptamer-based drug delivery systems have been developed; among them, aptamer-guided nanomaterials or nanomaterial-based carriers are the most popular as they can enhance the efficiency of both drug loading and release. In these systems, aptamers always serve as specific recognition ligands, while nanomaterials and nanomaterial-based carriers loaded with chemical drugs act as therapeutic agents. However, only one aptamer-based drug has been approved by the FDA, although there are several aptamer-based drugs being tested in clinic. The main reasons are the instability and nonspecific binding in vivo. Studies demonstrated that aptamers selected in vitro are very prone to nuclease in serum and can be easily degraded, resulting in instability in vivo [116]. Besides, the negative charge of aptamers can make them nonspecifically bind to molecules or ions with positive charge, when it comes to in vivo applications. Moreover, due to the small size and small molecular mass, aptamers can be filtrated rapidly by kidney and nonspecifically accumulated in non-target organs such as liver and spleen, which may lead to adverse effects and reduce therapeutic efficiency. Although the toxicity of aptamers is demonstrated to be low in vitro, their in vivo toxicity are still controversial, which needs more detailed investigation. Besides, the nonspecific binding and accumulation may cause serious immune activation and polyanionic effects. All the above problems and challenges of aptamers eventually decrease the physiological stability and efficiency of aptamers, which dramatically limits their in vivo and clinic applications.
To advance aptamer applications in clinic medicine, the above problems and challenges should be thoroughly solved. On one hand, high efficient SELEX methods with good stability, reproducibility and low cost-effectiveness ratio should be developed for screening aptamers with high specificity, binding affinity, physiologic stability, and low adverse effects in vivo. On the other hand, although various versatile nanomaterials with properties such as monodispersity, good biocompatibility, low toxicity, high entrapment efficiency, and response-mediated release have been synthesized and investigated [134–136], scarcely are they approved for clinic medicine. Therefore, designing and synthesizing new nanomaterials meeting the requirement of clinical utilization should be addressed, either. Finally, the in vivo stability of aptamer-based therapeutics should be increased and immunogenicity monitored should be brought to a minimum. For improving the stability of aptamers in physiological environment, chemical modifications are usually conducted with groups such as fluoro, amino, O-methyl and so on [137–139]. Besides, modified nucleotides have also been applied to aptamer selection, which plays a significant role in stabilizing oligonucleotides against nuclease-mediated degradation in vivo. Another important factor concerned with the stability of aptamers in vivo is renal filtration. The molecular mass of aptamers always ranges from 5 to 15 kDa, which are very prone to renal filtration, despite their good resistance to nuclease-mediated degradation in vivo. However, the incorporation of nanomaterials will dramatically reduce renal filtration rates of aptamers. Therefore, more aptamer-nanomaterial conjugates should be developed to decrease the renal filtration and improve therapeutic efficiency of aptamers. Finally, spiegelmers (l-aptamers), which are the enantiomers of wild-type RNA aptamers (d-aptamers), have also been widely investigated for therapeutic applications due to their excellent resistant to nuclease degradation in vivo [140–142]. Because l-aptamer sequences are intrinsically insensitive to nucleases that can not be degraded, thus possessing better biostability than d-aptamer sequences in vivo. However, to obtain spiegelmers, enantiomeric targets are always needed and only some correlated molecules could be applied, which limits their applications.
Authors’ contributions
ML accomplished most of the work and was a major contributor in writing the manuscript. XY, DY, QL and KD helped to find the references. ZC and TY helped to draw and modify the figures. BL, ZW, SL and YD helped to modify the English grammar. NH provided the idea and gave many suggestions in preparations of the manuscript. All authors read and approved the final manuscript.
Acknowledgements
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Availability of data and materials
Not applicable.
Consent for publication
Not applicable.
Ethics approval and consent to participate
Not applicable.
Funding
This research was financially supported by NSFC (61527806, 61401217, 61471168 and 61571187), Chinese 863 Project (2015AA020502), the National Key Research and Development Program of China (2017YFA0205300), Open Funding of State Key Laboratory of Oral Diseases (SKLOD2017OF04), China Postdoctoral Science Foundation (2016T90403), and the Economical Forest Cultivation and Utilization of 2011 Collaborative Innovation Center in Hunan Province [(2013) 448].
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Abbreviations
- SELEX
Systematic Evolution of Ligands by Exponential enrichment
- ER
estrogen receptor
- PR
progestrone receptor
- HER2
human epidermal growth factor receptor 2
- IHC
immunohistochemistry
- CTCs
circulating tumor cells
- MNPs
magnetic nanoparticles
- HBV
hepatitis B virus
- HBsAg
hepatitis B virus surface antigen
- HBeAg
hepatitis B virus e antigen
- ssDNA
single-stranded DNA
- CE-SELEX
capillary electrophoresis SELEX
- HT-SELEX
high-throughput sequencing-assisted SELEX
- MSD-SELEX
monoclonal surface display SELEX
- Hb
hemoglobin
- AgNCs
silver nanoclusters
- ROS
reactive oxygen species
- hTERT
human telomerase reverse transcriptase
- Apt-Dox
aptamer-doxorubicin complex
- Apt-pD-DTX/NPs
polydopamine (pD)-modified NP-aptamer bioconjugates
- GNRs
gold nanorods
- GNSs
gold nanospheres
- siRNA
small interfering RNA
- EpCAM
epithelial cell adhesion molecule
Contributor Information
Mei Liu, Email: 570613187@163.com.
Xiaocheng Yu, Email: xcyu95@163.com.
Zhu Chen, Email: chukeh@163.com.
Tong Yang, Email: 517782728@qq.com.
Dandan Yang, Email: 739961720@qq.com.
Qianqian Liu, Email: 904621306@qq.com.
Keke Du, Email: 1689661914@qq.com.
Bo Li, Email: 1321630453@qq.com.
Zhifei Wang, Email: zfwang@seu.edu.cn.
Song Li, Email: solisong@163.com.
Yan Deng, Email: hndengyan@126.com.
Nongyue He, Email: nyhe1958@163.com.
References
- 1.DeSantis C, Ma J, Bryan L, Jemal A. Breast cancer statistics, 2013. CA Cancer J Clin. 2014;64:52–62. doi: 10.3322/caac.21203. [DOI] [PubMed] [Google Scholar]
- 2.Liu M, Li Z, Yang J, Jiang Y, Chen Z, Ali Z, He N, Wang Z. Cell-specific biomarkers and targeted biopharmaceuticals for breast cancer treatment. Cell Prolif. 2016;49:409–420. doi: 10.1111/cpr.12266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65:87–108. doi: 10.3322/caac.21262. [DOI] [PubMed] [Google Scholar]
- 4.Azim HA, Ibrahim AS. Breast cancer in Egypt, China and Chinese: statistics and beyond. J Thorac Dis. 2014;6:864–866. doi: 10.3978/j.issn.2072-1439.2014.06.38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.DeSantis CE, Fedewa SA, Goding Sauer A, Kramer JL, Smith RA, Jemal A. Breast cancer statistics, 2015: convergence of incidence rates between black and white women. CA Cancer J Clin. 2016;66:31–42. doi: 10.3322/caac.21320. [DOI] [PubMed] [Google Scholar]
- 6.Lukong KE. Understanding breast cancer-The long and winding road. BBA Clin. 2017;7:64–77. doi: 10.1016/j.bbacli.2017.01.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Maleki S, Dorokhova O, Sunkara J, Schlesinger K, Suhrland M, Oktay MH. Estrogen, progesterone, and HER-2 receptor immunostaining in cytology: the effect of varied fixation on human breast cancer cells. Diagn Cytopathol. 2013;41:864–870. doi: 10.1002/dc.22973. [DOI] [PubMed] [Google Scholar]
- 8.Wu M, Ma J. Association between imaging characteristics and different molecular subtypes of breast cancer. Acad Radiol. 2017;24:426–434. doi: 10.1016/j.acra.2016.11.012. [DOI] [PubMed] [Google Scholar]
- 9.Toss A, Cristofanilli M. Molecular characterization and targeted therapeutic approaches in breast cancer. Breast Cancer Res. 2015;17:60. doi: 10.1186/s13058-015-0560-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Zardavas D, Irrthum A, Swanton C, Piccart M. Clinical management of breast cancer heterogeneity. Nat Rev Clin Oncol. 2015;12:381–394. doi: 10.1038/nrclinonc.2015.73. [DOI] [PubMed] [Google Scholar]
- 11.Lindenberg MA, Miquel-Cases A, Retel VP, Sonke GS, Wesseling J, Stokkel MP, van Harten WH. Imaging performance in guiding response to neoadjuvant therapy according to breast cancer subtypes: a systematic literature review. Crit Rev Oncol Hematol. 2017;112:198–207. doi: 10.1016/j.critrevonc.2017.02.014. [DOI] [PubMed] [Google Scholar]
- 12.Gunn S, Yaziji H, Sims C, Govender S, Moore M, Cotter P, Jones S. A clinically validated DNA microarray for high-resolution HER2 testing defines a new genomic subtype in high-risk breast cancer with equivocal results by IHC and FISH. Cancer Res. 2017;77:P1–P9. doi: 10.1158/1538-7445.SABCS16-P1-09-18. [DOI] [Google Scholar]
- 13.Harbeck N, Gnant M. Breast cancer. Lancet. 2017;389:1134–1150. doi: 10.1016/S0140-6736(16)31891-8. [DOI] [PubMed] [Google Scholar]
- 14.Tuerk C, Gold L. Systematic evolution of ligands by exponential enrichment: rNA ligands to bacteriophage T4 DNA polymerase. Science. 1990;249:505–510. doi: 10.1126/science.2200121. [DOI] [PubMed] [Google Scholar]
- 15.Ellington A, Szostak J. Selection in vitro of single-stranded DNA molecules that fold into specific ligand-binding structures. Nature. 1992;355:850–852. doi: 10.1038/355850a0. [DOI] [PubMed] [Google Scholar]
- 16.Fang X, Tan W. Aptamers generated from cell-SELEX for molecular medicine-A chemical biology approach. Acc Chem Res. 2010;43:48–57. doi: 10.1021/ar900101s. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Xi ZJ, Zheng B, Wang C. Synthesis, surface modification, and biolabeling with aptamer of Fe3O4@SiO2 magnetic nanoparticles. Nanosci Nanotechnol Lett. 2016;8:1061–1066. doi: 10.1166/nnl.2016.2246. [DOI] [Google Scholar]
- 18.Chen ZS, Ali Z, Li S, Liu B, He NY. Aptamers generated from cell-systematic evolution of ligands through exponential enrichment and their applications. J Nanosci Nanotechnol. 2016;16:9346–9358. doi: 10.1166/jnn.2016.12641. [DOI] [Google Scholar]
- 19.Sengupta B, Sinha SS, Garner BL, Arany I, Corley C, Cobb K, Brown E, Ray PC. Influence of aptamer-enclosed silver nanocluster on the prevention of biofilm by bacillus thuringiensis. Nanosci Nanotechnol Lett. 2016;8:1054–1060. doi: 10.1166/nnl.2016.2243. [DOI] [Google Scholar]
- 20.Zhao XQ, Huang YS, Yang CJ, Mao BW. Molecular behavior of the aptamer HJ24 self-assembled on highly oriented pyrolytic graphite (HOPG) Sci China Chem. 2015;58:1858–1865. doi: 10.1007/s11426-015-5465-z. [DOI] [Google Scholar]
- 21.Mayer G. The chemical biology of aptamers. Angew Chem Int Ed. 2009;48:2672–2689. doi: 10.1002/anie.200804643. [DOI] [PubMed] [Google Scholar]
- 22.Xi Z, Huang R, Deng Y, He N. Progress in selection and biomedical applications of aptamers. J Biomed Nanotechnol. 2014;10:3043–3062. doi: 10.1166/jbn.2014.1979. [DOI] [PubMed] [Google Scholar]
- 23.Li X, Zhou B, Zhao ZL, Hu ZX, Zhou SF, Yang N, Huang Y, Zhang ZH, Su J, Lan D, et al. A smart detection system based on specific magnetic and rolling cycle amplification signal-amplified dual-aptamers to accurately monitor minimal residual diseases in patients with T-ALL. J Biomed Nanotechnol. 2016;12:2151–2160. doi: 10.1166/jbn.2016.2302. [DOI] [PubMed] [Google Scholar]
- 24.Mo XD, Wang LX, Zheng Q, Wu GZ. Bi-specific aptamers on nanostructured substrates fail to capture cancer cells. J Nanosci Nanotechnol. 2016;16:8640–8647. doi: 10.1166/jnn.2016.11776. [DOI] [Google Scholar]
- 25.Guo FQ, Hu Y, Yu LY, Deng XY, Meng J, Wang C, Yang XD. Enhancement of thermal damage to adenocarcinoma cells by iron nanoparticles modified with MUC1 aptamer. J Nanosci Nanotechnol. 2016;16:2246–2253. doi: 10.1166/jnn.2016.10941. [DOI] [PubMed] [Google Scholar]
- 26.Wang K, He MQ, Zhai FH, He RH, Yu YL. A novel electrochemical biosensor based on polyadenine modified aptamer for label-free and ultrasensitive detection of human breast cancer cells. Talanta. 2017;166:87–92. doi: 10.1016/j.talanta.2017.01.052. [DOI] [PubMed] [Google Scholar]
- 27.Shahdordizadeh M, Taghdisi SM, Ansari N, Alebooye Langroodi F, Abnous K, Ramezani M. Aptamer based biosensors for detection of Staphylococcus aureus. Sensor Actuat B-Chem. 2017;241:619–635. doi: 10.1016/j.snb.2016.10.088. [DOI] [Google Scholar]
- 28.Bianco M, Sonato A, De Girolamo A, Pascale M, Romanato F, Rinaldi R, Arima V. An aptamer-based SPR-polarization platform for high sensitive OTA detection. Sensor Actuat B Chem. 2017;241:314–320. doi: 10.1016/j.snb.2016.10.056. [DOI] [Google Scholar]
- 29.Chang TJ, Wang LB, Zhao KX, Ge Y, He M, Li G. Duplex identification of Staphylococcus aureus by aptamer and gold nanoparticles. J Nanosci Nanotechnol. 2016;16:5513–5519. doi: 10.1166/jnn.2016.11656. [DOI] [PubMed] [Google Scholar]
- 30.Wang PJ, Wan Y, Ali A, Deng SY, Su Y, Fan CH, Yang SL. Aptamer-wrapped gold nanoparticles for the colorimetric detection of omethoate. Sci China Chem. 2016;59:237–242. doi: 10.1007/s11426-015-5488-5. [DOI] [Google Scholar]
- 31.Zhang Z, Guo C, Zhang S, He L, Wang M, Peng D, Tian J, Fang S. Carbon-based nanocomposites with aptamer-templated silver nanoclusters for the highly sensitive and selective detection of platelet-derived growth factor. Biosens Bioelectron. 2017;89:735–742. doi: 10.1016/j.bios.2016.11.019. [DOI] [PubMed] [Google Scholar]
- 32.Kang YR, Byun JS, Kim TJ, Park HG, Park JC, Barange N, Nam KH, Kim YD. Monitoring of the binding between egfr protein and egfr aptamer using in situ total internal reflection ellipsometry. J Nanosci Nanotechnol. 2016;16:6445–6449. doi: 10.1166/jnn.2016.12123. [DOI] [PubMed] [Google Scholar]
- 33.Lu D, He L, Zhang G, Lv A, Wang R, Zhang X, Tan W. Aptamer-assembled nanomaterials for fluorescent sensing and imaging. Nanophotonics. 2017;6:109–121. [Google Scholar]
- 34.Zhu G, Zhang S, Song E, Zheng J, Hu R, Fang X, Tan W. Building fluorescent DNA nanodevices on target living cell surfaces. Angew Chem Int Ed. 2013;52:5490–5496. doi: 10.1002/anie.201301439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Yan LA, Shi H, He DG, He XX, Wang KM, Xu FZ. A pH-responsive activatable aptamer probe for targeted cancer imaging based on i-motif-driven conformation alteration. Sci China Chem. 2016;59:802–808. doi: 10.1007/s11426-016-5575-2. [DOI] [Google Scholar]
- 36.Zhao Z, Xu L, Shi X, Tan W, Fang X, Shangguan D. Recognition of subtype non-small cell lung cancer by DNA aptamers selected from living cells. Analyst. 2009;134:1808–1814. doi: 10.1039/b904476k. [DOI] [PubMed] [Google Scholar]
- 37.Zhu G, Zheng J, Song E, Donovan M, Zhang K, Liu C, Tan W. Self-assembled, aptamer-tethered DNA nanotrains for targeted transport of molecular drugs in cancer theranostics. P Natl Acad Sci USA. 2013;110:7998–8003. doi: 10.1073/pnas.1220817110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Liu LS, Lu XL, Zhao YX. Aptamer-based strategies for cancer diagnosis and therapy. J Nanosci Nanotechnol. 2016;16:6611–6621. doi: 10.1166/jnn.2016.11366. [DOI] [Google Scholar]
- 39.Shandordizadeh M, Yazdian-Robati R, Ramezani M, Abnous K, Taghdisi S. Aptamer application in targeted delivery systems for diagnosis and treatment of breast cancer. J Mater Chem B. 2016;4:7766–7778. doi: 10.1039/C6TB02564A. [DOI] [PubMed] [Google Scholar]
- 40.Niazi J, Verma S, Niazi S, Qureshi A. In vitro HER2 protein-induced affinity dissociation of carbon nanotube-wrapped anti-HER2 aptamers for HER2 protein detection. Analyst. 2013;140:243–249. doi: 10.1039/C4AN01665C. [DOI] [PubMed] [Google Scholar]
- 41.Li J, Zhong X, Cheng F, Zhang JR, Jiang LP, Zhu JJ. One-pot synthesis of aptamer-functionalized silver nanoclusters for cell-type-specific imaging. Anal Chem. 2012;84:4140–4146. doi: 10.1021/ac3003402. [DOI] [PubMed] [Google Scholar]
- 42.Hua X, Zhou Z, Yuan L, Liu S. Selective collection and detection of MCF-7 breast cancer cells using aptamer-functionalized magnetic beads and quantum dots based nano-bio-probes. Anal Chim Acta. 2013;788:135–140. doi: 10.1016/j.aca.2013.06.001. [DOI] [PubMed] [Google Scholar]
- 43.Ahirwar R, Vellarikkal SK, Sett A, Sivasubbu S, Scaria V, Bora U, Borthakur BB, Kataki AC, Sharma JD, Nahar P. Aptamer-assisted detection of the altered expression of estrogen receptor alpha in human breast cancer. PLoS ONE. 2016;11:e0153001. doi: 10.1371/journal.pone.0153001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Kang H, Huh Y, Kim S, Lee D. Isolation of rna aptamers targeting HER-2-overexpressing breast cancer cells using cell-SELEX. Bull Korean Chem Soc. 2009;30:1827–1831. doi: 10.5012/bkcs.2009.30.8.1827. [DOI] [Google Scholar]
- 45.Li X, Zhang W, Liu L, Zhu Z, Ouyang G, An Y, Zhao C, Yang CJ. In vitro selection of DNA aptamers for metastatic breast cancer cell recognition and tissue imaging. Anal Chem. 2014;86:6596–6603. doi: 10.1021/ac501205q. [DOI] [PubMed] [Google Scholar]
- 46.Wu P, Gao Y, Zhang H, Cai C. Aptamer-guided silver-gold bimetallic nanostructures with highly active surface-enhanced raman scattering for specific detection and near-infrared photothermal therapy of human breast cancer cells. Anal Chem. 2012;84:7692–7699. doi: 10.1021/ac3015164. [DOI] [PubMed] [Google Scholar]
- 47.Liu Z, Duan J, Song Y, Ma J, Wang F, Lu X, Yang X. Novel HER2 aptamer selectively delivers cytotoxic drug to HER2-positive breast cancer cells in vitro. J Transl Med. 2012;10:148. doi: 10.1186/1479-5876-10-148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Zhang K, Sefah K, Tang L, Zhao Z, Zhu G, Ye M, Sun W, Goodison S, Tan W. A novel aptamer developed for breast cancer cell internalization. Chem Med Chem. 2012;7:79–84. doi: 10.1002/cmdc.201100457. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Song Y, Zhu Z, An Y, Zhang W, Zhang H, Liu D, Yu C, Duan W, Yang CJ. Selection of DNA aptamers against epithelial cell adhesion molecule for cancer cell imaging and circulating tumor cell capture. Anal Chem. 2013;85:4141–4149. doi: 10.1021/ac400366b. [DOI] [PubMed] [Google Scholar]
- 50.Ellington AD, Szostak JW. In vitro selection of RNA molecules that bind specific ligands. Nature. 1990;346:818–822. doi: 10.1038/346818a0. [DOI] [PubMed] [Google Scholar]
- 51.Darmostuk M, Rimpelova S, Gbelcova H, Ruml T. Current approaches in SELEX: an update to aptamer selection technology. Biotechnol Adv. 2015;33:1141–1161. doi: 10.1016/j.biotechadv.2015.02.008. [DOI] [PubMed] [Google Scholar]
- 52.Hamula CLA, Guthrie JW, Zhang HQ, Li XF, Le XC. Selection and analytical applications of aptamers. TracTrend Anal Chem. 2006;25:681–691. doi: 10.1016/j.trac.2006.05.007. [DOI] [Google Scholar]
- 53.Hamula CLA, Zhang HQ, Li F, Wang ZX, Le XC, Li XF. Selection and analytical applications of aptamers binding microbial pathogens. TracTrend Anal Chem. 2011;30:1587–1597. doi: 10.1016/j.trac.2011.08.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Yang KA, Pei RJ, Stojanovic MN. In vitro selection and amplification protocols for isolation of aptameric sensors for small molecules. Methods. 2016;106:58–65. doi: 10.1016/j.ymeth.2016.04.032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Wang HY, Cheng H, Wang J, Xu LJ, Chen HX, Pei RJ. Selection and characterization of DNA aptamers for the development of light-up biosensor to detect Cd(II) Talanta. 2016;154:498–503. doi: 10.1016/j.talanta.2016.04.005. [DOI] [PubMed] [Google Scholar]
- 56.Zhou JH, Rossi JJ. Cell-type-specific, aptamer-functionalized agents for targeted disease therapy. Mol TherNucl Acids. 2014;3:e169. doi: 10.1038/mtna.2014.21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Li S, Xu H, Ding H, Huang Y, Cao X, Yang G, Li J, Xie Z, Meng Y, Li X, et al. Identification of an aptamer targeting hnRNP A1 by tissue slide-based SELEX. J Pathol. 2009;218:327–336. doi: 10.1002/path.2543. [DOI] [PubMed] [Google Scholar]
- 58.Wang R, Zhao J, Jiang T, Kwon YM, Lu H, Jiao P, Liao M, Li Y. Selection and characterization of DNA aptamers for use in detection of avian influenza virus H5N1. J Virol Methods. 2013;189:362–369. doi: 10.1016/j.jviromet.2013.03.006. [DOI] [PubMed] [Google Scholar]
- 59.Heiat M, Ranjbar R, Latifi AM, Rasaee MJ. Selection of a high-affinity and in vivo bioactive ssDNA aptamer against angiotensin II peptide. Peptides. 2016;82:101–108. doi: 10.1016/j.peptides.2016.06.004. [DOI] [PubMed] [Google Scholar]
- 60.Mi J, Ray P, Liu J, Kuan CT, Xu J, Hsu D, Sullenger BA, White RR, Clary BM. In vivo selection against human colorectal cancer xenografts identifies an aptamer that targets RNA helicase protein DHX9. Mol Therucl Acids. 2016;5:e315. doi: 10.1038/mtna.2016.27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Zhou J, Rossi J. Aptamers as targeted therapeutics: current potential and challenges. Nat Rev Drug Discov. 2017;16:181–202. doi: 10.1038/nrd.2016.199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Zhang H, Zhou L, Zhu Z, Yang C. Recent progress in aptamer-based functional probes for bioanalysis and biomedicine. Chem Eur J. 2016;22:9886–9900. doi: 10.1002/chem.201503543. [DOI] [PubMed] [Google Scholar]
- 63.Paniel N, Istamboulie G, Triki A, Lozano C, Barthelmebs L, Noguer T. Selection of DNA aptamers against penicillin G using Capture-SELEX for the development of an impedimetric sensor. Talanta. 2017;162:232–240. doi: 10.1016/j.talanta.2016.09.058. [DOI] [PubMed] [Google Scholar]
- 64.Spiga FM, Maietta P, Guiducci C. More DNA-aptamers for small drugs: a capture-SELEX coupled with surface plasmon resonance and high-throughput sequencing. ACS Comb Sci. 2015;17:326–333. doi: 10.1021/acscombsci.5b00023. [DOI] [PubMed] [Google Scholar]
- 65.Xi Z, Huang R, Li Z, He N, Wang T, Su E, Deng Y. Selection of HBsAg-specific DNA aptamers based on carboxylated magnetic nanoparticles and their application in the rapid and simple detection of hepatitis B virus infection. ACS Appl Mater Interfaces. 2015;7:11215–11223. doi: 10.1021/acsami.5b01180. [DOI] [PubMed] [Google Scholar]
- 66.Bock L, Griffin L, Latham J, Vermaas E, Toole J. Selection of single-stranded DNA molecules that bind and inhibit human thrombin. Nature. 1992;355:564–566. doi: 10.1038/355564a0. [DOI] [PubMed] [Google Scholar]
- 67.Huang R, Xi Z, Deng Y, He N. Fluorescence based aptasensors for the determination of hepatitis B virus e antigen. Sci Rep. 2016;6:31103. doi: 10.1038/srep31103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Shangguan D, Li Y, Tang Z, Cao ZC, Chen HW, Mallikaratchy P, Sefah K, Yang CJ, Tan W. Aptamers evolved from live cells as effective molecular probes for cancer study. Proc Natl Acad Sci USA. 2006;103:11838–11843. doi: 10.1073/pnas.0602615103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Jin C, Qiu L, Li J, Fu T, Zhang X, Tan W. Cancer biomarker discovery using DNA aptamers. Analyst. 2016;141:461–466. doi: 10.1039/C5AN01918D. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Lu M, Zhou L, Zheng X, Quan Y, Wang X, Zhou X, Ren J. A novel molecular marker of breast cancer stem cells identified by cell-SELEX method. Cancer Biomark. 2015;15:163–170. doi: 10.3233/CBM-140450. [DOI] [PubMed] [Google Scholar]
- 71.Bing T, Shangguan D, Wang Y. Facile discovery of cell-surface protein targets of cancer cell aptamers. Mol Cell Proteomics. 2015;14:2692–2700. doi: 10.1074/mcp.M115.051243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Bayrac AT, Sefah K, Parekh P, Bayrac C, Gulbakan B, Oktem HA, Tan W. In vitro selection of DNA aptamers to glioblastoma multiforme. ACS Chem Neurosci. 2011;2:175–181. doi: 10.1021/cn100114k. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Li X, An Y, Jin J, Zhu Z, Hao L, Liu L, Shi Y, Fan D, Ji T, Yang CJ. Evolution of DNA aptamers through in vitro metastatic-cell-based systematic evolution of ligands by exponential enrichment for metastatic cancer recognition and imaging. Anal Chem. 2015;87:4941–4948. doi: 10.1021/acs.analchem.5b00637. [DOI] [PubMed] [Google Scholar]
- 74.Huang R, Chen Z, Liu M, Deng Y, Li S, He N. The aptamers generated from HepG2 cells. Sci China Chem. 2017;60:786–792. doi: 10.1007/s11426-016-0491-7. [DOI] [Google Scholar]
- 75.Keefe AD, Pai S, Ellington A. Aptamers as therapeutics. Nat Rev Drug Discov. 2010;9:537–550. doi: 10.1038/nrd3141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Mi J, Liu Y, Rabbani Z, Yang Z, Urban J, Sullenger B, Clary B. In vivo selection of tumor-targeting RNA motifs. Nat Chem Biol. 2010;6:22–24. doi: 10.1038/nchembio.277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Wang HL, Zhang YB, Chu C, Dai W, Liu RH, Jiang YP. Identification of a specific RNA aptamer targeting non-small cell lung cancer by in vivo SELEX. Cancer Res. 2016;76:2058. doi: 10.1158/1538-7445.AM2016-2058. [DOI] [Google Scholar]
- 78.Li Q, Zhao X, Liu H, Qu F. Low pH capillary electrophoresis application to improve capillary electrophoresis-systematic evolution of ligands by exponential enrichment. J Chromatogr A. 2014;1364:289–294. doi: 10.1016/j.chroma.2014.08.073. [DOI] [PubMed] [Google Scholar]
- 79.Birch CM, Hou HW, Han J, Niles JC. Identification of malaria parasite-infected red blood cell surface aptamers by inertial microfluidic SELEX (I-SELEX) Sci Rep. 2015;5:11347. doi: 10.1038/srep11347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Thiel WH, Giangrande PH. Analyzing HT-SELEX data with the Galaxy Project tools—a web based bioinformatics platform for biomedical research. Methods. 2016;97:3–10. doi: 10.1016/j.ymeth.2015.10.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Gotrik MR, Feagin TA, Csordas AT, Nakamoto MA, Soh HT. Advancements in aptamer discovery technologies. Acc Chem Res. 2016;49:1903–1910. doi: 10.1021/acs.accounts.6b00283. [DOI] [PubMed] [Google Scholar]
- 82.Zhu Z, Song Y, Li C, Zou Y, Zhu L, An Y, Yang CJ. Monoclonal surface display SELEX for simple, rapid, efficient, and cost-effective aptamer enrichment and identification. Anal Chem. 2014;86:5881–5888. doi: 10.1021/ac501423g. [DOI] [PubMed] [Google Scholar]
- 83.Hunniger T, Wessels H, Fischer C, Paschke-Kratzin A, Fischer M. Just in time-selection: a rapid semiautomated SELEX of DNA aptamers using magnetic separation and BEAMing. Anal Chem. 2014;86:10940–10947. doi: 10.1021/ac503261b. [DOI] [PubMed] [Google Scholar]
- 84.Dong L, Tan Q, Ye W, Liu D, Chen H, Hu H, Wen D, Liu Y, Cao Y, Kang J, et al. Screening and identifying a novel ssDNA aptamer against alpha-fetoprotein using CE-SELEX. Sci Rep. 2015;5:15552. doi: 10.1038/srep15552. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Lin HI, Wu CC, Yang CH, Chang KW, Lee GB, Shiesh SC. Selection of aptamers specific for glycated hemoglobin and total hemoglobin using on-chip SELEX. Lab Chip. 2015;15:486–494. doi: 10.1039/C4LC01124D. [DOI] [PubMed] [Google Scholar]
- 86.Ulrich H, Kim EY, Kim JW, Kim WK, Han BS, Park SG, Chung BH, Lee SC, Bae K-H. Selection of aptamers for mature white adipocytes by cell SELEX using flow cytometry. PLoS ONE. 2014;9:e97747. doi: 10.1371/journal.pone.0097747. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Moon J, Kim G, Lee S, Park S. Identification of Salmonella Typhimurium-specific DNA aptamers developed using whole-cell SELEX and FACS analysis. J Microbiol Methods. 2013;95:162–166. doi: 10.1016/j.mimet.2013.08.005. [DOI] [PubMed] [Google Scholar]
- 88.Gonzalez-Fernandez E, de-los-Santos-Alvarez N, Miranda-Ordieres AJ, Lobo-Castanon MJ. SPR evaluation of binding kinetics and affinity study of modified RNA aptamers towards small molecules. Talanta. 2012;99:767–773. doi: 10.1016/j.talanta.2012.07.019. [DOI] [PubMed] [Google Scholar]
- 89.Szeto K, Craighead HG. Devices and approaches for generating specific high-affinity nucleic acid aptamers. Appl Phys Rev. 2014;1:031103. doi: 10.1063/1.4894851. [DOI] [Google Scholar]
- 90.Jiao F, Fan H, Yang G, Zhang F, He P. Directly investigating the interaction between aptamers and thrombin by atomic force microscopy. J Mol Recognit. 2013;26:672–678. doi: 10.1002/jmr.2312. [DOI] [PubMed] [Google Scholar]
- 91.Ge L, Jin G, Fang X. Investigation of the interaction between a bivalent aptamer and thrombin by AFM. Langmuir. 2012;28:707–713. doi: 10.1021/la203954x. [DOI] [PubMed] [Google Scholar]
- 92.Ouellet E, Foley JH, Conway EM, Haynes C. Hi-Fi SELEX: a high-fidelity digital-PCR based therapeutic aptamer discovery platform. Biotechnol Bioeng. 2015;112:1506–1522. doi: 10.1002/bit.25581. [DOI] [PubMed] [Google Scholar]
- 93.Qureshi A, Gurbuz Y, Niazi JH. Label-free capacitance based aptasensor platform for the detection of HER2/ErbB2 cancer biomarker in serum. Sensor Actuat B Chem. 2015;220:1145–1151. doi: 10.1016/j.snb.2015.06.094. [DOI] [Google Scholar]
- 94.Sett A, Borthakur BB, Sharma JD, Kataki AC, Bora U. DNA aptamer probes for detection of estrogen receptor alpha positive carcinomas. Transl Res. 2017;183:104–120. doi: 10.1016/j.trsl.2016.12.008. [DOI] [PubMed] [Google Scholar]
- 95.Lan J, Li L, Liu Y, Yan L, Li C, Chen J, Chen X. Upconversion luminescence assay for the detection of the vascular endothelial growth factor, a biomarker for breast cancer. Microchim Acta. 2016;183:3201–3208. doi: 10.1007/s00604-016-1965-6. [DOI] [Google Scholar]
- 96.Guo Q, Li X, Shen C, Zhang S, Qi H, Li T, Yang M. Electrochemical immunoassay for the protein biomarker mucin 1 and for MCF-7 cancer cells based on signal enhancement by silver nanoclusters. Microchim Acta. 2015;182:1483–1489. doi: 10.1007/s00604-015-1471-2. [DOI] [Google Scholar]
- 97.Meirinho SG, Dias LG, Peres AM, Rodrigues LR. Development of an electrochemical aptasensor for the detection of human osteopontin. Procedia Engineer. 2014;87:316–319. doi: 10.1016/j.proeng.2014.11.671. [DOI] [Google Scholar]
- 98.Wu J, Wang C, Li X, Song Y, Wang W, Li C, Hu J, Zhu Z, Li J, Zhang W, et al. Identification, characterization and application of a G-quadruplex structured DNA aptamer against cancer biomarker protein anterior gradient homolog 2. PLoS ONE. 2012;7:e46393. doi: 10.1371/journal.pone.0046393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Li H, Bai X, Wang N, Chen X, Li J, Zhang Z, Tang J. Aptamer-based microcantilever biosensor for ultrasensitive detection of tumor marker nucleolin. Talanta. 2016;146:727–731. doi: 10.1016/j.talanta.2015.06.034. [DOI] [PubMed] [Google Scholar]
- 100.Jo H, Ban C. Aptamer-nanoparticle complexes as powerful diagnostic and therapeutic tools. Exp Mol Med. 2016;48:e230. doi: 10.1038/emm.2016.44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Liu Q, Jin C, Wang Y, Fang X, Zhang X, Chen Z, Tan W. Aptamer-conjugated nanomaterials for specific cancer cell recognition and targeted cancer therapy. NPG Asia Mater. 2014;6:e95. doi: 10.1038/am.2014.12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Meng H-M, Fu T, Zhang X-B, Tan W. Cell-SELEX-based aptamer-conjugated nanomaterials for cancer diagnosis and therapy. Natl Sci Rev. 2015;2:71–84. doi: 10.1093/nsr/nwv001. [DOI] [Google Scholar]
- 103.Zhu Y, Chandra P, Shim YB. Ultrasensitive and selective electrochemical diagnosis of breast cancer based on a hydrazine-Au nanoparticle-aptamer bioconjugate. Anal Chem. 2013;85:1058–1064. doi: 10.1021/ac302923k. [DOI] [PubMed] [Google Scholar]
- 104.Liu F, Zhang Y, Yu J, Wang S, Ge S, Song X. Application of ZnO/graphene and S6 aptamers for sensitive photoelectrochemical detection of SK-BR-3 breast cancer cells based on a disposable indium tin oxide device. Biosens Bioelectron. 2014;51:413–420. doi: 10.1016/j.bios.2013.07.066. [DOI] [PubMed] [Google Scholar]
- 105.Cai S, Li G, Zhang X, Xia Y, Chen M, Wu D, Chen Q, Zhang J, Chen J. A signal-on fluorescent aptasensor based on single-stranded DNA-sensitized luminescence of terbium (III) for label-free detection of breast cancer cells. Talanta. 2015;138:225–230. doi: 10.1016/j.talanta.2015.02.056. [DOI] [PubMed] [Google Scholar]
- 106.Jo H, Her J, Ban C. Dual aptamer-functionalized silica nanoparticles for the highly sensitive detection of breast cancer. Biosens Bioelectron. 2015;71:129–136. doi: 10.1016/j.bios.2015.04.030. [DOI] [PubMed] [Google Scholar]
- 107.Li T, Yang J, Ali Z, Wang Z, Mou X, He N, Wang Z. Synthesis of aptamer-functionalized Ag nanoclusters for MCF-7 breast cancer cells imaging. Sci China Chem. 2017;60:370–376. doi: 10.1007/s11426-016-0159-2. [DOI] [Google Scholar]
- 108.Wu X, Zhao Z, Bai H, Fu T, Yang C, Hu X, Liu Q, Champanhac C, Teng IT, Ye M, et al. DNA aptamer selected against pancreatic ductal adenocarcinoma for in vivo imaging and clinical tissue recognition. Theranostics. 2015;5:985–994. doi: 10.7150/thno.11938. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Pu Y, Liu Z, Lu Y, Yuan P, Liu J, Yu B, Wang G, Yang CJ, Liu H, Tan W. Using DNA aptamer probe for immunostaining of cancer frozen tissues. Anal Chem. 2015;87:1919–1924. doi: 10.1021/ac504175h. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Bauer M, Macdonald J, Henri J, Duan W, Shigdar S. The application of aptamers for immunohistochemistry. Nucleic Acid Ther. 2016;26:120–126. doi: 10.1089/nat.2015.0569. [DOI] [PubMed] [Google Scholar]
- 111.Bukari BA, Citartan M, Ch’ng ES, Bilibana MP, Rozhdestvensky T, Tang TH. Aptahistochemistry in diagnostic pathology: technical scrutiny and feasibility. Histochem Cell Biol. 2017;147:545–553. doi: 10.1007/s00418-017-1561-9. [DOI] [PubMed] [Google Scholar]
- 112.Gupta S, Thirstrup D, Jarvis TC, Schneider DJ, Wilcox SK, Carter J, Zhang C, Gelinas A, Weiss A, Janjic N, et al. Rapid histochemistry using slow off-rate modified aptamers with anionic competition. Appl Immunohisto M M. 2011;19:273–278. doi: 10.1097/PAI.0b013e3182008c29. [DOI] [PubMed] [Google Scholar]
- 113.Sun H, Zhu X, Lu PY, Rosato RR, Tan W, Zu Y. Oligonucleotide aptamers: new tools for targeted cancer therapy. Mol TherNucl Acids. 2014;3:e182. doi: 10.1038/mtna.2014.32. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Xiang D, Shigdar S, Qiao G, Wang T, Kouzani AZ, Zhou SF, Kong L, Li Y, Pu C, Duan W. Nucleic acid aptamer-guided cancer therapeutics and diagnostics: the next generation of cancer medicine. Theranostics. 2015;5:23–42. doi: 10.7150/thno.10202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Ng EW, Shima DT, Calias P, Cunningham ET, Jr, Guyer DR, Adamis AP. Pegaptanib, a targeted anti-VEGF aptamer for ocular vascular disease. Nat Rev Drug Discov. 2006;5:123–132. doi: 10.1038/nrd1955. [DOI] [PubMed] [Google Scholar]
- 116.Healy J, Lewis S, Kurz M, Boomer R, Thompson K, Wilson C, McCauley T. Pharmacokinetics and biodistribution of novel aptamer compositions. Pharm Res. 2004;21:2234–2246. doi: 10.1007/s11095-004-7676-4. [DOI] [PubMed] [Google Scholar]
- 117.Chen Y, Lin JS. The application of aptamer in apoptosis. Biochimie. 2017;132:1–8. doi: 10.1016/j.biochi.2016.10.008. [DOI] [PubMed] [Google Scholar]
- 118.Jian Y, Gao Z, Sun J, Shen Q, Feng F, Jing Y, Yang C. RNA aptamers interfering with nucleophosmin oligomerization induce apoptosis of cancer cells. Oncogene. 2009;28:4201–4211. doi: 10.1038/onc.2009.275. [DOI] [PubMed] [Google Scholar]
- 119.Kunz C, Borghouts C, Buerger C, Groner B. Peptide aptamers with binding specificity for the intracellular domain of the ErbB2 receptor interfere with AKT signaling and sensitize breast cancer cells to Taxol. Mol Cancer Res. 2006;4:983–998. doi: 10.1158/1541-7786.MCR-06-0046. [DOI] [PubMed] [Google Scholar]
- 120.Camorani S, Crescenzi E, Gramanzini M, Fedele M, Zannetti A, Cerchia L. Aptamer-mediated impairment of EGFR-integrin alphavbeta3 complex inhibits vasculogenic mimicry and growth of triple-negative breast cancers. Sci Rep. 2017;7:46659. doi: 10.1038/srep46659. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Chen K, Liu J, Tong GX, Liu B, Wang GD, Liu HX. Adipo8, a high-affinity DNA aptamer, can differentiate among adipocytes and inhibit intracellular lipid accumulation in vitro. Sci China Chem. 2015;58:1612–1620. doi: 10.1007/s11426-015-5367-0. [DOI] [Google Scholar]
- 122.Bala J, Bhaskar A, Varshney A, Singh AK, Dey S, Yadava P. In vitro selected RNA aptamer recognizing glutathione induces ROS mediated apoptosis in the human breast cancer cell line MCF 7. RNA Biol. 2011;8:101–111. doi: 10.4161/rna.8.1.14116. [DOI] [PubMed] [Google Scholar]
- 123.Soundararajan S, Chen W, Spicer EK, Courtenay-Luck N, Fernandes DJ. The nucleolin targeting aptamer AS1411 destabilizes Bcl-2 messenger RNA in human breast cancer cells. Cancer Res. 2008;68:2358–2365. doi: 10.1158/0008-5472.CAN-07-5723. [DOI] [PubMed] [Google Scholar]
- 124.Varshney A, Bala J, Santosh B, Bhaskar A, Kumar S, Yadava PK. Identification of an RNA aptamer binding hTERT-derived peptide and inhibiting telomerase activity in MCF7 cells. Mol Cell Biochem. 2017;427:157–167. doi: 10.1007/s11010-016-2907-7. [DOI] [PubMed] [Google Scholar]
- 125.Dai B, Hu Y, Duan J, Yang X. Aptamer-guided DNA tetrahedron as a novel targeted drug delivery system for MUC1-expressing breast cancer cells in vitro. Oncotarget. 2016;7:38257–38269. doi: 10.18632/oncotarget.9431. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Tao W, Zeng X, Wu J, Zhu X, Yu X, Zhang X, Zhang J, Liu G, Mei L. Polydopamine-based surface modification of novel nanoparticle-aptamer bioconjugates for in vivo breast cancer targeting and enhanced therapeutic effects. Theranostics. 2016;6:470–484. doi: 10.7150/thno.14184. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Reinemann C, Strehlitz B. Aptamer-modified nanoparticles and their use in cancer diagnostics and treatment. Swiss Med Wkly. 2014;144:w13908. doi: 10.4414/smw.2014.13908. [DOI] [PubMed] [Google Scholar]
- 128.Beqa L, Fan Z, Singh AK, Senapati D, Ray PC. Gold nano-popcorn attached SWCNT hybrid nanomaterial for targeted diagnosis and photothermal therapy of human breast cancer cells. ACS Appl Mater Interfaces. 2011;3:3316–3324. doi: 10.1021/am2004366. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Alkilany AM, Thompson LB, Boulos SP, Sisco PN, Murphy CJ. Gold nanorods: their potential for photothermal therapeutics and drug delivery, tempered by the complexity of their biological interactions. Adv Drug Deliv Rev. 2012;64:190–199. doi: 10.1016/j.addr.2011.03.005. [DOI] [PubMed] [Google Scholar]
- 130.Chandrasekaran R, Lee ASW, Yap LW, Jans DA, Wagstaff KM, Cheng W. Tumor cell-specific photothermal killing by SELEX-derived DNA aptamer-targeted gold nanorods. Nanoscale. 2016;8:187–196. doi: 10.1039/C5NR07831H. [DOI] [PubMed] [Google Scholar]
- 131.Malik M, O’Toole M, Casson L, Thomas S, Bardi G, Reyes-Reyes E, Ng C, Kang K, Bates P. AS1411-conjugated gold nanospheres and their potential for breast cancer therapy. Oncotarget. 2015;6:22270–22281. doi: 10.18632/oncotarget.4207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Thiel KW, Hernandez LI, Dassie JP, Thiel WH, Liu X, Stockdale KR, Rothman AM, Hernandez FJ, McNamara JO, Giangrande PH. Delivery of chemo-sensitizing siRNAs to HER2+-breast cancer cells using RNA aptamers. Nucleic Acids Res. 2012;40:6319–6337. doi: 10.1093/nar/gks294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133.Wang T, Gantier MP, Xiang D, Bean AG, Bruce M, Zhou SF, Khasraw M, Ward A, Wang L, Wei MQ, et al. EpCAM aptamer-mediated survivin silencing sensitized cancer stem cells to doxorubicin in a breast cancer model. Theranostics. 2015;5:1456–1472. doi: 10.7150/thno.11692. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Jiang F, Liu BA, Lu J, Li FF, Li DF, Liang C, Dang L, Liu J, He B, Badshah SA, et al. Progress and challenges in developing aptamer-functionalized targeted drug delivery systems. Int J Mol Sci. 2015;16:23784–23822. doi: 10.3390/ijms161023784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Liao J, Liu B, Liu J, Zhang JN, Chen K, Liu HX. Cell-specific aptamers and their conjugation with nanomaterials for targeted drug delivery. Expert Opin Drug Del. 2015;12:493–506. doi: 10.1517/17425247.2015.966681. [DOI] [PubMed] [Google Scholar]
- 136.Zhu HJ, Li J, Zhang XB, Ye M, Tan WH. Nucleic acid aptamer-mediated drug delivery for targeted cancer therapy. Chem Med Chem. 2015;10:39–45. doi: 10.1002/cmdc.201402312. [DOI] [PubMed] [Google Scholar]
- 137.Biesecker G, Dihel L, Enney K, Bendele RA. Derivation of RNA aptamer inhibitors of human complement C5. Immunopharmacology. 1999;42:219–230. doi: 10.1016/S0162-3109(99)00020-X. [DOI] [PubMed] [Google Scholar]
- 138.Siddiqui MAA, Keating GM. Pegaptanib in exudative age-related macular degeneration. Drugs. 2005;65:1571–1577. doi: 10.2165/00003495-200565110-00010. [DOI] [PubMed] [Google Scholar]
- 139.Burmeister PE, Lewis SD, Silva RF, Preiss JR, Horwitz LR, Pendergrast PS, McCauley TG, Kurz JC, Epstein DM, Wilson C, et al. Direct in vitro selection of a 2’-O-methyl aptamer to VEGF. Chem Biol. 2005;12:25–33. doi: 10.1016/j.chembiol.2004.10.017. [DOI] [PubMed] [Google Scholar]
- 140.Vater A, Klussmann S. Toward third-generation aptamers: spiegelmers and their therapeutic prospects. Curr Opin Drug Discov Dev. 2003;6:253–261. [PubMed] [Google Scholar]
- 141.Sczepanski JT, Joyce GF. Binding of a structured D-RNA molecule by an L-RNA aptamer. J Am Chem Soc. 2013;135:13290–13293. doi: 10.1021/ja406634g. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142.Sczepanski JT, Joyce GF. Specific inhibition of microRNA processing using L-RNA aptamers. J Am Chem Soc. 2015;137:16032–16037. doi: 10.1021/jacs.5b06696. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Not applicable.


